Abstract
Real-time quantitative polymerase chain reaction (RT-qPCR) is the most common method to determine mRNA expression, and Minimum Information for Publication of RT-qPCR Experiments (MIQE) proposes that a panel of reference genes for RT-qPCR is conducive to obtaining accurate results. This study aimed to screen and verify the optimal panel of reference genes in colorectal cancer (CRC) and normal colonic cell lines. In the study, eight candidate reference genes (GAPDH, ACTB, 18S, PPIA, B2M, SDHA, GUSB, and YWHAZ) were selected for RT-qPCR to detect their expression in NCM460, HT29, HCT116, SW480, SW620, DLD-1, LOVO and RKO cell lines. The stability of reference genes and the optimal panel were evaluated by geNorm, NormFinder, and BestKeeper software. As results, the expression levels of candidate reference genes differed in the colonic epithelial cell lines, and the number of optimal panel of reference genes is two. B2M and YWHAZ were the two most stable reference genes for NCM460, HCT116, SW620, LOVO, and RKO cell lines, while only one of B2M and YWHAZ was most stable in HT29 and SW480 cells. In DLD-1 cells, the stability of B2M and YWHAZ ranked 3rd and 6th, PPIA and GUSB were the most stable two. Furthermore, the YWHZA + B2M performed smaller intragroup differences than other panel or single reference gene. In conclusion, this study indicates the optimal panel of reference genes is YWHZA + B2M for the NCM460, HCT116, SW620, LOVO, RKO, SW480, and HT29 cell lines, but it is PPIA + GUSB in DLD-1 cell lines.
Similar content being viewed by others
Introduction
Real-time quantitative polymerase chain reaction (RT-qPCR) is currently frequently used for gene expression quantification, due to the advantage of rapidity, accuracy and sensibility1,2,3. A low number copies of target gene sequence are amplified based on the PCR technique which can’t be quantify by primary total RNA content2. Therefore, in order to obtain accuracy gene expression level of target gene, it’s important to use reference genes to normalize RT-qPCR data, which are supposed to stably express in cells or tissues under different experimental conditions4,5,6,7. Currently, housekeeping gene (HKG), including glyceraldehyde 3-phosphate dehydrogenase (GAPDH), beta actin (ACTB) as well as the 18S rRNA gene, are widely used for RT-qPCR as the reference genes8,9,10. However, it has been validated that HKG expression may be affected by different experimental conditions and pathological conditions such as bowel inflammation or cancer6,11,12. As a sensitive quantitative method, target gene expression may be incorrectly estimated as a result of variable reference genes. Therefore, screening and verifying reference genes that are stably expressed in different conditions is much more important.
Colorectal cancer (CRC) is the third most common cancer and the second cause of cancer death worldwide, RT-qPCR is widely used in the etiology, pathogenesis, pathological changes, and treatment of CRC13,14. However, it has been validated that ACTB and B2M in CRC can be affected by inflammation and be related to CRC stage, suggesting the expression of HKG are variable in CRC11. As a result, the differential gene expression of CRC will be wrongly estimated by RT-qPCR approach, owing to the use of reference genes that are not validated in CRC and relative cell lines. At present, there are several studies have aimed to demonstrate the best reference genes derived from CRC tissues and matched colonic epithelium. In a recent study held by the team of Malgorzata, 166 samples (35 CRC, 91 normal and 40 inflamed) were adopted and validated by geNorm and NormFinder, results showed that PPIA was the top-ranked reference gene for CRC tissue. And the pair RPLPO + PPIA was recommended as the best reference gene to normalize the RT-qPCR data for tissues from CRC and IBD. Moreover, ACTB, which is often used before was not recommended as a reference gene11. Other studies similar to Malgorzata, RPLPO, PPIA, HPRT1, and IPO8 are recommended as optimal reference genes11,15,16,17. However, there is no recommendation for commonly used in CRC research as candidate reference genes in different cell lines. Moreover, Minimum Information for Publication of RT-qPCR (MIQE) has shown that the use of a stable panel of reference genes, combination of more than 1 reference gene, is needed to obtain authentic results18,19, but there is no related report on human colonic epithelial and CRC cell lines.
In this study, the expression of eight candidate genes commonly used in CRC research (by consulting the literature) was assayed by RT-qPCR, and the stability of candidate genes and the optimal panel of reference genes were analyzed GeNorm, NormFinder and BestKeeper in seven CRC cell lines and one human normal colonic epithelial cell line. Furthermore, the reliability of the selected panel of reference genes was verified by normalizing the expression of APC, which is well known that the mRNA expression of APC is significantly decreased in HCT116 cells20. This study screened and validated the optimal panel of reference genes in the human normal colonic epithelial cell and CRC cell lines, which provides a methodological basis for studies related to CRC.
Materials and methods
Chemicals and reagents
TRIzol kit (CAS No. 5346994) was purchased from Invitrogen Co. (15596-026, Carlsbad, CA, USA); HiScript III RT SuperMix for qPCR (+ gDNA wiper) (CAS R323-01) and Taq Pro Universal SYBR qPCR Master Mix kits (CAS Q712-02) were purchased from Vazyme Biotech Co., Ltd. (Nanjing, China); All primers were designed and synthesized by Sangon Biotechnology Co., Ltd. (Shanghai, China). Isopropanol (CAS No. 67-63-0) and chloroform (CAS No. 67-66-3) were procured through Merck (Beijing, China). The other chemicals and agents were analytical grade.
Cell culture
Human CRC cell lines HT29 (HTB-38), HCT116 (CCL-247EMT), SW480 (CCL-228), SW620 (CCL-227), DLD-1 (CCL-221), and LOVO (CCL-229) were purchased from ATCC; RKO was obtained from Procell (CL-0196, China); and NCM460 was provided by Cellverse (iCell-h373, China). HT29, HCT116, DLD-1, LOVO, and NCM460 were cultured in RPMI1640 (SH30809.01, HyClone, USA), and SW480, SW620, and RKO were grown in DMEM medium (C11995500BT, Gibco, USA). The medium was supplemented with 10% fetal bovine serum (FBS) (C0235, Gibco, AUS) and penicillin–streptomycin 1 × (C0222, Beyotime, China). All cells were maintained at 37 °C, 5% CO2 in a humidified atmosphere. Cells less than passage 10 were harvested at 90% confluence for further experiments.
RNA extraction, integrity test and cDNA synthesis
The total RNA of cells was extracted using TRIzol Reagent (15596-026, Invitrogen, USA). In brief, the cells were washed with PBS, and 1 mL of trizol was added to the cell culture plastic. By pipetting 3–5 times, cell lysis was collected into 1.5 mL Centrifuge tubes (enzyme-free) and allowed to stand at room temperature for 10 min. Subsequently, 200 μL chloroform was added and mixed by a vortex. After placed for 5 min on ice, the samples were centrifuged in a centrifuge (Centrifuge 5430R, Eppendorf, Germany) at 12,000 rpm for 15 min at 4 °C. The supernatant (400 μL) of the upper layer was transferred into a new EP tube and blended with an equal volume of isopropyl alcohol, and then the mixture was allowed to precipitate at room temperature for 10 min. The samples were then centrifuged at 12,000 rpm for 10 min at 4 °C to precipitate the total RNA, which was washed with 1 mL of precooled 75% ethanol twice with a centrifuge at 9500 rpm for 5 min at 4 °C and dissolved with enzyme-free water. NanoDropOne (ThermoFisher Scientific, USA) was used to assay the RNA concentration and purity. It suggested the RNA purity was acceptable when A260/280 was between 1.8 and 2.1.
RNA integrity was assayed by agarose gel electrophoresis21. Briefly, 1% agarose gels containing 3 μL of 10 mg/ml ethidium bromide in each gel were used for most RNA integrity assessments. Marker (DL5000, Vazyme Biotech Co., Ltd., Nanjing, China) and 2 μL total RNA samples mixed with 2 × RNA loading buffer (R0215, Beyotime™, Shanghai, China) at a ratio of 1:1 were added to into separate wells of the gel. Electrophoresis was performed in TAE buffer at a condition of 100 voltage, and stopped at 2–3 cm from the edge of the gel. The gels were observed and photographed using a gel imaging system (JY04S-3C, Beijing JUNYI Electrophoresis Co., Ltd, China). 28S:18S band intensity ratios were calculated by Image J (n = 3 × 3), and ratios higher than 1:1 were considered eligible samples22. The eligible RNA samples were used to synthesize cDNA by HiScript III RT SuperMix for qPCR (+ gDNA wiper) kits (R323-01, Vazyme, China) following the manufacturer’s protocol. The genomic DNA was first removed by a mixture of 4 μL 4 × gDNA wiper Mix, 1 μg RNA and RNase-free ddH2O (added to 16 μL) and incubating at 42 °C for 2 min by a PCR instrument. Then, 4 μL 5 × HiScript II qRT SuperMix II was added for the reverse transcription step, which was performed at 50 °C for 15 min followed by 85 °C for 5 s. Finally, the samples were diluted with enzyme-free water at 1:5 and stored at − 20 °C for further analysis.
Candidate reference genes, primers design, and RT-qPCR
As shown in Table S1, 8 reference genes (GAPDH, ACTB, 18S, PPIA, B2M, SDHA, GUSB, YWHAZ) commonly used in colorectal research were selected as candidate genes by consulting the literature. The complete cDNA sequence of the candidate genes was obtained from NCBI's Entrez Nucleotides database. Primer Premier 5.0 (PREMIER Biosoft International) was used for primer design, and then the designed primers were input into the BLAST database for homology comparison. Melting curve and standard curve were used to verify primer specificity and amplification efficiency. As previously described23, the standard curve reaction program is at 95 °C for 10 min, 95 °C for 15 s and 60 °C for 1 min for 40 cycles, and the melting curve is at 95 °C for 15 min, 60 °C for 30 s, and 95 °C for 15 min. After sample gradient dilution of 28 times, the standard curve was analyzed by StepOnePlus™ Real-Time PCR System (AppliedBiosystems by ThermoFisher Scientific™, USA). The amplification efficiency (E) of primers was calculated by using the slope and R2 of the standard curve, which was acquired according to \(E(\%)=({10}^{-\frac{1}{slope}}-1)\times 100\). The specific primer sequences were shown in Table S2.
The expression levels of candidate genes were detected by RT-qPCR in NCM460, HT29, HCT116, SW480, SW620, DLD-1, LOVO and RKO cell lines using StepOnePlus™ Real-Time PCR System according to the manufacturer's protocol of Taq Pro Universal SYBR qPCR Master Mix kits. The 10 μL reaction system, containing 0.2 μL forward and reversed primers, 1 μL cDNA, 3.6 μL ddH2O and 5 μL SYBR, was used in the RT-qPCR assay of this study. The amplification curve reaction program was at 50 °C for 2 min, 95 °C for 2 min, 95 °C for 15 s and 60 °C for 1 min for 40 cycles. The 2−ΔΔCt mode in StepOnePlus™ Real-Time PCR System was used to obtain the Ct values of the candidate reference genes and the 2−ΔΔCt method was used for the statistical analysis of APC gene expression differences in the validation experiments. A single reference gene or the geometric mean of screened reference gene panel was used to normalize APC expression levels24. Three technical repetitions were performed in this study.
Software
GeNorm (version 3.5, http://medgen.ugent.be/~jvdesomp/genorm/)25,26, NormFinder (http://www.mdl.dk/publicati-onsnormfinder.htm)26,27, and BestKeeper (http://www.wzw.tum.de/gene-quantification/bestkeeper.html)28. are three commonly used reference gene stability analysis software based on statistics. GeNorm determines the stability of the reference gene by the average stability value of M. The smaller the M value is, the more stable the reference gene expression is. When M is less than 1.5, it indicates that the expression of the gene is stable, which can be used as a reference gene. In addition, geNorm can also calculate the optimal number of reference genes by judging the pairing coefficient of variation (the V values) of standardization factor after introducing a new reference gene. When Vn/(n + 1) < 0.15, the number is n23. BestKeeper directly analyzes the standard deviation (SD), correlation coefficient (r), and coefficient of variation (CV) of circulation threshold (Ct) value to judge the stability of gene expression; the smaller the SD and CV, the more stable the reference gene expression was; when the SD value is more than 1, it indicates that the gene expression is unstable and can’t be used as a reference gene29,30,31. The same as geNorm, normFinder obtains the average stable value (M) of gene expression by calculating the within-group and between-group variance of Ct value. The smaller the m value is, the more stable the gene expression is, but only one of the most stable reference genes can be screened29,30,31. In this study, the most stable reference genes in normal colonic epithelial cells and colonic cancer cell lines were analyzed by the above three software. The comprehensive ranking was obtained by calculating the geometric mean of each software ranking, and finally the optimal panel of reference gene in each cell line was screened.
Statistical analysis
Data statistics and graphing were operated in Prism 8 (Graph Pad, USA). Quantitative data were reflected as the mean ± SEM. An unpaired two-tailed Student's t-test was used for statistical test. P < 0.05 was considered statistically significant.
Results
RNA integrity and the specificity and amplification efficiency of the candidate gene primers
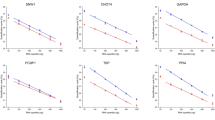
The integrity of total RNA samples, the specificity and the amplification efficiency of PCR primers are important factors affecting the accuracy of RT-qPCR results. Therefore, the above factors are verified first. The agarose gel electrophoresis showed that the integrity of total RNA was eligible (Fig. 1A, S1). All melt curves showed a single peak (Fig. 1B–I), indicating all the selected reference genes primers showed a good specificity. The standard curves showed R2 ≥ 0.993, suggesting there was a good linear relationship between template cDNA and CT values (Fig. 1J). Moreover, The amplification efficiency of the primers calculated by slope and R2 showed that the values were 97–103%, which is in line with the RT-PCR principle of MIQE32.
The integrity of total RNA and the specificity and amplification efficiency of the primers. (A) The cropped gels showing the total RNA integrity by RNA gel electrophoresis; The melt curves of GADPH (B), ACTB (C), 18S (D), PPIA (E), SDHA (F), GUSB (G), B2M (H) and YWHAZ (I); (J) The amplification efficiencies and standard curve parameters of the candidate reference genes.
Basic expression levels of the candidate reference genes in different cells
The reference genes have stable expression, and the general level is high. Our results showed that the average Ct values of the candidate genes were between 8 and 27 in the human colonic epithelial cell (NCM460) and cancer cell lines (HT29, HCT116, SW480, SW620, DLD-1, LOVO and RKO), and the expression of 18S was the highest while GUSB was the lowest in all cells except for LOVO cell line (Fig. 2A–H). Further analysis showed that the expression levels of reference genes in different cells are inconsistent. For example, the expression of YWHAZ in NCM460 cells is lower than that in B2M, which is the opposite in HT29 cells (Fig. 2A, B), indicating the stability of reference gene expression may be changed in human normal colonic epithelial cells and cancer cells.
The stability analysis of the candidate reference genes in colonic epithelial cells
First, we analyzed the stability of reference genes in normal colonic epithelial cell lines (NCM460). GeNorm showed the stability of the candidate reference genes from high to low was B2M > 18S > YWHAZ > SDHA > GAPDH > ACTB > GUSB > PPIA (Fig. 3A); NormFinder showed the order was B2M > 18S > GAPDH > YWHZA > SDHA > ACTB > GUSB > PPIA (Fig. 3B); BestKeeper showed the order was B2M > YWHAZ > SDHA > GAPDH > ACTB > PPIA > 18S > GUSB (Fig. 3C).
Stability of each reference gene in normal epithelial cells. The stability of the candidate reference genes was analyzed by geNorm (A), normFinder (B) and BestKeeper (C); (D) The comprehensive stability ranking of the candidate reference genes; (E) Paired coefficient of variation (CV) of standardization factor’s variation after introducing a new reference gene in NCM460 cells.
We calculated the geometric mean of ranking in three software to further analyze the comprehensive rank of reference genes in normal colonic epithelial NCM460 cell line. The results showed the comprehensive ranking of the most to least stable genes in NCM460 cells were B2M > YWHAZ > 18S > SDHA > GAPDH > ACTB > PPIA > GUSB (Fig. 3D). Furthermore, the paired coefficients of standardization factor’s variation after introducing a new reference gene calculated by geNorm showed V2/3 were less than 0.15 (Fig. 3E), suggesting the best number of optimal reference genes is two. Therefore, B2M + YWHAZ was the optimal panel of reference gene in NCM460 cell line.
The stability analysis of the candidate reference genes in colonic cancer cell lines
We further analyzed the stability of reference genes in colonic cancer cell lines (HT29, HCT116, SW480, SW620, DLD-1, LOVO and RKO). In HT29 cell line, geNorm showed the most to least stable genes were B2M > SDHA > 18S > YWHAZ > ACTB > GAPDH > GUSB > PPIA (Fig. 4A); normFinder showed the order was SDHA > B2M > 18S > YWHAZ > ACTB > GAPDH > GUSB > PPIA (Fig. 4B); BestKeeper showed the order was B2M > SDHA > 18S > ACTB > YWHAZ > GAPDH > GUSB > PPIA (Fig. 4C). In HCT116 cell line, geNorm showed the ranking of most to least stable genes were B2M > YWHAZ > PPIA > GUSB > 18S > GAPDH > SDHA > ACTB (Fig. 4D); normFinder showed the order was YWHAZ > PPIA > SDHA > B2M > 18S > GAPDH > GUSB > ACTB (Fig. 4E); BestKeeper showed the order was B2M > GUSB > YWHAZ > PPIA > SDHA > GAPDH > 18S > ACTB (Fig. 4F). In SW480 cell line, geNorm showed the ranking was GAPDH > YWHAZ > B2M > 18S > GUSB > ACTB > PPIA > SDHA (Fig. 4G); normFinder showed the order was YWHAZ > 18S > GAPDH > B2M > GUSB > ACTB > PPIA > SDHA (Fig. 4H); BestKeeper showed it was GAPDH > PPIA > GUSB > B2M > YWHAZ > ACTB > 18S > SDHA (Fig. 4I). In SW620 cell line, geNorm showed the ranking of most to least stable genes were B2M > YWHAZ > SDHA > 18S > GAPDH > GUSB > PPIA > ACTB (Fig. 4J); normFinder showed the ranking was SDHA > YWHAZ > B2M > 18S > GUSB > GAPDH > PPIA > ACTB (Fig. 4K); BestKeeper showed the ranking was YWHAZ > B2M > SDHA > PPIA > 18S > GUSB > GAPDH > ACTB (Fig. 4L).
In DLD-1 cell line, geNorm showed the ranking was PPIA > GUSB > B2M > 18S > SDHA > YWHAZ > ACTB > GAPDH (Fig. 5A); normFinder showed the ranking was PPIA > B2M > GUSB > 18S > SDHA > YWHAZ > ACTB > GAPDH (Fig. 5B); BestKeeper showed the ranking was PPIA > GUSB > B2M > SDHA > YWHAZ > GAPDH > 18S > ACTB (Fig. 5C). In LOVO cell line, geNorm showed the ranking was YWHAZ > B2M > GUSB > 18S > SDHA > PPIA > GAPDH > ACTB (Fig. 5D); normFinder showed the ranking was B2M > YWHAZ > GUSB > 18S > PPIA > SDHA > GAPDH > ACTB (Fig. 5E); BestKeeper showed it was B2M > YWHAZ > GUSB > 18S > SDHA > PPIA > GAPDH > ACTB (Fig. 5F). In RKO cell line, geNorm showed the ranking was B2M > YWHAZ > SDHA > GUSB > GAPDH > 18S > ACTB > PPIA (Fig. 5G); normFinder showed it was B2M > YWHAZ > SDHA > GAPDH > GUSB > 18S > ACTB > PPIA (Fig. 5H); BestKeeper showed it was B2M > SDHA > YWHAZ > GUSB > GAPDH > 18S > ACTB > PPIA (Fig. 5I).
Similarly, we further calculated the comprehensive ranking of the candidate reference genes. The ranking was shown in Table 1. The two most stable reference genes B2M and YWHAZ in normal colon epithelial cell lines were also the most stable in HCT116, SW620, LOVO and RKO cell lines, while only one of B2M and YWHAZ was most stable in HT29 and SW480 cells. In DLD-1 cell line, the stability of B2M and YWHAZ ranked 3rd and 6th respectively (Table 1). The same as NCM460, the paired coefficients of standardization factor’s variation after introducing a new reference gene V2/3 in all colonic cancer cell lines was less than 0.15 (Table S3), suggesting the number of optimal panel of reference gene was two. Due to the stability values M of YWHAZ and B2M were less than 1.5 and the stability ranked in the top four in HT29 and SW480 cell lines, B2M + YWHAZ can also be used as the optimal panel of reference gene in HT29, HCT116, SW480, SW620, LOVO and RKO cell line. But PPIA + GUSB was the optimal panel of reference gene in DLD-1 cell line.
Verification of stability and accurateness of the selected reference genes
To further verify the stability and believability of the selected panel of reference gene, we selected HCT116 cell line for further experiment. APC is a well-known tumor suppressor gene. Various experiments have suggested that the mRNA expression of APC in HCT116 cells is significantly decreased20. First, we standardized APC expression with GAPDH and/or ACTB, and the results showed that there was a more significant statistical difference normalized with the panel of reference genes (Fig. 6A–C). We then normalized APC expression with the screened reference genes (B2M and YWHAZ). The results showed that, consistently, there was also a more significant statistical difference normalized with the panel of reference genes compared to a single reference gene (Fig. 6D–F). Moreover, whether screened single or panel of reference genes, the results indicated a more significant statistical difference normalized with B2M and YWHAZ compared to GAPDH and ACTB (Fig. 6A–F).
Discussion
RT-qPCR is a robust tool for biomedical research, providing a sensitive and quantification method to determine gene expression. At present, geNorm, NormFinder, and BestKeeper are most common software used for gene expression stability analysis29,30,31. In this study, we use the three above software to estimate the stability of eight candidate reference genes in normal human colonic epithelial cell and seven CRC cell lines. Due to different algorithms in three tools, there was slight discrepancy in the stability ranking of candidate genes. For example, B2M was the best suitable reference gene in NCM460, HT29, HCT116, SW620, RKO cell lines according to geNorm. However, NormFinder showed that SDAH was the most stable reference gene for HT29 and SW620 cell lines, and B2M ranked second and third, respectively. YWHAZ ranked first in HCT116 and SW620 cell lines according to BestKeeper. PPIA was the best suitable reference for DLD-1 cell line according to three tools, but it was the lowest stable in all tools for RKO cell line. Therefore, comprehensive ranking was made, and YWHAZ and B2M were the two most stable reference genes in NCM460, HCT116, SW620, LOVO, and RKO cell lines, respectively, while only one of B2M and YWHAZ was the most stable in HT29 and SW480 cells. However, PPIA and GUSB were just the two most stable reference genes in DLD-1 cell line.
The optimal reference gene may change in different tissues, experiment designs, or diseases, even vary in cell lines and tissues of the same tumor25,33,34. Most recent research from the team of Sørby reported that PPIA was the most optimal reference gene to normalize gene expression in colon cancer even metastatic colon cancer15. In their research, PPIA was one of top two stable reference genes by using geNorm, and GUSB was ranked second by CtCV%, which was in concordance with our findings of DLD-1 cell line. However, our result showed that GUSB and PPIA was not the most stable reference gene when compared with other genes except in DLD-1 cell line. To our surprise, PPIA even was the lowest stable reference gene in some cell lines, such as RKO and HT29. Same as GUSB, in most cell lines, it was not also the most stable reference gene. This difference may be explained by different experiment designs. In Sørby’s research, they included tissues both cancer and matched normal mucosa from 18 non-metastatic colon cancer patients and 20 metastatic cancer patients, while we tested in diverse colon cancer cell lines. Furthermore, the percentage of cancer cells in the tissues they accepted was not assessed, indicating that total RNA was mixed with other RNA from colonic cells. Another study suggested that the panel of B2M was the most suitable reference genes to normalize RT-qPCR data in CRC tissue35. Identically, our result showed that B2M was also the most stable reference gene in majority cell lines, especially in NCM460, RKO, and LOVO. Although the study demonstrated that the expression of B2M gene was variable15, our result showed that B2M ranked top three in all cell lines, which further indicates that B2M is stably expressed in human colonic epithelial and colorectal cancer cell lines.
According to the Minimum Information for Publication of PCR Experiments (MIQE) guidelines in 2013 and updated in 202018,36, it is suggested that the accuracy of the RT-qPCR results can be affected by using only one single reference gene to standardize the expression of genes. In addition, Vandesompele et al. found that the error of using a single reference gene was 6.5-fold compared with using multiple genes data for normalization25. Therefore, to ensure the believability of RT-qPCR, it is important to screen the stable panel of reference genes in human colonic epithelial and colorectal cancer cell lines. Studies suggested that when Vn/(n + 1) < 0.15 in geNorm, the number of optimal panel of reference genes is n23. Our result showed that V2/3 in all cell lines we selected was less than 0.15, suggesting that the number of optimal panel of reference genes was two. According to the comprehensive ranking, YWHZA + B2M was naturally the optimal panel of reference genes in NCM460, HCT116, SW620, LOVO, and RKO cell lines, while it was PPIA + GUSB in DLD-1 cell line. In addition, because the stability values M of YWHAZ and B2M were less than 1.5 and one of B2M and YWHAZ was most stable and the stability ranked in the top four in HT29 and SW480 cells, it is worth noting that, considering the experimental simplicity, repeatability and economy, YWHZA + B2M can be also used as the optimal panel of reference gene in HT29 and SW480 cell lines.
Previous studies showed that transcriptomic and proteomic expression levels of APC also significantly reduced in HCT116 cell lines20. Therefore, APC expression was normalized by different reference genes in HCT116 cell lines to further verify the believability of reference genes screened in this study. The results showed there was more significant statistical difference in APC expression normalized by the most stable reference screened in this study than the commonly used reference genes. Moreover, there was a more significant statistical difference in APC expression normalized by whether the commonly used panel of reference genes (GAPDH + ACTB) or the optimal panel of reference genes (YWHZA + B2M) screened in this study than that of a single reference gene among them. These results suggested that the selected the optimal panel of reference genes reduced the differences within the group and was more conducive to obtaining an authentic experimental conclusion.
Conclusion
In this study, to identify the optimal panel of reference genes to normalize RT-qPCR data in CRC and colonic epithelial cell lines, we applied geNorm, NormFinder, and BestKeeper to estimate the stability of eight reference genes. Our finding showed that YWHAZ + B2M is the optimal panel of reference genes in the normal human colonic epithelial cell lines (NCM460) and some CRC cell lines (HCT116, SW620, LOVO, RKO, SW480, and HT29), while it is PPIA + GUSB in DLD-1 cell lines. This study provided a theoretical and experimental basis for the accuracy of studies on CRC.
Data availability
Data generated by this study could be asked from corresponding authors with reasonable request.
Abbreviations
- RT-qPCR:
-
Real-time quantitative polymerase chain reaction
- MIQE:
-
Minimum information for publication of RT-qPCR experiments
- CRC:
-
Colorectal cancer
- HKG:
-
Housekeeping gene
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- 18S:
-
18s ribosomal RNA
- PPIA:
-
Peptidylprolyl isomerase A
- ACTB:
-
Actin beta
- SDHA:
-
Succinate dehydrogenase complex flavoprotein subunit A
- GUSB:
-
Glucuronidase beta
- B2M:
-
Beta-2-microglobulin
- YWHAZ:
-
Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein zeta
References
Bustin, S. A., Benes, V., Nolan, T. & Pfaffl, M. W. Quantitative real-time RT-PCR–a perspective. J. Mol. Endocrinol. 34, 597–601. https://doi.org/10.1677/jme.1.01755 (2005).
Mocellin, S., Rossi, C. R., Pilati, P., Nitti, D. & Marincola, F. M. Quantitative real-time PCR: A powerful ally in cancer research. Trends Mol. Med. 9, 189–195. https://doi.org/10.1016/s1471-4914(03)00047-9 (2003).
Janik, M. E. et al. RT-qPCR analysis of human melanoma progression-related genes—A novel workflow for selection and validation of candidate reference genes. Int. J. Biochem. Cell Biol. 101, 12–18. https://doi.org/10.1016/j.biocel.2018.05.007 (2018).
Li, Y. L., Ye, F., Hu, Y., Lu, W. G. & Xie, X. Identification of suitable reference genes for gene expression studies of human serous ovarian cancer by real-time polymerase chain reaction. Anal. Biochem. 394, 110–116. https://doi.org/10.1016/j.ab.2009.07.022 (2009).
Nolan, T., Hands, R. E. & Bustin, S. A. Quantification of mRNA using real-time RT-PCR. Nat. Protoc. 1, 1559–1582. https://doi.org/10.1038/nprot.2006.236 (2006).
Dheda, K. et al. The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization. Anal. Biochem. 344, 141–143. https://doi.org/10.1016/j.ab.2005.05.022 (2005).
Schrader, A. J. et al. Application of real-time reverse transcriptase-polymerase chain reaction in urological oncology. J. Urol. 169, 1858–1864. https://doi.org/10.1097/01.ju.0000047363.03411.6b (2003).
de Jonge, H. J. et al. Evidence based selection of housekeeping genes. PLoS One 2, e898. https://doi.org/10.1371/journal.pone.0000898 (2007).
Kozera, B. & Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 54, 391–406. https://doi.org/10.1007/s13353-013-0173-x (2013).
Caradec, J. et al. “Desperate house genes”: The dramatic example of hypoxia. Br. J. Cancer 102, 1037–1043. https://doi.org/10.1038/sj.bjc.6605573 (2010).
Krzystek-Korpacka, M., Diakowska, D., Bania, J. & Gamian, A. Expression stability of common housekeeping genes is differently affected by bowel inflammation and cancer: Implications for finding suitable normalizers for inflammatory bowel disease studies. Inflamm. Bowel Dis. 20, 1147–1156. https://doi.org/10.1097/MIB.0000000000000067 (2014).
Blanquicett, C., Johnson, M. R., Heslin, M. & Diasio, R. B. Housekeeping gene variability in normal and carcinomatous colorectal and liver tissues: Applications in pharmacogenomic gene expression studies. Anal. Biochem. 303, 209–214. https://doi.org/10.1006/abio.2001.5570 (2002).
Medema, J. P. Cancer stem cells: The challenges ahead. Nat. Cell Biol. 15, 338–344. https://doi.org/10.1038/ncb2717 (2013).
Nassar, D. & Blanpain, C. Cancer stem cells: Basic concepts and therapeutic implications. Annu. Rev. Pathol. 11, 47–76. https://doi.org/10.1146/annurev-pathol-012615-044438 (2016).
Sørby, L. A., Andersen, S. N., Bukholm, I. R. & Jacobsen, M. B. Evaluation of suitable reference genes for normalization of real-time reverse transcription PCR analysis in colon cancer. J. Exp. Clin. Cancer Res. CR 29, 144. https://doi.org/10.1186/1756-9966-29-144 (2010).
Dydensborg, A. B., Herring, E., Auclair, J., Tremblay, E. & Beaulieu, J. F. Normalizing genes for quantitative RT-PCR in differentiating human intestinal epithelial cells and adenocarcinomas of the colon. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G1067-1074. https://doi.org/10.1152/ajpgi.00234.2005 (2006).
Zheng, G. et al. Identification and validation of reference genes for qPCR detection of serum microRNAs in colorectal adenocarcinoma patients. PLoS One 8, e83025. https://doi.org/10.1371/journal.pone.0083025 (2013).
Huggett, J. F. et al. The digital MIQE guidelines: Minimum information for publication of quantitative digital PCR experiments. Clin. Chem. 59, 892–902. https://doi.org/10.1373/clinchem.2013.206375 (2013).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Liu, P. et al. Oncogenic mutations in armadillo repeats 5 and 6 of β-catenin reduce binding to APC, increasing signaling and transcription of target genes. Gastroenterology 158, 1029-1043.e1010. https://doi.org/10.1053/j.gastro.2019.11.302 (2020).
Aranda, P. S., LaJoie, D. M. & Jorcyk, C. L. Bleach gel: A simple agarose gel for analyzing RNA quality. Electrophoresis 33, 366–369. https://doi.org/10.1002/elps.201100335 (2012).
Faraldi, M. et al. A novel methodological approach to simultaneously extract high-quality total RNA and proteins from cortical and trabecular bone. Open Biol. 12, 210387. https://doi.org/10.1098/rsob.210387 (2022).
Liu, L. et al. Selection and validation of the optimal panel of reference genes for RT-qPCR analysis in the developing rat cartilage. Front. Genet. https://doi.org/10.3389/fgene.2020.590124 (2020).
Gao, Z., Deng, W. & Zhu, F. Reference gene selection for quantitative gene expression analysis in black soldier fly (Hermetia illucens). PLoS One 14, e0221420. https://doi.org/10.1371/journal.pone.0221420 (2019).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, Research0034. https://doi.org/10.1186/gb-2002-3-7-research0034 (2002).
Chen, L. et al. Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta 234, 377–390. https://doi.org/10.1007/s00425-011-1410-3 (2011).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250. https://doi.org/10.1158/0008-5472.Can-04-0496 (2004).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515. https://doi.org/10.1023/b:bile.0000019559.84305.47 (2004).
Al-Sabah, A., Stadnik, P., Gilbert, S. J., Duance, V. C. & Blain, E. J. Importance of reference gene selection for articular cartilage mechanobiology studies. Osteoarthr. Cartel. 24, 719–730. https://doi.org/10.1016/j.joca.2015.11.007 (2016).
Lü, J. et al. Selection and validation of reference genes for RT-qPCR analysis of the Ladybird Beetle Henosepilachna vigintioctomaculata. Front. Physiol. 9, 1614. https://doi.org/10.3389/fphys.2018.01614 (2018).
Sarker, N. et al. Identification of stable reference genes for quantitative PCR in koalas. Sci. Rep. 8, 3364. https://doi.org/10.1038/s41598-018-21723-0 (2018).
Taylor, S., Wakem, M., Dijkman, G., Alsarraj, M. & Nguyen, M. A practical approach to RT-qPCR-Publishing data that conform to the MIQE guidelines. Methods (San Diego, Calif.) 50, 11–15. https://doi.org/10.1016/j.ymeth.2010.01.005 (2010).
Wisnieski, F. et al. Reference genes for quantitative RT-PCR data in gastric tissues and cell lines. World J. Gastroenterol. 19, 7121–7128. https://doi.org/10.3748/wjg.v19.i41.7121 (2013).
Zhai, Z., Yao, Y. & Wang, Y. Importance of suitable reference gene selection for quantitative RT-PCR during ATDC5 cells chondrocyte differentiation. PLoS One 8, e64786. https://doi.org/10.1371/journal.pone.0064786 (2013).
Kheirelseid, E. A., Chang, K. H., Newell, J., Kerin, M. J. & Miller, N. Identification of endogenous control genes for normalisation of real-time quantitative PCR data in colorectal cancer. BMC Mol. Biol. 11, 12. https://doi.org/10.1186/1471-2199-11-12 (2010).
d, M. G. & Huggett, J. F. The digital MIQE guidelines update: Minimum information for publication of quantitative digital PCR experiments for 2020. Clin. Chem. 66, 1012–1029. https://doi.org/10.1093/clinchem/hvaa125 (2020).
Author information
Authors and Affiliations
Contributions
Hong.Y. and L.L. conceived the study. Hu.Y. and Hong.Y. carried out the experiments. Hu.Y., J.Q., and L.L. analyzed the data and prepared figures. Hu.Y. and Z.X. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, Y., Jiang, Q., Zhai, X. et al. Screening and validation of the optimal panel of reference genes in colonic epithelium and relative cancer cell lines. Sci Rep 13, 17777 (2023). https://doi.org/10.1038/s41598-023-45174-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-45174-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.