Abstract
The COVID-19 pandemic in Korea has dynamically changed with the occurrence of more easily transmissible variants. A rapid and reliable diagnostic tool for detection of SARS-CoV-2 is needed. While RT-PCR is currently the gold standard for detecting SARS-CoV-2, the procedure is time-consuming and requires expert technicians. The rapid antigen detection test (RADT) was approved as a confirmatory test on 14 March 2022 due to rapid dissemination of the Omicron variant. The benefits of the RADT are speed, simplicity, and point-of-care feasibility. The aim of our study was to evaluate the clinical performance of RADT compared to RT-PCR in a single center over 15 months, fully covering the SARS-CoV-2 ‘Variants of Concern (VOC).’ A total of 14,194 cases was simultaneously tested by RT-PCR and RADT from January 2021 to March 2022 in Gangnam Severance Hospital and were retrospectively reviewed. PowerChek SARS-CoV-2, Influenza A&B Multiplex Real-time PCR Kit, and STANDARD Q COVID-19 Ag Test were used. Positive rates, sensitivities, specificities, positive predictive values (PPV), and negative predictive values (NPV) were estimated for five periods (3 months/period). Receiver operator characteristic curve (ROC) analysis was performed, and Spearman’s rank test assessed the correlation between RT-PCR Ct values and semi-quantitative RADT results. The overall positive rate of RT-PCR was 4.64%. The overall sensitivity and specificity were 0.577 [95% confidence interval (CI) 0.539–0.614] and 0.991 [95% CI 0.989–0.993], respectively. ROC analysis resulted in an area under the curve of 0.786 (P < 0.0001, Yuden’s index = 0.568). The PCR positive rates were estimated as 0.11%, 0.71%, 4.51%, 2.02%, and 13.72%, and PPV was estimated as 0.045, 0.421, 0.951, 0.720, and 0.798 in Periods 1, 2, 3, 4, and 5, respectively. A significant and moderate negative correlation between PCR Ct values and semi-quantitative RADT results was observed (Spearman’s ρ = − 0.646, P < 0.0001). The RADT exhibited good performance in specimens with low Ct values (Ct ≤ 25.00) by RT-PCR. The PPV was significantly higher in Periods 3 and 5, which corresponds to rapid dissemination of the Delta and Omicron variants. The high PPV implies that individuals with a positive RADT result are very likely infected with SARS-CoV-2 and would require prompt quarantine rather than additional RT-PCR testing. The sensitivity of 0.577 indicates that RADT should not replace RT-PCR. Nonetheless, given the high PPV and the ability to track infected persons through rapid results, our findings suggest that RADT could play a significant role in control strategies for further SARS-CoV-2 variants.
Similar content being viewed by others
Introduction
The World Health Organization declared a global pandemic of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection, a novel coronavirus disease 2019 (COVID-19), in March 2020. In accordance with global trends, the national situation of COVID-19 in Korea has changed dynamically. The Delta variant was disseminated in early August 2021, and the Omicron variant replaced the Delta variant in December 20211,2,3,4. The number of COVID-19 cases escalated in each event, with the appearance of variants of higher transmissibility; therefore, it is necessary to prepare a rapid and reliable diagnostic tool for detection of SARS-CoV-2. Real-time reverse transcription PCR (RT-PCR) has been the gold standard in detecting SARS-CoV-2 due to its high sensitivity and specificity5,6. Nevertheless, it is time-consuming and requires expert technicians to be performed accurately7,8,9. Rapid antigen detection tests could be an alternative option for diagnosis of COVID-19 infection.
Initial approval of a rapid SARS-CoV-2 Antigen Detection Test (RADT) was granted in Korea in November 2020. The RADT was approved for use in areas with urgent needs including emergency rooms, intensive care units, and long-term care facilities. The RADT exhibits lower sensitivity in detecting COVID-19 patients compared with PCR-based methods; however, the advantages of RADT are rapidity, simplicity, and point-of-care feasibility with a 15–20 min turnaround time10. There have been several studies on the diagnostic performance of RADTs compared with results obtained by RT-PCR; however, most of them were performed with small numbers of samples or were analyzed with inconsistent sample types, reagents, and test kits. Therefore, the clinical performance of RADTs has been reported in a variety of ranges with heterogeneity. In addition, there was a limitation in those prior studies that variants of SARS-CoV-2 could not be fully evaluated11,12,13,14. The aim of our study was to evaluate the clinical performance of the STANDARD Q COVID-19 Ag Test (SD Biosensor Inc., Gyeonggi-do, Korea) compared to RT-PCR with a large number of samples over a long period, covering the Alpha, Delta, and Omicron variant pandemics. We also analyzed the diagnostic performance of the STANDARD Q COVID-19 Ag Test according to the change of positive rates of SARS-CoV-2.
Materials and methods
Study population
To evaluate the clinical performance of RADT comparing with the results of RT-PCR, a retrospective chart review was performed. Inclusion criteria was the cases subjected for detection of SARS-CoV-2 by both RT-PCR and RADT in same day from January 2021 to March 2022. A total of 86,678 samples was tested for detection of SARS-CoV-2 by RT-PCR, and 14,194 cases simultaneously tested by RADT were included in this study (Fig. 1). The nasopharyngeal swab (NPS) and oropharyngeal swab (OPS) of the patients were collected and transported via viral transport medium (VTM) (AB Transport Medium (AB MEDICAL, Seoul, Korea)) and were used in performing RT-PCR and RADTs to detect SARS-CoV-2. All processes of this study were in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki declaration and comparable ethical standards. This study was approved by the Institutional Review Board of Yonsei University Gangnam Severance Hospital (approval number: 3-2023-0243) to waive the requirement for informed consents about the collection of clinical data.
SARS-CoV-2 RNA detection using RT-PCR
RNA was extracted from 300 \(\mathrm{\mu L}\) of VTM using the TANBead Nucleic Acid Extraction Kit (Taiwan Advanced Nanotech Inc., Taiwan) according to the manufacturer’s instructions. RT-PCR of SARS-CoV-2 was performed with PowerChek SARS-CoV-2, Influenza A&B Multiplex Real-time PCR Kit (KOGENEBIOTECH Co., Seoul, Korea). The PowerChek assay simultaneously targets the envelope (E) and open reading frame 1ab (ORF1ab) genes of SARS-CoV-2. One drop of 5 μL of extracted template RNA was added to 15 μL of PCR reaction mixture (2X RT-PCR Master Mix 10 \(\mathrm{\mu L}\) & Primer/Probe Mix 5 \(\mathrm{\mu L}\)). A total volume of 20 μL was loaded and amplified 40 cycles by the CFX96 Real-time PCR Detection System (Bio-Rad Laboratories, Inc., Hercules, CA). Exponential fluorescence curves that crossed the threshold line at or before 38 cycles (cycle threshold [Ct] ≤ 38) for both the SARS-CoV-2 E gene and ORF1ab gene were considered a positive result. A negative result was determined when Ct values for both genes were above 40 (Ct > 40). The indeterminate results were defined when Ct value of only one of the E gene and the ORF1ab gene was below 38 and when Ct values of either E gene or ORF1ab gene was between 38 and 40.
Rapid SARS-CoV-2 Antigen Detection Test (RADT)
The STANDARD Q COVID-19 Ag Test was approved by the MFDS of Korea in November 2020. This test is a lateral flow immunochromatographic assay (ICA) that detects the nucleocapsid protein (NP) antigen of SARS-CoV-2 semi-quantitatively. It has two pre-coated lines on the nitrocellulose membrane: control (C) and test (T) lines. A sample of 350 μL of VTM was mixed with the extraction buffer solution, and 2–3 drops of the sample mixture were applied on the sample area of the device. After 15–20 min of incubation at room temperature, results were determined manually with the naked eye. For a positive result, two colored C and T lines were present within 30 min. The test was interpreted as ‘trace’ if the T line was fainter than the C line; ‘1+ (one positive)’ if the T line showed similar intensity to the C line; and ‘2+ (two positive)’ if the T line was thicker than the C line. For a negative result, only the C line was present.
Statistical analysis
Analyse-it\(\circledR \) (version 5.68; Analyse-it Software Ltd., Leeds, UK) was used for all statistical analyses. Differences between groups were analyzed using Kruskal–Wallis tests for continuous variables and Chi-square tests for categorical variables. Spearman’s rank test was performed to assess the correlation between results by RT-PCR and RADT. Sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated for the results obtained by the STANDARD Q COVID-19 Ag Test compared with those by RT-PCR. The 95% confidence intervals (CIs) were estimated by Wilcoxon sum rank test. To evaluate the diagnostic performance of the STANDARD Q COVID-19 Ag Test, receiver operating characteristics (ROC) curve analysis was performed, and the area under curve (AUC) was estimated. Statistical significance was determined when the P value was less than 0.05.
Results
Characteristics of cases, PCR results, and ORF1ab gene Ct values by period
Among the 14,194 cases, 500 positive and 13,694 negative results were obtained. The positive results consisted of 258 results of 2+, 128 results of 1+, and 114 results of trace (Fig. 1).
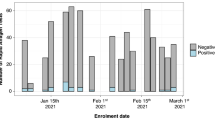
The evaluation period was divided into 5 periods (3 months per period), and the number of cases examined using the STANDARD Q COVID-19 Ag Test by period ranged from 1822 to 3217. The gender ratio, median age, PCR qualitative results, and ORF1ab gene Ct value distribution are shown in Table 1. The PCR positive rate was estimated to be 0.11% (2/1822) in Period 1, surged to 4.51% (145/3217) in Period 3 (the Delta variant epidemic), and peaked at 13.72% (429/3127) in Period 5 (the Omicron variant epidemic). The proportion of cases with Ct values less than 25.00, indicating a high viral load, was 0.05% in Period 1, increased to 2.55% in Period 3, and peaked at 6.78% in Period 5.
Correlation between Ct value of ORF1ab gene PCR and semiquantitative results obtained by STANDARD Q COVID-19 Ag Test
Table 2 summarizes the qualitative and semiquantitative results of the STANDARD Q COVID-19 Ag Test according to the Ct values of ORF1ab gene PCR, including indeterminate results. Most (329/348, 94.5%) of the cases with low CT values under 25.00 gave a positive result with the STANDARD Q COVID-19 Ag Test. In addition, most (137/157, 89.0%) of the cases with Ct values under 20.00 showed 2+, and more than half (102/191, 58.3) of the cases with Ct values of 20.00–24.99 also showed 2+ with the STANDARD Q COVID-19 Ag Test. When semiquantitative results of the STANDARD Q COVID-19 Ag Test were compared with Ct values obtained by RT-PCR, a significant and moderate negative correlation was observed (Spearman’s ρ = − 0.646, P < 0.0001) (Fig. 2).
Clinical performance of the RADT
The PCR positive rates, clinical performance of the STANDARD Q COVID-19 Ag Test compared with the PCR results, and PPV and NPV by period are summarized in Table 3. The overall sensitivity, specificity, PPV, and NPV were 0.577 (95% CI 0.539–0.614), 0.991 (95% CI 0.989–0.993), 0.760 (0.724–0.795), and 0.980 (95% CI 0.978–0.981), respectively. The PPV was lowest in Period 1 (0.045; 95% CI 0.011–0.169) and high in Period 3 (0.951; 95% CI 0.890–0.979) and Period 5 (0.798; 95% CI 0.751–0.838). The specificity was greater than 0.970 in all periods. ROC curve analysis resulted in an AUC of 0.786 (P < 0.0001) and Youden’s index of 0.568 (Fig. 3). The monthly trends of sensitivity, specificity, PPV, NPV, and PCR positive rates are described in Supplementary Figure S1.
Discussion
Antigen-based immunoassays for detecting SARS-CoV-2 were introduced for rapid diagnosis of COVID-19. In this study, the clinical performance of the STANDARD Q COVID-19 Ag Test, an immunochromatographic assay, was evaluated in comparison with results obtained by the RT-PCR method.
There have been previous studies evaluating the clinical performance of RADTs for diagnosing COVID-19 infection. In a German study, 2,028 samples collected from a variable population including patients with COVID-19 symptoms, individuals with SARS-CoV-2 exposure, and hospital staff members showed overall sensitivity and specificity of 42.86 and 99.89%, respectively15. In a study from Serbia, 120 symptomatic patients were tested by the RADT, and overall sensitivity was about 60%16. In another study from Egypt performed with COVID-19 suspected individuals, the sensitivity and specificity were 78.2% and 64.2%, respectively17. A previous study conducted in Korea with pre-hospital patients, emergency room visitors, and patients confirmed with COVID-19 by RT-PCR demonstrated an overall sensitivity of 17.5% and a specificity of 100%18. This study was performed with a large number of clinical samples over 15 months, provided more reliable results reflecting practical aspects compared with other evaluation studies, and minimized bias that might occur in selection of study target samples. Overall sensitivity and specificity of the RADT were 0.577 (95% CI, 0.539–0.614) and 0.991 (95% CI, 0.989–0.993) respectively, which are comparable to the results reported in previous studies16,17,18. In addition, three cases of false-negative results in the cases with Ct value < 20. Clerical errors, inappropriate specimen handling, or interference in specimens could be potential reasons19. A total 120 cases (0.89%) of false-positive results were identified in this study. According to the previous report in Canada, the false positive results were identified in 0.05% (462/903,408) of asymptomatic screening tests20, and a false positive rate in another study in Germany was reported to be 0.13% (1561/1,245,962)21. False positive results may lead to unnecessary quarantine and delay of appropriate procedure to patients. The AUC of RADT was suboptimal (0.781) due to relatively low sensitivity and confirmation test with RT-PCR assay should be performed for accurate diagnosis of SARS-CoV-2 infection.
Significant correlation was identified between the Ct values obtained by RT-PCR and semiquantitative results by RADT, and the accuracy of RADT was very high, particularly in cases with high viral loads, exhibiting low Ct values22,23. In the German study, higher sensitivities were noted as Ct values < 20, < 25, and < 30 produced results of 100%, 98.25%, and 88.64%, respectively15. Therefore, the RADT could be accurate in diagnosing SARS-CoV-2 infection in the early stage, in which a high viral load is usually identified.
PPV and NPV are the most important indexes to determine clinical relevance and could vary according to disease prevalence. During the study period, steep increases in prevalence were observed in Periods 3 and 5, which correspond to the Delta and Omicron variant epidemics, respectively. As the prevalence increased, PPVs were confirmed to increase in Period 3 (0.951). The high PPV in the COVID-19 epidemic indicates that an individual with a positive RADT result is highly suspected to be infected with SARS-CoV-2 and should be required to promptly quarantine rather than undergo additional RT-PCR testing. Meanwhile, the PCR positive rate in Period 5 (13.7%) was higher than that in Period 3 (4.5%), but PPV in Period 5 was lower than that in Period 3. During the Period 5, the number of positive cases were dramatically increased up to 4.1% (55/1336), and Ct value of more than half of these cases were ≥ 25.00.
Most RADTs target the nucleocapsid protein stably associated with RNA in the virion and is found at higher level than other viral structural proteins24. Prior to the introduction of VOCs, the majority of RADTs was developed with strains that lacked nucleocapsid mutations25, and there have only been a few clinical validation studies that have examined variants affecting antigen test performance. From our results, it can be inferred that the Omicron and Delta variants, which are characterized by many mutations in the spike protein, had little effect on the accuracy of the nucleocapsid targeting RADT. Nonetheless, Omicron and Delta variants also harbor nucleocapsid proteins involving mutations (i.e., Omicron; P13L, ∆31–33, R203K & Delta; D63G, R203M, D377Y), in conjunction with the spike protein's conspicuous alterations26,27. While there are some false-negative reports of specimens carrying nucleocapsid mutations that show a high viral load as confirmed by PCR28,29, it is considered not to affect the accuracy of the RADT since current tests target the C-terminus of the nucleocapsid protein, whereas the great majority of mutations is identified in the N-terminus. In our study, no substantial shift or inclination in specificity was identified according to variant, which indicates the variants did not affect the clinical performance of the RADT. However, we must remain cautious to detect changes in the sensitivity of RADT due to variations with specific mutation trends, and additional studies are necessary to validate this point.
One of the limitations of this study is that the samples were collected in a single center in a single country; therefore, ethnic or racial diversities could not be considered. In addition, samples tested for pre-hospitalization screening were included; therefore, some asymptomatic patients were inevitably included in our study, which would lower the positive rate. The lack of further investigation about false-positive and false-negative results due to the retrospective study design was another limitation of this study. Finally, STADNARD Q COVID-19 Ag was approved for testing the detection of SARS-CoV-2 by Ministry of Food and Drug Safety of the Republic of Korea in both specimens including nasopharyngeal swab and VTM. Even though the limit of detection of this diagnostic kit in both specimens were validated, possible dilution effect could be occurred when using VTM as a specimen.
In conclusion, the STANDARD Q COVID-19 Ag Test showed lower sensitivity compared with RT-PCR; however, this RADT kit showed high PPV in the pandemic situation with a high prevalence of COVID-19 infection, which suggests this RADT kit could be an alternative option for rapid detection of SARS-CoV-2 in a pandemic situation. Further investigation should be performed to compare the results of other RADTs to figure out the clinical usefulness of RADT in diagnosing SARS-CoV-2 infection.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
References
Park, S. et al. Clinical and virological characteristics of SARS-CoV-2 B.1617.2 (Delta) variant: a prospective cohort study. Clin. Infect. Dis. 75, e27 (2022).
Hwang, H. et al. Transmission dynamics of the delta variant of SARS-CoV-2 infections in South Korea. J. Infect. Dis. 225, 793–799 (2022).
Meo, S. A., Meo, A. S., Al-Jassir, F. F. & Klonoff, D. C. Omicron SARS-CoV-2 new variant: global prevalence and biological and clinical characteristics. Eur. Rev. Med. Pharmacol. Sci. 25, 8012–8018 (2021).
Kim, E. Y. et al. Community Transmission of SARS-CoV-2 Omicron Variant, South Korea, 2021. Emerg. Infect. Dis. 28, 898–900 (2022).
Lu, X. et al. US CDC real-time reverse transcription PCR panel for detection of severe acute respiratory syndrome coronavirus 2. Emerg. Infect. Dis. 26, 1654–1665 (2020).
Shen, M. et al. Recent advances and perspectives of nucleic acid detection for coronavirus. J. Pharm. Anal. 10, 97–101 (2020).
Russo, A. et al. Current Status of laboratory diagnosis for COVID-19: a narrative review. Infect. Drug Resist. 13, 2657–2665 (2020).
Corman, V. M. et al. Novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020, 25 (2019).
Kevadiya, B. D. et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. 20, 593–605 (2021).
Young, S. et al. Clinical evaluation of BD Veritor SARS-CoV-2 point-of-care test performance compared to PCR-based testing and versus the sofia 2 SARS antigen point-of-care test. J. Clin. Microbiol. 59, e02338 (2020).
Wagenhäuser, I. et al. Clinical performance evaluation of SARS-CoV-2 rapid antigen testing in point of care usage in comparison to RT-qPCR. EBioMedicine 69, 103455 (2021).
Brümmer, L. E. et al. Accuracy of novel antigen rapid diagnostics for SARS-CoV-2: A living systematic review and meta-analysis. PLoS Med. 18, e1003735 (2021).
Khandker, S. S., Nik Hashim, N. H. H., Deris, Z. Z., Shueb, R. H. & Islam, M. A. Diagnostic accuracy of rapid antigen test kits for detecting SARS-CoV-2: A systematic review and meta-analysis of 17,171 suspected COVID-19 patients. J. Clin. Med. 10, 3493 (2021).
Chaimayo, C. et al. Rapid SARS-CoV-2 antigen detection assay in comparison with real-time RT-PCR assay for laboratory diagnosis of COVID-19 in Thailand. Virol. J. 17, 177 (2020).
Korenkov, M. et al. Evaluation of a Rapid Antigen Test to detect SARS-CoV-2 infection and identify potentially infectious individuals. J. Clin. Microbiol. 59, e0089621 (2021).
Ristić, M., Nikolić, N., Čabarkapa, V., Turkulov, V. & Petrović, V. Validation of the STANDARD Q COVID-19 antigen test in Vojvodina, Serbia. PLoS ONE 16, e0247606 (2021).
Amer, R. M. et al. Diagnostic performance of rapid antigen test for COVID-19 and the effect of viral load, sampling time, subject’s clinical and laboratory parameters on test accuracy. J. Infect. Public Health 14, 1446–1453 (2021).
Oh, S. M. et al. Clinical application of the standard Q COVID-19 Ag test for the detection of SARS-CoV-2 infection. J. Korean Med. Sci. 36, e101 (2021).
Safiabadi Tali, S. H. et al. Tools and techniques for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)/COVID-19 detection. Clin. Microbiol. Rev. 34, e00228-e320 (2021).
Gans, J. S. et al. False-Positive results in Rapid Antigen Tests for SARS-CoV-2. JAMA 327(5), 485–486 (2022).
Kretschmer, A. et al. False positive rapid antigen tests for SARS-CoV-2 in the real-world and their economic burden. J. Infect. 84(2), 248–288 (2022).
La Scola, B. et al. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur. J. Clin. Microbiol. Infect. Dis. 39, 1059–1061 (2020).
Sarkar, B., Sinha, R. N. & Sarkar, K. Initial viral load of a COVID-19-infected case indicated by its cycle threshold value of polymerase chain reaction could be used as a predictor of its transmissibility - An experience from Gujarat, India. Indian J. Community Med. 45, 278–282 (2020).
Yao, H. et al. Molecular architecture of the SARS-CoV-2 virus. Cell 183, 730–8.e13 (2020).
Harcourt, J. et al. Severe acute respiratory syndrome coronavirus 2 from patient with coronavirus disease, United States. Emerg. Infect. Dis. 26, 1266–1273 (2020).
Suratekar, R. et al. High diversity in Delta variant across countries revealed by genome-wide analysis of SARS-CoV-2 beyond the Spike protein. Mol. Syst. Biol. 18, e10673 (2022).
Osterman, A. et al. Impaired detection of omicron by SARS-CoV-2 rapid antigen tests. Med. Microbiol. Immunol. 211, 105–117 (2022).
Bourassa, L. et al. A SARS-CoV-2 nucleocapsid variant that affects Antigen Test performance. J. Clin. Virol. 141, 104900 (2021).
Del Vecchio, C., Brancaccio, G., Brazzale, A.R., Lavezzo, E., Onelia, F., Franchin, E., et al.: Emergence of N antigen SARS-CoV-2 genetic variants escaping detection of antigenic tests. MedRxiv 2021.
Lalkhen, A. G. & McCluskey, A. Clinical tests: sensitivity and specificity. Contin. Educ. Anaesthesia Crit. Care Pain 8, 221–223 (2008).
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.P., D.K., and S.H.J.; Data curation: Y.P. and S.W.K.; Supervision: Y.P. and D.K.; Writing - original draft: S.W.K.. Writing - review & editing: D.K. and Y.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, S.W., Park, Y., Kim, D. et al. A single-center experience on long-term clinical performance of a rapid SARS-CoV-2 Antigen Detection Test, STANDARD Q COVID-19 Ag Test. Sci Rep 13, 20777 (2023). https://doi.org/10.1038/s41598-023-48194-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48194-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.