Abstract
In wheat (Triticum aestivum L.), the grain size varies according to position within the spike. Exposure to drought and high temperature stress during grain development in wheat reduces grain size, and this reduction also varies across the length of the spike. We developed the phenomics approach involving image-based tools to assess the intra-spike variation in grain size. The grains were arranged corresponding to the spikelet position and the camera of smart phone was used to acquire 333 images. The open-source software ImageJ was used to analyze features of each grain and the image-derived parameters were used to calculate intra-spike variation as standard deviation (ISVAD). The effect of genotype and environment were highly significant on the ISVAD of grain area. Sunstar and Raj 4079 contrasted in the ISVAD of grain area under late sown environment, and RNA sequencing of the spike was done at 25 days after anthesis. The genes for carbohydrate transport and stress response were upregulated in Sunstar as compared to Raj 4079, suggesting that these play a role in intra-spike assimilate distribution. The phenomics method developed may be useful for grain phenotyping and identifying germplasm with low intra-spike variation in grain size for their further validation as parental material in breeding.
Similar content being viewed by others
Introduction
The number of developed grains and the grain weight are the critical components of grain yield which have been extensively used for the improvement of productivity in wheat (Triticum aestivum L.). Variation exists in these two traits within the spike. The proximal florets and middle spikelets have larger grains than the distal florets and top spikelets1,2,3. The distal primordia are prominently delayed with respect to the rate of initiation exhibited by proximal primordia, however, the maturation of all grains is synchronous which may cause less assimilate supply to distal grains4,5. The uneven intra-spike assimilate distribution is demonstrated from the reports that approximately 44% of the total number of spikelets located in the lower half of the spike account for 64% of the grain yield6. The source or photosynthetic potential is underutilized without the efficient partitioning of assimilates to sink (grains)7,8,9, therefore, increasing the sink strength is critical for further improving wheat productivity10. Breeding for the sink strength (grain number or potential weight) has continuously boosted wheat yield previously11.
A uniform intra-spike grain filling is preferred to obtain higher yield and better wheat quality. Grain yield is greatly decreased because smaller grains are readily lost during the winnowing process before milling12,13. Grain size variation also increases the variability of seedling emergence, early groundcover, and initial biomass, indirectly reducing crop management efficiency and production14,15. Heterogeneity of grain mass deteriorates flour quality and end-use suitability16.
Individual grain size variance at intra-genotype level is influenced by specific chromosomal regions and the assimilate translocation to grains17. At a temperature above 35 °C, the grain size is reduced due to less starch content in grains18, and the reduction in single grain weight can be as high as 20.61% if the high temperature prevails from the start of grain filling until ripening19. Reduced photosynthesis and increased leaf senescence under post-anthesis high temperatures restrict the availability of starch for deposition and usually cause top and bottom spikelets and distal florets to either abort or produce small grains20,21,22. An earlier study reported that post anthesis temperature regime of 30/25 °C decreases grain size in all floret positions within the spike23. Later it was found that grains located on different parts of the spike and spikelets have distinct responses to post-anthesis high temperature24. Several studies on diverse germplasm observed genetic variability in grain size stability in response to heat stress25,26. In addition, soil moisture deficit during grain filling increases the small-sized grains; however, breeding can reduce this risk27.
The limitations of traditional genetic improvement methods primarily focused on enhancing grain yield have prompted the adoption of genomics-driven tools, including genomic selection. These tools hold the potential to revolutionize plant breeding processes by increasing precision and efficiency28. Integrating phenomics protocols is crucial for maximizing the benefits of genomic science in advancing crop improvement. Furthermore, the often overlooked intraspike assimilate distribution in wheat's grain yield dynamics emphasizes the significance of sophisticated imaging techniques such as X-ray CT (Computed Tomography) and NMR (Nuclear Magnetic Resonance)29,30,31,32. However, due to their high cost, these tools are not accessible to many crop scientists. In response to this limitation, efforts have been made to develop an affordable phenotyping approach for intraspike assimilate distribution. The phenotyping approach aims to provide a cost-effective alternative for studying this intricate plant trait, ensuring that a wider range of researchers can contribute to advancing crop science. Images of grains within the spike were captured using smartphone camera for sixteen diverse wheat genotypes grown under non-stressed, terminal drought and terminal high temperature environments and analyzed using an open source software ImageJ. Our approach accurately extracted many agronomically relevant grain size and shape parameters of each grain within the spike that was used to calculate the intra-spike variation as standard deviation (ISVAD). This method was used to quantify the genetic variation and effect of genotype-environment interaction on the intra-spike variation in grain size. The RNA sequencing of the spike of contrasting genotypes was conducted to identify the genes responsible for low intra-spike variation in grain size.
Results
Validation of ImageJ results
The known length in millimeters of the line drawn against the scale in the first image is entered in the set scale tool which converts the pixels into mm and this calibration applies to all the images processed in the session. The distance between the grains and the camera was fixed at 31 cm for all the images. The manually counted actual number of grains spike−1 was same as the number of grains values given by ImageJ. The ImageJ result parameters, Feret and MinFeret of each grain could explain the actual grain length and width in the selected genotypes. The Feret and MinFeret values obtained from ImageJ correlated to the actual length and width of 100 grains as measured by the vernier caliper (R2 = 0.892 and R2 = 0.77) (Figs. 1a, 2a). The correlation of actual grain length with Feret values and actual grain width with MinFeret values was the same in a repeat of the experiment. Therefore, we confirmed the consistency over grain size estimation by ImageJ. The predicted values were tenfold cross-validated using the cv.lm function of “DAAG” package of R33 (Figs. 1b, 2b). The function cv.lm carries out a k-fold cross-validation for a linear model (i.e. an 'lm' model). For each fold, an 'lm' model is fit to all observations that are not in the fold (the 'training set') and prediction errors are calculated for the observations in the fold (the 'test set'). The prediction errors are the absolute error and its square. The average prediction errors over the observations in the fold are calculated, and the square root of the average of the squared errors is taken.
Intra-spike variation in grain size
The genotypes were sown for the non-stress or control (C) and terminal drought (D) during the optimal sowing period, commonly known as timely sown in India (usually referred to as free from heat stress)34. The sowing between the mid of December to last week is referred to as late sown (L) in the India when the crop is naturally exposed to terminal high-temperature stress34. Heading and anthesis occurred in the third week of February under C and D environments, and in the first week of March under L environment. Physiological maturity occurred at the end of March under all environments. The mean daily maximum temperature from heading to maturity under timely sown conditions (C, D) and late sown condition (L) was 30.04 °C and 32.21 °C in 2018–2019; 30.72 °C and 31.79 °C in 2019–2020 and, 34.32 °C and 36.18 °C in 2020–2021 (Table 3). This was conducive to create measurable variations in the grain size for the optimization of methods. The grain yield plot−1 was correlated to grain yield spike−1 (R2 = 0.456) (Fig. S1). The measurements of the spike yield components (grains spike−1, grain yield spike−1, spikelets spike−1) and spike length were done using three replicates of main stem spikes only; however the grain yield plot−1 includes the spikes of main stems and tillers.
The grain area ranged from 3.04 to 30.15 mm2 across all the genotypes and environments. The environments C, D and L had a significant effect on the ISVAD of grain width (p ≤ 0.001) and area (p ≤ 0) in 2018–2019 and 2019–2020 (Table S1). The ISVAD of grain area and grain width were highly influenced by genotype and genotype-environment interaction during 2018–2019 and 2019–2020 (Table S1).The mean ISVAD of grain width across the genotypes was higher under C (0.366a) as compared to D (0.354b) and L (0.34c) in 2018–2019 and 2019–2020 (Table 1). The mean ISVAD of grain area also decreased under D (2.9b) and L (2.786c) as compared to C (3.21a). The grains spike−1, grain yield spike−1, single grain weight (SGW), spikelets spike−1 and spike length decreased significantly under D and L in comparison to C (Table 1). The mean grain length/width ratio increased under D (2.26b) and L (2.215b) as compared to C (2.137a) and the mean SGW (g) decreased under D (0.041b) and L (0.039b) compared to C (0.045a) (Table 1). The K mean cluster plot for 56% variation of spike length, spikelets spike−1, grains spike−1 and yield spike−1 under C, D and L environments of 2018–2019 and 2019–2020 gives four clusters of genotypes (Fig. 3). Within the cluster 4, Sunstar has significantly lower mean ISVAD of grain area as compared to Raj 4079 and Raj 4037 across C, D and L environments (Tables S2–S4). Under C, the mean SGW (g) of 2018–2019 and 2019–2020 of Sunstar (0.038d) is lower as compared to Raj 4079 (0.051ab), however under D the SGW (g) of Sunstar (0.037ab) is higher than Raj 4079 (0.025c). Under L environment, the SGW (g) of Sunstar (0.025ef) and Raj 4079 (0.035de) are comparable and the ISVAD of grain area of Sunstar (1.778e) is notably lower than Raj 4079 (2.696bcd).
Biplot accounting for 68.7% variation reveals that ISVAD of grain area of HD 2987, HPBW 01, PBW 175, AKAW 4627, HUW 234 and MP 4010 is highly influenced by the environments (Fig. 4). Among the 16, HD 2987 had the highest ISVAD of grain area under C (4.011) and the grain yield spike−1 of HD 2987 decreased significantly under D and L (Tables S2–S4). The grains spike−1 of HD 2643 and HD 2967 decreased markedly under D and L as compared to the C. The genotypes Chinese spring, PBW 396 and WH 730 have most stable ISVAD of grain area; however the three genotypes are different in spike yield components (Figs. 3, 4). PBW 396 is distinct from Chinese spring and WH 730 in the distance heat map of the ISVAD of grain width, length, area and perimeter across the environments in 2018–2019 and 2019–2020 (Fig. S2), as PBW 396 has a high ISVAD of grain area across these 6 environments (Tables S2–S4). The compact spike shaped genotypes Chinese spring, AKAW 4627, Sunstar and HUW 234 are similar in the ISVAD of grain width, length, area and perimeter as represented by the distance heat map. The K means-cluster plot of rachis area and rachis perimeter under C, D and L environments, accounting for 67.3% variation formed three different clusters of genotypes, where Raj 4079, PBW 175 and Raj 4037 are in cluster 2, whereas Sunstar, Chinese Spring, Halna, HD 2987, HD 2643 and MP 4010 are in cluster 3 (Fig. 5).
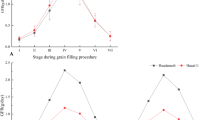
The top region of spike has relatively higher ISVAD of grain width and ISVAD of grain area than middle and bottom regions under C (Fig. S3a,b) and also across the C, D and L environments (Fig. S4a,b) of 2018–2019 and 2019–2020. The effect of environment was highly significant on the ISVAD of grain area within top (p < 0.001) and bottom (p < 0.001) regions of spike whereas it was less significant on the middle (p < 0.05) region of spike (Table S5). However, the genotype-environment interaction influenced only top and bottom ISVAD of grain area. The ISVAD of grain width was influenced by environment only in the middle (p < 0.05) and bottom (p < 0.01) regions of the spike. The genotype-environment interaction notably affects only the bottom region ISVAD of grain width (Table S6). The ISVAD of grain length was weakly associated with ISVAD of grain width; however, D and L impaired this relationship (Fig. 6). The ISVAD of grain width and length was not influenced by the spike length, spikelets spike−1, grains spike−1 and grain yield spike−1. The Fig. 6 also depicts that the environment has no effect on the association between ISVAD of grain length and spike yield components.
The experiments were repeated in 2020–2021 for five genotypes Sunstar, Halna, Raj 4079, HD 2987 and PBW 396. The crop season 2020–2021 had highest mean maximum daily temperature of 41.6 °C (Table 3). The ISVAD of grain area in 2020–2021 increased significantly under L (Fig. 7) although grains spike−1 decreased as compared to C (Fig. S5). The grain yield spike−1 decreased under D and L (Fig. S6). The ISVAD of grain area did not change significantly in D but the grains spike−1 was reduced. The mean ISVAD of grain area was significantly lower in Sunstar (2.401b), as compared to Raj 4079 (3.185a) (Fig. 7). The effect of environment was significant only on ISVAD of grain area of selected five genotypes in three crop seasons (Table S7).
Variation in intra-spike variation as standard deviation (ISVAD) of grain area. (a) The effect of late sown (L), terminal drought (D) and Control (C) environments on ISVAD of grain area in 2020–2021. Each bar in the environment effect represents 15 observations (3 replications and 5 genotypes). (b) The genotype effect across the environments on ISVAD of grain area. Each bar in the genotype effect represents 9 observations (mean values of three replications and three environments). (c) Genotype environment interaction effect on ISVAD of grain area. Each bar in the genotype environment interaction effect represents mean values of three replications. Letters represent the significance of differences among mean values as computed by the Duncan multiple range test at p ≤ 0.05. Genotypes with common letters are not significantly different.
In summary, the effects of genotype and environment were significant on the ISVAD of grain area across the three crop seasons (Tables S1, S7). The grains spike−1, grain yield spike−1, SGW, spikelets spike−1 and spike length decreased significantly under D and L in comparison to C. In the first two years, ISVAD of grain area also decreased under D and L (Table 1). The K-means cluster plot of spike length and spike yield components formed 4 clusters. Within cluster 4, Sunstar and Raj 4079 have significantly different ISVAD of grain area. The ISVAD of grain area was higher in the top region of the spike as compared to middle and bottom regions of spike across the environments (Fig. S4b). The effect of environment and genotype-environment interaction was significant on top and bottom regions of spike (Table S5). In 2020–2021, ISVAD of grain area increased, and grains spike−1 and grain yield spike−1 decreased (Figs. 7, S5, S6).
Sequence assembly based on cDNA libraries sequencing data
The RNA sequencing of three biological replicates of Sunstar under L (SL) and Raj 4079 under L (RL) generated 160 million and 170 million reads (Table S8). The sequencing raw data have been submitted to NCBI and the accession number is PRJNA918542. GC content and Q30 is 52.20 and 87.50% for SL, and 53.28% and 93.09% for RL respectively. Q30 refer to a 0.1% chance of error and 99.9% confidence. These results indicate that all the data were qualified for downstream analysis.
Differential expression of genes (DEGs)
A total of 2861 DEGs were identified out of which 1582 DEGs were downregulated in RL and 1279 were upregulated in RL as compared to SL at p < 0.05. The top 5% DEGs are shown by heat map (Fig. 8).
Heat map of top 5% differentially expressed genes log2 fold 2 at p < 0.05 between Sunsar and Raj 4079 under late sown (SL and RL). Heat map was generated with the R (Version 3.4.4) function heatmap.2() of the package gplots https://CRAN.R-project.org/package=gplots35.
Functional analysis of DEGs
Gene Ontology (GO) analysis was used to analyze functions of DEGs. These DEGs are classified into 54 functional groups in three main categories (cellular component, molecular function and biological process) (Figs. S7–S9). GO annotation results showed that the terms of cell anatomical structure and organelle were dominant in the cellular component, those of binding and catalytic activity were primary in molecular function, and those of metabolic process, and cellular process were foremost in biological process. These GO terms demonstrated that genes related to wheat late grain development encoded diverse regulators and protein. Among the 92 DEGs involved in carbohydrate transport and metabolism, 34 DEGs were involved in sucrose transport and sucrose metabolism. 97 transmembrane transporters genes were differentially expressed between Sunstar and Raj 4079 under L environment. The Ubiquitin-proteasome system plays an important role in grain development during later stage as reflected by 43 DEGs related to ubiquitin-mediated proteolysis and proteasome pathway. The expression of 18 DEGs of signal transduction was significantly lower in Raj 4079. A total of 25 genes were downregulated in Raj 4079 in response to heat stress, water stress and oxidative stress. The genes involved in transmembrane transporter activity were downregulated under high temperature stress in Raj 4079 such as 3 SWEET bidirectional sugar transporter genes. Other transporter genes downregulated were peptide transporter PTR3-A, 10 ABC transporter C family member, 2 low affinity nitrate transporter NPF7.1a, 3 cadmium/zinc-transporting ATPase and 7 oligopeptide transporter. The sucrose phosphate synthase involved in sucrose synthesis and metabolism during grain development was also downregulated in Raj 4079 under high temperature stress. The stress responsive gene transcripts, temperature stress induced lipocalin, and stress induced transcription factor SNAC1 expression were downregulated in Raj 4079 as compared to Sunstar.
Discussion
Phenomics for wheat grain size
The limited progress achieved through traditional genetic improvement methods, primarily centered around enhancing grain yield, has prompted the utilization of tools stemming from advancements in genomics. These tools focus on dissecting individual traits that are associated with specific genes. Genomic selection, among these approaches, presents an alternative means of predicting breeding values in plants by leveraging genome-wide markers. This strategy holds the promise of expediting genetic advancement through heightened selection precision, accuracy, and a reduction in the time required for breeding cycles28. Nonetheless, the successful implementation of genomic selection necessitates the availability of both phenotypic and genomic data for a designated set of genotypes within a given crop. Thus, phenomics protocols emerge as pivotal components in harnessing the benefits of genomic science to drive further progress in crop improvement. While considerable attention has been directed towards understanding wheat grain yield constituents such as grain number and weight, certain critical factors, notably the distribution of assimilates within the spike (intraspike), have not received thorough investigation. This facet has the potential to significantly influence grain yield outcomes particularly under abiotic stress environments. Measuring intraspike assimilate distribution requires a meticulous characterization of each individual grain within the spike. However, this task becomes notably more intricate when dealing with a substantial number of genotypes that need screening. To address this complexity, sophisticated and costly tools have been demonstrated. These tools, including X-ray CT or NMR, offer advanced imaging capabilities, enabling the non-destructive examination of plant structures and traits such as features of grains29,31,32. However, these tools are not affordable for many of the crop improvement scientists. Hence an attempt was made to employ affordable phenotyping approach for assessing genetic variability in traits such as ISVAD. Our method uses simple equipment to collect images and processing of images is quick with an open source software ImageJ. It is fully-automated as the thresholding and make binary tool ensure the accurate detection of grains against the background. ImageJ provides a large number of complex traits, while sequentially numbering each grain and thereby retaining positional information of each grain in spike and spikelet. In addition to the grain size parameters such as feret’s diameter, area and perimeter, it also measures the shape descriptors such as circularity, aspect ratio and roundness (https://imagej.nih.gov/ij/docs/guide/146-30.html). Haghshenas et al. (2022) used ImageJ to explore the image-based mean grain weight estimations in wheat by developing simple predictive linear models36. ImageJ has been previously used to measure the grain size and shape for QTL analysis in wheat37 and Arabidopsis38. It has also been used for measurement of grain traits of rice39 and different food grains40. The wheat grain characteristics have also been explored using other image-based phenotyping platforms such as WinSEEDLE (Regent Instruments, Quebec City, QC, Canada)41,42,43,44,45,46, SeedCount (Next Instruments, Condell Park, Australia)47,48,49,50,51,52,53 and Marvin seed analyzer54,55,56 however, these are associated with expensive specialized hardware. SmartGrain and GrainScan are similar free softwares that require low-cost equipment and are more commonly used for grain phenotyping57,58, to explore genetic underpinnings of grain size59,60,61,62,63,64,65, to help breeding and introgression66,67,68, to explore drought response69 and evaluate genebank accessions70. In this study, the analysis of variance (ANOVA) of 16 genotypes in the first two crop seasons and combined ANOVA of three crop seasons for five genotypes confirmed the significant effect of genotype and environment on ISVAD of grain area. Therefore, the grain area was the most reliable parameter for the analysis of intra-spike variation in grain size. We show that ImageJ assessment of quantitative grain characteristics can be an accurate and affordable method for the estimation of ISVAD of grain size for large sample sizes. In summary, the limitations encountered in traditional genetic improvement pathways, which predominantly target grain yield enhancement, have spurred the adoption of genomics-driven tools. These tools, exemplified by genomic selection, have the potential to revolutionize breeding processes through enhanced precision and efficiency. The integration of phenomics protocols is pivotal in maximizing the benefits of genomic science for the advancement of crop improvement. Moreover, the overlooked aspect of intraspike assimilate distribution in wheat's grain yield dynamics underscores the importance of sophisticated imaging techniques like X-ray CT and NMR, which facilitate the investigation of intricate plant traits. Since these are expensive and not affordable for many of the crop scientists, an attempt was made to develop an affordable phenotyping approach for ISVAD.
ISVAD as trait for assessment of sensitivity to high temperature and soil moisture stress
Genetic variation exists in ISVAD of grain width, length, area and perimeter within the spike. The ISVAD of grain area was highest in the top region of spike (Fig S3b, S4b), indicating that the assimilate distribution was uneven in top region. This is in agreement with an earlier report that increased temperature reduces grain volume at the top of the spike when water is non-limiting; however, a significant reduction in grain number occurs in the middle of the spike, with the top and bottom regions being less affected71. Under combined heat and drought stress, and drought stress alone, the grain weight reduces significantly at the top and bottom of the wheat spike72. The effect of genotype-environment interaction was significant on ISVAD of grain area of only the spike's top and bottom regions (Table S5), indicating that genotypes have distinct responses to grain size reduction in these regions under high temperature and drought.
The mean ISVAD of grain area across the genotypes decreased under D and L in the first two crop seasons (Table 1), indicating that grains were more uniformly filled under D and L than the control. An increase in grain length, width and depth is associated with a decrease in grains spike−171. Grain size increases to compensate for the reduction in yield due to reduced grains spike−1 for some varieties under terminal drought73,74. The mean grains spike−1, grain yield spike−1 and SGW decreased under D and L, and the grain length/width ratio increased compared to C, suggesting that grain width reduction contributed to grain weight reduction (Table 1). This is in agreement with a recent study that reported a 20% average reduction in grain area mainly due to a decrease in the grain width after heat shock treatment26. The grain length is set during early phase of grain filling (from anthesis to 10–15 days after anthesis (DAA) by mesocarp cell elongation75,76. During 2020–2021, plants experienced severe heat stress, therefore mean ISVAD of grain area increased under L (Fig. 7) despite a decrease in grains spike−1 due to poor floret and pollen fertility under high temperature77 and less grain setting shortly before or at anthesis78 (Figs. 7, S5), along with decreased grain yield spike−1 (Fig. S6). For some genotypes, high temperature at grain filling accelerates the grain filling rate upto 30 °C, presumably reflecting increased enzyme activity and metabolic processes79; however, it also causes truncation of the grain filling period, which may not be fully compensated by increased rate, depending on the genotype; therefore the overall effect is a reduction of grain size18. The spike length and spike yield components and their responses to terminal drought and high temperature were taken into account (Fig. 3) to identify similar genotypes for assessing the contrast in only ISVAD of grain size parameters.
The posthoc test shows that among the genotypes Sunstar and Raj 4079, which are similar in spike yield components (Fig. 3), the mean ISVAD of grain area of Sunstar was significantly lower than Raj 4079 under L environment of three crop seasons (Tables S2–S4; Fig. 7). The C environments of 2018–2019 and 2019–2020 were cooler and more favorable than the other 7 environments (D and L of 2018–2019 and 2019–2020; and C, D and L of 2020–2021). As Sunstar normally has smaller grains than Raj 4079 (Fig. S10), the mean SGW of Raj 4079 was significantly higher than Sunstar under the C of 2018–2019 and 2019–2020. The SGW of Raj 4079 and Sunstar became comparable under the less favorable 7 environments which suggest that lower ISVAD of grain area of Sunstar under stress could be associated with the more stable SGW (HSI = 0.35) in contrast to SGW of Raj 4079 (HSI = 1.75) which notably decreased under L (Tables S2–S4). However, further screening of large set of germplasm is required to confirm the hypothesis that genetic variability in ISVAD of grain area could be one of the opportunities to improve the grain weight spike−1.
Barrero et al. reported that grain size stability is less impacted by heat stress in the genotypes with smaller grains26. In the present study, the genotypes which have the smaller grain size Chinese spring (15.67 mm2), WH 730 (17.89 mm2) and PBW 396 (19.49 mm2) have a stable ISVAD of grain area and genotypes with larger grains such as PBW 175 (23.04 mm2), and HD 2987 (21.62 mm2) are highly unstable (Figs. S10, 4). The correlation (R2) between grain width and grain length across all the environments in three years was 0.45 (Fig. S11). Similarly, an earlier study reported that the grain width, length, and area quantified in 210 European wheat accessions showed strong positive correlations80. However, a low correlation (0.29) was found under non-stressed conditions by Philipp et al.6. Theoretical models suggest that improving grain size and shape could boost milling yield, with large and spherical grains being the optimum grain morphology81. Grain circularity across all the genotypes and environments in three crop seasons ranged from 0.15 to 0.92, and roundness ranged from 0.22 to 0.89. The mean circularity and roundness of grains under the C environment of three years were 0.72 ± 0.0009 and 0.48 ± 0.0006, respectively. Under D, grains' mean circularity (0.69 ± 0.0011) and roundness (0.44 ± 0.0008) decreased by 4.2% and 8.3%, respectively. Also, under L, grains' mean circularity (0.71 ± 0.0009) and roundness (0.45 ± 0.0007) decreased by 1.4% and 6.3%, respectively. The variation in grain size and shape has been supported by the genetic and phenotypic structure, where grain size has progressively increased through changes both in grain width and length, and later modifications in grain shape have primarily been made through changes to grain length54.
The photosynthate availability (source capacity) is adequate for grain development, and assimilating (sink) capacity determines the grain yield9,10,82. Under heat and drought stress, grain filling is supported by stem reserve mobilization contributing up to 70% of grain weight to the grains, compensating for reduced leaf photosynthesis depending on the genotype19,83. However, in a recent study, half spike removal increased, but defoliation decreased single grain weight, indicating that grain yield is co-limited by source and sink capacity82. Nonetheless, there is evidence for spikelet-to-spikelet competition for assimilates within a spike leading to the unbalanced distribution of grains per spikelet along the length of the spike80,85. Few studies have stated that the limiting factor for reaching maximum grain weight in distal spikelets could be due to the impediments in loading of photosynthates into grains and restrictions in the transport of assimilate to grains within spike and spikelets86,87,88. Barrero et al. reported that heat stress reduces grain size by affecting assimilate translocation and grain growth rate26. An earlier study also stated that the source is non-limiting under mild heat stress (30/25 °C), and the grain set and single grain weight reduction is probably due to the inhibition of H+-ATPase activity and active transport of hexose into the endosperm89. Rachis plays an important role in the translocation of assimilates within the spike. Rachis parameters were determined to find out the effect of rachis size on the intra-spike assimilate distribution to the grains. Therefore, the genetic variation in the rachis parameters could be one of the factors affecting the ISVAD of grain parameters. The K means cluster plot of rachis area and perimeter of the spikes in the present study grouped the low ISVAD of grain size genotypes (Sunstar, Chinese Spring, Halna, HD 2987 and HD 2643) in the same cluster (Fig. 5). Although the assimilate supply within the spike is governed by competition among the different parts of the spike90, the genetic advancement in yield gain has no direct association with the vasculature size of the spike91. In summary, the present study could differentiate sensitive and tolerant genotypes for their intra-spike variation in grain traits under high temperature and low soil moisture conditions relative to plants grown under optimum temperature and sufficient soil moisture. The reduction in grain width contributed to the reduction in grain area, SGW, grain circularity and roundness under stress. The genetic variation in the rachis size parameters could be one of the factors affecting the ISVAD of grain size parameters.
Genes associated with responses of distinct genotypes
RNA sequencing was carried out to identify the genes involved in uniform intra-spike assimilate distribution by comparing gene expression profiles of genotypes contrasting in ISVAD of grain area. Grain growth and development is divided into three phases92,93 lag-phase (from anthesis to about 10–15 DAA); filling-phase (from 15 to 35–40 DAA); and maturation phase. During the lag-phase, endosperm cells undergo cell division and cell differentiation75,76. The filling-phase consists of the accumulation of assimilates into the endosperm (mainly starch and proteins) and dry mass content increases. During filling-phase, cell layers of different maternal tissues exhibit degeneration processes at discrete time points94. The outer tissues such as vascular bundles and nucellar epidermis remain alive for the duration of starch accumulation as they play a role in nutrient transport and grain filling. Retaining live cell layers of these tissues at late stages of grain filling affect the final grain weight. Therefore, mid of filling-phase was selected for RNA sequencing of spike to identify the differentially expressed genes in genotypes contrasting in ISVAD of grain area.
As the ISVAD of grain area of Sunstar was significantly lower than Raj 4079 under L environment, the up and downregulated genes and isoforms at p < 0.05 are discussed for L only. The transcripts of three Bidirectional sugar transporter SWEET were downregulated in Raj 4079 as compared to Sunstar. The transcripts of three other proteins that are integral component of membrane involved in carbohydrate transport were downregulated in Raj 4079 as compared to Sunstar. These genes could be involved in a more efficient intra-spike assimilate distribution in Sunstar. Grain size controlling factor GS5 positively regulates grain size and weight by regulating grain width, and filling in rice95,96. Under L, GS5 transcripts decreased in Raj 4079 (log2 fold change − 2 and − 3.5) compared to Sunstar. Grain incomplete filling 1 which encodes a cell wall invertase that regulates the hydrolysis of sucrose and has an essential role in sucrose unloading, was also downregulated in Raj 4079 (log2 fold change − 2.6 and − 2.8). At 37 °C, the mRNA levels of the enzymes in the starch synthesis pathway are considerably reduced97. Sucrose phosphate synthase was highly expressed in Sunstar (Fig. 8). The transcript levels of sucrose synthase were also lower in Raj 4079 as compared to Sunstar (log2 fold − 0.7 and − 1.9). The mean SGW of Sunstar is more tolerant to high temperature (HSI = 0.35) than mean SGW of Raj 4079 (HSI = 1.75) across the three years. These could be some of the reasons for significant decrease of SGW and higher ISVAD of grain area in Raj 4079 as compared to Sunstar. Hutsch et al. postulated that the adverse effect of heat stress on the grain set and SGW reduction is probably due to the inhibition of H+-ATPase activity and the active transport of hexose into the endosperm89. The H+-ATPase establishes a pH gradient across the plasma membrane of the sink cell that energizes the carrier-mediated active transport of hexoses from the apoplast into developing grain by H+-cotransport98,99. However, in the present study, plasma membrane ATPase involved in proton-exporting ATPase activity across the plasma membrane is upregulated in Raj 4079 as compared to Sunstar. A total of 25 genes responsive to heat, water and oxidative stress were downregulated in Raj 4079. These results indicated that the above genes play essential roles in intra-spike assimilate distribution. In summary, the involvement of the sucrose and starch metabolism genes was a possible reason for distinct responses of genotypes differing in ISVAD. For candidates with a significant role in crop response to ISVAD of grain size under high temperature/drought, genome editing techniques can be applied to overexpress the gene and/or introduce beneficial alleles into an elite, high yielding cultivar. Understanding the genetic basis of diversity in wheat grain size and shape is critical to increasing yield potential and processing performance. Improvement of wheat grain yield should consider insight into the mechanisms of grain development and the supply of assimilates for developing grains.
Proposed phenomics protocol for ISVAD
We demonstrated an affordable plant phenomics approach for ISVAD using images acquired by smartphone and open-source software for image analysis. The results proved that it is possible to quantify intra-spike genetic variation in wheat grain size by employing an image-based phenomics approach.This method can be useful in routine selection as well as phenotyping for identification of genes and QTLs associated with intraspike assimilation and hence productivity of wheat under stress conditions (Fig. 9).
Materials and methods
Plant material and growth conditions
A set of 16 diverse wheat genotypes were seeded at the Farm Science Center, Banasthali Vidyapith, India situated at an elevation of 1010 ft. at 26°24′N, 75° 47′E (Table 2). We obtained the seeds of AKAW 4627, Chinese spring, Halna, HD 2643, HD 2967, HD 2987, HUW 234, MP 4010, PBW 175, PBW 396 and WH 730 from ICAR-Indian Institute of Wheat and Barley Research. The Department of Plant Breeding and Genetics, Punjab Agricultural University provided Sunstar. HPBW 01, Raj 3765, Raj 4037 and Raj 4079 were provided by Farm Science Centre, Banasthali Vidyapith, Tonk. The genotypes vary in spike morphology and are released for cultivation either for irrigated and timely sown environments (Raj 4037), restricted soil moisture environments (HD 2967, HD 2987, PBW 175 and PBW 396) and for delayed sowing (Halna, HD 2643, HPBW 01, HUW 234, MP 4010, Raj 3765, WH 730, AKAW 4627, Raj 4079 and Sunstar). The different spike shapes have different orientation of grains along the rachis of the spike which affects the assimilate distribution. Spike compactness has been reported to be closely related to grains spike−1 and thousand grain weight100,101.
The experiment was laid out in a split-plot arrangement and randomized complete block design in 2018–2019, 2019–2020 and 2020–2021 growing seasons. The genotypes were sown within the optimum sowing period, also known as timely sown in India (generally referred to as free from heat stress) from mid of November to the last week of November, for the non-stress or control (C) and terminal drought environment (D)34. The sowing was done on 23 November for C and D. The late flowering genotypes under C and D were sown 8–9 days before the early flowering genotypes to synchronize anthesis during drought. Under the timely sown C and D environments, the average temperature during the vegetative growth phase was same for the late flowering and early flowering genotypes. Synchronization of anthesis helped with ease of data collection and exposure of high temperature and drought treatment was same post-anthesis for all the genotypes. The sowing between the mid of December to last week is known as late sown (L) in the Indian context when the crop is naturally exposed to terminal high-temperature stress34. The sowing under the L environment was carried out on 15 December. Based on local practices102, C and L were given six irrigations through flooding provided at 20–25, 45–50, 65–66, 80–81, 90–95 and 110–115 days after sowing (DAS). Irrigation in D was given at 20–25, 45–50 and 65–66 DAS. In the K-Means cluster plot of spike length and spike yield components, Sunstar and Raj 4079 are in same cluster but significantly differ in ISVAD of grain area under C and L environments (Tables S3, S4; Fig. 3). Halna has the lowest ISVAD of grain area under D environment which is significantly lower than same cluster genotype Raj 4079 (Table S2). PBW 396 has a consistently significantly higher ISVAD of grain area than Sunstar across the 6 environments (Tables S4–S6). HD 2987 was selected for further experiments due to the highest ISVAD under C, and also a highly unstable ISVAD across the 6 environments (Fig. 4). The remaining 11 genotypes were not significantly varying in ISVAD of grain area across the 6 environments. The ISVAD of grain area values of remaining 11 genotypes are in between the extreme values of selected 5 genotypes. Therefore, five genotypes (Sunstar, Halna, Raj 4079, HD 2987 and PBW396) were selected for third year experiments in 2020–2021. Each experimental genotype was sown in triplicate on a 2 m2 plot; each plot had four rows with row spacing of 22.5 cm, a row length of 200 cm, and the seeding rate was 100 kg ha−1. The pH, electrical conductivity, and organic carbon of the top 15 cm of soil were 7.8, 0.76 sm−1, and 0.47%, respectively. The soil's N, P, and K contents were 224.3, 33.06, and 362.88 kg ha−1, respectively. The soil's micronutrients Fe, Mn, Cu, and Zn contents were 4.03, 3.60, 1.31, and 2.31 ppm, respectively. Recommended quantities of N (60 kg ha−1 pre-sowing and 60 kg ha−1 at 40 DAS) and P (60 kg ha−1 pre-sowing) were applied. Manual weeding was performed at 15 and 35 days after sowing to keep weeds below the economic threshold level.
Environment monitoring and plant measurements
The daily temperature in the field was recorded at 8:30 a.m. and 5:30 p.m. The ambient temperatures were obtained from the weather measuring equipment (Davis Vantage PRO2) installed at the adjacent field by the National Collateral Management Services Limited, Crop and Weather Intelligence Group (Table 3). The soil moisture was measured at heading stage when the irrigation was stopped in drought experiment to confirm that plants experience drought stress as compared to control. At the heading stage, the soil moisture was calculated by the gravimetric method. The soil moisture at heading stage in C, D and L was 18% (w/w), 4% (w/w) and 19% (w/w) averaged for 2018–2019 and 2019–2020. In 2020–2021, the soil moisture in C, D and L was 18% (w/w), 5% (w/w) and 16% (w/w).
At physiological maturity, when the spikes lost the green colour, three main shoot spikes were sampled from each plot. The number of spikelets spike−1, grains spike−1, and grain weight spike−1 (g) were determined. Spike length (cm) was measured, excluding awns. Grain yield plot−1 was measured in 2018–2019. Heads were visually classified into average, spelt and compact shapes103.
Image acquisition
At physiological maturity, three main shoot spikes of each genotype and environment were collected. Each grain of a spike was manually separated from the rachis and arranged corresponding to the spikelet position with the ventral grain surface downward and no contact between grains. Images were acquired with a Redmi Note 6 Pro smartphone with a fixed distance of 31 cm between the camera and the grains placed on a dark background with a scale at the side (Fig. S12). The total data set of three crop seasons consisted of 333 images (144 images for each of first two years, and 45 images in third year) of over 18,000 grains. Rachis images were also captured separately in the same way (Fig. S13).
Image analysis
Open-source software ImageJ version 1.46r was used to extract the features of grains from the images. A set of images (in JPEG format) were imported for processing simultaneously using the image sequence tool (File > Import > Image Sequence). The number of images processed in one session is limited as the available memory of ImageJ is 640 MB. A known length of a line was drawn along the scale in the first image. Using set scale tool (Analyze > Set Scale), the pixels were converted into millimeters by specifying the known length of the line in mm and clicking on checkbox “global” to apply this calibration to all the images. The RGB images of grains were subjected to thresholding and then converted into binary (Process > Binary > Make Binary) after clicking the checkbox calculate threshold of each image. The parameters of grains to be measured were selected using analyze > Set Measurements. The different parameters of each grain were measured using analyze particles tool (Analyze<Analyze Particles), by setting a lower limit of 2.5 mm2 to exclude any non-grain material. Each grain is numbered in sequence from the bottom left to the top right in an image, and an image containing numbered outlines of measured particles replaces the original image after the analysis. The grain parameters measured were area, perimeter, minimum (MinFeret) and maximum (Feret) caliper diameter, the major and minor axes of the best fitted ellipses to the grains (Major & Minor), Circularity (a value between 0 to 1 for an infinitely elongated shape to a perfect circle), etc. The rachis images were also analysed in same way. The results file were saved as CSV (comma-separated values) files for analysis.
Validation of grain length and width
The actual length and width of 100 reference grains was measured by using digital vernier calliper and these observed values were correlated with the values generated by image analysis (Figs. 1, 2). The Feret and MinFeret values from ImageJ were used as the length and width of grains. The experiment for correlation of actual grain length with Feret and actual grain width MinFeret was carried out twice to determine the consistency over estimation by ImageJ. Using the cv.lm function of the "DAAG" package of R, the predicted values were cross-validated tenfold33 (Figs. 1b, 2b). A k-fold cross-validation is performed for a linear model (also known as an "lm" model) via the function cv.lm. For each fold, prediction errors are computed for the observations (the "test set") and a "lm" model is fitted to all data that are not in the fold (the "training set"). The absolute error and its square make up the prediction errors. The square root of the average of the squared errors is calculated from the average prediction errors over the observations in the fold.
Measurement of intra-spike variation as standard deviation (ISVAD)
The MinFeret, Feret, area (projected surface area) and perimeter of each grain at different locations within the spike were used for calculating the ISVAD of grain width, ISVAD of grain length, ISVAD of grain area and ISVAD of grain perimeter within a spike and within top, middle and bottom regions of spike. ISVAD was derived from variance in statistics. The standard deviation was calculated using the following formula.
σ = Standard deviation in the grain size parameter within a spike (referred as ISVAD), N = Number of grains within a spike, xi = Value of the grain size parameter of each grain within a spike, µ = Mean value of all grains within a spike.
RNA sequencing
RNA extraction and high-throughput sequencing
The RNA sequencing of spike was done at 25 DAA to identify the genes involved in supply of assimilates to the developing grain and increase in dry mass content. Three biological replicates of the spike of the main stem of Sunstar and Raj 4079 were collected at 25 DAA stage under C, D and L environments in 2020–2021 and immediately snapped frozen in liquid nitrogen. Total RNA extraction was carried out using standard trizol protocol. The quality and quantity of RNA were confirmed by RNAse-free agarose gel electrophoresis, and concentration was measured using qubit, nanodrop and Agilent tapestation. High-quality RNA from all the samples was used for mRNA purification. The mRNA library was prepared using the TruSeq Stranded mRNA Kit_Plant. The libraries were then sequenced on Illumina Hiseq. X 10 platform at SciGenom Next-Gen sequencing facility, Cochin, India.
Preprocessing of RNA-Seq data and transcriptome profile analysis
In the preprocessing step of the raw reads, the adaptor sequences and low-quality bases were trimmed using AdapterRemoval-v2 (version 2.3.1). From the preprocessed reads, ribosomal RNA sequences were removed by aligning the reads with the silva database using bowtie2 (version 2.2.9) and subsequent workflow using samtools (version 1.9), sambamba (version 0.7.0), BamUtil (version 1.0.14) tools and in-house scripts. The preprocessed and rRNA removed reads were used for reference-based pair-wise alignment with Wheat Ensembl release 31 reference genome downloaded from Ensemble (http://ftp.ebi.ac.uk/ensemblgenomes/pub/plants/release-31/fasta/triticum_aestivum/dna/Triticum_aestivum.IWGSC1%2Bpopseq.dna.toplevel.fa.gz). The alignment was performed using the STAR program (version 2.7.6a).
Differential gene expression
After aligning the reads with the transcriptome, differential expression analysis for all pairs of samples comparisons was performed using the cuffdiff program of the cufflinks package (version 2.2.1). Absolute (Log2 fold change) _ 2 along with p-value < 0.05 separately were used as cut offs for identifying up and down regulated genes.
Statistical significance
The statistical significance of the data was calculated using “agricolae” package in R version 4.0.1104. The Duncan’s multiple range test (DMRT) was carried out at p < 0.05 and used as a posthoc test to separate the means where ANOVA indicated significant differences.
Statement
We confirm that experimental research and field studies on plants (either cultivated or wild), including the collection of plant material, comply with relevant institutional, national, and international guidelines and legislation.
Data availability
The datasets presented in this study can be found in online repositories and Supplementary material. The transcriptome data can be found in the National Genomics Data Centre (NGDC) database BioProject: PRJNA918542.
References
Li, Y. et al. Plant density effect on grain number and weight of two winter wheat cultivars at different spikelet and grain positions. PLoS ONE 5(11), e0155351. https://doi.org/10.1371/journal.pone.0155351 (2016).
Xie, Q., Mayes, S. & Sparkes, D. Carpel size, grain filling, and morphology determine individual grain weight in wheat. J. Exp. Bot. 66, 6715–6730 (2015).
Ferrante, A., Savin, R. & Slafer, G. A. Relationship between fruiting efficiency and grain weight in durum wheat. Field Crops Res. 177, 109–116. https://doi.org/10.1016/j.fcr.2015.03.009 (2015).
Feng, F. et al. The effect of grain position on genetic improvement of grain number and thousand grain weight in winter wheat in North China. Front. Plant Sci. 9, 1664–2462. https://doi.org/10.3389/fpls.2018.00129 (2018).
Prieto, P. et al. Dynamics of floret initiation/death determining spike fertility in wheat as affected by Ppd genes under field conditions. J. Exp. Bot. 69(10), 2633–2645. https://doi.org/10.1093/jxb/ery105 (2018).
Philipp, N. et al. Grain number and grain yield distribution along the spike remain stable despite breeding for high yield in winter wheat. PLoS ONE 13, e0205452 (2018).
McCormick, A., Cramer, M. D. & Watt, D. A. Sink strength regulates photosynthesis in sugarcane. New Phytol. 171, 759–770 (2006).
Acreche, M. M. & Slafer, G. A. Grain weight, radiation interception and use efficiency as affected by sink-strength in Mediterranean wheats released from 1940 to 2005. Field Crops Res. 110, 98–105 (2009).
Foulkes, M. J. et al. Raising yield potential of wheat. III. Optimizing partitioning to grain while maintaining lodging resistance. J. Exp. Bot. 62(2), 469–486. https://doi.org/10.1093/jxb/erq300 (2011).
Serrago, R. A. et al. Understanding grain yield responses to source–sink ratios during grain filling in wheat and barley under contrasting environments. Field Crops Res. 150, 42–51 (2013).
Aisawi, K. A. B. et al. The physiological basis of the genetic progress in yield potential of CIMMYT spring wheat cultivars from 1966 to 2009. Crop Sci. 55, 1749–1764 (2015).
Sharma, D. L. & Anderson, W. K. Small grain screenings in wheat: Interactions of genotypes with season, site, and management practices. Aust. J. Agric. Res. 55, 797. https://doi.org/10.1071/AR03265 (2004).
Nuttall, J. G. et al. Models of grain quality in wheat—A review. Field Crops Res. 202, 136–145 (2017).
Monneveux, P., Jing, R. & Misra, S. Phenotyping for drought adaptation in wheat using physiological traits. Front. Physiol. 3, 429. https://doi.org/10.3389/fphys.2012.00429 (2012).
Finch-Savage, W. E. & Bassel, G. W. Seed vigour and crop establishment: Extending performance beyond adaptation. J. Exp. Bot. 67, 567–591. https://doi.org/10.1093/jxb/erv490 (2016).
Evers, A. D. Grain size and morphology: Implications for quality. In Wheat Structure, Biochemistry and Functionality (ed. Schofield, D.) 19–24 (Royal Society of Chemistry, 2000).
Beral, A. et al. Wheat individual grain-size variance originates from crop development and from specific genetic determinism. PLoS ONE 15(3), e0230689. https://doi.org/10.1371/journal.pone.0230689 (2020).
Jenner, C. F. Starch synthesis in the kernel of wheat under high temperature conditions. Funct. Plant Biol. 21, 791–806 (1994).
Schittenhelm, S., Langkamp-Wedde, T., Kraft, M., Kottmann, L. & Matschiner, K. Effect of two-week heat stress during grain filling on stem reserves, senescence, and grain yield of European winter wheat cultivars. J. Agron. Crop Sci. 206, 722–733. https://doi.org/10.1111/jac.12410 (2020).
Bremner, P. M. Accumulation of dry matter and nitrogen by grains in different positions of the wheat war as influenced by shading and defoliation. Aust. J. Biol. Sci. 25, 657–668 (1972).
Asseng, S., Foster, I. & Turner, N. C. The impact of temperature variability on wheat yields. Glob. Change Biol. 17, 997–1012 (2011).
Farooq, M. et al. Heat stress in wheat during reproductive and grain-filling phases. Crit. Rev. Plant Sci. 30(6), 491–507. https://doi.org/10.1080/07352689.2011.615687 (2011).
Sofield, I. et al. Factors influencing the rate and duration of grain filling in wheat. Aust. J. Plant Physiol. 4(5), 785. https://doi.org/10.1071/pp9770785 (1977).
Tashiro, T. & Wardlaw, I. F. The effect of high temperature at different stages of ripening on grainset, grainweight and grain dimensions in the semi-dwarf wheat “banks”. Ann. Bot. 65, 51–61 (1990).
Zhao, H. et al. Leaf senescence and grain filling affected by post-anthesis high temperatures in two different wheat cultivars. Plant Growth Regul. 51, 149–158 (2007).
Barrero, J. et al. Evaluation of the impact of heat on wheat dormancy, late maturity α-amylase and grain size under controlled conditions in diverse germplasm. Sci. Rep. https://doi.org/10.1038/s41598-020-73707-8 (2020).
Sharma, D. L., Shackley, B. J. & Amjad, M. Use of grain size distribution parameters to explain variation in small grain screenings of wheat in multi-environment trials involving new cultivars. Crop Pasture Sci. 60, 658–666 (2009).
Meuwissen, T. H., Hayes B. J. & Goddard M. E. Prediction of total genetic value using genome-wide dense marker maps. Genetics. 157, 1819–1829. https://doi.org/10.1093/genetics/157.4.1819 (2001).
Le, T. D. Q., Alvarado, C., Girousse, C., Legland, D. & Chateigner-Boutin, A-L. Use of X-ray micro computed tomography imaging to analyze the morphology of wheat grain through its development. Plant Methods. 15(1), 84. https://doi.org/10.1186/s13007-019-0468-y (2019).
Xiong, B., Wang, B., Xiong, S., Lin, C. & Yuan, X. 3D morphological processing for wheat spike phenotypes using computed tomography images. Remote Sens. 11(9), 1110 (2019).
Schmidt, J. et al. Drought and heat stress tolerance screening in wheat using computed tomography. Plant Methods. 16(1), 15. https://doi.org/10.1186/s13007-020-00565-w (2020).
Zhou, H. et al. Determination of wheat spike and spikelet architecture and grain traits using X-ray Computed Tomography imaging. Plant Methods. 17(1), 26. https://doi.org/10.1186/s13007-021-00726-5 (2021).
Bay, T. F. & Schoney, R. A. Data analysis with computer graphics: Production functions. Am. J. Agric. Econ. 64, 289–297. https://doi.org/10.2307/1241138 (1982).
Kumar, P. V. et al. Optimizing sowing dates and selection of varieties of wheat through long-term crop and weather analysis. J. Agrometeorol. 15(Special Issue II), 67–72 (2013).
Warnes, G. R. et al. gplots: Various R Programming Tools for Plotting Data. https://CRAN.R-project.org/package=gplots (2019).
Haghshenas, A., Emam, Y. & Jafarizadeh, S. Wheat grain width: a clue for re-exploring visual indicators of grain weight. Plant Methods. 18, 58. https://doi.org/10.1186/s13007-022-00891-1 (2022).
Breseghello F. & Sorrells M. E. QTL analysis of kernel size and shape in two hexaploid wheat mapping populations. Field Crops Res. 101, 172–179. https://doi.org/10.1016/j.fcr.2006.11.008 (2007).
Herridge, R. P., Day, R. C., Baldwin, S. & Macknight, R. C. Rapid analysis of seed size in Arabidopsis for mutant and QTL discovery. Plant Methods. 7, 11–10. https://doi.org/10.1186/1746-4811-7-3 (2011).
Li, D. et al. Engineering OsBAK1 gene as a molecular tool to improve rice architecture for high yield. Plant Biotechnol J. 7, 791–806. https://doi.org/10.1111/j.1467-7652.2009.00444.x (2009).
Igathinathane, C., Pordesimo, L. O. & Batchelor, W. D. Major orthogonal dimensions measurement of food grains by machine vision using ImageJ. Food Res. Int. 42, 76–84. https://doi.org/10.1016/j.foodres.2008.08.013 (2009).
Liu, Y. et al. Identification of QTL for flag leaf length in common wheat and their pleiotropic effects. Mol. Breed. 38, 11. https://doi.org/10.1007/s11032-017-0766-x (2018).
Yang, J. et al. Molecular characterization of a novel TaGL3-5A allele and its association with grain length in wheat (Triticum aestivum L.). Theor. Appl. Genet. 132, 1799–1814. https://doi.org/10.1007/s00122-019-03316-1 (2019).
Desiderio, F. et al. Genomic regions from an Iranian landrace increase kernel size in durum wheat. Front. Plant Sci. 10, 448. https://doi.org/10.3389/fpls.2019.00448 (2019).
Ma, J. et al. Identification of quantitative trait loci for kernel traits in a wheat cultivar Chuannong16. BMC Genet. 20, 77. https://doi.org/10.1186/s12863-019-0782-4 (2019).
Liu, H. et al. Several stably expressed QTL for spike density of common wheat (Triticum aestivum) in multiple environments. Plant Breed. 139, 284–294. https://doi.org/10.1111/pbr.12782 (2020).
Ma, J. et al. Flag leaf size and posture of bread wheat: Genetic dissection, QTL validation and their relationships with yield-related traits. Theor. Appl. Genet. 133, 297–315. https://doi.org/10.1007/s00122-019-03458-2 (2020).
Magallanes-López, A. M. et al. Grain quality traits of commercial durum wheat varieties and their relationships with drought stress and glutenins composition. J. Cereal Sci. 75, 1–9. https://doi.org/10.1016/j.jcs.2017.03.005 (2017).
Guzmán, C. et al. Response to drought and heat stress on wheat quality, with special emphasis on bread-making quality, in durum wheat. Field Crops Res. 186, 157–165. https://doi.org/10.1016/j.fcr.2015.12.002 (2016).
Guzmán, C. et al. Use of rapid tests to predict quality traits of CIMMYT bread wheat genotypes grown under different environments. LWT 69, 327–333. https://doi.org/10.1016/j.lwt.2016.01.068 (2016).
Magallanes-López, A. M. Variability in iron, zinc and phytic acid content in a worldwide collection of commercial durum wheat cultivars and the effect of reduced irrigation on these traits. Food Chem. 237, 499–505. https://doi.org/10.1016/j.foodchem.2017.05.110 (2017).
Velu, G. et al. Genomic prediction for grain zinc and iron concentrations in spring wheat. Theor. Appl. Genet. 129, 1595–1605. https://doi.org/10.1007/s00122-016-2726-y (2016).
Fleitas, M. C. et al. Identification of CIMMYT spring bread wheat germplasm maintaining superior grain yield and quality under heat-stress. J. Cereal Sci. 93, 102981. https://doi.org/10.1016/j.jcs.2020.102981 (2020).
Hernández-Espinosa, N. et al. Preliminary characterization for grain quality traits and high and low molecular weight glutenins subunits composition of durum wheat landraces from Iran and Mexico. J. Cereal Sci. 88, 47–56. https://doi.org/10.1016/j.jcs.2019.05.007 (2019).
Gegas, V. C. et al. A genetic framework for grain size and shape variation in wheat. Plant Cell. 22(4), 1046–1056. https://doi.org/10.1105/tpc.110.074153 (2010).
Neuweiler, J. E., Maurer, H. P. & Würschum, T. Long-term trends and genetic architecture of seed characteristics, grain yield and correlated agronomic traits in triticale (× Triticosecale wittmack). Plant Breed. 139, 717–729. https://doi.org/10.1111/pbr.12821 (2020).
Sanchez-Bragado, R., Kim, J. W. & Rivera-Amado, C. Are awns truly relevant for wheat yields? A study of performance of awned/awnless isogenic lines and their response to source–sink manipulations. Field Crops Res. 254, 107827. https://doi.org/10.1016/j.fcr.2020.107827 (2020).
Tanabata, T., Shibaya, T., Hori, K., Ebana, K. & Yano, M. SmartGrain: High-throughput phenotyping software for measuring seed shape through image analysis. Plant Physiol. 160, 1871–1880. https://doi.org/10.1104/pp.112.205120 (2012).
Whan, A. P. et al. GrainScan: A low cost, fast method for grain size and colour measurements. Plant Methods 10, 23. https://doi.org/10.1186/1746-4811-10-23 (2014).
Okamoto, Y., Nguyen, A. T., Yoshioka, M., Iehisa, J. C. M. & Takumi, S. Identification of quantitative trait loci controlling grain size and shape in the D genome of synthetic hexaploid wheat lines. Breed. Sci. 63, 423–429. https://doi.org/10.1270/jsbbs.63.423 (2013).
Sestili, F. et al. Enhancing grain size in durum wheat using RNAi to knockdown GW2 genes. Theor. Appl. Genet. 132, 419–429. https://doi.org/10.1007/s00122-018-3229-9 (2019).
Kumari, S., Mir, R. R., Tyagi, S., Balyan, H. S. & Gupta, P. K. Validation of QTL for grain weight using MAS-derived pairs of NILs in bread wheat (Triticum aestivum L.). J. Plant Biochem. Biot. 28, 336–344. https://doi.org/10.1007/s13562-018-0485-3 (2019).
Yoshioka, M. et al. Genetic dissection of grain morphology in hexaploid wheat by analysis of the NBRP-Wheat core collection. Genes Genet. Syst. 94, 35–49. https://doi.org/10.1266/ggs.18-00045 (2019).
Kumari, S. et al. QTL mapping for some grain traits in bread wheat (Triticum aestivum L.). Physiol. Mol. Biol. Plants 24, 909–920. https://doi.org/10.1007/s12298-018-0552-1 (2018).
Bowerman, A. F. et al. Suppression of glucan, water dikinase in the endosperm alters wheat grain properties, germination and coleoptile growth. Plant Biotechnol. J. 14, 398–408. https://doi.org/10.1111/pbi.12394 (2016).
Alemu, A. et al. Genome-wide association mapping for grain shape and color traits in Ethiopian durum wheat (Triticum turgidum ssp. durum). Crop J. https://doi.org/10.1016/j.cj.2020.01.001 (2020).
Sakai, Y. et al. QTLs for agronomic traits detected in recombinant inbred lines derived from a bread wheat × spelt cross. Breed. Sci. 68, 587–595. https://doi.org/10.1270/jsbbs.18046 (2018).
Okada, M. et al. Phenotypic effects of the U-genome variation in nascent synthetic hexaploids derived from interspecific crosses between durum wheat and its diploid relative Aegilops umbellulata. PLoS ONE 15, e0231129 (2020).
Takumi, S. et al. Introgression of chromosomal segments conferring early heading date from wheat diploid progenitor, Aegilops tauschii Coss., into Japanese elite wheat cultivars. PLoS ONE 15, e0228397. https://doi.org/10.1371/journal.pone.0228397 (2020).
Abid, M. et al. Pre-drought priming sustains grain development under post-anthesis drought stress by regulating the growth hormones in winter wheat (Triticum aestivum L.). Planta 246, 509–524. https://doi.org/10.1007/s00425-017-2698-4 (2017).
Kehel, Z. et al. Predictive characterization for seed morphometric traits for genebank accessions using genomic selection. Front. Ecol. Evol. 8, 32 (2020).
Hughes, A. et al. Non-destructive, high-content analysis of wheat grain traits using X-ray micro computed tomography. Plant Methods 13, 76. https://doi.org/10.1186/s13007-017-0229-8 (2017).
Schmidt, J. et al. Drought and heat stress tolerance screening in wheat using computed tomography. Plant Methods 16, 15. https://doi.org/10.1186/s13007-020-00565-w (2020).
Semenov, M. A., Stratonovitch P., Alghabari, F. & Gooding, M. J. Adapting wheat in Europe for climate change. J. Cereal Sci. 59, 245–256 (2014).
Barber, H. M. et al. Temporally and genetically discrete periods of wheat sensitivity to high temperature. Front. Plant Sci. 8, 51 (2017).
Lizana, X. C. et al. Expansins expression is associated with grain size dynamics in wheat (Triticum aestivum L.). J. Exp. Bot. 61, 1147–1157. https://doi.org/10.1093/jxb/erp380 (2010).
Nadaud, I. et al. Proteomic and morphological analysis of early stages of wheat grain development. Proteomics. 10, 2901–2910. https://doi.org/10.1002/pmic.200900792 (2010).
Prasad, P. V. V. & Djanaguiraman, M. Response of floret fertility and individual grain weight of wheat to high temperature stress: Sensitive stages and thresholds for temperature and duration. Funct. Plant Biol. 41(12), 1261–1269. https://doi.org/10.1071/FP14061 (2014).
Dreccer, M. F., Barnesa, L. & Mederb, R. Quantitative dynamics of stem water soluble carbohydrates in wheat can be monitored in the field using hyperspectral reflectance. Field Crops Res. 159, 70–80 (2014).
Guedira, M. & Paulsen, G. M. Accumulation of starch in wheat grain under different shoot/root temperatures during maturation. Funct. Plant Biol. 29(4), 495–503. https://doi.org/10.1071/PP01006 (2002).
Guo, Z. & Schnurbusch, T. Variation of floret fertility in hexaploid wheat revealed by tiller removal. J. Exp. Bot. 66, 5945–5958 (2015).
Evers, A. D. et al. Predicting milling extraction rate by image analysis of wheat grains. Asp Appl. Biol. 25, 417–426 (1990).
Reynolds, M. et al. Addressing research bottlenecks to crop productivity. Trends Plant Sci. 26(6), 607–630 (2021).
Plaut, Z. et al. Transport of dry matter into developing wheat kernels and its contribution to grain yield under post-anthesis water deficit and elevated temperature. Field Crops Res. 86, 185–198 (2004).
Lv, X. et al. Source-sink modifications affect leaf senescence and grain mass in wheat as revealed by proteomic analysis. BMC Plant Biol. 20, 257. https://doi.org/10.1186/s12870-020-02447-8 (2020).
Ferrante, A., Savin, R. & Slafer, G. A. Floret development and grain setting differences between modern durum wheats under contrasting nitrogen availability. J. Exp. Bot. 64, 169–184 (2013).
Li, G. et al. Limitation of unloading in the developing grains is a possible cause responsible for low stem non-structural carbohydrate translocation and poor grain yield formation in rice through verification of recombinant inbred lines. Front. Plant Sci. 8, 1369 (2017).
Reynolds, M. et al. Raising yield potential in wheat. J. Exp. Bot. 60(7), 1899–1918. https://doi.org/10.1093/jxb/erp016 (2009).
Bremner, P. & Rawson, H. The weights of individual grains of the wheat ear in relation to their growth potential, the supply of assimilate and interaction between grains. Funct. Plant Biol. 5, 61–72 (1978).
Hütsch, B. W., Jahn, D. & Schubert, S. Grain yield of wheat (Triticum aestivum L.) under long-term heat stress is sink-limited with stronger inhibition of kernel setting than grain filling. J. Agron. Crop Sci. https://doi.org/10.1111/jac.12298 (2018).
Guo, Z. et al. Genome-wide association analyses of 54 traits identified multiple loci for the determination of floret fertility in wheat. New Phytol. 214, 257–270 (2017).
Lopez-Garrido, E. et al. Quantification of vascular tissues in the peduncle of durum wheat cultivars improved during the twentieth century. Int. J. Dev. Biol. 45(S1), S47–S48 (2001).
Egli, D. B. Seed growth and seed fill duration: Variation and regulation. Seed Biology and the yield of grain crops. 38–69 (CAB International, 1998).
Brinton, J. & Uauy, C. A. Reductionist approach to dissecting grain weight and yield in wheat. J. Integr. Plant Biol. 61, 337–358. https://doi.org/10.1111/jipb.12741 (2019).
Wu, X., Liu, J., Li, D. & Liu, C. M. Rice caryopsis development I: Dynamic changes in different cell layers. J. Integr. Plant Biol. 58(9), 772–785. https://doi.org/10.1111/jipb.12440 (2016) (Epub 2015 Dec 16).
Li, Y. et al. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat. Gen. 43, 1266–1269 (2011).
Xu, C. et al. Differential expression of GS5 regulates grain size in rice. J. Exp. Bot. 66(9), 2611–2623. https://doi.org/10.1093/jxb/erv058 (2015) (Epub 2015 Feb 24).
Hurkman, W. J. et al. Effect of temperature on expression of genes encoding enzymes for starch biosynthesis in developing wheat endosperm. Plant Sci. 164(5), 873–881. https://doi.org/10.1016/S0168-9452(03)00076-1 (2003).
Hütsch, B. W. & Schubert, S. Harvest index of maize (Zea mays L.): Are there possibilities for improvement? Adv. Agron. 146, 37–82 (2017).
Jung, S., Hütsch, B. W. & Schubert, S. Salt stress reduces kernel number of corn by inhibiting plasma membrane H+-ATPase activity. Plant Physiol. Biochem. 113, 198–207. https://doi.org/10.1016/j.plaphy.2017.02.009 (2017).
Wu, X. et al. Precise mapping of a quantitative trait locus interval for spike length and grain weight in bread wheat (Triticum aestivum L.). Mol. Breed. 33, 129–138 (2014).
Yao, H. et al. HL2 on chromosome 7D of wheat (Triticum aestivum L.) regulates both head length and spikelet number. Theor. Appl. Genet. 132(6), 1789–1797 (2019).
Kumar, S. et al. Effect of irrigation levels and moisture conserving polymers on growth, productivity and profitability of wheat. Indian J. Agric. Sci. 89(3), 133–138. https://doi.org/10.56093/ijas.v89i3.87597 (2019).
Konopatskaia, I. et al. Spike morphology genes in wheat species (Triticum L.). Proc. Latv. Acad. Sci. B: Nat. Exact Appl. Sci. 70, 345–355. https://doi.org/10.1515/prolas-2016-0053 (2016).
R Core Team R: a language and environment for statistical computing. https://www.R-project.org/ (Vienna, Austria: R Foundation for statistical Computing, 2016) (Accessed 3 Sept 2021).
Acknowledgements
The authors acknowledge the Director, ICAR-Indian Institute of Wheat & Barley Research, Karnal for providing studied wheat germplasm. We acknowledge the DST WOS-A scheme, Government of India for providing funding at the Department of Bioscience and Biotechnology, BanasthaliVidyapith, India. We thank the Department of Bioscience and Biotechnology, Banasthali Vidyapith and the Farm Science Centre, Banasthali Vidyapith, Tonk for providing the necessary facilities.
Funding
This research was funded by Women Scientist-A scheme by Department of Science and Technology, Government of India at the Department of Bioscience and Biotechnology, Banasthali Vidyapith (Grant number: SR/WOS-A/LS-162/2017).
Author information
Authors and Affiliations
Contributions
Conceptualization, V.T. and J.R.; methodology, V.T. and J.R.; software, V.T., J.R. and G.P.; validation, V.T. and J.R.; formal analysis, V.T. and J.R.; resources, V.T., G.P. and J.R.; sequence raw data submission on NCBI, S.Y.; writing—original draft preparation, V.T.; writing—review and editing, V.T. and J.R.; supervision, J.R. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thakur, V., Rane, J., Pandey, G.C. et al. Image facilitated assessment of intra-spike variation in grain size in wheat under high temperature and drought stress. Sci Rep 13, 19850 (2023). https://doi.org/10.1038/s41598-023-44503-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-44503-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.