Abstract
Colorectal cancer (CRC) involves epigenetic alterations. Irregular gene-methylation alteration causes and advances CRC tumor growth. Detecting differentially methylated genes (DMGs) in CRC and patient survival time paves the way to early cancer detection and prognosis. However, CRC data including survival times are heterogeneous. Almost all studies tend to ignore the heterogeneity of DMG effects on survival. To this end, we utilized a sparse estimation method in the finite mixture of accelerated failure time (AFT) regression models to capture such heterogeneity. We analyzed a dataset of CRC and normal colon tissues and identified 3406 DMGs. Analysis of overlapped DMGs with several Gene Expression Omnibus datasets led to 917 hypo- and 654 hyper-methylated DMGs. CRC pathways were revealed via gene ontology enrichment. Hub genes were selected based on Protein–Protein-Interaction network including SEMA7A, GATA4, LHX2, SOST, and CTLA4, regulating the Wnt signaling pathway. The relationship between identified DMGs/hub genes and patient survival time uncovered a two-component mixture of AFT regression model. The genes NMNAT2, ZFP42, NPAS2, MYLK3, NUDT13, KIRREL3, and FKBP6 and hub genes SOST, NFATC1, and TLE4 were associated with survival time in the most aggressive form of the disease that can serve as potential diagnostic targets for early CRC detection.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC), the third most common cancer worldwide, is a group of diseases characterized by genetic and epigenetic changes1,2. Despite being the second leading cause of cancer-related deaths, less attention has been paid to early detection due to the fact that patients do not adhere to invasive screening tests such as colonoscopy3. It has been shown that epigenetic alterations in solid and liquid biopsies can be used for early detection and thus prognosis and effective treatment4. DNA methylation at CpG sites (5mc) is an epigenetic mark that regulates gene expression through transcriptional silencing5. Aberrant DNA methylation plays a crucial role in the pathogenesis and progression of CRC and has emerged as a promising diagnostic marker for the disease6. In particular, aberrant DNA methylation can impact genes where their inactivation may exacerbate tumor formation through the induction of genomic instability or by directly silencing the methylated gene7.
Much research has been done to develop comprehensive panels of biomarkers based on DNA methylation that can facilitate accurate diagnosis of CRC8. While the genes SEPT9, NDRG4, and BMP3 are FDA-approved for CRC9,10, there are many other genes such as APC, SFRP1, TFPI2, and VIM that have not yet been approved8.
In order to detect and validate genes that are potential CRC biomarkers, the following steps should be taken. Firstly, a panel of biomarkers must be developed using accurate statistical methods with a deep understanding of the underlying biology of the disease and the molecular mechanisms that drive them. Secondly, the significant biomarkers must be validated via in silico validation using several other datasets; and thirdly, the effectiveness of top candidate biomarkers in improving patient health should be verified using survival models. Lack of adequate precision in each of the above steps leads to misleading conclusions. Among others, two issues affect precision: removing genomic positions with missing values or low read-depth and ignoring the heterogeneity of DMG effects on survival times.
To accurately predict the differentially methylated profiles in CRC, one must consider all biological and environmental factors such as dietary11, aging12, and hazardous behaviors13 (e.g., smoking), among others. Such factors are often ignored by most studies when predicting methylation profiles. In addition, methylation data always suffer from heavy missing values that can affect subsequent analyses. For instance, 68% of CpG sites have missing values in at least one sample in our dataset (Section 2). Almost all DNA methylation pipelines, except a few such as the DMCHMM method14, filter out such positions from the analysis. We used DMCHMM to not only account for extra covariates but also efficiently impute the missing values.
Having identified the differentially methylated genes (DMG) associated with CRC and validating them, it is crucial to identify their underlying signaling pathways that regulate gene expression15,16. The main known CRC pathways are Wnt17, MAPK18, TGF-\(\beta\)19, and TP5320. Although significant progress has been made in understanding the biology of CRC, there are still many unknown pathways and mechanisms involved in this disease. Identification of hub genes, also known as driver genes is the next step in the analysis of biomarker detection. Hub genes play a critical role in regulating several genes in the biological network and have the potential to be regarded as therapeutic targets in CRC21.
In the next step, the relationship between identified DMGs and the survival time of CRC patients should be evaluated. Most studies employ a limited panel of biomarkers selected through conventional univariate Cox proportional hazard regression models and overlook the potential effects of the rest of the biomarkers22,23,24. In a recent study25, the Cox-LASSO survival model was used to account for a larger set of biomarkers but ignored the heterogeneity of covariate effects. To the best of our knowledge, none of the studies have taken into account the heterogeneity of DMG effects on survival time. To address this problem, one may use the sparse estimation method in the finite mixture of accelerated failure time (AFT) regression models26. Prior to this step, it is common to screen the number of genes to a manageable magnitude. This process can be done by selecting the top highly correlated genes with survival time of the patients using the correlation-adjusted scoring method27.
This study aimed to identify CRC-related DMGs to serve as potential biomarkers for early detection by including all the available information in the data and avoiding the exclusion of any genomic position. To this end, we acquired a high-throughput DNA methylation dataset which consists of patients with CRC and healthy individuals. Information on age, history of smoking, and drug abuse was also collected. A description of the data is provided in Section “Methods”. Information on other datasets used for validation and survival analysis and all statistical and Bioinformatics methods are listed in this section. In Section “Results”, a comprehensive analysis of data is conducted. Section “Discussion” gives a discussion and some concluding remarks.
Methods
In this section, we outline the data analysis process we followed to detect DMGs, hub genes, and their effects on the survival time and enriched pathways of CRC. Figure 1 depicts the flowchart of this process.
Phase I (pre-processing of discovery samples)
To identify methylation-based CRC biomarkers, information on 6 patients with adenocarcinoma of CRC and 6 normal males was obtained. Two groups were matched based on age, and family history of cancer28.
The methylation profiles in our dataset are derived from a three-step pre-processing phase conducted through SureSelectXT Human Methyl-Seq. Initially, the purity and quantity of 12 DNA tissue samples were assessed using specific criteria, including a minimum concentration of \(50 ng/\upmu {\text{l}}\), a purity ratio \((A260/A280) \ge 1.7\), a volume of at least \(20 ng/\upmu {\text{l}}\), and a total amount exceeding \(3.0 \upmu {\text{g}}\). Subsequently, global methylation profiles of CRC and normal samples were analyzed using SureSelectXT Human Methyl-Seq.
Pre-sequencing tasks, such as sample collection and DNA extraction, were consistently carried out by a single technician. Experimental conditions for all samples remained constant both before and after sequencing. During the sequencing process, all sample runs were executed simultaneously using the same device, employing Next-generation sequencing technology, a highly parallel sequencing method. This approach minimized the potential introduction of batch effects attributable to non-biological factors such as variations in laboratory conditions, personnel, and equipment used in the experiment.
In the second step, a quality control assessment of total reads using FastQ29 was conducted. This step aimed to provide informative global and graphical representations of read quality in methylation sequencing, both pre and post-alignment. Notably, our data consistently exhibited high quality in raw sequencing reads across all samples. Subsequently, Trim Galore30 was utilized to process the raw sequencing reads. This involved the removal of sequencing adapters, specifically the Illumina universal adapter, and discarding the low-quality bases (those with quality scores below 67, as per Illumina standards) located at the 3ʹ end of reads. Additionally, any ambiguous bases found in both reads were removed.
Finally, the raw bisulfite sequencing data were aligned to the human reference genome (GRCh37/19) using Bismark31. Several comparisons and visualization confirmed minimal to no presence of batch effects in our data. This discovery dataset includes methylation read counts and read-depth for each CpG site, generating 57 to 76 million Illumina sequencing reads per subject. Between 88.5% and 89.8% of sequenced reads were mapped to either strand of the human genome (GRCh37/19). On average, each CpG site was sequenced between 19 × and 24 × per sample. The sequencing details for the subjects are presented in Table 1. Approximately 68% of the 19,530,818 CpG sites have missing information in at least one sample.
Phase II (identification of differentially methylated genes)
We utilized the DMCHMM pipeline32 to identify CpGs with differentially methylated patterns between CRC and normal discovery samples. We specifically did not remove any position with missing information or low read-depth. The missing information was imputed using DMCHMM via hidden Markov models14. Significant differentially methylated cytosines (DMCs) were selected based on the FDR threshold of 0.05. DMCs were aligned to the human reference genome (GRCh37/19) using the UCSC Genome Browser (https://genome.ucsc.edu). A gene whose promoter was mainly hypo- or hyper-methylated was classified as hypo- or hyper DMG, respectively.
Phase III (cross-platform validation)
To validate our result, several methylation profiles (GSE5305133, GSE7771834, GSE10176413, GSE4275235, GSE4868436) were extracted from the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/). Of these datasets, a total of 212 CRC and 242 normal mucosa tissue samples were selected based on setup conditions to minimize the confounding effect of other variables. These datasets have provided valuable insights into the molecular alterations that occur in CRC, and their findings have implications for the diagnosis and treatment of this disease. For the analysis of methyl array profiles of validation sets, the GEO2R (http://www.ncbi.nlm.nih.gov/geo/geo2r/) web tool and the limma R-package37 were used. To mitigate batch effects, we applied the ‘removeBatchEffect’ option from the package. A probe was considered differentially methylated if its adjusted p-value was less than 0.05, and the absolute of \(\log _2\) of methylation fold change was greater or equal to 1. The differentially methylated probes were aligned to the human reference genome (GRCh37/19) using the FDb.InfiniumMethylation.hg19 package38. In the last step, we compared the lists of DMGs based on the validation sets and our discovery samples to identify consistent hypo/hyper-methylated genes across different populations and platforms.
Phase IV (network construction and functional analysis)
In order to investigate the Protein-Protein Interaction (PPI) network and module analysis, we utilized the ‘Search Tool for the Retrieval of Interacting Genes’ (STRING) database. We set the interaction score threshold to 0.4 to screen for high-confidence interactions and visualized the resulting network using the Cytoscape39 software (Version 3.9.1). Next, we employed the Molecular Complex Detection (MCODE) algorithm to uncover densely connected substructures within the network. The MCODE score must be greater than 3 and the minimum number of nodes must be 4. In order to identify key hub genes within the network, we used the cytoHubba plugin and considered the degree of centrality as a parameter.
To gain insight into the biological mechanisms that are driving CRC and prioritize identified DMGs, we performed functional and pathway enrichment analysis using DAVID40 (https://david.ncifcrf.gov/). Gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG41) pathways were considered significantly enriched if the p-values were less than 0.05 and the q-values were less than 0.1. The visualization of the identified GO terms and KEGG pathways were done with the clusterProfiler42, pathfindR43, and ShinyGO44,45 (http://bioinformatics.sdstate.edu/go/) packages.
Phase V (Uncovering intangible heterogeneity of DMG effects on survival time)
To explore the relationship between identified DMGs and survival time, the DNA methylation profiles of 521 samples were obtained from The Cancer Genome Atlas (TCGA) network46. Complete information on clinical variables including days to follow-up and the status of the patient were analyzed.
We conducted several preliminary analyses on the overall survival time of patients with CRC. First, we estimated the density of the logarithm of the survival times using the Kaplan-Meier estimator. The density plot in Figure 2 shows a mixture distribution. Second, we applied mixture and non-mixture models of normal distributions using the mixtools package47. The BICs for the mixture of components \(K=1, 2, 3, 4\), and 5 were estimated as 777.73, 709.02, 712.31, 722.51, and 721.68, respectively, with the lowest BIC observed for the mixture with 2 components. Finally, we employed mixture and non-mixture models of semiparametric scaled data using the stochastic EM algorithm48 via mixtools. Please note that the package only outputs a mixture with two components, and the BICs for the non-mixture \(K=1\) and the mixture of 2 components were 1323.62 and 582.01, respectively. Similarly, the lowest BIC was observed for the mixture with 2 components. All of these preliminary analyses indicate heterogeneity in the overall survival time of patients with CRC.
Furthermore, a comprehensive literature review revealed numerous conflicting results. For example, researchers49 found a significant association between the methylation level of RASSF1A and the overall survival of CRC patients, while other studies50 did not observe such an association.
Moreover, several studies indicate stage-specific51 and age-specific52 effects of DNA methylation in certain genes on the survival outcomes of CRC patients. These results clearly suggest heterogeneity in the overall survival time of patients with CRC.
We hypothesize that the effect of identified DMGs and hub genes on the overall survival time varies in each subpopulation, but not all DMGs and hub genes have an effect in each subpopulation, implying that the underlying regression model is sparse. To capture such heterogeneity, we employed the sparse estimation method in the finite mixture of AFT regression models26. The details of the method are given in Section “Sparse finite mixture of AFT regression models to estimate the DMG effects on the survival times” below. The response variable is “Overall Survival Time”, and the independent variables are the log-transformed average methylation of identified DMGs or hub genes that we discovered through Phase II–IV. The goal of such a regression model is to estimate the effects of each gene in different sub-populations of the response variable, providing insights into the effects of each gene on the survival time of patients with CRC. It is important to note that the response variable (survival time) is subject to right-censoring. The sparse estimation method requires tuning parameters, which are estimated using a data-adaptive approach explained in Section “Sparse finite mixture of AFT regression models to estimate the DMG effects on the survival times”.
Sparse finite mixture of AFT regression models to estimate the DMG effects on the survival times
As hypothesized above, the overall survival time of patients with CRC is heterogeneous; thus, we hypothesize that the relationship between overall survival time and DMGs and hub genes found in Phase II-IV is heterogeneous. Such heterogeneity cannot be detected using a regular AFT regression model for censored data. Therefore, we employ the finite mixture of the AFT regression model to capture intangible DMG and hub gene effects on survival time. To this end, we use the finite mixture of AFT regression model:
where \(f_k\) and \(S_k\) are respectively the density of normal distribution and its survival function, \(y = \log (t)\), t is the overall survival time, \(\delta\) is an indicator representing right-censored (i.e., \(\delta = 0\) if time is censored and 1 if it is not censored), \(\textbf{X}\) is the vector of all DMGs and hub genes discovered in Phase II-IV, \(\varvec{\beta }_k\) is the vector of effects of these genes in Component k of the mixture model, \(\sigma _k^2\) is the variance, and \(\pi _k\) is the proportion of the kth component.
It is common to screen the number of genes prior to analysis in case of a large number of identified genes. To this end, we applied a correlation-adjusted score method using the carSurv package27 to screen the genes.
Next, we used the fmrs package53 to fit finite mixture and non-mixture of AFT regression models to the data. We employed the smoothly clipped absolute deviations (SCAD) penalty26. This sparse method requires K tuning parameters which are estimated via the data-adaptive component-wise BIC method proposed in Shokoohi et al.26.
Results
Differentially methylated cytosine detection
We identified 2,691,019 DMCs between CRC and normal groups of the discovery dataset while adjusting for the potential confounding effect of smoking history or drug abuse. Of these identified DMCs, 1,985,557 positions were hypo-methylated and 705,462 CpGs were hyper-methylated in CRC vs normal samples. The heatmaps (see R package pheatmap54) in Fig. 3a indicate a clear clustering pattern between the CRC and normal samples based on the predicted methylation levels of DMCs.
To explore the genomic location of the DMCs, we analyzed their distribution across different regions and summarized the results in Fig. 3b. Intergenic regions were found to harbor the majority of the detected DMCs both in the hypo and hyper categories. Notably, we observed that 32% of hyper-methylated DMCs were located in CpG islands, while only 9% of hypo-methylated DMCs were found in these regions. Additionally, the regions with the highest percentage of hyper-methylated DMCs were identified in introns, exons, and CGI shores. The Chord diagrams (see R package circlize55) in Fig. 3c gives a comprehensive overview of how hyper and hypo-methylated DMCs were distributed across different genomic regions. Our findings suggest that many DMCs in intergenic regions were expanded to intronic regions in both hypo and hyper-methylated categories.
Given the potential significance of promoter methylation in cancer development and progression, we focused our subsequent analysis on DMCs located on gene promoters, which encompassed 268,978 CpGs. These CpGs resided on 3406 gene promoters, of which 1394 were hyper-methylated and 2012 were hypo-methylated. The list of DMGs is available as supplementary material.
Robust DMGs in CRC
To verify the robustness of identified DMGs, we performed a cross-platform procedure with DMGs identified in selected GEO datasets as depicted in Fig. 4a (see R package venn56). The comparison revealed a total of 1571 overlapped DMGs that were consistently identified across multiple studies. As Fig. 4b (see R package karyoploteR57) illustrated, the identified DMGs were spread almost evenly across different chromosomes, with chromosomes 1 and 7 having some dense regions of CRC-related DMGs. Within this set, 917 genes were hypo-methylated, and 654 genes were hyper-methylated. We focused our subsequent analysis on these identified DMGs to gain a deeper understanding of their role in CRC pathogenesis.
GO enrichment KEGG pathway analysis
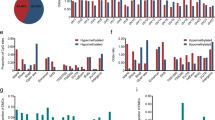
The analysis of robust DMGs in CRC utilizing the DAVID tool yielded a variety of enriched biological processes, molecular functions, and cellular components. Specifically, the hyper-methylated DMGs were found to be principally involved in ‘cell fate commitment’, ‘regionalization’, ‘embryonic organ morphogenesis’, ‘embryonic organ development’, ‘pattern specification process’, ‘animal organ morphogenesis’, ‘tube morphogenesis’, ‘tube development’, and ‘neurogenesis’ in the context of biological processes (Fig. 5a, see R Shiny package ShinyGO44). Enriched cellular components included ‘basement membrane’, ‘integral component of postsynaptic membrane’, and ‘Collagen-containing extracellular matrix’ (Fig. 5c). Additionally, KEGG pathway analysis indicated that hyper-methylated DMGs were significantly enriched in several pathways, including ‘signaling pathways regulating pluripotency of stem cells’, ‘axon guidance’, ‘morphine addiction’, ‘rap1 signaling pathway’, ‘circadian entrainment’, and ‘pathways in cancer’ (Fig. 5e and Table 2). Regarding biological processes, the hypo-methylated DMGs were found to be associated with a number of processes including ‘keratinization’, ‘keratinocyte differentiation’, ‘epidermal cell differentiation’, and ‘epithelial cell differentiation’ (Fig. 5b). Furthermore, analysis of the cellular component pathway revealed that the hypo-methylated DMGs were most significantly enriched in the ‘cornified envelope’, ‘integral component of the synaptic membrane’, and ‘integral component of the postsynaptic membrane’. Notably, these cellular components demonstrated the highest FDR and fold enrichment (Fig. 5d). Regarding molecular functions, the pathways with higher fold enrichment included ‘molecular transducer activity’, ‘signaling receptor activity’, and ‘transmembrane signaling receptor activity’. Notably, KEGG pathway analysis revealed that hypo-methylated DMGs were significantly enriched in several pathways, including the ‘oxytocin signaling pathway’, ‘glioma’, ‘adrenergic signaling in cardiomyocytes’, ‘MAPK signaling pathway’, ‘arrhythmogenic right ventricular cardiomyopathy’, and ‘cell adhesion molecules’ (Fig. 5f). These results offer valuable insights into the potential mechanisms of DMGs in CRC and identify possible therapeutic targets for this disease. A comprehensive summary of the KEGG pathways of hyper-methylated DMGs can be found in Table 2.
PPI network construction
We ran a PPI network to further investigate the complex interactions between DMGs and find important hub proteins. A total of 606 PPI nodes of the hyper-methylated DMGs were constructed on the basis of the STRING database (Fig. 6, see R Shiny package ShinyGo44). The 16 node proteins, including KIT, SEMA7A, BDNF, MEF2A, LDB2, GATA4, LHX2, SOST, CTLA4, NKX2-2, TLE4, BMP5, NFATC1, ZFPM1, DPYSL2, and ITGA2B that showed a close interaction with other node proteins were chosen as hub genes (Fig. 7a, see Cytoscape39). The most important biological process and KEGG pathways of hub genes are shown in Fig. 7b and c. One important module was selected when the number of nodes is greater than 4. The key module demonstrated functions enriched in pathways such as Wnt signaling58 (Table 2 and Fig. 8, see R Shiny package ShinyGo44)).
Wnt signaling pathway. The identified genes SOST, Gro/TLE, and NFAT are highlighted (R Shiny package ShineyGO, v0.77 & https://www.kegg.jp/pathway/hsa04310).
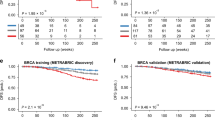
We performed a survival analysis using the TCGA-selected samples to investigate the association of selected hub genes with the survival time of CRC patients. Based on Fig. 9a–d (see GEPIA202159), those patients with gene SEMA7A (\(p=0.024\)), SOST (\(p=0.027\)), NFATC1 (\(p=0.017\)), and TLE4 (\(p=0.0061\)) being upregulated, had a significantly lower probability of survival. However, this conclusion is based on univariate analysis, and the effect of other genes and the potential heterogeneity of DMG effects were ignored. We reanalyzed these data by accounting for the heterogeneity of DMG effects and obtained different results as follows.
Overall survival of CRC patients stratified by their hub gene expression levels (http://gepia2021.cancer-pku.cn).
Intangible heterogeneity of DMG effects on survival time
We studied the relationship between the average promoter methylation of the identified DMGs and the survival time subject to right-censoring by accounting for the heterogeneity of gene effects using an independent set of 521 TCGA CRC samples. To this end, we screened all the 1571 candidate DMGs using the correlation-adjusted regression survival scores to obtain the list of top candidate covariates. This process led to the selection of 95 highly correlated DMGs. These genes were also dysregulated in the TCGA samples. In addition, 4 hub genes that were related to the survival time of CRC patients were added to the list of covariates.
Our analysis yielded a two-component mixture of AFT regression model. The estimated gene effects on the survival time are given in Table 3. The result showed that 46% of the subjects were classified into Component 1, which is the most aggressive form of the disease. Figure 10 (see R package fmrs60) depicts the posterior probability of a subject belonging to Component 1. From this figure, we noticed that all living patients were classified into Component 2, which is the less aggressive form of the disease. A total of 83 and 18 DMGs were active in Components 1 and 2, respectively. Twelve genes including HLA-F, MMP2, MT1A, RFPL4B, SIX6, ZFAT, BCKDK, AMOTL1, ADCY10, KCNK10, STAU2, and NOC4L were not related to survival time in either of the components. These findings demonstrate the heterogeneity of DMG effects in CRC data and justify using a sparse mixture modeling rather than a univariate one. In addition, the DMGs with active promoters in Component 1 can be considered biomarkers for CRC prognosis. The bioinformatics and biological information of selected DMGs are given in Table 4.
Discussion
Colorectal cancer is one of the deadliest cancers in the world. Given that early stages of CRC do not display symptoms, proactive screening is the only viable approach to identify the disease83. As DNA methylation changes are closely associated with cancer, their role in CRC biomarker detection in the early stages of cancer is of great importance. Although many CRC biomarkers have been detected in the literature, only a few are used in practice. Our findings resulted in identifying new biomarkers for CRC which can be used for diagnosis and prognosis.
We identified 1,571 DMGs most of which have been previously studied in the literature. Among them, SEPT9, NDRG4, VIM, APC, SFRP1, SFRP4, and SFRP5,84 are the most important CRC-related ones. We also explored CRC-related hub genes. Fourteen functional modules that may play important roles in the early detection of CRC were highlighted and the sub-network of hub genes KIT, SEMA7A, BDNF, MEF2A, LDB2, GATA4, LHX2, SOST, CTLA4, NKX2-2, TLE4, BMP5, NFATC1, ZFPM1, DPYSL2, and ITGA2B was extracted. These hub genes were flagged as potential diagnostic and therapeutic targets for CRC in our analysis.
In addition to the diagnostic role of our identified hub genes such as NKX2-2, KIT, BNDF, and TLE4 in CRC and its sub-types74,85,86,87, their roles in increasing CRC risk, tumor progression, and targeted therapy have been investigated. For instance, MEF2A88 and BMP589 increase the CRC risk. Up-regulation of the expression of ITGB7 and ITGA2B has been found to be significantly associated with death by sodium butyrate-induced CRC organoids90. Moreover, some studies91,92 have shown effective treatments by targeting CLT-4 and LDB2n.
There is a rich literature on the contribution of some of our identified hub genes in CRC and less evidence in support of some others such as LHX2, ZFPM1, and DPYSL2. For instance, the differences in tumor and corresponding adjacent benign tissues regarding LHX gene expressions have been investigated93. However, contrary to our findings, they did not find any statistical differences for LHX2 and LHX3 genes. Furthermore, the upregulation of ZFPM1 was revealed in molecular high-risk patients with cytogenetically normal acute myeloid leukemia94, yet its diagnostic value in CRC has not fully been confirmed95. SEMA7A is also one of our selected hub genes that play a key role in several cancers including pancreatic, breast, and lung cancers69,96,97,98. However, there has been less attention on the role of SEMA7A in CRC. Further investigation is required on our flagged DMGs.
Although there are many mechanisms that drive CRC, only a handful of them have been discovered in past studies. As researchers continue to genotype large panels of CRC tumors, it can be expected that additional new pathways of CRC carcinogenesis will be revealed. SOST, an identified hub gene in our study, plays a vital role in inhibiting the Wnt signaling pathway by binding to the Wnt co-receptor, LRP5/6, and preventing its activation99. Therefore, decreased SOST expression could lead to an increase in Wnt signaling, promoting CRC cell proliferation, migration, and survival. Another identified hub gene is TLE4 which is involved in the negative regulation of the canonical Wnt signaling pathway. Only a few investigations provided evidence of TLE4 upregulation in CRC biopsies, partially through regulation of the JNK/c-Jun pathway73. Moreover, recent studies that focus on the NFAT signaling pathway showed a promising strategy for CRC treatment64.
Heterogeneity is one of the key features of genomic data. Specifically, there is evidence of the heterogeneity of DMG effects on the survival of CRC patients in the literature and in our dataset. The finite mixture of the AFT regression model is a plausible method to uncover such intangible heterogeneity. Our analysis suggested a mixture of two-component mixture of the AFT regression model in which patients were separated into two subgroups based on their vital status. In this model, almost all of the deceased patients were classified into the most aggressive form of the disease (Component 1). In Component 1, 83 DMGs including NMNAT2, ZFP42, NPAS2, MYLK3, NUDT13, KIRREL3, and FKBP6 had an effect on the survival time of the patients. The relation between some of these DMGs and survival time has been previously reported100. On the other hand, there are a few discoveries regarding other genes. For instance, significantly higher expression of NMNAT2 in CRC tissues compared to normal ones have been found, yet this gene was not a prognostic factor for overall survival101. Note that, while the hub genes SOST, NFATC1, and TLE4 were associated with survival in the univariate Cox model, they were only associated with survival time in the most aggressive form of the disease in our study.
Our study does not exclusively depend on bioinformatics analysis, as we have employed several statistical and machine learning analyses. These include modeling methylation profiles, identifying DMCs via DMCHMM, conducting statistical tests, performing multiple validation analyses, and applying statistical learning algorithms to survival times via fmrs. One of the advantages of the DMCHMM method is that it does not require a large number of samples or matched samples, as it is highly flexible and can accommodate various experimental designs. It demonstrates significant power, particularly when dealing with moderate to low sample sizes.
Data availability
In this study methylation profiling datasets with accession numbers GSE53051, GSE77718, GSE101764, GSE42752, and GSE48684 were obtained from Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/), of the National Center for Biotechnology Information (NCBI). Additional DNA methylation datasets and expression profiles of CRC patients (TCGA-COAD, TCGA-READ, TCGA-SARC projects) were obtained from The Cancer Genome Atlas (TCGA, https://www.cancer.gov/ccg/research/genome-sequencing/tcga), of the National Cancer Institute (NCI). Our SureSelectXT Human Methyl-Seq dataset on methylation profiles of 6 patients with adenocarcinoma of CRC and 6 normal males is obtained from ‘Reza Radiotherapy and Oncology Center’ in Iran and is available upon request.
References
Sung, H. et al. Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Fearon, E. R. Molecular genetics of colorectal cancer. Annu. Rev. Pathol. Mech. Dis. 6, 479–507 (2011).
Andrew, A. et al. Risk factors for diagnosis of colorectal cancer at a late stage: A population-based study. J. Gen. Intern. Med. 33, 2100–2105 (2018).
Das, P. & Singal, R. DNA methylation and cancer. J. Clin. Oncol. 22, 4632–4642 (2004).
Moore, L., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Ashktorab, H. & Brim, H. DNA methylation and colorectal cancer. Curr. Colorectal Cancer Rep. 10, 425–430 (2014).
Grady, W. Epigenetic events in the colorectum and in colon cancer. Biochem. Soc. Trans. 33, 684–688 (2005).
Lam, K. et al. DNA methylation-based biomarkers in colorectal cancer: A systematic review. Biochim. Biophys. Acta 1866, 106–120 (2016).
Payne, S. R. From discovery to the clinic: The novel DNA methylation biomarker m SEPT9 for the detection of colorectal cancer in blood. Epigenomics 2, 575–585 (2010).
Imperiale, T. et al. Multitarget stool DNA testing for colorectal-cancer screening. N. Engl. J. Med. 370, 1287–1297 (2014).
Mathers, J., Strathdee, G. & Relton, C. Induction of epigenetic alterations by dietary and other environmental factors. Adv. Genet. 71, 3–39 (2010).
Issa, J.-P. et al. Methylation of the oestrogen receptor CpG island links aging and neoplasia in human colon. Nat. Genet. 7, 536–540 (1994).
Barrow, T. M. et al. Smoking is associated with hypermethylation of the APC 1A promoter in colorectal cancer: The ColoCare Study. J. Pathol. 243, 366–375 (2017).
Shokoohi, F. et al. A hidden Markov model for identifying differentially methylated sites in bisulfite sequencing data. Biometrics 75, 210–221 (2019).
Al-Sohaily, S. et al. Molecular pathways in colorectal cancer. J. Gastroenterol. Hepatol. 27, 1423–1431 (2012).
Ilyas, M., Straub, J., Tomlinson, I. & Bodmer, W. Genetic pathways in colorectal and other cancers. Eur. J. Cancer 35, 1986–2002 (1999).
Behrens, J. The role of the Wnt signaling pathway in colorectal tumorigenesis. Biochem. Soc. Trans. 33, 672–675 (2005).
Fang, J. & Richardson, B. The MAPK signaling pathways and colorectal cancer. Lancet Oncol. 6, 322–327 (2005).
Markowitz, S. et al. Inactivation of the type II TGF-\(\beta\) receptor in colon cancer cells with microsatellite instability. Science 268, 1336–1338 (1995).
Levine, A. & Oren, M. The first 30 years of p53: Growing ever more complex. Nat. Rev. Cancer 9, 749–758 (2009).
Gong, B. et al. Identification of hub genes related to carcinogenesis and prognosis in colorectal cancer based on integrated bioinformatics. Mediat. Inflamm. 2020, 1–11 (2020).
Huang, H. et al. Integrative analysis of identifying methylation-driven genes signature predicts prognosis in colorectal carcinoma. Front. Oncol. 11, 629860 (2021).
Hu, J. et al. An eight-CpG-based methylation classifier for preoperative discriminating early and advanced-late stage of colorectal cancer. Front. Genet. 11, 614160 (2021).
Feng, Z., Liu, Z., Peng, K. & Wu, W. A prognostic model based on nine DNA methylation-driven genes predicts overall survival for colorectal cancer. Front. Genet. 12, 2446 (2022).
Long, J. et al. DNA methylation-driven genes for constructing diagnostic, prognostic, and recurrence models for hepatocellular carcinoma. Theranostics 9, 7251 (2019).
Shokoohi, F., Khalili, A., Asgharian, M. & Lin, S. Capturing heterogeneity of covariate effects in hidden subpopulations in the presence of censoring and large number of covariates. Ann. Appl. Stat. 13, 444 (2019).
Welchowski, T., Zuber, V. & Schmid, M. Correlation-adjusted regression survival scores for high-dimensional variable selection. Stat. Med. 38, 2413–2427 (2019).
Kerachian, M. A. et al. Crosstalk between dna methylation and gene expression in colorectal cancer, a potential plasma biomarker for tracing this tumor. Sci. Rep. 10, 1–13 (2020).
Wingett, S. W. & Andrews, S. Fastq screen: A tool for multi-genome mapping and quality control. F1000 Res. 7, 1–10 (2018).
Krueger, F. Trim Galore!: A Wrapper Around Cutadapt and Fastqc to Consistently Apply Adapter and Quality Trimming to Fastq Files, with Extra Functionality for RRBS Data (Babraham Institute, 2015).
Krueger, F. & Andrews, S. R. Bismark: A flexible aligner and methylation caller for bisulfite-seq applications. Bioinformatics 27, 1571–1572 (2011).
Shokoohi, F. DMCHMM: Differentially methylated CpG using hidden Markov model. Bioconductorhttps://doi.org/10.18129/B9.bioc.DMCHMM (2023).
Timp, W. et al. Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors. Genome Med. 6, 1–11 (2014).
McInnes, T. et al. Genome-wide methylation analysis identifies a core set of hypermethylated genes in CIMP-H colorectal cancer. BMC Cancer 17, 1–11 (2017).
Naumov, V. A. et al. Genome-scale analysis of DNA methylation in colorectal cancer using infinium human methylation 450 bead chips. Epigenetics 8, 921–934 (2013).
Luo, Y. et al. Differences in DNA methylation signatures reveal multiple pathways of progression from adenoma to colorectal cancer. Gastroenterology 147, 418–429 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Tim Triche, J. FDb.InfiniumMethylation.hg19: Annotation package for Illumina Infinium DNA methylation probes. R package version 2.2.0. (2014).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Sherman, B. T. et al. David: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. Clusterprofiler: An r package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Ulgen, E., Ozisik, O. & Sezerman, O. U. pathfindr: An r package for comprehensive identification of enriched pathways in omics data through active subnetworks. Front. Genet. 10, 858 (2019).
Ge, S. X., Jung, D. & Yao, R. Shinygo: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Luo, W. & Brouwer, C. Pathview: An r/bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Network, C. G. A. et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330 (2012).
Benaglia, T., Chauveau, D., Hunter, D. R. & Young, D. mixtools: An R package for analyzing finite mixture models. J. Stat. Softw. 32, 1–29 (2009).
Bordes, L. & Chauveau, D. Stochastic em algorithms for parametric and semiparametric mixture models for right-censored lifetime data. Comput. Stat. 31, 1513–1538 (2016).
Matthaios, D. et al. Methylation status of the apc and rassf1a promoter in cell-free circulating dna and its prognostic role in patients with colorectal cancer. Oncol. Lett. 12, 748–756 (2016).
Chen, S.-P. et al.\(\beta\)-catenin and k-ras mutations and rassf1a promoter methylation in Taiwanese colorectal cancer patients. Genet. Test. Mol. Biomark. 16, 1277–1281 (2012).
Muthamilselvan, S., Raghavendran, A. & Palaniappan, A. Stage-differentiated ensemble modeling of dna methylation landscapes uncovers salient biomarkers and prognostic signatures in colorectal cancer progression. PLoS ONE 17, e0249151 (2022).
Joo, J. E. et al. Dna methylation signatures and the contribution of age-associated methylomic drift to carcinogenesis in early-onset colorectal cancer. Cancers 13, 2589 (2021).
Shokoohi, F. Sparse estimation in finite mixture of accelerated failure time and mixture of regression models with R package fmrs. N. Engl. J. Stat. Data Sci. 1, 1–18. https://doi.org/10.51387/23-NEJSDS49 (2023).
Kolde, R. pheatmap: Pretty Heatmaps (2019). R package version 1.0.12.
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. Circlize implements and enhances circular visualization in r. Bioinformatics 30, 2811–2812 (2014).
Dusa, A. venn: Draw Venn Diagrams (2022). R package version 1.11.
Gel, B. & Serra, E. karyoploter: An r / bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33, 3088–3090. https://doi.org/10.1093/bioinformatics/btx346 (2017).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. Kegg for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Tang, Z. et al. Gepia: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Shokoohi, F. fmrs: Variable Selection in Finite Mixture of AFT Regression and FMR. Bioconductor Open Source Software for Bioinformatics. https://doi.org/10.18129/B9.bioc.fmrs (2023). R package version 2.0.1.
Pérez, A. G. et al. N-glycosylation and receptor tyrosine kinase signaling affect claudin-3 levels in colorectal cancer cells. Oncol. Rep. 44, 1649–1661 (2020).
Cherradi, S., Martineau, P., Gongora, C. & Del Rio, M. Claudin gene expression profiles and clinical value in colorectal tumors classified according to their molecular subtype. Cancer Manag. Res. 1, 1337–1348 (2019).
Chuvpilo, S. et al. Multiple nf-atc isoforms with individual transcriptional properties are synthesized in t lymphocytes. J. Immunol. 162, 7294–7301 (1999).
Shen, T. et al. NFATc1 promotes epithelial-mesenchymal transition and facilitates colorectal cancer metastasis by targeting SNAI1. Exp. Cell Res. 408, 112854 (2021).
Wu, B. et al. Development of an immune infiltration-related eight-gene prognostic signature in colorectal cancer microenvironment. BioMed. Res. Int. 2020 (2020).
Mazzoni, S. M. & Fearon, E. R. Axin1 and axin2 variants in gastrointestinal cancers. Cancer Lett. 355, 1–8 (2014).
Mastrantonio, R., You, H. & Tamagnone, L. Semaphorins as emerging clinical biomarkers and therapeutic targets in cancer. Theranostics 11, 3262 (2021).
Fijneman, R. J. et al. Proximal fluid proteome profiling of mouse colon tumors reveals biomarkers for early diagnosis of human colorectal cancer. Clin. Cancer Res. 18, 2613–2624 (2012).
Liu, Y., Guo, C., Li, F. & Wu, L. LncRNA LOXL1-AS1/miR-28-5p/SEMA7A axis facilitates pancreatic cancer progression. Cell Biochem. Funct. 38, 58–65 (2020).
Long, N. P. et al. Novel biomarker candidates for colorectal cancer metastasis: A meta-analysis of in vitro studies. Cancer Inform. 15, 40301 (2016).
Matchett, E. C., Ambrose, E. C. & Kornbluth, J. Characterization of uridine-cytidine kinase like-1 nucleoside kinase activity and its role in tumor growth. Biochem. J. 479, 1149–1164 (2022).
Wang, F., Qin, G., Liu, J., Wang, X. & Ye, B. Integrated genome-wide methylation and expression analyses reveal key regulators in osteosarcoma. Comput. Math. Methods Med. 2020 (2020).
Wang, S.-Y. et al. TLE4 promotes colorectal cancer progression through activation of JNK/c-Jun signaling pathway. Oncotarget 7, 2878 (2016).
Yu, H. et al. DNA methylation profile in CpG-depleted regions uncovers a high-risk subtype of early-stage colorectal cancer. JNCI J. Natl. Cancer Inst. 115, 52–61 (2023).
Regan, J. L. et al. Identification of a neural development gene expression signature in colon cancer stem cells reveals a role for egr2 in tumorigenesis. Science 25, 104498 (2022).
De Robertis, M. et al. Novel insights into notum and glypicans regulation in colorectal cancer. Oncotarget 6, 41237 (2015).
Danielsen, S. A. et al. Phospholipase c isozymes are deregulated in colorectal cancer-insights gained from gene set enrichment analysis of the transcriptome. PLoS ONE 6, e24419 (2011).
Sandberg, A. et al. Fibrillation and molecular characteristics are coherent with clinical and pathological features of 4-repeat tauopathy caused by mapt variant g273r. Neurobiol. Dis. 146, 105079 (2020).
Wang, C. et al. Mapt promoter cpg island hypermethylation is associated with poor prognosis in patients with stage ii colorectal cancer. Cancer Manag. Res. 1, 7337–7343 (2019).
Li, T., Huang, S., Yan, W., Zhang, Y. & Guo, Q. Foxf2 regulates prune2 transcription in the pathogenesis of colorectal cancer. Technol. Cancer Res. Treatm. 21, 15330338221118716 (2022).
Hauptman, N., Jevšinek Skok, D., Spasovska, E., Boštjančič, E. & Glavač, D. Genes cep55, foxd3, foxf2, gnao1, gria4, and kcna5 as potential diagnostic biomarkers in colorectal cancer. BMC Med. Genomics 12, 1–17 (2019).
Barderas, R. et al. In-depth characterization of the secretome of colorectal cancer metastatic cells identifies key proteins in cell adhesion, migration, and invasion. Mol. Cell. Proteom. 12, 1602–1620 (2013).
Imperiale, T. et al. Fecal DNA versus fecal occult blood for colorectal-cancer screening in an average-risk population. N. Engl. J. Med. 351, 2704–2714 (2004).
Mueller, D. & Győrffy, B. DNA methylation-based diagnostic, prognostic, and predictive biomarkers in colorectal cancer. Biochim. Biophys. Acta 1877, 1–12 (2022).
Gutierrez, A., Demond, H., Brebi, P. & Ili, C. Novel methylation biomarkers for colorectal cancer prognosis. Biomolecules 11, 1722 (2021).
He, Y. et al. NK homeobox 2.2 functions as tumor suppressor in colorectal cancer due to DNA methylation.. J. Cancer 11, 4791 (2020).
Küçükköse, E. et al. KIT promotes tumor stroma formation and counteracts tumor-suppressive TGF\(\beta\) signaling in colorectal cancer. Cell Death Dis. 13, 617 (2022).
Xiao, Q. et al. MEF2A transcriptionally upregulates the expression of ZEB2 and CTNNB1 in colorectal cancer to promote tumor progression. Oncogene 40, 3364–3377 (2021).
Pellatt, A. J. et al. The TGF\(\beta\)-signaling pathway and colorectal cancer: Associations between dysregulated genes and miRNAs. J. Transl. Med. 16, 1–22 (2018).
Li, F. et al. Transcriptomic landscape of sodium butyrate-induced growth inhibition of human colorectal cancer organoids. Mol. Omics 18, 754–764 (2022).
Rotte, A. Combination of CTLA-4 and PD-1 blockers for treatment of cancer. J. Exp. Clin. Cancer Res. 38, 1–12 (2019).
Yuan, C., Wu, C., Xue, R., Jin, C. & Zheng, C. Suppression of human colon tumor by EERAC through regulating Notch/DLL4/Hes pathway inhibiting angiogenesis in vivo. J. Cancer 12, 5914 (2021).
Cha, N. et al. Oncogenicity of LHX4 in colorectal cancer through Wnt/\(\beta\)-catenin/TCF4 cascade. Tumor Biol. 35, 10319–10324 (2014).
Marcucci, G. et al. Prognostic significance of, and gene and microRNA expression signatures associated with, CEBPA mutations in cytogenetically normal acute myeloid leukemia with high-risk molecular features: A Cancer and Leukemia Group B Study. J. Clin. Oncol. 26, 5078 (2008).
Ge, W. et al. A novel 4-gene prognostic signature for hypermutated colorectal cancer. Cancer Manag. Res. 11, 1985 (2019).
Rehman, M. & Tamagnone, L. Semaphorins in cancer: Biological mechanisms and therapeutic approaches. Semin. Cell Dev. Biol. 24, 179–189 (2013).
Crump, L. S. et al. Hormonal regulation of Semaphorin 7a in ER+ breast cancer drives therapeutic resistance. Cancer Res. 81, 187–198 (2021).
Kinehara, Y. et al. Semaphorin 7A promotes EGFR-TKI resistance in EGFR mutant lung adenocarcinoma cells. JCI Insight 3, 1–17 (2018).
Katoh, M. & Katoh, M. Molecular genetics and targeted therapy of Wnt-related human diseases. Int. J. Mol. Med. 40, 587–606 (2017).
Yang, S.-F., Xu, M., Yang, H.-Y., Li, P.-Q. & Chi, X.-F. Expression of circadian gene NPAS2 in colorectal cancer and its prognostic significance. Nan Fang Yi Ke Da Xue Xue Bao 36, 714–718 (2016).
Cui, C. et al. Nicotinamide mononucleotide adenylyl transferase 2: A promising diagnostic and therapeutic target for colorectal cancer. BioMed Res. Int. 2016, 1–8 (2016).
Acknowledgements
The authors would like to express their gratitude to the editor, the associate editor, and the two anonymous reviewers for their thoughtful and constructive comments, which significantly improved the manuscript. The authors also wish to extend their thanks to Prof. Kazem Taghva, the Chair of the Department of Computer Science, and Prof. Martin Schiller, the Director of the Nevada Institute for Personalized Medicine (NIPM) at the University of Nevada, Las Vegas (UNLV), for their financial support and assistance. Furthermore, the authors appreciate the generosity of the Reza Radiotherapy and Oncology Center in Iran, Mashhad University of Medical Sciences, for sharing the CRC dataset. The publication fees for this article were supported by the UNLV University Libraries Open Article Fund.
Funding
Farhad Shokoohi is supported by Start-up Grant number PG18929 and ‘In Support of Research Scholar’ Grant Number PG18494, University of Nevada, Las Vegas. This research is partially supported by NIPM, the Department of Computer Science at UNLV, and the Center of Biomedical Research Excellence through COBRE Pilot Grant Number P20GM121325.
Author information
Authors and Affiliations
Contributions
S.H.K. and F.S. analyzed the data and wrote the paper. H.E. and M.A.K. edited the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hajebi Khaniki, S., Shokoohi, F., Esmaily, H. et al. Analyzing aberrant DNA methylation in colorectal cancer uncovered intangible heterogeneity of gene effects in the survival time of patients. Sci Rep 13, 22104 (2023). https://doi.org/10.1038/s41598-023-47377-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-47377-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.