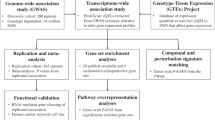
Abstract
Anthracycline-induced cardiomyopathy is a leading cause of late morbidity in childhood cancer survivors. Aberrant DNA methylation plays a role in de novo cardiovascular disease. Epigenetic processes could play a role in anthracycline-induced cardiomyopathy but remain unstudied. We sought to examine if genome-wide differential methylation at ‘CpG’ sites in peripheral blood DNA is associated with anthracycline-induced cardiomyopathy. This report used participants from a matched case–control study; 52 non-Hispanic White, anthracycline-exposed childhood cancer survivors with cardiomyopathy were matched 1:1 with 52 survivors with no cardiomyopathy. Paired ChAMP (Chip Analysis Methylation Pipeline) with integrated reference-based deconvolution of adult peripheral blood DNA methylation was used to analyze data from Illumina HumanMethylation EPIC BeadChip arrays. An epigenome-wide association study (EWAS) was performed, and the model was adjusted for GrimAge, sex, interaction terms of age at enrollment, chest radiation, age at diagnosis squared, and cardiovascular risk factors (CVRFs: diabetes, hypertension, dyslipidemia). Prioritized genes were functionally validated by gene knockout in human induced pluripotent stem cell cardiomyocytes (hiPSC-CMs) using CRISPR/Cas9 technology. DNA-methylation EPIC array analyses identified 32 differentially methylated probes (DMP: 15 hyper-methylated and 17 hypo-methylated probes) that overlap with 23 genes and 9 intergenic regions. Three hundred and fifty-four differential methylated regions (DMRs) were also identified. Several of these genes are associated with cardiac dysfunction. Knockout of genes EXO6CB, FCHSD2, NIPAL2, and SYNPO2 in hiPSC-CMs increased sensitivity to doxorubicin. In addition, EWAS analysis identified hypo-methylation of probe ‘cg15939386’ in gene RORA to be significantly associated with anthracycline-induced cardiomyopathy. In this genome-wide DNA methylation profile study, we observed significant differences in DNA methylation at the CpG level between anthracycline-exposed childhood cancer survivors with and without cardiomyopathy, implicating differential DNA methylation of certain genes could play a role in pathogenesis of anthracycline-induced cardiomyopathy.
Similar content being viewed by others
Introduction
Anthracycline-induced cardiomyopathy is a leading cause of premature death in children with cancer1. The risk for cardiomyopathy increases with anthracycline dose2,3. Other risk factors include young age (< 5 years) at anthracycline exposure, female sex, chest radiation and presence of cardiovascular risk factors (CVRFs: diabetes, hypertension, dyslipidemia)4,5. The considerable inter-individual variability in the dose-dependent association between anthracycline exposure and cardiomyopathy risk suggest a need for a better understanding of the underlying molecular mechanisms. Several studies have examined the role of single nucleotide variants (SNVs) associated with anthracycline-induced cardiomyopathy6,7,8. There is emerging evidence to suggest that anthracycline-induced myocardial injury may be related to oxidative stress, mitochondrial damage, cardiomyocyte apoptosis, necrosis and autophagy9. However, there is paucity of information regarding the functional significance of most SNVs, precluding a clear understanding of the pathogenesis of anthracycline-induced cardiomyopathy, as well as hindering the development of potential preventive or therapeutic strategies.
Phenotypic traits are determined not only by genetic variants of coding sequences, but also by mechanisms that regulate how genes are expressed. Epigenetic modifications (DNA methylation, histone modifications and RNA-associated gene silencing)10 regulate gene expression, thereby contributing to the functional state of the genome. Indeed, aberrant DNA methylation plays a role in human disease, including de novo cardiovascular disease11,12,13,14,15,16. Epigenome-wide association studies (EWAS) have identified regions harboring variation in DNA methylation associated with disease phenotypes17. Recently, Robinson et al.18 showed that there is genome wide loss of DNA methylation and a gain in hydroxymethylation in endomyocardial biopsies from patients with early and late cardiotoxicity due to anthracyclines. While there is some evidence of an association between microRNAs and anthracycline-induced cardiomyopathy19,20, the contribution of epigenetic changes remains largely unexplored. In the present study, we examined the peripheral blood methylome at single CpG resolution in anthracycline-exposed childhood cancer survivors with and without cardiomyopathy to address this gap.
Methods
Study design and population
Study used a matched case–control design to understand the pathogenesis of cardiomyopathy in childhood cancer survivors. Participants were enrolled to a Children’s Oncology Group (COG) study ALTE03N1 (Key Adverse Events after Childhood Cancer). Thirty-one COG institutions (see supplement for full names) contributed participants to the study after obtaining approval from local institutional review boards. Written informed consent/assent was obtained from patients and/or parents/legal guardians. The University of Alabama at Birmingham Institutional Review Board (IRB-150115006) approved all experimental protocols and methods. All methods were performed in accordance with the ethical standards of University of Alabama at Birmingham Institutional Review Board and with the 1964 Helsinki Declaration. Cases consisted of childhood cancer survivors who developed cardiomyopathy. For each case, patients with no signs or symptoms of cardiomyopathy were randomly selected as controls from the same childhood cancer survivor cohort, matched on primary cancer diagnosis, year of diagnosis (± 5 years) and race/ethnicity. The selected controls also needed to have a longer duration of cardiomyopathy-free follow-up compared with time from cancer diagnosis to cardiomyopathy for the corresponding case. Participants provided peripheral blood in K2EDTA tubes for germline DNA. For this report, we restricted the participants to anthracycline-exposed non-Hispanic White patients to ensure homogeneous populations of sufficient size.
Cases fulfilled American Heart Association criteria for cardiac compromise by presenting with signs/symptoms (dyspnea, orthopnea, fatigue, edema, hepatomegaly and/or rales); or, in the absence of signs/symptoms, had echocardiographic features of left ventricular dysfunction [ejection fraction (EF) ≤ 40% and/or fractional shortening (SF) ≤ 28%]. Lifetime anthracycline exposure was calculated by multiplying the cumulative dose (mg/m2) of individual anthracyclines by a factor that reflects the drug’s cardiotoxic potential21 and then summing the results. Exposure to chest radiation was treated as a yes/no variable. Cardiovascular Risk factors (CVRFs) included the presence of any of the following: diabetes, hypertension, or dyslipidemia.
Genome-wide DNA methylation analysis
DNA methylation status of 850,000 CpG (5'–C–phosphate–G–3') sites across the whole genome was analyzed using the Illumina HumanMethylation EPIC BeadChip arrays (Supplementary methods). The methylation score for each CpG was represented as a ‘β’ value according to the fluorescent intensity ratio [range: 0 (non-methylated) to 1 (completely methylated)]. Raw intensity data (IDAT files) were assessed for quality using BeadArray Controls Reporter (Illumina). Probe location and gene annotation used Illumina reference files (GRCh37/hg19).
Raw IDAT files were analyzed in R (v4.0.2, https://www.r-project.org/), using the Bioconductor packages minfi (v.1.34.0) and ChAMP (Chip Analysis Methylation Pipeline, v.2.18.3) for EPIC arrays22,23. Initial probe filtering steps used a threshold of p > 0.01 to remove 8511 probes with poor detection values. Next, we removed 2927 probes represented by < 3 beads, 2829 non-CpG probes, 95,316 probes that overlapped with SNVs, 24 probes that mapped to multiple locations and 16,593 probes that were on X or Y chromosomes24. Data were normalized using Noob25 for within-array normalization, correcting for background fluorescence and dye bias (Supplementary Fig. 1A). To account for DNA methylation differences due to cell-type composition in whole blood, we performed reference based deconvolution using FlowSorted.Blood.EPIC (v.1.6.1)26 (Supplementary Fig. 1B), and corrected the beta matrix in ChAMP. Following this, the singular value decomposition method (SVD) was used to detect significant components of variation, including valid covariates for statistical analysis. Champ.runCombat27 was used to eliminate technical and biological sources of variation from normalized and cell-type corrected methylation data (Supplementary Figs. 2 and 3). Following QC, 740,636 probes were analyzed in 104 samples (52 cases; 52 matched controls). The epigenetic age estimates, including Horvath’s28, Hannum’s29, DNAmAge, PhenoAge30 and GrimAge31 were obtained using an online calculator (https://dnamage.genetics.ucla.edu/new)28.
Differential methylation analysis
Paired differentially-methylated probe (DMP) and differentially-methylated region (DMR) analyses were performed in ChAMP23, to identify differences in methylation that distinguished cardiomyopathy cases from matched controls. Default Bonferroni-corrected alpha threshold was set to < 0.05 in ChAMP.
DAVID enrichment analysis
To further investigate the function of DMPs, we performed gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis using DAVID bioinformatics resource tool. Terms with kappa value > 0.5 were considered significantly enriched.
Epigenome-wide association study (EWAS)
EWAS was performed to identify the association between DNA methylation at CpG loci and anthracycline-induced cardiomyopathy. Normalized, cell composition- and batch-effect-corrected ‘β’ values were used as the dependent variable. The following covariates were examined: age at primary cancer diagnosis, sex, age at study, anthracycline dose, chest radiation, CVRFs, DNAmGrimAge and PhenoAge. Collinearity statistics was used to filter covariates with a correlation coefficient > 0.7 (Supplementary Fig. 4). The following covariates were retained in the EWAS model: DNAmGrimAge, sex*age at study, chest radiation, age at primary cancer diagnosis squared and matching pair identifier. A series of linear regression models were built, one CpG locus at a time, with methylation β-values (0–1) as the dependent variable32 and cardiomyopathy case–control status as the key independent variable. Probes with a p-value < 10–8 were considered epigenome-wide significant associations. A Manhattan plot was generated for visualization of EWAS results (CMplot R package). We also used ‘CpGassoc’ R package to verify results obtained by the EWAS model33.
Functional analyses
Criteria for prioritizing candidate genes for functional analyses
We used the following criteria to prioritize genes for functional analyses: (i) significant association of CpG probes with cardiomyopathy in paired ChAMP analysis (DMP, DMR) and EWAS, (ii) expression of genes harboring significant probes in adult human heart and hiPSC-CMs34,35, and (iii) mechanistic plausibility and published association with cardiomyopathy.
hiPSC line 19c3 generated from peripheral blood mononuclear cells from a healthy individual using the CytoTune-iPS 2.0 Sendai Reprogramming Kit (Invitrogen, A1651836) was used (Supplementary methods). To generate gene knockout gRNA expression vectors, one to two gRNAs targeting all splicing variants of the targeted genes were designed using an online CRISPR design tool (IDT) with high predicted on-target score and minimal predicted off-target effect. DNA oligos (IDT) encoding each gRNA with BbsI ligation overhangs were annealed and inserted into the BbsI restriction site of a pSpCas9(BB)-2A-Puro (PX459, Addgene 62988) plasmid (Supplementary Table 1A). The constructed gRNA expression plasmids were confirmed by Sanger sequencing (Eurofins) with the LKO1_5_primer (5′-GACTATCATATGCTTACCG-3′). Supplementary Table 1B includes sgRNA oligo sequences and sequencing primers. Details of the CRISPR/Cas9-mediated knockout of candidate genes are summarized in the Supplementary methods. We used qRT-PCR to assess candidate gene knockout (Supplementary methods and Supplementary Fig. 5). All PCR reactions were performed in triplicates (Supplementary Fig. 6) in a 384-well plate format using TaqMan Gene Expression Master Mix (Applied Biosystems, 4444557) in a QuantStudio 5 Real-Time PCR System (Applied Biosystems, A28140). Supplementary Table 2 summarizes TaqMan probes. Differentiation into cardiomyocytes was completed by using a hiPSC line expressing an exogenous TNNT2 promoter-driven Zeocin antibiotic selection resistance cassette for cardiomyocyte purification37 (Supplementary methods).
Day 30 hiPSC-CMs were treated for 72 h with doxorubicin (0.01–100 μM). Cell viability was assessed after 72 h using a resazurin assay. Fluorescence was measured using a VarioSkan Lux Multi-Mode Reader (Thermo Scientific). Data were analyzed using Excel and graphed using Prism 7.0 software (GraphPad) depicting standard dose–response guidelines. Data were presented as mean ± SEM. Comparisons were conducted via one way-ANOVA test, an unpaired two-tailed Student’s t-test, or F-test.
Results
Demographic and clinical characteristics
The case–control set included 104 non-Hispanic White, anthracycline-exposed childhood cancer survivors (52 cases; 52 matched controls). As shown in Table 1, the median age at primary cancer diagnosis for the cases and controls was 7.5 years and 10.7 years, respectively. Cases received a higher cumulative anthracycline dose (308 mg/m2 vs. 253 mg/m2, P = 0.04) and were more likely to have CVRFs (44.2% vs. 7.7%, P < 0.001). The median time between cancer diagnosis and cardiomyopathy was 6.3 years; controls were followed for a significantly longer period (median, 11.9 years, P < 0.001).
Differentially-methylated probes
Of the 5,500 probes with an adjusted P-value of < 0.05, 32 probes showed an absolute difference in β-values ≥ ± 0.05 (Table 2 and Supplementary Excel, Table A); 22 of the 32 probes were associated with known genes. Of the 22 genes 17 (77%) are linked to heart diseases (Table 2). Ten genes (SLC18A2/VMAT2, PDXK, SPTBN4, SYNPO2/CHAP, SLC9A2/NHE2, AHRR, GPR139, GNAO1, TRABD2A/TIKI1 and MAP9) harbored hypermethylated (in cases compared to controls) CpGs and twelve genes (NIPAL2/SLC57A4, OR4D10, FARP1/PLEKHC2, ZFAND6, TEX9, SLMAP, HDAC9, POU6F2, RAPGEF6, FCHSD2, PLEKHN1 and EXOC6B) harbored hypomethylated (in cases compared to controls) CpG probes.
Of the 22 known genes that harbored DMPs, EXOC6B, FCHSD2, NIPAL2, SYNPO2, PDXK, and SLMAP are expressed in the adult heart tissue and hiPSC-CMs38. We examined whether the loss of function of these genes altered sensitivity to doxorubicin in an isogenic hiPSC line (ISO). Cell viability assays showed that the ISO-EXOC6B KO (LD50 = 0.18 μM), ISO-FCHSD2 ko (LD50 = 0.67 μM), ISO-NIPAL2 ko (LD50 = 0.97 μM), and ISO-SYNPO2 ko (LD50 = 1.45 μM) hiPSC-CMs were 24-, 6.5-, 4.5-, and 3-fold more sensitive to doxorubicin as compared to ISO (LD50 = 4.40 μM, P < 0.0001), respectively (Fig. 1). PDXK, SLMAP and RORA KO- hiPSC-CMs did not show significant sensitivity to doxorubicin compared to the isogenic line.
The top hypermethylated (cg15417294) and hypomethylated (cg23280506) probes show a Δβ of 0.09 and −0.08, respectively (Table 2 and Supplementary Fig. 7). cg15417294 is associated with SLC18A2/VMAT2 (Solute Carrier Family 18 Member A2/Vesicular Monoamine Transporter), whereas cg23280506 is located in the intergenic region and overlaps with a CpG island.
DAVID analysis showed a GADD_DISEASE_CLASS representation of “cardiovascular”, “chemodependency” and “metabolic” terms for genes that overlap with DMPs (Supplementary Excel, Table B).
Differentially-methylated regions
Using the default P-value threshold of 0.05, we identified 354 paired DMRs (spanning 3 to 1396 base pairs) distributed over 1520 probes (Supplementary Excel, Table C); four probes were classified as DMPs in the above analysis. Overall, 33 DMRs were located on chromosome 1, 29 on chromosome 12, 27 on chromosome 6, and 23 each on chromosomes 2 and 5. The DMRs represented 323 annotated genes, of which 3 genes were included among the DMP genes above. The top two significant DMR-associated genes HS3ST3B1 and PNPO/SP2-AS1 are shown in Table 3 and Supplementary Fig. 8. Paired DMR_1 (chr17:14206572–14207968) was hypomethylated in cases and Paired DMR_2 (chr17:46018654–46019184) was hypermethylated in cases. Knockout of PNPO resulted in defective differentiation of hiPSCs to cardiomyocytes, suggesting a likely critical role of PNPO in cardiomyocyte differentiation and maturation.
Epigenome wide association analysis
As shown in Fig. 2, two CpG probes exceeded the epigenome-wide significance threshold (P < 1 × 10–8); both were located in ‘open sea’. cg15939386 (P = 5.32 × 10–9) is located in the intron of RORA (Retinoic acid-related orphan receptor α) (Supplementary Fig. 9) and overlaps with distal enhancer-like signature EH38E1766996. The probe was hypomethylated in cases and had the smallest P-value 2.8 × 10–8 in the DMP analysis but showed a Δβ of −0.008 (cutoff for DMPs was ± 0.05). cg26610307 (P = 9.70 × 10–9) is in the X450k enhancer region, but does not overlap with any known genes.
Discussion
We identified ten genes with hypermethylated DMPs and twelve genes with hypomethylated DMPs in childhood cancer survivors with anthracycline-induced cardiomyopathy when compared with survivors without. Seventy seven percent of these genes are linked to heart disease. Knockouts in hiPSC-CMs of four of these genes (EXOC6B, FCHSD2, NIPAL2, and SYNPO2) demonstrated significantly increased sensitivity to doxorubicin. We identified 354 DMRs; HS3ST3B1 and PNPO/SP2-AS1 ranked as the top two DMR-associated genes. In the EWAS, we identified cg15939386 in the RORA gene to be significantly associated with anthracycline-induced cardiomyopathy.
Knockouts of DMP-harboring genes EXOC6B, FCHSD2, NIPAL2 and SYNPO2 in hiPSC-CMs demonstrated significant sensitivity to doxorubicin, suggesting that these genes play a role in protecting cardiomyocytes from anthracycline-induced toxicity. Knockout of EXOC6B in hiPSC-CMs showed the highest (24-fold) sensitivity to doxorubicin compared to the control. EXOC6B encodes for the evolutionarily conserved exocyst, a multimeric protein complex necessary for exocytosis. It also encodes for a circular RNA cEXOC6B. Circular RNAs (circRNAs) are a class of long non-coding RNAs that play a role in cardiac hypertrophy, acute myocardial infarction, cardiac cell senescence, diabetic cardiomyopathy and heart failure39,40,41,42,43. Recently, a patent filed in 2018 (United States Patent Application 20200188356) showed that cEXOC6B was significantly dysregulated in left ventricular tissue in subjects with failing hearts compared to subjects with non-failing hearts. Although not well studied, there is evidence to suggest circRNAs can regulate gene expression by controlling methylation and that aberrant DNA methylation might regulate circRNA expression44,45. FCHSD2 encodes a protein termed Carom that regulates cell growth, migration and adhesion46. FCHSD2 also regulates F-actin polymerization, suggesting involvement in insulin exocytosis47. The locus has been associated with systemic lupus erythematosus48 which is frequently complicated by aggressive atherosclerosis. NIPAL2/SLC57A4 is predicted to be involved in magnesium ion transport. SYNPO2/Myopodin is an actin- and α-actinin-binding member of the synaptopodin family49. It is localized in the sarcomeric Z-disc, but shuttles to the nucleus in cardiomyocytes in a stress- and differentiation- dependent fashion50,51. Pyle et al. proposed that complexes within the sarcomeric Z-disc drive downstream events in response to mechanical load of the heart, leading to cardiac hypertrophy52.
The top two DMR-associated genes were HS3ST3B1 and PNPO. HS3ST3B1 is downregulated in rybp−/−mice embryonic stem cells (ESC)53. Ring1 and Yy1 binding protein (Rybp) is a critical regulator of heart development and rybp null mice ESCs do not form normally functioning cardiomyocytes. In the rybp null cardiomyocytes, gene expression profiles revealed a downregulation of cardiac terminal markers. PNPO (pyridoxamine 5'-phosphate oxidase) is involved in the vitamin B6 metabolic pathway and is crucial to heart development during embryogenesis54. Zebrafish Pnpo morphants display a defective circulatory system55,56. Interestingly, cg19815989, a significantly hyper-methylated DMP in our analysis is on gene PDXK (pyridoxal kinase), also a key enzyme in vitamin B6 metabolism. Vitamin B6 is a required co-factor for enzymes involved in homocysteine metabolism. High plasma levels of homocysteine increase the risk of cardiovascular disease and stroke. Both PDXK (DMP) and PNPO (DMR) are hypermethylated in cases, possibly resulting in lower expression of these key genes with consequent aberrant vitamin B6 metabolism and increased risk of cardiovascular disease57. However, PDXK KO did not demonstrate sensitivity to doxorubicin in our study. Knockout of PNPO was incompatible with hiPSC differentiation to cardiomyocytes, indicating that this gene has a fundamental role in cardiac differentiation in human cells.
The CpG ‘cg15939386’ on RORA was significantly associated with cardiomyopathy in our EWAS model. RORA, also known as the nuclear melatonin receptor, plays an important role in the regulation of circadian rhythm, and protects against angiotensin II-induced cardiac hypertrophy and heart failure58. RORA is an endogenous protective receptor against diabetic cardiomyopathy by inhibiting oxidative stress and apoptosis in mouse models59. RORA protein is robustly expressed in non-failing human ventricular myocardium but is decreased in heart failure tissues, suggesting that RORA is a cardioprotective nuclear receptor60. lncRNA RORA-AS1 (RORA Antisense RNA 1) has a regulatory role in the expression of RORA. The hypomethylated probe ‘cg15939386’ is present in an intron that overlaps both with RORA (‘−’ strand) and RORA-AS1 (‘+’ strand). In our study, hiPSC-CM knockout for RORA, however, was not differentially sensitive to doxorubicin compared to isogenic cells.
We need to consider the findings in the context of certain limitations. We did not consider other factors that affect methylation, such as socioeconomic status, health behaviors and environmental exposures. Our study focused only on non-Hispanic White survivors of childhood cancer. We performed methylation analysis in blood and not cardiac tissue. There is concern regarding the use of blood in epigenetic investigations in that it contains multiple cell types, each having a characteristic methylation profile. However, we performed a reference-based deconvolution to correct for proportions of different cell types in peripheral blood. DNA methylation patterns are often tissue-specific, and hence the concern that the peripheral blood may not reflect the methylation pattern in the heart. However, use of peripheral blood as a surrogate for cardiac tissue, has practical significance as ‘disease-affected tissues’ are not readily accessible for sampling. Meder et al.61 report cross-tissue conservation of epigenetic patterns occurring during heart failure. DNA methylation is thought to control transcriptional programs in a time-specific manner. We obtained samples from cases after they had developed cardiomyopathy. For future studies, it will be important to systematically evaluate DNA methylation markers in longitudinal cohorts of anthracycline-induced cardiomyopathy and heart failure.
Conclusions
We performed the first epigenome wide analysis in peripheral blood derived DNA of childhood cancer survivors, with and without cardiomyopathy, following anthracyline exposure. We identified differentially methylated CpG sites and regions that are associated with genes that have previously been implicated in the pathogenesis of cardiovascular diseases and novel biologically relevant targets. Sensitivity to doxorubicin in hiPSC-CMs carrying the gene knock-outs of EXOC6B, FCHSD2, NIPAL2, SYNPO2 need further investigation as potential mechanistic and/or therapeutic targets.
Data availability
The data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus and are accessible through GEO Series accession number GSE224359 https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE224359.
Code availability
Scripts to perform reference-based deconvolution of whole-blood biospecimens using FlowSorted.Blood.EPIC package and correct the beta matrix in ChAMP have been deposited in a public GitHub repository and are available with the identifier https://github.com/lucylz/methylation.
Abbreviations
- CVRF:
-
Cardiovascular risk factors
- EWAS:
-
Epigenome-wide association study
- hiPSC-CM:
-
Human induced pluripotent stem cell-derived cardiomyocyte
- KO:
-
Knockout
- ISO:
-
Isotype
- LD50 :
-
Lethal dose 50
References
Armstrong, G. T. et al. Late mortality among 5-year survivors of childhood cancer: A summary from the Childhood Cancer Survivor Study. J. Clin. Oncol. 27, 2328–2338. https://doi.org/10.1200/JCO.2008.21.1425 (2009).
Lipshultz, S. E. et al. Chronic progressive cardiac dysfunction years after doxorubicin therapy for childhood acute lymphoblastic leukemia. J. Clin. Oncol. 23, 2629–2636. https://doi.org/10.1200/JCO.2005.12.121 (2005).
Adams, M. J. & Lipshultz, S. E. Pathophysiology of anthracycline- and radiation-associated cardiomyopathies: Implications for screening and prevention. Pediatr. Blood Cancer 44, 600–606. https://doi.org/10.1002/pbc.20352 (2005).
Armenian, S. H. et al. Cardiovascular disease in survivors of childhood cancer: Insights into epidemiology, pathophysiology, and prevention. J. Clin. Oncol. 36, 2135–2144. https://doi.org/10.1200/JCO.2017.76.3920 (2018).
Olsen, M. et al. Cardiovascular disease risk in childhood cancer survivors. Am. J. Epidemiol. 180, 120–123. https://doi.org/10.1093/aje/kwu144 (2014).
Leong, S. L., Chaiyakunapruk, N. & Lee, S. W. Candidate gene association studies of anthracycline-induced cardiotoxicity: A systematic review and meta-analysis. Sci. Rep. 7, 39. https://doi.org/10.1038/s41598-017-00075-1 (2017).
Aminkeng, F. et al. Recommendations for genetic testing to reduce the incidence of anthracycline-induced cardiotoxicity. Br. J. Clin. Pharmacol. 82, 683–695. https://doi.org/10.1111/bcp.13008 (2016).
Linschoten, M., Teske, A. J., Cramer, M. J., van der Wall, E. & Asselbergs, F. W. Chemotherapy-related cardiac dysfunction: A systematic review of genetic variants modulating individual risk. Circ. Genom. Precis. Med. 11, e001753. https://doi.org/10.1161/CIRCGEN.117.001753 (2018).
Zhang, Y. W., Shi, J., Li, Y. J. & Wei, L. Cardiomyocyte death in doxorubicin-induced cardiotoxicity. Arch. Immunol. Ther. Exp. (Warsz) 57, 435–445. https://doi.org/10.1007/s00005-009-0051-8 (2009).
Bhan, A. & Mandal, S. S. Long noncoding RNAs: Emerging stars in gene regulation, epigenetics and human disease. ChemMedChem 9, 1932–1956. https://doi.org/10.1002/cmdc.201300534 (2014).
Feinberg, A. The key role of epigenetics in human disease. N. Engl. J. Med. 379, 400–401. https://doi.org/10.1056/NEJMc1805989 (2018).
Bhattacharjee, D., Shenoy, S. & Bairy, K. L. DNA methylation and chromatin remodeling: The blueprint of cancer epigenetics. Scientifica (Cairo) 6072357, 2016. https://doi.org/10.1155/2016/6072357 (2016).
Zoghbi, H. Y. & Beaudet, A. L. Epigenetics and human disease. Cold Spring Harb. Perspect. Biol. 8, a019497. https://doi.org/10.1101/cshperspect.a019497 (2016).
Handy, D. E., Castro, R. & Loscalzo, J. Epigenetic modifications: Basic mechanisms and role in cardiovascular disease. Circulation 123, 2145–2156. https://doi.org/10.1161/CIRCULATIONAHA.110.956839 (2011).
Udali, S., Guarini, P., Moruzzi, S., Choi, S. W. & Friso, S. Cardiovascular epigenetics: from DNA methylation to microRNAs. Mol. Aspects Med. 34, 883–901. https://doi.org/10.1016/j.mam.2012.08.001 (2013).
Mahmoud, S. A. & Poizat, C. Epigenetics and chromatin remodeling in adult cardiomyopathy. J. Pathol. 231, 147–157. https://doi.org/10.1002/path.4234 (2013).
Rakyan, V. K., Down, T. A., Balding, D. J. & Beck, S. Epigenome-wide association studies for common human diseases. Nat. Rev. Genet. 12, 529–541. https://doi.org/10.1038/nrg3000 (2011).
Robinson, E. L. et al. Differential expression of epigenetic modifiers in early and late cardiotoxic heart failure reveals DNA methylation as a key regulator of cardiotoxicity. Front. Cardiovasc. Med. 10, 884174. https://doi.org/10.3389/fcvm.2023.884174 (2023).
Tony, H., Yu, K. & Qiutang, Z. MicroRNA-208a silencing attenuates doxorubicin induced myocyte apoptosis and cardiac dysfunction. Oxid. Med. Cell Longev. 2015, 597032. https://doi.org/10.1155/2015/597032 (2015).
Wang, J. X. et al. MicroRNA-532-3p regulates mitochondrial fission through targeting apoptosis repressor with caspase recruitment domain in doxorubicin cardiotoxicity. Cell Death Dis. 6, e1677. https://doi.org/10.1038/cddis.2015.41 (2015).
Feijen, E. A. M. et al. Derivation of anthracycline and anthraquinone equivalence ratios to doxorubicin for late-onset cardiotoxicity. JAMA Oncol. 5, 864–871. https://doi.org/10.1001/jamaoncol.2018.6634 (2019).
Morris, T. J. et al. ChAMP: 450k chip analysis methylation pipeline. Bioinformatics 30, 428–430. https://doi.org/10.1093/bioinformatics/btt684 (2014).
Tian, Y. et al. ChAMP: Updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984. https://doi.org/10.1093/bioinformatics/btx513 (2017).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 45, e22. https://doi.org/10.1093/nar/gkw967 (2017).
Triche, T. J. Jr., Weisenberger, D. J., Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of Illumina Infinium DNA methylation BeadArrays. Nucleic Acids Res. 41, e90. https://doi.org/10.1093/nar/gkt090 (2013).
Salas, L. A. et al. An optimized library for reference-based deconvolution of whole-blood biospecimens assayed using the Illumina HumanMethylationEPIC BeadArray. Genome Biol. 19, 64. https://doi.org/10.1186/s13059-018-1448-7 (2018).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127. https://doi.org/10.1093/biostatistics/kxj037 (2007).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115. https://doi.org/10.1186/gb-2013-14-10-r115 (2013).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367. https://doi.org/10.1016/j.molcel.2012.10.016 (2013).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY) 10, 573–591. https://doi.org/10.18632/aging.101414 (2018).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY) 11, 303–327. https://doi.org/10.18632/aging.101684 (2019).
Karlsson Linner, R. et al. An epigenome-wide association study meta-analysis of educational attainment. Mol. Psychiatry 22, 1680–1690. https://doi.org/10.1038/mp.2017.210 (2017).
Barfield, R. T., Kilaru, V., Smith, A. K. & Conneely, K. N. CpGassoc: An R function for analysis of DNA methylation microarray data. Bioinformatics 28, 1280–1281. https://doi.org/10.1093/bioinformatics/bts124 (2012).
Burridge, P. W. et al. Human induced pluripotent stem cell-derived cardiomyocytes recapitulate the predilection of breast cancer patients to doxorubicin-induced cardiotoxicity. Nat. Med. 22, 547–556. https://doi.org/10.1038/nm.4087 (2016).
Magdy, T. et al. RARG variant predictive of doxorubicin-induced cardiotoxicity identifies a cardioprotective therapy. Cell Stem Cell 28, 2076-2089e2077. https://doi.org/10.1016/j.stem.2021.08.006 (2021).
Kuo, H. H. et al. Negligible-cost and weekend-free chemically defined human iPSC culture. Stem Cell Rep. 14, 256–270. https://doi.org/10.1016/j.stemcr.2019.12.007 (2020).
Magdy, T. et al. Identification of drug transporter genomic variants and inhibitors that protect against doxorubicin-induced cardiotoxicity. Circulation 145, 279–294. https://doi.org/10.1161/CIRCULATIONAHA.121.055801 (2022).
Human Protein Atlas proteinatlas.org.
Du, W. W. et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur. Heart J. 38, 1402–1412. https://doi.org/10.1093/eurheartj/ehw001 (2017).
Wang, K. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 37, 2602–2611. https://doi.org/10.1093/eurheartj/ehv713 (2016).
Xia, L. & Song, M. Role of non-coding RNA in diabetic cardiomyopathy. Adv. Exp. Med. Biol. 1229, 181–195. https://doi.org/10.1007/978-981-15-1671-9_10 (2020).
Zhang, Q. et al. The circular RNA hsa_circ_0007623 acts as a sponge of microRNA-297 and promotes cardiac repair. Biochem. Biophys. Res. Commun. 523, 993–1000. https://doi.org/10.1016/j.bbrc.2019.12.116 (2020).
Sun, C., Ni, M., Song, B. & Cao, L. Circulating circular RNAs: Novel biomarkers for heart failure. Front. Pharmacol. 11, 560537. https://doi.org/10.3389/fphar.2020.560537 (2020).
Kristensen, L. S. et al. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 20, 675–691. https://doi.org/10.1038/s41576-019-0158-7 (2019).
Zhang, C. et al. Interactions of circRNAs with methylation: An important aspect of circRNA biogenesis and function (review). Mol. Med. Rep. https://doi.org/10.3892/mmr.2022.12685 (2022).
Liu, S., Xiong, X., Zhao, X., Yang, X. & Wang, H. F-BAR family proteins, emerging regulators for cell membrane dynamic changes-from structure to human diseases. J. Hematol. Oncol. 8, 47. https://doi.org/10.1186/s13045-015-0144-2 (2015).
Cao, H. et al. FCHSD1 and FCHSD2 are expressed in hair cell stereocilia and cuticular plate and regulate actin polymerization in vitro. PLoS ONE 8, e56516. https://doi.org/10.1371/journal.pone.0056516 (2013).
Lessard, C. J. et al. Identification of a systemic lupus erythematosus risk locus spanning ATG16L2, FCHSD2, and P2RY2 in Koreans. Arthritis Rheumatol. 68, 1197–1209. https://doi.org/10.1002/art.39548 (2016).
Chalovich, J. M. & Schroeter, M. M. Synaptopodin family of natively unfolded, actin binding proteins: Physical properties and potential biological functions. Biophys. Rev. 2, 181–189. https://doi.org/10.1007/s12551-010-0040-5 (2010).
Weins, A. et al. Differentiation- and stress-dependent nuclear cytoplasmic redistribution of myopodin, a novel actin-bundling protein. J. Cell Biol. 155, 393–404. https://doi.org/10.1083/jcb.200012039 (2001).
Frank, D. & Frey, N. Cardiac Z-disc signaling network. J. Biol. Chem. 286, 9897–9904. https://doi.org/10.1074/jbc.R110.174268 (2011).
Pyle, W. G. & Solaro, R. J. At the crossroads of myocardial signaling: The role of Z-discs in intracellular signaling and cardiac function. Circ. Res. 94, 296–305. https://doi.org/10.1161/01.RES.0000116143.74830.A9 (2004).
Ujhelly, O. et al. Lack of Rybp in mouse embryonic stem cells impairs cardiac differentiation. Stem Cells Dev. 24, 2193–2205. https://doi.org/10.1089/scd.2014.0569 (2015).
Alghamdi, M. et al. Phenotypic and molecular spectrum of pyridoxamine-5’-phosphate oxidase deficiency: A scoping review of 87 cases of pyridoxamine-5’-phosphate oxidase deficiency. Clin. Genet. 99, 99–110. https://doi.org/10.1111/cge.13843 (2021).
Friedrichs, F. et al. HBEGF, SRA1, and IK: Three cosegregating genes as determinants of cardiomyopathy. Genome Res. 19, 395–403. https://doi.org/10.1101/gr.076653.108 (2009).
Tayal, U., Prasad, S. & Cook, S. A. Genetics and genomics of dilated cardiomyopathy and systolic heart failure. Genome Med. 9, 20. https://doi.org/10.1186/s13073-017-0410-8 (2017).
Friso, S. et al. Low plasma vitamin B-6 concentrations and modulation of coronary artery disease risk. Am. J. Clin. Nutr. 79, 992–998. https://doi.org/10.1093/ajcn/79.6.992 (2004).
Zoccarato, A. & Shah, A. M. RORalpha nuclear receptors in protection against angiotensin II-induced cardiac hypertrophy. Am. J. Physiol. Heart Circ. Physiol. 316, H357–H359. https://doi.org/10.1152/ajpheart.00732.2018 (2019).
Zhao, Y. et al. Novel protective role of the circadian nuclear receptor retinoic acid-related orphan receptor-alpha in diabetic cardiomyopathy. J. Pineal Res. https://doi.org/10.1111/jpi.12378 (2017).
Beak, J. Y. et al. The nuclear receptor RORalpha protects against angiotensin II-induced cardiac hypertrophy and heart failure. Am. J. Physiol. Heart Circ. Physiol. 316, H186–H200. https://doi.org/10.1152/ajpheart.00531.2018 (2019).
Meder, B. et al. Epigenome-wide association study identifies cardiac gene patterning and a novel class of biomarkers for heart failure. Circulation 136, 1528–1544. https://doi.org/10.1161/CIRCULATIONAHA.117.027355 (2017).
Uhl, G. R. et al. The VMAT2 gene in mice and humans: Amphetamine responses, locomotion, cardiac arrhythmias, aging, and vulnerability to dopaminergic toxins. FASEB J. 14, 2459–2465. https://doi.org/10.1096/fj.00-0205rev (2000).
Barlow, G. M. et al. Down syndrome congenital heart disease: A narrowed region and a candidate gene. Genet. Med. 3, 91–101. https://doi.org/10.1097/00125817-200103000-00002 (2001).
Howey, R. A. J. & Cordell, H. J. Application of Bayesian networks to GAW20 genetic and blood lipid data. BMC Proc. 12, 19. https://doi.org/10.1186/s12919-018-0116-y (2018).
Kraja, A. T. et al. Simulation of a medication and methylation effects on triglycerides in the Genetic Analysis Workshop 20. BMC Proc. 12, 25. https://doi.org/10.1186/s12919-018-0115-z (2018).
Nassal, D. et al. Regulation of cardiac conduction and arrhythmias by ankyrin/spectrin-based macromolecular complexes. J. Cardiovasc. Dev. Dis. https://doi.org/10.3390/jcdd8050048 (2021).
Faul, C., Dhume, A., Schecter, A. D. & Mundel, P. Protein kinase A, Ca2+/calmodulin-dependent kinase II, and calcineurin regulate the intracellular trafficking of myopodin between the Z-disc and the nucleus of cardiac myocytes. Mol. Cell Biol. 27, 8215–8227. https://doi.org/10.1128/MCB.00950-07 (2007).
Odunewu-Aderibigbe, A. & Fliegel, L. The Na(+) /H(+) exchanger and pH regulation in the heart. IUBMB Life 66, 679–685. https://doi.org/10.1002/iub.1323 (2014).
Reynolds, L. M. et al. DNA methylation of the aryl hydrocarbon receptor repressor associations with cigarette smoking and subclinical atherosclerosis. Circ. Cardiovasc. Genet. 8, 707–716. https://doi.org/10.1161/CIRCGENETICS.115.001097 (2015).
Wang, J., Gareri, C. & Rockman, H. A. G-protein-coupled receptors in heart disease. Circ. Res. 123, 716–735. https://doi.org/10.1161/CIRCRESAHA.118.311403 (2018).
Inazumi, H. et al. 4968 increased Gao expression underlies cardiac dysfunction and lethal arrhythmias accompanied with abnormal Ca2+ handling. Eur. Heart J. https://doi.org/10.1093/eurheartj/ehz746.0027 (2019).
Welsh, P. et al. Cardiac troponin T and troponin I in the general population. Circulation 139, 2754–2764. https://doi.org/10.1161/CIRCULATIONAHA.118.038529 (2019).
Dewey, S., Lai, X., Witzmann, F. A., Sohal, M. & Gomes, A. V. Proteomic analysis of hearts from Akita mice suggests that increases in soluble epoxide hydrolase and antioxidative programming are key changes in early stages of diabetic cardiomyopathy. J. Proteome Res. 12, 3920–3933. https://doi.org/10.1021/pr4004739 (2013).
Shendre, A. et al. Local ancestry and clinical cardiovascular events among African Americans from the atherosclerosis risk in communities study. J. Am. Heart Assoc. https://doi.org/10.1161/JAHA.116.004739 (2017).
Chang, S. et al. Histone deacetylases 5 and 9 govern responsiveness of the heart to a subset of stress signals and play redundant roles in heart development. Mol. Cell Biol. 24, 8467–8476. https://doi.org/10.1128/MCB.24.19.8467-8476.2004 (2004).
Lehmann, L. H., Worst, B. C., Stanmore, D. A. & Backs, J. Histone deacetylase signaling in cardioprotection. Cell Mol. Life Sci. 71, 1673–1690. https://doi.org/10.1007/s00018-013-1516-9 (2014).
Zhang, D., Hu, X., Henning, R. H. & Brundel, B. J. Keeping up the balance: Role of HDACs in cardiac proteostasis and therapeutic implications for atrial fibrillation. Cardiovasc. Res. 109, 519–526. https://doi.org/10.1093/cvr/cvv265 (2016).
Guzzo, R. M., Salih, M., Moore, E. D. & Tuana, B. S. Molecular properties of cardiac tail-anchored membrane protein SLMAP are consistent with structural role in arrangement of excitation–contraction coupling apparatus. Am. J. Physiol. Heart Circ. Physiol. 288, H1810-1819. https://doi.org/10.1152/ajpheart.01015.2004 (2005).
Nader, M. et al. SLMAP-3 is downregulated in human dilated ventricles and its overexpression promotes cardiomyocyte response to adrenergic stimuli by increasing intracellular calcium. Can. J. Physiol. Pharmacol. 97, 623–630. https://doi.org/10.1139/cjpp-2018-0660 (2019).
Nader, M. The SLMAP/Striatin complex: An emerging regulator of normal and abnormal cardiac excitation–contraction coupling. Eur. J. Pharmacol. 858, 172491. https://doi.org/10.1016/j.ejphar.2019.172491 (2019).
Mlynarova, J. et al. SLMAP3 isoform modulates cardiac gene expression and function. PLoS ONE 14, e0214669. https://doi.org/10.1371/journal.pone.0214669 (2019).
Chamberlain, A. A. et al. DNA methylation is developmentally regulated for genes essential for cardiogenesis. J. Am. Heart Assoc. 3, e000976. https://doi.org/10.1161/JAHA.114.000976 (2014).
Kong, X. et al. The association of type 2 diabetes loci identified in genome-wide association studies with metabolic syndrome and its components in a Chinese population with type 2 diabetes. PLoS ONE 10, e0143607. https://doi.org/10.1371/journal.pone.0143607 (2015).
Ndiaye, F. K. et al. Expression and functional assessment of candidate type 2 diabetes susceptibility genes identify four new genes contributing to human insulin secretion. Mol. Metab. 6, 459–470. https://doi.org/10.1016/j.molmet.2017.03.011 (2017).
Barison, M. J., Pereira, I. T., Waloski Robert, A. & Dallagiovanna, B. Reorganization of metabolism during cardiomyogenesis implies time-specific signaling pathway regulation. Int. J. Mol. Sci. https://doi.org/10.3390/ijms22031330 (2021).
Chen, P. Y., Tu, H. C., Schirch, V., Safo, M. K. & Fu, T. F. Pyridoxamine supplementation effectively reverses the abnormal phenotypes of zebrafish larvae with PNPO deficiency. Front. Pharmacol. 10, 1086. https://doi.org/10.3389/fphar.2019.01086 (2019).
Ciapaite, J. et al. Pyridox(am)ine 5’-phosphate oxidase (PNPO) deficiency in zebrafish results in fatal seizures and metabolic aberrations. Biochim. Biophys. Acta Mol. Basis Dis. 1866, 165607. https://doi.org/10.1016/j.bbadis.2019.165607 (2020).
Acknowledgements
Yuan Tian for help optimizing the ChAMP pipeline for ‘paired’ analysis. Wanding Zhou for his feedback on filtering the SNP-related probes on the EPIC Arrays. Lucas Salas for his feedback on integrating the FlowSorted.Blood.EPIC package into the ChAMP pipeline. Richard Karlsson Linner for help with optimizing the statistical pipeline for EWAS analysis. Ake T Lu for help with GrimAge calculations.
Funding
NCI (R35CA220502; PI: S Bhatia), LLS (6563-19; PI: S Bhatia), The V Foundation (DT2019-010; PI: S Bhatia). This work was supported in part by the American Cancer Society Institutional Research Grant (#IRG-60-001-53-IRG) and the Kaul Pediatric Research Institute Research Grant (to P.S.). The Children’s Oncology Group study (COG-ALTE03N1; NCT00082745; PI-Bhatia) reported here is supported by the National Clinical Trials Network (NCTN) Operations Center Grant (U10CA180886; PI-Hawkins); the NCTN Statistics & Data Center Grant (U10CA180899; PI-Alonzo); the Children’s Oncology Group Chair’s Grant (U10CA098543; PI-Adamson); The COG Statistics & Data Center Grant (U10CA098413; PI-Anderson); the NCI Community Oncology Research Program (NCORP) Grant (UG1CA189955; PI-Pollock); and the Community Clinical Oncology Program (CCOP) Grant (U10CA095861; PI-Pollock), and the St Baldrick’s Foundation through an unrestricted grant.
Author information
Authors and Affiliations
Contributions
S.B. had full access to all data in the study. She takes responsibility for the integrity of the data and accuracy of the data analysis. S.B., P.S. and P.W.B. contributed to the study conception and design. P.S., L.H., S.H.A., F.M.B., D.S.H., F.G.K., M.M.H., J.P.N., A.K.R., J.P.G., W.L., collected and assembled the data. L.Z., P.S., and D.K.C. designed the methodology for data analysis. P.S., L.Z., D.A.S., R.B.C., M.J., D.K.C., T.M., S.H., X.W., N.S., D.E.M., P.W.B. and S.B. analyzed and interpreted the data. P.S., S.B., L.Z., D.A.S., and P.W.B. drafted the manuscript. All authors critically reviewed the manuscript for important intellectual content and approved the final manuscript. S.B., P.S., D.C., L.Z., W.L. and L.H. provided administrative, technical, or material support. S.B., P.S., L.H. and W.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, P., Zhou, L., Shah, D.A. et al. Identification of novel hypermethylated or hypomethylated CpG sites and genes associated with anthracycline-induced cardiomyopathy. Sci Rep 13, 12683 (2023). https://doi.org/10.1038/s41598-023-39357-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-39357-2
This article is cited by
-
Impact of anthracycline-based chemotherapy on RB1 gene methylation in peripheral blood leukocytes and biomarkers of oxidative stress and inflammation in sarcoma patients
Clinical and Translational Oncology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.