Abstract
The serotonin transporter (5-HTT) critically shapes serotonin neurotransmission by regulating extracellular brain serotonin levels; it remains unclear to what extent 5-HTT levels in the human brain are genetically determined. Here we applied [11C]DASB positron emission tomography to image brain 5-HTT levels and evaluated associations with five common serotonin-related genetic variants that might indirectly regulate 5-HTT levels (BDNF rs6265, SLC6A4 5-HTTLPR, HTR1A rs6295, HTR2A rs7333412, and MAOA rs1137070) in 140 healthy volunteers. In addition, we explored whether these variants could predict in vivo 5-HTT levels using a five-fold cross-validation random forest framework. MAOA rs1137070 T-carriers showed significantly higher brain 5-HTT levels compared to C-homozygotes (2–11% across caudate, putamen, midbrain, thalamus, hippocampus, amygdala and neocortex). We did not observe significant associations for the HTR1A rs6295 and HTR2A rs7333412 genotypes. Our previously observed lower subcortical 5-HTT availability for rs6265 met-carriers remained in the presence of these additional variants. Despite this significant association, our prediction models showed that genotype moderately improved prediction of 5-HTT in caudate, but effects were not statistically significant after adjustment for multiple comparisons. Our observations provide additional evidence that serotonin-related genetic variants modulate adult human brain serotonin neurotransmission.
Similar content being viewed by others
Introduction
Serotonin neurotransmission mediates a multitude of brain functions, including neurodevelopment, behavior and cognition1. As such, identifying sources of individual variation in brain serotonin neurotransmission is relevant to identify mechanisms contributing to variation in behavior and possibly associated risks for disease. Dysregulation in brain serotonin signaling is implicated in a number of neuropsychiatric disorders such as anxiety and depression, which are known to have a prominent genetic component2,3,4,5. However, the contributions of genetic factors to individual differences in in vivo serotonin neurotransmission are not well understood.
The serotonin transporter (5-HTT) critically shapes serotonin neurotransmission as it facilitates serotonin reuptake, thereby regulating extracellular serotonin levels and associated receptor signaling1. It is also the pharmacological target for selective serotonin reuptake inhibitors (SSRIs), the most commonly prescribed class of antidepressants and anxiolytics6. 5-HTT availability can be visualized in humans in vivo using [11C]DASB positron emission tomography (PET)7,8.
Previous research has reported a link between 5-HTT levels and both healthy and pathological behavioral phenotypes. Increased 5-HTT availability (expressed in terms of binding potential, 5-HTT BP) has been linked to depressive symptoms severity in seasonal affective disorder9, greater negative affective bias10,11 and reduced amygdala reactivity to threat in healthy individuals12,13, whereas low 5-HTT availability has been associated with childhood abuse in patients with depression vs patients who did not experience childhood abuse14. Notably, some studies have speculated that 5-HTT availability may be used as a marker of serotonin tone and a histochemical marker for serotonergic projections15,16. Thus, identifying sources of variation in 5-HTT availability is relevant to advance our understanding of mechanisms driving individual variation in human behavior.
Evidence from PET studies supports a genetic effect on the brain distribution of receptors involved in serotonin function17,18,19. Similarly, genetic variation is likely to contribute to 5-HTT levels and activity. Multiple genetic variants have been hypothesized to affect brain serotonin signaling and behavior as well as brain function. Whereas human functional neuroimaging genetics with, e.g., BOLD fMRI, is challenging due to the inherently complex processes underlying brain function20, PET imaging provides a direct estimate of brain protein levels that is highly reproducible, making it a valuable target for relating to genetic variation.
5-HTTLPR, a common insertion/deletion polymorphism occurring in the promoter region of the 5-HTT gene (SLC6A4) is the most widely investigated 5-HTT variant. Nonetheless, although 5-HTTLPR was shown to affect 5-HTT gene expression in vitro in early studies3, PET studies in humans reported conflicting in vivo results21,22,23,24. In contrast, single nucleotide polymorphisms (SNP) in genes coding for proteins whose activity is related to that of 5-HTT, such as rs6265 (or Val66Met) in the neurotrophic factor (BDNF) gene23 and rs7333412 in the serotonin receptor 2A (5-HT2A) gene (HT2AR) were found to be associated with 5-HTT availability in healthy adults25,26. Specifically, the BDNF rs6265 met-carriers, who presumably have lower BDNF levels, showed increased 5-HTT availability in subcortical regions23, whereas the rs7333412 A-carriers showed reduced 5-HTT levels. BDNF is a common neurotrophin mediating neurodevelopmental, survival and plasticity functions whose activity levels can affect serotonin release 27 as well as 5-HTT expression in animal models28, while 5-HT2A is a core regulator of excitatory serotonin neurotransmission. These findings suggest that genetic variants other than those in the SLC6A4 gene may indirectly modulate 5-HTT availability by affecting serotonergic signaling via other pathways.In this framework, serotonin receptor 1A (5-HT1A, encoded by gene HTR1A), which is the principal inhibitory serotonin receptor that can inhibit serotonin release29, and monoamine oxidase A (MAOA, encoded by gene MAOA), the main enzyme involved in serotonin degradation30 can directly affect serotonin levels, which may in turn affect 5-HTT levels via its downregulation 31. Previous evidence reports that HTR1A rs6295 G-carriers have increased 5-HT1A protein levels in the dorsal raphe, pointing to decreased serotonin tone that may in turn affect 5-HTT levels in humans 34. MAOA rs1137070 has been linked to increased MAOA mRNA levels and higher enzymatic activity, suggesting that this variant may directly affect serotonin levels35 and it was associated to SSRI treatment outcome36,37, suggesting a possible interaction with 5-HTT.
For explanatory purposes, an overview of the relationship between the serotonin-related genetic variants mentioned above and clinical or behavioral phenotypes is provided in Supplementary Table 1.
Previous studies have focused on the association between genetic variant and 5-HTT availability. However, although observing associations informs on group differences, it does not establish the ability to predict features at the individual level, which could be relevant for, e.g., personalized medicine strategies. To this end, exploring whether genetic variants assist in predicting 5-HTT availability can be of complementary value.
In this study we used the largest currently available dataset (N = 140) of [11C]DASB PET scans from the Cimbi database38 to explore the role of BDNF rs6265 and SLC6A4 5-HTTLPR(previously investigated in the same cohort23) rs7333412 in HT2AR25,26, rs6295 in HTR1A32,39,40, and rs1137070 in MAOA36,41 in 5-HTT availability in the healthy brain. First, we evaluated whether the genotypes examined were associated with 5-HTT availability. Next, we used random forest to determine whether genetic information predicted regional 5-HTT availability.
Methods
Participants
Cross-sectional data were included that was collected previously and were available from the Cimbi database38.
We selected healthy participants based on the following inclusion criteria: (1) availability of BDNF val66met (rs6265) and SLC6A4 (5-HTTLPR and SNP rs23351) genotypes, (2) availability of blood samples for additional genotyping, (3) availability of a [11C]DASB PET scan before any intervention, 4) ⩽60 years of age (to avoid age-related biases in brain volumes, as partial volume effects become stronger after 60), 5) self-identification with European ancestry. In addition, we excluded participants who had the following: (1) diagnosis of a severe neurological or systemic disease; (2) diagnosis of a primary psychiatric disease; (3) substance or drug abuse, based on self-reported clinical history and neurological/physical examination.
We identified 140 healthy participants, 84 females and 56 males (mean age: 26.7 ± 7.2; range: 18–51). The sex imbalance is partly because some studies from which data are drawn included only males or females. Demographic data are described in Table 1.
Subsets of the PET and genetic data included in the current study were collected as parts of multiple previous studies and have been included in previous publications. PET scans included in the current analyses were acquired between 2005 and 201511,18,42,43,44,45,46. All research protocols were approved by the Ethics Committee of Copenhagen and Frederiksberg, Denmark ((KF) 01–124/04, (KF) 01–156/04, (KF) 01 2006–20, H-1–2010-085, H-1–2010-91, H-2–2010-108, amendments included). All participants provided written informed consent after receiving a detailed description of the respective study. All experimental procedures were carried out in compliance with the declaration of Helsinki.
Data included in the current study has been utilized in previous studies9,10,23,45,47, some of which focused on the relation between 5-HTTLPR and/or BDNF rs6265 [11C]DASB binding23,45.
Genotyping
No additional genotyping for the 5-HTTLPR (including rs25531) and BDNF rs6265 was performed beyond that which has been described previously23,23,48,48,49.
The three additional variants were determined from whole-blood derived genomic DNA using QIAamp DNA Blood Mini Kit (Qiagen, Valencia, CA). DNA concentration and purity levels were estimated using an UV–Vis spectrophotometer (Nanodrop 2000, Thermo Scientific).
The SNPs were determined using TaqMan SNP Genotyping Assay (Applied Biosystems, Foster City, CA) with genotype-specific probes (BDNF rs6265: C_11592758_10, HTR1A rs6295: C__11904666_10, HTR2A rs7333412: C__29235757, MAOA rs1137070: C___8878813_20). We performed real-time polymerase chain reaction for allelic discrimination using the LightCycler 480 Real-Time PCR System (Roche Diagnostics, IN).
MRI data acquisition
For each participant, high-resolution T1-weighted structural brain scans were acquired on either a Siemens Magnetom Trio 3 T (N = 81) or a Siemens Verio 3 T (N = 59) (Siemens, Erlangen, Germany magnetic resonance imaging (MRI) scanner. We used structural MRI scans for segmentation and to delineate regions of interest in the PET scans.
[11C]DASB PET data acquisition
We acquired PET scans for each participant on one of two PET scanners: 1) a Siemens ECAT high-resolution research tomography (HRRT) scanner operating in 3D-acquisition mode with an in-plane resolution of approximately 2 mm (N = 98) or 2) an 18-ring GE-Advance scanner (General Electric, Milwaukee, WI, USA) with a three-dimensional (3D) acquisition mode and an in-plane resolution of approximately 6 mm (N = 42). PET images were acquired on the HRRT scanner using a 6-min transmission scan followed by an intravenous bolus injection of [11C]DASB that was given over 20 s while a dynamic 90-min emission scan was acquired over 36 frames (6 × 10 s, 3 × 20 s, 6 × 30 s, 5 × 60 s, 5 × 120 s, 8 × 300 s, 3 × 600 s). Dynamic PET images were reconstructed with an iterative OP-OSEM3D method using resolution modeling (10 iterations, 16 subsets)50,51. Images acquired on the GE-Advance scanner involved a 10-min transmission scan followed by a bolus injection given over 12 s while a 90-min dynamic emission scan was acquired over 36 frames (same time frames as HRRT acquisitions). Dynamic PET images from the GE-Advance scanner were reconstructed using filtered back projection and were corrected for attenuation, dead-time and scatter with a 6-mm Hann filter.
For both scanners, we determined single-subject PET scan motion and realignment using an automatic image registration algorithm52. Next, we smoothed the PET images using a 10 mm (HRRT) or a 12 mm (GE-Advance) within-frame Gaussian filter and subsequently aligned the volumes. Using the scaled least squares cost function, we estimated the rigid translation/rotation parameters that align each PET frame to a reference frame with sufficient structural information (frame 26: 20-25 min post injection). We resliced non-filtered PET images and co-registered the high-resolution MR with PET using SPM (HRRT) or automatic image registration (GE-Advance). Co-registration was based on the mean of frames 10–26, i.e., a flow-weighted image; the accuracy of this step was confirmed visually. We automatically delineated brain regions on structural MRI scans using Pvelab53. We determined the time-activity curves across gray matter voxels within each region except for the midbrain region, where we used the mean time-activity across all voxels. We determined regional 5-HTT non-displaceable binding potential (BPND) using kinetic modeling of regional time-activity curves in PMOD (Zurich, Switzerland) We applied the multilinear reference tissue model (MRTM/MRTM2) with a fixed k2’ estimated for each individual in a high binding region (caudate, putamen, and thalamus) and with cerebellum as reference region54. We calculated regional BPND values bilaterally, computing a volume-weighted mean from the right and left hemisphere.
Data analysis
We carried out the statistical analyses in R v4.1.2 (https://cran.r-project.org/). Consistent with previously related analyses, we considered regional 5-HTT BPND within caudate, putamen, midbrain, thalamus, hippocampus, amygdala and neocortex as our regions of interest23. We selected a single neocortical region (including orbitofrontal-, parietal- and occipital cortex and superior frontal-, pre/post central-, superior temporal- and middle/inferior frontal gyrus) because of previous evidence of high correlation between different cortical areas55. For all analyses, we grouped genotypes as follows: SLC6A4 5-HTTLPR and rs23351: LALA versus S’- carriers (individuals carrying at least one S- or one LG allele); HTR1A rs6295: G-carriers (carrying at least one G-allele) vs CC homozygotes; HTR2A rs7333412: G- carriers vs AA homozygotes; MAOA rs1137070: T-carriers versus CC; BDNF rs6265: met-carriers versus val/val. In addition, we included age, sex, MRI and PET scanner type as covariates, consistent with previous findings23. We mean-centered all continuous variables. Although previous evidence showed a seasonal and body mass index (BMI) effect on 5-HTT availability45,55, no statistically significant effect of neither daylight minutes nor BMI on 5-HTT BPND was previously observed on the same cohort23 so these variables were not included as covariates. We considered additional lifestyle factors, including alcohol consumption (i.e., alcohol units per week), smoking status, total scores of the Pittsburgh Sleep Quality Index56 and of the Cohen Perceived Stress Scale57. None of these measures were significantly associated with regional 5-HTT BPND (p > 0.05) and were not included as covariates in the main analyses. For all the models, we set the statistical significance threshold to p < 0.05 (two-sided tests).
Association analyses
We fit a linear latent variable model (LVM) to evaluate associations between genotypes and 5-HTT BPND within our set of pre-defined brain regions (i.e., caudate, amygdala, hippocampus, putamen, thalamus, midbrain and neocortex) as described previously23. LVMs are a type of multivariate linear regression that effectively models associations in the presence of inter-correlated variables. In this case, 5-HTT BPND is highly intercorrelated between the brain regions that we considered23. Thus, using the lava package v 1.6.1058 in R, we modeled this shared correlation of regional 5-HTT BPND values with a latent variable (5-HTTLV). Next, we modeled all the genotype and covariate effects on 5-HTTLV. We included additional covariance links (caudate-putamen, amygdala-hippocampus and thalamus-midbrain) based on the model framework previously reported23. In addition, we used Wald tests of improvement in model fit to find additional paths. To control the false-discovery rate across all possible paths, we included the paths with a false-discovery rate of pFDR < 0.05 calculated using the Benjamini–Hochberg test across all paths.
We used caudate as a reference region, so that the covariate effects reported here can be interpreted as effects on caudate BPND (corresponding to the reference scale).
In addition, we estimated multiple linear regression models including all genotypes and covariates (age, sex, PET and MR scanner type) for each brain region and we used them to report percent differences in 5-HTT BPND between genotype groups. We reported all results with parameter estimates and 95% confidence intervals within brackets, including the associated units (Fig. 1).
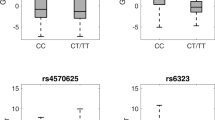
(A) Latent variable model (LVM) used to compute the association between each genotype independently from the other genotypes and 5-HTT BPND in caudate, putamen, midbrain, thalamus, hippocampus, amygdala and neocortex. PET and MRI scanner are not shown but were included as covariates. Yellow hatched boxes to the left represent the genetic variants and the covariates. The genetic variants depicted correspond to the following genes respectively: 5-HTTLPR- > SLC6A4, rs6295- > HTR1A, rs7333412- > HTR2A, rs1137070- > MAOA, rs6265- > BDNF. The orange boxes represent the covariate effects on the latent variable (5-HTTLV), which is represented by the central blue ellipse. Light blue boxes show the loadings on the latent variable of observed regional 5-HTT BPND (in the blue solid boxes to the right). β values in light blue and orange boxes indicate the parameter estimates for each model parameter with either its respective p-value (orange boxes) or 95% confidence interval (light blue). Hatched lines between regions indicate interregional shared correlations. Hatched circles on the brain regions represent the included error estimates. Arrows from the yellow to the blue boxes (sex- > caudate, rs6265- > neocortex) represent direct covariate effects on binding. All brain regions significantly loaded on to 5-HTTLV (p < 1 × 10–12). (B) Boxplots showing representative effects of HTR1A rs6295, HTR2A rs7333412 and MAOA rs1137070 on caudate 5-HTT BPND. The y axis represents 5-HTT BPND, adjusted for age, sex, MRI scanner, and PET scanner. Gray dots represent 5-HTT BPND from each participant adjusted for age, sex, MRI and PET scanner. The larger solid dots and lines represent respective group means and ± 1 SD. The boxes represent datapoints from the 25% to the 75% quantile.5-HTT: serotonin transporter; SLC6A4: 5-HTT gene; HTR1A: serotonin receptor 1A gene; HTR2A: serotonin receptor 2 gene; MAOA: monoamine oxidase A gene; BDNF: brain-derived neurotrophic factor gene; MRI: magnetic resonance imaging (MRI); PET: positron emission tomography; BPND: non-displaceable binding potential.
Finally, we tested for a main effect of each variant using a likelihood ratio test comparing a model including all covariates and genotypes with a model including all covariates but not genotypes.
Prediction analyses
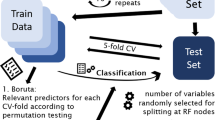
To determine whether genotype information predicts regional 5-HTT BPND, we trained and tested a random forest model59. We used the randomForest package v.4.7–1.1 in R with default data sampling and model fitting parameters; this included p/3 sampled features per tree (where p represents the total number of features) and 500 trees per forest. As we were specifically interested in how well genetic information predicted 5-HTT BPND, above and beyond other covariates, we constrained our feature set to include only the five genotypes and we evaluated the prediction of regional 5-HTT BPND, adjusted for relevant covariates. First, we fitted a multiple linear regression model regressing each region, e.g., caudate 5-HTT BPND, against age, sex, PET-scanner, and MR-scanner. The residuals from this multiple linear regression model were used as the outcome in our random forest machine learning models with genotype status for rs6265, 5-HTTLPR and rs23351, rs6295, rs7333412, and rs1137070 as model features. Prediction models were estimated using five-fold cross-validation; prediction of 5-HTT BPND on held out datasets (test data) within each fold was used to determine unbiased, predictive model performance. Model performance was calculated as the root mean-squared error (RMSE) of prediction on test data, across all folds. To account for fold-assignment related variance, model performance was assessed by repeating the five-fold cross-validation 10 times; overall performance was the mean of these 10 resamples. Statistical significance of performance was calculated from an empirical null distribution derived from 10,000 permutations of the resampled residuals. Notably, we performed five-fold cross-validation with 10 resamples within each of the 10,000 permutations so that each permutation more closely reflects our observed model structure. We fitted models for 5-HTT BPND in each of our seven regions of interest separately, i.e., seven different prediction models. In addition to expressing model performance in terms of RMSE, we express the percent change in RMSE of the prediction models accounting for genotype information (RMSEgenotype) compared to the RMSE computed on residual 5-HTT BPND values (RMSEresidual, i.e., ΔRMSE = 100*(RMSEresidual − RMSEgenotype)/RMSEresidual). Statistical significance (p-value) of model performance is expressed both uncorrected (punc) and corrected (pFWE) for the seven models estimated, using Bonferroni-Holm, which controls the family-wise type-I error rate60.
Results
Genotyping
Genotype distribution and allelic frequencies are depicted in Table 1. rs6265, rs1137070, rs7333412, rs6295 and rs1137070 were in Hardy–Weinberg equilibrium (all p > 0.16). MAOA is x-linked so the allele frequency for rs1137070 in men was compared with the frequency in women using a chi-squared test to determine whether there were significant differences between allele frequencies in males and females. We did not find a statistically significant difference in allele frequencies between males and females (P = 0.67). Assessment of Hardy–Weinberg for the 5-HTTLPR is not valid because this polymorphism was an inclusion criterion for some studies from which these data are derived, i.e., participants were not sampled independent of 5-HTTLPR genotype9,43.
Association analyses
The likelihood ratio showed that the LVM including genotype information was statistically significantly different from the LVM not including this information (p = 0.002), suggesting that genotype significantly contributed to the model.
The results from the LVM are depicted in Fig. 1. Consistent with previous observations, we observed that all regional 5-HTT BPND values loaded strongly on to 5-HTTLV (all p < 10–12). After adding genetic variants and covariates (sex, age, PET and MRI scanner) to the model a Wald test did not support the inclusion of additional paths to the model (pFDR > 0.05).
Within our final model, we found a statistically significant association between MAOA rs1137070 T-carriers (vs CC homozygotes) and 5-HTTLV (estimate: 0.07, 95% CI: [8.08 × 10–6, 0.14], p = 0.039). Across regions, rs1137070 T-carriers showed ~ 2–11% higher 5-HTT BPND compared to CC individuals, the largest differences were observed in caudate (~ 11%) and putamen (~ 9%) and the lowest in amygdala (~ 2%).
As expected based on our previous evaluation of this model, we observed: (1) higher caudate 5-HTT BPND in males vs females (estimate: 0.14, 95% CI: [0.06, 0.21], p = 3.88 × 10–4); (2) a negative association between age and 5-HTTLV (estimate: − 0.009, 95% CI: [− 0.015, − 0.0038], p < 0.001); (3) increased subcortical 5-HTT BPND in BDNF rs6265 met-carriers vs val/val individuals (estimate: − 0.06, 95% CI: [-0.04, -0.01], p = 0.07), corresponding to a 2–6% increase in 5-HTT BPND across subcortical areas. Conversely, an additional direct path from BDNF rs6265 to neocortex 5-HTT BPND effectively nullified the genetic effect on this brain region, which corresponds to the sum of indirect (BDNF rs6265—> 5-HTTLV—> neocortex BPND) and direct (BDNF rs6265—> neocortex BPND) effects. Although the estimate for statistical significance for rs6265 decreased in the presence of the three other genotypes, the effect size remained very similar, suggestive of independent main effects of BDNF rs6265 and MAOA rs1137070.
We did not observe evidence for an association between 5-HTTLPR (p = 0.35), HTR1A rs6295 (p = 0.33), nor HTR2A rs7333412 (p = 0.13) and 5-HTTLV.
Prediction analyses
Across the seven regions evaluated, we observed that the set of genetic variant features slightly improved prediction of caudate 5-HTT BPND compared to the model not including genetic information (caudate: RMSEresidual = 0.262, RMSEgenotype = 0.266, ΔRMSE = 1.6%, punc = 0.036) (Table 2). However, this effect, and the effect in all other regions was not statistically significant after correction for seven models (Table 2).
Discussion
We observed that MAOA rs1137070 T-carriers had higher 5-HTT availability compared to CC individuals. T-carriers showed higher 5-HTT availability in all the seven brain regions examined, with the highest binding in caudate (~ 11%) and the lowest in amygdala (~ 2%). Conversely, variants in the HTR1A and HTR2A genes were not associated with 5-HTT availability. Despite observing evidence for a statistically significant association, the genetic variants were not significantly informative for predicting brain 5-HTT available above chance. Taken together, our findings support that genetic variation in the MAOA contributes to variation in brain 5-HTT availability in the healthy adult human brain.
MAOA degrades monoamines in the brain, including serotonin, for which it has a preferential affinity compared to its other substrates61. MAOA knock-out rodents have increased extracellular serotonin levels and abnormal affective behavior30,62,63,64 as well as reduced 5-HTT expression63,65, suggesting that genetically altered MAOA signaling can affect regulation of 5-HT levels, which may in turn modulate 5-HTT levels e.g. via downregulation31. Similarly, inhibition of MAO activity by monoamine oxidase inhibitors increases extracellular serotonin levels62 and is associated with reduced 5-HTT BPND in rhesus monkey and rats66. Notably, reductions in 5-HTT BPND following an acute pharmacologically-induced serotonin increase may not only reflect a downregulation of 5-HTT but also increased serotonin levels competing for the radioligand rather than a change in 5-HTT gene expression67.
The rs1137070 T-allele has been previously associated with lower MAOA enzymatic activity compared to the C-allele both in human fibroblasts in vitro and in post-mortem brains35,68. Studies on clinical populations have reported mixed findings41,69,70. Although some studies suggest a link between the T-allele and increased MAOA mRNA expression in peripheral blood of patients with depression compared to healthy controls41, as well as increased vulnerability to depression41,69, other studies reported an association between the C-allele and impaired antidepressant treatment outcome in women70.
Conversely, we found that healthy human rs1137070 T-carriers, previously associated with low MAOA activity compared to C-carriers, had greater 5-HTT availability. In this case, putatively lower MAOA activity would correspond to greater amounts of intra- and extracellular serotonin. We can speculate that increased 5-HTT availability reflects increased 5-HTT levels, which might be a compensatory mechanism put in place to reuptake the excess serotonin and maintain extracellular serotonin levels constant. Nonetheless, findings from preclinical research point towards an effect opposite to what we observed, whereas the studies in humans provided mixed findings. Thus, the ambiguity provided by previous evidence in humans does not allow us to draw conclusions about the relationship that we detected between MAOA rs1137070 and 5-HTT BPND.
We did not observe evidence for an effect of HTR2A rs7333412 on 5-HTT availability. A previous study in patients with major depressive disorder, bipolar disorder, and healthy participants reported an effect on thalamus 5-HTT levels25. We did not replicate this effect in our study, as indicated by a comparison of Fig. 2 of their manuscript and our observed group differences. This is possibly because our study is based on a larger and more homogeneous cohort of healthy participants, whereas the previous study included patients with major depressive disorder and bipolar disorder as well as individuals with varying ethnic backgrounds. Taken together, our findings do not support that this HTR2A variant is associated with changes in 5-HTT availability in healthy adults.
Random forest model performance. The light blue dots represent the individual RMSE values obtained from resampling (for display purposes, the distribution of the RMSE values from resampling is derived from a model run with 100 instead of 10 resamples) of the model including genotype information (RMSEgenotype). The dark blue error plot displays the mean ± standard deviation of the distribution. The red hatched line indicates the 2.5% quantile of the average RMSE value derived from 10,000 permutations in the model that did not include genotype information (RMSEresidual). Dark blue dots below the red hatched line indicate that the model performed significantly better than chance upon adding genotype information.
Similarly, we did not observe evidence that HTR1A rs6295 is associated with 5-HTT availability, suggesting that whatever effects this polymorphism may have directly on the serotonin 1A receptor, those effects do not significantly modulate 5-HTT availability in healthy adults as measured with [11C]DASB PET.
Notably, we previously reported an association between BDNF rs6265 and 5-HTT BPND, such that met-carriers showed reduced binding in subcortical areas compared to val-homozygotes23. Although this effect was marginally above our threshold for statistical significance in the current model, the effect size was very similar, suggesting independent contributions of MAOA rs1137070 and BDNF rs6265 to 5-HTT availability in healthy humans. As we reported previously, 5-HTTLPR was not significantly associated with 5-HTT BPND in this cohort23,23.
Regarding our prediction model, we observed that using genotype information led to a marginal improvement in predicting caudate 5-HTT BPND vs not using genotype information, but this effect was not significant after correcting for multiple comparisons. In addition, we could not predict 5-HTT BPND in any other brain regions. This limited performance may be because we evaluated only five variants, whereas genetically induced variation in 5-HTT levels likely stems from many variants.
Previous studies underscore the limited extent to which candidate variants exert main effects on complex behavioral traits or related features of brain activity20,71. Direct measures of discrete neurobiological features, e.g., serotonin transporter protein levels, may however be more susceptible to genetic variants that modulate the relevant neurotransmission pathways72,73. Nevertheless, alternative genetic analysis strategies such as GWAS would undoubtedly provide a more comprehensive evaluation of genetic contributions to 5-HTT levels in the human brain. However, an exploratory GWAS requires either thousands of datasets or very large effect sizes (i.e., > 20% difference in 5-HTT BPND, Cohen’s d > 1) to establish statistical significance. GWAS-based polygenic risk scores, e.g., for psychiatric disorders or independent-dataset, hypothesis-generation via, e.g., expression quantitative trait loci (eQTL) databases may instead provide informative and statistically viable strategies for resolving genetic contributions to variation in brain serotonin neurotransmission measured with PET. Our cohort of 140 healthy participants stands as the largest single database of 5-HTT PET brain scans in the world. Future studies probing genetic contributions to brain serotonin-related PET scans would likely benefit from pooling data via, e.g., OpenNeuro (https://openneuropet.github.io/).
Although genetic variation is plausible and partially supported by our findings, environmental factors are also likely to contribute to 5-HTT levels67. A previous study based on the same cohort used for the present study reported no association between 5-HTT levels and daylight minutes or body mass index, contrarily to what reported by earlier studies based on smaller cohorts, part of which are included in our analyses42,55. In addition, we could not find any effect of variables reflecting lifestyle measures such as smoking, alcohol consumption, sleep and perceived stress on 5-HTT levels, suggesting that such environmental contributions in this cohort did not confound the observed genetic effects.
All participants self-identified with European ancestry. Self-reports of ancestry can be inaccurate74 and the lack of ethnic diversity in our sample limits the generalizability of our findings.
Notably, we exclude that the association between 5-HTT availability and the rs6265 and rs1137070 genotypes is due to a direct effect of genotype on BPND. BPND is proportional to the amount of target proteins available for binding (Bavail), i.e. 5-HTT, the affinity constant of the radioligand for its target (KD) and the free fraction of ligand in the non-displaceable tissue compartment (fND)75. We infer the observed genetic effects to be primarily related to change in Bavail, although we cannot rule out effects on KD or fND. Nonetheless, this seems unlikely because rs6265 and rs1137070 are proximal to SLC6A4, but a two scan study structure could more directly disentangle effects of Bavail and KD76.In conclusion, we report evidence for the association between MAOA rs1137070 genotype and brain 5-HTT availability. We did not observe evidence for an effect of HTR1A and HTR2A variants previously associated with brain serotonin markers, suggesting that their contribution may not be relevant to 5-HTT availability in the healthy adult human brain. Future studies considering additional genetic variants as well as environmental factors in larger datasets are critical for improving our understanding of the factors shaping serotonergic neurotransmission in health and disease.
Data availability
The R code employed for statistical analyses can be made available upon request to the corresponding author (patrick.fisher@nru.dk). Data can be made available upon reasonable request via this form (https://cimbi.dk/index.php/documents/category/3-cimbi-database) and with an appropriate data sharing agreement.
Code availability
The R codes used for the latent variable model and the random forest model described in this manuscript can be made available upon request.
References
Olivier, B. Serotonin: A never-ending story. Eur. J. Pharmacol. 753, 2–18 (2015).
Albert, P. R., Benkelfat, C. & Descarries, L. The neurobiology of depression-revisiting the serotonin hypothesis. I. Cellular and molecular mechanisms. Philos. Trans. R. Soc. B Biol. Sci. 367, 2378–2381 (2012).
Lesch, K.-P. Association of anxiety-related traits. Science 42, 2482–2488 (1996).
Baselmans, B. M. L., Yengo, L., van Rheenen, W. & Wray, N. R. Risk in relatives, heritability, SNP-based heritability, and genetic correlations in psychiatric disorders: A review. Biol. Psychiatr. 89, 11–19 (2021).
Kendall, K. M. et al. The genetic basis of major depression. Psychol. Med. 51, 2217–2230 (2021).
Perlman, K. et al. A systematic meta-review of predictors of antidepressant treatment outcome in major depressive disorder. J. Affect. Disord. 243, 503–515 (2019).
Frankle, W. G. et al. Comparative evaluation of serotonin transporter radioligands 11C-DASB and 11C-McN 5652 in healthy humans. J. Nucl. Med. 45, 682–694 (2004).
Houle, S., Ginovart, N., Hussey, D., Meyer, J. H. & Wilson, A. A. Imaging the serotonin transporter with positron emission tomography: Initial human studies with [11C]DAPP and [11C]DASB. Eur. J. Nucl. Med. 27, 1719–1722 (2000).
Mahon, M. B. et al. Seasonal difference in brain serotonin transporter binding predicts symptom severity in patients with seasonal affective disorder. Brain 139, 1605–1614 (2016).
Armand, S. et al. Brain serotonin transporter is associated with cognitive-affective biases in healthy individuals. Hum. Brain Mapp. 43, 4174–4184 (2022).
Kalbitzer, J. et al. The personality trait openness is related to cerebral 5-HTT levels. Neuroimage 45, 280–285 (2009).
Schneck, N. et al. Relationship of the serotonin transporter gene promoter polymorphism (5-HTTLPR) genotype and serotonin transporter binding to neural processing of negative emotional stimuli. J. Affect. Disord. 190, 494–498 (2016).
Rhodes, R. A. et al. Human 5-HT transporter availability predicts amygdala reactivity in vivo. J. Neurosci. 27, 9233–9237 (2007).
Miller, J. M. et al. Reported childhood abuse is associated with low serotonin transporter binding in vivo in major depressive disorder. Synapse 63, 565–573 (2009).
Marner, L. et al. Loss of serotonin 2A receptors exceeds loss of serotonergic projections in early Alzheimer’s disease: A combined [11C]DASB and [18F]altanserin-PET study. Neurobiol. Aging 33, 479–487 (2012).
Nielsen, K., Brask, D., Knudsen, G. M. & Aznar, S. Immunodetection of the serotonin transporter protein is a more valid marker for serotonergic fibers than serotonin. Synapse 59, 270–276 (2006).
Fisher, P. M. D. et al. Fluctuations in [11C]SB207145 PET binding associated with change in threat-related amygdala reactivity in humans. Neuropsychopharmacology 40, 1510–1518 (2015).
Fisher, P. M. et al. 5-HTTLPR status predictive of neocortical 5-HT 4 binding assessed with [11C]SB207145 PET in humans. Neuroimage 62, 130–136 (2012).
Pinborg, L. H. et al. The 5-HT2A receptor binding pattern in the human brain is strongly genetically determined. Neuroimage 40, 1175–1180 (2008).
Bogdan, R. et al. Imaging genetics and genomics in psychiatry: A critical review of progress and potential. Biol. Psychiatr. 82, 165–175 (2017).
Praschak-Rieder, N. et al. Novel 5-HTTLPR allele associates with higher serotonin transporter binding in putamen: A [11C] DASB positron emission tomography study. Biol. Psychiatr. 62, 327–331 (2007).
Reimold, M. et al. Central serotonin transporter levels are associated with stress hormone response and anxiety. Psychopharmacology (Berl). 213, 563–572 (2011).
Fisher, P. M. et al. BDNF val66met association with serotonin transporter binding in healthy humans. Transl. Psychiatr. 7, e1029–e1036 (2017).
Murthy, N. V. et al. Serotonin transporter polymorphisms (SLC6A4 insertion/deletion and rs25531) do not affect the availability of 5-HTT to [11C] DASB binding in the living human brain. Neuroimage 52, 50–54 (2010).
Laje, G. et al. Genetic variation in HTR2A influences serotonin transporter binding potential as measured using PET and [11C]DASB. Int. J. Neuropsychopharmacol. 13, 715–724 (2010).
Qesseveur, G. et al. Genetic dysfunction of serotonin 2A receptor hampers response to antidepressant drugs: A translational approach. Neuropharmacology 105, 142–153 (2016).
Goggi, J., Pullar, I. A., Carney, S. L. & Bradford, H. F. Modulation of neurotransmitter release induced by brain-derived neurotrophic factor in rat brain striatal slices in vitro. Brain Res. 941, 34–42 (2002).
Homberg, J. R., Molteni, R., Calabrese, F. & Riva, M. A. The serotonin-BDNF duo: Developmental implications for the vulnerability to psychopathology. Neurosci. Biobehav. Rev. 43, 35–47 (2014).
Pucadyil, T. J., Kalipatnapu, S. & Chattopadhyay, A. The serotonin1A receptor: A representative member of the serotonin receptor family. Cell. Mol. Neurobiol. 25, 553–580 (2005).
Bortolato, M. & Shih, J. C. Behavioral outcomes of monoamine oxidase deficiency: Preclinical and clinical evidence. Int. Rev. Neurobiol. 100, 13–42 (2011).
Jørgensen, T. N., Christensen, P. M. & Gether, U. Serotonin-induced down-regulation of cell surface serotonin transporter. Neurochem. Int. 73, 107–112 (2014).
Lemonde, S. et al. Impaired repression at a 5-hydroxytryptamine 1A receptor gene polymorphism associated with major depression and suicide. J. Neurosci. 23, 8788–8799 (2003).
Kautzky, A. et al. The influence of the rs6295 gene polymorphism on serotonin-1A receptor distribution investigated with PET in patients with major depression applying machine learning. Transl. Psychiatr. 7, e1150 (2017).
Pernhorst, K. et al. Rs6295 promoter variants of the serotonin type 1A receptor are differentially activated by c-Jun in vitro and correlate to transcript levels in human epileptic brain tissue. Brain Res. 1499, 136–144 (2013).
Hotamisligil, G. S. & Breakefield, X. O. Human monoamine oxidase a gene determines levels of enzyme activity. Am. J. Hum. Genet. 49, 383–392 (1991).
Bi, Y. et al. DRDInfluence and interaction of genetic, cognitive, neuroendocrine and personalistic markers to antidepressant response in Chinese patients with major depression. Prog. Neuro Psychopharmacol. Biol. Psychiatr. 104, 110036 (2021).
Xu, Z. et al. Influence and interaction of genetic polymorphisms in catecholamine neurotransmitter systems and early life stress on antidepressant drug response. J. Affect. Disord. 133, 165–173 (2011).
Knudsen, G. M. et al. The center for integrated molecular brain imaging (cimbi) database. Neuroimage 124, 1213–1219 (2016).
Drago, A., Ronchi, D. D. & Serretti, A. 5-HT1A gene variants and psychiatric disorders: A review of current literature and selection of SNPs for future studies. Int. J. Neuropsychopharmacol. 11, 701–721 (2008).
Wu, C. S. & D.E. A_common_C_1018G_polymorphism_in_the_human_5_HT1A.10.pdf. Psychiatr. Genet. 9, 105–106 (1999).
Zhang, J. et al. A cis-phase interaction study of genetic variants within the MAOA gene in major depressive disorder. Biol. Psychiatr. 68, 795–800 (2010).
Erritzoe, D. et al. Cerebral serotonin transporter binding is inversely related to body mass index. Neuroimage 52, 284–289 (2010).
Frokjaer, V. G. et al. Role of serotonin transporter changes in depressive responses to sex-steroid hormone manipulation: A positron emission tomography study. Biol. Psychiatr. 78, 534–543 (2015).
Frokjaer, V. G. et al. Prefrontal serotonin transporter availability is positively associated with the cortisol awakening response. Eur. Neuropsychopharmacol. 23, 285–294 (2013).
Kalbitzer, J. et al. Seasonal changes in brain serotonin transporter binding in short serotonin transporter linked polymorphic region-allele carriers but not in long-allele homozygotes. Biol. Psychiatr. 67, 1033–1039 (2010).
Klein, A. B. et al. Cerebral 5-HT 2A receptor and serotonin transporter binding in humans are not affected by the val66met BDNF polymorphism status or blood BDNF levels. J. Cereb. Blood Flow Metab. 30, 1890–1896 (2010).
Nørgaard, M. et al. Different preprocessing strategies lead to different conclusions: A [11C]DASB-PET reproducibility study. J. Cereb. Blood Flow Metab. 40, 1902–1911 (2020).
Madsen, M. K. et al. Threat-related amygdala functional connectivity is associated with 5-HTTLPR genotype and neuroticism. Soc. Cogn. Affect. Neurosci. 11, 140–149 (2015).
Fisher, P. M. et al. BDNF Val66met and 5-HTTLPR polymorphisms predict a human in vivo marker for brain serotonin levels. Hum. Brain Mapp. 36, 313–323 (2015).
Sureau, F. C. et al. Impact of image-space resolution modeling for studies with the high-resolution research tomograph. J. Nucl. Med. 49, 1000–1008 (2008).
Comtat, C. et al. Image based resolution modeling for the HRRT OSEM reconstructions software. IEEE Nucl. Sci. Symp. Conf. Rec. https://doi.org/10.1109/NSSMIC.2008.4774188 (2008).
Woods, R. P., Cherry, S. R. & Mazziotta, J. C. Rapid automated algorithm for aligning and reslicing PET images. J. Comput. Assist. Tomograph. 16(4), 620–633. https://doi.org/10.1097/00004728-199207000-00024 (1992).
Svarer, C. et al. MR-based automatic delineation of volumes of interest in human brain PET images using probability maps. Neuroimage 24, 969–979 (2005).
Ichise, M. et al. Linearized reference tissue parametric imaging methods: Application to [11C]DASB positron emission tomography studies of the serotonin transporter in human brain. J. Cereb. Blood Flow Metab. 23, 1096–1112 (2003).
Erritzoe, D. et al. A nonlinear relationship between cerebral serotonin transporter and 5HT-2A receptor binding: An in vivo molecular imaging study in humans. J. Neurosci. 30, 3391–3397 (2010).
Carpenter, J. S. & Andrykowski, M. A. Psychometric evaluation of the pittsburgh sleep quality index. J. Psychosom. Res. 45, 5–13 (1998).
Roberti, J. W., Harrington, L. N. & Storch, E. A. Further psychometric support for the 10-item version of the perceived stress scale. J. Coll. Couns. 9, 135–147 (2006).
Holst, K. K. & Budtz-Jørgensen, E. Linear latent variable models: The lava-package. Comput. Stat. 28, 1385–1452 (2013).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Holm, S. Board of the foundation of the scandinavian journal of statistics a simple sequentially rejective multiple test procedure a simple sequentially rejective multiple test procedure. Source Scand. J. Stat. Scand J Stat. 6, 65–70 (1979).
Bortolato, M., Chen, K. & Shih, J. C. The Degradation of Serotonin: Role of MAO. Handbook Behav. Neurosci. 21, 203–218 (2010).
Cases, O. et al. Aggressive behavior and altered amounts of brain serotonin and norepinephrine in mice lacking MAOA. Science 268, 1763–1766 (1995).
Evrard, A. et al. Altered regulation of the 5-HT system in the brain of MAO-A knock-out mice. Eur. J. Neurosci. 15, 841–851 (2002).
Scott, A. L., Bortolato, M., Chen, K. & Shih, J. C. Novel monoamine oxidase a knock out mice with human-like spontaneous mutation. Neuroreport 19, 739–743 (2008).
Godar, S. C. et al. The aggression and behavioral abnormalities associated with monoamine oxidase A deficiency are rescued by acute inhibition of serotonin reuptake. J. Psychiatr. Res. 56, 1–9 (2014).
Lundquist2007–Effect on 11C DASB binding after tranylcypromine‐induced increase in serotonin concentration.pdf.
Spies, M., Knudsen, G. M., Lanzenberger, R. & Kasper, S. The serotonin transporter in psychiatric disorders: Insights from PET imaging. Lancet Psychiatr. 2, 743–755 (2015).
Pinsonneault, J. K., Papp, A. C. & Sadée, W. Allelic mRNA expression of X-linked monoamine oxidase a (MAOA) in human brain: Dissection of epigenetic and genetic factors. Hum. Mol. Genet. 15, 2636–2649 (2006).
Huang, S. Y. et al. Association of monoamine oxidase A (MAOA) polymorphisms and clinical subgroups of major depressive disorders in the Han Chinese population. World J. Biol. Psychiatr. 10, 544–551 (2009).
Bi, Y. et al. DRDInfluence and interaction of genetic, cognitive, neuroendocrine and personalistic markers to antidepressant response in Chinese patients with major depression. Prog. Neuro Psychopharmacol. Biol. Psychiatr. 104, 110036 (2021).
Flint, J. & Munafò, M. R. Candidate and non-candidate genes in behavior genetics. Curr. Opin. Neurobiol. 23, 57–61 (2013).
Jennings, K. A. et al. Genetic variation in 5-hydroxytryptamine transporter expression causes adaptive changes in 5-HT4 receptor levels. Int. J. Neuropsychopharmacol. 15, 1099–1107 (2012).
Licht, C. L. et al. Changes in 5-HT4 receptor and 5-HT transporter binding in olfactory bulbectomized and glucocorticoid receptor heterozygous mice. Neurochem. Int. 56, 603–610 (2010).
Burnett, M. S. et al. Reliability of self-reported ancestry among siblings: Implications for genetic association studies. Am. J. Epidemiol. 163, 486–492 (2006).
Innis, R. B. et al. Consensus nomenclature for in vivo imaging of reversibly binding radioligands. J. Cereb. Blood Flow Metab. 27, 1533–1539 (2007).
Hirvonen, M. M. et al. C957T polymorphism of dopamine D2 receptor gene affects striatal DRD2 in vivo availability by changing the receptor affinity. Synapse 63, 907–912 (2009).
Acknowledgements
This study was supported by the Lundbeck Foundation and the Danish National Research Council, which funded the Center for Integrated Molecular Brain Imaging (10.1016/j.neuroimage.2015.04.025), the European Union’s Horizon 2020 research and innovation programme under Grant Agreement No. 953327 (Serotonin & Beyond), the Research Council of Rigshospitalet (Grant ID A6594), the Augustinusfonden (Grant ID 19‐0489) and the Lundbeck Foundation (Grant R279-2018-1145: BrainDrugs). We acknowledge the John and Birthe Meyer Foundation for donating the PET scanner and the Cyclotron. We thank G. Thomsen, B. Dall, L. Freyr, S. V. Larsen and A. Dyssegaard for their help with the data collection with both the PET and MR scanners.
Author information
Authors and Affiliations
Contributions
S.E.P.B.: Data analysis, writing of the original draft, figures design and execution, manuscript editing and reviewing. A.N.: Data analysis, manuscript editing and reviewing. S.S.A.: Data analysis, manuscript editing and reviewing. M.S.: Conceptualization, funding, manuscript editing and reviewing. B.O.: Data analysis, manuscript editing and reviewing. P.S.J.: Data management, manuscript editing and reviewing. G.M.K.: Funding, manuscript editing and reviewing, supervision. V.G.F.: Data acquisition, manuscript editing and reviewing, supervision. P.M.F.: Conceptualization, funding, data analysis, data acquisition, manuscript editing and reviewing, supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bruzzone, S.E.P., Nasser, A., Aripaka, S.S. et al. Genetic contributions to brain serotonin transporter levels in healthy adults. Sci Rep 13, 16426 (2023). https://doi.org/10.1038/s41598-023-43690-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43690-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.