Abstract
Regulation of cell division is crucial for the development of multicellular organisms, and in plants, this is in part regulated by the D-type cyclins (CYCD) and cyclin-dependent kinase A (CDKA) complex. Cell division regulation in Physcomitrium differs from other plants, by having cell division checks at both the G1 to S and G2 to M transition, controlled by the CYCD1/CDKA2 and CYCD2/CDKA1 complexes, respectively. This led us to hypothesize that upregulation of cell division could be archived in Bryophytes, without the devastating phenotypes observed in Arabidopsis. Overexpressing lines of PpCYCD1, PpCYCD2, PpCDKA1, or PpCDKA2 under Ubiquitin promotor control provided transcriptomic and phenotypical data that confirmed their involvement in the G1 to S or G2 to M transition control. Interestingly, combinatorial overexpression of all four genes produced plants with dominant PpCDKA2 and PpCYCD1 phenotypes and led to plants with twice as large gametophores. No detrimental phenotypes were observed in this line and two of the major carbon sinks in plants, the cell wall and starch, were unaffected by the increased growth rate. These results show that the cell cycle characteristics of P. patens can be manipulated by the ectopic expression of cell cycle regulators.
Similar content being viewed by others
Introduction
Bryophytes, including the moss Physcomitrium patens (syn. Physcomitrella patens), spend the majority of their life as photoautotrophic haploid gametophytes. Under appropriate environmental conditions, diploid sporophytes develop from leafy gametophores and produce spores and continue the dominant haploid life cycle. Haploid gametophyte cells in P. patens exist in two physiologically distinct cell types: the filamentous juvenile form of protonema, consisting of apically growing chloronema and caulonema cells and the adult leafy structure of gametophores. Protonema cells differentiate into adult leafy gametophores by initiating the division of three-faced apical cells (bud)1,2,3.
A dividing cell goes through four stages: G1, S, G2, and M phase, followed by cytoplasm division to separate the two daughter cells. The DNA replicates in S phase, with subsequent segregation in the M phase. The progression from G1 to S and G2 to M phases is controlled by cyclins and cyclin-dependent kinases (CDK) to ensure the completion of the previous S and M phases4. Plant cells are under normal conditions arrested at the G1-S transition and re-enter cell cycle progression upon proper stimuli4,5. In P. patens, D-type cyclins and CDKA forms complexes upon external or internal stimuli by sucrose, auxins, or cytokinin’s to progress the cell cycle through the G1 phase. Activation of the CYC/CDK complex phosphorylates the negative cell cycle regulator known as retinoblastoma (Rb) proteins to relieve the repression of E2F transcription factors, which is required for the S phase entry4. To follow the progression of the cell division an array of genes has been identified as G1-S and G2-M phase markers. Among these are the early S phase marker HFO (histone H4 gene), which is expressed just at the start of the S phase and marks the progression through the G1 to S. For the G2-M two classic markers are CYCB1 + 2 that both are expressed during the G2 to M phase. All of these have been used to study the cell division in plants and are good markers of the phase changes6. All three genes are used in this study.
P. patens protonema cells display a distinctive tissue-specific cell cycle alteration as compared to other plants. The faster-growing caulonema cells are arrested at G1-S transition controlled by CYCD1/CDKA2, whereas the slower-growing chloronema cells7 are arrested at G2-M transition controlled by CYCD2/CDKA13,5. This unique cell cycle arrest facilitates the highly efficient homologous recombination which is unique for bryophytes and not seen among other land plants8,9,10,11.
Fast cell proliferation is normally beneficial in industrial biotechnology, and P. patens is today used for biopharmaceutical production10,12,13,14. Although a lot of work has been done on ectopic expression of CYCD and CDKA proteins in vascular plants15,16,17, these efforts have led to detrimental phenotypes. Here we enhanced cell proliferation through ectopic expression of endogenous PpCDKA1, PpCDKA2, PpCYCD1 and PpCYCD2 genes individually and all four together. This led to an increase in gametophore size, which offer new opportunities for industrial biotechnology. Additionally, this showed that the transition from G1 to S is controlled by CYCD1/CDKA2, and the transition from G2 to M is controlled by CYCD2/CDKA1.
Results and discussion
Effects of PpCDKA1 overexpression
PpCDKA1 overexpression promoted protonema cell growth in BCDAT medium though with fewer as well as 10% shorter gametophores (Fig. 1A). The cells of the gametophores were 15% smaller, possibly explaining the observed size difference (Table 1A). When ZmUbi:PpCDKA1 lines were grown on solid BCD media it led to some differentiation of protonema into gametophores. The gametophores were 6% shorter and had 12% larger cell size, thus shorter with bigger and fewer cells. This indicates that ZmUbi:PpCDKA1 influences cell division negatively (Fig. 1B, Table 1B).
Phenotypic analysis of WT and transgenic lines. (A) Cell lines were grown in BCDAT medium for 35 days and photographed using Miotic SMZ-171 microscope. (B) Cells grown in similar conditions as above in BCD media. The cell lines are ZmUbi:PpCDKA1 (CDKA1), ZmUbi:PpCDKA2 (CDKA2), ZmUbi:PpCYCD1 (CYCD1), ZmUbi:PpCYCD2 (CYCD2), ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1 (CDK/CYC) and wildtype (WT). Same scale bar for all pictures = 1.5 mm.
An isolated ZmUbi:PpCDKA1 gametophyte grown for 35 days in liquid BCD medium was dominant in protonema cells and without larger gametophores, unlike WT that developed abundant amounts of gametophores (Fig. 1, 2, and Supplementary Fig. S1). The lesser gametophore differentiation in the ZmUbi:PpCDKA1 line prompted us to look at the expression of PpPIN, involved in auxin efflux18,19, as well as auxin gradient formation in the apical tip. PIN is essential for the initiation of protonemal differentiation, and it is established that auxin transport by PIN regulates gametophyte differentiations in moss2,19,20. qPCR analysis of PpPINA and PpPINB transcripts in ZmUbi:PpCDKA1 showed a > 80% reduction in both BCD and BCDAT media (Fig. 2). The lack of protonema differentiation towards gametophores as seen in Figs. 1, 2 and supplementary figures and stunted and lesser gametophore development in ZmUbi:PpCDKA1 lines can among other things be attributed to down-regulation of PIN and thereby change in auxin levels. However, the relationship between CDKA1 overexpression and PIN down-regulation is still unknown. No apparent change in the expression of auxin-induced transcription factors like RSL221 was observed in ZmUbi:PpCDKA1 in either of the two media (Fig. 2).
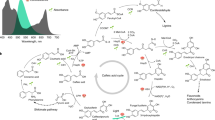
Heat Map of relative expression of cell cycle and cell development markers in mutant lines normalized to actin and tubulin. All cell lines, ZmUbi:PpCDKA1 (CDKA1), ZmUbi:PpCDKA2 (CDKA2), ZmUbi:PpCYCD1 (CYCD1), ZmUbi:PpCYCD2 (CYCD2), ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1 (CDK/CYC), are grown in BCDAT and BCD media, respectively. The figure also illustrates the cell cycle phases with the important checkpoints. The genes tested are CDKA1, CDKA2, Cyclin B1 and B2 (CYCB1, CYCB2), CYCD1, CYCD2, histone H4 gene (HFO), KIP-RELATED PROTEIN (KRP), auxin efflux carriers PINA and PINB, and auxin-induced transcription factor RSL2. The figure also illustrates the cell cycle phases with the important checkpoints. The G1/S marker is the HFO gene, and G2/M marker is the CYCB1 + 2.
Transcript analysis in BCDAT media showed that the transcript levels of the G2 to M phase markers, CYCB1 and CYCB24,5 were increased by 80% and 121%. Thus, CDKA1 is likely to be involved in the G2 to M transition as previously suggested5,7.
Effects of PpCDKA2 overexpression
Like ZmUbi:PpCDKA1, single ZmUbi:PpCDKA2 gametophores grown on BCD media were dominant in protonema cells and with small gametophores (Fig. 1, and Supplementary Fig. S1),. Furthermore, PpPIN expression was suppressed, but not as severely as in ZmUbi:PpCDKA1, possibly explaining the development of small gametophores as was not seen with ZmUbi:PpCDKA1 (Figs. 1 and 3). This indicates a different mechanism of the two cyclin-dependent kinases (Table 1).
The transcript analysis of ZmUbi:PpCDKA2, showed effects on cell cycle progression through G1 phase by increased expression of PpCDKA1, PpCYCD1, PpCYCD2 and down-regulation of PpKRP (Fig. 2). Signs of cell cycle advancement through G2-M phase are observed with increased expression of PpCYCB1 and PpCYCB2. Considering the protonema-dominant phenotype of the ZmUbi:PpCDKA2 lines, transcript analysis of cells grown in BCDAT suggests that ectopic expression of PpCDKA2 leads to protonema-specific gene expression and cell cycle proliferation. The up-regulation of G1-S and G2-M markers were more pronounced when cells were grown in BCDAT than in BCD media (Fig. 2). This shows that the ectopic expression of PpCDKA2 counteracts the ammonium tartrate G1-S inhibition, and links the PpCDKA2 to the G1 to S transition.
It has been demonstrated that ent-kaurenoic acid derived from ent-kaurene positively regulates the chloronema to caulonema differentiation as well as spore development in P. patens22,23. The two protonemal dominant lines ZmUbi:PpCDKA1 and ZmUbi:PpCDKA2 were grown on BCDAT for 28 days and compared to WT by a GC–MS analysis for the ent-kaurene level in the whole of the moss culture, thus both protonemal and the small possible gametophore tissue. The analysis showed for both ZmUbi:PpCDKA1 and ZmUbi:PpCDKA2 a significant increase in the levels of ent-kaurenes and ent-kaurenoic acid (Fig. 3). This indicates that the plants would try to compensate for the suppressed cell differentiation with the expression of other differentiation signals and hormones such as the ent-kaurene, highlighting the plasticity of plant cells to utilize several different signal pathways to obtain the same phenotype.
Effects of CYCD1 overexpression
Ectopic expression of PpCYCD1 promoted gametophore development and less protonema tissue in both BCD and BCDAT (Figs. 1, 2). The ZmUbi:PpCYCD1 gametophores grown in BCD and BCDAT were 1.7 and 1.6 times taller; and with larger leaf cell area on BCD media when compared to the WT (Table 1). The significant growth in gametophore height can in part be related to the increased expression of PpPINA and PpRSL2 transcript levels (Fig. 2), which regulates the differentiations into gametophores and more new buds as also seen in Arabidopsis17.
The G2-M markers CYCB1 + 2, the CDKA/CYCD complex suppresser KRP (KIP-RELATED PROTEIN) and early S phase marker HFO (histone H4 gene) were significantly downregulated in ZmUbi:PpCYCD1 cells grown on BCD (Fig. 2), whereas no significant differences were observed in cells on BCDAT (Fig. 2). A previous finding of P. patens gametophores hypothesized that gametophore cells tend to arrest in the late S phase5, and the observed increase in the size and numbers of gametophore cells together with low expression of CYCB1 + 2, HFO supports the previous finding. Overall these observations show that PpCYCD1 is likely to be involved in the G1 to S phase transition.
The gametophore cell area increased for all ZmUbi:PpCDKAs and ZmUbi:PpCYCD1 lines grown in BCD media. The ZmUbi:PpCYCD1 line had the most significant increase in gametophore cell area of 35% (Table 1). The cell area growth in plants could be under a cell cycle-related or cell cycle-independent regulation16. The increase in cell size suggests that CYCD1 might be able to influence the gametophore cell area growth in a cell cycle-related manner, but again this requires further studies.
Effects of CYCD2 overexpression
Similar to ZmUbi:PpCYCD1, the induction of gametophores in the ZmUbi:CYCD2 line was not impaired (Fig. 1). However, the height and cell area were reduced as compared to all the other lines in this study (Table 1). On average ZmUbi:PpCYCD2 grown on BCADT had 38% smaller gametophore cell size and 28% shorter plants (Table 1). When grown on BCD the cell size was reduced by 10%. No significant differences were observed in PIN transcript levels on any of the media, whereas RSL2 was upregulated on BCD but not BCDAT (Fig. 2). This indicates a link between auxin differentiation signals like RSL221 and cell cycle progression with the inhibition by ammonium tartrate. The data also suggest that Cyclin D’s influence the gametophore cell size in a cell-cycle dependent manner as previously speculated16.
The early S phase marker (Histone H4 gene, HFO) was upregulated on both BCDAT and BCD by 92% and 70% in ZmUbi:PpCYCD2 line (Fig. 2). This shows a relative increase in early S phase cells and that the gametophores were not arrested in the late S phase as seen in WT and ZmUbi:PpCYCD1, suggesting that PpCYCD2 induce the G2 to M transition.
Over-expression of AtCYCD3;1 in Arabidopsis has similar phenotypes as induced by exogenous cytokinin24. Previously, it has been shown that the daily replacement of the media stunted the differentiation of P. patens cells due to the removal of secreted auxins, cytokinins and other hormones3. To observe if overexpression of either PpCYCD1 or PpCYCD2 have any such phenotypic effect, we replaced the growth medium every day for two weeks. WT, ZmUbi:PpCYCD1 and ZmUbi:PpCYCD2 lines were all equally delayed when media was replaced every day (data not shown) indicating that PpCYCD1 and PpCYCD2 overexpression cannot replace the effect of secreted hormones.
Overexpression of CDKA1, CDKA2, CYCD1 and CYCD2 in the same cell line (CDK/CYC)
The unique functions of CDKAs and CYCDs prompted us to introduce CDKA1, CDKA2, CYCD1 and CYCD2 in the same transformation event to study the additive effects in P. patens and the potential for industrial application. The ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1 line showed a pronounced effect in both gametophore height (twofold) and gametophore cell area independent of growth media (Tables 1 and 2).
Analysis of cell cycle-related transcripts showed over 86% increase in both PpCYCB1 + 2 and 63% relatively low expression of PpKRP when grown on BCDAT, but not in BCD for PpCYCB1 + 2 though still with a 30% reduction PpKRP. For both media, the early S phase PpHFO was reduced, indicating fewer cells in the S phase. When all four genes are overexpressed in CDK/CYC the results suggest that transgenic cells advance faster through all the stages of the cell cycle (Fig. 2, Supplementary Fig. S2).
The dominating effects of CDKA2 and CYCD1 masked the dwarf gametophore phenotype of CDKA1 and CYCD2 when the CDK/CYC transgenic line was grown on both BCD and BCDAT mediums (Fig. 4A,B). Due to the large gametophore size of this transgenic line (Fig. 4C), the 90 days old “leaves” size were measured and the “leaves” were up to 70% broader (Table 2). Growing the CDK/CYC transgenic line for 90 days line in liquid BCD media yielded about half the biomass of WT when grown in BCD (Table 2). The major source for the difference between the two lines is the number and size of the rhizoids. Unlike with the gametophytes we observed only very few rhizoids in the CDK/CYC transgenic moss, and the length (Table 2, Fig. 4D) and numbers of rhizoids are significantly lower for the CDK/CYC transgenic line when grown in liquid culture (Figs. 1B, 4D and Supplementary Figs. S1 and S2).
(A) Comparison of a single gametophore of WT and the transgenic line grown in BCD liquid medium for 35 days (scale bar = 0.6 mm). (B) Homogenized cell lines studied in this work were grown for 90 days on BCDAT and BCD solid media for phenotypic comparison. (C) Comparison of gametophore phenotype of WT and ZmUbi:PpCKA1–PpCDKA2–OsAct:PpCYCD2–PpCYCD1 biological triplicate lines grown under same conditions for 35 days (scale bar = 0.6 mm). (D) Comparison of a single gametophore of WT and ZmUbi:PpCKA1–PpCDKA2–OsAct:PpCYCD2–PpCYCD1 grown in BCD liquid medium for 90 days, in six well microtiter plates.
This large phenotype of gametophores of ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1 (CDK/CYC) prompted us to analyse the cell wall composition and the starch content of all the transgenic lines reported in this study. No significant differences in the cell wall composition were observed between WT and the transgenic lines. The subsequent analysis of starch content showed that none of the engineered strains had reduced starch content when compared to WT. The ZmUbi:PpCDKA1 line had a surprising 50% increase in starch content over WT (Supplementary Figs. S3, S4 and Table S1). Thus, none of the transgenic lines appeared energy starved despite the growth phenotypes.
Ubiquitous expression of CYCD1, CYCD2, CDKA1 and CDKA2 led to more pleiotropic effects, for example, differences in cell differentiation leading to fewer gametophores for CDKA1, as discussed above. Based on these findings we tested combined overexpression of the four P. patens genes CDKA1, CDKA2, CYCD1, and CYCD2 and could observe that it leads to a twofold increase in the height of the gametophores but significantly smaller rhizoids and thus less accumulation of biomass. It remains to be investigated how the increase in gametophore size can be adopted in the rhizoids and this way provide significantly more biomass. The study confirms the link between the cyclins and cyclin-dependent kinases and the cell cycle phases. This was previously investigated through knock-outs, but our study confirms this through overexpression of the genes. The study shows that upregulation of the cell cycle does not always lead to dwarf mutants when the upregulation is performed through several genes, and not only one target. Similar studies remain to be done in other plants due to the technical challenges with the overexpression of multiple genes in vascular plants. Overall the study opens up for more questions on how the regulation of the cell cycle can be manipulated and show that with several genes uncoupled from the native regulation the cell cycle can be positively manipulated.
Conclusion
Combined overexpression was tested of the four P. patens genes CDKA1, CDKA2, CYCD1, and CYCD2 showed that it leads to a twofold increase in the height of the plants and increased accumulation of biomass in the early phase, but over a short time to less biomass, as the rhizoids growth was stunted. Individual overexpressing lines of PpCYCD1, PpCYCD2, PpCDKA1, or PpCDKA2 provided transcriptomic and phenotypical data that confirmed their involvement cell division cycle, and that the CYCD1/CDKA2 and CYCD2/CDKA1 complexes regulate the G1 to S and G2 to M phase transition, respectively.
Material and methods
Plant material, growth conditions, and transformation
Wild-type Physcomitrium patens (syn. Physcomitrella patens)25 (Gransden ecotype) was obtained from the International Moss Stock Center at the University of Freiburg (http://www.moss-stock-center.org/). All material used comply with relevant institutional, national, and international guidelines and legislation. Cells were grown in continuous white and red light at 23 °C, in BCD liquid or solid media with nitrate as the nitrogen source or ammonium tartrate as the additional nitrogen source (BCDAT). Further information on growth conditions and transformation processes could be found in previous publications11,26,27.
Strain construction
P. patens cells were transformed according to previously published protocols with minor modifications11,26 and yielded following moss cell mutant lines. ZmUbi:PpCDKA1, ZmUbi:PpCDKA2, ZmUbi:PpCycD1, ZmUbi:PpCycD2, ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1.
DNA fragments and genes
The coding sequence (CDS5) of PpCDKA1 (AJ515321), PpCDKA2 (AB547329), PpCYCD1 (AJ428953) and PpCYCD2 (AB547332) was PCR amplified from the cDNA of P. patens and stably transformed into the Pp108 neutral locus individually with the ubiquitin promoter and the OCS terminator. Further, CDS of PpCDKA1 and PpCDKA2, combined with translationally cleaved LP4/2A linker28,29 was synthesized as one construct using Thermo Fisher Scientific gene synthesis services. Similarly, PpCYCD1 and PpCYCD2 combined with LP4/2A linker were synthesized as PpCYCD2–PpCYCD1. Subsequently, all four genes were transformed into the Pp108 neutral locus using the ubiquitin and actin promoters. The Zea mays ubiquitin promoter, rice actin promoter, and the G418 antibiotic cassette was amplified from the pMP1355 and pZAG1 vector28,29. Stable transformants for each construct were selected according to previously published methods11,26,28. The list of the primers used is listed in Supplementary Table S2, S3, S4 and S5.
Phenotypic analysis
ZmUbi:PpCDKA1, ZmUbi:PpCDKA2, ZmUbi:PpCYCD1, ZmUbi:PpCYCD2, ZmUbi:PpCDKA1, ZmUbi:PpCDKA1-PpCDKA2-OsAct:PpCYCD2-PpCYCD1 and WT cell lines were blended and continuously grown in triplicates in BCD and BCDAT medium for 35 days. The media was changed at 17 days. The phenotype of each cell line was assessed by comparing images obtained with a Moticam 5.0 MP camera mounted on a Miotic SMZ-171 microscope. A selection of the pictures are found in Figs. 1 and 2, and Supplementary Figs. S1 and S2.
Cell size, gametophore height and “leaf” perimeter measurements
Cell size and gametophore height were measured by imaging leafy gametophore tissues of cells grown in continuous BCD and BCDAT liquid medium, the measurements were based on triplicates. For cell size measurements two trials were conducted with two different timelines, one at 35 days for all mutants (Table 1) and one at 90 days for ZmUbi:PpCDKA1–PpCDKA2–OsAct:PpCYCD2–PpCYCD1 (Table 2). Wild type was included in both experiments.
The mid-section of the larger “leaves” of the gametophores was imaged under the 200× magnification with Nikon ECLIPSE E1000 microscope. The tissue is not actual leaves but looks like it and as such referred to as “leaf” or leafy structures. Further, the micrographs were processed with the ImageJ software to calculate the area of the gametophore cells. From each gametophore, at least eight cells were measured. Gametophore height, rhizoid length and “leaf” length and width were measured by imaging with a Miotic SMZ-171 microscope equipped with a Moticam 5.0 MP camera. The single gametophytes were placed in a drop of media for easier analysis in the microscope. The images were further processed by image J to measure the length of the plants and the perimeter of the leaves. Perimeters of the “leaves” grown in BCD were measured by drawing a line from the base of the stem to the tip of the leaf using the freehand tool function of Image J to obtain the “leaf” area. Pictures and all data used for these measurements can be found here at this repository: https://doi.org/10.11583/DTU.20050499.
Cell biomass measurement
Following this, a second experiment was conducted also with a single gametophore from the ZmUbi:PpCDKA1–PpCDKA2–OsAct:PpCYCD2–PpCYCD1 line and the WT was grown in 6 well microplate in liquid BCD media. After 90 days of cultivation, cells were dried at 65 °C for 24 h and weighed for dry weight measurements.
qRT-PCR analysis
RNA transcription levels for ZmUbi:PpCDKA1, ZmUbi:PpCDKA2, ZmUbi:PpCYCD1, ZmUbi:PpCYCD2, ZmUbi:PpCDKA1–PpCDKA2–OsAct:PpCYCD2–PpCYCD1 and WT lines were measured by extracting the total RNA from cell lines (in triplicates) after 10 days of inoculation in BCD and BCDAT liquid mediums, using Spectrum™ Plant Total RNA Kit (Sigma, STRN250). A total of 500 ng of each RNA sample was reverse transcribed to cDNA with a iScriptTM cDNA Synthesis kit (BIO-RAD, Copenhagen, Denmark). RT-PCR was performed using QuantiFast® SYBR® Green PCR (Qiagen, Copenhagen, Denmark) according to the manufacturer’s protocol. The following amplification program was used: 95 °C 5 min, 40 cycles at 95 °C 10 s followed by 60 °C 30 s. Samples were amplified in duplicate from the same RNA isolation. RT-PCRs were performed using three biological triplicates with qPCR primers designed for the necessary genes and the actin and tubulin controls (Supplementary Table S5). RT-PCR efficiency, E, was estimated for each gene by generating standard curves by plotting quantification cycle (Cq) values (y) against the log of a serial cDNA dilution (x). For this, a cDNA sample was used as a template in a range of 25, 5, 2.5, 1.2 and 0.6 ng. The RT-PCR efficiencies were calculated from the slope of the linear regression equations of the standard curves, along with the regression coefficient (R2). The equation used was E = 10(–1/a), E values in a range of 1.90–2.10 (PCR efficiency between 90 and 110%) with a regression coefficient below 0.02 are acceptable. The ectopic expression of the four genes was also verified (Supplementary Fig. S5).
Cell wall and starch composition
All the transgenic lines and WT lines were grown in triplicates in BCD medium, changed every two weeks, for 6 weeks and freeze-dried for 48 h. Dried samples were ball milled in 70% EtOH in a Genogrinder (SPEX, NJ, USA) for 2 min. at 1500 rpm, centrifuged for 10 min. at 10,000g and the supernatant was discarded. The samples were first extracted with chloroform/MeOH (1:1 v/v) by adding the chloroform/MeOH, vortexing vigorously followed by centrifugation at 10,000g for 10 min. The supernatant was discarded and the pellet was extracted with 100% acetone as above. The supernatant was again discarded and the samples were then left to air dry in the fume hood. The sample weight was noted and starch was digested as described in Mosele et al., 2011 with Termamyl (Novozymes, Denmark) and amyloglucosidase (Megazyme, Ireland). The EtOH wash containing the Glc, maltose, and smaller dextrins, was dried in a speed-vac and the pellet was hydrolysed in 2 M TFA as previously described30 and the glucose quantified using a Dionex ICS 5000 + DC system (ThermoFisher, MA, USA) as described in Jensen et al., 2018. The pellet after starch removal was dried and analysed by both CoMPP as previously described31,32, and by sugar composition analysis after TFA hydrolysis as described above for starch.
GC/MS analysis of the ent-kaurene
Blended ZmUbi:PpCDKA1, ZmUbi:PpCDKA2, and WT lines were grown for 18 days in BCDAT media. Cell lines were dried at 98 °C for 24 h, and 10 mg of dry matter was extracted with 1 mL of ethyl acetate using a bead beater for 120 s at maximum power with 0.5 g of glass beads. Lysed cells were further extracted with ethyl acetate in an ultrasonic bath for 1 h. Completely extracted cells were centrifuged for 3 min at 12,000×g and 0.75 ml of the top layer containing hydrophobic molecules were isolated. 1 µl of the extracted ethyl acetate layer was injected into a GC/MS equipped with a ZB-5 ms column (30 m × 0.25 mm × 0.25 μm). The samples were injected in a non-split mode using the following program: Initial temperature of the oven was held for 1 min at 50 °C. Then, the temperature was increased to 320 °C at 15 °C/min and held at 325 °C for 5 min. The total run took 24 min. The injector temperature was set at 250 °C. The ion source temperature of the mass spectrometer was 230 °C. The data obtained was analysed for ent-kaurene and ent-kaurenoic acid using the mass spectra data published previously33,34.
Data availability
Data not represented online are available from the corresponding author upon reasonable request. The genes PpCDKA1 (AJ515321), PpCDKA2 (AB547329), PpCYCD1 (AJ428953) and PpCYCD2 (AB547332), are available from the NCBI database (https://www.ncbi.nlm.nih.gov/). The data used for broader image analysis, dry weight data, cell wall measurements, images of different setting of moss growth not used in the paper are available in the DTU repository. Simonsen, Henrik Toft (2023): Data for Overexpression of Physcomitrium patens Cell Cycle Regulators Leads to Larger Gametophytes. Technical University of Denmark. Dataset. https://doi.org/10.11583/DTU.20050499.
References
Reski, R. Development, genetics and molecular biology of mosses. Bot. Acta 111, 1–15 (1998).
Thelander, M., Landberg, K. & Sundberg, E. Auxin-mediated developmental control in the moss Physcomitrella patens. J. Exp. Bot. 69, 277–290. https://doi.org/10.1093/jxb/erx255 (2018).
Schween, G., Gorr, G., Hohe, A. & Reski, R. Unique tissue-specific cell cycle in Physcomitrella. Plant Biol. 5, 50–58. https://doi.org/10.1055/s-2003-37984 (2003).
Inzé, D. & De Veylder, L. Cell cycle regulation in plant development. Annu. Rev. Genet. 40, 77–105. https://doi.org/10.1146/annurev.genet.40.110405.090431 (2006).
Ishikawa, M. et al. Physcomitrella cyclin-dependent kinase a links cell cycle reactivation to other cellular changes during reprogramming of leaf cells. Plant Cell 23, 2924–2938 (2011).
Echevarría, C., Gutierrez, C. & Desvoyes, B. Tools for assessing cell-cycle progression in plants. Plant Cell Physiol. 62, 1231–1238. https://doi.org/10.1093/pcp/pcab066 (2021).
Menand, B., Calder, G. & Dolan, L. Both chloronemal and caulonemal cells expand by tip growth in the moss Physcomitrella patens. J. Exp. Bot. 58, 1843–1849. https://doi.org/10.1093/jxb/erm047 (2007).
Schaefer, D. G. & Zrÿd, J. P. Efficient gene targeting in the moss Physcomitrella patens. Plant J. 11, 1195–1206. https://doi.org/10.1046/j.1365-313X.1997.11061195.x (1997).
Nishiyama, T. et al. Tagged mutagenesis and gene-trap in the moss, Physcomitrella patens by shuttle mutagenesis. DNA Res. 7, 9–17. https://doi.org/10.1093/dnares/7.1.9 (2000).
Horn, A. et al. Natural products from bryophytes: From basic biology to biotechnological applications. CRC Crit. Rev. Plant Sci. 40, 191–217. https://doi.org/10.1080/07352689.2021.1911034 (2021).
King, B. C. et al. In vivo assembly of DNA-fragments in the moss, Physcomitrella patens. Sci. Rep. 6, 25030. https://doi.org/10.1038/srep25030 (2016).
Reski, R., Bae, H. & Simonsen, H. T. Physcomitrella patens, a versatile synthetic biology chassis. Plant Cell Rep. 37, 1409–1417. https://doi.org/10.1007/s00299-018-2293-6 (2018).
Simonsen, H. T., Drew, D. P. & Lunde, C. Perspectives on using Physcomitrella patens as an alternative production platform for thapsigargin and other terpenoid drug candidates. Perspect Med. Chem. 2009, 1–6. https://doi.org/10.4137/PMC.S2220 (2009).
Sabovljević, M. S. et al. Bryophytes—An emerging source for herbal remedies and chemical production. Plant Genet. Resour. 14, 314–327. https://doi.org/10.1017/S1479262116000320 (2016).
Iwakawa, H., Shinmyo, A. & Sekine, M. Arabidopsis CDKA1, a cdc2 homologue, controls proliferation of generative cells in male gametogenesis. Plant J. 45, 819–831. https://doi.org/10.1111/j.1365-313X.2005.02643.x (2006).
Qi, R. & John, P. C. L. Expression of genomic AtCYCD2;1 in Arabidopsis induces cell division at smaller cell sizes: Implications for the control of plant growth. Plant Physiol. 144, 1587–1597. https://doi.org/10.1104/pp.107.096834 (2007).
Cho, J. W. et al. Cyclin D1 and p22ack1 play opposite roles in plant growth and development. Biochem. Biophys. Res. Commun. 324, 52–57. https://doi.org/10.1016/j.bbrc.2004.08.233 (2004).
Kovrizshnykh, V. V., Omelyanchuk, N. A., Pasternak, T. P. & Mironova, V. V. The key role of PIN proteins in auxin transport in Arabidopsis thaliana roots. Russ. J. Genet. Appl. Res. 5, 279–285. https://doi.org/10.1134/S2079059715030089 (2015).
Bennett, T. A. et al. Plasma membrane-targeted PIN proteins drive shoot development in a moss. Curr. Biol. 24, 2776–2785. https://doi.org/10.1016/j.cub.2014.09.054 (2014).
Ashton, N. W., Schulze, A., Hall, P. & Bandurski, R. S. Estimation of indole-3-acetic acid in gametophytes of the moss, Physcomitrella patens. Planta 164, 142–144. https://doi.org/10.1007/BF00391040 (1985).
Jang, G. & Dolan, L. Auxin promotes the transition from chloronema to caulonema in moss protonema by positively regulating PpRSL1and PpRSL2 in Physcomitrella patens. New Phytol. 192, 319–327. https://doi.org/10.1111/j.1469-8137.2011.03805.x (2011).
Vesty, E. F. et al. The decision to germinate is regulated by divergent molecular networks in spores and seeds. New Phytol. 211, 952–966. https://doi.org/10.1111/nph.14018 (2016).
Hayashi, K. et al. Endogenous diterpenes derived from ent-Kaurene, a common gibberellin precursor, regulate protonema differentiation of the moss Physcomitrella patens. Plant Physiol. 153, 1085–1097. https://doi.org/10.1104/pp.110.157909 (2010).
Riou-Khamlichi, C. Cytokinin activation of Arabidopsis cell division through a D-type cyclin. Science 283, 1541–1544. https://doi.org/10.1126/science.283.5407.1541 (1999).
Medina, R. et al. Phylogenomic delineation of Physcomitrium (Bryophyta: Funariaceae) based on targeted sequencing of nuclear exons and their flanking regions rejects the retention of Physcomitrella, Physcomitridium and Aphanorrhegma. J. Syst. Evol. 57, 404–417. https://doi.org/10.1111/jse.12516 (2019).
Bach, S.S., King, B.C., Zhan, X. et al. Heterologous stable expression of terpenoid biosynthetic genes using the moss Physcomitrella patens. in Plant Isoprenoids. Methods in Molecular Biology (Methods and Protocols) (Rodríguez-Concepción, M. ed.). 257–271 (Humana Press, 2014).
Batth, R., Cuciurean, I.S., Kodiripaka, S.K. et al. Cellulase and Macerozyme—PEG-mediated transformation of moss protoplasts. BioProtocol (2020).
Khairul Ikram, N. K. B. et al. Stable production of the antimalarial drug artemisinin in the moss Physcomitrella patens. Front. Bioeng. Biotechnol. 5, 1–8. https://doi.org/10.3389/fbioe.2017.00047 (2017).
Pan, X. W. et al. Sclareol production in the moss Physcomitrella patens and observations on growth and terpenoid biosynthesis. Plant Biotechnol. Rep. 9, 149–159. https://doi.org/10.1007/s11816-015-0353-8 (2015).
Øbro, J., Harholt, J., Scheller, H. V. & Orfila, C. Rhamnogalacturonan I in Solanum tuberosum tubers contains complex arabinogalactan structures. Phytochemistry 65, 1429–1438. https://doi.org/10.1016/j.phytochem.2004.05.002 (2004).
Moller, I. et al. High-throughput mapping of cell-wall polymers within and between plants using novel microarrays. Plant J. 50, 1118–1128. https://doi.org/10.1111/j.1365-313X.2007.03114.x (2007).
Pedersen, H. L. et al. Versatile high resolution oligosaccharide microarrays for plant glycobiology and cell wall research. J. Biol. Chem. 287, 39429–39438 (2012).
Hayashi, K. et al. Identification and functional analysis of bifunctional ent-kaurene synthase from the moss Physcomitrella patens. FEBS Lett. 580, 6175–6181. https://doi.org/10.1016/j.febslet.2006.10.018 (2006).
Zhan, X. et al. Additional diterpenes from Physcomitrella patens synthesized by copalyl diphosphate/kaurene synthase (PpCPS/KS). Plant Physiol. Biochem. 96, 110–114. https://doi.org/10.1016/j.plaphy.2015.07.011 (2015).
Acknowledgements
AP, HB, JH and HTS were supported by The Danish Council for Independent Research (#4005-00158B). CQ was supported by funding from the People Program (Marie Curie Actions) of the European Union's Seventh Framework Program FP7/2007-2013/ under REA grant agreement n° [607011]. The Carlsberg Foundation and the Villum Foundation's Young Investigator Program are gratefully acknowledged for their support of JUF and JH. RB was supported by Villum Foundation Experiment grant (00022855). FJAR was supported by a stipend from Agencia Nacional de Investigaciõn y Desarollo, Chile.
Author information
Authors and Affiliations
Contributions
H.T.S. and J.H. conceived and directed the project. A.P., F.R., R.B., H.B., J.F., C.Q., J.H. and H.T.S. designed the experiments; A.P., F.R., R.B., H.B., J.F., J.H. and C.Q. performed the experiments and analysed the data; A.P., J.H. and H.T.S. wrote the draft manuscript; all authors revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peramuna, A., López, C.Q., Rios, F.J.A. et al. Overexpression of Physcomitrium patens cell cycle regulators leads to larger gametophytes. Sci Rep 13, 4301 (2023). https://doi.org/10.1038/s41598-023-31417-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31417-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.