Abstract
The annual goatgrass, Aegilops biuncialis is a rich source of genes with considerable agronomic value. This genetic potential can be exploited for wheat improvement through interspecific hybridization to increase stress resistance, grain quality and adaptability. However, the low throughput of cytogenetic selection hampers the development of alien introgressions. Using the sequence of flow-sorted chromosomes of diploid progenitors, the present study enabled the development of chromosome-specific markers. In total, 482 PCR markers were validated on wheat (Mv9kr1) and Ae. biuncialis (MvGB642) crossing partners, and 126 on wheat-Aegilops additions. Thirty-two markers specific for U- or M-chromosomes were used in combination with GISH and FISH for the screening of 44 Mv9kr1 × Ae. biuncialis BC3F3 genotypes. The predominance of chromosomes 4M and 5M, as well as the presence of chromosomal aberrations, may indicate that these chromosomes have a gametocidal effect. A new wheat-Ae. biuncialis disomic 4U addition, 4M(4D) and 5M(5D) substitutions, as well as several introgression lines were selected. Spike morphology and fertility indicated that the Aegilops 4M or 5M compensated well for the loss of 4D and 5D, respectively. The new cytogenetic stocks represent valuable genetic resources for the introgression of key genes alleles into wheat.
Similar content being viewed by others
Introduction
Bread wheat (Triticum aestivum L; 2n = 6x = 42; AABBDD) is the third most important crop after rice and maize and is essential in the human diet. Thousands of years of cultivation and modern breeding in the twentieth century resulted in improved yield and quality1. However, it reduced the available genetic diversity within elite germplasm, making it difficult to identify the best allele combinations to produce superior cultivars.
Goatgrasses (Aegilops) are close relatives of Triticum, making them promising gene sources for increasing wheat genetic diversity through interspecific hybridization2. In the 23 species of the genus, seven different genomes (C, D, M, N, S, T, U) were identified, and 12 of them contain U and/or M genomes3. The allotetraploid Ae. biuncialis Vis. (2n = 4x = 28, UbUbMbMb) has a high ability for ecological adaptation3,4,5. Together with its diploid progenitors Ae. comosa Sm. in Sibth. & Sm. (2n = 2x = 14; MM) and Ae. umbellulata Zhuk. (2n = 2x = 14; UU), they are attractive sources of agronomically important genes. Ae. biuncialis has been reported as gene source to improve wheat drought and salt tolerance6,7,8, as well as resistance to pests and disseases such as barley yellow dwarf luteovirus9, powdery mildew10, yellow rust11, brown rust12, leaf rust13, and stem rust14. Numerous data are also available for improving grain quality traits such as dietary fibre, β-glucan, protein content, Zeleny sedimentation value, wet gluten content, grain hardness, and micronutrient content15,16,17. Despite the high genetic potential, only a few wheat-Ae. biuncialis hybrids and addition lines have been developed18,19,20. Recently, a set of winter wheat (Mv9kr1)-Ae. biuncialis addition lines carrying chromosomes 1Ub, 2Ub, 3Ub, 2Mb, 3Mb and 7Mb18,21,22 has been developed. Tan et al.23 produced a wheat (‘Chuannong 19’)-Ae. biuncialis partial amphiploid and used it to create addition lines representing chromosomes 6Ub24, 1Ub15 and 2Mb10, while Song et al.25 developed a disomic addition line 5Mb. The production of wheat-Ae. biuncialis substitution and translocation lines has been reported even less frequently. Farkas et al.16 produced a wheat-Ae. biuncialis 3Mb(4B) substitution line and a 3Mb·4BS translocation line. It is estimated that only 4–5 Ae. biuncialis accessions are represented in the previously published wheat-Ae. biuncialis introgression lines. However, recent two diversity analyses of Ae. biuncialis accessions from different ecogeographical habitats revealed that this species has a high genetic variation for heading date, seed quality traits, and stem rust resistance available for the introgression breeding programs4,5,14.
The ability to screen large populations for the presence of introgressed chromatin is essential for producing wheat-alien translocations. Molecular cytogenetic methods such as genomic in situ hybridization (GISH) and fluorescence in situ hybridization (FISH) are probably the most popular approaches for characterizing alien chromatin in the wheat genetic background. Thus, GISH and FISH have been widely used to identify alien introgressions, including wheat-Ae. biuncialis addition and translocation lines10,16,22,26. Although molecular cytogenetic methods cannot be excluded from the selection process, their disadvantages, such as time and labor intensity, make screening large populations difficult. Furthermore, due to the low resolution of FISH and GISH, small alien chromosome segments (micro-introgressions) responsible for the trait of interest are undetectable 27.
Selection assisted by molecular markers would facilitate the much higher throughput identification of M- and U-genome chromatin. Earlier, testing wheat Amplified Fragment Length Polymorphism (AFLP), Restriction Fragment Length Polymorphism (RFLP), and Simple Sequence Repeat (SSR) molecular markers on Aegilops species was a suitable approach to detect introgressed chromatin28,29. Sequence-Specific Amplified Polymorphism (S-SAP) markers, which take advantage of the diversity of the integration sites of specific retroelements, provided an additional opportunity to identify transferred Aegilops chromosomes in wheat30. The use of gene-based markers can be even more effective, as demonstrated by Howard et al.31, who used wheat Conserved Ortholog Set (COS) markers to map QTLs determining the B-type starch granules content in Ae. peregrina. The markers specific for conserved orthologous genes facilitated gene introgression from U and M genome Aegilops species into wheat as well as the investigation of wheat-Aegilops macrosynteny at chromosomal level32,33. Molnár et al.32 used wheat-Aegilops chromosome addition lines to map a series of COS markers to the U and M genomes, demonstrating the utility of COS markers. Because the markers have known positions on the deletion bin map of wheat genomes, they allowed researchers to study wheat-Aegilops homeologous relationships34. In the last decade, PCR-based Landmark Unique Gene (PLUG) markers, another gene specific marker family, have become increasingly popular for detecting Aegilops chromatin in the wheat15,33,35. Until now, the majority of Ae. biuncialis markers tested were also wheat-specific, limiting their effectiveness. The development of Aegilops-specific markers would greatly aid in the detection of introgressed Aegilops chromatin in wheat.
Recent advances in flow cytometric sorting of plant mitotic chromosomes linked to next generation sequencing technologies36 provide an efficient strategy for producing chromosome-specific genomic resources in the case of large Triticeae genomes of agronomically important species such as wheat, barley and rye37,38,39.
Chromosome sorting and sequencing from wild species have also been reported40,41, including Aegilops species with U and M genomes34,42,43,44. Tiwari et al.45 flow sorted chromosome 5Mg from a wheat-Ae. geniculata disomic substitution line and sequenced it with Illumina. Using sequence assembly, the authors were able to annotate 4236 genes and develop 4538 SNPs on chromosome 5Mg. Bivariate flow cytometry has also been used to dissect the genomes of the diploid progenitors Ae. umbellulata (U) and Ae. comosa (M) into individual chromosomes34. Draft sequence assemblies of the U and M chromosomes allowed the authors to develop molecular markers for specific cytogenetic positions of orthologous genes previously mapped by single gene FISH on the chromosomes43. Although these markers have been PCR validated on wheat-Aegilops addition lines, they have not been tested on prebreeding populations to select wheat-Ae. biuncialis introgression lines.
Motivated by the need to develop wheat-Ae. biuncialis introgression lines, our study aims to develop gene specific markers for Aegilops chromosomes and use them to screen a winter wheat BC3F3 pre-breeding population for the presence of alien chromatin.
With Ae. umbellulata chromosome-specific draft assemblies, three marker design strategies were used: (1) a set of deletion-bin mapped wheat Expressed Sequence Tag (EST) sequences was used to design markers specific for InDel regions polymorphic between wheat and Aegilops (InDEL markers). (2) The annotated gene models of Ae. umbellulata were also used for sequence similarity searches against wheat in order to develop Intron Targeting (IT) markers. (3) We also tested previously developed markers for specific cytogenetic positions on U- and M-genome chromosomes using single-gene FISH. Finally, a combination of PCR markers validated on wheat-Aegilops chromosome addition lines and molecular cytogenetic methods were used to detect alien chromatin in a winter wheat (Mv9kr1)-Ae. biuncialis MvGB642 BC3 population to identify and characterize new addition, substitution, and translocation lines.
Results
Development of PCR markers
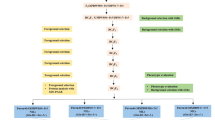
Sequence assemblies of Ae. umbellulata chromosomes enabled the development of new gene-specific markers that can support the Ae. biuncialis gene introgression into wheat. In this study, we used molecular markers developed by three different strategies (Fig. 1A–C).
Development of gene-based molecular markers using Ae. umbellulata chromosome sequences. (A) EST-based markers: wheat EST sequences were aligned to Ae. umbellulata chromosome sequences and primers were designed for the polymorphic exon regions. (B) Intron-targeting (IT) markers: annotated gene models of Ae. umbellulata were mapped to hexaploid wheat pseudomolecules. Primer pairs were designed for the last 100 bp of the first exon and the first 500 bp of the first intron. (C) cDNA-based markers: the physically mapped cDNA sequences of wheat orthologous genes 43 were aligned to the Ae. umbellulata and Ae. comosa chromosome sequences, and PCR primers were designed for the polymorphic (> 2 bp InDel) exon and intron regions. Aegilops chromosome sequences are highlighted in green, wheat sequences in ocher yellow, and polymorphic regions in white. F forward primer, R reverse primer.
The first strategy was based on the polymorphism between physically mapped wheat EST sequences and Ae. umbellulata chromosome sequences of (Fig. 1A). To generate EST markers, we aligned wheat EST sequences covering the wheat group I-VII chromosomes (sum of aligned ESTs = 4514; mean number of ESTs/chromosome: 644.8) to Ae. umbellulata chromosome contigs of Data S1. Using pairwise alignment, we identified polymorphic (> 5 bp InDels) hits with 338 ESTs on Ae. umbellulata chromosomes (1U:34, 2U:73, 3U:40, 4U:44, 5U:32, 6U:68, 7U:47). A total of 124 primer pairs out of the 313 primer pairs designed for the polymorphic regions of 204 ESTs were tested by PCR on the wheat (Mv9kr1), Ae. biuncialis (MvGB642), and Ae. umbellulata (AE740/03) (Fig. S1A, Table 1). Fifty markers (40.3%) produced polymorphic amplicons between wheat and Aegilops accessions (Data S2). In order to determine the chromosomal localization of the markers, we assigned the polymorphic markers to Aegilops chromosomes using wheat-Aegilops chromosome addition lines representing chromosomes 1U–7U and 1M–7M (Fig. S1B). Thirty-nine polymorphic markers were assigned to Aegilops chromosomes (Data S2.). Twenty-three markers were single-chromosome specific (1U: 9, 2U:1, 3U: 2, 4U:4, 5U:6, 6U:5, 7U: 7, 1M: 1, 5M: 2, 7M: 2), while three markers were detected on both the U and M genomes (1U/1M: BE497808, 7U/7M: BF483072, BE423703). These markers are considered suitable for selecting wheat lines with the corresponding Aegilops chromosomes during pre-breeding programs. (Table 1, Data S2).
In order to develop Aegilops chromosome-specific, Intron-targeting (IT) markers (Fig. 1B), the annotated gene models of Ae. umbellulata were mapped to the hexaploid wheat reference sequence (RefSeq v1.037). When suitable (> 10 bp) wheat/Aegilops polymorphism was found in the first exon and intron regions, a primer pair was designed within the last 100 bp of the exon (forward primer) and the first 500 bp of the intron (reverse primer). Finally, 15,475 primer pairs (5 primer pairs/contig) were designed using 3095 Ae. umbellulata contigs (Table 1, Data S3).
Eighty-four randomly selected primer pairs (12 markers per chromosome) were tested by PCR on the parental genotypes (Mv9kr1 and Ae. biuncialis MvGB642), and 29 (34.5%) of them showed presence/absence (21) or length (8) polymorphism between Aegilops and wheat (Data S2). Twenty polymorphic markers were located on Aegilops chromosomes using wheat Chinese Spring (CS)-Ae. umbellulata 1U-7U and CS-Ae. geniculata 1M–7M addition lines (1U: 2, 2U: 3, 3U: 2, 4U: 4, 5U: 3, 6U: 2, 7U: 3, 1M: 1, 2M: 2, 4M: 1, 7M: 1) (Table 1); sixteen were single-chromosome specific, three were located on both the U- and M-genome chromosomes (2U/2M: Ae2U14986.1, 7U/7M: Ae7U16619.2, 5U/4M: Ae5U23507.1), and the marker Ae4U242236.1 was detected on both the 4U and 6U chromosomes (Data S2).
The third type of markers was developed by Said et al.43 using wheat cDNAs that were single-gene FISH mapped to the U and M chromosomes of Ae. umbellulata and Ae. comosa, respectively. In this study, 19 markers were selected to characterize the Mv9kr1 × Ae. biuncialis MvGB642 BC3F3 generation (Data S2, Table 1).
In total, 16,062 primer pairs were designed using the three marker development strategies. 215 of the 482 markers tested on wheat and Aegilops parents showed polymorphisms. The chromosomal localization of 98 markers was determined using wheat-Aegilops addition lines. In order to trace the Aegilops chromosomes in the wheat background, 32 of these markers (3 EST, 10 IT, and 19 cDNA-based markers) were used to detect U or M chromosomes in the wheat-Ae. biuncialis BC3F3 population.
Marker-assisted selection of wheat-Ae. biuncialis introgression lines
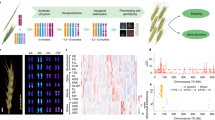
For tracking the individual Aegilops chromosomes in the wheat background, we chose markers that produced amplicons only on Aegilops chromosomes and tested them in 44 wheat-Ae. biuncialis BC3F3 hybrid progenies originated from crosses of hexaploid winter wheat line Mv9kr1 and Ae. biuncialis accession MvGB642 (Fig. S2). In order to validate the results of the 32 markers, sequential mcGISH and FISH were also performed on the BC3F3 population (results summarized in Fig. 2.).
Chromosomal constitution of wheat-Aegilops BC3F3 genotypes, as well as the amphiploid (Amph) and their parental wheat (Mv9kr1) and Ae. biuncialis MvGB642 (Ae.biu642) genotypes determined using GISH, FISH and molecular markers. Abbreviations for GISH and FISH results: Mon monosomic, Dis disomic, T translocation, Rob Robertsonian translocation (centric fusion), wheat unidentified wheat chromosome. The green and red colors represent the presence or absence of the Aegilops chromosome specific PCR amplicon, respectively.
Based on the markers, five of the 14 Aegilops chromosomes were detected in the pre-breeding population (Fig. 2; Data S4). Chromosomes 4M and 5M were most frequent in the population with 20 and 19 lines, respectively, containing these chromosomes. In contrast, the frequency of U-genome chromosomes was lower, with chromosomes 1U, 2U and 4U present in 1, 3 and 8 lines, respectively.
Regarding the lines with 4U chromatin, two genotypes (201106 and 201107) contained more 4U chromatin than the remaining six as they produced PCR amplicons with all of the five 4U-specific markers. The presence of a pair of 4U chromosomes in these lines was confirmed by molecular cytogenetic analysis (the chromosomes were metacentric with strong telomeric pSc119.2 signals on both arms). Becase the lines contained 42 wheat and two U chromosomes, they were identified as wheat-Ae. biuncialis 4U disomic addition lines (Fig. 3A, B). The remaining six lines (201077, 201079, 201081, 201082, 201087, and 201088) produced amplicons by only two 4U-specific markers, Ae4U15448.1 and Ae4U20523, indicating the presence of a 4U fragment (Fig. 2). Based on the specific Afa-family signals of normal wheat chromosome 1D (strong signals on subtelomeric and telomeric regions on the long and short arms, respectively), which are absent in these lines, the wheat-Aegilops translocation was identified as 1DL·1DS-4U translocation (Fig. 4A, B). In additio to the 1DL·1DS-4U translocation, Aegilops chromosome 5M was detected in five of these lines, while line 201079 contained the translocation in disomic form with no additional Aegilops chromosomes.
Molecular cytogenetic identification of Mv9kr1-Ae. biuncialis MvGB642 4U addition (A,B), 4M(4D) substitution (C,D), and 5M(5D) substitution (E,F) in the BC3F3 lines 201107, 201028 and 201195, respectively. FISH (A,C,E) was performed with Afa-family (red), pSc119.2 (green), and pTa71 (yellow) DNA repeat probes. The FISH pattern of the missing 4D (C) and 5D (E) chromosomes is shown in the inset. In the case of GISH (B,D,F), U- and M-genomic DNA probes were visualized with green and red fluorescence, respectively, while DAPI-stained wheat chromosomes were visualized in blue. Scale bar = 10 μm.
Identification of Mv9kr1-Ae. biuncialis MvGB642 1DL·1DS-4U (A,B), 1US·2BL (C,D), and 2DS·2DL-U (E,F) disomic translocations in the 201081, 201179, and 201114 BC3F3 lines, respectively. FISH (A,C,E) was performed using Afa-family (red), pSc119.2 (green), and pTa71 (yellow) repetitive DNA probes. On the GISH images (B,D,F), U- and M-chromosome fragments were visualized with green and red fluorescence, respectively, while DAPI-stained wheat chromosomes were visualized with blue. The presence of a wheat 1BL·1BS-6BS translocation (#) was detected (C,D). Scale bar = 10 μm.
The line 201179 was identified as a 1US·2BL disomic centric fusion by the presence of a satellite, a faint pTa71 signal on the U chromosome arm typical for 1US, and the pSc119.2 FISH pattern on the wheat chromosome arm. The line also carried two 4M chromosomes (Fig. 4C, D). The line 201114 carried a pair of 2DS·2DL-U translocation. As the 2U-specific marker Ae2U14986.1 (Fig. 3) produced amplicons in this line, the translocation was identified as a 2DS·2DL-2U (Fig. 4E, F).
The application of GISH in combination with the marker specific for chromosome 4M (4S-4_M_4) indicated that six of the 20 lines 4M (201027, 201028, 201045, 201061, 201174, and 201182) contained the Aegilops chromosome in disomic form. One of the six lines (201027) had 42 chromosomes. After rehybridization with DNA repeat probes (pSc119.2, Afa-family, pTa71), the presence of the 4M chromosome in disomic form was confirmed, as was the absence of wheat chromosome 4D, which has a typical hybridization pattern with pSc119.2 and Afa family repeats (Fig. 3C, D). This genotype was thus identified as a 4M(4D) disomic substitution line.
Using the 5M chromosome-specific marker BE499071, two genotypes (201195, 201196) were selected that contained two 5M but also had 42 chromosomes according to GISH. FISH confirmed the presence of disomic chromosome 5M (pTa71 signal in the telomeric region on the short arm and a relatively weak Afa-family signal in the telomeric and subtelomeric regions on the long arm) and the absence of chromosome 5D in these lines. These lines were thus identified as 5M(5D) disomic substitution lines (Fig. 3E, F). It should also be noted that instead of chromosome 7B, only one 7BL telosome was present in these lines.
GISH analysis revealed that a short M-genome segment had recombined with the terminal part of a wheat chromosome in four plants (201008, 201012, 201019 and 201020) (Figs. 2, 5A). The wheat segment showed strong telomeric Afa-family signals in the short arm, pericentromeric, weak interstitial and subtelomeric signals in the long arm, and very weak Afa-family signals in the the Aegilops segment (Fig. 5B), indicating that these lines carry a disomic 5DS·5DL-M translocation.
Molecular cytogenetic identification of Mv9kr1-Ae. biuncialis MvGB642 disomic 5DS·5DL-M (A,B) and 4DS·4DL-M translocations (C,D) in the 201020 and 201137 BC3F3 lines, respectively. FISH (A,C) was performed with Afa-family (red), pSc119.2 (green), and pTa71 (yellow) repetitive DNA probes. On the GISH images (B,D), M-chromosome fragments were visualized with red fluorescence, while DAPI-stained wheat chromosomes were visualized with blue. Asterisk: chromosome 5M. Scale bar = 10 μm.
GISH and FISH also detected a 5MS·5ML-D translocation in five lines, two of which (201044, 201053) contained the translocation in disomic form (Fig. S3 A-B). Three of the five lines (201049, 201051, 201053) with the 5MS·5ML-D translocation also carried an unidentified centric fusion between a wheat and an M-genome chromosome arm (Fig. S3A, B). These lines also contained a 4M chromosome.
Finally, a disomic 4DS·4DL-M translocation was also identified by GISH and FISH in five BC3F3 plants (not supported by markers), which also contained an extra chromosome 5M in disomic (201133, 201137, 201146) or monosomic (201127, 201143) form (Figs. 3, 5C, D).
Morphological traits of wheat-Ae. biuncialis chromosome lines
Traits related to plant and spike architecture, as well as yield, were investigated under glasshouse conditions (Table 2) for each genotype, and under field condition for addition, substitution, and the 2DS.2DL-2U translocation lines under field condition (Table S1). The spikes of the 4U addition lines have a compact structure but are significantly shorter than the spikes of the wheat parent (Mv9kr1) (Fig. 6, Table 2). These spikes have fewer spikelets than Mv9kr1, resulting in significantly lower grain numbers per spike and per plant. A similar trend was also observed under field conditions. The plant height (Ph) of the 4M(4D) substitution line is significantly shorter than that of the wheat parent under both glasshouse and field conditions. Its spikes are shorter, but only the number of spikelets per main spikes (Spms) differs significantly from wheat. The number of seeds per main spikes (Sepms) and per plant (Sepp), however, is comparable to the Mv9kr1 parent, indicating good fertility. These differences in morphological traits were eliminated under high-input field conditions (Table S1), with the exceptions of plant height and TKW.
The morphological traits of the 5M(5D) substitution line differ from those of the wheat parent in some ways, but due to the small number of samples (3 plants), they did not reach statistical significance (Fig. 6, Table 2). However, in the glasshouse, the substitution line appears to be shorter than the wheat parent, but it is longer in the field. It has compact, awnless, and slightly longer spikes than wheat, with similar or better fertility in terms of number of grains per spike and per plant in both environments.
Two of the Mv9kr1-Ae. biuncialis translocation lines (represented by single plants) deserve special attention. The T1US·2BL centric fusion is shorter than the wheat parent. It has shorter spikes with fewer spikelets than wheat, but similar fertility and seed set. T2DS·2DL-2U is also shorter than the wheat parent in both environments. Although its conical spikes have fewer spikelets, its fertility exceeds that of wheat in both glasshouse and field conditions.
Except for the good tillering ability of the T4DS·4DL-M and T5MS·5ML-DL translocations, the other translocation lines yielded less than the wheat parent in terms of yield components. At the same time, their fertility provides enough seeds to maintain these translocations and stabilize the genetic background of wheat through backcrossing. Following the propagation of these lines, the agronomic traits can be studied in details.
Discussion
Our results confirmed that sequence assemblies of the flow-sorted Aegilops chromosomes facilitate marker development to support gene introgression from Ae. biuncialis via the production of new wheat-Aegilops chromosome addition, substitution, and translocation lines.
Development of chromosome specific markers
The importance of flow-sorted chromosomes in marker development for mapping alien introgressions was previously demonstrated by Tiwari et al.46 who flow sorted 5MgS telosomes from wheat-Ae. geniculata ditelosomic addition line and identified 2,178 unique SNPs allowing the identification of T5D·5Mg translocation. The present work produced gene specific PCR markers suitable to use in laboratories equipped just by a PCR machine and ability for fragment analysis.
Only 7.7% of the 39 EST-based markers tested on the addition lines could be used to select introgression lines, compared to 50% (10 markers) for IT markers and 28% (19 markers) for cDNA-based markers. Our results are in line with previous findings that intron regions of genes have sufficient polymorphism (SNP, InDel) to develop markers suitable for use on various species with similar genomes47. COS and PLUG (PCR-based Landmark Unique Gene) markers specific for the intron regions of conserved ortholog genes have previously been used to identify wheat-Aegilops introgression lines31,48,49. These markers have also been used successfully to identify and trace alien chromosomes or to perform comparative analysis of Aegilops species with U and M genomes32,33,34.
The results showed that nearly 40%, 34.5% and 53% of the PCR-tested EST, IT and cDNA markers, respectively, produced amplicons polymorphic between the wheat (Mv9kr1) and Ae. biuncialis MvGB642 crossing partners. One reason for the low PCR validation ratio may be sequence variations between wheat (CS) or Aegilops (Ae. umbellulata AE740/03) genotypes used for in silico primer design and the parental wheat (Mv9kr1) or Ae. biuncialis (MvGB642) lines used for the PCR reactions. Another reason for the markers’ reduced validation ratio may be the use of low quality, highly fragmented draft sequence assemblies of Ae. umbellulata chromosomes43 for marker development. Combining contig assemblies from Illumina short read data with long-read sequencing technologies for scaffolding and data from Hi-C chromatin conformation capture libraries to prepare chromosome pseudomolecules would be a promising pipeline in the future to produce high quality reference assemblies for Ae. umbellulata genome, as it was demonstrated recently for Ae. sharonensis40.
We found that the combined application of molecular markers and genomic in situ hybridization (GISH) is the most optimal approach for selecting wheat genotypes with alien chromosomes in high throughput. The cost of the marker selection could be further reduced by converting PCR markers into Kompetitive Allele Specific PCR (KASP) markers, which do not require fragment analyzing step50. Combining GISH with multiple SNP genotyping systems, such as a 35 K SNP chip51 and Genotype-by-sequencing (GBS) platforms 52, can further increase the resolution and accuracy of the selection process.
The method we used enabled us to identify some translocations, such as the T2DS·2DL-2U or the T1DL·1DS-4U translocations. However, in other cases, the markers used were unable to detect the presence of cytogenetically detectable translocations (e.g., T5DS·5DL-M), implying that the generated markers were not localized on the given M genomic fragment. Additional markers specific to the aforementioned chromosomal regions will need to be developed.
Development of new wheat-Ae. biuncialis genetic materials
M-genome chromosomes were more abundant in the BC3F3 population than the U-genome chromosomes. This phenomenon may be related to the wheat-Aegilops cross-genome homology, which is higher in the case of the M genome than those of the U genome, which contains multiple re-arrangements relative to D genome of wheat32,34,43,53. The predominant pairing affinity of wheat chromosomes with M-genome chromosomes relative to U-genome chromosomes at meiotic metaphase I of wheat × Ae. biuncialis F1 hybrids 54 also indicated a closer wheat-M macrosyntenic relationship. During the crossing programs, a lower frequency of wheat-U chromosome associations resulted in abundant univalent formation of U chromosomes, which can be eliminated with higher probability from the offsprings.
The present study also indicated that chromosomes 4M and 5M were dominantly present in the population. The preferred inheritance of some alien chromosomes, which is common in wheat × Aegilops hybrid progenies, is frequently associated with the effect of gametocidal genes present in the given chromosome. Gametocidal genes ensure their existence in the host by causing selective abortion of gametes that do not carry them; as a result, they are preferentially transmitted to the offspring55,56,57,58. Gametocidal genes were detected on several chromosomes of different Aegilops species, including chromosome 4M of Ae. geniculata, another allotetraploid species with U and M genomes (UgUgMgMg)59,60,61,62 indicating that chromosome 4M of Ae. biuncialis may have a similar effect. The gametocide effect of chromosome 4M was also supported by the frequent observation of aberrant chromosomes resulted from random chromosome breakages.
From breeding point of view, gametocidal genes have received special attention as several cytogenetic materials have been developed using the gametocidal system, including a set of deletion lines for hexaploid wheat56 and wheat-alien introgression lines63,64,65. Kwiatek et al.62 produced wheat-rye translocations using triticale lines containing the 4Mg chromosome of the Ae. geniculata in the monosomal state. However, the confirmation of the presence of the gametocidal gene in our wheat-Ae. biuncialis 4Mb and/or 5Mb addition lines needs further evidences.
The present work developed new hexaploid wheat-Ae. biuncialis 4U disomic addition, as well as 4M(4D) and 5M(5D) disomic substitution lines. We also produced six wheat-Ae. biuncialis chromosome translocations (T4DS·4DL-M, T5DS·5DL-M, T5MS·5ML-DL, T1US·2BL, T2DS·2DL-2U, T1DL·1DS-4U), which occurred in disomic form in separate lines. These cytogenetic stocks significantly contribute to increase wheat genetic variability.
The significant macro collinearity of 4M and 5M chromosomes with the chromosomes of wheat homeologous groups 4 and 5, respectively43, implies that gene composition and order of these Aegilops and wheat chromosomes are similar. In line with this, the similar spike structure and fertility of the 4M(4D) and 5M(5D) substitution lines to the wheat parent also indicate that the 4M and 5M chromosomes compensate well for the loss of genes localized on the 4D and 5D chromosomes, respectively. In this respect, several QTLs have been mapped to wheat homeologous groups 4 and 5 chromosomes, which play a key role in crop yield formation 66, environmental adaptation67,68,69 and abiotic stress tolerance70. QTLs associated with yield components, including tillering ability and grain number per spike have been mapped on 4D and 5D chromosomes71,72 and on chromosomes 4A, 5B, and 5D73,74,75, respectively.
Plant height also has an impact on wheat yield, and exploiting the role of dwarfing or reduced height (Rht) loci resulted in one of the most significant breakthroughs in modern plant breeding during the Green Revolution76 of the twentieth century. The Rht1 and Rht2, are two of the most commonly used dwarfing genes in wheat, have been located on the 4BS and 4DS chromosome arms, respectively77,78. In the present study, replacement of the chromosome 4D with 4M caused a 10 cm reduction in plant height in the substitution line, which may be attributed to the effect of a putative Aegilops Rht gene variant present on the 4M chromosome.
The regulation of flowering time is one of the crucial elements in wheat's adaptation to various ecogeographical environments79,80. The most essential factors in determining flowering time in cereals are the Vrn genes that regulate the need for vernalization (the need for the cold period that triggers the transition from the vegetative to the generative phase). The major Vrn genes in wheat are Vrn-A1, Vrn-B1, and Vrn-D1, which are located on the long arms of chromosomes 5A, 5B, and 5D, respectively67,81,82.
Our previous research on wheat-Ae. geniculata addition lines suggests that chromosomes 5Ug and 5Mg have a positive effect on the grain dietary fiber composition and β-glucan content of hexaploid wheat17, implying that Mv9kr1-Ae. biuncialis 5Mb(5D) substitution line could also be useful gene source for the production of functional foods.
These examples indicates that new allelic variants of agriculturally important QTLs and genes can be introduced into wheat by ph-induced homoeologous recombination of chromosomes 4D and 5D with the collinear chromosomes 4M and 5M, respectively. The future use of ph1b mutant in wheat Mv9kr1 line may allow for the production of 4D/4M and 5D/5M recombinations without changing the wheat genetic background54.
Conclusion
The sequences of flow sorted chromosomes allowed the development of a significant amount of gene-based markers specific for the U and M genome chromosomes of Aegilops species. Using markers PCR-validated on the parental genotypes and wheat-Aegilops chromosome addition lines, we selected new wheat-Ae. biuncialis disomic 4U addition, 4M(4D), and 5M(5D) substitutions, as well as several introgression lines. The morphology and fertility of the spikes indicated that the Aegilops 4M or 5M chromosomes compensated well for the loss of wheat chromosomes 4D and 5D, respectively. The new cytogenetic stocks may serve as a suitable genetic resource to introgress wild alleles of key genes determining some important agronomic traits into wheat through chromosome engineering.
Methods
Plant material
Ae. umbellulata (2n = 2x = 14, UU) accession AE740/03 provided by the Institute of Plant Genetics and Crop Plant Research (Gatersleben, Germany) and Ae. comosa (2n = 2x = 14, MM) accession MvGB1039 maintained in the Martonvásár Cereal Gene Bank (Martonvásár, Hungary) were used for chromosome sorting, sequencing and marker design.
The chromosomal positions of the markers were validated by PCR on the following wheat-Aegilops chromosome addition lines: CS-Ae. umbellulata 1U, 2U, 4U, 5U, 6U, 7U (kindly provided by Dr. Steve Reader (John Innes Centre, Norwich, UK).); Mv9kr1-Ae. biuncialis MvGB642 3U22 and CS-Ae. geniculata 1M–7M addition lines 57 were kindly provided by Dr. Bernd Friebe (Kansas State University, Manhattan, Kansas, USA).
The Ae. biuncialis (2n = 4x = 28, UbUbMbMb) accession MvGB642 was collected from an area 1160 m above the sea level near Slinfah, Syria.
The winter wheat (T. aestivum) line Mv9kr1 carrying a recessive crossability allele kr1 83 was crossed with Ae. biuncialis MvGB642 to develop a BC3F3 population (Fig. S2). The BC3F3 progeny plants were characterized by GISH, FISH, and molecular markers.
In situ hybridization
Mitotic chromosome preparations were made from the root tips of BC3F3 plants according to Molnár et al. 42. Genomic DNA was isolated using the Quick Gene-Mini80 system (FujiFilm, Osaka, Japan) according to the manufacturer’s instructions. Total genomic DNA from Ae. umbellulata (2n = 2x = 14; UU) was labeled with biotin (BioPrime™ DNA Labeling System, Invitrogen™, Carlsbad, USA) and used as the U-genomic probe. Total genomic DNA from Ae. comosa (2n = 2x = 14; MM) was labeled with digoxigenin-11-dUTP (Roche) using the Random Primed DNA Labeling Kit (Roche) following the manufacturer’s instructions, and used as the M-genomic probe. The repetitive DNA sequences of Afa-family and pSc119.2 were amplified using PCR and labelled with digoxigenin-16-dUTP (Roche) and biotin-11-dUTP (Roche), respectively 84,85. The 18S-5.8S-26S rDNA clone pTa71 was labeled in equal proportions with biotin-11-dUTP and digoxigenin-16-dUTP by nick translation 86. Digoxigenin and biotin were detected using anti-digoxigenin-rhodamine Fab fragments (Roche) and streptavidin–FITC (Roche), respectively.
Sequential GISH and FISH on BC3F3 lines were performed as described by Molnár et al. 26. Briefly, the hybridization solution (25 μl per slide) for GISH experiments contained 70 ng of each U- and M-genomic labeled probe together with unlabeled wheat genomic DNA to block unspecific hybridization at a concentration of 50× that of the probe. After the hybridization (42 °C, overnight) and post treatments, the chromosomes were counterstained with 2 μg/ml DAPI (4’-6-diamino-2-phenylindole) and mounted in antifade solution. Following GISH, FISH was performed using probes for DNA repeats Afa-family, pSc119.2, and pTa71. Fluorescence signals were investigated by an AxioImager M2 epifluorescence microscope (Zeiss, Oberkochen, Germany) equipped with filter sets for detecting DAPI, FITC, and Rhodamine signals. Images were captured with a Zeiss AxioCam MRm CCD camera and processed with Zeiss AxioVision 4.8.2. software (Zeiss).
Flow cytometric chromosome sorting, sequencing and marker design
Flow cytometric sorting of mitotic metaphase chromosomes from Ae. comosa MvGB1039 and Ae. umbellulata AE740/03 was carried out as previously described by Molnár et al. 34 and Said et al. 87. The chromosomal DNA was purified according to Šimková et al. 88, sequenced on an Illumina NovaSeq 6000 (Ae. comosa) or HiSeq2000 (Ae. umbellulata), and assembled as described by Said et al. 43. In the present study, assembled chromosome contigs of Ae. umbellulata AE740/03 (https://doi.org/10.5061/dryad.70rxwdbwc) and Ae. comosa MvGB1039 (https://doi.org/10.5061/dryad.wpzgmsbk9) were used to design molecular markers.
Three strategies were used to design PCR markers specific for the U- and M-chromosomes of Aegilops. For the first strategy (1) ESTs specific for the D genome chromosomes of hexaploid wheat were selected from publicly available database (https://wheat.pw.usda.gov/cgi-bin/westsql/map_locus.cgi). A total of 4514 D-genome specific wheat EST sequences (1D: 633; 2D: 742; 3D: 719; 4D: 586; 5D: 622; 6D: 485; 7D: 727) were aligned to the genomic sequences of the Ae. umbellulata AE740/03 (Data S1) using BLASTn with the program Ragged Genes Genome Explorer (version 2.2.24) which is a graphic interface for NCBI’s BLAST command line tool 89. BLASTn hits that met certain criteria (InDel polymorphism ≥ 5 bp and wheat-Aegilops homology ≥ 70%) were selected, and identification of InDels to design primers to the polymorphic regions was carried out using pairwise alignments of the selected ESTs and Aegilops contigs using UGENE (v.1.27.0) and Primer3. The selected ESTs polymorphic between wheat and Aegilops together with their PCR primers were summarized in Data S2.
The second marker development strategy (2) was based on the polymorphism between the intron regions of the Ae. umbellulata genes and the homologous wheat genes (Intron Targeting markers: IT markers). During marker development, we used annotated gene models of Ae. umbellulata. For automated primer design, bedtools (v.2.26.0), primer3 (v.2.3.6) and a custom R script with the package devtools were used. In the first step, BLASTn was used to search wheat homologues of the Ae. umbellulata genes (Aegilops genes were used as query sequences for BLASTn searches in the wheat reference genome sequence [RefSeq v1.037], and then the positions of the introns were determined). The polymorphic 600 bp region of genes that contains the last 100 bp of the first exon and the first 500 bp of the first intron (> 500 bp) was selected and saved in a multifasta file. Primer pairs were generated for these sequences. The forward primer was located on the first 100 bp (exon-specific) region and the reverse primer was located on the subsequent 500 bp (intron-specific) region. A total of 84 primer pairs (12 markers per chromosome) were randomly selected. The oligonucleotide sequences were synthesized by Integrated DNA Technologies (Coralville, Iowa, USA).
The third marker development strategy was based on a polymorphism between cDNA sequences of wheat genes and Ae. umbellulata and Ae. comosa sequences. In a previous study, Said et al.43 mapped forty-three ortholog genes on M- and U-genome progenitors, Ae. comosa and Ae. umbellulata, respectively, using single-gene FISH with fluorescently labeled cDNA sequences as probes. In order to validate the cytogenetic maps, PCR markers were developed for the cytogenetically mapped genes. From this set of markers, 19 were selected and used to identify or confirm the cytological results in the BC3F3 plants.
The synthesized primer pairs for both EST-based markers and IT markers were tested by PCR on the parental genotypes Mv9kr1, Ae. biuncialis MvGB642, and Ae. umbellulata AE740/03. The primer pairs that produced Aegilops-specific amplicons were tested further by PCR on wheat Aegilops addition lines (CS-Ae. umbellulata 1U, 2U, 4U, 5U, 6U, 7U, Mv9kr1-Ae. biuncialis MvGB642 3U, and CS-Ae. geniculata 1M–7M addition lines) representing complete sets of U- and M-genome chromosomes in wheat genetic background, and on the Mv9kr1-Ae. biuncialis MvGB642 amphiploid. The reaction mixture (15 μl) containing 0.2 μmol/l of forward and reverse primers, as well as the thermal reaction profile used for the PCR reactions, were described by Molnár et al. 90. PCR amplicons were detected along with a 75–400 bp Range DNA Ladder using a Fragment Analyzer Automated CE System (Advanced Analytical Technologies, Ames, USA) and analyzed with PROsize v2.0 software (Advanced Analytical Technologies).
Statistical analysis
The wheat (Mv9kr1)-Ae. biuncialis MvGB642 addition, substitution and translocation lines developed by the present study were grown in glasshouse as described by Rakszegi et al. 17.
The wheat (Mv9kr1)-Ae. biuncialis MvGB642 4U addition, 4M(4D) and 5M(5D) substitution, and 2DS·2DL-2U translocation lines were also grown in a high-input field nursery (Bulgárföld, Martonvásár, Hungary, geographic coordinates: 47°19′39′′ N, 18°47′01′′ E) in the 2022–2023 season. In the field experiment, 250 grains of each genotype were sown in 2 m2 plots (5 × 2 m rows; 50 grains per row, row distance: 0.15 m). The trial plots were machine-planted (HEGE-80 plot driller) and hand-harvested. The clayey chernozem texture soil (pH 7.25) contained 2.8% w/w humus, 210 mg/kg P2O5 and 210 mg/kg K2O.
The results for the glasshouse experiments are the means ± standard deviation of three [5M(5D)], five [4M(4D)] and ten (Mv9kr1) measurements per genotype for plant height (Ph), tillering (T), length of main spike (Lms), spikelets/main spike (Spms), seeds/main spike (Sepms), seeds/plant (Sepp) and thousand kernel weight (TKW). Differences between the substitution lines and the wheat parental line Mv9kr1 were determined by means of Student’s t test for paired data at the P = 0.05 and P = 0.01 significance levels. In the case of field experiments, the results are the means ± standard deviations of ten plants per genotypes. The significant differences between the genotypes were calculated using the Tukey’s post hoc test of the IBM SPSS Statistics 20.0 software (SPSS Inc., Chicago, IL, USA).
Data availability
All data is contained within the present article and the supplementary material.
References
Tadesse, W. et al. Genetic gains in wheat breeding and its role in feeding the world. Crop Breed. Genet. Genome 1, e190005 (2019).
Friebe, B., Jiang, J., Raupp, W. J., McIntosh, R. A. & Gill, B. S. Characterization of wheat-alien translocations conferring resistance to diseases and pests: Current status. Euphytica 91, 59–87 (1996).
van Slageren, M. W. Wild Wheats. A Monograph of Aegilops L. and Amblyopyrum (Jaub. & Spach) Eig (Poaceae). (Wageningen Agricultural University/ICARDA, 1994).
Ivanizs, L. et al. Unlocking the genetic diversity and population structure of a wild gene source of wheat, Aegilops biuncialis Vis., and its relationship with the heading time. Front. Plant Sci. 10, 1531 (2019).
Ivanizs, L. et al. Identification of new QTLs for dietary fiber content in Aegilops biuncialis. Int. J. Mol. Sci. 23, 3821. https://doi.org/10.3390/ijms23073821 (2022).
Molnár, I. et al. Physiological and morphological responses to water stress in Aegilops biuncialis and Triticum aestivum genotypes with differing tolerance to drought. Funct. Plant Biol. 31, 1149–1159 (2004).
Dulai, S. et al. Wheat-Aegilops biuncialis amphiploids have efficient photosynthesis and biomass production during osmotic stress. J. Plant Phys. 171, 509–517 (2014).
Darko, E. et al. Addition of Aegilops biuncialis chromosomes 2M or 3M improves the salt tolerance of wheat in different way. Sci. Rep. 10, 22327. https://doi.org/10.1038/s41598-020-79372-1 (2020).
Makkouk, K. M., Comeau, A. & Ghulam, W. Resistance to barley yellow dwarf luteovirus in Aegilops species. Can. J. Plant Sci. 74, 631–634 (1994).
Li, H. et al. Discovery of powdery mildew resistance gene candidates from Aegilops biuncialis chromosome 2Mb based on transcriptome sequencing. PLoS One 14, e0220089. https://doi.org/10.1371/journal.pone.0220089 (2019).
Damania, A. B. & Pecetti, L. Variability in a collection of Aegilops species and evaluation for yellow rust resistance at two locations in Northern Syria. J. Genet. Breed. 44, 97–102 (1990).
Dimov, A., Zaharieva, M. & Mihova, S. Rust and powdery mildew resistance in Aegilops accessions from Bulgaria. In Biodiversity and Wheat Improvement (ed. Damania, A. B.). 165–169 (Wiley, 1993).
Kwiatek, M., Błaszczyk, L., Wiśniewska, H. & Apolinarska, B. Aegilops-Secale amphiploids: Chromosome categorisation, pollen viability and identification of fungal disease resistance genes. J. Appl. Genet. 53, 37–40 (2012).
Olivera, P. D., Rouse, M. N. & Jin, Y. Identification of new sources of resistance to wheat stem rust in Aegilops spp. in the tertiary genepool of wheat. Front. Plant Sci. 9, 1719 (2018).
Zhou, J. et al. Characterization of a new wheat-Aegilops biuncialis addition line conferring quality-associated HMW glutenin subunits characterization of a wheat. Genet. Mol. Res. 13, 660–669 (2014).
Farkas, A. et al. Increased micronutrient content (Zn, Mn) in the 3Mb(4B) wheat-Aegilops biuncialis substitution and 3Mb.4BS translocation. Genome 57, 61–67 (2014).
Rakszegi, M. et al. Addition of Aegilops U and M chromosomes affects protein and dietary fiber content of wholemeal wheat flour. Front. Plant Sci. 8, 1529. https://doi.org/10.3389/fpls.2017.01529 (2017).
Schneider, A., Molnár, I. & Molnár-Láng, M. Utilisation of Aegilops (goatgrass) species to widen the genetic diversity of cultivated wheat. Euphytica 163, 1–19 (2008).
Kishii, M. An update of recent use of Aegilops species in wheat breeding. Front. Plant Sci. 10, 585. https://doi.org/10.3389/fpls.2019.00585 (2019).
Zhang, P. et al. Wheat-Aegilops introgressions. In Alien Introgression in Wheat (eds. Molnár-Láng, M., Ceoloni, C. & Doležel, J.). 221–243 (Springer, 2015). https://doi.org/10.1007/978-3-319-23494-6.
Logojan, A. A. & Molnár-Láng, M. Production of Triticum aestivum–Aegilops biuncialis chromosome additions. Cereal Res. Commun. 28, 221–228 (2000).
Schneider, A., Linc, G., Molnár, I. & Molnár-Láng, M. Molecular cytogenetic characterization of Aegilops biuncialis and its use for the identification of 5 derived wheat-Aegilops biuncialis disomic addition lines. Genome 48, 1070–1082 (2005).
Tan, F. et al. Characterization of a new synthetic wheat-Aegilops biuncialis partial amphiploid. J. Biotechnol. 8, 3215–3218 (2009).
Zhao, H. et al. Comparative study on drought tolerance of wheat and wheat-Aegilops biuncialis 6Ub addition lines. J. Food Agricult. Environ. 11, 1046–1052 (2013).
Song, L. et al. Molecular cytogenetic identification of wheat-Aegilops biuncialis 5Mb disomic addition line with tenacious and black glumes. Int. J. Mol. Sci. 21, 4053. https://doi.org/10.3390/ijms21114053 (2020).
Molnár, I., Benavente, E. & Molnár-Láng, M. Detection of intergenomic chromosome rearrangements in irradiated Triticum aestivum –Aegilops biuncialis amphiploids by multicolour genomic in situ hybridization. Genome 52, 156–165 (2009).
Kuraparthy, V. et al. Characterization and mapping of cryptic alien introgression from Aegilops geniculata with new leaf rust and stripe rust resistance genes Lr57 and Yr40 in wheat. Theor. Appl. Genet. 114, 1379–1389 (2007).
Peil, A. et al. The application of wheat microsatellites to identify disomic Triticum aestivum–Aegilops markgrafii addition lines. Theor. Appl. Genet. 96, 138–146 (1998).
Schneider, A., Molnár, I. & Molnár-Láng, M. Selection of U and M genome-specific wheat SSR markers using wheat-Aegilops biuncialis and wheat-Ae. geniculata addition lines. Euphytica 175, 357–364 (2010).
Nagy, E. D., Molnár, I., Schneider, A., Kovács, G. & Molnár-Láng, M. Characterization of chromosome-specific S-SAP markers and their use in studying genetic diversity in Aegilops species. Genome 49, 289–296 (2006).
Howard, T. et al. Identification of a major QTL controlling the content of B-type starch granules in Aegilops. J. Exp. Bot. 62, 2217–2228 (2011).
Molnár, I. et al. Syntenic relationships between the U and M genomes of Aegilops, wheat and the model species Brachypodium and rice as revealed by COS markers. PLoS One 8, e70844. https://doi.org/10.1371/journal.pone.0070844 (2013).
Liu, C. et al. Characterization, identification and evaluation of a set of wheat-Aegilops comosa chromosome lines. Sci. Rep. 9, 4773. https://doi.org/10.1038/s41598-019-41219-9 (2019).
Molnár, I. et al. Dissecting the U, M, S and C genomes of wild relatives of bread wheat (Aegilops spp.) into chromosomes and exploring their synteny with wheat. Plant J. 88, 452–467 (2016).
Men, W. et al. Mapping of the novel powdery mildew resistance gene Pm2Mb from Aegilops biuncialis based on ph1b-induced homoeologous recombination. Theor. Appl. Genet. 135, 2993–3003 (2022).
Zwyrtková, J., Šimková, H. & Doležel, J. Chromosome genomics uncovers plant genome organization and function. Biotechnol. Adv. 46, 107659 (2021).
International Wheat Genome Sequencing Consortium (IWGSC), et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361, eaar7191 (2018).
Martis, M. M. et al. Reticulate evolution of the rye genome. Plant Cell 25, 3685–3698 (2013).
International Barley Genome Sequencing Consortium (IBGSC), et al. A physical, genetic and functional sequence assembly of the barley genome. Nature 491, 711–716 (2012).
Yu, G. et al. Aegilops sharonensis genome-assisted identification of stem rust resistance gene Sr62. Nat. Commun. 13, 1607. https://doi.org/10.1038/s41467-022-29132-8 (2022).
Zwyrtková, J. et al. Draft sequencing crested wheatgrass chromosomes identified evolutionary structural changes and genes and facilitated the development of SSR markers. Int. J. Mol. Sci. 23, 3191. https://doi.org/10.3390/ijms23063191 (2022).
Molnár, I., Cifuentes, M., Schneider, A., Benavente, E. & Molnár-Láng, M. Association between simple sequence repeat-rich chromosome regions and intergenomic translocation breakpoints in natural populations of allopolyploid wild wheats. Ann. Bot. 107, 65–76 (2011).
Said, M. et al. Development of DNA markers from physically mapped loci in Aegilops comosa and Aegilops umbellulata using single-gene FISH and chromosome sequences. Front. Plant. Sci. 12, 689031. https://doi.org/10.3389/fpls.2021.689031 (2021).
Said, M. et al. Flow karyotyping of wheat-Aegilops additions facilitate dissecting the genomes of Ae. biuncialis and Ae. geniculata into individual chromosomes. Front. Plant. Sci. 13, 1017958. https://doi.org/10.3389/fpls.2022.1017958 (2022).
Tiwari, V. K. et al. Exploring the tertiary gene pool of bread wheat: sequence assembly and analysis of chromosome 5Mg of Aegilops geniculata. Plant J. 84, 733–746 (2015).
Tiwari, V. K. et al. SNP discovery for mapping alien introgressions in wheat. BMC Genomics 15, 273. https://doi.org/10.1186/1471-2164-15-273 (2014).
Poczai, P. et al. Advances in plant gene-targeted and functional markers: A review. Plant Methods 9, 6 (2013).
Burt, C. & Nicholson, P. Exploiting colinearity among grass species to map the Aegilops ventricosa-derived Pch1 eyespot resistance in wheat and establish its relationship to Pch2. Theor. Appl. Genet. 123, 1387–1400 (2011).
Gong, W. et al. Agronomic traits and molecular marker identification of wheat-Aegilops caudata addition lines. Front. Plant. Sci. 8, 1743. https://doi.org/10.3389/fpls.2017.01743 (2017).
King, J. et al. A step change in the transfer of interspecific variation into wheat from Amblyopyrum muticum. Plant Biotechnol. J. 15, 217–226 (2016).
Grewal, S. et al. Development of Wheat-Aegilops caudata introgression lines and their characterization using genome-specific KASP markers. Front. Plant. Sci. 11, 606. https://doi.org/10.3389/fpls.2020.00606 (2020).
Ando, K. et al. Introgression of a novel Ug99-effective stem rust resistance gene into wheat and development of Dasypyrum villosum chromosome-specific markers via genotyping-by-sequencing (GBS). Plant Dis. 103, 1068–1074 (2019).
Zhang, H., Jia, J., Gale, M. D. & Devos, K. M. Relationships between the chromosomes of Aegilops umbellulata and wheat. Theor. Appl. Genet. 96, 69–75 (1998).
Türkösi, E. et al. Transfer of the ph1b deletion chromosome 5B from Chinese Spring wheat into a winter wheat line and induction of chromosome rearrangements in wheat-Aegilops biuncialis hybrids. Front. Plant Sci. 13, 875676. https://doi.org/10.3389/fpls.2022.875676 (2022).
Endo, T. R. Gametocidal chromosomes and their induction of chromosome mutations in wheat. Jpn. J. Genet. 65, 135–152 (1990).
Endo, T. R. & Gill, B. S. The deletion stocks of common wheat. J. Hered. 87, 295–307 (1996).
Friebe, B. R., Tuleen, N. A. & Gill, B. S. Development and indentification of a complete set of Triticum aestivum–Aegilops geniculata chromosome addition lines. Genome 42, 374–380 (1999).
Endo, T. R. Gametocidal genes. In Alien Introgression in Wheat: Cytogenetics, Molecular Biology, and Genomics. 121–131 (2015). https://doi.org/10.1007/978-3-319-23494-6_5/COVER.
Tsujimoto, H. & Tsunewaki, K. Gametocidal genes in wheat and its relatives. I. Genetic analyses in common wheat of a gametocidal gene derived from Aegilops speltoides. Can. J. Genet. Cytol. 26, 78–84 (1984).
Endo, T. R. The gametocidal chromosome as a tool for chromosome manipulation in wheat. Chrom. Res. 15, 67–75 (2007).
Kwiatek, M. et al. Transmission of the Aegilops ovata chromosomes carrying gametocidal factors in hexaploid triticale (×Triticosecale Wittm.) hybrids. J. Appl. Genet. 57, 305–315 (2016).
Kwiatek, M. T. et al. Gametocidal factor transferred from Aegilops geniculata Roth can be adapted for large-scale chromosome manipulations in cereals. Front. Plant. Sci. 8, 409. https://doi.org/10.3389/fpls.2017.00409 (2017).
Shi, F. & Endo, T. R. Production of wheat–barley disomic addition lines possessing an Aegilops cylindrica gametocidal chromosome. Genes Genet. Syst. 72, 243–248 (1997).
Shi, F. & Endo, T. R. Genetic induction of structural changes in barley chromosomes added to common wheat by a gametocidal chromosome derived from Aegilops cylindrica. Genes Genet. Syst. 74, 49–54 (1999).
Shi, F. & Endo, T. R. Genetic induction of chromosomal rearrangements in barley chromosome 7H added to common wheat. Chromosoma 109, 358–363 (2000).
Guan, P. et al. Global QTL analysis identifies genomic regions on chromosomes 4A and 4B harboring stable loci for yield-related traits across different environments in wheat (Triticum aestivum L.). Front. Plant Sci. 9, 529. https://doi.org/10.3389/fpls.2018.00529 (2018).
Galiba, G., Quarrie, S. A., Sutka, J., Morgounov, A. & Snape, J. W. RFLP mapping of the vernalization (Vrn1) and frost resistance (Fr1) genes on chromosome 5A of wheat. Theor. Appl. Genet. 90, 1174–1179 (1995).
Snape, J. W. et al. Mapping frost tolerance loci in wheat and comparative mapping with other cereals. Acta Agron. Hung. 45, 265–270 (1997).
Tóth, B., Galiba, G., Fehér, E., Sutka, J. & Snape, J. W. Mapping genes affecting flowering time and frost resistance on chromosome 5B of wheat. Theor. Appl. Genet. 107, 509–514 (2003).
Cattivelli, L. et al. Chromosome regions and stress-related sequences involved in resistance to abiotic stress in Triticeae. Plant Mol. Biol. 48, 649–665 (2002).
Liu, G. et al. Mapping QTLs of yield-related traits using RIL population derived from common wheat and Tibetan semi-wild wheat. Theor. Appl. Genet. 127, 2415–2432 (2014).
Wang, Z. et al. Identification and validation of novel low-tiller number QTL in common wheat. Theor. Appl. Genet. 129, 603–612 (2016).
Börner, A. et al. Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor. Appl. Genet. 105, 921–936 (2002).
Gao, F. et al. Genome-wide linkage mapping of QTL for yield components, plant height and yield-related physiological traits in the Chinese wheat cross Zhou 8425B/Chinese spring. Front. Plant Sci. 6, 1099. https://doi.org/10.3389/fpls.2015.01099 (2015).
Guo, J. et al. Association and validation of yield-favored alleles in Chinese cultivars of common wheat (Triticum aestivum L.). PLoS One 10, 1–18 (2015).
Hedden, P. The genes of the green revolution. Trends Genet. 19, 5–9 (2003).
Peng, J. et al. ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400, 256–261 (1999).
Achard, P. et al. Integration of plant responses to environmentally activated phytohormonal signals. Science 311, 91–94 (2006).
Kiss, T. et al. Allele frequencies in the VRN-A1, VRN-B1 and VRN-D1 vernalization response and PPD-B1 and PPD-D1 photoperiod sensitivity genes, and their effects on heading in a diverse set of wheat cultivars (Triticum aestivum L.). Mol. Breed. 34, 297–310 (2014).
Kiss, T. et al. Effects of ambient temperature in association with photoperiod on phenology and on the expressions of major plant developmental genes in wheat (Triticum aestivum L.). Plant Cell Environ. 40, 1629–1642 (2017).
Law, C. N., Worland, A. J. & Giorgi, B. The genetic control of ear-emergence time by chromosomes 5A and 5D of wheat. Heredity 36, 49–58 (1976).
Barrett, B., Bayram, M. & Kidwell, K. Identifying AFLP and microsatellite markers for vernalization response gene Vrn-B1 in hexaploid wheat using reciprocal mapping populations. Plant Breed. 121, 400–406 (2002).
Molnár-Láng, M., Linc, G. & Sutka, J. Transfer of the recessive crossability allele kr1 from Chinese Spring into the winter wheat variety Martonvásári 9. Euphytica 90, 301–305 (1996).
Nagaki, K., Tsujimoto, H., Isono, K. & Sasakuma, T. Molecular characterization of a tandem repeat, Afa family, and its distribution among Triticeae. Genome 38, 479–486 (1995).
Contento, A., Heslop-Harrison, J. S. & Schwarzacher, T. Diversity of a major repetitive DNA sequence in diploid and polyploid Triticeae. Cytogenet. Genome Res. 109, 34–42 (2005).
Gerlach, W. L. & Bedbrook, J. R. Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res. 7, 1869–1885 (1979).
Said, M. et al. Uncovering homeologous relationships between tetraploid Agropyron cristatum and bread wheat genomes using COS markers. Theor. Appl. Genet. 132, 2881–2898 (2019).
Šimková, H. et al. Coupling amplified DNA from flow-sorted chromosomes to high-density SNP mapping in barley. BMC Genomics 9, 294. https://doi.org/10.1186/1471-2164-9-294 (2008).
Zhang, Z., Schwartz, S., Wagner, L. & Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 7, 203–214. https://doi.org/10.1089/10665270050081478 (2000).
Molnár, I. et al. Flow cytometric chromosome sorting from diploid progenitors of bread wheat, T. urartu, Ae. speltoides and Ae. tauschii. Theor. Appl. Genet. 127, 1091–1104 (2014).
Acknowledgements
The authors would like to thank Fanni Tóth and Ildikó Lakner Könyvesné for the excellent technical assistance. Zdeňka Dubská, Romana Šperková, and Jitka Weiserová are also acknowledged for their assistance with chromosome sorting. This work has been supported by the Hungarian National Research, Development and Innovation Office (K135057 and TKP2021-NKTA-06), by ERDF project Plants as a Tool for Sustainable Global Development (No.CZ.02.1.01/0.0/0.0/16_019/0000827), and the Marie Curie Fellowship Grant award AEGILWHEAT (H2020-MSCA-IF-2016-746253).
Author information
Authors and Affiliations
Contributions
Conceptualization I.M.; methodology I.M., M.S., N.B., J.B.; investigation A.F., E.T., L.I., E.T., E.G., M.S., É.D., K.S.P., K.K., P.K.; resources I.M., É.S.Z., K.H.; data curation I.M., N.B., J.B., É.D., T.S.; writing-original draft preparation A.F., I.M., É.S.Z.; writing-review A.F., I.M., M.S., J.D.; visualization E.T., L.I., I.M.; project administration I.M.; funding administration I.M., J.D. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Farkas, A., Gaál, E., Ivanizs, L. et al. Chromosome genomics facilitates the marker development and selection of wheat-Aegilops biuncialis addition, substitution and translocation lines. Sci Rep 13, 20499 (2023). https://doi.org/10.1038/s41598-023-47845-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-47845-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.