Abstract
Gender identity refers to the consciousness of being a man, a woman or other condition. Although it is generally congruent with the sex assigned at birth, for some people it is not. If the incongruity is distressing, it is defined as gender dysphoria (GD). Here, we measured whole-genome DNA methylation by the Illumina © Infinium Human Methylation 850k array and reported its correlation with cortical thickness (CTh) in 22 transgender men (TM) experiencing GD versus 25 cisgender men (CM) and 28 cisgender women (CW). With respect to the methylation analysis, TM vs. CW showed significant differences in 35 CpGs, while 2155 CpGs were found when TM vs. CM were compared. With respect to correlation analysis, TM showed differences in methylation of CBLL1 and DLG1 genes that correlated with global and left hemisphere CTh. Both genes were hypomethylated in TM compared to the cisgender groups. Early onset TM showed a positive correlation between CBLL1 and several cortical regions in the frontal (left caudal middle frontal), temporal (right inferior temporal, left fusiform) and parietal cortices (left supramarginal and right paracentral). This is the first study relating CBLL1 methylation with CTh in transgender persons and supports a neurodevelopmental hypothesis of gender identity.
Similar content being viewed by others
Introduction
Gender identity is the consciousness of being male, female, or another gender identity. This inner sense may or may not align with a person’s sex as assigned at birth or to a person’s primary or secondary sex characteristics1. Transgender persons can fall within the binary male/female system [transgender men or transgender women (TM, TW, respectively)] or as non-binary (genderqueer, gender nonconforming, gender neutral, etc.). Gender identity emerges in the individual because of a constellation of interacting genetic, epigenetic, hormonal, and environmental variables. Some transgender people may experience gender incongruence2 and gender dysphoria (GD)1 and look for hormonal, and sometimes surgical, gender affirmation treatment.
Since the pioneering work by Swaab3 there has been an increase in the exploration of the role of biological variables in gender identity. Structural and functional MRI brain studies show the existence of brain endophenotypes associated to male and female cisgender individuals (CM and CW), people who show congruence between their gender identity and sex assigned at birth , and to transgender persons. Endophenotypes have been reported with respect to cortical thickness (CTh)4 and cortical surface area5. Moreover, white matter microstructure also shows differences among these four binary groups6,7,8. So, based on the sexual differentiation of the brain, a neurodevelopmental cortical hypothesis was suggested to explain the emergence of these endophenotypes9. This hypothesis has found further support by recent functional stationary10 and dynamic11 fMRI studies, as well as whole-brain dynamic approaches12, which also documented brain endophenotypes associated with CM and CW and binary transgender people.
Although cortical surface area is a sensitive measure to distinguish brain endophenotypes between cisgender and transgender populations5, CTh measure is less heritable than surface area13,14,15, more dynamic through the lifespan and suitable to search the genetic-environmental interplay, as previously reviewed in clinical and non-clinical populations16. Cerebral CTh, which reflects the number of cortical cells, their physiological state, and myelination, depends on age and sex. There is a monotonic thinning of the cortex from early childhood until senescence, but the trajectories of this process are different in different structures17 and CTh increases as well as decreases have been reported around puberty18, 19. Sex differences in CTh have been reported at all ages20, 21. Moreover, the thinning process is related to the efficiency of the androgen receptor22. Age- and sex-related changes in CTh suggest a neurodevelopmental process controlled by genetic, epigenetic, and hormonal variables.
The sexual differentiation of the brain depends on genetic and hormonal mechanisms as well as on epigenetic processes. In fact, sex chromosomes, gonadal hormones, and epigenetic mechanisms (DNA and RNA methylation) intertwine to shape brain development and influence behavior in mammals23. A growing number of studies demonstrate that there are sex differences in the brain epigenome and that brain sex differentiation requires the interaction between sex hormones and different epigenetic mechanisms24, 25.
Genetic studies in transgender people are only recently gaining attraction. For example, such studies have found an association between the four binary gender identities and polymorphisms in androgen and estrogen steroid receptors and enzymes implicated in the sexual differentiation of the brain26,27,28,29,30. Moreover, steroid receptor coactivators (SRC1 and SRC2) of estrogen receptors are influential in the masculinization of the brain31 and SRC1 and SRC2 polymorphisms were recently associated with GD32.
Epigenetic studies disentangle the relationships between genes and the influence of the internal and external environment on gene expression. In mammals DNA methylation, as well as the methylation of RNA messenger (m6A mRNA), play an important role in brain sexual differentiation and behavior23, 25, 33. To date, only two studies of whole-genome DNA methylation have been performed in a transgender population, before and during gender affirming hormone treatment (GAHT). These studies demonstrated the existence of characteristic epigenetic marks in the transgender population even before GAHT34 as well as the existence of specific epigenetic marks due to GAHT after 6 and 12 months of treatment35. Furthermore, methylation analysis of the regulatory region of ESR1 demonstrated sex differences in the methylation degree and that both TW and TM had significantly lower levels of methylation in this region compared to cisgender men prior to receiving GAHT36, 37.
In the present study, we assessed the relationship between DNA methylation and brain CTh in transgender and cisgender people. To this aim, we performed an epigenome-wide association analysis (DNA methylation) and correlated the findings with CTh in transgender men experiencing GD before GAHT and compared these with cisgender men and women. Because of the cortical neurodevelopmental hypothesis9, we anticipated that genes related to development would emerge and correlate with brain cortical regions.
Results
Demographic data
Data on age, ethnicity and smoking are presented in Table 1.
Global methylation analysis
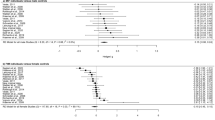
Before correlation analysis, a differential methylation analysis was performed from our 75 individuals (22 TM, 25 CM, and 28 CW) where a pair-wise comparison between individual groups (CW-TM, CM-TM, and CW-CM) was made to obtain significant CpG sites (p-value < 0.05). Thus, 35 CpG sites were significant for CW-TM comparison (Fig. 1), 2155 for CM-TM (Fig. 2) and 5684 for CM-CW (Fig. 3) of a total of 813199 CpG sites tested. These genes were involved in numerous biological processes relating to neuroactive ligand-receptor interaction, gap junction, glutamatergic synapse, serotonergic synapse, GABAergic synapse, inflammatory mediator regulation, among others (Supplemental Material Tables S1, S2 and S3).
Boxplot representation of the top four most relevant differentially methylated CpG sites in the cisgender women (CW)—transgender men (TM) comparison. Methylation β-values were represented for the top four differentially methylated CpG sites in the CW-TM comparison. Vertical lines in each box represent β-values standard deviation between individuals of the same group, whereas horizontal lines correspond to average β-values of the groups for the probe. Outliers mean individuals whose β-values are extreme in comparison to the average. The samples are coloured according to the variable “group” (purple for cisgender men, green for cisgender women and yellow for transgender men).
Boxplot representation of the top four most relevant differentially methylated CpG sites in the cisgender men (CM)—transgender men (TM) comparison. Methylation β-values were represented for the top four differentially methylated CpG sites in the CM-TM comparison. Vertical lines in each box represent β-values standard deviation between individuals of the same group, whereas horizontal lines correspond to average β-values of the groups for the probe. Outliers mean individuals whose β-values are extreme in comparison to the average. The samples are coloured according to the variable “group” (purple for cisgender men, green for cisgender women and yellow for transgender men).
Boxplot representation of the top four most relevant differentially methylated CpG sites in the cisgender men (CM)—cisgender women (CW) comparison. Methylation β-values were represented for the top four differentially methylated CpG sites in the CM-CW comparison. Vertical lines in each box represents β-values standard deviation between individuals of the same group, whereas horizontal lines correspond to average β-values of the groups for the probe. Outliers mean individuals whose β-values are extreme in comparison to the average. The samples are coloured according to the variable “group” (purple for cisgender men, green for cisgender women and yellow for transgender men).
Due to the existence of different significant CpG sites, we included variables of cortical regions in our methylation study to make a correlation analysis between these characteristics and patrons of methylation (Fig. 4).
Principal components analysis (PCA) showing methylation profiles of the study samples. Each sample is represented by a dot, the axes are the first two PCs, the percentages indicate the fraction of variance explained by each PC. The number at the top (32.3%) is the variance explained by the first two PCs. The samples are coloured according to the variable “group” (blue for cisgender men, red for cisgender women and yellow for transgender men).
MRI data: CTh differences
Regionally statistically significant results emerged in some cortical regions (see Supplemental Tables S4 and S5). Cisgender men showed a thicker cortex with respect to CW in bilateral fusiform areas, the left medial orbitofrontal and pars opercularis regions as well as the right middle and transverse temporal regions. However, none of these areas survived Bonferroni correction. Small to medium effect sizes were present in all these analyses.
Regarding early or late onset of GD in TM individuals, defined as an onset of GD that occurs during childhood or pubertal/adult period respectively, regional analyses pointed out the following (see Supplemental Tables S6 and S7): (1) CM compared with CW had a thicker cortex in the left medial orbitofrontal and pars opercularis region as well as in the right fusiform area; (2) CM showed higher CTh values in comparison with early onset TM in the left pars opercularis and postcentral areas; (3) late onset when compared to early onset TM had a thicker cortex in the left pars triangularis and postcentral regions and right superior temporal and transverse temporal regions and also in the latter region in comparison with CW. All the above-mentioned results were statistically significant (p < 0.05), although none remained significant after Bonferroni correction.
Methylation & MRI data
The ANOVA test comparing methylation by groups and by CTh (CM vs. CW, CM vs. TM, and CW vs. TM) showed statistically significant differences in two genes: CBLL1 and DLG1 (Table 2; Fig. 5). According to the Enrichment Score study, these genes are involved in neuronal myelination, CBLL1 in the CNS and DLG1 in the peripheral nervous system. Specifically, CBLL1 is involved in messenger RNA methylation (see Supplemental Material S5).
Dotplot showing M-value data for the CBLL1 gene. Each sample is represented by a dot, which corresponds to the overall degree of methylation (M-value data). The samples are colored according to the levels of the variable “group”: blue for cisgender men, red for cisgender women and yellow for transgender men. The middle line is the median; the box represents the upper and the lower quartile, while the whiskers correspond to the 90th and 10th percentiles of the data.
ANOVA test results: CBLL1
With respect to the CBLL1 gene, statistically significant differences after FDR correction were found in total CTh and in the left hemisphere, but not in the right hemisphere.
Total CTh
In the CBLL1 gene, the methylation degree was higher in cisgender men than in cisgender women (CM > CW) (p < 0.001) (Table 2). Similarly, the difference in CBLL1 methylation was also significantly higher in CM than in TM (CM > TM) (p < 0.001) (Table 2).
However, a difference also emerged between CW and TM indicating a higher methylation degree in CW than in TM (CW > TM) (p < 0.001) (Table 2). Thus, the CBLL1 methylation degree followed the sequence CM > CW > TM. The highest methylation degree was found in CM, while TM showed the lowest. Therefore, this gene was hypermethylated in cisgender men and women relative to transgender men, where it was hypomethylated (Table 2).
Left hemisphere CTh
The pattern in the left hemisphere reflected that of the total CTh, namely, a statistically significant overall effect (p < 0.001) (Table 2) following the same sequence: CW > CM > TM. However, while the comparisons between CM vs.TM (p < 0.001) and CW vs. TM (p < 0.001) were statistically significant, the differences between the two cisgender groups (CM vs. CW; p = 0.80) was not (Table 2).
ANOVA test results: DLG1
Left hemisphere CTh
After FDR correction, the DLG1 gene showed a slightly different pattern. First, statistically significant differences were only found in the left hemisphere. The difference in methylation was statistically significant (p = 0.003) when we compared CM to CW, with a higher methylation degree in CW (Table 2). When we compared CM to TM, the difference was also significant (p = 0.01), with a higher methylation degree in CM than in TM (Table 2). Finally, the difference was also significant (p < 0.001) when we compared CW to TM. The methylation degree was higher in CW than in TM. Therefore, this gene was hypermethylated in the cis population (CM and CW) and hypomethylated in TM, but with higher methylation in CW than CM (CW > CM > TM).
Correlation between regional CTh and Methylation data
Neither cisgender participants nor late onset TM showed a statistically significant correlation that survived Bonferroni correction (p ≤ 0.001) between CTh regions and methylation data. The early onset TM group showed a positive association that survived Bonferroni correction (p ≤ 0.001) between CBLL1 methylation and left caudal middle frontal (r = 0.84) fusiform (r = 0.87) and supramarginal (r = 0.89), as well as with the right inferior temporal (r = 0.90) and paracentral regions (r = 0.87). No statistically significant results were found in respect to the DLG1 gene.
Discussion
This study is the first, to our knowledge, to demonstrate associations between wide DNA methylation and brain structural measures in transgender people. Two main findings emerged. First, TM showed differences in the methylation degree in two genes, CBLL1 and DLG1 that correlated with global and left hemisphere CTh. Both genes were hypomethylated in TM with respect to CW and CM. Second, early onset TM showed a positive correlation between CBLL1 methylation and several CTh regions. Methylation of CBLL1 correlated positively with regional CTh in some regions of the frontal (left caudal middle frontal), temporal (right inferior temporal and left fusiform) and parietal cortices (left supramarginal and right paracentral). As was predicted, a gene related with cortical development, CBLL1, showed differences in the methylation degree between cisgender and transgender groups.
To study the cortical development of the brain, both volume and CTh values are commonly measured and reported by MRI studies. Regarding the latter, CTh is an extremely dynamic measurement that presents changes across the lifespan19. CTh increases in almost all brain areas from the neonatal period to childhood38, whereas there is a progressive cortical thinning during adolescence19. With respect to GD, a variety of results have been reported in TM before GAHT. On the one hand, some studies have found a thicker cortex in TM with respect to cisgender populations4, 39 while a mega-analysis did not find statistically significant differences in TM5.
Evaluation of CBLL1 methylation showed differences between groups in respect to total and left hemisphere CTh, being hypomethylated in TM respect to cisgender groups. There are reports in the literature that relate CBLL1 with cortical development through complex mechanisms that involve, among others, estrogen receptors. These receptors play a key function in the sexual differentiation of the brain of mammals.
CBLL1, which encodes the HAKAI protein, is characterized as an E-cadherin-binding protein that is predominantly expressed in the brain40 and has regulatory actions on the alfa estrogen receptor (ERα). It is known that HAKAI modulates ERα transactivation through competitive interference with the coactivators SRC1 and SRC2 by binding to ERα41, thereby regulating the expression of ERα target genes. Interestingly, in previous studies, we have found that some polymorphisms in the ESR1 gene, and in SRC1 and SCR2 coactivators, were associated with gender variants32.
Moreover, HAKAI have other mechanisms to influence brain development and differentiation through mRNA methylation. This protein is a conserved component of the N6-methyladenosine (m6A) transferase complex for mRNA methylation. This modification plays a central role in almost every aspect of mRNA metabolism and is essential for several biological processes such as myelination, cell differentiation, DNA repair, neurogenesis, sex determination, and silencing the second X chromosome in females42,43,44. Even more, the functioning of the m6A molecular machinery in the nervous system is essential for the life of mammals since m6A marks the "tempo" of corticogenesis45 and is also implicated in oligodendrocyte plasticity during adulthood46.
The three binary variants also showed differences in DLG1 methylation in relation to the CTh of the left hemisphere. DLG1 gene is fully involved in brain development intervening in cellular intimate processes and myelination. Respect to the later, we should underscore that sex differences were reported respect to white matter microstructure in our species44, 48.
The DLG1 gene, also called SAP97, is a scaffolding protein that participates in the control of key cellular processes such as polarity, proliferation, and migration as well as in the trafficking of membrane proteins49, 50. This gene is widely expressed throughout the brain, concentrating at the postsynaptic space in the cerebral cortex51. In addition, SAP97 is widely distributed in excitatory synapses, where it is strongly involved in the trafficking and localization of NMDA and AMPA-type glutamate receptors, regulating synaptic plasticity52. Cotter et al.53 showed that Dlg1 acts as an inhibitor of Schwann cell myelination, thus Dlg1 silencing during mouse SNP myelination generates a hypermyelination phenotype. In contrast, Dlg1 overexpression reduces myelination53. In Schwann cells, Dlg1 interacts with PTEN (phosphatase and tensin homolog deleted on chromosome 10)54 to inhibit stimulation of axonal myelination. This mechanism limits the thickness of the myelin sheath and prevents over myelination.
Early and late onset comparison can provide a view of gender identity development and the organizational and activational influences of sex hormones. Puberty marks the early or the late onset GD classification. Early GD emerges before puberty while late GD onset occurs during or after puberty55, 56.
The age of onset of GD is a variable of interest unaddressed in MRI studies. However, it was recently reported that age of onset of GD plays a role in the modulation of cerebral metabolite concentrations in TM57. In the present study, we found that those TM with early onset GD had a thinner cortex compared to late onset TM and CM. However, these differences did not maintain statistical significance after the Bonferroni correction and so may be due to the small sample size analysed. Nevertheless, early onset TM showed a positive relationship between CBLL1 methylation and CTh in several cortical regions. A possible explanation is that this gene modulates its activity in early onset GD TM to achieve optimal CTh values in some specific regions of the brain. For example, the supramarginal region, one of the cortical regions in which CTh positively correlates with CBLL1 methylation only in early onset TM participants, is a crucial region for GD in TM. Resting state functional MRI studies have involved the parietal lobe in own body self-referential processing58 and it is a key piece to understand TM stationary and dynamic functional connectivity10, 11. As we saw above, CBLL1 functions are mainly developmental, our findings support two different developmental pathways for early and late GD onset TM and signal the possible implication of specific regions of the cortex in these differences. Our research design precludes to determine if either internal (i.e., hormones), or external (familial/social) variables or their interaction influence CBLL1 differential methylation at different stages of brain development in early and late onset transgender men.
Some limitations of the current study were the total sample size and the sample size of each subgroup (early vs. late onset). The study we have carried out aimed to obtain very homogeneous groups, so some individuals have been eliminated, which has resulted in even smaller groups. A major challenge in the study of DNA methylation is the study of a non-stratified population59 that can have a significant impact on the results, as they can lead to false positives or false negatives. In our study, a comprehensive sample selection based on ethnicity and geographic origin was applied, but individuals with drug abuse or associated psychiatric illnesses from the study were removed. This has resulted in a homogenous population, eliminating or reducing the effects of population stratification, but at the same time has resulted in a smaller population. Besides, our results only can apply to binary cisgender heterosexuals and transgender men diagnosed with GD and sexually attracted to women. Moreover, finding an association between DNA methylation and brain structure does not prove that DNA methylation markers imply a cause or consequence of brain findings or are due to other confounding variables16.
However, the main strength of our work is that CBLL1 and DLG1 have been identified as candidate genes in TM due to its relationship with CTh. Moreover, early, and late onset TM were epigenetically and cortically differentiated. These robust differences remain after rigorous standards were applied with a very conservative statistical correction (i.e. Bonferroni and FDR corrections).
Concluding remarks
What do we know with respect to the variables that could intervene in brain development of binary cis and transgender persons. At the genetic level we have shown that either the alfa or the beta estrogen receptor polymorphisms28, 29 as well as SCR1 and SCR2 polymorphisms are related to TM32. Moreover, epigenetic analyses show that methylation of the promoter region of the alpha estrogen receptor is also associated with TM37. It seems, at least in our studies, that any genetic or epigenetic changes that could affect normative functioning of transcription by estrogens is associated with TM. The present findings with respect to CBLL1 support SCR1 and SCR2 implication in TM and envision the possible participation of mechanisms implicating mRNA methylation in the regulation of the thickness of the cortex in our species.
Finally, our findings suggest that CBLL1 and DLG1 may participate in the modulation of CTh in humans and they distinguish three cortical endophenotypes related to gender identity (i.e., CM, CF, TM). Moreover, they also suggest epigenetic differences related to the cortex between early and late GD onset TM. All these results support a cortical hypothesis that suggests that different rates of development, in specific cortical regions, could underlie gender identity and its variants 9, 38. Future studies should include larger and homogeneous transgender samples with different onsets of GD. All these findings support a neurodevelopmental hypothesis9 to explain the development of the binary male and female gender identities and point out they have a biological counterpart.
Methods
Participants’ characteristics
One hundred and twelve individuals, belonging to a larger study, participated in the current investigation. For this study, the inclusion criteria for transgender people were: (1) being aged 17 years or older; (2) the presence of GD according to DSM-5; (3) identification with the gender other than the one assigned at birth (male or female), and (4) having no prior history of hormonal treatment. The exclusion criteria for cisgender and transgender populations were: (a) current or past drug consumption; (b) the presence of psychiatric, neurological and/or hormonal disorders, or a major medical condition as assessed by anamnesis and (c) history of cranial contusion or injury. In addition, the Mini-International Neuropsychiatric Interview60 was carried out by a clinical psychologist, assessing possible undiagnosed psychiatric symptoms in participants. During their first clinic visit to enrol for GAHT, self-reported sexual orientation was solicited and only people attracted to the opposite gender relative to their gender identity were included in the study. TM also self-reported the age of GD onset55, the stage of life they have experienced gender incongruent feelings (i.e., childhood, pubertal age, and adult life). Early onset GD corresponds to the childhood period and late onset GD to pubertal or adult life55.
From the initial 112 participants 37 were excluded after the inclusion/exclusion criteria were applied for the following reasons: 32 individuals did not have an MRI study; 4 reported drug consumption and 1 had no sociodemographic data. Therefore, 22 TM (mean age: 29.32 ± 11.52) before GAHT and 53 cisgender individuals [25 CM (mean age: 26.52 ± 5.86) and 28 CW (mean age: 31.61 ± 9.08)] were recruited at the Center for Sexology and Gender, Department of Endocrinology at Ghent University Hospital (Belgium). Cisgender individuals were enrolled via media advertising and by word of mouth. All participants in the study received 20 Euros as compensation. Written informed consent was obtained from all participants after a full explanation of the procedures. The study was approved by the ethical committee of Ghent University Hospital (Belgium) and it was performed in accordance with relevant regulations and guidelines.
DNA methylation analysis
Genomic DNAs were extracted from peripheral blood using the DNeasy Blood & Tissue Kit (Qiagen), and an aliquot of 1 µg DNA per subject was processed for bisulphite conversion (Zymo Research EZ Methylation Kit), according to the manufacturer’s instructions. Subsequently, DNA methylome was analysed using the Illumina © Infinium Human Methylation 850k BeadChip array (Illumina, San Diego, CA, USA) that assesses 862,927 cytosine–phosphate–guanine (CpG) sites throughout the genome, covering 99% of RefSeq genes, 95% of CpG islands and offers high coverage of enhancer regions. Studies have shown a correlation between DNA methylation in blood and brain tissue61. The array was scanned with the Illumina iScan SQ system and the image intensities were extracted with Genome Studio (2011.1). DNA quality controls, data normalization and statistical filters were performed with the Partek® Genomics Suite® v7.19.1018. Methylation of the X and Y chromosomes were excluded from the analysis in order to compare XX and XY populations. Functional normalization, NOOB background correction, and dye correction were applied.
All analyses were done by the Partek® Genomics Suite® software, version 7.0 and are supported by RStudio. The human reference genome (GRCh37/hg19 assembly) was used to determine the location and features of the gene region using the UCSC Genome Browser62.
MRI acquisition and analysis
The three-dimensional MRI data sets were acquired at Ghent University Hospital (Belgium). High-resolution T1-weighted images were acquired for all participants on a 3-Tesla Siemens Prisma Fit MRI scanner (Siemens Healthcare, Erlangen, Germany) with a 64-channel head coil. The following parameters were used: a MPRAGE sequence with TR/TE = 2300/2.96 ms; TI = 900 ms; 192 slices; flip angle 9 degrees; 256 × 256 matrix and 1 mm3 isotropic voxel.
Automated cortical reconstruction of the T1-weighted images was performed using the FreeSurfer (version 6.0.0) image analysis suite (http://surfer.nmr.mgh.harvard.edu). This method was used to create a cortical surface three-dimensional model of CTh using intensity and continuity information previously described in detail63. Processing of T1 high resolution images includes several procedures: motion correction, removal of non-brain tissue using a hybrid watershed/surface deformation procedure64, automated Talairach transformation, intensity normalization65, tessellation of the gray matter/white matter boundary, automated topology correction66, 67, and surface deformation to detect gray matter/white matter and gray matter/cerebrospinal fluid boundaries where the greatest shift in intensity defines the transition to the other tissue class63. Moreover, the cerebral cortex was divided into different regions according to gyral and sulcal structure information68. The resulting representation of CTh is calculated as the distance between tissue boundaries (gray matter/white matter and gray matter/cerebrospinal fluid63. All surface models in our study were visually inspected for accuracy.
Analysis procedure
Firstly, after a quality control and filtering, a wise-pair differential methylation study was performed to compare our groups (CW-TM, CM-TM and CW-CM) to ensure there are significant CpG sites before continuing the analysis. In second place, CTh values (total, left and right hemisphere) were introduced as new variables to apply an ANOVA test to obtain genes which were specially related to CTh and TM. These candidate genes were used to perform a correlation analysis with regional CTh values based on the Desikan atlas68. Complementary correlations were performed to investigate the effect of age of onset of GD in the relationship between methylation data and regional CTh. More detailed information is described in the following paragraphs.
Statistical analysis
Sociodemographic data (age, smoking, drugs, race, onset)
Age was tested for normality and homogeneity. Age differences between the three groups were tested using a Kruskal–Wallis test. Categorical variables such as smoking and age of onset were compared using a X2 test of independence. When complementary analyses were performed in a subset of participants (early vs. late onset transgender men), the age variable was compared using Mann–Whitney U statistics. All statistical analyses were performed using IBM SPSS Statistics v. 29.
Methylation data and functional and regulatory enrichment analysis
To detect the differential methylation in CpG sites that varies across all groups, we compared CW-TM, CM-TM, CW-CM. Then, we performed an ANOVA test comparing groups by CTh: CW-TM, CM-TM, CW-CM.
For each contrast, a p-value, Beta difference (Δβ), and M difference (ΔM) were generated. P values were calculated using false discovery rate correction (FDR, p < 0.05) and fold change ≥ ± 2 was applied. The distribution of significant CpG sites was examined across functional and regulatory annotations. The GO enrichment analysis was carried out with the Partek® Pathway program (Supplemental Material Table S8).
MRI data: regional CTh values
Freesurfer‐generated surfaces were used to calculate CTh estimates according to the 68 regions contemplated in the Desikan atlas66. Regional CTh in the above-mentioned areas (34 areas per hemisphere) were compared between the three groups of interest (TM, CW and CM) using a multivariate analysis of covariance. Complementary analyses were performed in order to study the role of age of onset of GD. The reason for this complementary analysis is a previous report where we found metabolomic differences within the TM group regarding early vs. late onset of GD57. Bonferroni’s post-hoc test was employed to assess CTh differences between groups. Partial squared eta was employed to calculate the effect sizes; around 0.01 is a small size effect, 0.06 is medium, and above 0.14 is large.
Methylation correlation with MRI data
Regional CTh correlations with methylation data were conducted, specifically with those genes that showed a different methylation degree between groups. As a secondary analysis, the relationship between CTh and methylation degree was explored considering age at GD onset (early vs. late onset). Partial correlations between CTh and genetic data were also performed in early vs. late TM. The Smoking variable was introduced as a covariate in all analyses. MRI data: regional CTh values and Methylation and MRI data sections’ analysis were performed by means of IBM SPSS Statistics v. 29.
Multiple comparisons correction
To be statistically conservative, we applied a multi-layered approach of correcting for multiple comparisons at several levels. First, to correct for multiple genes, we used the common FDR correction (p < 0.05). Second, to correct for brain regions, we used Bonferroni correction; significance was set at p ≤ 0.001 (CTh areas (left/right) p = 0.05/34 = 0.001).
Ethical approval
This study was reviewed and approved by the Ethical Committees of Gent University Hospital, Belgium. The participants provided their written informed consent to participate in this study.
Data availability
The datasets presented in this study can be found in GEO online repository. The names of the repository and accession number(s) can be found below: GSE237955. The samples were randomised across EPIC slides.
References
American Psychiatric Association, Diagnostic and Statistical Manual of Mental Disorders (DSM-5). (2013).
The ICD-11 classification of mental and behavioural disorders: clinical description and diagnostic guidelines. World Health Organization, (2018).
Zhou, J. N., Hofman, M. A., Gooren, L. J. & Swaab, D. F. A sex difference in the human brain and its relation to transsexuality. Nature. 378(6552), 68–70 (1995).
Zubiaurre-Elorza, L. et al. Cortical thickness in untreated transsexuals. Cereb Cortex. 23(12), 2855–2862 (2013).
Mueller, S. C. et al. The neuroanatomy of transgender identity: Mega-analytic findings from the ENIGMA transgender persons working group. J. Sex Med. 18(6), 1122–1129 (2021).
Rametti, G. et al. The microstructure of white matter in male to female transsexuals before cross-sex hormonal treatment. A DTI study. J. Psychiatr. Res. 45(7), 949–954 (2011).
Rametti, G. et al. White matter microstructure in female to male transsexuals before cross-sex hormonal treatment A diffusion tensor imaging study. J. Psychiatr. Res. 45(2), 199–204 (2011).
Kranz, G. S. et al. White matter microstructure in transsexuals and controls investigated by diffusion tensor imaging. J. Neurosci. 34(46), 15466–15475 (2014).
Guillamon, A., Junque, C. & Gómez-Gil, E. A review of the status of brain structure research in transsexualism. Arch. Sex. Behav. 45(7), 1615–1648 (2016).
Uribe, C. et al. Brain network interactions in transgender individuals with gender incongruence. Neuroimage. 211, 116613 (2020).
Uribe, C., Junque, C., Gómez-Gil, E., Díez-Cirarda, M. & Guillamon, A. Brain connectivity dynamics in cisgender and transmen people with gender incongruence before gender affirmative hormone treatment. Sci. Rep. 11(1), 21036 (2021).
Uribe, C. et al. Whole-brain dynamics differentiate among cisgender and transgender individuals. Hum. Brain Mapp. 43(13), 4103–4115 (2022).
Panizzon, M. S. et al. Distinct genetic influences on cortical surface area and cortical thickness. Cereb Cortex. 19(11), 2728–3735 (2009).
Winkler, A. M. et al. Cortical thickness or grey matter volume? The importance of selecting the phenotype for imaging genetics studies. Neuroimage 53(3), 1135–1146 (2010).
Jha, S. C. et al. Genetic influences on neonatal cortical thickness and surface area. Hum. Brain Mapp. 39(12), 4998–5013 (2018).
Walton, E., Baltramonaityte, V., Calhoun, V. D., Heijmans, B., Thompson, P. & Cecil, C. A systematic review of neuroimaging epigenetic research: Calling for an increased focus on development [Internet]. PsyArXiv; psyarxiv.com/4a8xn (2020).
Fjell, A. M. et al. Development and aging of cortical thickness correspond to genetic organization patterns. Proc. Natl. Acad. Sci. USA. 112(50), 15462–15467 (2015).
Sowell, E. R. et al. Longitudinal mapping of cortical thickness and brain growth in normal children. J. Neurosci. 24(38), 8223–8231 (2004).
Shaw, P. et al. Neurodevelopmental trajectories of the human cerebral cortex. J. Neurosci. 28(14), 3586–3594 (2008).
Benavides, A. et al. Sex-specific alterations in preterm brain. Pediatr. Res. 85(1), 55–62 (2019).
Sowell, E. R. et al. Sex differences in cortical thickness mapped in 176 healthy individuals between 7 and 87 years of age. Cereb Cortex. 17(7), 1550–1560 (2007).
Raznahan, A. et al. Longitudinally mapping the influence of sex and androgen signaling on the dynamics of human cortical maturation in adolescence. Proc. Natl. Acad. Sci. USA. 107(39), 16988–16993 (2010).
Cortes, L. R. & Forger, N. G. Role of epigenetics in shaping sex differences in brain development and behavior. In: Perinatal and Developmental Epigenetics. Elsevier. p. 209–239 (2023).
Cortes, L. R. et al. DNA methylation and demethylation underlie the sex difference in estrogen receptor alpha in the arcuate nucleus. Neuroendocrinology 112(7), 636–648 (2022).
Cisternas, C. D., Cortes, L. R., Golynker, I., Castillo-Ruiz, A. & Forger, N. G. Neonatal inhibition of DNA methylation disrupts testosterone-dependent masculinization of neurochemical phenotype. Endocrinology 161(1), bqz022 (2020).
Henningsson, S. et al. Sex steroid-related genes and male-to-female transsexualism. Psychoneuroendocrinology 30(7), 657–664 (2005).
Hare, L. et al. Androgen receptor repeat length polymorphism associated with male-to-female transsexualism. Biol. Psychiatry. 65(1), 93–96 (2009).
Fernández, R. et al. The (CA)n polymorphism of ERβ gene is associated with FtM transsexualism. J. Sex Med. 11(3), 720–728 (2014).
Fernández, R. et al. Molecular basis of gender dysphoria: androgen and estrogen receptor interaction. Psychoneuroendocrinology. 98, 161–167 (2018).
Foreman, M. et al. Genetic link between gender dysphoria and sex hormone signaling. J. Clin. Endocrinol. Metab. 104(2), 390–396 (2018).
Auger, A. P., Tetel, M. J. & Mccarthy, M. M. Steroid receptor coactivator-1 (SRC-1) mediates the development of sex-specific brain morphology and behavior. Proc. Natl. Acad. Sci. USA. 97(13), 7551–7555 (2000).
Ramírez, K. et al. Implications of the estrogen receptor coactivators SRC1 and SRC2 in the biological basis of gender incongruence. Sex Med. 9(3), 100368 (2021).
McCarthy, M. M. & Nugent, B. M. At the frontier of epigenetics of brain sex differences. Front. Behav. Neurosci. 9, 221 (2015).
Ramirez, K. et al. Epigenetics is implicated in the basis of gender incongruence: An epigenome-wide association analysis. Front. Neurosci. 15, 701017 (2021).
Shepherd, R. et al. Gender-affirming hormone therapy induces specific DNA methylation changes in blood. Clin. Epigenet. 14(1), 24 (2022).
Aranda, G. et al. Effects of sex steroids on the pattern of methylation and expression of the promoter region of estrogen and androgen receptors in people with gender dysphoria under cross-sex hormone treatment. J. Steroid Biochem. Mol. Biol. 172, 20–28 (2017).
Fernández, R. et al. Gender-affirming hormone therapy modifies the CpG methylation pattern of the ESR1 gene promoter after six months of treatment in transmen. J. Sex Med. 17(9), 1795–1806 (2020).
Lyall, A. E. et al. Dynamic development of regional cortical thickness and surface area in early childhood. Cereb Cortex. 25(8), 2204–2212 (2015).
Burke, S. M. et al. Testosterone effects on the brain in transgender men. Cereb Cortex. 28(5), 1582–1596 (2018).
Cao, M. et al. Upregulation of CBLL1 in rat brain cortex after lipopolysaccharide treated. J. Mol. Histol. 44(2), 135–145 (2013).
Gong, E. Y., Park, E. & Lee, K. Hakai acts as a coregulator of estrogen receptor alpha in breast cancer cells. Cancer Sci. 101(9), 2019–2025 (2010).
Bawankar, P. et al. Hakai is required for stabilization of core components of the m6A mRNA methylation machinery. Nat. Commun. 12(1), 3778 (2021).
Livneh, I., Moshitch-Moshkovitz, S., Amariglio, N., Rechavi, G. & Dominissini, D. The m6A epitranscriptome: transcriptome plasticity in brain development and function. Nat. Rev. Neurosci. 21, 36–51 (2020).
Patil, D. P. et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 537(7620), 369–373 (2016).
Yoon, K. J. et al. Temporal control of mammalian cortical neurogenesis by m6A methylation. Cell. 171(4), 877-889.e17 (2017).
Xu, H. et al. m6A mRNA methylation is essential for oligodendrocyte maturation and CNS myelination. Neuron. 105(2), 293-309.e5 (2020).
Giedd, J. N., Raznahan, A., Mills, K. L. & Lenroot, R. K. Review: Magnetic resonance imaging of male/female differences in human adolescent brain anatomy. Biol. Sex Differ. 3(1), 19 (2012).
Tamnes, C. K., Roalf, D. R., Goddings, A. L. & Lebel, C. Diffusion MRI of white matter microstructure development in childhood and adolescence: Methods, challenges and progress. Dev. Cogn. Neurosci. 33, 161–175 (2018).
Underhill, S. M., Wheeler, D. S. & Amara, S. G. Differential regulation of two isoforms of the glial glutamate transporter EAAT2 by DLG1 and CaMKII. J. Neurosci. 35(13), 5260–5270 (2015).
Braithwaite, S. P., Meyer, G. & Henley, J. M. Interactions between AMPA receptors and intracellular proteins. Neuropharmacology. 39(6), 919–930 (2000).
Valtschanoff, J. G. et al. SAP97 concentrates at the postsynaptic density in cerebral cortex. Eur. J. Neurosci. 12(10), 3605–3614 (2000).
Fourie, C., Li, D. & Montgomery, J. M. The anchoring protein SAP97 influences the trafficking and localisation of multiple membrane channels. Biochim Biophys Acta. 1838(2), 589–594 (2014).
Cotter, L. et al. Dlg1-PTEN interaction regulates myelin thickness to prevent damaging peripheral nerve overmyelination. Science. 328(5984), 1415–1418 (2010).
Adey, N. B. et al. Threonine phosphorylation of the MMAC1/PTEN PDZ binding domain both inhibits and stimulates PDZ binding. Cancer Res. 60(1), 35–37 (2000).
Smith, Y. L. S., Van Goozen, S. H. M., Kuiper, A. J. & Cohen-Kettenis, P. T. Transsexual subtypes: Clinical and theoretical significance. Psychiatry Res. 137(3), 151–160 (2005).
Van Goozen, S. H. M., Slabbekoorn, D., Gooren, L. J. G., Sanders, G. & Cohen-Kettenis, P. T. Organizing and activating effects of sex hormones in homosexual transsexuals. Behav. Neurosci. 116(6), 982–988 (2002).
Collet, S. et al. Characterization of the 1H-MRS metabolite spectra in transgender men with gender dysphoria and cisgender people. J. Clin. Med. 10(12), 2623 (2021).
Feusner, J. D. et al. Intrinsic network connectivity and own body perception in gender dysphoria. Brain Imaging Behav. 11(4), 964–976 (2017).
Barfield, R. T. et al. Accounting for population stratification in DNA methylation studies. Genet. Epidemiol. 38(3), 231–241 (2014).
Sheehan, D. V. et al. The Mini-International Neuropsychiatric Interview (MINI): the development and validation of a structured diagnostic psychiatric interview for DSM-IV and ICD-10. J. Clin. Psychiatry. 59, 22–33 (1998).
Lunnon, K. et al. Methylomic profiling implicates cortical deregulation of ANK1 in Alzheimer’s disease. Nat. Neurosci. 17(9), 1164–1170 (2014).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12(6), 996–1006 (2002).
Fischl, B., Fischl, B., Dale, A. M. & Dale, A. M. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc. Natl. Acad. Sci. USA. 97(20), 11050–11055 (2000).
Ségonne, F. et al. A hybrid approach to the skull stripping problem in MRI. Neuroimage. 22(3), 1060–1075 (2004).
Sled, J. G., Zijdenbos, A. P. & Evans, A. C. A nonparametric method for automatic correction of intensity nonuniformity in MRI data. IEEE Trans. Med. Imaging. 17(1), 87–97 (1998).
Fischl, B., Liu, A. & Dale, A. M. Automated manifold surgery: constructing geometrically accurate and topologically correct models of the human cerebral cortex. IEEE Trans. Med. Imaging. 20(1), 70–80 (2001).
Ségonne, F., Pacheco, J. & Fischl, B. Geometrically accurate topology-correction of cortical surfaces using nonseparating loops. IEEE Trans. Med. Imaging. 26(4), 518–529 (2007).
Desikan, R. S. et al. An automated labelling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage. 31(3), 968–980 (2006).
Acknowledgements
We are grateful to everyone who directly or indirectly contributed to the study.
Funding
This work was supported by grants: Ministerio de Ciencia, Innovación y Universidades: PGC2018-094919-B-C21 & PDI21-127547NB-C21 (A.G.), PGC2018-094919-B-C22 and PID2021-127547NB-C22 (R.F., E.P.), Xunta de Galicia ED431B 2022/16 (E.P.) and Special Research Fund (BOF) of Ghent University (IOP003- 18) (S.M.C. and G.T.S).
Author information
Authors and Affiliations
Contributions
R.F., A.G., E.P. and S.M.C. made the design of the study; M.K. and S.C. collected the M.R.I. data and DNAs; R.F., L.Z.E., N.O. and A.S. organized and analysed the data; A.G., R.F., L.Z.E., A.S., S.M.C. and E.P. wrote the main manuscript text; G.T.S. provided clinical advice. All authors (R.F., L.Z.E., A.S., N.O., S.C., M.K., G.T.S., S.M.C., A.G., E.P.) reviewed the manuscript and have approved the submitted version. All authors (R.F., L.Z.E., A.S., N.O., S.C., M.K., G.T.S., S.M.C., A.G., E.P.) have agreed both to be personally accountable for the author's own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernández, R., Zubiaurre-Elorza, L., Santisteban, A. et al. CBLL1 is hypomethylated and correlates with cortical thickness in transgender men before gender affirming hormone treatment. Sci Rep 13, 21609 (2023). https://doi.org/10.1038/s41598-023-48782-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48782-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.