Abstract
A total of 187 lactic acid bacteria were isolated from four types of grains collected in South Korea. The bacterial strains were assigned as members of Levilactobacillus brevis, Latilactobacillus curvatus, Lactiplantibacillus plantarum, Lactococcus taiwanensis, Pediococcus pentosaceus, and Weissella paramesenteroides based on the closest similarity using 16S rRNA gene sequence analysis. The strains belonging to the same species were analyzed using RAPD-PCR, and one or two among strains showing the same band pattern were selected. Finally, 25 representative strains were selected for further functional study. Inhibitory effects of lipid accumulation were observed in the strains tested. Pediococcus pentosaceus K28, Levilactobacillus brevis RP21 and Lactiplantibacillus plantarum RP12 significantly reduced lipid accumulation and did not show cytotoxicity in C3H10T1/2 cells at treatment of 1–200 μg/mL. The three LAB strains decreased significantly expression of six adipogenic marker genes, PPARγ, C/EBPα, CD36, LPL, FAS and ACC, in C3H10T1/2 adipocytes. The three strains survived under strong acidity and bile salt conditions. The three strains showed adhesion to Caco-2 cells similar to a reference strain LGG. The resistance of the three strains to several antibiotics was also assessed. Strains RP12 and K28 were confirmed not to produce harmful enzymes based on API ZYM kit results. Based on these results, strains K28, RP21 and RP12 isolated from grains had the ability to inhibit adipogenesis in adipocytes and potentially be useful as probiotics.
Similar content being viewed by others
Introduction
Diet-induced obesity is a major factor in many chronic diseases such as cardiovascular disease, non-alcoholic fatty liver disease (NAFLD) and type2 diabetes, and is considered a serious health concern1,2. Currently, drug treatment and surgery with mechanisms that inhibit lipid absorption and suppress appetite are used worldwide to prevent and treat obesity. However, in the case of drugs, continuous administration is limited due to side effects, and a risk of complications in surgery is possible3,4. Therefore, non-drug therapies that are safer and improve the balance of metabolism are needed to treat and prevent obesity and probiotics have been proposed as an alternative5. Probiotics are defined as live microorganisms that provide the host with health benefits when administered in proper quantities6. It has been shown in many studies that the probiotics are effective in alleviation of intestinal inflammation, allergy prevention, reduced total serum cholesterol and inflammation and anti-cancer activity7,8,9,10. Among them, several probiotics are also known to have potential anti-obesity effects such as reduced body fat, blood sugar control and cholesterol improvement11,12. Lactic acid bacteria (LAB) is the main bacteria in probiotics13. According to the American Food and Drug Administration, LAB are generally regarded as safe (GRAS). LAB is widely used in the food industry, and studies of LAB functionality are growing in popularity. In several recent studies, LAB was shown to alleviate obesity by decreasing body weight, improving inflammatory state or glucose tolerance, and altering gut microbiota in diet-induced obese mice14,15,16.
Grains are a major staple food in Asia. Grains mainly contain carbohydrates including fiber and oligosaccharides, which makes them an excellent source of prebiotics defined as “a nondigestible food ingredient that beneficially affects the host by selectively stimulating the growth and/or activity of one or a limited number of bacteria in the colon, and thus improves host health”17,18. A microbial community composed of LAB was shown to exist in grains18. In a previous study, LAB isolated from grains exerted antimicrobial properties19; however, few studies about useful properties, including anti-obesity effects, have been performed.
The objectives of the present study were to isolate LAB from various grains collected in the Republic of Korea and to screen lactic acid bacteria with inhibition of adipogenesis. The selected strains with anti-adipogenic effects were investigated to determine whether they have useful properties as probiotic candidates for development as functional food.
Material and methods
Isolation of LAB strains and growth conditions
Four types of grains, rice, brown rice, black rice and hulled barley, were collected in the Republic of Korea. The grains were ground using blender and enriched for 7 days with distilled water under aerobic conditions. The grain samples were serially diluted with 0.85% (w/v) saline solution and spread on De Man-Rogosa-Sharpe (MRS; BD Difco, Sparks, MD, USA) agar. After incubation for 48 h at 30 ℃ or 37 ℃, LAB were isolated from the MRS agar plates and cultivated aerobically for 18 h at 30 ℃ or 37 °C in MRS agar. Lacticaseibacillus rhamnosus LGG (KCTC 5033), which was used as the experimental control strain for comparative analyses, was purchased from the Korean Collection for Type Cultures (Daejeon, Republic of Korea), and cultured at 37 ℃. All strains, including the experimental control strain, were stored at − 80 ℃ after suspension in 20% (w/v) glycerol solution (Georgiachem, GA, USA).
16S rRNA gene sequence analysis and random amplified polymorphic DNA-polymerase chain reaction (RAPD-PCR) analysis
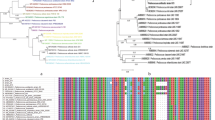
Genomic DNA was extracted using a G-spin genomic extraction kit (iNtRON, Seongnam, Republic of Korea), according to the manufacturer’s protocol. Polymerase chain reaction (PCR) amplification, purification, and sequencing of 16S rRNA gene were performed as described previously20. Identification of the closest phylogenetic species based on 16S rRNA gene sequence was performed using the EzBioCloud server (https://www.ezbiocloud.net/)21.
Random amplified polymorphic DNA-PCR (RAPD-PCR), which was used to exclude replicates among LAB strains, was performed using two primers, ERIC2 (5′-AAGTAAGTGACTGGGGTGAGCG-3′) and ERIC1R (5′-ATGTAAGCTCCTGGGGATTCAC-3′) as described previously22. The PCR products were electrophoresed on 1.5% (w/v) agarose (LPS Solution, Daejeon, Republic of Korea) gel for 60 min, and after electrophoresis, the gel was stained with RedSafe (iNtRON, Seongnam, Republic of Korea).
Preparation of cell extract from LAB
Cell extracts of LAB were prepared as described previously23 with minor modifications. The cell mass was harvested using centrifugation and washed twice with phosphate-buffered saline (PBS, pH 7.2). The washed cells were resuspended in distilled water at a concentration of 100 mg/mL and sonicated using the method described previously16. The sonicated cell extracts were centrifuged at 13,000 rpm for 15 min at 4 °C and the supernatants were filtered using a 0.45 µM syringe filter (Sartorius Stedim Biotech GmbH, Göttingen, Germany) and lyophilized. The resulting powder was dissolved in sterile water to appropriate concentrations.
Cell culture, adipocyte differentiation and intracellular triglyceride content
C3H10T1/2 cells were purchased from the American Type Culture Collection (Manassas, VA, USA) and cultured following the method described previously24. A subculture of C3H10T1/2 cells was performed with Dulbecco's modified Eagle's medium (DMEM) containing 10% fetal bovine serum (FBS; Hyclone, Logan, UT, USA) and antibiotics (penicillin and streptomycin, Hyclone). After seeding in 12-well plates, C3H10T1/2 cells were cultured in DMEM containing 10% FBS and antibiotics until confluency. Confluent cells were induced into adipocytes in DMEM supplemented with 10% FBS, antibiotics, 20 nM GW1929 (Sigma), 0.5 mM 3-isobutyl-1-methylxanthine (Sigma-Aldrich, St. Louis, MO, USA), 1 μM dexamethasone (Sigma-Aldrich), and 10 μg/mL insulin (Sigma-Aldrich). After 48 h, the differentiating cells were refreshed with media containing DMEM, 10% FBS, 20 nM GW1929, and 10 μg/mL insulin.
Cell extracts of LAB were adjusted by suspending to a concentration of 25, 50, and100 μg/mL with sterile distilled water to create the same conditions. LAB cell extracts were treated during adipocyte differentiation of C3H10T1/2 cells, and sterile distilled water was treated as control. Then, the differentiated C3H10T1/2 cells were fixed with 4% formaldehyde (Sigma-Aldrich) in PBS (Hyclone) at room temperature overnight and stained with Oil Red O (Sigma-Aldrich). To quantify intracellular triglyceride content, stained cells from at least two independent experiments were resolved in isopropanol (Sigma-Aldrich) and measured with a spectrophotometer at 520 nm.
Cell viability assay
Cell viability was determined using methyl thiazolyl tetrazolium salt (MTT) colorimetric assays (ab197010, Abcam). C3H10T1/2 cells were seeded at 1.5 × 104 cells per well in 96-well plates, then treated with various strain doses (1, 12.5, 25, 50, 100, and 200 μg/mL) and sterile distilled water as control in triplicate. After 24 h, MTT (20 μL/100 μL in medium) was added into the media and cells incubated for 4 h at 37 ℃. The absorbance of formazan dye was measured at 490 nm using a microplate reader (BioTek, Winooski, VT, USA).
Quantitative real-time polymerase chain reaction (RT PCR) analysis
Total RNA was extracted from C3H10T1/2 cells using QIAzol lysis reagent (QIAGEN, Germantown, MD, USA). First-strand complementary DNA was synthesized from 0.5 μg of total RNA using ReverTra Ace Master Mix (Toyobo, Osaka, Japan) according to the manufacturer’s instructions. Quantitative RT PCR was performed in 25 μL final reaction volume containing Power SYBR Premix ExTaq (RP041A; Takara, Shiga, Japan), primers, and cDNA using thermal cycler machine (Takara). The primer sequences used for the PCR were described previously24.
Tolerance assays against acid and bile salts
Tolerance to acid was measured as described previously25 with minor modifications. LAB strains were cultured overnight (18 h) at 30 °C (for RP21 and K28) or 37 °C (for RP12 and LGG), harvested for 10 min at 7000 rpm at 4 °C, and washed twice with PBS buffer (pH 7.2). Bacterial cells (approximately 109 CFU/mL) were resuspended in liquid MRS medium (pre-adjusted to pH 1.0, 2.0, 2.5, and 3.0) and incubated for 3 h at 30 or 37 °C. Viability was determined in triplicate in terms of viable colony counts using the plate count method. Tolerance to bile salts was measured as described previously26 with minor modifications. LAB strains were suspended in liquid MRS medium containing 0.3, 0.5, 1.0, and 2.0% oxgall (Sigma-Aldrich) at a concentration of approximately 109 CFU/mL. After incubation for 6 h at 30 °C or 37 °C, suspension was poured into MRS agar plates and incubated at optimum growth temperatures for 48 h. Tolerance assays against acid and bile salts were performed in triplicate and LGG was used as the comparative strain.
In vitro adhesion assays
The adherence assay was performed according to the method described previously27 with minor modifications. Caco-2 cells used for the adherence assay were purchased from the Korean Cell Line Bank (Seoul, Korea). The Caco-2 cells were cultured in high glucose DMEM supplemented with 10% (v/v) FBS (Hyclone) and 1% (v/v) penicillin–streptomycin at 37 °C in 5% CO2 atmosphere. The Caco-2 cells were seeded at 2 × 105 cells/well in 6-well tissue culture plates. The adherence assay was performed at post-confluence. The monolayer was washed with sterile PBS (Hyclone) twice. LAB cells were diluted with DMEM to approximately 109 CFU/mL and added to the wells. Plates were incubated for 90 min at 37 °C in 5% CO2 atmosphere. The Caco-2 monolayers were washed three times with sterile PBS (Hyclone) and treated with EDTA-trypsin solution for 3 min. The cell suspensions were serially diluted and spread on MRS agar plates. Cell viability was counted after incubation for 48 h. The adhesion ability of LAB was calculated as the percentage between remaining bacteria and initial bacteria per well. The same passage Caco-2 cells were used in adhesion assays and assays were repeated in triplicate.
Antibiotic susceptibility
Susceptibility to antibiotics was examined using the disc-diffusion method with application of modified agar diffusion method described previously28,29. LAB inoculated in MRS agar were adjusted to approximately 108 CFU/mL and paper discs (Advantec, Tokyo, Japan) were dispensed. Each disc was treated with 10 μL of specific antibiotic. The concentrations of antibiotics tested are listed in Supplementary Table 1. The inhibition zone diameters were measured and evaluated in terms of sensitive, intermediate sensitive, and resistant according to the interpretative standard table (Supplementary Table 1). The 2013 Clinical and Laboratory Standards Institute criteria30 were used for interpretation.
Enzyme activity test
Enzyme activity of the LAB was investigated as described previously16 using the API ZYM kit (BioMérieux, Marcy l’Etoile, France).
Statistical analysis
Results are presented as mean ± standard error of the mean (SEM) of three independent experiments. Significance differences between groups in triglyceride content were determined using Duncan's multi-range test. Significance differences in gene expression and adhesion ability were determined by comparison with control using two-tailed unpaired Student’s t-test. A p-value < 0.05 was considered statistically significant. Statistical analyses were performed using SPSS Inc. software (version 19.0).
Results and discussion
Isolation and identification of LAB strains from grains
Bacterial strains were isolated from four types of grains collected in the Republic of Korea and a total of 187 LAB strains were obtained through 16S rRNA gene sequencing followed by identification. From the 16S rRNA gene sequence analyses of the LAB strains, 20 strains had the closest similarities to the type strain of Levilactobacillus (previously Lactobacillus) brevis, 3 strains had the closest similarities to the type strain of Latilactobacillus (previously Lactobacillus) curvatus, 22 strains had the closest similarities to the type strain of Lactiplantibacillus (previously Lactobacillus) plantarum, 17 strains had the closest similarities to the type strain of Lactococcus taiwanensis, 82 strains had the closest similarities to the type strain of Pediococcus pentosaceus, and 43 strains had the closest similarities to the type strain of Weissella paramesenteroides (Table 1). The genus Lactobacillus has been recently reclassified as 25 genera including Levilactobacillus, Latilactobacillus, and Lactiplantibacillus31. Levilactobacillus brevis, Latilactobacillus curvatus, Lactiplantibacillus plantarum, Pediococcus pentosaceus, and Weissella paramesenteroides have been shown to be isolated from grains32,33.
In RAPD-PCR analysis, six different band patterns were assigned to 82 strains with the closest 16S rRNA gene sequence similarities to Pediococcus pentosaceus, and four different band patterns were assigned to 43 strains with the closest 16S rRNA gene sequence similarities to Weissella paramesenteroides (Supplementary Fig. 1). The strains assigned as Levilactobacillus brevis, Latilactobacillus curvatus, Lactiplantibacillus plantarum, and Lactococcus taiwanensis each showed only one type of band pattern (Supplementary Fig. 1). Finally, two representative strains from each group, except for the three groups having only one strain, were randomly selected, and used for further functional characterization (Supplementary Fig. 1; Table 1).
Screening of strains with anti-adipogenic effects
Inhibitory effects of lipid accumulation were tested by treating the LAB cell extract on C3H10T1/2 cells. Because the strains assigned to Lactococcus taiwanensis and Weissella paramesenteroides caused cell damage during the treatment process, they were excluded from the test. A wide range of inhibitory effects of lipid accumulation was observed in the strains selected. Among the strains tested, five strains (Pediococcus pentosaceus K28; Levilactobacillus brevis RP20 and RP21; Lactiplantibacillus plantarum RP11 and RP12) reduced lipid accumulation by more than 20% compared with the control, indicating that they have anti-adipogenic effects (Fig. 1). The two strains (RP20 and RP21) assigned to Levilactobacillus brevis and the two strains (RP11 and RP12) of Lactiplantibacillus plantarum showed similar results, respectively (Fig. 1). Thus, one strain from RP20 and RP21 and one strain from RP11 and RP12 were selected, and the three strains (K28, RP21 and RP12) were used for further experiments. The above results indicate that the components of LAB cell extract might influence the adipocyte differentiation process, thereby suppressing fat production.
Inhibitory effect on lipid accumulation of LAB strains isolated from grains. Each strain was treated as cell extract at a concentration of 50 μg/mL. Values of each sample were determined relative to control. Values are presented as the mean ± SEM of three independent experiments. Significant differences between selected strains and control are indicated as **p < 0.01, and ***p < 0.001.
Significant diversity exists among LAB strains regarding functional characteristics that benefit health, such as antioxidant, antitumor, immunomodulatory, and hypocholesterolemic activities34,35,36,37,38,39,40. In several studies, the cellular components of LAB have shown beneficial effects on improving health38,41,42. It is not clear which substance(s) in LAB cell extract induce the anti-adipogenic effects. Exopolysaccharide (EPS) has been known to have anti-adipogenic effects23. The EPS, a cell wall component of LAB cells, is loosely associated with the cell envelope and easily released into the surrounding environment43,44.
Effects of LAB strains on cell viability of C3H10T1/2
The cytotoxicity at various concentrations of strains LGG, K28, RP21, and RP12 on C3H10T1/2 cells was investigated by measuring cell viability using the MTT assay. C3H10T1/2 cells were found viable at all treatment concentrations of the four strains (Fig. 2). These results indicate that strains LGG, K28, RP21, and RP12 cause no damage to C3H10T1/2 cells45.
Effects of LAB treatment at different concentrations on viability of C3H10T1/2 cells. The C3H10T1/2 cells were treated with sterile distilled water (control) or LAB strains and their viability was determined using the MTT assay. Data are presented as the mean ± SEM from three independent experiments.
Inhibition of adipogenic gene expression by LAB extract during adipocyte differentiation
Inhibition of adipogenesis by strains K28, RP21 and RP12 was investigated by measuring expression of six adipogenic genes using quantitative RT PCR (Fig. 3). PPARγ and C/EBPα are transcription factors that regulate the process of adipocyte differentiation46,47. In addition, the activation of PPARγ promotes the expression of adipogenic genes, such as CD36 and LPL, which are important for the uptake and storage of triglycerides48. The down regulation of these adipogenic genes may affect decreased lipid accumulation in cells. Fatty acid synthase (FAS) gene is a downstream adipocyte gene that contributes to fatty acid synthesis49. Acetyl-coenzyme A carboxylase (ACC) is another key enzyme for fatty acid synthesis that catalyzes the synthesis of malonyl-CoA50.
Effects of selected LAB treatment on expression of adipogenic genes in C3H10T1/2 cells. The C3H10T1/2 cells were treated with the indicated concentrations during differentiation for 6 days and expression of adipocyte markers was measured. The mRNA expression levels of peroxisome proliferator-activated receptor γ (PPARγ), CCAAT-enhancer-binding protein-α (C/EBPα), lipoprotein lipase (LPL), fatty acid synthase (FAS), cluster of differentiation 36 (CD36), and acetyl-coenzyme A carboxylase (ACC) were measured using quantitative real-time polymerase chain reaction. Data are expressed as mean ± SEM of three independent experiments. Significant differences between selected strains (Pediococcus pentosaceus K28, Levilactobacillus brevis RP21 and Lactiplantibacillus plantarum RP12) and control are indicated as *p < 0.05, **p < 0.01, and ***p < 0.001.
Cell extracts from strains K28, RP21, and RP12 decreased the expression of adipocyte-related genes in C3H10T1/2 cells (Fig. 3). The expression of the six genes decreased proportionally with increasing concentrations of the extracts of the three strains (Fig. 3). The three strains significantly reduced (p < 0.01 or 0.001) the expression of PPARγ and C/EBPα in all concentrations tested. In addition, expression of four other genes associated with adipogenesis, was significantly reduced (p < 0.05 or 0.01) in the three strains, except for LPL expression in 25 μg/mL treatment of strains K28 and RP12. Strain K28, which showed the lowest lipid accumulation, was analyzed to have the lowest values in expressions of the six genes after 100 μg/mL treatment (Fig. 3). These results indicate that the three strains may have anti-adipogenic effects by inhibiting the expression of adipogenesis-related genes.
Tolerance against acid and bile salts
To have specific functionality, a probiotic must reach the intestines alive with resistance to acid and bile salts51. The acid tolerance of the selected strains and LGG as a reference strain was examined after incubation for 3 h in pH 3.0, 2.5, 2.0, and 1.0 (Table 2). The three strains and LGG maintained the values of more than 9 log CFU/mL at pH 3. Under pH 2.5 condition, the survival rates of strains K28 and RP21 decreased more than 2 log and approximately 1 log, respectively, whereas strains RP12 and LGG showed decreases of more than 3 log and 2 log, respectively (Table 2). Under pH 2 condition, the survival rate of strains K28 and RP21 decreased approximately 3 log and 2 log, respectively, and the survival rate of strains RP12 and LGG decreased approximately 6 log (Table 2). The three strains, except RP21 with approximately 3 log CFU/mL, showed low viability of less than 2 log CFU/mL at pH 1 (Table 2). Strain RP21 was also found to have higher acid resistance, as a strain of Levilactobacillus (Lactobacillus) brevis was shown highly acid-resistant in a previous study52. The pH of gastric fluid in the body is maintained at approximately 3.0, and probiotics are generally known to be highly acid-resistant if they are maintained at pH 3 for approximately 3 h53. Thus, the three strains were concluded to be highly tolerant to acid. Because food matrix can help the survival of LAB in the gastrointestinal tract due to its buffering capacity, the strains are expected to have stronger viability when used with carrier foods54.
Bile salts are another factor that can reduce bacterial survival in the gastrointestinal tract by destroying cell membranes51. Strains K28, RP21 and RP12 were found to survive after 6 h exposure to 0.3, 0.5, 1.0, and 2.0% bile salts, similar to LGG which is known to be highly resistant to bile salts (Table 2). Although the in vitro assay cannot provide the same conditions as the gastrointestinal tract, it is recognized as an effective evaluation method to select potential strains when using proper criteria27.
Adherence to Caco-2 cells
The adhesion ability of probiotics is a main factor that can increase the possibility of their survival and colonization in the gastrointestinal tract55. Adhesion is also required to prevent attachment of pathogenic bacteria through competition in intestinal epithelium56. Thus, the adherence ability has been considered an important biological property for the selection of useful probiotic strains57. In the present study, the adhesion ability of the three strains was evaluated using Caco-2 cells, which have morphological and physiological properties of human enterocytes, and their adhesion abilities were compared with that of the reference strain LGG (Fig. 4). Strain K28 had stronger adhesion ability than those of LGG and the two other strains (Fig. 4). The adhesion ability of strain K28 was highest at 1.95%, followed by LGG (1.79%), RP12 (1.67%), and RP21 (1.46%).
Antibiotic susceptibility
Probiotics have been widely used in various fields including food and medical industries. Antimicrobial sensitivity for evaluation of probiotics is considered important for safety, because the resistant genes can be horizontally transferred to pathogenic bacteria, which can become a serious threat58. Sensitivity results of the strains for nine antibiotics used in this study are listed in Supplementary Table 1. For the nine antibiotics tested, strains K28, RP21 and RP12 showed sensitivity patterns similar to strain LGG. In this study, all four strains were equally sensitive to chloramphenicol and rifampicin, whereas strains K28 and RP21 were intermediate sensitive to tetracycline and strains RP12 and LGG were sensitive to tetracycline (Supplementary Table 1). Sensitivity or intermediate sensitivity of Lactobacillus species and Pediococcus species to chloramphenicol and tetracycline has been previously reported59,60. Strains K28, RP21, RP12 and LGG were resistant to gentamycin, kanamycin, and streptomycin, which are known to inhibit protein synthesis targeting Gram-negative bacteria. The resistance to aminoglycoside antibiotics is an intrinsic property among Lactobacillus species and Pediococcus species61. Therefore, the three strains are unlikely to cause safety problems based on antibiotic susceptibility profile tested.
Enzyme production
For the safety of probiotic strains, it may be required to assess whether the strains produce harmful enzyme. β-glucuronidase is known as the carcinogen enzyme, which may increase the likelihood of tumor induction in the colon62,63. When the three strains were evaluated using API ZYM kit, strains RP12 and K28 did not produce any harmful enzymes such as β-glucuronidase, but strain RP21 was observed to produce β-glucuronidase (Supplementary Table 2).
Conclusion
In the present study, 187 LAB strains were isolated from four types of grains and identified using 16S rRNA gene sequence analysis. The 25 strains selected based on RAPD-PCR analysis were subjected to functional characterization. Among the strains tested, Pediococcus pentosaceus K28, Levilactobacillus brevis RP21, and Lactiplantibacillus plantarum RP12 had the potential to be useful probiotic candidates based on several characteristic analyses. The three strains exerted inhibitory effects on lipid accumulation and adipocyte differentiation by decreasing the expression of adipocyte-related genes. In addition, the three strains showed good tolerance against acid and bile salts, good intestinal cell adhesion, and were sensitive to chloramphenicol and rifampicin. In particular, strains RP12 and K28 did not produce β-glucuronidase. Therefore, Pediococcus pentosaceus K28, Levilactobacillus brevis RP21 and Lactiplantibacillus plantarum RP12 were concluded to have potential as probiotic candidates for use as functional neutraceutical foods.
Data availability
16S rRNA gene sequences of strains K28, RP21 and RP12 have been deposited in the National Centre for Biotechnology Information (NCBI) under GenBank accession numbers ON724233, ON724232 and ON724263, respectively.
References
Ford, N. D., Patel, S. A. & Narayan, K. M. Obesity in low- and middle-income countries: burden, drivers, and emerging challenges. Annu. Rev. Public Health 38, 145–164 (2017).
Mazzotti, A. et al. Which treatment for type 2 diabetes associated with non-alcoholic fatty liver disease?. Dig. Liver Dis. 49, 235–240 (2017).
Bessesen, D. H. & Van Gaal, L. F. Progress and challenges in anti-obesity pharmacotherapy. Lancet Diabetes Endocrinol. 6, 237–248 (2018).
Castellini, G. et al. Psychological effects and outcome predictors of three bariatric surgery interventions: A 1-year follow-up study. Eat. Weight Disord. 19, 217–224 (2014).
Ji, Y. et al. Amelioration of obesity-related biomarkers by Lactobacillus sakei CJLS03 in a high-fat diet-induced obese murine model. Sci. Rep. 9, 6821 (2019).
FAO. WHO Guidelines for the Evaluation of Probiotics in Food. Vols. 1–11 (2002).
Harb, H. et al. Neonatal supplementation of processed supernatant from Lactobacillus rhamnosus GG improves allergic airway inflammation in mice later in life. Clin. Exp. Allergy 43, 353–364 (2013).
Lin, P. W. et al. Lactobacillus rhamnosus blocks inflammatory signaling in vivo via reactive oxygen species generation. Free Radic. Biol. Med. 47, 1205–1211 (2009).
Ooi, L. G. & Liong, M. T. Cholesterol-lowering effects of probiotics and prebiotics: A review of in vivo and in vitro findings. Int. J. Mol. Sci. 11, 2499–2522 (2010).
Schultz, M. & Sartor, R. B. Probiotics and inflammatory bowel diseases. Am. J. Gastroenterol. 95, S19–S21 (2000).
Kobyliak, N. et al. Probiotics in prevention and treatment of obesity: A critical view. Nutr. Metab. 13, 14 (2016).
Shen, Y.-L. et al. Advances in the role and mechanism of lactic acid bacteria in treating obesity. Food Eng. 1, 101–115 (2022).
Ljungh, A. & Wadström, T. Lactic acid bacteria as probiotics. Curr. Issues Intest. Microbiol. 7, 73–89 (2006).
Li, X. et al. Lactobacillus plantarum prevents obesity via modulation of gut microbiota and metabolites in high-fat feeding mice. J. Funct. Foods 73, 104103 (2020).
Park, J. E., Oh, S. H. & Cha, Y. S. Lactobacillus brevis OPK-3 from kimchi prevents obesity and modulates the expression of adipogenic and pro-inflammatory genes in adipose tissue of diet-induced obese mice. Nutrients 12, 604 (2020).
Won, S. M. et al. Isolation of lactic acid bacteria from kimchi and screening of Lactobacillus sakei ADM14 with anti-adipogenic effect and potential probiotic properties. LWT-Food Sci. Technol. 126, 109296 (2020).
Gibson, G. R. & Roberfroid, M. B. Dietary modulation of the human colonic microbiota: Introducing the concept of prebiotics. J. Nutr. 125, 1401–1412 (1995).
Panghal, A. et al. Potential non-dairy probiotic products—A healthy approach. Food Biosci. 21, 80–89 (2018).
Ekwem, O. H. Isolation of antimicrobial producing lactobacilli from akamu (a Nigerian fermented cereal gruel). Afr. J. Microbiol. Res. 8, 718–720 (2014).
Yoon, J. H., Lee, S. T. & Park, Y. H. Inter- and intraspecific phylogenetic analysis of the genus Nocardioides and related taxa based on 16S rDNA sequences. Int. J. Syst. Bacteriol. 48, 187–194. https://doi.org/10.1099/00207713-48-1-187 (1998).
Yoon, S. H. et al. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67, 1613–1617 (2017).
Versalovic, J. et al. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 19, 6823–6831 (1991).
Zhang, Z. et al. Isolated exopolysaccharides from Lactobacillus rhamnosus GG alleviated adipogenesis mediated by TLR2 in mice. Sci. Rep. 6, 36083 (2016).
Song, N. J. et al. Butein is a novel anti-adipogenic compound. J. Lipid Res. 54, 1385–1396 (2013).
Zheng, Y. et al. Probiotic properties of Lactobacillus strains isolated from Tibetan kefir grains. PLoS ONE 8, e69868 (2013).
Bao, Y. et al. Screening of potential probiotic properties of Lactobacillus fermentum isolated from traditional dairy products. Food Control 21, 695–701 (2010).
Jacobsen, C. N. et al. Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl. Environ. Microbiol. 65, 4949–4956 (1999).
Charteris, W. P. et al. Antibiotic susceptibility of potentially probiotic Lactobacillus species. J. Food Prot. 61, 1636–1643 (1998).
Morelli, L. In vitro assessment of probiotic bacteria: From survival to functionality. Int. Dairy J. 17, 1278–1283 (2007).
CLSI. Performance standards for antimicrobial susceptibility testing; twenty-third informational supplement. Clin. Lab. Standards Inst. 33, 44–49 (2013)
Zheng, J. et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 70, 2782–2858 (2020).
Milanovic, V. et al. Selection of cereal-sourced lactic acid bacteria as candidate starters for the baking industry. PLoS ONE 15, e0236190 (2020).
Sáez, G. D. et al. Identification and biotechnological characterization of lactic acid bacteria isolated from chickpea sourdough in northwestern Argentina. LWT 93, 249–256 (2018).
Dilna, S. V. et al. Characterization of an exopolysaccharide with potential health-benefit properties from a probiotic Lactobacillus plantarum RJF4. LWT-Food Sci. Technol. 64, 1179–1186 (2015).
Fuentes, M. C. et al. Cholesterol-lowering efficacy of Lactobacillus plantarum CECT 7527, 7528 and 7529 in hypercholesterolaemic adults. Br. J. Nutr. 109, 1866–1872 (2013).
Jeun, J. et al. Hypocholesterolemic effects of Lactobacillus plantarum KCTC3928 by increased bile acid excretion in C57BL/6 mice. Nutrition 26, 321–330 (2010).
Kim, S. W. et al. Lactobacillus rhamnosus GG improves insulin sensitivity and reduces adiposity in high-fat diet-fed mice through enhancement of adiponectin production. Biochem. Biophys. Res. Commun. 431, 258–263 (2013).
Lee, J. W. et al. Immunomodulatory and antitumor effects in vivo by the cytoplasmic fraction of Lactobacillus casei and Bifidobacterium longum. J. Vet. Sci. 5, 41–48 (2004).
Liu, C. F. et al. Immunomodulatory and antioxidant potential of Lactobacillus exopolysaccharides. J. Sci. Food Agric. 91, 2284–2291 (2011).
Miyoshi, M. et al. Anti-obesity effect of Lactobacillus gasseri SBT2055 accompanied by inhibition of pro-inflammatory gene expression in the visceral adipose tissue in diet-induced obese mice. Eur. J. Nutr. 53, 599–606 (2014).
Jeung, W. H. et al. Lactobacillus curvatus HY7601 and Lactobacillus plantarum KY1032 cell extracts inhibit adipogenesis in 3T3-L1 and HepG2 cells. J. Med. Food 21, 876–886 (2018).
Kim, J. Y. et al. Screening for antiproliferative effects of cellular components from lactic acid bacteria against human cancer cell lines. Biotechnol. Lett. 24, 1431–1436 (2002).
Bhat, B. & Bajaj, B. K. Hypocholesterolemic potential and bioactivity spectrum of an exopolysaccharide from a probiotic isolate Lactobacillus paracasei M7. Bioact. Carbohydr. Diet. Fibre. 19, 100191 (2019).
Kleerebezem, M. et al. The extracellular biology of the lactobacilli. FEMS Microbiol. Rev. 34, 199–230 (2010).
van Meerloo, J., Kaspers, G. J. L. & Cloos, J. Cell sensitivity assays: The MTT assay. in Cancer Cell Culture. Methods in Molecular Biology (Cree, I. eds.). Vol 731. 237–245. (Humana Press, 1999).
Gregoire, F. M., Smas, C. M. & Sul, H. S. Understanding adipocyte differentiation. Physiol. Rev. 78, 783–809 (1998).
Rosen, E. D. et al. PPARγ is required for the differentiation of adipose tissue in vivo and in vitro. Mol. Cell 4, 611–617 (1999).
Tontonoz, P. & Spiegelman, B. M. Fat and beyond: The diverse biology of PPARγ. Annu. Rev. Biochem. 77, 289–312 (2008).
Liou, C. J. et al. Protective effects of licochalcone A ameliorates obesity and non-alcoholic fatty liver disease via promotion of the Sirt-1/AMPK pathway in mice fed a high-fat diet. Cells 8, 47 (2019).
Li, K. K. et al. Cocoa tea (Camellia ptilophylla) water extract inhibits adipocyte differentiation in mouse 3T3-L1 preadipocytes. Sci. Rep. 6, 20172 (2016).
Succi, M. et al. Bile salt and acid tolerance of Lactobacillus rhamnosus strains isolated from Parmigiano Reggiano cheese. FEMS Microbiol. Lett. 244, 129–137 (2005).
Wu, C. H. et al. Characterization of a potential probiotic Lactobacillus brevis RK03 and efficient production of γ-aminobutyric acid in batch fermentation. Int. J. Mol. Sci. 19, 143 (2018).
Guo, X. H. et al. Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 16, 321–326 (2010).
Linares, D. M. et al. Lactic acid bacteria and bifidobacteria with potential to design natural biofunctional health-promoting dairy foods. Front. Microbiol. 8, 846 (2017).
García-Cayuela, T. et al. Adhesion abilities of dairy Lactobacillus plantarum strains showing an aggregation phenotype. Food Res. Int. 57, 44–50 (2014).
Monteagudo-Mera, A. et al. Adhesion mechanisms mediated by probiotics and prebiotics and their potential impact on human health. Appl. Microbiol. Biotechnol. 103, 6463–6472 (2019).
Tuomola, E. M. & Salminen, S. J. Adhesion of some probiotic and dairy Lactobacillus strains to Caco-2 cell cultures. Int. J. Food Microbiol. 41, 45–51 (1998).
Gueimonde, M. et al. Antibiotic resistance in probiotic bacteria. Front. Microbiol. 4, 202 (2013).
Hummel, A. S. et al. Antibiotic resistances of starter and probiotic strains of lactic acid bacteria. Appl. Environ. Microbiol. 73, 730–739 (2007).
Maragkoudakis, P. A. et al. Probiotic potential of Lactobacillus strains isolated from dairy products. Int. Dairy J. 16, 189–199 (2006).
Singla, V. et al. Antibiotic susceptibility profile of Pediococcus spp. from diverse sources. 3 Biotech 8, 489 (2018).
Hatakka, K. et al. The influence of Lactobacillus rhamnosus LC705 together with Propionibacterium freudenreichii ssp. shermanii JS on potentially carcinogenic bacterial activity in human colon. Int. J. Food Microbiol. 128, 406–410 (2008).
Monteagudo-Mera, A. et al. Characterization of certain bacterial strains for potential use as starter or probiotic cultures in dairy products. J. Food Prot. 74, 1379–1386 (2011).
Acknowledgements
This work was supported by "Cooperative Research Program for Agriculture Science and Technology Development (Project No. PJ015247)" of Rural Development Administration, Republic of Korea and BK21 plus project of the Ministry of Education, Republic of Korea.
Author information
Authors and Affiliations
Contributions
M.J.S. and J.H.Y. conceived and designed the study. S.M.W., M.J.K., J.H.S., E.B.L., and J.H.C. performed the experiments. M.J.S. and J.H.Y. analyzed the data and wrote the manuscript. K.W.P. supported and discussed the study. J.H.Y. reviewed and edited manuscript. K.W.P. and J.H.Y. supervised the study. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seo, M.J., Won, SM., Kwon, M.J. et al. Screening of lactic acid bacteria with anti-adipogenic effect and potential probiotic properties from grains. Sci Rep 13, 11022 (2023). https://doi.org/10.1038/s41598-023-36961-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-36961-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.