Abstract
Mood disorders, including depression and anxiety, affect almost one-fifth of the world’s adult population and are becoming increasingly prevalent. Mutations in circadian clock genes have previously been associated with mood disorders both directly and indirectly through alterations in circadian phase, suggesting that the circadian clock influences multiple molecular pathways involved in mood. By targeting previously identified single nucleotide polymorphisms (SNPs) that have been implicated in anxiety and depressive disorders, we use a combination of statistical and machine learning techniques to investigate associations with the generalized anxiety disorder assessment (GAD-7) scores in a UK Biobank sample of 90,882 individuals. As in previous studies, we observed that females exhibited higher GAD-7 scores than males regardless of genotype. Interestingly, we found no significant effects on anxiety from individual circadian gene variants; only circadian genotypes with multiple SNP variants showed significant associations with anxiety. For both sexes, severe anxiety is associated with a 120-fold increase in odds for individuals with CRY2_AG(rs1083852)/ZBTB20_TT(rs1394593) genotypes and is associated with a near 40-fold reduction in odds for individuals with PER3-A_CG(rs228697)/ZBTB20_TT(rs1394593) genotypes. We also report several sex-specific associations with anxiety. In females, the CRY2/ZBTB20 genotype combination showed a > 200-fold increase in odds of anxiety and PER3/ZBTB20 and CRY1 /PER3-A genotype combinations also appeared as female risk factors. In males, CRY1/PER3-A and PER3-B/ZBTB20 genotype combinations were associated with anxiety risk. Mediation analysis revealed direct associations of CRY2/ZBTB20 variant genotypes with moderate anxiety in females and CRY1/PER3-A variant genotypes with severe anxiety in males. The association of CRY1/PER3-A variant genotypes with severe anxiety in females was partially mediated by extreme evening chronotype. Our results reinforce existing findings that females exhibit stronger anxiety outcomes than males, and provide evidence for circadian gene associations with anxiety, particularly in females. Our analyses only identified significant associations using two-gene combinations, underscoring the importance of combined gene effects on anxiety risk. We describe novel, robust associations between gene combinations involving the ZBTB20 SNP (rs1394593) and risk of anxiety symptoms in a large population sample. Our findings also support previous findings that the ZBTB20 SNP is an important factor in mood disorders, including seasonal affective disorder. Our results suggest that reduced expression of this gene significantly modulates the risk of anxiety symptoms through direct influences on mood-related pathways. Together, these observations provide novel links between the circadian clockwork and anxiety symptoms and identify potential molecular pathways through which clock genes may influence anxiety risk.
Similar content being viewed by others
Introduction
Mood disorders, including depression and anxiety are becoming increasingly prevalent and affect greater than twenty percent of the world’s adult population1,2. Estimates in Europe suggest that anxiety disorders alone affect nearly one-fifth of the population and that current interventions are ill-equipped to treat this complex disorder3. Every year, anxiety and depression account for over 60 million disability-adjusted life years (DALYs), threatening work productivity and social relationships, among other economic and personal costs4. Efforts to better understand the etiology of mood disorders have uncovered genetic and environmental links between circadian clock-related genes and mood disorders including major depressive disorder (MDD)5,6,7,8,9,10, schizophrenia11, bipolar disorder (BD)7,8,9,10, seasonal affective disorder (SAD)12, and anxiety13,14. While clinical studies have reinforced the roles of circadian rhythm and clock genes in depressive disorders, less is known about how the circadian clockwork influences anxiety risk15.
Circadian rhythms are responsible for the 24-h sleep–wake cycle that revolves around natural and/or artificial light–dark cycles. These rhythms are facilitated by the molecular clock, which regulates over one-third of all transcribed genes responsible for physiological cycles throughout the body through feedback loops in core clock genes and their associated transcription factors16,17. While most tissues in the body individually express these genes, the suprachiasmatic nucleus (SCN), located in the hypothalamus, plays a crucial role in synchronizing and modulating expression in response to environmental cues. For example, the SCN integrates light/dark input from the eyes18,19 and relays this information to the peripheral clock systems via the secretion of neurotransmitters, neuropeptides, growth factors, and cytokines20,21. The transcription factor CLOCK is involved in the core feedback loop, regulating the transcription of Period (PER1, PER2, PER3) and Cryptochrome (CRY1 and CRY2) gene families to maintain the 24-h circadian rhythm cycle. The CLOCK protein forms a heterodimer with brain and muscle ARNT-like protein-1 (BMAL1), which activates the transcription of additional core clock genes, including PER and CRY. PER/CRY heterodimers then inhibit CLOCK/BMAL1 activity, forming a negative feedback loop22. Mutations in these core clock genes may affect mood directly by altering the downstream physiological pathways involved in mood regulation, or indirectly through the disruption of circadian phase.
Many studies have demonstrated an indirect effect of the circadian clock on mood disorders by linking chronotype or diurnal preferences, differences in sleep–wake timing and activity patterns, with negative affect13,14,23,24,25,26,27. These studies support the ‘social jetlag’ hypothesis which posits that alterations in circadian rhythms influence mood by creating a misalignment between an individual’s sleep–wake activity and their social routine7,26,28,29,30,31,32,33,34. Moreover, genome-wide association studies (GWAS) and candidate gene studies have provided strong evidence that circadian clock genes are associated with diurnal preference, chronotype, and sleep disturbance [35,36,37,38,39]. Insights from GWAS studies also suggest overlapping genetic profiles between individuals with altered chronotype and individuals with mood disorders35,39. However, GWAS studies on anxiety have not yet identified significant associations between circadian clock genes and risk40,41. Core circadian genes may also have direct effects as transcriptional regulators, influencing neurotransmitter signaling in serotonin, dopamine, and glucocorticoid mood pathways42. Such links suggest the potential for direct and indirect influences of circadian clock genes on mood disorders, although the molecular mechanisms that facilitate these effects remain poorly understood. In addition, clock genes often affect neurotransmitter signaling and circadian alignment simultaneously, making it difficult to disentangle the indirect and direct effects of clock genes on mood.
Recent studies have demonstrated the involvement of clock genes in mood disorders (Table 1). For example, recent studies have found that the knockdown of CRY116, CRY243, and PER344 clock genes affected anxiety, depression behaviors, and circadian timing. Population-level candidate gene studies9,13,45 and mathematical modeling25 have also identified clock genes important to mood disorders. Although GWAS studies have struggled to identify genotypes associated with complex mood disorder phenotypes14, Ho and colleagues (2018) found that an intronic SNP variant in ZBTB20 (rs1394593) exhibited a strong association with seasonal affective disorder46. This study also found strong association signals for candidate gene targets of ZBTB20, suggesting that ZBTB20-regulated pathways have an important role in SAD etiology46. In addition, methylome changes in ZBTB20 have previously been associated with major depressive disorder47. However, candidate gene studies have yet to investigate the influence of ZBTB20 on circadian and mood pathways.
In this study, we investigate associations of clinical features and SNP variants in core clock genes, including ZBTB20, with anxiety (Generalized Anxiety Disorder 7-item scale (GAD-7)) scores in a UK Biobank population of 90,882 individuals, aged 40–69 years old. We explore the effects of single-gene variants and multi-gene variant combinations utilizing a complementary approach of machine learning and statistical methods. Here, we report strong sex-specific associations of circadian gene variants with anxiety symptoms and propose that a variant of ZBTB20 (rs1394593) influences anxiety risk via amplification or attenuation of anxiety symptoms in combination with other clock variants.
Methods
Data sourcing
All data used in the study originates from the UK Biobank, a large prospective study comprised of approximately 500,000 individuals ages 40–69 that were recruited from 2006 to 2010. UK Biobank is a resource for studies investigating the genetic, environmental, and lifestyle determinants of a wide range of diseases present in middle and later life48. At the beginning of the initial assessment, written consent was obtained from each study participant48. Individuals underwent body measurements, provided lifestyle information and medical history, and donated blood, urine, and saliva samples for genetic and phenotypic investigations49. We did not exclude by ethnicity in this study; 94.6% of UK Biobank participants are recorded as ‘White,’ and 5.4% are recorded as ethnic minorities106.
UK biobank features
Supplementary Fig. 1 provides a summary of methods employed in this study. UK Biobank data was translated into a readable form using ukbb_parser50. We analyzed the following single-nucleotide polymorphisms (SNPS): CLOCK (rs1801260), PER2 (rs10462023), PER3 variants PER3-A (rs228697), PER3-B (rs17031614) and PER3-C (rs10462020), CRY1 (rs2287161), CRY2 (rs1083852), and ZBTB20 (rs1394593), which were all collected by UK Biobank researchers. In addition, we examined six behavioral/clinical features: age (ID: 21003), sex (ID:31), chronotype (ID: 1180), household income (ID: 738), substance addiction (ID: 20457), and Townsend Material Deprivation score (ID: 22189). Substance addiction was defined by self-report answers to a survey question asking participants if they had an ongoing addiction or dependence on illicit or recreational drugs, and they were given the answer options “No”, “Yes”, and “Prefer not to answer”. Chronotype was defined by self-report responses where individuals classified themselves as definite morning or evening-types, partial morning or evening-types, or had the option of selecting “I don’t know” or “prefer not to answer”. Household income was also collected through self-report where individuals were asked to report their average household income by selecting one of five pre-specified ranges, also with “I don’t know” or “Prefer not to answer” options. Individuals who selected “I don’t know” or “Prefer not to answer” for any of the covariates of interest were removed from the analysis. Townsend Material Deprivation score was calculated by participant postal code.
Anxiety was assessed using the Generalized Anxiety Disorder 7-item scale51. Additional measures related to anxiety are available from the UK Biobank database, including the ICD-10 (International Classification of Disease for anxiety disorder), clinical diagnoses of anxiety, the use of anxiolytic medicine, and self-reported measures of anxiety-related doctor visits and symptoms of anxiety. Although these variables, particularly the clinical diagnoses of and treatments for anxiety, are excellent measures of anxiety disorders, using these measures significantly reduces the sample size for statistical analysis of gene variant associations. Thus, we chose to include the well-supported GAD-7 instrument as the anxiety measure to maximize the sample size for our analyses.
Feature engineering and selection
To account for the effects of population structure and batch-based genotyping, UK Biobank researchers utilized several stages of quality control [52,53]. First, several different SNP-based metrics were used identify and eliminate less reliable genotyping results. If SNPs were missing in multiple batches, then they were removed from analysis [53]. SNPs with a minor allele frequency less than 1 percent (MAF < 1%) were removed from analysis [53]. Next, researchers focused exclusively on high-quality SNPs to identify poor-quality samples [53]. Finally, principal components analysis and relatedness inference were used for sample-based inference. From these quality control steps, UK Biobank researchers identified few SNPs and samples to be removed [53]. These researchers performed whole-genome imputation with IMPUTE2 using a diverse reference panel. Imputation information scores were used to assess imputation, and these scores revealed effective imputation for SNPs of varying MAFs [52]. In our analysis, any individuals lacking relevant data pertaining to any of our selected features or outcome were removed from the population analyzed; no imputation was performed for these missing values. This left 90,882 UK Biobank participants for analysis.
One-hot encoding was performed to transform categorical variables into numeric values that can be read for machine learning analysis. For categorical variables with n categories, n-1 new columns were created. One of the categories was considered the reference category and excluded, since it could be inferred from the other columns. With SNP data, for example, two new columns were created for the two less frequent genotypes, and the common genotype column was considered as the reference category (Supplementary Fig. 2). SNP data was also imputed for CLOCK, PER2, PER3-A, and PER3-C, consistent with the imputation that UK Biobank researchers had already performed on CRY1, CRY2, PER3-B, and ZBTB20 using which were imputed by UK Biobank researchers using the Haplotype Reference Consortium54, and UK10K and 1000 Genomes reference panels55,56. These 6 clinical features and 8 single-genotype features were examined for associations with anxiety. Also, we investigated gene combinations involving two genotypes and their respective variants to examine pairwise interactions. This generated an additional 8C2*8 two-way features; of eight total genes, any two can exist in a pair and there are eight potential genotypic combinations that paired genes may have, resulting in 224 total genotype combinations.
The GAD-7 is a self-report scale that ranges from 0 to 21. GAD-7 scores have commonly been broken into thresholds where scores ≤ 4 indicate minimal anxiety, 5–9 suggest mild anxiety, 10–14 suggest moderate anxiety, and ≥ 15 indicate severe anxiety. Since a score of 8 is a commonly held cutoff for symptoms of mild anxiety57,58, we used this supported cutoff to establish anxiety presence. From here, we adhered to the cutoffs for anxiety severity and used the following thresholds: mild anxiety (8–11), moderate anxiety (12–15), and severe anxiety (≥ 16). Individuals with a GAD-7 score < 8 were used as controls for the data analysis. We split these categories into three different binary outcomes for multiple multivariate logistic regression analyses. Our data was reprocessed four times into four total separate datasets where the data either was or was not one-hot encoded and the outcome variable was either binary or continuous, so that multivariate linear regression, multivariate logistic regression, the Sheirer Ray Hare Test, and mediation analysis could be performed with the correct types of data.
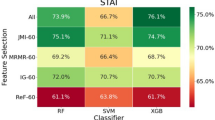
We used a combination of feature selection methods to determine which features to use in our subsequent analyses. We used a combination of multiple ranking-based and subset-based feature selection methods to mitigate the inherent bias of individual feature selection methods, choosing features that were ranked highly by all feature selection algorithms. Chi-square, InfoGain (IG), and ReliefF (ReF) are ranking-based feature selection methods that rank features by their contribution to the disorder outcome. The chi-square method calculates the association between features and anxiety outcomes using the chi-squared score59. InfoGain (IG) ranks features based on the amount of entropy each feature explains60,61. ReliefF (ReF) scores features based on their value, relative to their nearest-neighbor instance62. Joint mutual information (JMI) and minimum redundancy maximum relevance (MRMR) are subset-based feature selection methods. Both of these methods determine subsets in the feature space, and select feature subsets that have the strongest relationships with an outcome and the weakest relationship with other features by evaluating and comparing these two different interactions. Joint mutual information (JMI) selects features for a subset that maximize the cumulative sum of joint mutual information when added to the subset63. Minimum redundancy maximum relevance (MRMR) iteratively selects a subset of features that have the most correlation with the class, and the least correlation with other features64. Bootstrapping was performed for each feature selection method by running it 50 times and taking features that appeared in the top 50 features at least 70 percent of the time. Then, the results across these five methods were compared using a sum of ranks and the 25 features that performed the best over the five techniques were used in our statistical analysis.
Statistical analyses
All statistical analyses were performed in R [65] and the Scikit-learn library in Python66. Our code is available at: https://github.com/cventresca/ukbb_analyzer.
Since there was still high dimensionality in the dataset following feature selection and we observed initial overfitting of our model, we used Variance Inflation Factor (VIF) and Akaike Information Criterion (AIC) as we performed our regressions to identify important features. VIF was used to identify multicollinearity and variables with a VIF score > 10 were excluded from the analysis67. We also used AIC, which is a model selection algorithm that uses sequential replacement to identify features with low multicollinearity and strong association with GAD-768. Features that were deemed important by both selection methods, in addition to machine learning feature selection methods described above, were used in subsequent multivariate analyses.
Multivariate linear and logistic regression analyses were performed using the Statsmodels library in R69. We performed multivariate linear regression to predict continuous GAD-7 scores with genotypic and clinical independent variables. First, a Durbin-Watson statistic was obtained to check the independence of residuals assumption70, and then a scatterplot was constructed to confirm a linear relationship between each independent feature and the collective of independent features with GAD-7 scores. Studentized residuals were plotted against the unstandardized predicted values to check that the assumption of homoscedasticity was met. No outliers, high leverage points, or highly influential points were detected during analysis, and the normality of residuals was confirmed via a histogram with a superimposed normal curve and a P–P Plot. After performing multivariate linear regression, P-value corrections were performed using the Benjamini Hochberg (BH) correction, due to unplanned pairwise comparisons between features71. Next, multivariate logistic regression was performed to assess genotypic and clinical predictors for mild, moderate, and severe anxiety outcomes. A linear relationship was confirmed between continuous independent variables and the logit-transformed GAD-7 outcome for all three anxiety classifications, and no outliers, high leverage points, or highly influential points were detected during analysis. Following the analysis, p-values were also adjusted for multivariate logistic regression using the Benjamini Hochberg (BH) procedure71.
We sought to analyze the two-way interactions (genotype and sex) of the SNP combinations that appeared in our multivariate analysis using two-way ANOVA. However, the Shapiro–Wilk test of normality showed that our data was not normally distributed72. Therefore, to identify sex-specific differences in average GAD-7 scores for two-way gene combinations, we performed the Scheirer Ray Hare Test in R73, which instead compares the median GAD-7 scores across groups. Bar plots of these significant combinations revealed that GAD-7 score distributions were similar enough in shape across groups to have their medians compared.
Mediation analysis was performed to examine whether features were directly associated with anxiety or indirectly associated with anxiety through extreme morning or extreme evening chronotype. Our mediation analysis was conducted in the Mediation library in R74, and this analysis was completed for SNP combinations that were significant in the multivariate logistic regression analysis and their associated anxiety outcome(s). This analysis was performed two times for each combination: once with extreme morning type as the mediator and once with extreme evening type as the mediator. Clinical variables including addiction, age, income, and Townsend deprivation index scores were used as confounders when determining the mediation effect of chronotype.
Results
Risk and protective factors for anxiety
Average GAD-7 scores were higher in females than males for several combinations, regardless of genotype (Fig. 1; CRY2_AG/ZBTB20_TT: H1,60584 = 827.94, p < 0.0001; PER2_AG/ZBTB20_TT: H1,60584 = 827.15, p < 0.0001; PER3-B_GG/ZBTB20_TT: H1,60584 = 828.47, p < 0.0001; CLOCK_AA/ZBTB20_TT: H1,60584 = 828.46, p < 0.0001). Extreme evening type behavior was associated with increased odds of severe anxiety (Table 2; OR 1.4(1.16–1.79) adj. p = 0.0021), mild anxiety (OR 1.1(1.02–1.27) adj. p = 0.028) and was identified as a risk factor by linear regression (estimate 0.01(0.0060–0.015) adj. p = 0.0000036).
Scheirer-Ray-Hare test reveals the effects of genotype and gender on GAD-7 scores. (a) There was an effect of genotype and sex on GAD7 score for CRY2-AG/ZBTB20-TT (Genotype: H1,60584 = 4.66, p = 0.031, Gender: H1,60584 = 827.94, p < 0.0001, Genotype x Gender: H1,60584 = 2.09, p = 0.15). (b) There was an effect of genotype and sex on GAD7 score for CLOCK-AA/ZBTB20-TT (Genotype: H1,60584 = 4.40, p = 0.036, Gender: H1,60584 = 828.46, p < 0.0001, Genotype x Gender: H1,60584 = 0.51, p = 0.48). (c) There was an effect of genotype and sex on GAD7 score for PER2-AG/ZBTB20-TT (Genotype: H1,60584 = 10.8, p = 0.0010, Gender: H1,60584 = 827.15, p < 0.0001, Genotype x Gender: H1,60584 = 0.10, p = 0.75). (d) There was an effect of genotype and sex on GAD7 score for PER3-B-GG/ZBTB20-TT (Genotype: H1,60584 = 3.42, p = 0.064, Gender: H1,60584 = 828.47, p < 0.0001, Genotype x Gender: H1,60584 = 1.20, p = 0.27).
For both sexes, CRY2_AG/ZBTB20_TT was associated with a 120-fold increase in odds of severe anxiety (OR 120(10.1–1420) adj. p = 0.00029), a sixfold increase in odds of moderate anxiety (OR 6.1(1.99–18.50) adj. p = 0.0020), and a 13-fold increase in odds of mild anxiety (OR 13.4(2.02–89.20) adj. p = 0.011). PER3-A_CG/ZBTB20_TT was associated with a near 40-fold reduction in odds of severe anxiety (OR 0.02(0.0011–0.57) adj. p = 0.034), and a ten-fold reduction in odds of mild anxiety (OR 0.1(0.013–0.87) adj. p = 0.045). CLOCK_AA/ZBTB20_TT was associated with a five-fold reduction in odds of mild anxiety (OR 0.2(0.05–0.80) adj. p = 0.09). PER2_AG/ZBTB20_TT was revealed as a protective factor by linear regression (estimate − 0.03(− 0.054 to − 0.0040) adj. p = 0.021).
Sex-specific risk and protective factors for anxiety in males and females
In females, being an extreme evening type was associated with 1.6 times the odds of severe anxiety (Table 3; OR 1.6(1.22–2.08) adj. p = 0.0011), and eveningness appeared as a risk factor in multivariate linear regression (estimate 0.01(0.005–0.018) adj. p = 0.00088). Being an extreme morning type was associated with reduced odds of moderate anxiety (OR 0.8(0.68–0.86) adj. p = 0.0000045), mild anxiety (OR 0.8(0.77–0.92) adj. p = 0.00040) and was also identified as a protective factor by multivariate linear regression (estimate − 0.006(− 0.010 to − 0.0030) adj. p = 0.0012).
In females, CRY2_AG/ZBTB20_TT was associated with an > 200-fold increase in odds of moderate anxiety (OR 219(9.02–5.32E + 03) adj. p = 0.0016), and a > 40 times increase in odds of mild anxiety (OR 43.4(2.34–807) adj. p = 0.017). CRY2_AG/ZBTB20_TT also appeared as a risk factor in multivariate linear regression (estimate 0.060(0.012–0.109) adj. p = 0.017). CRY2_GG/ZBTB20_TT was associated with ~ 25-fold increased odds of moderate anxiety (OR 24.6(1.28–471) adj. p = 0.039). CRY1_GG/PER3-A_GG was associated with a 2.5-fold increase in odds of severe anxiety (OR 2.5(1.32–4.86) adj. p = 0.0093.
Also, logistic regression revealed that PER3-A_CG/ZBTB20_TT was associated with a near five-fold reduction in odds of moderate anxiety (OR 0.02(0.00057–0.47) adj. p = 0.02), and a greater than 20-fold decrease in odds of mild anxiety in females (OR 0.04 (95% CI) adj. p = 0.049). PER3-A_CC/ZBTB20_TT was associated with a near five-fold decrease in odds of moderate anxiety (OR 0.02(0.0011–0.49) adj. p = 0.02).
For males, multivariate linear regression identified extreme evening type as a risk factor (Table 4; estimate 0.01(0.0040–0.016) adj. p = 0.0016). Also, being an extreme morning type was identified as a protective factor for mild anxiety by logistic regression (OR 0.9(0.75–0.98) adj. p = 0.028) and was revealed as protective by linear regression in males (estimate − 0.004(− 0.0080–0.000) adj. p = 0.033). In addition, CRY2_AG/ZBTB20_TT was associated with a ~ 20-fold increase in odds of severe anxiety (OR 22.5(2.72–186) adj. p = 0.0078). CRY1_CC/PER3-A_GG was associated with nearly three times the odds of severe anxiety (OR 2.8(1.22–6.49) adj. p = 0.028). PER3-B_GG/ZBTB20_TT was associated with a ~ sevenfold increase in odds of mild anxiety (OR 7.3(1.47–36.60) adj. p = 0.024).
Mediation analyses reveal direct and mediated effects on anxiety symptoms
In females, CRY2_AG/ZBTB20_TT exhibited a direct association with moderate anxiety (Fig. 2b; morning type estimate = 0.047(0.0054–0.12) p = 0.020; evening type estimate = 0.054(0.0063–0.15) p = 0.020). CRY1_GG/PER3-A_GG displayed two associations with severe anxiety: a direct association (Fig. 2a; morning type estimate = 0.017(0.0034–0.040) p = 0.02; evening type estimate = 0.018(0.0040–0.040) p = 2.0E−16), and one mediated by extreme evening type (Fig. 3a; evening type estimate = − 0.00056(− 0.0011–0.00) p = 2.0E-16). In males, CRY1_CC/PER3-A_GG exhibited a direct association with severe anxiety (Fig. 3; morning type estimate = 0.012(0.0011–0.040) p = 0.02; evening type estimate = 0.017(0.00098–0.050) p = 0.04).
Mediation analysis reveals a combination directly associated with anxiety and a combination whose association is partially mediated by chronotype in females. Mediation analysis was performed in females for mild, moderate, and severe anxiety classifications with extreme morning type and extreme evening type entered as potential mediators. The black line from genotypic combination to outcome indicates a direct effect. Red dotted lines indicate a mediation effect through extreme evening type, and black dotted lines that are crossed indicate no mediation effect. (a) A direct effect was observed between CRY1_GG/PER3-A_GG and severe anxiety with extreme morning type as the mediator (p = 0.02), and a partial mediation effect was observed with extreme evening type as the mediator (direct p = 0.04; indirect p = 2E-16). (b) A direct effect between CRY2_AG/ZBTB20_TT with moderate anxiety was observed (direct morning p = 0.02; direct evening p = 0.04).
Mediation analysis reveals CRY1_CC/PER3-A_GG is directly associated with severe anxiety in males. Mediation analysis was performed in males for mild, moderate, and severe anxiety classifications with extreme morning type and extreme evening type entered as potential mediators. The black line from genotypic combination to outcome indicates a direct effect. Red dotted lines indicate a mediation effect through extreme evening type, and black dotted lines that are crossed indicate no mediation effect.
Decision trees summarize genotypic associations with anxiety symptoms
For both sexes, ZBTB20 variants occurred in a risk combinations with CRY2_AG and protective combinations with PER3-A_CG (Fig. 4). We constructed association networks to provide a visual summary of our analysis. For females, the CRY1_GG/PER3-A_GG combination was associated with severe anxiety both directly and indirectly through extreme evening type (Fig. 5). ZBTB20 was associated with increased anxiety risk in combinations with CRY2_AG and CRY2_GG and was associated with decreased anxiety risk in combinations with PER3-A_CG and PER3-A_CC (Fig. 4). In males, the CRY1_CC/PER3-A_GG combination was directly associated with severe anxiety, and ZBTB20 variants occurred in risk combinations with CRY2_AG and PER3-B_GG (Fig. 6).
Decision tree summarizes multivariate and mediation analysis findings for genotypes associated with anxiety in both sexes. A decision tree was constructed to visualize associations between clock genes and anxiety that appeared in the overall dataset. Red and blue ovals represent genotypes belonging to risk and protective combinations, respectively, while gray ovals represent genotypes belonging to both risk and protective combinations. Dashed lines represent associations with anxiety, supported by multivariate regression, while solid lines represent effects found by mediation analysis.
Decision tree summarizes multivariate and mediation analysis findings for genotypes associated with anxiety in females. A decision tree was constructed to visualize associations between clock genes and anxiety that appeared in females. Red and blue ovals represent genotypes belonging to risk and protective combinations, respectively, while gray ovals represent genotypes belonging to both risk and protective combinations. Dashed lines represent associations with anxiety, supported by multivariate regression, while solid lines represent effects found by mediation analysis (*Exhibited partial mediation by extreme evening type).
Decision tree summarizes multivariate and mediation analysis findings for genotypes associated with anxiety in males. A decision tree was constructed to visualize associations between clock genes and anxiety that appeared in males. Red ovals represent genotypes belonging to risk combinations. Dashed lines represent associations with anxiety, supported by multivariate regression, while solid lines represent effects found by mediation analysis.
Discussion
There is a growing body of evidence supporting the roles of clock gene variants and circadian disruption in anxiety. However, many of these studies have utilized GWAS and PheWAS approaches making it difficult to detect synergistic effects between genotypes or to explore whether significant genotypes are directly or indirectly associated with anxiety. Novel machine learning approaches have shown promise in illuminating these synergistic effects and proposing potential mechanisms of clock pathways that may influence anxiety14 and sleep disturbance38. Utilizing similar machine learning approaches to analyze clock gene associations with anxiety in a large UK Biobank dataset, we report three main findings: (1) Clock genotype combinations including ZBTB20 variants exhibit combination-specific effects on anxiety, (2) Clock variant combinations associated with anxiety tend to display sex-specific effects, and (3) Circadian-related variants linked to anxiety risk have both direct (chronotype-independent) influences and indirect (chronotype-mediated) influences on anxiety symptoms.
Genotype combinations with ZBTB20 exhibit diverse associations with anxiety
In this study, we did not observe any single-gene associations with anxiety; only genotype combinations were identified as significant predictors for GAD-7 outcomes. ZBTB20_TT was present in nearly every risk combination—with CRY2 and PER3-B and protective combination—with CLOCK and PER3-A. These findings reinforce results from previous GWAS and target gene studies and suggest that ZBTB20_TT may have an important regulatory effect on clock genes involved in anxiety. ZBTB20 is a zinc finger transcriptional repressor protein that is abundant in the hippocampus and is known to have an important role in hippocampal development75,76. Previous research from Ho et al.46 revealed that the minor T-allele of ZBTB20 was associated with lower ZBTB20 mRNA expression and an increased risk for seasonal affective disorder (SAD)46. These authors also found that 32 genes associated with SAD were enriched when ZBTB20 levels were reduced46 suggesting that ZBTB20 plays an important role in the regulation of clock gene expression. Indeed, other studies have found that ZBTB20 loss is associated with impaired circadian rhythms77, and that epigenetic changes inhibiting ZBTB20 expression are associated with MDD47.
Our findings provide further support for ZBTB20 as a regulator for circadian clock genes and demonstrate that genotype combinations that include ZBTB20 variants can exhibit sex-specific outcomes on anxiety symptoms. In previous studies, the A-allele of CRY2 has been associated with chronicity patterns characteristic of depressive symptoms78, and CRY2_AG has appeared in a risk combination for anxiety14. Decreases in CRY2 mRNA have previously been observed in depressed bipolar patients6, suggesting that ZBTB20 could act as a repressor of another gene that represses CRY2 transcription. Therefore, reductions in ZBTB20 could indirectly contribute to lower CRY2 expression.
This trend was also observed in females who exhibited protective PER3-A combinations with ZBTB20_TT (PER3-A_CC/ZBTB20_TT and PER3-A_CG/ZBTB20_TT). The PER3-A G-allele and GG genotype have previously been associated with MDD25,79, anxiety13, and eveningness25,80. Previous mathematical modeling insights suggest that greater PER3 stability contributes to slight increases in period and large reductions in clock amplitude, contributing to circadian misalignment25. Thus, ZBTB20 could serve as a repressor for the transcription of PER3.
In males, PER3-B_GG/ZBTB20_TT was associated with increased odds of mild anxiety. However, the A-allele of PER3-B has previously been the allele associated with increased odds of MDD and anxiety14,79, suggesting a potential combination-specific effect. Indeed, increases in PER3-B expression have been shown to associate with circadian disruption19,81. Loss of ZBTB20 repression activity may lead to increased PER3-B expression and greater circadian disruption, influencing the likelihood of anxiety symptoms.
Finally, we observed that the CLOCK_AA/ZBTB20_TT and PER2_AG/ZBTB20_TT genotype combinations were protective factors for both sexes. The C-allele of CLOCK has been associated with MDD79, seasonal depression24, and evening chronotype82, supporting a protective association for CLOCK_AA/ZBTB20_TT in both sexes. The G-allele of PER2 has previously been associated with depression vulnerability5, so this protective effect may function through alterations in PER2 expression. As the transcription of PER2 is finely tuned in response to environmental light83 and increased PER2 stabilization leads to circadian disruption in mice84, ZBTB20 may act to inhibit a repressor of the PER2 gene.
Genotype combinations exhibit sex-specific associations with anxiety
Our main findings in the regression analyses provide support for sex-specific associations of circadian genotypes with anxiety. CRY1_GG/PER3-A_GG, CRY2_GG/ZBTB20_TT, and CRY2_AG/ZBTB20_TT were risk factors that showed stronger associations in females. Also, PER3-A_CG/ZBTB20_TT and PER3-A_CC/ZBTB20_TT were protective for mild and moderate anxiety in females. In males, CRY1_CC/PER3-A_GG was a risk factor for severe anxiety and PER3-B_GG/ZBTB20_TT was a risk factor for mild anxiety. Sex-specific associations with circadian genes have previously been observed for anxiety14 and major depressive disorder (MDD)79.
Sex-specific associations of CRY2/ZBTB20 and PER3-A/ZBTB20 with anxiety in females suggest that these combinations are involved in sex-specific pathways. We observed that CRY2 combinations (CRY2_GG/ZBTB20_TT and CRY2_AG/ZBTB20_TT) showed significantly stronger associations with anxiety in females than in males. Because CRY2_AG/ZBTB20_TT also appeared as a risk factor in males, this combination may exert effects through a shared pathway in both sexes. The sex-specific association of PER3-A in females is supported by a previous association with MDD79. We also observed that CRY1_GG/PER3-A_GG was associated with severe anxiety in females. Since the C-allele of CRY1 is the risk allele associated with depression25,27,85, and the G-allele of PER3-A is the risk allele associated with MDD25,79, anxiety13, and eveningness25,80, these findings provide further support for the sex-specific involvement of PER3-A in female anxiety risk.
In males, CRY1_CC/PER3-A_GG was a risk factor for severe anxiety. Since this combination includes the risk alleles for both genes, this combination could indicate that both genotypes affect anxiety symptoms independently, or in a combination-specific manner. Also, PER3-B_GG/ZBTB20_TT appeared as a risk factor for mild anxiety in males and this genotype combination does not appear to be a risk factor for females.
There are multiple pathways by which clock gene variants may exert sex-specific effects. Glucocorticoid regulation may be a potential sex-dependent pathway by which these genotypes modulate one’s odds of anxiety86. The glucocorticoid pathway has previously been implicated in mood disorders87 and clock gene pathways modulate the release of and sensitivity to glucocorticoids88. In addition, PER3-A and PER3-B have been identified in several associations with the sleep–wake cycle and diurnal preference13,38,80, which are hypothesized to alter mood through the regulation of serotonin89,90,91. As the function of the 5-HT serotonin system is intertwined with the circadian system92,93, and this system affects mood94, serotonin regulation has been implicated as a pathway by which circadian disruption can lead to effects on mood82,89,95,96. Furthermore, anxiety symptoms have been shown to associate closely with shifts in serotonin activity97,98, and sex differences have been observed in serotonergic transmission99,100. Therefore, circadian disruptions due to PER3-A and PER3-B variants could affect mood in a sex-specific manner through alterations in 5-HT signaling. These suggested pathways are supported by previous GWAS studies on anxiety, which have identified other genes known to function in neurotransmitter signaling pathways40,41. However, previous GWAS on anxiety have not yet identified associations of clock genes with anxiety.
Effects on anxiety may be direct or mediated through chronotype
We observed that CRY2_AG/ZBTB20_TT was directly associated with moderate anxiety in females, suggesting this genotypic combination exerts direct effects on mood in females. Previously, Zafar et al.14 also found that CRY2_AG was directly associated with anxiety symptoms14. The results of the current study suggest that decreases in the transcriptional repression of CRY2 by ZBTB20 may lead to greater transcription of CRY2, which exerts direct effects on anxiety through mood-related pathways.
For both sexes, our multivariate analyses revealed that extreme evening type behavior was associated with an increased risk of anxiety, while extreme morning type was protective against anxiety. Our findings are supported by large-scale GWAS studies that identified clock genes involved in the core feedback loop to be associated with alterations in sleep/wake timing [35,37]. Silva and colleagues (2020) suggested that genotypic variants associated with shifts in chronotype may indirectly affect one’s odds of anxiety through the development of symptoms characteristic of various mood disorders101. Interestingly, we found that the association between CRY1_GG/PER3-A_GG and severe anxiety in females was partially mediated by extreme evening type behavior. CRY genes activate the circadian loop and function in the retina as light-independent inhibitors of CLOCK/BMAL heterodimers102,103, suggesting their role in circadian rhythm maintenance. As stated above, this PER3-A variant (rs228697) has previously been associated with evening type behavior13,25, and a significantly higher risk of anxiety13. Altogether, these findings suggest that the modulation of CRY1 expression may lead to alterations in sex-specific mood pathways and diurnal preference pathways in ways that are conducive to anxiety in females. Interestingly, the anxiety risk associated with the co-occurrence of CRY1_GG with PER3-A_GG in females is similar in magnitude to the male-specific association for CRY1_CC/ PER3-A_GG genotypes, suggesting that CRY1 homozygotes, in the presence of PER3-A_GG, may affect anxiety via distinct sex-specific mechanisms.
Limitations
Previous studies have suggested that the UK Biobank population may have a “healthy volunteer” selection bias because only 5% of recruited individuals responded to the invitation. Thus, the Biobank cohort study may not be representative of the UK population104,105. For example, UK Biobank study participants were less likely to be socioeconomically deprived, obese, smoke, drink alcohol on a daily basis, and have self-reported health conditions106. To counteract the healthy volunteer bias present in the UK Biobank, we controlled for issues of economic status and sex to maximize the generalizability of our results. The UK population and, accordingly, the UK Biobank participants, are predominantly of European Caucasian descent with nearly 95% of the database identifying as ‘White’. Although we did not exclude by ethnicity, our results may not be generalizable to populations of non-Caucasian descent given the small representation of ethnic minorities in the analyses. In addition, the UK Biobank offers additional measures of anxiety, including clinical diagnoses, that could be used to test for associations of clock genes with anxiety. However, these measures offered smaller sample sizes relative to the GAD-7 instrument. Thus, we chose to utilize the well-supported GAD-7 instrument to maximize the power of our analyses. Finally, to minimize computational requirements, we selected circadian gene variants that had been linked to chronotype and/or mood disorders in previous studies and did not study all possible circadian-related variants. Therefore, this study may be missing important circadian features that influence anxiety.
Conclusions
In this study, we report sex-dependent, combination-specific, and indirect and direct effects of circadian genotypes on anxiety. ZBTB20 was a feature in several risk and protective genotypic combinations, occurring with CRY2 and PER3-B in risk combinations, and with CLOCK and PER3-A in protective combinations. Several additional clock-related genes were involved in sex-specific associations with anxiety and these polymorphisms likely influence pathways involved in glucocorticoid and serotonin regulation. Together, these observations reinforce previous GWAS insights into the associations of ZBTB20 with mood pathways and suggest that ZBTB20 may have a critical regulatory role, both as a repressor and indirect activator, in the transcription of clock genes. In females, we found that the CRY2_AG/ZBTB20_TT genotype, our strongest predictor of anxiety, was directly associated with anxiety. The CRY1_GG/PER3-A_GG genotype in females exhibited effects partially mediated by extreme evening-type behavior, suggesting that circadian effects on anxiety can be both direct and/or mediated by chronotype.
Data availability
UK Biobank data are available upon an application process, accessed at: https://www.ukbiobank.ac.uk/enable-your-research.
References
Nochaiwong, S. et al. Global prevalence of mental health issues among the general population during the coronavirus disease-2019 pandemic: A systematic review and meta-analysis. Sci. Rep. 11, 10173. https://doi.org/10.1038/s41598-021-89700-8 (2021).
Steel, Z. et al. The global prevalence of common mental disorders: A systematic review and meta-analysis 1980–2013. Int. J. Epidemiol. 43, 476–493. https://doi.org/10.1093/ije/dyu038 (2014).
Wittchen, H. U. et al. The size and burden of mental disorders and other disorders of the brain in Europe 2010. Eur. Neuropsychopharmacol. 21, 655–679. https://doi.org/10.1016/j.euroneuro.2011.07.018 (2011).
Ferrari, A. et al. Global, regional, and national burden of 12 mental disorders in 204 countries and territories, 1990–2019: A systematic analysis for the Global Burden of Disease Study 2019. The Lancet Psychiatry https://doi.org/10.1016/S2215-0366(21)00395-3 (2022).
Lavebratt, C., Sjöholm, L. K., Partonen, T., Schalling, M. & Forsell, Y. PER2 variantion is associated with depression vulnerability. Am. J. Med. Genet. B Neuropsychiatr. Genet. 153, 570–581. https://doi.org/10.1002/ajmg.b.31021 (2010).
Lavebratt, C. et al. CRY2 is associated with depression. PLoS ONE 5, e9407. https://doi.org/10.1371/journal.pone.0009407 (2010).
McCarthy, M. J. & Welsh, D. K. Cellular circadian clocks in mood disorders. J. Biol. Rhythms 27, 339–352. https://doi.org/10.1177/0748730412456367 (2012).
Nievergelt, C. M. et al. Examination of the clock gene Cryptochrome 1 in bipolar disorder: Mutational analysis and absence of evidence for linkage or association. Psychiatr. Genet. 15, 45–52. https://doi.org/10.1097/00041444-200503000-00008 (2005).
Partonen, T. Clock gene variants in mood and anxiety disorders. J. Neural Transm. (Vienna) 119, 1133–1145. https://doi.org/10.1007/s00702-012-0810-2 (2012).
Roybal, K. et al. Mania-like behavior induced by disruption of CLOCK. Proc. Natl. Acad. Sci. USA 104, 6406–6411. https://doi.org/10.1073/pnas.0609625104 (2007).
Ashton, A. & Jagannath, A. Disrupted sleep and circadian rhythms in schizophrenia and their interaction with dopamine signaling. Front. Neurosci. 14, 636. https://doi.org/10.3389/fnins.2020.00636 (2020).
Garbazza, C. & Benedetti, F. Genetic factors affecting seasonality, mood, and the circadian clock. Front. Endocrinol. 9, 481. https://doi.org/10.3389/fendo.2018.00481 (2018).
Liberman, A. R. et al. Circadian clock model supports molecular link between PER3 and human anxiety. Sci. Rep. 7, 9893. https://doi.org/10.1038/s41598-017-07957-4 (2017).
Zafar, A., Overton, R., Attia, Z., Ay, A. & Ingram, K. Machine learning and expression analyses reveal circadian clock features predictive of anxiety. Sci. Rep. 12, 5508. https://doi.org/10.1038/s41598-022-09421-4 (2022).
Ozburn, A. R. et al. Functional implications of the CLOCK 3111T/C single-nucleotide polymorphism. Front. Psychiatry 7, 67. https://doi.org/10.3389/fpsyt.2016.00067 (2016).
Evans, J. A., Pan, H., Liu, A. C. & Welsh, D. K. Cry1-/- circadian rhythmicity depends on SCN intercellular coupling. J. Biol. Rhythms 27, 443–452. https://doi.org/10.1177/0748730412461246 (2012).
Walker, W. H., Walton, J. C., DeVries, A. C. & Nelson, R. J. Circadian rhythm disruption and mental health. Transl. Psychiatry 10, 28. https://doi.org/10.1038/s41398-020-0694-0 (2020).
Buhr, E. D. & Takahashi, J. S. Molecular components of the Mammalian circadian clock. Handb. Exp. Pharmacol. https://doi.org/10.1007/978-3-642-25950-0_1 (2013).
Takahashi, J. S., Hong, H.-K., Ko, C. H. & McDearmon, E. L. The genetics of mammalian circadian order and disorder: Implications for physiology and disease. Nat. Rev. Genet. 9, 764–775. https://doi.org/10.1038/nrg2430 (2008).
Abrahamson, E. E. & Moore, R. Y. Suprachiasmatic nucleus in the mouse: Retinal innervation, intrinsic organization and efferent projections. Brain Res. 916, 172–191. https://doi.org/10.1016/s0006-8993(01)02890-6 (2001).
Lee, J. E. et al. Endogenous peptide discovery of the rat circadian clock: A focused study of the suprachiasmatic nucleus by ultrahigh performance tandem mass spectrometry. Mol. Cell Proteomics 9, 285–297. https://doi.org/10.1074/mcp.M900362-MCP200 (2010).
Cao, X., Yang, Y., Selby, C. P., Liu, Z. & Sancar, A. Molecular mechanism of the repressive phase of the mammalian circadian clock. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2021174118 (2021).
Archer, S. N. & Oster, H. How sleep and wakefulness influence circadian rhythmicity: Effects of insufficient and mistimed sleep on the animal and human transcriptome. J. Sleep Res. 24, 476–493. https://doi.org/10.1111/jsr.12307 (2015).
Kim, H. I. et al. Association of CLOCK, ARNTL, and NPAS2 gene polymorphisms and seasonal variations in mood and behavior. Chronobiol. Int. 32, 785–791. https://doi.org/10.3109/07420528.2015.1049613 (2015).
Liberman, A. R., Halitjaha, L., Ay, A. & Ingram, K. K. Modeling strengthens molecular link between circadian polymorphisms and major mood disorders. J. Biol. Rhythms 33, 318–336. https://doi.org/10.1177/0748730418764540 (2018).
Nguyen, C., Murray, G., Anderson, S., Filipowicz, A. & Ingram, K. K. In vivo molecular chronotyping, circadian misalignment, and high rates of depression in young adults. J. Affect. Disord. 250, 425–431. https://doi.org/10.1016/j.jad.2019.03.050 (2019).
Soria, V. et al. Differential association of circadian genes with mood disorders: CRY1 and NPAS2 are associated with unipolar major depression and CLOCK and VIP with bipolar disorder. Neuropsychopharmacology 35, 1279–1289. https://doi.org/10.1038/npp.2009.230 (2010).
Chaput, J. P. et al. Sleep timing, sleep consistency, and health in adults: A systematic review. Appl. Physiol. Nutr. Metab 45, s232–s247. https://doi.org/10.1139/apnm-2020-0032 (2020).
Henderson, S. E. M., Brady, E. M. & Robertson, N. Associations between social jetlag and mental health in young people: A systematic review. Chronobiol. Int. 36, 1316–1333. https://doi.org/10.1080/07420528.2019.1636813 (2019).
Islam, Z. et al. Social jetlag is associated with an increased likelihood of having depressive symptoms among the Japanese working population: The Furukawa Nutrition and Health Study. Sleep https://doi.org/10.1093/sleep/zsz204 (2020).
Levandovski, R. et al. Depression scores associate with chronotype and social jetlag in a rural population. Chronobiol. Int. 28, 771–778. https://doi.org/10.3109/07420528.2011.602445 (2011).
Murray, J. M. et al. Prevalence of circadian misalignment and its association with depressive symptoms in delayed sleep phase disorder. Sleep https://doi.org/10.1093/sleep/zsw002 (2017).
Slavish, D. C., Taylor, D. J. & Lichstein, K. L. Intraindividual variability in sleep and comorbid medical and mental health conditions. Sleep https://doi.org/10.1093/sleep/zsz052 (2019).
Taylor, B. J. & Hasler, B. P. Chronotype and mental health: Recent advances. Curr. Psychiatry Rep. 20, 59. https://doi.org/10.1007/s11920-018-0925-8 (2018).
Dashti, H. S. et al. Genome-wide association study identifies genetic loci for self-reported habitual sleep duration supported by accelerometer-derived estimates. Nat. Commun. 10(1), 1100. https://doi.org/10.1038/s41467-019-08917-4 (2019).
Enoch, M. A. et al. Association between seasonal affective disorder and the 5-HT2A promoter polymorphism, -1438G/A. Mol. Psychiatry 4, 89–92. https://doi.org/10.1038/sj.mp.4000439 (1999).
Jones, S. E. et al. Genome-wide association analyses of chronotype in 697,828 individuals provides insights into circadian rhythms. Nat. Commun. 10(1), 343. https://doi.org/10.1038/s41467-018-08259-7 (2019).
Overton, R., Zafar, A., Attia, Z., Ay, A. & Ingram, K. K. Machine learning analyses reveal circadian features predictive of risk for sleep disturbance. Nat. Sci. Sleep 14, 1887–1900. https://doi.org/10.2147/nss.S379888 (2022).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681. https://doi.org/10.1038/s41588-018-0090-3 (2018).
Meier, S. M. et al. Genetic variants associated with anxiety and stress-related disorders: A genome-wide association study and mouse-model study. JAMA Psychiatry 76(9), 924–932. https://doi.org/10.1001/jamapsychiatry.2019.1119 (2019).
van der Walt, K. et al. Systematic review of genome-wide association studies of anxiety disorders and neuroticism. World J. Biol. Psychiatry 24(4), 280–291. https://doi.org/10.1080/15622975.2022.2099970 (2023).
McClung, C. A. How might circadian rhythms control mood? Let me count the ways. Biol. Psychiatry 74, 242–249. https://doi.org/10.1016/j.biopsych.2013.02.019 (2013).
Savalli, G. et al. Anhedonic behavior in cryptochrome 2-deficient mice is paralleled by altered diurnal patterns of amygdala gene expression. Amino Acids 47, 1367–1377. https://doi.org/10.1007/s00726-015-1968-3 (2015).
Zhang, L. et al. A PERIOD3 variant causes a circadian phenotype and is associated with a seasonal mood trait. Proc. Natl. Acad. Sci. 113, E1536–E1544. https://doi.org/10.1073/pnas.1600039113 (2016).
Ozburn, A. R. et al. NPAS2 regulation of anxiety-like behavior and GABAA receptors. Front. Mol. Neurosci. https://doi.org/10.3389/fnmol.2017.00360 (2017).
Ho, K. W. D. et al. Genome-wide association study of seasonal affective disorder. Transl. Psychiatry 8, 190. https://doi.org/10.1038/s41398-018-0246-z (2018).
Davies, M. N. et al. Hypermethylation in the ZBTB20 gene is associated with major depressive disorder. Genome Biol. 15, R56. https://doi.org/10.1186/gb-2014-15-4-r56 (2014).
Trehearne, A. Genetics, lifestyle and environment. UK Biobank is an open access resource following the lives of 500,000 participants to improve the health of future generations. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz 59, 361–367. https://doi.org/10.1007/s00103-015-2297-0 (2016).
Sudlow, C. et al. UK biobank: An open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779. https://doi.org/10.1371/journal.pmed.1001779 (2015).
Brandes, N., Linial, N. & Linial, M. PWAS: Proteome-wide association study—Linking genes and phenotypes by functional variation in proteins. Genome Biol. 21, 173. https://doi.org/10.1186/s13059-020-02089-x (2020).
Spitzer, R. L., Kroenke, K., Williams, J. B. W. & Löwe, B. A brief measure for assessing generalized anxiety disorder: The GAD-7. Arch. Intern. Med. 166, 1092–1097. https://doi.org/10.1001/archinte.166.10.1092 (2006).
Marchini, J. UK Biobank Phasing and Imputation Documentation. 1–13 (2015).
Genotyping and quality control of UK Biobank, a large-scale, extensively phenotyped prospective resource. 1–21 (2015).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283. https://doi.org/10.1038/ng.3643 (2016).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65. https://doi.org/10.1038/nature11632 (2012).
Walter, K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90. https://doi.org/10.1038/nature14962 (2015).
Johnson, S. U., Ulvenes, P. G., Øktedalen, T. & Hoffart, A. Psychometric properties of the general anxiety disorder 7-item (GAD-7) scale in a heterogeneous psychiatric sample. Front. Psychol. 10, 1713. https://doi.org/10.3389/fpsyg.2019.01713 (2019).
Plummer, F., Manea, L., Trepel, D. & McMillan, D. Screening for anxiety disorders with the GAD-7 and GAD-2: A systematic review and diagnostic metaanalysis. Gener. Hosp. Psychiatry 39, 24–31. https://doi.org/10.1016/j.genhosppsych.2015.11.005 (2016).
Senan, E. M., Abunadi, I., Jadhav, M. E. & Fati, S. M. Score and correlation coefficient-based feature selection for predicting heart failure diagnosis by using machine learning algorithms. Comput. Math. Methods Med. 2021, 8500314. https://doi.org/10.1155/2021/8500314 (2021).
Fan, R. et al. Entropy-based information gain approaches to detect and to characterize gene–gene and gene-environment interactions/correlations of complex diseases. Genet. Epidemiol. 35, 706–721. https://doi.org/10.1002/gepi.20621 (2011).
Quinlan, J. R. Induction of decision trees. Mach. Learn. 1, 81–106. https://doi.org/10.1007/BF00116251 (1986).
Robnik-Sikonja, M. & Kononenko, I. An adaptation of Relief for attribute estimation in regression. ICML '97: Proceedings of the Fourteenth International Conference on Machine Learning (2000).
Yang, H. H. & Moody, J. E. Feature Selection Based on Joint Mutual Information. 1999.
Ding, C. & Peng, H. in Computational Systems Bioinformatics. CSB2003. Proceedings of the 2003 IEEE Bioinformatics Conference. CSB2003. 523–528.
R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna, Austria, 2021).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Akinwande, M. O., Dikko, H. G. & Samson, A. Variance inflation factor: As a condition for the inclusion of suppressor variable(s) in regression analysis. Open J. Stat. 05, 754–767 (2015).
Bozdogan, H. Model selection and Akaike’s information criterion (AIC): The general theory and its analytical extensions. Psychometrika 52, 345–370. https://doi.org/10.1007/BF02294361 (1987).
Seabold, S. & Perktold, J. Statsmodels: Econometric and Statistical modeling with Python. Proceedings of the 9th Python in Science Conference 2010 (2010).
Durbin, J. & Watson, G. S. Testing for serial correlation in least squares regression. II. Biometrika 38, 159–177. https://doi.org/10.2307/2332325 (1951).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 57, 289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x (1995).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples). Biometrika 52, 591–611 (1965).
Sokal, R. & Rohlf, F. Biometry: the principles and practice of statistics in biological research/Robert R. Sokal and F. James Rohlf. SERBIULA (sistema Librum 2.0) (2013).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. Mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38. https://doi.org/10.18637/jss.v059.i05 (2014).
Nielsen, J. V., Blom, J. B., Noraberg, J. & Jensen, N. A. Zbtb20-induced CA1 pyramidal neuron development and area enlargement in the cerebral midline cortex of mice. Cereb. Cortex 20, 1904–1914. https://doi.org/10.1093/cercor/bhp261 (2009).
Nielsen, J. V., Thomassen, M., Møllgård, K., Noraberg, J. & Jensen, N. A. Zbtb20 defines a hippocampal neuronal identity through direct repression of genes that control projection neuron development in the isocortex. Cereb. Cortex 24, 1216–1229. https://doi.org/10.1093/cercor/bhs400 (2014).
Qu, Z. et al. Loss of ZBTB20 impairs circadian output and leads to unimodal behavioral rhythms. Elife https://doi.org/10.7554/eLife.17171 (2016).
Fiedorowicz, J. G., Coryell, W. H., Akhter, A. & Ellingrod, V. L. Chryptochrome 2 variants, chronicity, and seasonality of mood disorders. Psychiatr. Genet. 22, 305–306. https://doi.org/10.1097/YPG.0b013e3283539594 (2012).
Shi, S. Q. et al. Molecular analyses of circadian gene variants reveal sex-dependent links between depression and clocks. Transl. Psychiatry 6, e748. https://doi.org/10.1038/tp.2016.9 (2016).
Hida, A. et al. Screening of clock gene polymorphisms demonstrates association of a PER3 polymorphism with morningness–eveningness preference and circadian rhythm sleep disorder. Sci. Rep. 4, 6309. https://doi.org/10.1038/srep06309 (2014).
Chen, C.-Y. et al. Effects of aging on circadian patterns of gene expression in the human prefrontal cortex. Proc. Natl. Acad. Sci. 113, 206–211. https://doi.org/10.1073/pnas.1508249112 (2016).
Katzenberg, D. et al. A CLOCK polymorphism associated with human diurnal preference. Sleep 21, 569–576. https://doi.org/10.1093/sleep/21.6.569 (1998).
Hughes, S., Jagannath, A., Hankins, M. W., Foster, R. G. & Peirson, S. N. Photic regulation of clock systems. Methods Enzymol. 552, 125–143. https://doi.org/10.1016/bs.mie.2014.10.018 (2015).
Reischl, S. et al. Beta-TrCP1-mediated degradation of PERIOD2 is essential for circadian dynamics. J. Biol. Rhythms 22, 375–386. https://doi.org/10.1177/0748730407303926 (2007).
Hua, P. et al. Cry1 and Tef gene polymorphisms are associated with major depressive disorder in the Chinese population. J. Affect. Disord. 157, 100–103. https://doi.org/10.1016/j.jad.2013.11.019 (2014).
Halbreich, U. & Lumley, L. A. The multiple interactional biological processes that might lead to depression and gender differences in its appearance. J. Affect. Disord. 29, 159–173. https://doi.org/10.1016/0165-0327(93)90030-n (1993).
Zunszain, P. A., Anacker, C., Cattaneo, A., Carvalho, L. A. & Pariante, C. M. Glucocorticoids, cytokines and brain abnormalities in depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 35, 722–729. https://doi.org/10.1016/j.pnpbp.2010.04.011 (2011).
Landgraf, D., McCarthy, M. J. & Welsh, D. K. Circadian clock and stress interactions in the molecular biology of psychiatric disorders. Curr. Psychiatry Rep. 16, 483. https://doi.org/10.1007/s11920-014-0483-7 (2014).
Daut, R. A. & Fonken, L. K. Circadian regulation of depression: A role for serotonin. Front. Neuroendocrinol. 54, 100746. https://doi.org/10.1016/j.yfrne.2019.04.003 (2019).
Gressier, F., Calati, R. & Serretti, A. 5-HTTLPR and gender differences in affective disorders: A systematic review. J. Affect. Disord. 190, 193–207. https://doi.org/10.1016/j.jad.2015.09.027 (2016).
Songtachalert, T., Roomruangwong, C., Carvalho, A. F., Bourin, M. & Maes, M. Anxiety disorders: Sex differences in serotonin and tryptophan metabolism. Curr. Top. Med. Chem. 18, 1704–1715. https://doi.org/10.2174/1568026618666181115093136 (2018).
Malek, Z. S., Dardente, H., Pevet, P. & Raison, S. Tissue-specific expression of tryptophan hydroxylase mRNAs in the rat midbrain: Anatomical evidence and daily profiles. Eur. J. Neurosci. 22, 895–901. https://doi.org/10.1111/j.1460-9568.2005.04264.x (2005).
Malek, Z. S., Sage, D., Pévet, P. & Raison, S. Daily rhythm of tryptophan hydroxylase-2 messenger ribonucleic acid within raphe neurons is induced by corticoid daily surge and modulated by enhanced locomotor activity. Endocrinology 148, 5165–5172. https://doi.org/10.1210/en.2007-0526 (2007).
Lowry, C. A. et al. Serotonergic systems, anxiety, and affective disorder: Focus on the dorsomedial part of the dorsal raphe nucleus. Ann. N. Y. Acad. Sci. 1148, 86–94. https://doi.org/10.1196/annals.1410.004 (2008).
Benedetti, F. et al. Actimetric evidence that CLOCK 3111 T/C SNP influences sleep and activity patterns in patients affected by bipolar depression. Am. J. Med. Genet. B Neuropsychiatr. Genet. 144b, 631–635. https://doi.org/10.1002/ajmg.b.30475 (2007).
Benedetti, F. et al. Influence of CLOCK gene polymorphism on circadian mood fluctuation and illness recurrence in bipolar depression. Am. J. Med. Genet. B Neuropsychiatr. Genet. 123b, 23–26. https://doi.org/10.1002/ajmg.b.20038 (2003).
Albert, P. R., Vahid-Ansari, F. & Luckhart, C. Serotonin-prefrontal cortical circuitry in anxiety and depression phenotypes: Pivotal role of pre- and post-synaptic 5-HT1A receptor expression. Front. Behav. Neurosci. 8, 199. https://doi.org/10.3389/fnbeh.2014.00199 (2014).
Zhou, Y. S. et al. Regular aerobic exercise attenuates pain and anxiety in mice by restoring serotonin-modulated synaptic plasticity in the anterior cingulate cortex. Med. Sci. Sports Exerc. 54, 566–581. https://doi.org/10.1249/mss.0000000000002841 (2022).
Haase, J. et al. Sex and brain region-specific regulation of serotonin transporter activity in synaptosomes in guanine nucleotide-binding protein G(q) alpha knockout mice. J. Neurochem. 159, 156–171. https://doi.org/10.1111/jnc.15482 (2021).
Jones, M. D. & Lucki, I. Sex differences in the regulation of serotonergic transmission and behavior in 5-HT receptor knockout mice. Neuropsychopharmacology 30, 1039–1047. https://doi.org/10.1038/sj.npp.1300664 (2005).
Silva, V. M., Magalhaes, J. E. M. & Duarte, L. L. Quality of sleep and anxiety are related to circadian preference in university students. PLoS ONE 15, e0238514. https://doi.org/10.1371/journal.pone.0238514 (2020).
Dardente, H., Fortier, E. E., Martineau, V. & Cermakian, N. Cryptochromes impair phosphorylation of transcriptional activators in the clock: A general mechanism for circadian repression. Biochem. J. 402, 525–536. https://doi.org/10.1042/bj20060827 (2007).
Hsu, D. S. et al. Putative human blue-light photoreceptors hCRY1 and hCRY2 are flavoproteins. Biochemistry 35, 13871–13877. https://doi.org/10.1021/bi962209o (1996).
Allen, N. et al. UK Biobank: Current status and what it means for epidemiology. Health Policy Technol. 1, 123–126. https://doi.org/10.1016/j.hlpt.2012.07.003 (2012).
Manolio, T. A. & Collins, R. Enhancing the feasibility of large cohort studies. Jama 304, 2290–2291. https://doi.org/10.1001/jama.2010.1686 (2010).
Fry, A. et al. Comparison of sociodemographic and health-related characteristics of UK biobank participants with those of the general population. Am. J. Epidemiol. 186, 1026–1034. https://doi.org/10.1093/aje/kwx246 (2017).
Author information
Authors and Affiliations
Contributions
C.V., W.M., A.A., and K.I. conceived the analysis. C.V. and W.M. performed the statistical and machine learning analysis. C.V., W.R., A.A., and K.I. interpreted the results. W.R. wrote the original manuscript text. C.V. and W.R. created figures and tables. W.R., A.A., and K.I. revised the manuscript text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ventresca, C., Mohamed, W., Russel, W.A. et al. Machine learning analyses reveal circadian clock features predictive of anxiety among UK biobank participants. Sci Rep 13, 22304 (2023). https://doi.org/10.1038/s41598-023-49644-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-49644-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.