Abstract
Lens lamottei is a member of the Fabaceae family and the second gene pool of the genus Lens. The environmental factors that drove the divergence among wild and cultivated species have been studied extensively. Recent research has focused on genomic signatures associated with various phenotypes with the acceleration of next-generation techniques in molecular profiling. Therefore, in this study, we provide the complete sequence of the chloroplast genome sequence in the wild Lens species L. lamottei with a deep coverage of 713 × next-generation sequencing (NGS) data for the first time. Compared to the cultivated species, Lens culinaris, we identified synonymous, and nonsynonymous changes in the protein-coding regions of the genes ndhB, ndhF, ndhH, petA, rpoA, rpoC2, rps3, and ycf2 in L. lamottei. Phylogenetic analysis of chloroplast genomes of various plants under Leguminosae revealed that L. lamottei and L. culinaris are closest to one another than to other species. The complete chloroplast genome of L. lamottei also allowed us to reanalyze previously published transcriptomic data, which showed high levels of gene expression for ATP-synthase, rubisco, and photosystem genes. Overall, this study provides a deeper insight into the diversity of Lens species and the agricultural importance of these plants through their chloroplast genomes.
Similar content being viewed by others
Introduction
Lentils are nutritionally important protein-rich crops that provide essential and nonessential amino acids1, carbohydrates, lipids, dietary fiber, minerals, and water-soluble vitamins2,3. Lentils are self-pollinating legumes belonging to Fabaceae (Leguminosae), the third-largest plant family4. This family includes plants with economic value5. The genus Lens consists of seven species belonging to four gene pools. Lens culinaris is the cultivated form, while Lens lamottei, Lens orientalis, Lens tomentosus, Lens ervoides, Lens odemensis, and Lens nigricans are wild species6. Their natural habitats are distributed in the Mediterranean region and Central Asia7.
Lens lamottei belongs to the second gene pool of Lens and is a member of the Papillionodeae subfamily3. Czefronova and colleagues were the first to describe L. lamottei, which originated from Spain, Italy, and Turkey8. L. lamottei has a small seed, at approximately 1.92 mm, with a round shape9. Its stipules are horizontal, and the leaves are less dentate9. L. lamottei has one of the highest average harvest indices, the highest number of seeds per plant, the highest seed yield, and the highest biomass10. It is highly resistant to various biotic stressors, such as Stremphylium blight, Fusarium wilt, and Bruchid weevils. It ranks second among wild species in terms of resistance to anthracnose and Ascochyta blight11. It also has enhanced tolerance to drought, salinity, and alkalinity9.
The chloroplast (cp) is a crucial organelle that plays a major role in photosynthesis, starch production, and metabolism of amino acids, fatty acids, pigments, vitamins, sulfur, and nitrogen12,13. It harnesses energy that can be supplied to the plant cell14. Cp-based studies are vital in plant biotechnology, especially in plant breeding studies15. Cp genomes can be used to construct phylogenetic trees to elucidate distinctive plant features, such as maternal inheritance, high copy numbers, and self-replication ability16,17. It provides valuable information about the structural variations and exposes the molecular divergence of related species18.
The cp has an independent genome that is around 120–170 kb in size and consists of 120–130 genes mainly involved in photosynthesis19. The recently published cp genome of L. ervoides, one of the close relatives of L. lamottei, is also approximately 120 kb20. Cp genome size is more conversed across species as opposed to the mitochondrial genome21. It is stable and highly conserved, with the mutation rate of the cp genome lower than the nuclear genome13,22. The cp genome has a quadripartite structure with a small single copy region (SSC), a large single copy region (LSC), and two inverted repeat regions (IRs)22. Species carrying a single copy IR region in their cp genome structure are classified under the Inverted Repeat Lacking Clade (IRLC)23, which includes Cicer arietinum, Pisum sativum, Vicia faba, L. culinaris, and L. ervoides20,24. The IRLC is a useful model system for examining how the evolution of the plastome is impacted by the loss of a single IR region25.
Divergence patterns, substitution rates, and phylogenetic relations across land plants depend heavily on the comparative analyses of their genomic sequences26,27. Expansion or contraction of simple sequence repeats (SSRs) can be potential DNA markers for identification of species18. Genomic data is also utilized in studying gene expression and regulation. Recently, comparisons of transcriptomic profiles have been reported to determine differential gene expression28. Likewise, RNA editing sites and codon usage bias have been studied to identify the role of cp genes and these gene’s regulatory mechanisms29,30.
The lack of a cp genome sequence for L. lamottei limits its phylogenetic examination across related species and its potential to associate with wild-type features that can be agriculturally beneficial. In the current study, we report the cp sequence of L. lamottei and compare it with L. culinaris.
Results
Genomic features and organization of L. lamottei cp DNA
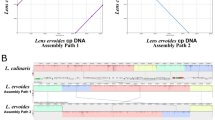
The BGISEQ-500 platform was used to sequence the cp genome of L. lamottei. The de novo cp genome of L. lamottei was assembled using the GetOrganelle pipeline via two different assembly paths. To examine genome rearrangement, the Mauve aligner was used to align L. culinaris and L. lamottei’s cp genome. The alignment calculated the locally collinear blocks (LCB), which represent highly similar, or conserved regions. When these assemblies were compared, certain regions of assembly path 1 were detected to be inverted without any missing parts in reference to L. culinaris (Fig. 1a). On the other hand, assembly path 2 was performed by carrying the swapped blocks on the same strand. After manually reorienting the major collinear blocks for both of these assembly options, we found that the L. lamottei genome has the same structures and gene content as the cp genome of L. culinaris (Fig. 1b).
Gene arrangement analysis using Locally Colinear Blocks (LCBs). MAUVE software was used to compare the gene order of L. lamottei with the genome of L. culinaris cp as a reference. (a) Comparison of two possible haplotypes of L. lamottei without manual correction. (b) Comparison of two possible haplotypes of L. lamottei after manual correction.
As a result of our de novo genome construction analysis, the cp genome of L. lamottei was assembled with a size of 122,855 bp and includes the LSC, SSC, and one IR region. Therefore, L. lamottei can be classified under the IRLC. Cross-species plastome comparison was done to identify genic regions, including protein-coding, tRNA, and rRNA loci. Figure 2 shows the structural organization of both Lens cp genomes with the same orientation of the genic regions. The L. lamottei and L. culinaris cp genomes were similar in genic content and structural organization. However, due to ambiguities in the annotation approaches, a few genes detected in L. lamottei were not found in L. culinaris. These genes are pafII(ycf4), rps18, and rpl22. Another difference is that the pafI gene was annotated as a protein-coding gene in L. lamottei but was defined as a pseudogene in the annotation of L. culinaris. This is because the annotation of the L. culinaris genome was performed using a different tool (DOGMA). When the cp genome of L. culinaris was reannotated using the GeSeq tool, the two cp genomes contained the same set of genes. L. lamottei and L. culinaris have 108 and 105 genes, respectively, that make up the cp genome, including 77, and 73 protein-coding genes, respectively. They have the same number of tRNA (27) and rRNA (4) genes (Supplementary Table S1). These genes have various functions in the cp, including photosynthesis, transcription, and translation (Supplementary Table S2).
The circular cp genome map of (a) L. lamottei (b) L.culinaris. Genes inside the circle are transcribed in the clockwise direction, whereas genes outside the circle are transcribed in the counterclockwise direction. The darker gray on the inside refers to GC content, while the lighter gray on the outside corresponds to AT content. Genes are color-coded according to their function.
Introns play a crucial role in regulating gene expression. Overall, there are 17 intron-containing genes. Among these, six are tRNA genes (trnK-UAA, trnF-AAA, trnL-UAA, trnI-GAU, trnE-UUC, and trnA-UGC), and ten are protein-coding genes (rpoC1, atpF, clpP1, petB, petD, rpl16, rpl2, ndhB, rps12, and ndhA). PafI (ycf3) is the only gene containing two introns. The longest intron (2423 bp) was found in the trnK-UUU gene, which also contained the matK gene. The smallest intron (39 bp) was found in the trnA-UGC gene.
Comparative genome analysis
To study the degree of sequence divergence between the cp genomes of L. lamottei and L. culinaris mVISTA was used to visualize the overall sequence identities of the two cp genomes (Fig. 3). Comparative analyses showed a high sequence similarity with no discernible differences between the wild and cultivated species. We observed similar gene orders and organizations between the two species and highly conserved protein-coding genes with limited substitutions.
Visualization of alignment of the L. lamottei cp genome using L. culinaris as a reference sequence using mVISTA. The vertical axis shows the percentage of identity, which goes from 50 to 100%. In the reference plastome, each arrow represents annotated genes and the direction of transcription. Exons are purple, the untranslated region (UTR) is blue, and conserved non-coding sequences (CNS) are pink in the plastome coding areas. A decrease in purple/pink shadowing indicates a decrease in sequence identity (white spaces).
The hotspot regions in the cp genomes of L. lamottei and L. culinaris were determined by comparing the nucleotide variation in the coding and noncoding regions. Compared to protein-coding regions, noncoding regions showed higher levels of divergence. Genomic hotspots were detected. High nucleotide variation was found in the intergenic region between the trnQ-accD genes with a dissimilarity of more than 1.0%. We identified a genomic hotspot in an intergenic region between 50,800 and 52,000 bp (Fig. 4).
Simple sequence repeats analysis
Repetitive region were summarized for both lens species in Fig. 5. SSRs, commonly referred to as microsatellite repeats, are shorter tandem repeats ranging from 1 to 6 bp in length and are found throughout the cp genome31. A total of 67 and 68 SSRs, respectively, were found in L. lamottei and L. culinaris with MISA (MIcroSAtellite Identification Tool). Of these, there were 49 and 47 mononucleotides, 6 dinucleotides in both, 2 and 4 trinucleotides, 6 and 5 tetranucleotides, and 4, and 5 pentanucleotides, respectively (Fig. 5a). One hexanucleotide repeat motif was detected in L. culinaris but not in L. lamottei. The most common type of SSR was the A/T mononucleotide repeat motif (Fig. 5b). There were differences between the two species in terms of trinucleotide, pentanucleotide, and hexanucleotide motifs. ATC/ATG, AAAAT/ATTTT, and AGATAT/ATATCT 2,1,1 repeat motifs were detected in L. culinaris, respectively. However, these motifs were not found in L. lamottei (Fig. 5b). Repetitions can be classified under four types: forward, reverse, palindromic, and complementary. Using REPuter, we found 50 forward repeats in both L. lamottei and L. culinaris cp genomes. There were 15 and 16 palindromic repeats detected in L. lamottei and L. culinaris, respectively (Fig. 5c). One reverse repeat was found in L. lamottei but none in L. culinaris. No complementary repeats were found in either species (Fig. 5c). In addition, 31 tandem repeats were found in L. lamottei and 36 in L. culinaris.
Using L. culinaris (NC_027152) as a reference sequence, synonymous and non synonymous variations were compared and evaluated to further investigate the divergence of nucleotides. Variations in protein-coding and non-coding regions in the cp genome were examined. Synonymous and nonsynonymous nucleotide changes in protein-coding regions were detected in the L. lamottei genome (Table 1). Out of 73 protein-coding genes, at least one variation was detected in 18 protein-coding genes. Of these, 8 protein-coding genes (ndhB, ndhF, ndhH, petA, rpoA, rpoC2, rps3, and ycf2) were found to have nonsynonymous changes, i.e., variation resulting in an amino acid replacement. In particular, more variations (L429F, M1290R, H1324D, F1436L, and I1751M) were identified in the ycf2 gene region compared to other genes. In addition, the only synonymous changes occurred in the other 10 genes in the cp genome. In addition, 32 noncoding regions were found (Supplementary Table S3). These genes consist of rRNA and tRNA genes. Only two single nucleotide variations were detected in the rrn23 gene.
RNA editing is a post-transcriptional modification that results in base conversion. The PREP-Cp (Predictive RNA Editor for plants) program was used to identify putative RNA editing sites in L. lamottei. We found 41 predicted RNA editing sites in 17 genes (Supplementary Table S4). The most frequent editing site predicted the change of serine to leucine (%29.3). From nucleotide base C to T, there were all replacement sites. The ndhB (7) gene has the highest number of editing sites. At the second codon position, the editing event was most prevalent. In addition, RNA editing sites of L. culinaris which were cultivated species of L. lamottei were also predicted (Supplementary Table S5), with 41 RNA editing sites in 17 genes and the ndhB (7) gene with the highest abundance of sites. The most frequent editing site in L. culinaris was also the change of serine to leucine (29.3%).
Codon preference and expression levels of protein-coding genes
Synonymous codons are multiple codons that encode one amino acid, and synonymous codons typically exhibit varied usage preferences as a result of selection pressure during plant evolution. All 20 amino acid and stop codons in the protein-coding genes of the L. lamottei’s cp genome were evaluated for codon content and relative synonymous codon usage (RSCU) values (Supplementary Fig. S1; Supplementary Table S6). A total of 19,509 codons was detected in the cp genome of L. lamottei. Looking at the distribution of these codons in amino acids, leucine was the most abundant, while cysteine was the least abundant. Among those amino acids prevalent ATT, which encodes isoleucine (Ile), was the most abundant codon in L. lamottei, with 910 TGC, which encodes cysteine (Cys), was the least frequently used codon with 48. Most amino acids show codon biases, while methionine (ATG) and tryptophan (TGG) were expressed by only one codon and have no codon bias. TTA (leucine) had the highest RSCU value, whereas CTC (leucine) had the lowest.
Prediction of gene expression based on codon usage requires defining a set of reference genes, which includes all genes encoding ribosomal proteins. Higher MELP (MILC-based Expression Level Predictor) values indicate higher gene expression levels and codon usage bias. The log10(FPKM) (Fragments per Kilobase Million) value is expected to increase with the MELP value. Genes with MELP values higher than 1.0 (Fig. 6a) were psbN, psbI, psbE, psbK, psaI, and ndhC, which play roles in photosynthesis, and accD.
(a) The correlation between each protein-coding gene's MELP score and the percentage of codons used that are preferred (RSCU > 1.0). Green indicates ribosomal genes, while red indicates other genes. Gene names are only assigned to non-ribosomal genes with MELP > 1.0. The linear regression is shown by the blue line. (b) The value of mean FPKM is used to represent cp gene expression. The X-axis shows the gene in which category they are involved. The Y-axis shows the mean FPKM values of biological function.
Using RNA-seq data from L. lamottei, we analyzed the expression of the 76 genes that code for proteins in the cp32. Reads were mapped to the L. lamottei cp genome we assembled in this study. Taking gene lengths into account, the numbers of reads corresponding to coding genes were computed and normalized. Gene expression values in FPKM were calculated for 76 protein-coding genes (Supplementary Table S7). We found that genes encoding ATP synthase showed the highest expression among all genes (Fig. 6b). This was followed by rubisco and photosystem genes.
Phylogenetic analysis
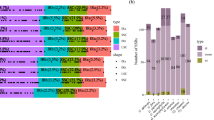
In this study, 14 species were included in the phylogenetic analysis to determine the relationships of L. lamottei with the other members of the Papilionoideae subfamily (Fig. 7). Arabidopsis thaliana was selected as an outgroup. Five species were chosen from Papilionoideae, which belongs to IRLC (L. culinaris, L. ervoides, C. ariethinum, Cicer echinospermum, Cicer bijigum), and two species from Papilionoideae (Cajanus cajan, Glycine max), two species from the Detarionoideae subfamily (Intsia bijuga, Crudia hamsiana), and two species from Cercidoideae subfamily (Cercis glabra, Tylosema esculentum). Phylogenetic analysis of the 13 cp genomes was performed to provide a better resolution with 1000 bootstraps. In the phylogenetic tree, L. lamottei formed a sister clade with L. culinaris, indicating that these two species are monophyletic and closely related.
Discussion
Lentils are richer in protein, carbohydrate, and dietary fiber content compared to other legumes. Therefore, they are consumed at higher rates worldwide due to their importance for human health1,2,3. The cp is a highly conserved organelle, both structurally and genetically, and is involved in many biological functions, particularly in photosynthesis16,17. Therefore, the cp genomes of various have been widely sequenced with the development of technologies22,33. In comparison to the nuclear genome, the cp genome is inherited maternally and can conveniently be used for phylogenetic investigations due to its short size and conserved structure34. We presented the first assembly and annotation of the cp genome of L. lamottei and compared it to the cultivated lentil species L. culinaris. By examining the genome rearrangements, we found highly similar regions between L. lamottei and L. culinaris. Two different haplotypes in L. lamottei have emerged. Two structural haplotypes of cp genomes that differ in the direction of single-copy regions have been identified in previously published studies35. Flip-flop recombination is a plausible theory to explain the existence of structural heteroplasmy and has been reported previously35,36. Gene loss, changes in the intergenic region, and expansion, or contraction of the IR region are all variables that affect genome size37. The cp genome of L. lamottei (122.8 kb) of the Fabaceae family has a similar size to that of the close wild relative L. ervoides20, the cultivated form L. culinaris, and other Fabaceae species such as O. arctobia38. The configuration of cp genomes in the species of this family is similar, with sizes ranging from 107 to 218 kb39.
The chloroplast genome of L. lamottei has an LSC, SSC region and a single copy IR region. In other plant cp genomes, a pair of IRs are separated by one LSC and one SSC22. Some species of the Fabaceae family, including L. culinaris contain a single copy of the IR region and belong to the IRLC26,38. IRs stabilize the plastome structure and affect its size40. However, expansion or contraction, gene loss, and genome rearrangement occurred in the IR regions of some species41. Therefore, the cp genomes of L. lamottei and L. culinaris are highly conserved and similar in genome organization.
The cp gene content was also highly conserved between L. lamottei and L. culinaris, except for some genes. Some genes annotated in L. lamottei could not be identified in the annotation of L. culinaris. This could be due to the limitations of DOGMA, the annotation tool used for the reference genome L. culinaris. When the cp genome was reannotated using the GeSeq tool, the genes on the cp genome matched with each other. The L. lamottei cp genome contains 107 genes, but some genes such as rps16 and infA were not identified. Many IRLC species plastomes contain regions with notable changes and rearrangements, such as the loss of introns from the rps12 and clpP genes and the lack of the rps16 gene42. The absence of these genes has been reported in other species20,23. Introns serve a critical function in gene expression regulation43. L. lamottei contains 17 genes that have an intron. The absence of one of the clpP1 introns in L. lamottei is consistent with the loss of the clpP1 intron in the genomes of highly conserved papilionoid IRLC members20,44.
The sliding window and mVISTA analysis demonstrated high sequence similarity between L. lamottei and L. culinaris, implying a highly conserved evolutionary model. The intergenic region between the trnQ and accD genes has several nucleotide variations in the cp genome of L. lamottei. Furthermore, the highly divergent areas were mostly found in the noncoding rather than coding regions, as previously shown in the Fabaceae family20,45. These hypervariable intergenic regions can serve as candidate regions for creating genetic markers46. These hotspots as markers can be used in the discrimination of species in the Lens genus for phylogenetic and identification studies.
Repetitive regions in the genome are essential for genome rearrangements47. SSRs can also serve as useful molecular markers for studying genetic diversity and evolutionary relationships in L. lamottei and similar species. Nucleotide and tandem repeats in cp genomes are associated with gene duplication, rearrangement, and expansion and can be particularly helpful markers for classifying populations or species46,48. SSRs are another type of molecular marker that consists of 1–6 nucleotide repeat units and are commonly employed in population genetics31. For both species, mononucleotide SSR is the most prominent motif. Although the numbers of repeat motifs are similar, they differ from each other in some motifs. While L. culinaris has one hexanucleotide repeat motif, L. lamottei has none. Of these repeat motifs, mononucleotide repeats are the most common SSR motif discovered in the cp genome49. Among these mononucleotides, the most common SSR motif found in the L. lamottei cp genome is the A/T motif. The common consensus is that plastid SSRs are mostly composed of A/T repeats, with G/C repeats considered as rare37. Similar results were found in the wild chickpea species belonging to Fabaceae family and in the L. ervoides cp genome20,50,51. SSRs have been identified in various organelle genomes and have great application potential for studies on the molecular evolution of plant population genetics and crop breeding52.
Understanding the mechanisms underlying molecular evolution critically depends on the estimation of synonymous and nonsynonymous nucleotide mutations53. Eight protein-coding genes (ndhB, ndhF, ndhH, petA, rpoA, rpoC2, rps3, and ycf2) in the cp genome of L. lamottei had nonsynonymous changes. Nonsynonymous mutations are those in which base mutation results in an alteration of the amino acid in the encoded protein21. Ycf1 and matK genes have been determined as molecular markers due to the detection of nonsynonymous substitutions in these genes in other studies53,54. However, in this study, while no substitution occurred in matK and ycf1 genes, several mutations were detected for the ycf2 gene (in the position of L429F, M1290R, H1324D, F1436L, I1751M). Numerous nonsynonymous changes were detected in the ycf2 gene for L. ervoides20, a close relative of L. lamottei. Among noncoding genes, only two single nucleotide substitutions were found in the rrn23 gene. Therefore, markers that can be developed using the ycf2 and rrn23 genes may shed light on future studies.
RNA editing is a fundamental post-transcriptional process that commonly occurs in the cp protein-coding regions to restore the evolutionarily conserved amino acid sequence24,55. The ndhB gene, which had seven RNA editing sites, contained the highest number of editing sites. The NDH protein complex serves an important function in photosynthesis. RNA editing may enhance photosynthetic efficiency and exhibit positive selection during evolution41. In 29.3% of the editing sites, serine aminoacids were converted to leucine. Many RNA editing sites can convert the encoded amino acid from polar to nonpolar in both species. In previous studies, unlike hydrophilic mutations, which can affect the secondary structure and function of proteins and improve their genetic information, hydrophobic mutations produce a more stable structure in proteins24,55,66. Therefore, the presence of RNA editing sites in the cp genome may elucidate evolutionary mechanisms.
Codon usage significantly influences the evolution of the cp genome and can also be used to determine gene functions, gene expression, and levels of mRNA and proteins21,56. The L. lamottei cp genome has 19,509 codons, with cysteine having the lowest codon usage and leucine having the highest. In Coleanthus subtilis, which also belongs to the legume family, leucine amino acid has the highest codon usage, while cysteine amino acid has the lowest codon usage56. In L. lamottei, one codon each encodes the amino acids methionine and tryptophan. On the other hand, the remaining amino acids are encoded by two to six codons. Similar codon usage results are observed in wild lentil and chickpea species20,50,51. Codon usage could be helpful for phylogenetic association research as codon use can influence the manner of gene mutation31.
For cp growth and photosynthesis, appropriate expression of cp genes is essential57. Therefore, the expression levels of cp genes in L. lamottei were investigated in this study. MELP values of protein-coding genes were obtained using ribosomal genes as reference. In particular, the MELP values of genes involved in photosynthesis were high. In addition, FPKM values were calculated using RNA-seq data to examine the gene expression of cp genes. We found high gene expression levels, especially for the ATP-synthase, rubisco, and photosynthesis-related genes were classified according to their biological functions. Plants synthesize ATP during electron and proton transport. These processes include photosystem I, photosystem II, cytochrome b6/f complex, and ATP synthase58. Also, the rbcL gene is another gene that is related to the photosystem59. Therefore, the expression levels of these genes are expected to be higher. We found that the expression levels of these genes were not positively correlated with the MELP value, which could be due to the highly increased expression of ATP synthase.
To assess the phylogenetic relationships of L. lamottei, the complete cp genomes of 14 plant species were used to construct a phylogenetic tree using the Jukes-Cantor model. We used A. thaliana as an outgroup to confirm the phylogeny study. For the phylogenetic analysis, the representative species were selected from three main subfamilies of the Fabaceae family: the Papilionoideae, Detarionoideae, and Cercidoideae subfamilies.In this study, results show that L. lamottei was closely related to L. culinaris and L. ervoides. In a previously published study, L. ervoides provided the same branching as L. culinaris20. Thus, these three Lens species are expected to be closer to each other than to other species. Therefore, it should present close branching with L. lamottei, and cicer species belonging to the IRLC family24,50,51. Therefore, as a result of the phylogenetic tree, these two species are expected to be closer to each other than the other species. C. cajan and G. max are members of the Papilionoideae family12,24. The other species are included in the Detarionoideae and Cercidoideae subfamilies60,61,62,63. Therefore, these species are expected to be more phylogenetically distant from L. culinaris in Fig. 7. The strong bootstrap and posterior probability scores in this study suggested that the whole cp genomes could be beneficial in determining the phylogenetic locations and relations of the Papilionoideae tribe for L. lamottei.
Methods
All the methods were carried out in accordance with relevant local/national/international guidelines and regulations.
Organelle isolation and whole chloroplast genome sequencing of L. lamottei
In collaboration with Akdeniz University (Antalya), L. lamottei seeds were obtained and genotyped for use in this study. It was registered by the Department of Field Crops herbarium at Akdeniz University, and identified by Prof. Dr. Cengiz Toker. It was deposited with the voucher ID of L. lamottei-01. Rows of seeds were planted in February 2020 and collected from Antalya (Turkey)(36.89722° N, 30.71414° E). Plants were grown at an average temperature of 12 °C. Plants were harvested when their leaves reached the full green stage. For 72 h, leave samples were stored at 4 °C to prevent starch accumulation. 20 g of fresh leaves were harvested and extracted from the cp genome according to the cp DNA isolation method published by Shi et al.64. DNA isolation purity was assessed using NanoDrop (Thermo Scientific) and agarose gel electrophoresis.
The cp genomic DNA of the plant was sequenced using the DNA nanoball technique at Beijing Genome Institute (BGI, Hong Kong, China). Briefly, the Agencourt AMPure XP-Medium kit was used. DNA fragments in the 200–400 bp range were chosen after random cpDNA fragmentation and purified the PCR products. With the help of a splint oligosequence, heat-denatured DNAs were converted into single-stranded circular DNA (ssCir DNA) for library preparation. These libraries were sequenced on the BGISEQ-500 platform using whole-genome sequencing. The ssCir DNA molecules were transformed into DNA nanoball (DNB). These DNBs were loaded into nanoarrays with a certain pattern using DNA nanochip technology. Paired-end 100-bp reads were obtained using combinatorial Probe-Anchor Synthesis (cPAS)65.
The sequencing qualities were assessed using FastQC66 utility. The de novo genome assembly tool GetOrganelle67, which was specifically designed for plastid genomes, was used. This pipeline uses the assembly algorithm Spades68 to construct contigs and scaffolds from raw sequencing reads. During the assembly process, the algorithm was fed with the L. culinaris (NC_027152.1) cp genome as a reference seed and the database of known plastid genomes ('embplant pt'). The constructed scaffolds were visually inspected using Bandage69 software to consider possible structural rearrangements. To inspect the assembly quality and read support, raw sequencing reads were aligned back to scaffolds using bowtie270 short read alignment software.
Gene annotations
The cp genome of L. culinaris, which was retrieved from NCBI with accession NC_027152, was used as a reference. MAUVE alignment was performed to analyze gene homology between the two species (L. lamottei and L. culinaris). Gene order, rearrangements, and structural changes were also identified71. The GeSeq72 annotation tool was used to determine the locations of protein-coding, tRNA, and rRNA genes. The annotation of tRNA genes was confirmed using tRNAscan-SE v.2.0.7 and ARAGORN v1.2.3872. To visualize the cp genome sequence map of L. lamottei, Organelle Genomes DRAW (OGDRAW)73 was used. The complete cp genome of L. lamottei was deposited in the European Nucleotide Archive (ENA) databases with the accession number ERS7635406.
Comparative genome analysis
Comparison of the cp genomes of the two species (L. lamottei and L. culinaris) was performed using mVISTA74 in Shuffle-LAGAN mode. Using pairwise genome alignment, the divergent areas between the cp genomes of L. lamottei and L. culinaris were identified and repetitive sequence regions detected by miropeats (version 2.02)75 were masked. Custom python scripts were used to determine the sequence single nucleotide substitutions and indels between the two species and their genome-wide hot spots using 1000-bp windows with 200-bp sliding steps. Using the canonical amino acid codon table, synonymous, and nonsynonymous substitutions for the genomic regions were identified. The online PREP-Cp76 tool was used to predict RNA editing sites of all protein-coding sequences of L. lamottei, with a cutoff value of 0.8.
Simple sequence repeats analysis
The REPuter77 program was used for the detection of the forward, reverse, palindromic, and complementary repeats in the L. lamottei cp genome with a minimum repeat size of 30 bp, sequence identity of more than 90%, and hamming distance of 3. Tandem Repeat Finder78 was used to identify tandem repeats. MISA79 was used for detecting simple sequence repeats within the cp genome. The parameters used were 10 for mononucleotide, 5 for dinucleotide, 4 for trinucleotide, and 3 for tetranucleotide, pentanucleotide, and hexanucleotide.
Codon usage bias
Functions from the "coRdon" R package (v1.1.3) were used to determine codon usage statistics in the protein-coding regions. Along with the fundamental codon usage frequencies, these statistics also include Measure Independent of Length Composition Value (MILC) and MELP80,81. Based on the concept of RSCU, the ratio of observed codon frequency to expected uniform codon usage values was calculated.
Gene expression analysis
Lens lamottei RNA sequencing data were retrieved from NCBI SRA using accession number PRJNA625627. The reads were mapped using STAR82 to the assembled cp genome of L. lamottei. The featureCounts83 program was used to quantify gene expression. FPKM84 normalization of expression values for protein-coding genes were calculated.
Phylogenetic analysis
Following multiple sequence alignment using MAFFT85, MEGA 1186 was used for the phylogenetic analysis of the aligned sequences. The Jukes–Cantor87 model was used to create a maximum likelihood tree for phylogenetic analysis of whole cp genomes of different species. A. thaliana was used as an outgroup in the construction of a phylogenetic tree based on the complete cpDNA sequences of 13 species from the Leguminosae subfamilies Papilionoideae (3), IRLC (6), Detarioideae (2), and Cercidoideae (2). Construction and calculation of the phylogenetic tree were performed with 1000 bootstraps. GenBank data of the species used were obtained from the NCBI database. The species that were used were L. culinaris (NC_027152.1), L. ervoides (OU707010.1), C. arietinum (NC_011163.1), C. echinospermum (OU707005.1), C. bijigum (OU707008.1), C. cajan (NC_031429.1), G. max (NC_007942.1), I. bijuga (NC_047336.1), C. hamsiana (NC_036743.1), C. glabra (NC_036762.1), T. esculentum (NC_067756.1), and A. thaliana (NC_000932.1).
Data availability
The whole chloroplast genome raw sequence of L. lamottei are available in the ENA (European Nucleoide Archive) of EMBL-EBI under the accession number PRJEB47534 and sample identification number ERS7635406.
References
Faris, M. A. I. E., Takruri, H. R. & Issa, A. Y. Role of lentils (Lens culinaris L.) in human health and nutrition: A review. Mediterr. J. Nutr. Metab. 6, 3–16. https://doi.org/10.1007/s12349-012-0109-8 (2013).
Vidal-Valverde, C. et al. New functional legume foods by germination: Effect on the nutritive value of beans, lentils and peas. Eur. Food Res. Technol. 215, 472–477 (2002).
Yadav, S. S., McNeil, D., & Stevenson, P. C. (eds.). Lentil: An Ancient Crop for Modern Times (Springer Science & Business Media, 2007).
Ogutcen, E., Ramsay, L., von Wettberg, E. B. & Bett, K. E. Capturing variation in Lens (Fabaceae): Development and utility of an exome capture array for lentil. Appl. Plant Sci. 6, e01165 (2018).
Catarino, S., Duarte, M. C., Costa, E., Carrero, P. G. & Romeiras, M. M. Conservation and sustainable use of the medicinal Leguminosae plants from Angola. PeerJ 7, 6736 (2019).
Dikshit, H. K. et al. Genetic diversity in lens species revealed by EST and genomic simple sequence repeat analysis. PLoS ONE 10, e0138101 (2015).
Sonnante, G., Galasso, I. & Pignone, D. ITS sequence analysis and phylogenetic inference in the genus Lens mill. Ann. Bot. 91, 49–54 (2003).
Czefranova, Z. Review of species of the genus Lens mill. Nov. Cyst. P1 Vas. 8, 197–204 (1971).
Tehseen Azhar, M. & Hussain Wani, S. Wild Germplasm for Genetic Improvement in Crop Plants (Academic Press, 2021).
Ferguson, M. E., Maxted, N., Van Slageren, M. & Robertson, L. D. A re-assessment of the taxonomy of Lens mill. (Leguminosae, Papilionoideae, Vicieae). Bot. J. Linn. Soc. 133(1), 41–59 (2000).
Tullu, A., Diederichsen, A., Suvorova, G. & Vandenberg, A. Genetic and genomic resources of lentil: Status, use and prospects. Plant Genet. Resour. Charact. Util. 9, 19–29 (2011).
Saski, C. et al. Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol. Biol. 59, 309–322 (2005).
Bausher, M. G., Singh, N. D., Lee, S. B., Jansen, R. K. & Daniell, H. The complete chloroplast genome sequence of Citrus sinensis (L.) Osbeck var ‘Ridge Pineapple’: Organization and phylogenetic relationships to other angiosperms. BMC Plant Biol. 6, 1–11 (2006).
Zha, X., Wang, X., Li, J., Gao, F. & Zhou, Y. Complete chloroplast genome of Sophora alopecuroides (Papilionoideae): Molecular structures, comparative genome analysis and phylogenetic analysis. J. Genet. 99, 1–4 (2020).
Abdullah, et al. Comparison of chloroplast genomes among species of unisexual and bisexual clades of the monocot family Araceae. Plants. 9, 1–16 (2020).
Asaf, S., Khan, A. L., Khan, A. & Al-Harrasi, A. Unraveling the chloroplast genomes of two prosopis species to identify its genomic information, comparative analyses and phylogenetic relationship. Int. J. Mol. Sci. 21, 3280 (2020).
Tan, W. et al. The complete chloroplast genome of Chinese medicine (Psoralea corylifolia): Molecular structures, barcoding and phylogenetic analysis. Plant Gene. 21, 100216 (2020).
Wang, M. et al. Comparative analysis of Asteraceae chloroplast genomes: Structural organization, RNA editing and evolution. Plant Mol. Biol. Rep. 33, 1526–1538 (2015).
Zhang, D. et al. Comparative and phylogenetic analysis of the complete chloroplast genomes of six Polygonatum species (Asparagaceae). Sci. Rep. https://doi.org/10.21203/rs.3.rs-1669614/v1 (2022).
Tayşi, N. et al. Complete chloroplast genome sequence of Lens ervoides and comparison to Lens culinaris. Sci. Rep. 12, 15068 (2022).
Liang, D., Wang, H., Zhang, J., Zhao, Y. & Wu, F. Complete chloroplast genome sequence of Fagus longipetiolata Seemen (Fagaceae): Genome structure, adaptive evolution, and phylogenetic relationships. Life. 12, 92 (2022).
Antunes, A. M. et al. The chloroplast genome sequence of Dipteryx alata Vog. (Fabaceae: Papilionoideae): Genomic features and comparative analysis with other legume genomes. Revista Brasileira de Botanica. 43, 271–282 (2020).
Jansen, R. K., Wojciechowski, M. F., Sanniyasi, E., Lee, S. B. & Daniell, H. Complete plastid genome sequence of the chickpea (Cicer arietinum) and the phylogenetic distribution of rps12 and clpP intron losses among legumes (Leguminosae). Mol. Phylogenet. Evol. 48, 1204–1217 (2008).
Kaila, T. et al. Chloroplast genome sequence of pigeonpea (Cajanus cajan (L.) Millspaugh) and Cajanus scarabaeoides (L.) thouars: Genome organization and comparison with other legumes. Front. Plant Sci. 7, 1847 (2016).
Xia, X. et al. Comparison of chloroplast genomes of compound-leaved maples and phylogenetic inference with other Acer species. Tree Genet. Genomes. 18, 11 (2022).
Feng, J. et al. Analysis of complete chloroplast genome: Structure, phylogenetic relationships of Galega orientalis and evolutionary inference of Galegeae. Genes (Basel). 14, 176 (2023).
Jiang, D. et al. Complete chloroplast genomes provide insights into evolution and phylogeny of Zingiber (Zingiberaceae). BMC Genom. 24, 30 (2023).
Zhou, T. et al. Comparative transcriptome and chloroplast genome analyses of two related Dipteronia species. Front. Plant Sci. 7, 1512 (2016).
Parvathy, S. T., Udayasuriyan, V. & Bhadana, V. Codon usage bias. Mol. Biol. Rep. 49, 539–565. https://doi.org/10.1007/s11033-021-06749-4 (2022).
Somaratne, Y., Guan, D. L., Wang, W. Q., Zhao, L. & Xu, S. Q. The complete chloroplast genomes of two Lespedeza species: Insights into codon usage bias, rna editing sites, and phylogenetic relationships in desmodieae (Fabaceae: Papilionoideae). Plants. 9, 51 (2020).
Yu, T. et al. Analysis of the chloroplast genomes of four Pinus species in Northeast China: Insights into hybrid speciation and identification of DNA molecular markers. J. For. Res. (Harbin). 33, 1881–1890 (2022).
Dissanayake, R., Braich, S., Cogan, N. O., Smith, K. & Kaur, S. Characterization of genetic and allelic diversity amongst cultivated and wild lentil accessions for germplasm enhancement. Front. Genet. 11, 546 (2020).
Park, J. H., Lee, M., Lee, Y. & Lee, J. The complete chloroplast genome of Viola grypoceras (Violaceae). Mitochondrial DNA B Resour. 8, 42–44 (2023).
Wang, L. et al. The comparative studies of complete chloroplast genomes in Actinidia (Actinidiaceae): Novel insights into heterogenous variation, clpP gene annotation and phylogenetic relationships. Mol. Genet. Genom. 297, 535–551 (2022).
Wang, W. & Lanfear, R. Long-reads reveal that the chloroplast genome exists in two distinct version in most plants. Genome Biol. Evol. 11, 3372–3381 (2019).
Lin, Z. et al. Comparative analysis of chloroplast genomes in Vasconcellea pubescens A.DC. and Carica papaya L. Sci. Rep. 10, 15799 (2020).
Tang, Y. et al. New insights into the evolutionary characteristic between the New World and Old World Lupinus species using complete chloroplast genomes. All Life. 14, 414–427 (2021).
Tavares, J. C. C., Achakkagari, S. R., Archambault, A. & Strömvik, M. V. The plastome of Arctic Oxytropis arctobia (Fabaceae) is significantly different from that of O. splendens and other related species. Genome 65, 301–313 (2022).
Duan, H. et al. Complete chloroplast genome of Calligonum mongolicum Turcz. and comparative analysis with other Calligonum species. J. Appl. Res. Med. Aromat. Plants. 27, 100370 (2022).
Hao, Z. et al. The complete chloroplast genome sequence of a relict conifer Glyptostrobus pensilis: Comparative analysis and insights into dynamics of chloroplast genome rearrangement in cupressophytes and pinaceae. PLoS ONE 11, e0161809 (2016).
Nawae, W. et al. The genome and transcriptome analysis of the Vigna mungo chloroplast. Plants. 9, 1–17 (2020).
Moghaddam, M., Ohta, A., Shimizu, M., Terauchi, R. & Kazempour-Osaloo, S. The complete chloroplast genome of Onobrychis gaubae (Fabaceae-Papilionoideae): Comparative analysis with related IR-lacking clade species. BMC Plant Biol. 22, 75 (2022).
Wang, X., Dorjee, T., Chen, Y., Gao, F. & Zhou, Y. The complete chloroplast genome sequencing analysis revealed an unusual IRs reduction in three species of subfamily Zygophylloideae. PLoS ONE 17, e0263253 (2022).
Dugas, D. V. et al. Mimosoid legume plastome evolution: IR expansion, tandem repeat expansions, and accelerated rate of evolution in clpP. Sci. Rep. 5, 16958 (2015).
Zhou, T. et al. Complete chloroplast genome of Ilex dabieshanensis: Genome structure, comparative analyses with three traditional Ilex tea species, and its phylogenetic relationships within the family Aquifoliaceae. PLoS ONE 17, e0268679 (2022).
Hong, Z. et al. Comparative Analyses of 35 complete chloroplast genomes from the genus Dalbergia (Fabaceae) and the identification of DNA barcodes for tracking illegal logging and counterfeit rosewood. Forests 13, 626 (2022).
Wu, S. et al. Extensive genomic rearrangements mediated by repetitive sequences in plastomes of Medicago and its relatives. BMC Plant Biol. 21, 1–6 (2021).
Yang, X. et al. Structural characterization and comparative analysis of the chloroplast genome of Ginkgo biloba and other gymnosperms. J. For. Res. (Harbin). 32, 765–778 (2021).
Wu, H. Y. et al. Comparative analysis of chloroplast genomes of Dalbergia species for identification and phylogenetic analysis. Plants. 11, 1109 (2022).
Mehmetoglu, E., Kaymaz, Y., Ates, D., Kahraman, A. & Tanyolac, M. B. The complete chloroplast genome sequence of Cicer echinospermum, genome organization and comparison with related species. Sci. Hortic. 296, 110912 (2022).
Temel, M., Kaymaz, Y., Ateş, D., Kahraman, A. & Tanyolaç, M. B. The Complete Chloroplast genome sequence of Cicer bijugum, genome organization, and comparison with related species. Curr. Genom. 23, 50–65 (2022).
Cay, S. B. et al. Genome skimming approach reveals the gene arrangements in the chloroplast genomes of the highly endangered Crocus L. species: Crocus istanbulensis (B. Mathew) Rukšāns. PLoS ONE 17, e0269747 (2022).
Li, B., Lin, F., Huang, P., Guo, W. & Zheng, Y. Complete chloroplast genome sequence of Decaisnea insignis: Genome organization, genomic resources and comparative analysis. Sci. Rep. 7, 10073 (2017).
Liu, X., Xu, D., Hong, Z., Zhang, N. & Cui, Z. Comparative and phylogenetic analysis of the complete chloroplast genome of santalum (Santalaceae). Forests 12, 1303 (2021).
Jiao, Y. et al. Complete chloroplast genomes of 14 subspecies of D. glomerata: Phylogenetic and comparative genomic analyses. Genes (Basel). 13, 1621 (2022).
Ren, T. et al. Complete chloroplast genome of Cnidium monnieri (Apiaceae) and comparisons with other tribe selineae species. Diversity (Basel). 14, 323 (2022).
Zhang, Y., Zhang, A., Li, X. & Lu, C. The role of chloroplast gene expression in plant responses to environmental stress. Int. J. Mol. Sci. 21, 1–16. https://doi.org/10.3390/ijms21176082 (2020).
Gomes Pacheco, T. et al. The complete plastome of Passiflora cirrhiflora A. Juss.: Structural features, RNA editing sites, hotspots of nucleotide diversity and molecular markers within the subgenus Deidamioides. Revista Brasileira de Botanica. 43, 839–853 (2020).
Yan, L. et al. Analyses of the complete genome and gene expression of chloroplast of sweet potato [Ipomoea batata]. PLoS ONE 10, e0124083 (2015).
Wang, Y. H., Wang, H., Yi, T. S. & Wang, Y. H. The complete chloroplast genomes of Adenolobus garipensis and Cercis glabra (Cercidoideae, Fabaceae). Conserv. Genet. Resour. 9, 635–638 (2017).
Tosso, F. et al. Evolution in the Amphi-Atlantic tropical genus Guibourtia (Fabaceae, Detarioideae), combining NGS phylogeny and morphology. Mol. Phylogenet. Evol. 120, 83–93 (2018).
Kim, Y. & Cullis, C. A novel inversion in the chloroplast genome of marama (Tylosema esculentum). J. Exp. Bot. 68, 2065–2072 (2017).
U-thoomporn, S. et al. The complete chloroplast genome sequence of Intsia bijuga (Colebr.) Kuntze (Fabaceae: Detaroideae: Afzelieae). Mitochondrial DNA B Resour. 7, 1814–1816 (2022).
Shi, C. et al. An improved chloroplast DNA extraction procedure for whole plastid genome sequencing. PLoS ONE 7, e31468 (2012).
Fehlmann, T. et al. cPAS-based sequencing on the BGISEQ-500 to explore small non-coding RNAs. Clin. Epigenet. 8, 1–11 (2016).
Andrews, S. FastQC: A quality control tool for high throughout sequence data. (2010).
Jin, J. J. et al. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 1–31 (2020).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Wick, R. R., Schultz, M. B., Zobel, J. & Holt, K. E. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 31, 3350–3352 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9, 357–359 (2012).
Darling, A. C. E., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403 (2004).
Tillich, M. et al. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 45, W6–W11 (2017).
Greiner, S., Lehwark, P. & Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 47, W59–W64 (2019).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004).
Parsons, J. D. Miropeats: graphical DNA sequence comparisons. CABIOS. 11 https://academic.oup.com/bioinformatics/article/11/6/615/187092 (1995).
Mower, J. P. The PREP suite: Predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 37, W253–W259 (2009).
Kurtz, S. et al. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 29, 4633–4642 (2001).
Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Supek, F. & Vlahoviček, K. Comparison of codon usage measures and their applicability in prediction of microbial gene expressivity. BMC Bioinform. 6, 463 (2005).
Elek, A., Kuzman, M. & Vlahovicek, K. Package “coRdon” type package title codon usage analysis and prediction of gene expressivity. (2022).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis cersion 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Jukes, T. H. & Cantor, C. R. Evolution of protein molecules. In Mammalian Protein Metabolism (ed. Munro, H. N.) 21–132 (Academic Press, 1969).
Acknowledgements
This current study was funded by Ege University Scientific Research Projects Coordinatorship (EGE-BAP) with the project number FOA-2020-20981. We also thank Coordination of Organizational Development and Directorate of Library and Documentation for their support in editing and proofreading of this study. We are also greatly acknowledged to Prof Cengiz Toker who identified and provide the L. lamottei species.
Author information
Authors and Affiliations
Contributions
S.K.: data analyses, writing manuscript, Y.K.: data interpretation, writing and editing manuscript, D.A.: experimental analysis of chloroplast isolation, M.B.T.: conceived the research, project supervisor, editing-review original draft. All the authors have approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurt, S., Kaymaz, Y., Ateş, D. et al. Complete chloroplast genome of Lens lamottei reveals intraspecies variation among with Lens culinaris. Sci Rep 13, 14959 (2023). https://doi.org/10.1038/s41598-023-41287-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-41287-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.