Abstract
Seagrasses are important primary producers in oceans worldwide. They live in shallow coastal waters that are experiencing carbon dioxide enrichment and ocean acidification. Posidonia oceanica, an endemic seagrass species that dominates the Mediterranean Sea, achieves high abundances in seawater with relatively low concentrations of dissolved inorganic nitrogen. Here we tested whether microbial metabolisms associated with P. oceanica and surrounding seawater enhance seagrass access to nitrogen. Using stable isotope enrichments of intact seagrass with amino acids, we showed that ammonification by free-living and seagrass-associated microbes produce ammonium that is likely used by seagrass and surrounding particulate organic matter. Metagenomic analysis of the epiphytic biofilm on the blades and rhizomes support the ubiquity of microbial ammonification genes in this system. Further, we leveraged the presence of natural carbon dioxide vents and show that the presence of P. oceanica enhanced the uptake of nitrogen by water column particulate organic matter, increasing carbon fixation by a factor of 8.6–17.4 with the greatest effect at CO2 vent sites. However, microbial ammonification was reduced at lower pH, suggesting that future ocean climate change will compromise this microbial process. Thus, the seagrass holobiont enhances water column productivity, even in the context of ocean acidification.
Similar content being viewed by others
Introduction
Associations between marine species and microbes have been increasingly identified with genomic methodologies, revealing many hidden species interactions between eukaryotic and prokaryotic partners1,2,3. In addition to affecting host fitness, these interactions may affect global biogeochemical cycles, through the use and cycling of carbon, nitrogen and other elements4,5. Seagrasses are marine angiosperms and are foundational species in coastal ecosystems that are critical in structuring these ecosystems and modulating the flux of nutrients and energy. They provide habitat for other species6,7, are key primary producers8,9, and also act as carbon sinks by sequestering carbon10,11. Marine angiosperms are some of the most widely distributed plants on the planet12, and recent studies suggest they host a diversity of microbial taxa13,14, with some functional partnerships demonstrated15,16.
Among seagrasses, a number of microbial taxa have been identified as important to nutrient cycling, including microbes capable of nitrogen fixation15, ammonification16 and sulfur oxidation13,14. Posidonia oceanica, the seagrass species that dominates the Mediterranean Sea, achieves high abundances in coastal habitats with relatively low seawater dissolved inorganic nitrogen concentrations. Microbial partnerships may partly explain its productivity. First, the low oxygen environment surrounding the roots provides a microenvironment for nitrogen fixation15,17 and other reducing metabolisms. Second, seagrasses might have access to the dissolved organic nitrogen from the surrounding seawater. A related species of Posidonia in Australia has microbial associates that ammonify amino acids—resulting in dissolved inorganic nitrogen that is taken up as ammonium16. Although it has been claimed that seagrasses can directly take up dissolved organic nitrogen sources (e.g.18), we know of no studies that show this while controlling for the role of bacteria.
Bacteria that deaminate amino acids to produce ammonium are likely abundant in nature, given that there is a diversity of enzymes that cleave carbon–nitrogen bonds and produce ammonium14. Whether these bacteria are consistently associated with hosts is relatively little studied, including what determines their abundance. Studies that focus only on the taxonomy of microbes (e.g. amplicon sequencing) may not reveal the functional capacity of microbes19. Information on amino acids in seawater in the environments where marine macrophytes are abundant are also lacking and infrequently quantified. Nonetheless, dissolved organic compounds can be abundant in coastal waters and can be a key part of nutrient recycling20.
While the discovery of host-microbe interactions is still nascent, populations of many marine macrophytes are in decline6,21,22. For seagrasses, global environmental change, including ocean acidification, warming, pollution, and physical disturbances such as increased turbidity represent key threats23. Future ocean conditions will include a continued decrease of pH and a change in carbonate system parameters. While marine angiosperms such as P. oceanica may benefit from increased bicarbonate concentrations24, little is known about how their associations with microbes may be altered. If increased bicarbonate access improves carbon fixation and fitness, host fitness might increase and microbial associates may benefit. If the availability of inorganic carbon increases while nitrogen remains the same, the release of excess carbon via stoichiometric overflow, could increase25. While this carbon release could benefit heterotrophic host-associated microbes, nitrogen limitation of microbial growth might also result. Low pH could also be a stressor to the host and host health, with potential consequences for the diversity and/or functional roles of host-associated microbes. The seagrass Cymodocea nodosa showed taxonomic differences in rhizome microbes but not leaf microbes across a pH gradient driven by the natural release of carbon dioxide (CO2) at vents in the Mediterranean26, though the functional differences were unexplored. The mechanisms underlying when microbes are resistant or sensitive to changes in host biogeochemistry from ocean acidification needs further understanding.
At CO2 vent sites in the Mediterranean the seagrass P. oceanica shows a number of differences compared to areas without the influence of venting CO2. Benthic communities near vents show a decline in calcified species27,28, and accordingly, calcareous epiphytes of P. oceanica are much reduced in proximity to the vents8,29. Additionally, herbivorous fish are more abundant near the vents30. In proximity to the vents, P. oceanica has a shorter stature, fewer epiphytes, increased tissue nitrogen content31, and increased shoot density29,32. Proximity to vents can increase P. oceanica productivity at the scale of individual leaves, but shows variable results at the community level8. Seawater pH is variable in the areas surrounding CO2 vent sites and P. oceanica is often in pH conditions around 7.728.
The foundational role that P. oceanica plays in the Mediterranean30, its economic importance33, and concern about its health status34 makes this species important for investigating host-associated microbial function in a global change context, particularly under ocean acidification. We thus investigated microbial function at natural CO2 venting areas and reference areas with no venting and ambient pH. We focused on nitrogen dynamics, a relatively little studied aspect of ocean acidification (but see35). Populations of P. oceanica may have spent generations in association with decreased pH, allowing tests of community interactions and functional changes, including host-microbe interactions.
Here, we quantified whether ammonification, or the ability of microbes to make dissolved organic nitrogen available as ammonium, occurred in microbes associated with P. oceanica, and microbes that were free-living, and whether ammonification differed in areas of low pH at CO2 vents. We tested carbon fixation rates, dissolved organic carbon release rates, and tissue nitrogen values as possible plant traits that differed among sites. We used metagenomics to describe the microbial taxa in association with Posidonia blades and roots, and probe their functions, both at a site with CO2 venting with low pH and at a nearby site unaffected by vent activity with ambient pH.
Results
The movement of 15N amino acids in P. oceanica and seawater
There were differences in the fate of the 15N amino acids that were incubated with P. oceanica versus seawater only (Fig. 1), with enrichment of seawater ammonium greatest at the control site and greatest when in association with P. oceanica and at night (Fig. 2a). The 15N in ammonium in seawater (δ15N of NH4) increased during the first 7 daylight hours (‘day’) and increased again overnight (‘night’). Initial conditions for the incubation bottles are in Table S1; statistics and all measurements of P. oceanica are in Supplementary Tables 2 and 3, respectively.
Illustration of the possible paths of δ15N amino acids in with the seagrass Posidonia oceanica or seawater only. Enlarged arrows on the left indicate processes that were greater at the control site than the CO2 venting site. All processes were also greater with Posidonia present, even if only during either the day or the night, indicated with either a sun or moon icon. All processes correspond to Fig. 2a–d and the statistical tests can be found in Table S2.
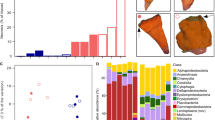
The fate of 15N amino acids added to incubation bottles with and without P. oceanica. In a, δ15N ammonium (‰) in seawater in incubation bottles with P. oceanica or seawater only. A comparison of the values during the day and night, showed a significant increase in the bottles at the end of the night, with increased values at control site and with P. oceanica. There was also an interaction between time and treatment; the increase at night was highest with P. oceanica. In b. The ammonification rate in nmol per hour across the two sites and attributed to P. oceanica and seawater versus seawater only based on Eq. (1). The control site had greater ammonification compared with the vent site (p = 0.002) and P. oceanica was nearly associated with greater ammonification rates than seawater (p = 0.060), while rates did not differ overall between night and day (p = 0.732). Examining P. oceanica separate from seawater during day versus night, P.oceanica has no diel pattern, while the POM in seawater is associated with greater ammonification during the day. In c, ammonium uptake rate in umol N per g dry mass per hour in the Posidonia meristem during both day and night periods using Eq. (2). Uptake was greater during the day (p = 0.002), with a significant interaction (p = 0.020), indicating a relatively greater uptake of 15N ammonium at night in the control site. In d., the uptake of ammonium by POM was greater when P. oceanica was absent from the incubation bottles during the day, and this effect was greatest at the control site, based on the interaction between site and treatment (p = 0.033). At night, the situation is reversed and the incubation bottles with P. oceanica resulted in a greater incorporation of 15N into the POM in the seawater compared with bottles that had no P. oceanica (p = 0.013). All values were log transformed prior to statistical analyses shown in Table S2.
The ammonification rate in nmol per hour across the two sites (Eq. 1, Methods), revealed greater ammonification rates at the control site with ambient pH compared with the vent site, both in seawater and when P. oceanica was present (Fig. 2b, p = 0.002). Across all treatments, ammonification rates were more than 4 times greater at control sites. There was a trend for greater ammonification with P. oceanica compared with seawater (p = 0.045), and ammonification with seagrass was 2.5× greater when averaged across both sites. Rates did not differ between night and day (p = 0.512). Comparing the ammonification rates where P. oceanica was in the incubation bottles, there was no diel pattern, while incubations with seawater only were associated with greater ammonification by particulate organic matter (POM) during the day (p = 0.002).
The very newest Posidonia tissue growth in the meristem showed almost 3× greater uptake of 15N-enriched ammonium during the day compared to night (p = 0.002, Fig. 2c), but the increase seen at the vent during the day switched at night to greater uptake in control sites (interaction between time of the experiment and site, p = 0.020, Fig. 2c). Rates of ammonium uptake by Posidonia reached up to 6–8 μmol N per g dry mass per hour based on estimates of 15N enrichment and using Eq. (2) (Fig. 2c, Table S3). P. oceanica plants also showed similar patterns of 15N enrichment in the midblade and on the blade under epiphytes (Table S3).
The nitrogen content in Posidonia tissue did not differ between the vent and control site, either in the meristem (t = 0.993, p = 0.360) or midblade (t = 0.517, p = 0.623), and averaged 2.1% in the meristem and 1.5% in the midblade region overall (Fig. S1).
The daylight carbon fixation rates per unit tissue mass of P. oceanica did not differ whether Posidonia was in proximity to CO2 vents or in control areas (p = 0.646, Fig. 3a, Table S2). At night, respiration rates too were indistinguishable (Fig. 3a). The lack of difference in carbon fixation by Posidonia and the nitrogen content of the blades were consistent with a statistically indistinguishable C:N ratio of P. oceanica tissue among sites, whether we considered the meristem tissue or the midblade underneath epiphytes (Fig. S1a,b).
Carbon uptake rates by a. P. oceanica and b. seawater. In a, carbon fixation or respiration, in mg C per hour per g P. oceanica dry mass in incubation bottles during daylight and nighttime hours. P. oceanica carbon fixation was based on oxygen change, following subtraction of water column rates. P. oceanica carbon fixation was greater during the day, but did not differ between control and vent sites. Two-way ANOVA (site effect 0.646, diel effect p < 0.001, interaction p = 0.728). In b, carbon uptake by unfiltered seawater (in mg C per L per hour) is shown for both sites and in incubation bottles with and without P. oceanica. POM carbon uptake differed between day and night (p < 0.001) and with P. oceanica (p = 0.002); there were significant interactions with time of day and site (p = 0.004) and time of day and treatment (p < 0.001). Boxplots show mean values and delimit 25th and 75th percentiles; all points are shown, statistical tests in Table S2.
Dissolved and inorganic nutrient dynamics
The dynamics of dissolved organic nutrients was affected by P. oceanica, proximity to CO2 vents, and time of day. Dissolved organic carbon (DOC) was relatively unaffected by P. oceanica, with a nearly significant interaction between time of day (day or night), likely reflecting the increase DOC at the vents at night (p = 0.017) (Fig. S2a). DON release in association with P. oceanica showed no difference between control and vent site as a main effect (p = 0.220) but there was an interaction over the diel cycle, with DON increased during the day when P. oceanica was fixing the most carbon (p = 0.003, Fig. S2b). The pattern of dissolved organic matter release in seawater only showed that DOC was greater at the vent site (p = 0.005) and greater at night (p = 0.041). DON release was also greater at the vent site, though the interaction between site and time of day (p < 0.001) indicated that DON release was greater during the day at the vent site and greater at night at the control site (p < 0.001). DOC and DON tended to have net production during the day and net uptake at night. In contrast, the use of inorganic nutrients was greater during daylight hours and is illustrated by decreased concentrations of ammonium, nitrite, nitrate and silica (Fig. S2).
The effect of P. oceanica on surrounding POM
The presence of P. oceanica was associated with increased uptake of 15N in POM during nighttime hours, an effect that was greatest at the control site, based on the interaction between site and treatment (p = 0.001, Fig. 2d). Combining the day and night interval, POM 15N uptake was greater with P. oceanica (Table S2), resulting in 1.6 times more nitrogen uptake in chambers with P. oceanica compared with seawater at the control site, and 2.5× higher at the vent site.
Carbon uptake by POM was also enhanced with P. oceanica though only during the day; POM carbon uptake decreased with P. oceanica at night (Fig. 3b). The mean C:N ratio of POM was 5.09 at the control site and 5.97 at the vent site (Table S3). If the carbon:nitrogen uptake by POM was relatively constant, water column carbon uptake was increased 8.6 times (1.6 × 5.09) at the control site and 17.4 times (2.5 × 5.97) at the vent site due to the contribution of bacteria that metabolize amino acids in association with P. oceanica.
Metagenome analyses
After quality control and filtering, we obtained an average of 33.7 million sequence reads per sample (range 28.6–43.8 million). Metagenomic short reads were assembled into contigs of at least 1000 nucleotides in length, resulting in 26,460–123,697 contigs per sample with a mean of 68,852 across all 6 samples (Table S4). Across all the metagenomes in our study, the taxonomy of Posidonia blades had the greatest alpha diversity, with rhizomes at the vent having the fewest (Fig. 4, Table S4), though due to lack of replication we are precluded from statistical tests of diversity.
The taxonomic assignments of 67 metagenome-assembled genomes (MAGs) in the 6 samples from the blade or rhizome surface of Posidonia as (a). Phylum, (b). Class and (c). Family. There were 22 high quality MAGs and 42 medium quality MAGs, while taxonomy was not found for 3. In (c), alpha diversity in each sample is shown and based on ‘anvi-estimate-scg-taxonomy’. See Supplemental Table S4 and S5.
The metagenomic reads allowed us to construct metagenome-assembled genomes (MAGs), ranging from only a single MAG on rhizomes at the vent site to 21 MAGs on a blade metagenome at the control site. We used Bowers et al.36 criteria to define a MAG as high quality if it had a competition score of > 90% and a redundancy (or contamination) of < 10% (Table S5), yielding 22 high quality MAGs. A remaining 42 were medium quality MAGs with completion scores between 42 and 90% and redundancy scores between 0 and 11% (Table S5). All MAGs were bacterial and represented 5 phyla (Fig. 4a). A notable difference among blades versus rhizomes was the presence of the Cyanobacteria and Bacteroidota phyla on blades but not on rhizomes. In three cases, we were unable to taxonomically resolve a metagenome-assembled genome. Taxonomic diversity at the class and family level was higher on blades compared with rhizomes (Fig. 4b,c).
The metabolism of these metagenomes was diverse, with some functional differences (Fig. 5). Photosynthetic and anoxygenic photosystem II were present only on blades, an expected result given the presence of Cyanobacteria only on blades. The KEGG module for nitrogen fixation was partially annotated within three MAGs, and especially the two MAGs in the genus Thiodiazotropha. The third MAG was an unidentified Gammaproteobacteria and was only 33% complete for nitrogen fixation genes (Table S7). Further, nitrogen fixation was indicated only on rhizome tissue and at the control site based on these three MAGs (Fig. 5, Table S7). Ammonification was a prevalent metabolism across all bacterial MAGs on P. oceanica. Of the 67 MAGs, all had enzymes classified as EC:1.4.* or EC:3.5.*, while 61 MAGs had enzyme function classified as EC:4.3.1* (Table S6). Nitrate reduction capability was present in the rhizome of seagrass from control areas (MAG 1 and MAG4), including denitrification in control rhizomes (MAG4).
The metabolisms in 67 microbial metagenome assembled genomes (MAGs) assembled on P. oceanica at Ischia, Italy across 5 Phyla and on four different tissue types. The completion (black = complete, white = absent) in each microbial MAG for metabolic modules that showed differences across many of the MAGs. A full list of MAG metabolisms is in Table S7. Every MAG had some ammonification genes (Table S6). The figure was generated from anvi’o76.
Other microbial metabolisms that could be beneficial to P. oceanica included the presence of B vitamin metabolism on both tissue types at both vent and control areas. Vitamin B6 (pyridoxal) biosynthesis was only present in control rhizomes, while thiamin biosynthesis, vitamin B1, was present in all samples except vent rhizomes and vitamin B7 (biotin) biosynthesis was in multiple MAGS in both tissue types and sites. Vitamin B12 (cobalamin) synthesis was suggested in multiple MAGs at both sites, with aerobic biosynthesis only on the blades and anaerobic biosynthesis suggested in MAGs from all samples. Multiple MAGs on blades and rhizomes at both sites had sulfur oxidizing capability.
Discussion
Proximity to CO2 vents changed microbial processes in association with the seagrass P. oceanica, reducing rates of ammonification. Carbon and nitrogen uptake by POM was also increased at the vent sites. Thus, water column processes differed between the control and vent area, possibly due to microbial nitrogen processing in association with seagrass. Despite differences in plant morphology29,32 and grazing pressure30 at vent sites, we measured similar rates of carbon fixation by P. oceanica.
Benthic-pelagic coupling and enhancement of pelagic productivity through the host-associated microbes on P. oceanica have implications due to global declines in seagrass cover37 and the prevalence of seagrass wasting diseases38. Marine macrophytes have increasingly been shown to host a species rich microbiome39, with diverse functions that include not only ammonification, but also nitrogen fixation8,35,40, nitrification41 and nitrate reduction14,42. Seagrasses are important to carbon sequestration due to carbon fixation43 and the carbon storage in the root phyllosphere44 and sediment. Seagrasses also ameliorate pH decreases45. Here we show that P. oceanica enhanced carbon fixation in the water column by over eightfold at the control site and over 17-fold at the vent site, suggesting that the contribution of seagrass to the carbon cycle is likely greater than previously recognized due to water column effects. The presence of seagrass also enhanced water column nitrogen uptake to 2.5 times at the vent site and 1.6 times at the ambient site when we integrated over both day and night intervals (Fig. 2d). A possible reason for enhanced POM carbon and nitrogen uptake is the increased dissolved organics available to heterotrophic bacteria when P. oceanica is present (Fig. S2). Nitrogen cycling in association with P. oceanica has been shown previously to be elevated near vents35. P. oceanica mirrors the findings for other foundational species, such as corals and sponges4,46,47 where there are indirect effects through a food web due to the nutrient provisioning by particular species. Our findings with P. oceanica suggest that its role as a host for an active microbiome amplifies its role in these shallow Mediterranean systems.
Ammonification occurred during day and night and with or without P. oceanica, suggesting that there is an abundance of microbes that metabolize amino acids and enhance carbon fixation and primary productivity in seagrass ecosystems. Further, every genome that we assembled had the functional capacity for deamination of amino acids to ammonium (Table S6). The use of unfiltered, in situ seawater allowed us to quantify ammonification in the water column, both with and without P. oceanica. Bacteria in the water column was likely a significant contributor to ammonification. 15N uptake to water column POM was detected in all incubation bottles, though an unknown amount of this uptake could be due to bacteria or eukaryotic phototrophs.
Regardless of whether the 15N went into bacteria first, then into water column phytoplankton, the presence of Posidonia enhanced the uptake of 15N into POM, suggesting that seagrass-associated bacteria, or even fungi, enhance water column productivity both at vent and control sites. The comparison of light versus dark uptake of 15N into POM demonstrated that at least some of this uptake can be attributed to bacteria given the continued incorporation of 15N into POM at night (Fig. 2d).
One enigmatic aspect of our results is ammonification rates in units of nmol per hour, while 15NH4 uptake rates could be in umol per hour. We can think of several explanations for this, including that amino acid concentrations that were greater than the 1uM that we assumed would have overestimated enrichment (Rsource) and underestimated ammonification in Eq. (1). A second reason is that these processes could be spatially restricted, where microbial production in a biofilm could be immediately followed by host use, causing us to underestimate ammonification in the water column. Thirdly, if amino acid metabolisms are rapid and amino acid concentrations have high flux, we may have underestimated the processing and recycling of nitrogen during incubation. Ammonium concentrations are often not measured to the extent that nitrate concentrations are measured, despite preferential use of the former by macrophytes48,49. Yet, our estimates of ammonium uptake by P. oceanica during the day from the addition of 15N enriched amino acids ranged from 1.17 to 15.33 umol per g dry mass per hour, comparable to the 2.77 umol rate estimated in50 for P. oceanica in the Mediterranean and suggesting ammonification may contribute to ammonium fluxes and uptake rates here and in other areas.
Estimates of ammonium use and turnover in nature can be high50, though bacterial ammonification has been measured only rarely51. Also measured rarely is the concentration of DON, including any component parts such as amino acids. Amino acid concentrations have been estimated at 05–1.9 uM in other areas in the Mediterranean52,53, but a single flux estimate in the Caribbean suggests the flux could be high51. DON could be a persistent source of ammonium, and DON metabolisms warrant increased understanding of their contribution to nitrogen concentrations and fluxes.
Our demonstration of increased δ15N in seagrass tissue does not conclusively prove that these plants are taking up 15NH4 from microbial processes. Only pulse-chase experiments with more precise imaging, for example using NanoSIMS15,16, could show that microbial metabolism was the intermediate between 15N-amino acids and the higher δ15N of macrophyte tissue. However, ammonification genes were ubiquitous across MAGs here (Table S4) and in other studies14,54, and suggests that microbes may mediate the availability of ammonium. Several previous studies that suggest seagrasses directly take up DON did not control or account for microbial activity18, and thus direct DON uptake by marine angiosperms remains unknown.
Amino acid concentrations in the ocean are rarely reported, though there are several studies in the Mediterranean in proximity to P. oceanica beds that report 0.5 to 1.9 uM dissolved free amino acids (DFAA) concentrations52,53, several orders of magnitude greater than those reported in open ocean settings55,56. DFAA can be a reduced source of nitrogen for both phytoplankton and bacteria20, and may be rapidly renewed via the release from animals57,58, phytoplankton59, and possibly the host themselves, such as P. oceanica here52. Concentrations can be greater in the sediment surrounding P. oceanica when compared to the water column52. Although estimates are that DFAA is only from 1 to 10% of DON in coastal areas60, the estimates are few and DFAA and DON may have rapid turnover rates which make their contribution to overall nitrogen use difficult to estimate.
Posidonia oceanica has other means of acquiring dissolved inorganic nitrogen from microbial associates. Nitrogen fixation genes were found in MAGs from the rhizome at the control site, which is consistent with other studies that have identified nitrogen fixation61, nifH genes62, or nitrogen-fixing taxa in association with P. oceanica15 in the low-oxygen environment of the rhizomes. Mohr et al.15 assembled a MAG for a novel taxon that fixes nitrogen within the cells of P. oceanica roots. We did not find the ‘Candidatus Celerinatantimonas neptuna’ they described in any of the MAGs we assembled, nor in the family Celerinatantimonadaceae, perhaps due to either incomplete DNA sequencing or to our use of surface swabs rather than grinding the root tissue of P. oceanica, as they did. Our analysis of P. oceanica blades did not reveal nifH genes. Dissimilatory nitrate reduction to ammonium is an additional bacterial metabolism that may help the seagrass host access nitrogen. We found genes for this function in rhizomes in control areas, consistent with previous studies of denitrification in association with seagrass63.
Ambient nitrate values were similar between the two sites at the single timepoint when we estimated them; previously published values also cite similar low nitrate concentrations regardless of venting that are < 1 uM64,65. Both vent and control sites had microbes that could increase nitrogen to enhance P. oceanica primary production. In addition to the abundance of enzymes with deaminating function (Table S4), nitrate reduction genes were indicated, as was nitrogen fixation. Even though estimates of ammonification were greater in control areas, P. oceanica carbon fixation did not differ between the control area and the CO2 vent area. Carbon fixation rates were similar to the rates estimated at the same location previously8 for blades without epiphytes. The NPP values reported by Berlinghof et al. 8 in μmol O2 convert to ~ 0.7 to 0.9 mg C per g dry mass per hour, assuming a photosynthetic quotient of 1.0 (a 1:1 molar ratio of oxygen release to carbon uptake), a range nearly identical to what we estimated, though they show increased net primary production in proximity to vents. Here, our estimates of primary production included the rhizome tissue which may have obscured carbon fixation differences due to respiration.
There were microbial metabolisms that were unique to P. oceanica tissue types, such as cyanobacteria and anoxygenic photosynthesis only on seagrass blades. B vitamin synthesis was also prevalent, but the rhizomes in vent sites lacked Vitamin B1 and B6 synthesis. Vitamin B12, suggested to be auxotrophic for marine eukaryotic hosts66,67 was in some MAGs in both tissue types and at both control and vent sites. Whether it plays the critical role in host fitness that has been demonstrated for other marine macrophytes (e.g.1) remains to be investigated. In general, alpha diversity and the number of high quality MAGs was low in vent samples. Whether this reflects lower bacterial abundance in these areas is unclear, though certain functions were absent.
Our finding that P. oceanica and its microbial associates stimulate water column carbon fixation adds an important dimension to the role of seagrass beds in the carbon cycle. Seagrasses are suggested to alter the dissolved organic nitrogen51 and seaweeds can influence water column microbes68. As ocean acidification continues, our results suggest that microbial ammonification rates in association with seagrass will decrease. However, the demonstrated role that P. oceanica plays in enhancing carbon fixation by surrounding POM, particularly in low pH seawater, suggests an underappreciated role for resilient foundational species in a changing ocean.
Methods
Study system
Underwater CO2 vent systems are a powerful tool for exploring the effects of low pH on the features of species and biological systems. In Ischia, Italy, CO2 vent-influenced P. oceanica meadows (Castello South, “vent”, 40.73068047, 13.96303709) are separated by less than a km from areas unaffected by CO2 vents and (”control”, Sant' Anna, 40.73053186, 13.96102293). The vent sites at Castello South have a mean pH near 7.7, while the control pH site at Sant' Anna has ambient pH ~ 7.9528. Seawater temperature and other environmental variables (e.g. depth, light, salinity) are similar in CO2 vent and in control pH sites28. While carbon dioxide is the main gas from the vents, other gasses include N2 (3.2–6.6%), O2 (0.6–0.8%), Ar (0.08–0.1%), and CH4 (0.2–0.8%) but not sulfur gas32. Trace metal concentrations can be elevated near vents, although detrimental effects on P. oceanica have not been recorded69.
15N amino acid incubation with P. oceanica
P. oceanica was collected in situ at the control pH site at 1000 h and the CO2 vent site at 1040 h on 14 Sept 2021. At each site, two blades attached to a single rhizome unit were selected via SCUBA to have both a new blade developing as well as a corresponding older blade that was generally free of epiphytes. The rhizome was removed so that the blades (from 4 to 6) remained within a sheath. P. oceanica wet mass was quantified with a Pesola and ranged from 2.0 to 3.5 g (mean = 2.75 g). At each site, 8 P. oceanica shoots were collected. While we collected plants that were approximately a half meter distant from each other, we do not know if individuals were genetically distinct.
Simultaneously, gas-tight 500 ml polycarbonate bottles (actual volume is 660 ml) were filled at 1–2 m depth at each site, holding the bottle underwater above the P. oceanica beds, filling to eliminate all bubbles, and capping underwater. For each site, P. oceanica was immediately added to 8 of the bottles, while 4 held seawater only to quantify water column processes. Oxygen and temperature were recorded immediately with 3 mm optical fiber probe (#OXROB3, Pyroscience, Germany) and temperature sensor (Firesting FS02-4, Pyroscience, Germany) assuming 38 ppt Salinity.
We amended each of the 16 bottles with P. oceanica and the 8 with seawater only with amino acids that were enriched in 15N (98%, Cambridge Isotope, NLM-2151, Lot#PR-24163). We assumed a concentration of dissolved free amino acids (DFAA) of 1 uM based on previous work in Mediterranean Posidonia beds that ranged from 0.5 to 1.9 uM52,53. We added 50 uL of a 0.05 M solution to an assumed existing concentration of 1 uM to achieve an enrichment of ~ 0.76 at%. Each bottle was gently inverted 5 times to distribute the isotope, then attached to a line with dive weights at either end and deployed at 2 m depth at noon. All 32 bottles were deployed across 4 lines in the same embayment (Cartaromana Bay), approximately 0.5 km shoreward from where both sets of seagrasses originated, though in waters that were unaffected by the vent activity. The bottles rested on the benthos and they received natural light and some gentle water surge throughout the experiment.
To quantify nitrogen metabolic processes during the daylight versus darkness, we analyzed half of the P. oceanica and seawater contained within the bottles after 6 h at 1800 h; the remaining 16 experimental units were removed from the water immediately after sunrise the following day on 15 Sept 2021 (0720 h).
For each of the end of day and end of night censuses, the oxygen concentration in each bottle was measured and seawater was preserved for dissolved organic matter concentration, inorganic nutrient concentrations, including the concentration of 15N-ammonium. We also filtered 360–420 ml of seawater through a 0.7 uM gf/f filter (25 mm, Whatman) to quantify the 15N of particulate organic matter (POM) in the seawater and dried at 50 °C for 48 h. Dissolved inorganic nutrient samples were filtered through a 0.2 um PE filter and frozen at − 20 °C. Seawater samples for dissolved organic C and N analysis were filtered through pre-combusted GF/F filters into acid-washed HDPE vials, immediately fixed with 160 uL of 18.5% HCl and stored at 4 °C until analysis on a total organic carbon analyzer (TOC-L with TNM-L Unit, Shimadzu Corporation, Japan). DON was obtained from TDN subtracting total dissolved inorganic N (nitrate, nitrite and ammonium), determined on seawater samples preserved frozen and analyzed on a Continuous Flow Analyzer (Flowsys, SYSTEA SpA., Italy).
We analyzed the changes to both organic nutrient (DOC and DON) and inorganic nutrients (nitrate, nitrite, ammonium, phosphorus, silica) when incubation bottles had seawater only or P. oceanica, using a univariate three-way ANOVA, followed by two-way ANOVAs with site and treatment and their interaction for both daytime and nighttime.
Quantifying nitrogen dynamics
We quantified the amino acid mineralization rate (ammonification) using the source-sink model from Lipschultz70. The model quantifies the transfer from the source (15N amino acids) at the beginning of the experiment to the 'sink', which is the 15NH4 in the seawater, either at the end of the day or the end of the night. The rate from T0 to Tf was based on the following difference equation:
where R is the isotopic ratio of the sink or source, t is the length of the incubation, and \(\left[\overline{{NH }_{4}}\right]\) represents the average concentration of ammonium over the course of the experiment. By quantifying the metabolism of 15N from amino acids (the source) to ammonium in the water column (the sink), we estimated the hourly rate in nmol for the approximately 7 h daytime incubation, as well as the entire 20 h until the next morning. We then separated the ammonification rate for the night only by assuming the R(0)sink was the mean value at the end of the day. We quantified Eq. (1) for all 8 bottles with unfiltered seawater and 16 incubation bottles with both P. oceanica and unfiltered seawater. We used unfiltered seawater from immediately adjacent to the P. oceanica used in the experiment to quantify activity by water column versus host-associated microbes at each site.
Because we needed a concentration of 15NH4 in seawater that we could detect, given the relatively low concentrations of [NH4], we added 77.6 μL of 0.05 M NH4Cl, effectively adding 1.4 mg/L of NH4. The δ15NH4 value of the NH4Cl was known, allowing for reliable detection of ammonium without diluting the 15NH4 signal54.
We tracked the uptake of 15N to P. oceanica tissue to estimate ammonium uptake rates, following methods in71. This method uses the following equation:
where tissue 15N atom% excess is the final 15N atom% minus the initial 15N atom%, R is the mean 15N atom% enrichment in the NH4+ pool, t is the duration of the incubation, and NT is the amount of nitrogen in P. oceanica in μmols. We also used Eq. (2) to estimate 15N uptake by particulate organic matter (POM); for POM the 15N could have been in the form of ammonium or amino acids. The possible pathways for the addition of 15N amino acids across our incubations is illustrated in Fig. 1.
The percent carbon, nitrogen and δ15N and δ13C of P. oceanica tissue was measured from a 1 cm section of the midblade where we scraped off all epiphytes, as well as new meristematic tissue at the base that was always free of epiphytes. P. oceanica tissue was sampled prior to enrichment, at the end of the daylight hours, and following the nighttime period. The tissue was dried, ground to a fine powder in a GenoGrinder Spex (Metuchen, NJ), and analyzed on an elemental-analyzer–isotope-ratio mass spectrometer at Northwestern University Stable Isotope Biogeochemistry Laboratory (NUSIBL).
All statistical analyses were performed in R (www.R-project.org), version 4.2.2 (2022-10-31). For rate estimates that were highly variable and non-zero, we logged to reduce variance, prior to ANOVA.
Metagenome analyses
The surface microbiome of P. oceanica was sampled with a sterile cotton swab on the upper rhizome tissue; the mid blade region (corresponding to 2 above) and the older tissue with epiphytes (3. above) was sampled with both a cotton swab and Dentek brush to ensure a complete surface sample on blade tissue. All were placed in − 20 °C and shipped to storage at − 80° C, then shipped frozen to the University of Chicago. DNA was extracted with a Qiagen PowerSoil Kit and multiple samples were pooled for each of 6 metagenome samples to increase DNA quantity and possible discovery. The concentration of DNA was less on vent site samples compared with the control site, so we pooled all 6 sampled individuals into a single sample for each of the vent blade and vent rhizome. Thus, P. oceanica blade samples were pooled from 6 plants at the vent site and 3 plants each at control site; rhizome samples were also pooled from 6 plants at the vent site and 3 plants each at the control site. We thus had duplicated samples for control blades and rhizomes but only a single sample for vent tissue type.
The above 6 samples were run over 1 lane on a HiSeq 2500 (2 × 150) with TruSeq DNA library preps at Argonne National Laboratory. For each sample, resulting DNA sequences were first quality filtered72, then assembled with IDBA-UD v1.1.373 with a minimum scaffold length of 1 kbp. Metagenomic short reads from each sample were then recruited back to their corresponding assembled contigs using Bowtie274. Samtools75 was used to generate sorted and indexed BAM files. Anvi’o v7.076 was used as the command line environment for all downstream analyses. ‘anvi-gen-contigs-database’ was used to generate anvi’o contigs databases, during which Prodigal v2.6.377 identified open reading frames, and ‘anvi-run-hmms’ was used to identify genes matching to archaeal and bacterial single-copy core gene collections using HMMER78. We quantified overall alpha diversity with 'anvi-estimate-scg-taxonomy'. We then reconstructed genomes from the assembled metagenomes, we used a combination of automatic binning via CONCOCT v1.1.079, followed by a manual curation of each MAG that had greater than 50% completion, following guidance by80 Genome taxonomy was determined using the GTDB (v.1.3.0,81) database and 'anvi-run-scg-taxonomy'. We also inferred gene-level taxonomy using Centrifuge v1.0.482 to aid manual curation.
We examined the metabolic capability of the bacterial MAGs associated with P. oceanica using ‘anvi-estimate-metabolism’ on the manually refined bins that were > 50% complete and < 10% redundancy36. We classified a metabolic module as complete if 75% percent of the genes within the module were complete.
We further quantified the presence of genes that could increase host access to nitrogen. Our experiment specifically tested whether amino acid deamination occurred, so we searched within our assembled MAGs for ammonification hydrolases, including ureases and ammonia-lyases that cleave the C–N bonds in amino acids and make ammonium available to the host. We identified these enzymes through the Enzyme Commission numbers, and searched for EC numbers classified as acting on CH–NH2 bonds (EC:1.4.*), acting on carbon–nitrogen bonds other than peptides (EC:3.5.*), or ammonia lyases (EC:4.3.1*), where * indicates any subset of these classifications, and determined whether a MAG had each of these 3 classes of enzymes (e.g.14). The Kegg Orthologs83 associated with each of these EC numbers is shown in Supplemental Table 5. We searched for evidence of nitrogen fixation by assessing the completion of module 00175, as well as other nitrogen cycling genes and functions that could benefit the host.
All methods on the marine angiosperm, Posidonia oceanica, were carried out in situ and in accordance with local and national guidelines.
Data availability
Environmental data have been archived at Zenodo, https://doi.org/10.5281/zenodo.7834043;metagenomic data are archived at NCBI, http://www.ncbi.nlm.nih.gov/bioproject/967736 and at Figshare: https://figshare.com/collections/Posidonia_oceanica_metagenomics_and_ammonification_Scientific_Reports/6918289.
References
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Matthews, J. L. et al. Symbiodiniaceae-bacteria interactions: rethinking metabolite exchange in reef-building corals as multi-partner metabolic networks. Environ. Microbiol. 22, 1675–1687 (2020).
Wichard, T. et al. The green seaweed Ulva: A model system to study morphogenesis. Front. Plant Sci. 6, 72 (2015).
de Goeij, J. M. et al. Surviving in a marine desert: The sponge loop retains resources within coral reefs. Science 342, 108–110 (2013).
Dittami, S. M. et al. A community perspective on the concept of marine holobionts: Current status, challenges, and future directions. PeerJ 9, e10911 (2021).
Duffy, J. E. et al. Toward a coordinated global observing system for seagrasses and marine macroalgae. Front. Mar. Sci. 6, 317 (2019).
Shelton, A. O. Temperature and community consequences of the loss of foundation species: Surfgrass (Phyllospadix spp., Hooker) in tidepools. J. Exp. Mar. Biol. Ecol. 391, 35–42 (2010).
Berlinghof, J. et al. The role of epiphytes in seagrass productivity under ocean acidification. Sci. Rep. 12, 6249 (2022).
Pfister, C. A., Altabet, M. A. & Weigel, B. L. Kelp beds and their local effects on seawater chemistry, productivity, and microbial communities. Ecology https://doi.org/10.1002/ecy.2798 (2019).
Duarte, C. M. Reviews and syntheses: Hidden forests, the role of vegetated coastal habitats in the ocean carbon budget. Biogeosciences 14, 301–310 (2017).
Duarte, C. M., Losada, I. J., Hendriks, I. E., Mazarrasa, I. & Marbà, N. The role of coastal plant communities for climate change mitigation and adaptation. Nat. Clim. Change 3, 961–968 (2013).
Pfister, C. A. The evolutionary past and the uncertain future of foundational species. Proc. Natl. Acad. Sci. U.S.A. 119, e2211134119 (2022).
Crump, B. C., Wojahn, J. M., Tomas, F. & Mueller, R. S. Metatranscriptomics and amplicon sequencing reveal mutualisms in seagrass microbiomes. Front. Microbiol. 9, 388 (2018).
Miranda, K. et al. The diversity and functional capacity of microbes associated with coastal macrophytes. mSystems https://doi.org/10.1128/msystems.00592-22 (2022).
Mohr, W. et al. Terrestrial-type nitrogen-fixing symbiosis between seagrass and a marine bacterium. Nature https://doi.org/10.1038/s41586-021-04063-4 (2021).
Tarquinio, F. et al. Microorganisms facilitate uptake of dissolved organic nitrogen by seagrass leaves. ISME J. 12, 2796–2800 (2018).
Agawin, N. et al. Significant nitrogen fixation activity associated with the phyllosphere of Mediterranean seagrass Posidonia oceanica: First report. Mar. Ecol. Prog. Ser. 551, 53–62 (2016).
Vonk, J. A., Middelburg, J. J., Stapel, J. & Bouma, T. J. Dissolved organic nitrogen uptake by seagrasses. Limnol. Oceanogr. 53, 542–548 (2008).
Wang, Y. et al. RNA-based amplicon sequencing is ineffective in measuring metabolic activity in environmental microbial communities. Microbiome 11, 131 (2023).
Bronk, D. A., See, J. H., Bradley, P. & Killberg, L. DON as a source of bioavailable nitrogen for phytoplankton. Biogeosciences 4, 283–296 (2007).
Evans, S. M., Griffin, K. J., Blick, R. A. J., Poore, A. G. B. & Vergés, A. Seagrass on the brink: Decline of threatened seagrass Posidonia australis continues following protection. PLoS ONE 13, e0190370 (2018).
Krumhansl, K. A. et al. Global patterns of kelp forest change over the past half-century. Proc. Natl. Acad. Sci. 113, 13785–13790 (2016).
van Katwijk, M. M., Bos, A. R., Kennis, P. & de Vries, R. Vulnerability to eutrophication of a semi-annual life history: A lesson learnt from an extinct eelgrass (Zostera marina) population. Biol. Conserv. 143, 248–254 (2010).
Russell, B. D. et al. Future seagrass beds: Can increased productivity lead to increased carbon storage?. Mar. Pollut. Bull. 73, 463–469 (2013).
Weigel, B. L. & Pfister, C. A. The dynamics and stoichiometry of dissolved organic carbon release by kelp. Ecology 102, 03221 (2021).
Banister, R. B., Schwarz, M. T., Fine, M., Ritchie, K. B. & Muller, E. M. Instability and stasis among the microbiome of seagrass leaves, roots and rhizomes, and nearby sediments within a natural pH gradient. Microb. Ecol. https://doi.org/10.1007/s00248-021-01867-9 (2021).
Kroeker, K. J., Micheli, F., Gambi, M. C. & Martz, T. R. Divergent ecosystem responses within a benthic marine community to ocean acidification. Proc. Natl. Acad. Sci. U.S.A. 108, 14515–14520 (2011).
Teixidó, N. et al. Functional biodiversity loss along natural CO2 gradients. Nat. Commun. 9, 5149 (2018).
Mecca, S., Casoli, E., Ardizzone, G. & Gambi, M. C. Effects of ocean acidification on phenology and epiphytes of the seagrass Posidonia oceanica at two CO2 vent systems of Ischia (Italy). Mediterr. Mar. Sci. 21, 70 (2020).
Mirasole, A., Badalamenti, F., Di Franco, A., Gambi, M. C. & Teixidó, N. Boosted fish abundance associated with Posidonia oceanica meadows in temperate shallow CO2 vents. Sci. Total Environ. 771, 145438 (2021).
Scartazza, A. et al. Carbon and nitrogen allocation strategy in Posidonia oceanica is altered by seawater acidification. Sci. Total Environ. 607–608, 954–964 (2017).
Foo, S. A., Byrne, M., Ricevuto, E. & Gambi, M. C. The carbon dioxide vents of Ischia, Italy, a natural system to assess impacts of ocean acidification on marine ecosystems: An overview of research and comparisons with other vent systems. In Oceanography and Marine Biology (eds Hawkins, S. J. et al.) 237–310 (CRC Press, 2018). https://doi.org/10.1201/9780429454455-4.
Jackson, E. L., Rees, S. E., Wilding, C. & Attrill, M. J. Use of a seagrass residency index to apportion commercial fishery landing values and recreation fisheries expenditure to seagrass habitat service: Seagrass contribution to fishery value. Conserv. Biol. 29, 899–909 (2015).
Telesca, L. et al. Seagrass meadows (Posidonia oceanica) distribution and trajectories of change. Sci. Rep. 5, 12505 (2015).
Berlinghof, J. et al. Accelerated nitrogen cycling on seagrass leaves in a high-CO2 world. In review.
The Genome Standards Consortium et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Marbà, N., Díaz-Almela, E. & Duarte, C. M. Mediterranean seagrass (Posidonia oceanica) loss between 1842 and 2009. Biol. Conserv. 176, 183–190 (2014).
Groner, M. et al. Warming sea surface temperatures fuel summer epidemics of eelgrass wasting disease. Mar. Ecol. Prog. Ser. 679, 47–58 (2021).
Park, J., Davis, K., Lajoie, G. & Parfrey, L. W. Alternative approaches to identify core bacteria in Fucus distichus microbiome and assess their distribution and host-specificity. Environ. Microbiome 17, 55 (2022).
Capone, D. G. Nitrogen fixation (acetylene reduction) by rhizosphere sediments of the Eelgrass Zostera marina. Mar. Ecol. Prog. Ser. 10, 67–75 (1982).
Capone, D. G. & Taylor, B. F. Microbial nitrogen cycling in a seagrass community. In Estuarine Perspectives (ed. Kennedy, V. S.) 153–161 (Academic Press, 1980). https://doi.org/10.1016/B978-0-12-404060-1.50021-6.
Weigel, B. L., Miranda, K. K., Fogarty, E. C., Watson, A. R. & Pfister, C. A. Functional insights into the kelp microbiome from metagenome-assembled genomes. mSystems https://doi.org/10.1128/msystems.01422-21 (2022).
Duarte, C. M. & Krause-Jensen, D. Export from seagrass meadows contributes to marine carbon sequestration. Front. Mar. Sci. 4, 13 (2017).
Sogin, E. M. et al. Sugars dominate the seagrass rhizosphere. Nat. Ecol. Evol. https://doi.org/10.1038/s41559-022-01740-z (2022).
Hendriks, I. E. et al. Photosynthetic activity buffers ocean acidification in seagrass meadows. Biogeosciences 11, 333–346 (2014).
Allgeier, J. E., Burkepile, D. E. & Layman, C. A. Animal pee in the sea: consumer-mediated nutrient dynamics in the world’s changing oceans. Glob. Change Biol. 23, 2166–2178 (2017).
Rädecker, N., Pogoreutz, C., Voolstra, C. R., Wiedenmann, J. & Wild, C. Nitrogen cycling in corals: The key to understanding holobiont functioning?. Trends Microbiol. 23, 490–497 (2015).
Bracken, M. E. S. & Stachowicz, J. J. Seaweed diversity enhances nitrogen uptake via complementary use of nitrate and ammonium. Ecology 87, 2397–2403 (2006).
Herbert, R. A. Nitrogen cycling in coastal marine ecosystems. FEMS Microbiol. Rev. 23, 563–590 (1999).
Lepoint, G., Gobert, S., Dauby, P. & Bouquegneau, J.-M. Contributions of benthic and planktonic primary producers to nitrate and ammonium uptake fluxes in a nutrient-poor shallow coastal area (Corsica, NW Mediterranean). J. Exp. Mar. Biol. Ecol. 302, 107–122 (2004).
Smith, G. W. Influence of microbial deamination on ammonium pools in marine waters. Sci. Total Environ. 75, 319–324 (1988).
Jørgensen, N. O. G., Blackburn, T. H., Henriksen, K. & Bay, D. The importance of Posidonia oceanica and Cymodocea nodosa as contributors of free amino acids in water and sediment of seagrass beds. Mar. Ecol. 2, 97–112 (1981).
Velimirov, B. DOC dynamics in a Mediterranean seagrass system. Mar. Ecol. Prog. Ser. 28, 21–41 (1986).
Hochroth, A. & Pfister, C. A. Ammonification by kelp associated microbes increases ammonium availability. In review.
Keil, R. G. & Kirchman, D. L. Dissolved combined amino acids: Chemical form and utilization by marine bacteria. Limnol. Oceanogr. 38, 1256–1270 (1993).
Suttle, C. A., Chan, A. M. & Fuhrman, J. A. Dissolved free amino acids in the Sargasso Sea: Uptake and respiration rates, turnover times, and concentrations. Mar. Ecol. Prog. Ser. 70, 189–199 (1991).
Maas, A. E. et al. Migratory zooplankton excreta and its influence on prokaryotic communities. Front. Mar. Sci. 7, 573268 (2020).
Nagata, T. & Kirchman, D. L. Release of dissolved free and combined amino acids by bacterivorous marine flagellates. Limnol. Oceanogr. 36, 433–443 (1991).
Sarmento, H. et al. Phytoplankton species-specific release of dissolved free amino acids and their selective consumption by bacteria. Limnol. Oceanogr. 58, 1123–1135 (2013).
Sipler, R. E. & Bronk, D. A. Dynamics of dissolved organic nitrogen. In Biogeochemistry of Marine Dissolved Organic Matter 127–232 (Elsevier, 2015). https://doi.org/10.1016/B978-0-12-405940-5.00004-2.
Agawin, N. S. R., Ferriol, P., Sintes, E. & Moyà, G. Temporal and spatial variability of in situ nitrogen fixation activities associated with the Mediterranean seagrass Posidonia oceanica meadows: Nitrogen fixation in Posidonia oceanica. Limnol. Oceanogr. 62, 2575–2592 (2017).
Garcias-Bonet, N., Arrieta, J. M., Duarte, C. M. & Marbà, N. Nitrogen-fixing bacteria in Mediterranean seagrass (Posidonia oceanica) roots. Aquat. Bot. 131, 57–60 (2016).
Welsh, D. et al. Denitrification in an intertidal seagrass meadow, a comparison of 15 N-isotope and acetylene-block techniques: Dissimilatory nitrate reduction to ammonia as a source of N2O?. Mar. Biol. 139, 1029–1036 (2001).
Kumar, A. et al. Ocean acidification affects biological activities of seaweeds: A case study of Sargassum vulgare from Ischia volcanic CO2 vents. Environ. Pollut. 259, 113765 (2020).
Lorenti, M., Buia, M. C., Di Martino, V. & Modigh, M. Occurrence of mucous aggregates and their impact on Posidonia oceanica beds. Sci. Total Environ. 353, 369–379 (2005).
Brawley, S. H. et al. Insights into the red algae and eukaryotic evolution from the genome of Porphyra umbilicalis (Bangiophyceae, Rhodophyta). Proc. Natl. Acad. Sci. U.S.A. 114, E6361–E6370 (2017).
Helliwell, K. E., Wheeler, G. L., Leptos, K. C., Goldstein, R. E. & Smith, A. G. Insights into the evolution of vitamin B12 auxotrophy from sequenced algal genomes. Mol. Biol. Evol. 28, 2921–2933 (2011).
Chen, M. Y. & Parfrey, L. W. Incubation with macroalgae induces large shifts in water column microbiota, but minor changes to the epibiota of co-occurring macroalgae. Mol. Ecol. 27, 1966–1979 (2018).
Mishra, A. K., Santos, R. & Hall-Spencer, J. M. Elevated trace elements in sediments and seagrasses at CO2 seeps. Mar. Environ. Res. 153, 104810 (2020).
Lipschultz, F. Isotope tracer methods for studies of the marine nitrogen cycle. In Nitrogen in the Marine Environment 2nd edn (eds Capone, D. G. et al.) 1345–1384 (Academic Press, 2008).
Pather, S., Pfister, C. A., Post, D. M. & Altabet, M. A. Ammonium cycling in the rocky intertidal: Remineralization, removal, and retention. Limnol. Oceanogr. 59, 361–372 (2014).
Minoche, A. E., Dohm, J. C. & Himmelbauer, H. Evaluation of genomic high-throughput sequencing data generated on Illumina HiSeq and genome analyzer systems. Genome Biol. 12, R112 (2011).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Eren, A. M. et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat. Microbiol. 6, 3–6 (2021).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Alneberg, J. et al. Binning metagenomic contigs by coverage and composition. Nat. Methods 11, 1144–1146 (2014).
Shaiber, A. et al. Functional and genetic markers of niche partitioning among enigmatic members of the human oral microbiome. Genome Biol. 21, 292 (2020).
Parks, D. H. et al. GTDB: An ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab776 (2021).
Kim, D., Song, L., Breitwieser, F. P. & Salzberg, S. L. Centrifuge: Rapid and sensitive classification of metagenomic sequences. Genome Res. 26, 1721–1729 (2016).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y., Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic. Acids Res. 45, D353–D361. https://doi.org/10.1093/nar/gkw1092 (2017).
Acknowledgements
We thank the University of Chicago's France and Chicago Collaborating in the Sciences (FACCTS), and the National Research Agency Investments for the Future (“4Oceans-Make Our Planet Great Again” grant, ANR-17-MOPGA-001) for funding. We thank Pietro Sorvino and family for hospitality and logistical support throughout the study. S. Owens and S. Greenwald at Argonne National Lab facilitated metagenome sequencing. We thank Francesca Margiotta for the inorganic and organic nutrient analyses, M. Khamla for assistance with Fig. 1, and J. Berlinghof and 2 anonymous reviewers for comments that improved the manuscript.
Funding
This study was funded by The University of Chicago's France and Chicago Collaborating in the Sciences (FACCTS) and the National Research Agency Investments for the Future ("4Oceans-Make Our Planet Great Again" grant, ANR-17-MOPGA-001).
Author information
Authors and Affiliations
Contributions
C.A.P., N.T., U.C., A.M., J.P.G. designed the study and C.A.P., N.T., U.C., A.M., L.M. carried out all field and laboratory analyses. C.A.P., U.C., I.V. analyzed data. All individuals contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pfister, C.A., Cardini, U., Mirasole, A. et al. Microbial associates of an endemic Mediterranean seagrass enhance the access of the host and the surrounding seawater to inorganic nitrogen under ocean acidification. Sci Rep 13, 19996 (2023). https://doi.org/10.1038/s41598-023-47126-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-47126-4
This article is cited by
-
Accelerated nitrogen cycling on Mediterranean seagrass leaves at volcanic CO2 vents
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.