Abstract
Ligilactobacillus agilis is a motile lactic acid bacterium found in the gastrointestinal tracts of animals. The findings of our previous study suggest that the motility of L. agilis BKN88 enables gut colonization in murine models. However, the chemotactic abilities of motile lactobacilli remain unknown. This study aimed to identify the gut-derived chemoeffectors and their corresponding chemoreceptors in L. agilis BKN88. Chemotaxis assays with chemotactic and non-chemotactic (ΔcheA) L. agilis strains revealed that low pH, organic acids, and bile salts served as repellents. L. agilis BKN88 was more sensitive to bile and acid than the gut-derived non-motile lactobacilli, implying that L. agilis might utilize motility and chemotaxis instead of exhibiting stress tolerance/resistance. L. agilis BKN88 contains five putative chemoreceptor genes (mcp1–mcp5). Chemotaxis assays using a series of chemoreceptor mutants revealed that each of the five chemoreceptors could sense multiple chemoeffectors and that these chemoreceptors were functionally redundant. Mcp2 and Mcp3 sensed all tested chemoeffectors. This study provides further insights into the interactions between chemoreceptors and ligands of motile lactobacilli and the unique ecological and evolutionary features of motile lactobacilli, which may be distinct from those of non-motile lactobacilli.
Similar content being viewed by others
Introduction
Bacterial chemotaxis is the movement of motile bacteria toward favorable chemicals or away from unfavorable chemicals and is mediated by chemotactic signaling pathways1,2,3. The molecular mechanisms underlying chemotactic signaling have been intensely studied in Escherichia coli and Salmonella enterica serovar Typhimurium4,5. Chemotaxis is triggered by the binding of ligands (chemoeffectors) to chemoreceptors called methyl-accepting chemotaxis proteins (MCPs) and transducer-like proteins (Tlps). Subsequently, chemoreceptors transduce signals through a series of cytoplasmic chemotactic proteins and modulate flagellar motor rotation6,7. This chemotactic signaling pathway is relatively conserved across bacteria, whereas chemoeffectors and chemoreceptors vary among bacterial species/strains.
In the last few decades, several chemoeffectors have been identified, and various chemoreceptors have been functionally characterized. For example, urea8,9,10,11, lactate12, various amino acids8,13, and mucin14 serve as chemoattractants for Helicobacter pylori, whereas low pH11,15,16, reactive oxygen species17 and autoinducer-218 serve as chemorepellents. Four chemoreceptors (TlpA, TlpB, TlpC, and TlpD) are mediated by these chemotactic responses, which are required to colonize the gastric mucosa efficiently19,20. Ten chemoreceptors have been identified in the gram-positive model bacterium Bacillus subtilis, 10 chemoreceptors have been found, and some have been characterized. McpB and McpC mediate chemotaxis toward amino acids21,22,23, whereas McpA mediates chemoattraction toward glucose and α-methylglucoside22. Although most studies have examined the chemoeffectors and chemoreceptors of pathogenic bacteria and some bacterial species6,24, limited studies have focused on beneficial gut microbes such as lactic acid bacteria.
Some lactobacilli, considered beneficial commensals, are found in the gastrointestinal tract of humans and animals. Most lactobacilli are non-motile and do not possess motility genes, although a small proportion of lactobacilli are motile25,26. The ecology of non-motile lactobacilli has been relatively well studied27,28,29,30, whereas that of motile lactobacilli is poorly characterized. Previously, we demonstrated that Ligilactobacillus agilis (previously named Lactobacillus agilis) BKN88, a motile strain isolated from chicken, utilizes motility for gut colonization in murine models31. Thus, the chemotaxis of motile lactobacilli may be involved in gut colonization. However, this has not been demonstrated owing to a lack of information on the chemoeffectors and chemoreceptors of motile lactobacilli. This study aimed to identify gut-derived chemoeffectors and their corresponding chemoreceptors in L. agilis BKN88 to gain insights into its ecology and evolution.
Results
L. agilis BKN88 has only one histidine kinase (CheA) that plays a central role in chemotactic signaling. Thus, a motile but non-chemotactic derivative of L. agilis BKN88 was constructed by deleting cheA and was used as a negative control in the chemotaxis assay. Deletion of cheA was confirmed by polymerase chain reaction (PCR). PCR analysis of the cheA deletion mutant (ΔcheA) revealed an amplicon with the expected size (Supplementary Fig. S1a, b). The swimming behavior of the ΔcheA mutant was examined using optical microscopy. As shown in Supplementary Fig. S1c, the ΔcheA mutant exhibited motility with a higher frequency of tumbling than the wild-type (WT) strain, similar to B. subtilis32. Preliminary analysis of the chemotaxis of chemotactic and non-chemotactic (ΔcheA) L. agilis strains was performed using the capillary and microscopic agar-drop assays31,33. Among these assays, the microscopic agar-drop assay was chosen for further studies on chemotaxis in L. agilis owing to its high reproducibility (Supplementary Fig. S2).
Chemotactic responses of L. agilis to pH
L. agilis may respond to acidic environments in the presence of stomach acids or organic acids produced by the gut microbes to survive in the gastrointestinal tract. To examine the chemotaxis of L. agilis to low pH, chemotactic or non-chemotactic L. agilis strains were subjected to a microscopic agar-drop assay. Figure 1a, b shows that the non-chemotactic mutant (ΔcheA) did not respond significantly to the agar with a pH adjusted to 3.0 with HCl. In contrast, the chemotactic L. agilis (WT) escaped from the acidic agar. pH taxis have also been observed in some pathogenic bacteria, such as E. coli, S. enterica, and H. pylori11,15,34,35,36,37,38,39. E. coli exhibits repellent responses to acidic and basic pH conditions, resulting in its migration to neutral environments 39. H. pylori escapes acidic environments via attractant responses to basic pH and repellent responses to acidic pH 40. To examine these bidirectional responses of L. agilis to pH, the chemotactic responses of L. agilis to the agar with pH values ranging from 3.0 to 12.0 were analyzed. Chemotactic L. agilis responded only to acidic agar, whose pH was adjusted to 3.0 or 5.0 (Fig. 1c). This repellent response was stronger at lower pH (Fig. 1c). Several gut microbes, including lactobacilli, exhibit tolerance to acid41,42. To investigate the acid tolerance of L. agilis, L. agilis BKN88 and three strains of non-motile lactobacilli isolated from animals and humans were incubated under acidic conditions (pH 3.0), and the number of viable cells was counted over time. The sensitivity of L. agilis to acid was higher than that of the gut-derived non-motile lactobacilli (Fig. 1d). Some acid tolerance systems have been identified in non-motile lactobacilli 42. In L. agilis, only the F1F0-ATPase system, which plays a vital role in pH homeostasis in bacterial cells 43, was found among other systems (Supplementary Fig. S3). However, other systems, such as the malolactic fermentation (MLF) pathway (mleRPS genes), glutamate decarboxylase (GAD) system (gadBC genes), arginine deiminase (ADI) system (arcABC genes), and urease system (ureABC genes) were not detected in L. agilis.
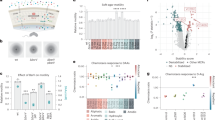
Chemotactic responses of Ligilactobacillus agilis to pH. (a, b) Chemotaxis toward acidic pH was observed in non-chemotactic (ΔcheA) and chemotactic [wild-type (WT)] L. agilis strains using the microscopic agar-drop assay. Details of the assay are described in Supplementary Fig. S2. (a) Representative microscopic images of the L. agilis ΔcheA or WT cells near the agar drop with pH adjusted to 3.0 at 0 or 5 min. (b) The relative number of cells represents the ratio of cells near the agar drop at 5 min to that at 0 min. Values are represented as the mean + standard error (SE) (n = 4). Significant difference is indicated using an asterisk (*P < 0.01; Student’s t-test). (c) Time-course chemotactic responses of L. agilis BKN88 (WT) to pH values of 3.0, 5.0, 7.0, and 12.0. The relative number of cells represents the ratio of cells near the agar drop at each time point to that at 0 min. Values are represented as the mean ± SE (n = 6). Significant difference between the pH 7.0 and other pH values is indicated using an asterisk (*P < 0.05; Dunnett’s test). (d) Susceptibility of L. agilis to acid. Colony-forming units (CFUs) of L. agilis BKN88, L. johnsonii NRIC 0220 T, L. reuteri PTL371, and L. acidophilus NCFM incubated in MRS with pH adjusted to 3.0 at 37 °C were counted once every 30 min. Values are represented as mean ± standard deviation (SD) (n = 3).
Chemotactic responses of L. agilis to bile
Bile exerts antimicrobial effects and is a major stressor for gut microbes. Several pathogens, including H. pylori, Campylobacter jejuni, and Vibrio cholerae, exhibit chemotaxis toward bile and its components8,44,45,46. Bile and some conjugated bile acids serve as repellents for H. pylori8, whereas bile and taurocholic acids serve as attractants for V. cholerae 46. To investigate bile chemotaxis in L. agilis, chemotactic and non-chemotactic L. agilis strains were subjected to a microscopic agar-drop assay. In contrast, to the non-chemotactic mutant, chemotactic L. agilis moved away from the bile salts (Fig. 2a, b) and exhibited dose-dependent repellent responses to bile salts (Fig. 2c). To identify the constituents of bile that elicit a repellent response, chemotaxis of L. agilis to major constituents of bile, such as sodium deoxycholate (SDC), sodium cholate (SC), and sodium taurocholate (STC), was examined. Because L. agilis exhibits repellent responses to low pH, the pH of the bile constituents tested in the microscopic agar-drop assay was adjusted to 7.0. Figure 2d–f shows that all tested bile constituents served as repellents. Among the bile constituents, the strongest and weakest repellent responses were elicited by SDC and STC, respectively. The repellent response to SC was between SDC and STC (Fig. 2d–f). Because these bile constituents are sodium salts, chemotaxis to sodium chloride was investigated. L. agilis did not exhibit chemotaxis toward NaCl (data not shown). The bile tolerance of L. agilis BKN88 and three non-motile lactobacillus strains was also examined. Although bile did not significantly inhibit the growth of non-motile lactobacilli, bile salt concentration-dependently decreased the viability of L. agilis BKN88 (Fig. 2g). The growth of L. agilis BKN88 was inhibited in the presence of high concentrations of bile salts [1.0–2.0% (w/v)]. The mechanisms involved in bile tolerance in gut-derived non-motile lactobacilli are mainly mediated by bile efflux, bile salt hydrolysis, and changes in cell membrane/wall architecture and composition29,47. Next, genes involved in bile tolerance in L. agilis BKN88 were examined. As shown in Supplementary Table S1, several putative genes encoding bile efflux pumps and bile salt hydrolases were identified in the genome of L. agilis BKN88.
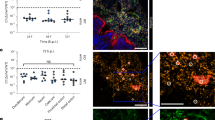
Chemotactic responses of L. agilis to bile. (a–b) Chemotaxis toward bile was observed in non-chemotactic (ΔcheA) and chemotactic (WT) L. agilis strains using the microscopic agar-drop assay. (a) Representative microscopic images of the L. agilis ΔcheA or WT cells near the agar drop containing 0.1% (w/v) bile salt at 0 or 5 min. (b) The relative number of cells represents the ratio of cells near the agar drop at 5 min to that at 0 min. Values are represented as the mean + SE (n = 4). Significant difference is indicated using an asterisk (*P < 0.01; Student’s t-test). (c–f) Time-course chemotactic responses of L. agilis BKN88 (WT) to various concentrations of bile salt (c) and its constituents [sodium deoxycholate (SDC), sodium cholate (SC), and sodium taurocholate (STC)]. The relative number of cells represents the ratio of cells near the agar drop at each time point to that at 0 min. An agar drop without test chemicals was used as a control. Values are represented as the mean ± SE (n = 6). Significant difference between the values of the control group and groups involving various concentrations of test chemicals is indicated using an asterisk (*P < 0.05; Dunnett’s test). (g) Susceptibility of L. agilis BKN88 to bile salt. CFUs of L. agilis BKN88, L. johnsonii NRIC 0220 T, L. reuteri PTL371, and L. acidophilus NCFM cultured on an MRS plate containing 0–2.0% (w/v) bile salts were counted. Values are represented as the mean ± SD (n = 3).
Chemotactic responses of L. agilis to organic acids produced by gut microbes
Some organic acids, such as lactic acid, butyric acid, and acetic acid, are the main metabolites produced by gut microbes that compete with L. agilis. Therefore, in this study, we examined the chemotaxis of L. agilis in response to these organic acids. As L. agilis exhibits repellent responses to low pH, organic acid salts with a pH value adjusted to 7.0 were used to perform the microscopic agar-drop assay. The non-chemotactic mutant did not exhibit significant responses to organic acid salts, whereas the chemotactic strain showed repellent responses (Fig. 3a–f). Furthermore, the chemotactic response of L. agilis to various concentrations of organic acid salts was examined. The repellent responses to the organic acid salts were dose-dependent (Fig. 3g–i). A repellent response to acetate was only observed at high concentrations (> 50 mM) (Fig. 3i). It has been reported that most chemicals, such as various sugars and inorganic compounds, at high concentrations (> 1 M) serve as repellents for E. coli, which may be due to high osmolarity48. In contrast, L. agilis did not exhibit chemotactic responses to 500 mM d-galactose, 500 mM l-fucose, or 100 mM NaCl (data not shown) but was repelled by 25 mM lactate, 25 mM butyrate, and 50 mM acetate. Therefore, the repellent response of L. agilis to organic acids may not be attributable to high osmolarity.
Chemotactic responses of L. agilis to organic acids produced by gut microbes. (a–f) Chemotaxis toward 100 mM lactate (a, d), 100 mM butyrate (b, e), and 500 mM acetate (c, f) were observed in non-chemotactic (ΔcheA) and chemotactic (WT) L. agilis strains using the microscopic agar-drop assay. (a–c) Representative microscopic images of the L. agilis ΔcheA or WT cells near the agar drop containing organic acid salts at 0 or 5 min. (d–f) The relative number of cells represents the ratio of cells near the agar drop at 5 min to that at 0 min. Values are represented as the mean + SE (n = 4). Significant difference is indicated using an asterisk (*P < 0.01; Student’s t-test). (g–i) Time-course chemotactic responses of L. agilis BKN88 (WT) to various concentrations of organic acid salts. The relative number of cells represents the ratio of cells near the agar drop at each time point to that at 0 min. An agar drop without test chemicals was used as a control. Values are represented as the mean ± SE (n = 6). Significant difference between the values of the control group and groups involving various concentrations of organic acid salts is indicated using an asterisk (*P < 0.05; Dunnett’s test).
Chemotaxis-related genes and the features in L. agilis BKN88
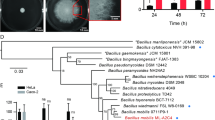
As illustrated in Fig. 4a, the draft genome sequence analysis of L. agilis BKN88 revealed the presence of only one set of chemotaxis-related genes, except for two cheW genes, in a single gene cluster (motility operon) and five putative MCP-encoding genes (mcp1–mcp5). mcp1 and mcp2 are encoded in the motility operon, whereas mcp3, mcp4, and mcp5 are encoded elsewhere in the genome. Chemoreceptors sense ligands directly or indirectly via interactions with periplasmic ligand-binding proteins (LBPs)49. Genes encoding LBPs are often located adjacent to chemoreceptor-encoding genes. In the genome of L. agilis BKN88, the LBP-encoding gene, which encodes the sugar-binding protein, was located in the vicinity of mcp5 but not in the vicinity of mcp1, mcp2, mcp3, and mcp4 (Figs. 4a and S4). Domain architecture analysis of the five chemoreceptors revealed that Mcp2, Mcp3, Mcp4, and Mcp5 have the following typical MCP domains: two transmembrane regions, a variable periplasmic ligand-binding domain (LBD), and a conserved cytoplasmic signaling domain. The LBDs of these four chemoreceptors contain a dual calcium channel and chemotaxis receptor (dCache_1) domain, which is the predominant extracellular sensory domain in bacteria50. In contrast, Mcp1 has two transmembrane regions and a cytoplasmic signaling domain but no identifiable LBD (Fig. 4b). The primary structures of the sensory domains (LBDs) are highly variable because they have evolved to recognize specific ligands. To gain insight into the correlation between chemoreceptors and their ligands, the amino acid sequences of the LBDs of Mcp2 (255 amino acids; residues 34–288), Mcp3 (154 amino acids; residues 39–192), Mcp4 (249 amino acids; residues 40–288), and Mcp5 (246 amino acids; residues 33–278) were analyzed using BLASTP with default settings and a cutoff of 45% identity and 90% query coverage. Although LBDs exhibiting high similarity (> 45% identity) to the LBDs of Mcp2 or Mcp3 were not detected, the LBD of Mcp4 was highly similar to that of the chemoreceptor from Ligilactobacillus ruminis (65% identity), which is the most prevalent Lactobacillus species in the human gut51. The LBD of Mcp5 shared sequence similarity with that of chemoreceptors from several butyrate-producing bacteria found in the human gut [belonging to the genera Eubacterium, Lachnospira, and Roseburia (approximately 45–50% identity)], as well as with that of L. ruminis (65% identity). The genes encoding sugar-binding proteins were also located near the genes encoding these chemoreceptors in the butyrate-producing bacteria and L. ruminis, similar to mcp5 in L. agilis (Supplementary Fig. S4). The similarity between the LBDs of the four L. agilis chemoreceptors, except Mcp1, was less than 30%. Reverse transcription PCR (RT-PCR) analysis, performed to validate the transcription of chemoreceptor-encoding genes in L. agilis BKN88, revealed that all chemoreceptor genes were expressed during the exponential phase (Fig. 4c).
Chemoreceptors of L. agilis BKN88. (a) A genetic map of chemotaxis-related genes in the L. agilis BKN88 genome. (b) Domain architectures of the five chemoreceptors were predicted using InterPro81 and CDvist82. Transmembrane domains are shown as gray rectangles. MCP signal, cytoplasmic signaling domain; dCache_1 (a dual calcium channel and chemotaxis receptor), one of the ligand-binding domains; HAMP (histidine kinases, adenylyl cyclases, methyl binding proteins, and phosphatases), linker domain. (c) Expression of the MCP-encoding genes was evaluated using RT-PCR. PCR was performed with chromosomal DNA isolated from bacterial cells (upper). RT-PCR was performed with total RNA isolated from bacterial cells (lower).
Chemotactic responses of L. agilis mutants lacking one or all of the five MCP genes
To identify the chemoreceptors responsible for sensing chemoeffectors in L. agilis, L. agilis mutants lacking one or all of the five putative MCP-encoding genes were constructed. PCR analysis using primers specific to the sequences flanking each MCP region revealed the deletion of MCP-encoding genes in L. agilis mutants (Supplementary Fig. S5a). The motility of MCP deletion mutants was also assessed in MRS soft-agar culture. In contrast, to the non-motile L. agilis BKN134 strain, all the MCP deletion mutants exhibited motility (Supplementary Fig. S5b). Next, the chemotactic responses of the MCP deletion mutants were determined using a microscopic agar drop assay. The L. agilis mutant deficient in all the five MCP-encoding genes (Δmcp1-5) did not exhibit chemotactic responses to all tested chemoeffectors, while the L. agilis mutants lacking one of the five MCP-encoding genes showed repellent responses to chemoeffectors at the same levels as the WT L. agilis, except for the constituents of bile (Fig. 5a). These results indicate that multiple MCPs (Mcp1–Mcp5) may be involved in the chemotactic responses to low pH and organic acid salts in a complementary manner. Chemotactic responses to STC, SDC, and SC were attenuated in the Δmcp1 and Δmcp2 mutants. Meanwhile, chemotactic responses to SC were attenuated in the Δmcp5 mutants (Fig. 5a). These results suggest that Mcp1 and Mcp2 mediate chemotactic responses to STC, SDC, and SC and that Mcp5 is involved in chemotactic responses to SC.
Chemotactic responses of the putative methyl-accepting chemotaxis protein (MCP) deletion mutants. (a–b) Chemotaxis toward pH 3.0, 100 mM lactate, 100 mM butyrate, 500 mM acetate, 20 mM sodium taurocholate (STC), 10 mM sodium deoxycholate (SDC), and 10 mM sodium cholate (SC) was observed in L. agilis strains lacking one or all of the MCP-encoding genes (a) and L. agilis strains expressing individual MCP-encoding genes (b). The relative number of cells represents the ratio of cells near the agar drop at 5 min to that at 0 min. Values are represented as the mean + SE (n = 6). Different superscripts indicate significant differences (P < 0.05; Tukey’s multiple comparison test). The dotted lines represent the values of L. agilis WT (a) or L. agilis Δmcp1-5 (b).
Chemotactic responses of L. agilis mutants expressing single MCP genes
Next, L. agilis mutants expressing a single chemoreceptor were constructed. The chemotaxis of these mutants was examined to identify the chemoreceptors that sense chemoeffectors. L. agilis mutants expressing each of the five MCP-encoding genes were first constructed by introducing a plasmid carrying one of the MCP-encoding genes into the Δmcp1-5 mutant, which lacks all five MCP-encoding genes. However, the constructed mutants exhibit decreased motility, which hinders the observation of their chemotaxis. Thus, L. agilis mutants expressing a single MCP-encoding gene with four other MCPs deleted were constructed (Supplementary Fig. S6). As shown in Fig. 5b, the microscopic agar-drop assay with these mutants revealed that the mutant with Mcp1 as the sole chemoreceptor was defective in chemotaxis to low pH, and organic acid salts could swim away from the three tested bile constituents. This was consistent with the results of the chemotaxis assay performed using the Δmcp1 mutant (Fig. 5a). L. agilis mutants expressing only Mcp2 or Mcp3 exhibited repellent responses to all tested chemoeffectors. Among the mutants, the Mcp2-expressing strain exhibited the strongest repellent response to all chemoeffectors. This suggests that Mcp2 is a major chemoreceptor of gut-derived substances in L. agilis. Mutant strains expressing Mcp4 or Mcp5 exhibited chemotaxis to all chemoeffectors except for lactate or organic acid salts, respectively. These data suggest that each of the five MCPs responds to some or all of the chemoeffectors and that these MCPs are redundantly involved in chemotaxis to each chemoeffectors (Fig. 6).
Discussion
We previously reported that L. agilis BKN88 colonized the gut in murine models via motility behavior31. This suggests that chemotaxis is also involved in gut colonization by L. agilis. The role of chemotaxis in host colonization has been demonstrated in some pathogenic bacteria24. However, few studies have examined the role of chemotaxis in the gut colonization of beneficial gut microbes, such as lactobacilli, owing to a lack of information on their chemoeffectors and chemoreceptors. In this study, we performed a chemotaxis assay using chemotactic and non-chemotactic L. agilis strains and a series of chemoreceptor mutants to identify chemoeffectors and their corresponding chemoreceptors in L. agilis BKN88.
Gut microbes sense and respond to gastrointestinal stress conditions, such as acidic and bile stress, to survive under harsh conditions in the gastrointestinal tract. Some pathogens, including E. coli, S. enterica, H. pylori, and C. jejuni, escape low pH and bile via motility and chemotaxis 8,11,15,34,35,36,37,38,39,44,45. Chemotaxis has been reported to be involved in host colonization and cell invasion in these bacteria 24,52,53,54,55,56,57,58,59. This study demonstrated that L. agilis BKN88 swims away from low pH conditions and bile. This suggests that L. agilis, a commensal microbe, survives and persists in the gastrointestinal tract via repellent responses to acids and bile. L. agilis is more sensitive to acid and bile than gut-derived non-motile lactobacilli. Therefore, the repellent response to these stress conditions may be an important factor for the survival of L. agilis in the gut. These findings suggest that stress tolerance is not essential for L. agilis, which can escape acids and bile via motility and chemotaxis. Several gut microbes, including lactobacilli, exhibit acid and bile tolerance41,47, whereas L. agilis likely utilizes motility and chemotaxis instead of stress tolerance/resistance. This also hypothesizes that lactobacilli might have chosen one of the following two evolutionary strategies: establishing a sedentary lifestyle that does not require motility or a nomadic lifestyle based on motility and chemotaxis.
L. agilis BKN88 exhibited repellent responses to organic acids, lactate, acetate, and butyrate. Organic acids are the main metabolites produced by gut microbes and exert toxic effects on bacterial cells at high concentrations. In addition, high acid concentrations can lead to decreased pH in the intestine, resulting in bacterial growth inhibition60,61. Hence, the chemotactic response to organic acids is important for escape from toxic acids and a low pH environment and may confer survival advantages in the intestine. Gut microbes that produce organic acids compete with L. agilis. Therefore, another possible benefit of chemotactic responses to organic acids may be the ability to escape competitive gut microbiota, resulting in their migration to a specific niche in the gastrointestinal tract. Previously, we demonstrated that the effect of the presence or absence of motility on gut colonization by L. agilis in specific pathogen-free mice was higher than that in germ-free mice31. This finding suggests that motility is beneficial when L. agilis is surrounded by other gut microbes. In this case, the benefits of motility may be attributed to the repellent responses to organic acids and their producers. Thus, the repellent response to organic acids may play an important role in the survival and competition of L. agilis in the gut.
L. agilis BKN88 contains five putative chemoreceptor genes (mcp1–mcp5). RT-PCR analysis confirmed the expression of these genes in L. agilis BKN88. A chemotaxis assay with chemoreceptor deletion mutants revealed that the single MCP deletion mutants exhibited repellent responses to the chemoeffectors. In contrast, mutants deficient in all five MCP-encoding genes did not exhibit repellent responses to all tested chemoeffectors. These results indicate that the chemoreceptors of L. agilis are functionally redundant and may be involved in chemotactic responses to chemoeffectors in a complementary manner. This is consistent with the results of a chemotaxis assay using mutants expressing a single chemoreceptor, in which chemotaxis to each effector was observed in some or all mutants. Functional redundancy of chemoreceptors has also been described in many bacteria and may be a common survival strategy for motile bacteria in their environment6. However, the need for multiple chemoreceptors to detect the same ligands in bacteria remains unclear. One possible reason for this redundancy is that the chemotaxis of bacteria is not impaired, even if there is a loss or mutation of any chemoreceptor. The chemotaxis of L. agilis to chemoeffectors was not completely impaired in L. agilis mutants lacking one of the five chemoreceptors. In particular, the chemotaxis of the mutants in response to low pH and organic acids was comparable to that of the WT. Thus, the functional redundancy of L. agilis chemoreceptors for gut-derived substances may contribute to the robust chemotactic ability of L. agilis in the environment.
The findings of this study also indicate that each of the five chemoreceptors in L. agilis can sense multiple chemoeffectors. Mcp2, Mcp3, and Mcp4, which are involved in the responses to low pH, some organic acids, and bile components, may sense a broad range of ligands. Domain architecture analysis of the chemoreceptors revealed that the LBDs (sensing domains) of L. agilis chemoreceptors, except for Mcp1, contained a dCache_1 domain. Several dCache-containing chemoreceptors can sense multiple ligands62,63,64,65,66,67,68. Our findings are consistent with those of previous studies.
Several chemoreceptors of the chemoeffectors tested in this study have been previously reported (Supplementary Table S2). To our knowledge, this is the first study to report chemoreceptors in the SC. Mcp2, Mcp3, and Mcp4 are the only identified dCache-containing chemoreceptors for butyrate and acetate. Therefore, the findings of this study may provide additional information on bacterial chemoreceptor-ligand interactions. However, the affinity of chemoreceptors for effectors and the ability of the receptors to directly sense the effectors are unknown. Thus, further studies (e.g., isothermal titration calorimetry analysis) are required. Some motile bacteria monitor their intracellular energy levels via chemoreceptors and migrate to favorable niches for energy generation69,70. This bacterial behavior is known as energy taxis, and the effectors of energy taxis include electron acceptors, light, redox chemicals, and metabolizable substrates. For example, E. coli senses the proton motive force and the redox state of the electron transport system via two chemoreceptors, Tsr and Aer71,72. In H. pylori, low pH, which directly influences the proton motive force, and changes in the electron transport chain are sensed by TlpB and TlpD, respectively15,73. Although the chemotactic behavior of L. agilis is not completely understood, further studies may reveal that it includes energy taxis.
In conclusion, this study demonstrated that L. agilis, which is highly sensitive to gut-associated stress conditions, escapes the stress and metabolites of competitive gut microbiota via chemotaxis. This suggests that L. agilis can utilize chemotaxis to facilitate its survival and persistence in the gastrointestinal tract. Multiple L. agilis chemoreceptors mediate the chemotaxis of each gut-derived substance in a complementary manner. This functional redundancy of chemoreceptors may confer a robust chemotactic ability to L. agilis in the environment. The findings of this study provide further insights into the interactions between chemoreceptors and ligands of motile lactobacilli and the unique ecological features and evolutionary strategies of motile lactobacilli, which may be distinct from those of non-motile lactobacilli.
Methods
Bacterial strains and growth conditions
The bacterial strains and plasmids used in this study are listed in Table 1. L. agilis BKN88 74 and its derivative strains were anaerobically propagated in MRS broth or agar (Difco, BD, USA) with or without 5 μg mL−1 of erythromycin at 37 °C. The motility of L. agilis strains was determined by observing strains cultured in a semi-solid MRS medium with 0.3% agar. Bacterial motility was observed using a BZ-X710 microscope (KEYENCE, Osaka, Japan). E. coli EC101 and derivative strains were aerobically grown in Brain Heart Infusion (Difco, BD, USA) broth or agar with or without 200 μg mL−1 of erythromycin and 40 μg mL−1 of kanamycin at 37 °C.
Microscopic agar-drop assay
The microscopic agar-drop assay was performed according to the protocols described by Islarm et al.33 with minor modifications (Supplementary Fig. S2). A flow chamber containing an agar drop was prepared as follows: Test reagent in distilled water containing 1.5% (w/v) agar was dropped onto the center of a long cover glass (NEO Micro Cover Glass, 24 × 60 mm, No. 1:0.12–0.17 mm in thickness, Matsunami, Osaka, Japan), followed by the attachment of a small cover glass (Micro Cover Glass, 18 × 18 mm, No. 1:0.12–0.17 mm in thickness, Matsunami, Osaka Japan) to the long cover glass using double-sided tape. The agar drop in the flow chamber was washed with chemotaxis buffer (10 mM potassium phosphate, 1 mM glucose, and 0.1 mM EDTA in pure water, pH 7.0). Bacterial cells in the exponential phase were harvested via centrifugation at 3900 rpm at 25 °C for 5 min and suspended in an MRS medium. The bacterial suspension was diluted 1:15 with a chemotaxis buffer and infused into the flow chamber. Motile cells near the agar drop were immediately observed, and images were captured using a time-lapse microscope (10 × objective, BZ-X710, Keyence, Osaka, Japan) once every 1 min. As shown in Supplementary Fig. S2, the number of cells in the red dotted area (500 µm × 500 µm area) was counted using ImageJ software (National Institutes of Health, Bethesda, MD, USA)75. The relative number of cells represents the ratio of the number of cells in the area at each time point to the number at 0 min. An agar drop without the test chemicals was used as a control.
Test chemicals
The following chemicals were used in the chemotaxis assay: bile salts (Sigma‐Aldrich, USA), SDC (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), SC (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), STC (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), mucin from the porcine stomach (Type II, Sigma-Aldrich, USA), lactic acid (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan), butyric acid (Tokyo Chemical Industry Co., Ltd, Tokyo, Japan), and acetic acid (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). The pH of the test reagents used in the chemotaxis assays (excluding the pH taxis assay) was adjusted to 7.0 with HCl or NaOH solution. In the pH taxis assay, the pH of the agar drop was adjusted to 3.0, 5.0, 7.0, or 12.0, using HCl or NaOH.
Expression analysis of MCP-encoding genes
Transcription of the five putative MCP-encoding genes was determined by RT-PCR using the specific primers listed in Supplementary Table S2. Total RNA was isolated from L. agilis BKN88 in the exponential phase using NucleoSpin RNA (Macherey–Nagel, Germany), following the manufacturer’s instructions. To digest the contaminated genomic DNA, RNA samples were treated with deoxyribonuclease (RT grade, Nippon Gene, Tokyo, Japan). RT-PCR was performed using a PrimeScript One Step RT-PCR Kit (Takara, Japan). DNA contamination in the RNA samples was tested by PCR using the same primers as those used for RT-PCR and Ex. Taq DNA polymerase (Takara, Japan).
Construction of MCP-encoding or CheA-encoding gene deletion mutants
Putative MCP-encoding or CheA-encoding genes in L. agilis BKN88 were deleted via double-crossover using the thermosensitive plasmid pG+host5. The DNA fragments of the upstream and downstream regions of the target gene were amplified by PCR using the primers listed in Supplementary Table S3. The amplicons were fused by overlap PCR. The resulting PCR products were digested with appropriate restriction endonucleases, ligated into a similarly digested pG+host5 vector, and transformed into E. coli EC101 as the cloning host. Transformants harboring plasmids with the desired insert were selected using colony PCR and Sanger sequencing with the M13 primers DOKJ78 (5′-GTAAAACGACGGCCAGT-3′) and DOKJ79 (5′-CAGGAAACAGCTATGAC-3′). The constructed plasmids were introduced into L. agilis BKN88 via electroporation as described previously31. Erythromycin-resistant transformants were selected and cultured in MRS containing erythromycin at 28 °C. To integrate the plasmids into the genome, the transformants were then sub-cultured in MRS containing erythromycin at 42 °C, plated on MRS agar plates containing erythromycin, and incubated at 42 °C until colonies were visible. PCR confirmed the integration. The integrants were cultured in MRS without erythromycin at 37 °C, followed by seven to nine transfers in MRS without erythromycin. After plating the bacterial cultures on MRS agar plates, single colonies were replicated on MRS agar plates and MRS agar plates containing erythromycin. Deletion of target genes in erythromycin-sensitive transformants was validated by PCR using the primers listed in Supplementary Table S3.
Analysis of swimming behavior
The swimming behavior of the WT and non-chemotactic (ΔcheA) L. agilis strains at the exponential phase was observed. Individual motile cells (n = 10) were tracked for 10 s (15 frames per second) under a BZ-X710 microscope (10 × objective; Keyence, Osaka, Japan) using video editing analysis software VW-H2MA (Keyence, Osaka, Japan). All coordinates (x,y) of the starting point of each track were set to (0,0).
Bile tolerance
The bile tolerances of L. agilis BKN88 and the three non-motile Lactobacillus species listed in Table 1 were examined as previously described, with minor modifications76. Bacterial cells from the exponential phase were serially diluted, plated onto MRS agar plates containing 0%, 0.25%, 0.5%, 1.0%, and 2.0% (w/v) oxgall (Difco, USA), and incubated at 37 °C anaerobically for enumeration. Oxgall concentrations were used to simulate intestinal conditions.
Acid tolerance
The acid tolerance of L. agilis BKN88 and the three non-motile Lactobacillus species listed in Table 1 was examined following the method described by Liong and Shah77. Bacterial cells from the exponential phase were suspended in MRS broth with pH adjusted to 3.0, with HCl, and incubated at 37 °C for 120 min. The bacterial cells were collected once every 30 min, plated onto MRS agar plates, and incubated at 37 °C anaerobically for enumeration.
Data availability
All data described in the manuscript are presented in the main text, figures, and Supporting Information.
References
Adler, J. Chemotaxis in bacteria motile Escherichia coli migrate in bands that are influenced by oxygen and organic nutrients. Adv. Sci. 153, 708–716 (2008).
Wadhams, G. H. & Armitage, J. P. Making sense of it all: Bacterial chemotaxis. Nat. Rev. Mol. Cell Biol. 5, 1024–1037 (2004).
Porter, S. L., Wadhams, G. H. & Armitage, J. P. Signal processing in complex chemotaxis pathways. Nat. Rev. Microbiol. 9, 153–165 (2011).
Parkinson, J. S., Hazelbauer, G. L. & Falke, J. J. Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol. 23, 257–266 (2015).
Sourjik, V. & Wingreen, N. S. Responding to chemical gradients: Bacterial chemotaxis. Curr. Opin. Cell Biol. 24, 262–268 (2012).
Ortega, Á., Zhulin, I. B. & Krell, T. Sensory repertoire of bacterial chemoreceptors. Microbiol. Mol. Biol. Rev. 81, 1–28 (2017).
Hazelbauer, G. L., Falke, J. J. & Parkinson, J. S. Bacterial chemoreceptors: High-performance signaling in networked arrays. Trends Biochem. Sci. 33, 9–19 (2008).
Worku, M. L., Karim, Q. N., Spencer, J. & Sidebotham, R. L. Chemotactic response of Helicobacter pylori to human plasma and bile. J. Med. Microbiol. 53, 807–811 (2004).
Mizote, T., Yoshiyama, H. & Nakazawa, T. Urease-independent chemotactic responses of Helicobacter pylori to urea, urease inhibitors, and sodium bicarbonate. Infect. Immun. 65, 1519–1521 (1997).
Huang, J. Y. et al. Chemodetection and destruction of host urea allows Helicobacter pylori to locate the epithelium. Cell Host Microbe 18, 147–156 (2015).
Goers Sweeney, E. et al. Structure and proposed mechanism for the pH-sensing Helicobacter pylori chemoreceptor TlpB. Structure 20, 1177–1188 (2012).
Machuca, M. A. et al. Helicobacter pylori chemoreceptor TlpC mediates chemotaxis to lactate. Sci. Rep. 7, 1–15 (2017).
Cerda, O., Rivas, A. & Toledo, H. Helicobacter pylori strain ATCC700392 encodes a methyl-accepting chemotaxis receptor protein (MCP) for arginine and sodium bicarbonate. FEMS Microbiol. Lett. 224, 175–181 (2003).
Yoshiyama, H., Nakamura, H., Kimoto, M., Okita, K. & Nakazawa, T. Chemotaxis and motility of Helicobacter pylori in a viscous environment. J. Gastroenterol. 34, 18–23 (1999).
Croxen, M. A., Sisson, G., Melano, R. & Hoffman, P. S. The Helicobacter pylori chemotaxis receptor tlpB (HP0103) is required for pH taxis and for colonization of the gastric mucosa. J. Bacteriol. 188, 2656–2665 (2006).
Sanders, L., Andermann, T. M. & Ottemann, K. M. A supplemented soft agar chemotaxis assay demonstrates the Helicobacter pylori chemotactic response to zinc and nickel. Microbiology (United Kingdom) 159, 46–57 (2013).
Collins, K. D. et al. The Helicobacter pylori CZB cytoplasmic chemoreceptor TlpD forms an autonomous polar chemotaxis signaling complex that mediates a tactic response to oxidative stress. J. Bacteriol. 198, 1563–1575 (2016).
Rader, B. A. et al. Helicobacter pylori perceives the quorum-sensing molecule AI-2 as a chemorepellent via the chemoreceptor TlpB. Microbiology 157, 2445–2455 (2011).
Johnson, K. S. & Ottemann, K. M. Colonization, localization, and inflammation: the roles of H. pylori chemotaxis in vivo. Curr. Opin. Microbiol. 41, 51–57 (2018).
Keilberg, D. & Ottemann, K. M. How Helicobacter pylori senses, targets and interacts with the gastric epithelium. Environ. Microbiol. 18, 791–806 (2016).
Glekas, G. D. et al. The Bacillus subtilis chemoreceptor McpC senses multiple ligands using two discrete mechanisms. J. Biol. Chem. 287, 39412–39418 (2012).
Hanlon, D. W. & Ordal, G. W. Cloning and characterization of genes encoding methyl-accepting chemotaxis proteins in Bacillus subtilis. J. Biol. Chem. 269, 14038–14046 (1994).
Müller, J., Schiel, S., Ordal, G. W. & Saxild, H. H. Functional and genetic characterization of mcpC, which encodes a third methyl-accepting chemotaxis protein in Bacillus subtilis. Microbiology 143, 3231–3240 (1997).
Matilla, M. A. & Krell, T. The effect of bacterial chemotaxis on host infection and pathogenicity. FEMS Microbiol. Rev. 42, 40–67 (2018).
Cousin, F. J. et al. Detection and genomic characterization of motility in Lactobacillus curvatus: Confirmation of motility in a species outside the Lactobacillus salivarius clade. Appl. Environ. Microbiol. 81, 1297–1308 (2015).
Suzuki, S. et al. PCR-based screening, isolation, and partial characterization of motile lactobacilli from various animal feces. BMC Microbiol. 20, 1–11 (2020).
Gaucher, F. et al. Review: Adaptation of beneficial propionibacteria, lactobacilli, and bifidobacteria improves tolerance toward technological and digestive stresses. Front. Microbiol. 10, 1–22 (2019).
Goh, Y. J. & Klaenhammer, T. R. Insights into glycogen metabolism in Lactobacillus acidophilus: Impact on carbohydrate metabolism, stress tolerance and gut retention. Microb. Cell Fact. 13, 1–12 (2014).
Ruiz, L., Margolles, A. & Sánchez, B. Bile resistance mechanisms in Lactobacillus and Bifidobacterium. Front. Microbiol. 4, 1–8 (2013).
Duar, R. M. et al. Lifestyles in transition: Evolution and natural history of the genus Lactobacillus. FEMS Microbiol. Rev. 41, S27–S48 (2017).
Kajikawa, A., Suzuki, S. & Igimi, S. The impact of motility on the localization of Lactobacillus agilis in the murine gastrointestinal tract. BMC Microbiol. 18, 1–7 (2018).
Rao, C. V., Kirby, J. R. & Arkin, A. P. Design and diversity in bacterial chemotaxis: A comparative study in Escherichia coli and Bacillus subtilis. PLoS Biol. 2, 239–252 (2004).
Islam, M. S., Takabe, K., Kudo, S. & Nakamura, S. Analysis of the chemotactic behaviour of Leptospira using microscopic agar-drop assay. FEMS Microbiol. Lett. 356, 39–44 (2014).
Larsen, S. H., Reader, R. W., Kort, E. N., Tso, W. W. & Adler, J. Change in direction of flagellar rotation is the basis of the chemotactic response in Escherichia coli. Nature 249, 74–77 (1974).
Krikos, A., Conley, M. P., Boyd, A., Berg, H. C. & Simon, M. I. Chimeric chemosensory transducers of Escherichia coli. Proc. Natl. Acad. Sci. U. S. A. 82, 1326–1330 (1985).
Kihara, M. & Macnab, R. M. Cytoplasmic pH mediated pH taxis and weak-acid repellent taxis of bacteria. J. Bacteriol. 145, 1209–1221 (1981).
Repaske, D. R. & Adler, J. Change in intracellular pH of Escherichia coli mediates the chemotactic response to certain attractants and repellents. J. Bacteriol. 145, 1196–1208 (1981).
Umemura, T., Matsumoto, Y., Ohnishi, K., Homma, M. & Kawagishi, I. Sensing of cytoplasmic pH by bacterial chemoreceptors involves the linker region that connects the membrane-spanning and the signal-modulating helices. J. Biol. Chem. 277, 1593–1598 (2002).
Yang, Y. & Sourjik, V. Opposite responses by different chemoreceptors set a tunable preference point in Escherichia coli pH taxis. Mol. Microbiol. 86, 1482–1489 (2012).
Huang, J. Y., Goers Sweeney, E., Guillemin, K. & Amieva, M. R. Multiple acid sensors control helicobacter pylori colonization of the stomach. PLoS Pathog. 13, 1–28 (2017).
Van de Guchte, M. et al. Stress responses in lactic acid bacteria. Antonie van Leeuwenhoek Int. J. Gen. Mol. Microbiol. 82, 187–216 (2002).
Papadimitriou, K. et al. Stress physiology of lactic acid bacteria. Microbiol. Mol. Biol. Rev. 80, 837–890 (2016).
Kullen, M. J. & Klaenhammer, T. R. Identification of the pH-inducible, proton-translocating F 1 F 0 -ATPase (atpBEFHAGDC) operon of Lactobacillus acidophilus by differential display: Gene structure, cloning and characterization. Mol. Microbiol. 33, 1152–1161 (1999).
Hugdahl, M. B., Beery, J. T. & Doyle, M. P. Chemotactic behavior of Campylobacter jejuni. Infect. Immun. 56, 1560–1566 (1988).
Li, Z. et al. Methyl-accepting chemotaxis proteins 3 and 4 are responsible for Campylobacter jejuni chemotaxis and jejuna colonization in mice in response to sodium deoxycholate. J. Med. Microbiol. 63, 343–354 (2014).
Nishiyama, S. I. et al. Identification of a Vibrio cholerae chemoreceptor that senses taurine and amino acids as attractants. Sci. Rep. 6, 1–11 (2016).
Begley, M., Gahan, C. G. M. & Hill, C. The interaction between bacteria and bile. FEMS Microbiol. Rev. 29, 625–651 (2005).
Li, C., Boileau, A. J., Kung, C. & Adler, J. Osmotaxis in Escherichia coli. Proc. Natl. Acad. Sci. U. S. A. 85, 9451–9455 (1988).
Matilla, M. A., Ortega, Á. & Krell, T. The role of solute binding proteins in signal transduction. Comput. Struct. Biotechnol. J. 19, 1786–1805 (2021).
Upadhyay, A. A., Fleetwood, A. D., Adebali, O. & Finn, R. D. Cache domains that are homologous to, but different from PAS domains comprise the largest superfamily of extracellular sensors in prokaryotes. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1004862 (2016).
Ghosh, T. S. Metagenomic analysis reveals distinct patterns of gut lactobacillus prevalence, abundance, and geographical variation in health and disease. Gut Microbes 12, 1–19 (2020).
Szymanski, C. M., King, M., Haardt, M. & Armstrong, G. D. Campylobacter jejuni motility and invasion of Caco-2 cells. Infect. Immun. 63, 4295–4300 (1995).
Golden, N. J. & Acheson, D. W. K. Identification of motility and autoagglutination Campylobacter jejuni mutants by random transposon mutagenesis. Infect. Immun. 70, 1761–1771 (2002).
O’Toole, P. W., Lane, M. C. & Porwollik, S. Helicobacter pylori motility. Microbes Infect. 2, 1207–1214 (2000).
Aihara, E. et al. Motility and chemotaxis mediate the preferential colonization of gastric injury sites by Helicobacter pylori. PLoS Pathog. 10, e1004275 (2014).
Josenhans, C. & Suerbaum, S. The role of motility as a virulence factor in bacteria. Int. J. Med. Microbiol. 291, 605–614 (2002).
Andino, A. & Hanning, I. Salmonella enterica: Survival, colonization, and virulence differences among serovars. Sci. World J. https://doi.org/10.1155/2015/520179 (2015).
Millet, Y. A. et al. Insights into Vibrio cholerae intestinal colonization from monitoring fluorescently labeled bacteria. PLoS Pathog. 10, e1004405 (2014).
Butler, S. M. & Camilli, A. Both chemotaxis and net motility greatly influence the infectivity of Vibrio cholerae. Proc. Natl. Acad. Sci. U. S. A. 101, 5018–5023 (2004).
Ricke, S. C. Perspectives on the use of organic acids and short chain fatty acids as antimicrobials. Poult. Sci. 82, 632–639 (2003).
Sun, Y. & O’Riordan, M. X. D. Chapter three - regulation of bacterial pathogenesis by intestinal short-chain fatty acids. In (eds. Sariaslani, S. & Gadd, G. M. B. T.-A. in A. M.) vol 85, 93–118 (Academic Press, 2013).
Feng, H. et al. Signal binding at both modules of its dCache domain enables the McpA chemoreceptor of Bacillus velezensis to sense different ligands. Proc. Natl. Acad. Sci. 119, e2201747119 (2022).
Ud-din, A. I. M. S., Khan, M. F. & Roujeinikova, A. Broad specificity of amino acid chemoreceptor CtaA of pseudomonas fluorescens is afforded by plasticity of its amphipathic ligand-binding pocket. Mol. Plant-Microbe Interact. 33, 612–623 (2020).
Webb, B. A., Compton, K. K., Ray, W. K., Helm, R. F. & Scharf, B. E. Sinorhizobium meliloti chemotaxis to quaternary ammonium compounds is mediated by the chemoreceptor McpX. Mol. Microbiol. 103, 333–346 (2017).
Fernández, M., Morel, B., Corral-Lugo, A. & Krell, T. Identification of a chemoreceptor that specifically mediates chemotaxis toward metabolizable purine derivatives. Mol. Microbiol. 99, 34–42 (2016).
Gavira, J. A. et al. Structural basis for polyamine binding at the dCACHE domain of the McpU Chemoreceptor from Pseudomonas putida. J. Mol. Biol. 430, 1950–1963 (2018).
Corral Lugo, A. et al. High-Affinity Chemotaxis to Histamine Mediated by the TlpQ chemoreceptor of the human pathogen Pseudomonas aeruginosa. MBio 9, 1–15 (2018).
Johnson, K. S. et al. The dCache chemoreceptor TlpA of helicobacter pylori binds multiple attractant and antagonistic ligands via distinct sites. MBio 12, e0181921. https://doi.org/10.1128/mBio.01819-21 (2021).
Alexandre, G., Greer-Phillips, S. & Zhulin, I. B. Ecological role of energy taxis in microorganisms. FEMS Microbiol. Rev. 28, 113–126 (2004).
Josenhans, C. & Schweinitzer, T. Bacterial energy taxis: A global strategy?. Arch. Microbiol. 192, 507–520 (2010).
Rebbapragada, A. et al. The Aer protein and the serine chemoreceptor Tsr independently sense intracellular energy levels and transduce oxygen, redox, and energy signals for Escherichia coli behavior. Proc. Natl. Acad. Sci. U. S. A. 94, 10541–10546 (1997).
Edwards, J. C., Johnson, M. S. & Taylor, B. L. Differentiation between electron transport sensing and proton motive force sensing by the Aer and Tsr receptors for aerotaxis. Mol. Microbiol. 62, 823–837 (2006).
Schweinitzer, T. et al. Functional characterization and mutagenesis of the proposed behavioral sensor TlpD of Helicobacter pylori. J. Bacteriol. 190, 3244–3255 (2008).
Kajikawa, A. et al. Characterization of flagellins isolated from a highly motile strain of Lactobacillus agilis. BMC Microbiol. 16, 49 (2016).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Pfeiler, E. A., Azcarate-Peril, M. A. & Klaenhammer, T. R. Characterization of a novel bile-inducible operon encoding a two-component regulatory system in Lactobacillus acidophilus. J. Bacteriol. 189, 4624–4634 (2007).
Liong, M. T. & Shah, N. P. Acid and bile tolerance and cholesterol removal ability of lactobacilli strains. J. Dairy Sci. 88, 55–66 (2005).
Law, J. et al. A system to generate chromosomal mutations in Lactococcus lactis which allows fast analysis of targeted genes. J. Bacteriol. 177, 7011–7018 (1995).
Altermann, E. et al. Complete genome sequence of the probiotic lactic acid bacterium Lactobacillus acidophilus NCFM. Proc. Natl. Acad. Sci U. S. A. 102, 3906–3912 (2005).
Biswas, I., Gruss, A., Ehrlich, S. D. & Maguin, E. High-efficiency gene inactivation and replacement system for gram-positive bacteria. J. Bacteriol. 175, 3628–3635 (1993).
Mitchell, A. L. et al. InterPro in 2019: Improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res. 47, D351–D360 (2019).
Adebali, O., Ortega, D. R. & Zhulin, I. B. CDvist: A webserver for identification and visualization of conserved domains in protein sequences. Bioinformatics 31, 1475–1477 (2015).
Acknowledgements
We thank Kazuho Kondo and Naoto Eguchi for their help with preliminary work, which could not be included in the final manuscript. This work was supported by the Tokyo University of Agriculture. We thank Editage [http://www.editage.com] for editing and reviewing the manuscript for the English language.
Funding
This work was supported by the Tokyo University of Agriculture. The funding body played no role in the study, article preparation, or publication decision.
Author information
Authors and Affiliations
Contributions
S.S. and A.K. designed the study and prepared the manuscript. S.S. performed and analyzed the experiments. All the authors interpreted the data and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suzuki, S., Yokota, K., Igimi, S. et al. Negative chemotaxis of Ligilactobacillus agilis BKN88 against gut-derived substances. Sci Rep 13, 15632 (2023). https://doi.org/10.1038/s41598-023-42840-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-42840-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.