Abstract
The perennial western ragweed (Ambrosia psilostachya DC.) arrived from North America to Europe in the late nineteenth century and behaves invasive in its non-native range. Due to its efficient vegetative propagation via root suckers, A. psilostachya got naturalized in major parts of Europe forming extensive populations in Mediterranean coastal areas. The invasion history, the spreading process, the relationships among the populations as well as population structuring is not yet explored. This paper aims to give first insights into the population genetics of A. psilostachya in its non-native European range based on 60 sampled populations and 15 Simple Sequence Repeats (SSR). By AMOVA analysis we detected 10.4% of genetic variation occurring among (pre-defined) regions. These regions represent important harbors for trading goods from America to Europe that might have served as source for founder populations. Bayesian Clustering revealed that spatial distribution of genetic variation of populations is best explained by six groups, mainly corresponding to regions around important harbors. As northern populations show high degrees of clonality and lowest levels of within-population genetic diversity (mean Ho = 0.40 ± 0.09), they could preserve the initial genetic variation levels by long-lived clonal genets. In Mediterranean populations A. psilostachya expanded to millions of shoots. Some of those were obviously spread by sea current along the coast to new sites, where they initiated populations characterized by a lower genetic diversity. For the future, the invasion history in Europe might get clearer after consideration of North American source populations of western ragweed.
Similar content being viewed by others
Introduction
Invasive species increase the local species richness1 but can also threaten native biodiversity by competition at various levels2. Beyond biodiversity threats, invasive plants could also reduce yield from agricultural fields3 and directly affect human health4. The latter holds for species of the genus Ambrosia that produce highly allergenic pollen5. Particularly, A. artemisiifolia L. (common ragweed) is well-documented to cause health problems like allergic rhinitis (hay fever) and asthma in its native range (North America6) as well as in invaded continents like Europe, Asia, Australia, and Africa7. All Ambrosia species are wind-pollinated and produce high numbers of pollen during summer8. Common ragweed is most striking for its release of allergic pollen5,9, and some other species (i.e. A. trifida L., A. psilostachya DC., A. confertiflora DC.) are also known to cause allergic diseases10. At the moment, five non-native Ambrosia species are documented from Europe11: A. artemisiifolia, A. psilostachya, A. tenuifolia Schreb., A. trifida, and A. tomentosa Nutt. A sixth species, A. maritima L., is supposed to be native to the Old World8,12,13. Only one of the alien ragweed species is widespread throughout temperate Europe (i.e. A. artemisiifolia12,14), continuously enlarging its distribution area towards northern regions–possibly favored by climate warming15,16,17. Western (= perennial) ragweed, A. psilostachya, is documented from several continents18 and occurs in many European countries19 behaving invasive in some regions3,16,20,21. It is treated taxonomically as conspecific to some other taxa from North America, i.e. to A. coronopifolia Torrey & Gray6,8. In its native range in Northern Mexico, USA (in 42 out of 50 federal states6) and all southern states of Canada6, A. psilostachya prefers open vegetation on sandy soils, i.e. prairie grasslands, but also alluvial areas and ruderal places6,8,18,21. This perennial herb is co-dominant to other grassland herbs and grasses, specifically in the Great Plains22,23, but prefers ruderal places towards the Great Lakes area and South-eastern Canada24. Naturalization of western ragweed happened even over a wider geographical range than in common ragweed due to the formation of root sprouts that survive cold winters belowground and allow clonal growth. An extensive review of specimens in European herbarium collections by the first author uncovered about 10% misidentifications by the collectors. Problems with identification resulted mostly from less accurate determination keys in floras like those by Hansen25, Sell & Murrell26, or Pignatti27. A. psilostachya is perennial with an extensive belowground root and shoot system8,11. Extensive populations are known from the Mediterranean coast of Spain28, France3, and Italy11,21,29 but also from the easternmost continental part of Europe (Russia20). A. psilostachya populations along the Northern European coastlines and inland sand habitats in Central Europe (Germany and Poland) are scattered. Perennation by root sprouts allows this invasive alien to survive in stabile populations on sandy substrates even in rather northern climates like Sweden (Falkenberg, 50 km S of Gothenburg, G. Karrer, pers. obs.) or north-western England (Birkdale near Southport, P. Ashton, pers. comm.). In such regions the annual A. artemisiifolia cannot naturalize due to failure of seed production12. The northernmost ephemeral finding of western ragweed is from Kurtto, Finland (pers. herbarium collection J. Särkkä, 2007, https://laji.fi/en/taxon/MX.42782).
However, invasion history of A. psilostachya to Europe is not well-documented; the first specimens were reported from Great Britain in 188030, France in 189131, Germany in 1894 (herbarium Leiden: L 3,003,673), Italy in 1924 (herbarium Pisa), Spain in 194732, and Russia in 1934 (specimen in the herbarium of Ufa). In Northern Africa, the first specimen was documented in 1918 from Algiers (herbarium Leiden: L 3,004,391). It is assumed that A. psilostachya arrived to Europe by traded crops21, i.e. potatoes from the Americas may have been contaminated by root fragments of western ragweed. No data is available about the likely source regions in America for the introductions to Europe. In Southern Europe, the invasion or expansion process is ongoing. In contrast, towards the north, there is no documentation about any increase of population sizes or further spread from initial places of introduction. Several accidental introductions in Central or Northern Europe (Austria, Hungary, Czech Republic, Denmark, Estonia, Finland, Norway) turned out to be ephemeral19,33.
Important biological traits of A. psilostachya like the prevailing asexual reproduction by root sprouts were documented mostly from its native range in North America31,34. Wagner & Beals24 described a rapid increase of single shoot spatial expansion up to 2 m2 within the second year of establishment. Consequently, clonal growth might result in phenotypically uniform populations. In the native as well as in the (European) invasive range, A. psilostachya regenerates rarely from seeds3,35. From Europe there is only few data available about biology and ecological preferences. E.g. Fried et al.3 reported biological traits of French A. psilostachya seed lots and established populations’ habitat as well as soil preferences. Obviously, belowground spreading rates are quite high what can be deduced from the huge dominant stands in Italian coastal dunes21. Djemaa35 documented germination rates of only 3% in French seed lots. This can be due to unfavorable pollination conditions or to population genetics (e.g. inbreeding depression).
While population genetics and invasion history of A. artemisiifolia is relatively well analyzed36,37,38,39,40,41,42,43,44,45,46,47,48,49, no research group has yet studied the population genetics of the clonal A. psilostachya neither in the native nor in its invasive range. No detailed taxonomic study was performed on this difficult diploid-polyploid complex since the monograph by Payne13. Greatly varying chromosome counts at diploid, tetraploid, hexaploid and octoploid levels are documented from North American populations of A. psilostachya50,51,52. In addition, reports from North America showed considerable morphological13,24 and phytochemical variation51 of A. psilostachya. But none of the authors linked this trait variation to ploidy levels.
In case of clonal species like A. psilostachya, one has to adapt methods when analyzing population genetic structure53,54. By setting the ranges for screening multilocus genotypes and adjusting the admixture models55, the outcome of the analyses (e.g. what is accepted as clonal offspring) may be influenced. Newly introduced populations of partially clonal plants tend to sustain the initial within-population genetic structure56. Bottleneck effects as well as gene flow have less or even almost no consequence on the population genetics of clonal species57.
Meyer et al.45 developed nuclear SSR-markers (Simple Sequence Repeats) for A. artemisiifolia and tested them in a population of A. psilostachya. These authors claimed that most of these markers are useful in describing western ragweed population genetics. In the present paper, we utilized these SSR markers to dissect the population genetic structure of A. psilostachya in Europe, providing insights into the genetic differences considering varying population sizes and local introduction histories.
We therefore aim to answer the following questions: (1) How strong is the genetic differentiation of A. psilostachya populations in Europe? (2) Is there an effect of different degrees of clonality within the populations on the invasion process and success across Europe? (3) How are genetic diversity patterns correlated to presumed age of populations (i.e. arrival date) and/or population sizes? (4) Are there different possible centers of introduction detectable based on the genetic structuring? (5) How much do population genetic characteristics differ between the two successful Ambrosia invaders, i.e. the annual A. artemisiifolia and the perennial A. psilostachya?
Results
Based on extensive studies on literature and herbarium collections reviewed by the first author, we sampled 60 populations from Middle Sweden to Southern Italy and from Spain to Croatia. From 1005 analyzed individuals representing 60 European populations, 50 individual samples were excluded from further analysis due to unreliable genotyping fragment patterns and/or missing data. Therefore, running 955 remaining individuals, we could avoid that some R applications treat missing states as novel alleles which would lead to bias in our results58. The frequencies of null alleles estimated over all populations ranged between 0.01 and 0.27 (mean over all loci: 0.09 with original as well as clone corrected data). Significant linkage disequilibrium was detected among loci as well as among populations and regions.
MLG diversity and clonality
The 15 SSR markers utilized showed an average of 2.56 ± 0.74 alleles per locus (= Na, ranging from 1.45 to 3.98). After clone correction, we gained 2.45 ± 0.63 alleles per locus (Table 1). Based on a minimum genetic dissimilarity threshold of 0.5, we identified 792 unique multilocus genotypes (MLGs) among the 955 originally analyzed individuals. Based on Bonin et al.59, our 15 loci should be adequate to detect a reliable number of unique MLGs. When comparing the genetic parameters calculated with and without clone correction (Supplementary Table S1) it is evident that there are hardly serious differences. Consequently, we discuss preferably the results of the analyses based on clone corrected (cc) data.
Assuming that individuals belonging to the same MLG within a given population belong to the same genet, the ratio of the number of MLGs to the number of individuals genotyped in a population (G:N) gives an estimate of the degree of clonality within the populations (see Table 1). Mean G:N was 0.84 ± 0.19; the lowest value (0.33) was found in population Psi-33 (Foro di Ortona2), whereas Psi-34 (Marina Lesina) from the same region (Southern Adria, Bari) showed a ratio of 1.0 (Table 1). The highest mean G:N ratio at the regional level was observed in populations surrounding Montpellier (0.98 ± 0.04) and along the northern Adriatic coastline (= region Venice, 0.91 ± 0.12). The populations from Northern Europe were measured by an average G:N of 0.66 ± 0.21, indicating higher levels of clonality towards the north.
In 37 out of 60 populations we could identify identical MLGs within the population (Supplementary Table S2). Mostly, few ramets produced few clonal offspring, but some MLGs were represented by five and more sampled ramets. In case of Psi-57 (Bydgoszcz, Poland) one single MLG proliferated to even ten ramets sampled on an area of 1000 m2. Identical MLGs could be detected either in populations in the close neighborhood (Psi-52 and 53: Gerwisch 1 and 2, or Psi-46 and 47: Barcelona: Martorello and Montmelo) or in populations up to a geographical distance of 170 km (Psi-8 and 9: Eraclea Mare and Mali Lošinj, the latter indicating long-distance dispersal of vegetative propagules.
Genetic diversity
Table 1 gives the details, and Supplementary Table S1 and Fig. S1 some additional statistics about basic regional differences of genetic diversity. The Shannon diversity index (H in Table 1) as a unifying measure of genetic diversity varies from 0.99 to 3.00. Evenness (E) as an alternative descriptor of genetic diversity is highly correlated with the Shannon index (Spearman’s r = 0.642; p < 0.001) and ranged from 0.52 to 1.0. Regional differences of H and E were significant in total (p = 0.005, and p = 0.011 resp.), but differences in H as well as E were significant only between N Europe (H = 2.08 ± 0.57, and E = 0.82 ± 0.14 resp.) and Venice (H = 2.64 ± 0.28 and E = 0.95 ± 0.08, resp.) (see Supplementary Table S1).
The mean allelic richness (Na) across all populations was 2.45 ± 0.62 when only unique MLGs were analyzed (cc, Table 1), and 2.65 ± 0.74 when all individuals were included (ori, Supplementary Table S2), being not significantly different (Mann–Whitney-U-Test: p = 0.350, see Supplementary Fig. S1). Na was not correlated (Pearson) with presumed population age, total population size or latitude, but proved to differ significantly by regions (Supplementary Fig. S1, all over Kruskal–Wallis-Test: p < 0.001), i.e. mean Na for the populations from Northern Europe was significantly (p < 0.001) lower than the mean for all other populations, regardless, if the calculations were either based on the original or the clone corrected data set.
The mean observed heterozygosity (Ho, Table 1) ranged from 0.28 (Psi-55: Falkenberg in Sweden) to 0.75 (Psi-44: Platy el Prat, Barcelona). Pearson correlation coefficients showed that Ho was negatively correlated to latitude (R = − 0.327, p = 0.011). ANOVA and consecutive Tukey-HSD-Test for significant group median differences revealed no effect of regions (p = 0.371), population age (p = 0.457), or population size (p = 0.069) on Ho.
Average expected gene diversity (He) across all populations was 0.43 ± 0.13 with the lowest within-population diversity (He = 0.18) observed in the Swedish population (Psi-55, Falkenberg) and the highest (He = 0.63) observed in population Psi-6 (Alberoni, Lido di Venice) (Table 1). Regional differences in He were significant in general (p < 0.001), and specifically differing when He median values of Northern Europe populations are compared with those from Montpellier (p = 0.026) and from Venice (p < 0.001). He was also negatively correlated to latitude in both data sets (cc: R = − 0.365, p = 0.004; ori: R = − 0.354, p = 0.006). There were no differences of He with respect to population size (p = 0.103), but differences by age classes (all over p = 0.049; Supplementary Table S1). However, specific age classes showed no significant pairwise differences.
The mean inbreeding coefficient (Fis) was − 0.19 (± 0.30) (with negative Fis values appearing in 38 out of 60 populations (Table 1). When analyzing median Fis values with respect to regions significant differences (Kruskal–Wallis-Test: p < 0.001) were detected; populations of Venice differed significantly from those of Northern Europe (p = 0.004) as well as from those of Barcelona (p = 0.014). Fis showed no difference with respect to population size (p = 0.987). Age matters for Fis in general (Kruskal–Wallis-Test: all over p = 0.037); but the differences between the specific age classes were not significant with respect to median Fis.
All populations showed significant deviations from Hardy–Weinberg equilibrium what indicates deficiencies of heterozygotes all over our sampled populations.
Genetic structure
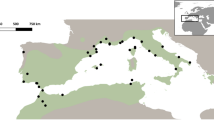
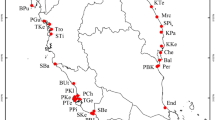
Bayesian clustering analysis performed on either the original and the clone corrected data set revealed that populations could be assigned in both cases basically to six genetic groups (Fig. 1 & Supplementary Fig. S2). Both, the original as well as the clone corrected data set, revealed K = 6 and K = 14 as most likely number of groups what was supported by the log likelihood lnP(D) calculations (Supplementary Fig. S3). Interestingly, the six genetic groups correspond largely with geographical locations, which could be partly affiliated to predefined regions (Figs. 1, 2, column “region” in Table 1). The centres of these regions were characterized as either important European trading harbors in Southern Europe (i.e. Barcelona, Bari, Genoa, Montpellier, and Venice) probably functioning as primary starting points of introductions, or the whole non-Mediterranean “Northern Europe” region (including Central Europe). The populations in Northern Europe (corresponding largely to the yellow genetic group in Fig. 2) are scattered over four different countries and could not be affiliated to any specific trading port. However, three out of eight northern populations (Psi-50, Hoyerswerda in Germany and Psi-56, Szczecin in Poland, both marked blue; and Psi-55, Falkenberg in Sweden, marked orange;) showed a completely different genetic constitution than the rest of the Northern European populations. They obviously share genetic variation mainly distributed in the Mediterranean regions of Barcelona and Bari, respectively.
Assignment of 792 European Ambrosia psilostachya individuals to color coded genetic cluster levels K = 6 (a) and K = 14 (b) based on clone corrected SSR-data analyzed in STRUCTURE. Populations are arranged to six geographic regions (main harbor regions: Genoa, Montpellier, Barcelona, Bari, Northern Europe, Venice).
Map of sampled Ambrosia psilostachya populations in Europe. Pie charts show proportions of membership of individuals for the K = 6 groups of Bayesian Clustering (sizes of the pie charts refer to the population sample size; 8 to 20 individuals). The base map was generated using the free of charge software QGIS 3.22 (https://www.qgis.org/en/site/forusers/visualchangelog322/index.html); pie charts were created using MS Excel included in MS Office Professional Plus 2019.
Generally, the degree of admixture at the population level differs fairly between regions and clusters. The populations in the North and in Spain are commonly most homogenous, i.e. dominated by individuals affiliated to a single genetic group. Most populations along the coastline of Tuscany (east of Genoa) also showed relatively low levels of admixture. The populations around Montpellier, in the Venice region (including the Po valley), and along the Southern Adriatic coastline (Bari region) are characterized by high degrees of admixture on average. Given these high levels of admixture in Bayesian Clustering, Principal Coordinate Analysis not surprisingly indicated relatively weak genetic structure (Supplementary Fig. S4) with the first two axes explaining only 8.6%, and 6.4% resp., of the total genetic variation.
Mantel tests imply significant isolation by distance (Supplementary Fig. S5). Spatial autocorrelation analysis revealed significant geographic correlation among all 60 populations (p = 0.001) as well as among the populations within regions (each with p = 0.001). Spatial autocorrelation test of the whole data set (Supplementary Fig. S6) revealed that the smallest distance classes (± 200 km) showed a clear positive relatedness of the individuals (r-values higher than expected, going far beyond the 95% confidence level) whereas at the intermediate geographical distances the relationship was rather lower than expected. However, towards one of the highest distance classes (about 1800 km) the relationship got positive again. The spatial autocorrelation analysis within the single regions (Supplementary Fig. S7) gave again positive relatedness of the individuals at low geographical distances within the respective regions.
Genetic differentiation among geographic regions
Hierarchical AMOVA on the full data set and all 955 individuals (ori data set) affiliated to the predefined regions showed that 10.4% of genetic variation occurred among these regions, and 40.3% of the genetic variation was found among populations within regions (Table 2). Most genetic variation was observed among individuals within populations (49.2%). The AMOVA performed on unique MLGs (cc data set) showed similar results: 9.9% of the genetic variation was found among regions, 34.0% among populations and 56.1% among individuals. Particularly within populations that are geographically associated to the surrounding of Barcelona as well as within the populations from Northern Europe (Netherlands, Germany and Poland) genetic variation among populations was significantly higher than among individuals (Table 2).
Discussion
Clonality remarkably affects the population genetics of introduced A. psilostachya
We identified substantial clonal reproduction within populations of A. psilostachya, but also dispersal of clonal offspring between populations throughout Europe. Duplicated MLGs were found in 62% of the analyzed populations. This can be expected from the morphological traits of creeping roots and added buds producing shoot sprouts from belowground34. We provide prove for this obvious species trait based on genetic data for the first time. Expecting an important role of clonal reproduction in A. psilostachya our sampling strategy was to collect individual ramets at a minimum distance of one meter. Due to our sampling strategy, we expected that our sampling might catch enough ramets from different genets to perform a reasonable analysis of genetic variation. Wagner & Beals24 stated, from their explorative analysis of A. psilostachya in the field, that clonal exploration of genets might cover many meters distance. We also observed secondary thickening of the horizontal roots as well as expansion of still connected roots of three meters length. This allows the assumption of a relatively long-lived clonal integration.
Besides clonal propagation, sexual reproduction plays a role in population dynamics of A. psilostachya34. The degree of sexual reproduction can vary from one individual clone to another by many to no fruits produced31. Furthermore, the latter authors stated that individual clones may differ also by specific, but in their study not detected, morphological characters. In general, identification of different genets can be indicated from MLG heterogeneity at the population level. Interestingly, our analysis provided evidences in various directions. Based on Wagner & Beals24 clonal growth can expand up to hundreds of m2 in A. psilostachya. Consequently, we expected to find clonal duplicates, especially in older spatially structured populations that had enough time to expand by hectares. Interestingly, this phenomenon could be found only in one case: Psi-51 and Psi-52 (two locations about 200 m apart, in a former military training area in Germany), but not in the case of subpopulations along the coast near Venice (Isola Verde 1, 2, 3; Rosolina Mare 1, 2; Lido di Volano 1, 2) – each population at only about 100–200 m apart from its neighbor. Lack of human disturbance due to conservancy regulations may have limited these small parceled coastal populations in the Venice region to spread one hundred or more meters or more. In contrast, the military training area in Germany was formerly utilized by tanks that can easily transport contaminated sandy soil containing ragweed root fragments60 over hundreds of meters. Dispersal of either seeds or clonal fragments of A. psilostachya over long distances of up to 170 km from and to coastal dunes was also proved by our data (Supplementary Table S2). As carrying vector, we are considering contaminated vehicles used for construction of infrastructure along beaches. Successful transportation of seeds by vehicles is well documented, at least for A. artemisiifolia in Europe61,62,63.
Another likely mode of dispersal is transportation by Adriatic Sea currents: the main direction of sea current along the Italian Adriatic coastline from North to South64 could have transported plant fragments. This would fit to the fact that the first occurrences of A. psilostachya were documented in areas surrounding Venice in the late thirties of the twentieth century whereas the Bari region was occupied decades later (Supplementary Table S3). Unfortunately, nothing is known about the survival of seeds or root and shoot fragments of our study species in sea water. However, Fumanal et al.65 showed that seeds of the related taxon A. artemisiifolia partially succeeded drifting in water and were still germinable after several days of inundation. Dispersal of clonal fragments of the invasive Carpobrotus edulis in floating sea water by sea currents up to 250 km within a few days was modelled comprehensibly based on the high survival rates of its clonal fragments in sea water66. Consequently, additional physiological experiments might enlighten the potential for transportation of plant fragments by sea current in case of A. psilostachya. However, our genetic data are not in conflict with this potential dispersal vector.
Genetic diversity varies along spatial and/or temporal trajectories
Parameters on genetic diversity can elucidate processes and directions of invasions. Invasion processes of plants with at least partial clonal reproduction tend to conserve diversity phenomena of initial colonization and subsequent local dispersal67 due to successful reproduction without seeds. Even local spread by ramet fragments can be indicated from such genetic data, since we proved local spread of unique MLGs over distances of up to 170 km (Supplementary Table S2).
Population genetic parameters may differ when MLG duplicates are either involved or removed from the data set53,67,68. In contrast, we found no significant differences at the population level when comparing allelic richness, Ho, He, and Fis for the whole data set calculated either for the original or the clone corrected data. However, all these parameters except for observed heterozygosity (ori: p = 0.090, cc: p = 0.069) differed significantly by regions (Supplementary Fig. S1a, Kruskal–Wallis-Test, p = 0.005) and partially − only for young populations − by age of the population, but not by population size (p = 0.971). Interestingly, mean observed heterozygosity Ho was marginally not significantly lower in Northern Europe populations than in all Mediterranean populations. Therefore, arrival and long-lasting persistence of genets from different source populations seems to be typical for A. psilostachya, but repeated/multiple introductions happened more frequently towards the south.
Our values for observed heterozygosity (mean Ho: 0.41 ± 0.09) of the perennial A. psilostachya correspond to Ho-estimate levels of the annual A. artemisiifolia (0.21–0.76) given by various authors (i.e. 38,44,45,49,69). Van Boheemen et al.46 gained on average lower Ho-values (0.16–0.30) for A. artemisiifolia, likely due to the use of SNPs (not SSRs) and to admixture processes. Mean values of expected heterozygosity of A. psilostachya were similar for the original and the clone corrected data. This corresponds to invasive Carpobrotus acinaciformis showing also no effect of clone correction in this respect70. Differences of Ho and He in A. psilostachya were significant when comparing the populations of Northern Europe, Bari and Barcelona regions (Mann–Whitney-U-Test: p < 0.001). Thereby, Northern Europe and Barcelona cover populations with low levels of admixture (Fig. 2).
High levels of heterozygosity excess in small populations
We found strikingly low values of Fis, far below zero for both the original as well as the clone corrected data in most regions except for Venice and Montpellier. Such negative values indicate excess of heterozygosity. Stoeckel et al.71 demonstrated some slight but not significant difference of original and clone corrected Fis -values for Prunus avium in France. Even more strikingly negative Fis population means were found in Carpobrotus acinaciformis that reproduces indeed almost exclusively by clonal fragmentation: Suehs et al.70 found very strong heterozygous excess (− 0.659, on average) for the original data set (calculated for all sampled ramets) and still significantly negative values (− 0.371, on average) for clone corrected data. In comparison, the related C. edulis showed rather homozygosity excess possibly due to partial selfing.
The strictly negative Fis -values of many A. psilostachya populations are in contrast to those of the annual A. artemisiifolia that turned out to have significantly positive deviation from Hardy–Weinberg equilibrium in practically all publications available38,39,40,43,49. A. psilostachya populations in Europe are exposed to various effects that may cause distinct excess of heterozygosity. In case of small absolute population sizes, Balloux72 stated that there is a clear tendency to heterozygosity excess due to small effective population size (i.e. few sexually reproducing individuals). Such could hold for our small and old populations (relicts) in Northern Europe, where we have to assume problems with seed production and the dominancy of few well-adapted clones. But very small-sized Mediterranean populations with high genetic variation possibly represent remnants of formerly bigger populations that eroded along the sandy coastline (i.e. Psi-15, Psi-31, and Psi-34). The longevity of successful, but few genets in isolated populations can go along preference of favoring basically heterozygous individuals over homozygous due to heterosis73 and inducing an excess of heterozygosity also in neutral loci of the genome by hitch-hiking71.
Balanced Fis values were found in larger populations. However, in this case low sexual reproduction rates and spatially restricted dispersal may cause sub-structuring of even large populations71. Such could have happened with three nearby populations Psi-10, -11, and -12 (Isola Verde, dunes south of Venice, subpopulations only 40 and 100 m apart) that possibly developed from one founder population a long time ago, because they show ± the same degree of admixture (Fig. 2), but interestingly no clonal duplicate in common (Supplementary Table S2); on the other hand, our sampling could not detect any clonality within the single samples of these very nearby populations. Nevertheless, we found significant linkage disequilibria among loci, populations, and regions, which is – together with negative Fis—a main characteristic of asexual or partially asexual (clonal) populations57,68. As self-incompatible species produce no offspring through selfing, this can lead to a significant deviation from random mating (i.e. in small populations), significantly influencing Fis and generating heterozygote excess72. This factor might have caused genetic erosion of some few older populations of A. psilostachya. Distinct negative Fis means at population level may also be indicative of an insufficient number of MLGs represented in the samples74.
Population genetics reflect the age of small populations (old relicts versus young founders)
There is no clear mono-factorial picture in our data. In a very isolated small population with very few ramets (Psi-34, Lesina Marina, Bari region) all 20 sampled ramets (out of a total of 60 ramets) turned out to represent unique MLGs (G/N = 1, Fis = 0.13). Comparable situations were observed in Psi-31 (Senigallia, Venice region) and Psi-15 (Boccasette, Venice) with population totals of 80 and 60 ramets, and 18 sampled ramets each that were almost all unique MLGs (G/N = 1, and 0.89 resp.; Fis = 0.15, and 0.03 resp.). In all three cases the sites suffered from coastal erosion at the front line of old grey dunes. This argues for historical (old) admixture at place. Other small populations like Psi-27 (Cerasomma, Genoa region, 15 out of 40 ramets sampled), Psi-33 (Foro di Ortona2, Bari, 8/80 ramets) and Psi-49 (Tavernes de Valldigna, Barcelona, 16/100 ramets) represent spatially delimitable populations of only few m2 each, at young roadsides that were obviously of recent age. While Psi-27 was represented only by unique MLGs (G/N = 1), the other two populations comprised of several multiplied genets (G/N = 0.33, and 0.50 resp.). In consequence, the mean Fis of Psi-27 was moderate (− 0.29) whereas the other two populations showed strongly negative values of Fis = − 0.70 and − 0.79, respectively. These very low population means of Fis might indicate significant undersampling74 or an initial population undergoing a bottleneck phase67,72, respectively. Based on our data we could not identify any bottleneck effects, in general. Small but older populations like Psi-15, Psi-31 and Psi-34 obviously consisted of several MLGs that may have survived since foundation due to clonal persistence and may therefore still represent the genetic structure of the initial colonization stage.
Population genetics comparison between the perennial and the annual invader
Most population genetic studies on the annual A. artemisiifolia were performed at the continental35,38,39,42,45,46,48,49 or regional40,41,43,44 scale using different SSRs, AFLPs or SNPs. These macro-scale analyses, as well as the local analysis by Kropf et al.47, based on AFLPs, demonstrated high admixture rates for nearby populations due to probable high levels of gene flow and repeated introductions from multiple sources. Only Kočiš Tubić et al.43, based on SSRs, found some decrease of genetic differentiation between populations in Serbia that might be correlated to geographical distance. The macro-scale analyses of A. artemisiifolia population genetic studies cited above deduced repeated introductions including several genetic consequences75,76 and secondary spread at high rates for the invasive range of the annual A. artemisiifolia across Europe, resulting in the lack of isolation by distance. However, we found that perennial A. psilostachya also represents an invasive species that shows no clear isolation by distance considering all populations and therefore, also a highly dynamic, ongoing invasion history, especially in Southern Europe. Isolation by distance was found in some regions only; although the respective R-values were low (Supplementary Fig. S5).
In case of A. psilostachya, we found relatively high degrees of admixture in some Mediterranean regions but not for the North European and Spanish populations (Figs. 1 and 2). The degree of admixture is specifically high in the Venice region (see inset of Fig. 2) where almost all sandy beaches are populated by A. psilostachya. While A. artemisiifolia is well known as agricultural weed transported throughout Europe in crop seed containments12,77,78, therefore, resulting in high degrees of admixture all over Europe, A. psilostachya, in contrast, rarely occurs on crop fields3, but in more natural habitat types. Therefore, and due to its main vegetative reproduction mode, current genetic diversity of populations represents rather the diversity of the founder population. This is most plausible for North European introductions that have survived for decades clonally. In case of Southern Europe, the older populations might have added slightly differing genotypes generated from locally produced seeds. At least in populations from southern France, Fried et al.3 showed that A. psilostachya is able to produce viable seeds at rates of 10 − 20%.
In general, founder populations of species expanding their range undergo selection of genotypes resistant to environmental constraints. Additionally, surviving clones do not need to reproduce by seeds due to bottleneck situations and unfavorable environmental stressors. Northern European initial populations of A. psilostachya might have faced this problem, indicated by their low Na and He, but also by highly significant negative Fis-values. To form extant populations via vegetative expansion takes time and our invasive study species expands in space mostly vegetatively. This is why all populations that comprise ≥ 50,000 individual ramets are at least at an age of 60 years or older. Genetic differentiation among populations within a region is lowest for Venice region where there is a high density of populations or subpopulations. If populations are geographically closer in this region, more dispersal events of single clonal fragments among them is to be expected. This is indeed the case for several adjacent populations around Venice (Supplementary Table S3).
Conclusions
Parameters describing population genetics of the invasive A. psilostachya did not differ with respect to calculations based on all sampled ramets or on unique genets only. This indicates that the sampling was sufficient for respective estimates. The strikingly negative inbreeding coefficient of small populations indicates the long-term persistence of well-adapted heterozygous clones. Under harsh environmental conditions, survival and expansion at the population level by clonal offspring can, therefore, be advantageous against the establishment by seeds79,80. Establishment and, specifically, spatial expansion can be less risky if root sprouts are produced that overcome stressful periods by resource allocations from the connected belowground mother individual organs81. Seedlings are commonly less costly but they easily face severe problems along with germination and juvenile growth if the environment is too harsh. In case of A. psilostachya, the vegetation period in Northern Europe is simply too short to produce ripened seeds, but long enough to assimilate and store carbohydrates belowground for renewal next season. In consequence, population genetics of the clonal A. psilostachya resembles the historical constitution of the founder populations. Change rates of population genetic parameters during the invasion process may be rather low, either with respect to expansion or reduction of clonal patches. Local extinction of such clonal species may take far longer than in short lived outcrossing annuals79. If genets get very old, they can accumulate somatic mutations, pathogens and epigenetic traits that may also contribute to genetic variation, hitchhiking even in neutral markers81. A future comparative detailed study of big and small, as well as, old and young populations would help to elucidate such processes of clonal population dynamics using population genetic parameters.
We learned that in clonal invaders, like western ragweed, even small populations may have a conservative history in that successfully established clones persist long time at the same place. Furthermore, bottleneck effects may be overcome easier than in outcrossing populations. We found a clear difference in naturalization dynamics and secondary spread throughout Europe. The perennial A. psilostachya forms long-lived local populations in Northern Europe but, recently, also larger populations towards the south. It performs far better with respect to recent spreading either by root fragments or seeds in the Mediterranean. But western ragweed is bound to instable habitat types everywhere, like river beds and coastal dunes. Prospectively, the invasion history in Europe might become clearer after consideration of North American source populations of western ragweed.
Methods
Study species and population sampling
In 2015 and 2016 a group of specialists evaluated several herbarium specimens of A. psilostachya from European herbaria11. Based on this experience we draw a map of ragweed occurrences in Europe to select for representative populations stretching the whole European invasive distribution range. Karrer et al.29 and Montagnani et al.21 report that some occurrence data from literature were due to misidentifications, named “A. maritima” in the south (Italy, Spain) or “A. artemisiifolia” in the north, respectively. Our sampling of verified A. psilostachya finally resulted in 60 populations listed in Table 1 & Supplementary Table S4. A. coronopifolia Torr. & A. Gray that was and is still in use in some databases82,83 and older literature is synonym to our taxon. Ten of the selected populations were tested for nuclear DNA-contents providing graphs that indicate identical ploidy levels relative to the respective reference (Supplementary Table S5).
Herbarium and literature studies revealed data on the first documented introductions to different European countries (Supplementary Table S3). The population size of sampled populations was estimated roughly in the field based on the number of visible shoots (Table 1). Furthermore, the prospective age of the populations was estimated based on the data about first introductions to the ‘regions’ in the surrounding of important European harbors (Table 1 and Supplementary Table S3). All ‘regions’ were geographically distinct except for ‘Northern Europe’ that includes all introductions into temperate Europe. Italy was invaded at different times; along the Tyrrhenian and North Adriatic coast line in the first half of the twentieth century, but along the South Adriatic coast line in the second half of the twentieth century or even later. Consequently, the Italian populations were affiliated to three different regions (Supplementary Table S4).
In each population we took maximum three leaves per individual stem and dried them quickly using silica gel. Commonly, we sampled leaves from maximum 20 individual stems per population at a distance of at least one meter from each other, except for very small populations that comprised only from few stems mostly nearby. In total, leaves from 1005 individuals were sampled from eight European countries. Furthermore, we collected few complete specimens for documentation. Specimens from all populations were identified by the first author and deposited in the herbarium collection of the University of Natural Resources and Life Sciences Vienna (WHB, cf. Supplementary Table S4). We confirm that the authors sampled and handled the collected plant material in accordance with the relevant institutional, national, and international guidelines and legislation. Our species of interest is alien to Europe and therefore we naturally comply with the IUCN Policy Statement on Research Involving Species at Risk of Extinction and the Convention on the Trade in Endangered Species of Wild Fauna and Flora.
SSR genotyping and statistical analysis
For DNA extractions, one small fragment of each leaf sample was placed in a 2 ml microtube containing one steel bead and 400 µl extraction buffer84. Microtubes were agitated in a bead mill homogenizer, then incubated in a water bath 5 min at 95 °C, cooled on ice and centrifugated 2 min at 20,000 g. The resulting DNA extracts were stored at − 20 °C. Genotyping was performed at GENTYANE (INRAE, Clermont-Ferrand, France) based on 15 microsatellite loci (AMBELssr-EST71, -ill75, -ill101, -EST114, -EST150, -EST13, -ill64, -EST69, -ill55, -ill35, -ill02, -EST111, -ill18, -ill20, -EST54) previously developed at INRAE Dijon, France for A. artemisiifolia45. PCR products were labelled with one of four fluorescent tags (i.e. 6-FAM, NED, VIC or PET) and loaded on an ABI 3730XL capillary DNA analyzer (Applied Biosystems) using the size standard GS500 LIZ. We used Peakscanner version 1.0 (Applied Biosystems) to read allele sizes. Poorly performing individuals with missing genotype call were excluded from further analysis to avoid bias in the data analysis58.
The presence and frequency of null alleles was checked for each marker using R-package “PopGenReport” version 3.0.485 estimating a bootstrap confidence interval for each locus using the methods of Brookfield86.
For identification of unique multilocus genotypes (MLGs) and genotypic (clonal) diversity of A. psilostachya the minimum genetic distance was calculated from a relative dissimilarity distance matrix (threshold = 0.558). The number of 15 loci used in our study fits to the needs for detecting unique MLGs59. Using these identified MLGs per population (“original” [ori] data set), we estimated the following components of genotypic diversity in each population: Shannon’s diversity index (H), genotypic evenness (E), and the ratio of number of MLGs to the number of individuals analyzed (genets to ramets = G:N). G:N indicates the degree of clonality within the population. A value of one indicates purely sexual reproduction, while a value near zero indicates purely clonal reproduction.
In clonal organisms such as A. psilostachya, asexual reproduction can result in a single genet being represented by multiple physiologically independent ramets87. Thus, we identified shared multilocus genotypes (MLGs) within or (even) across different populations using R-package “poppr” version 2.8.258, which can be used to infer clonal membership to the same genet87. We kept only one of each MLG per population in the “clone corrected” (cc) data set that served finally for additional calculations of the genetic diversity measures H, E and allelic richness.
As the concept of a population representing a group of interbreeding individuals is not straight forwardly applicable to species with clonal as well as sexual reproduction significant deviations from Hardy–Weinberg equilibrium (HWE) can be expected, as sexual and asexual reproduction may occur in parallel88. To determine deviations from HWE in the 60 European populations of A. psilostachya GenAlEx version 6.50289 was used. Tests for conformity with HWE were based on Chi-square tests, determining whether the observed genotypic frequencies are deviating from expected frequencies by chance or due to a lack of random mating.
Furthermore, we checked for linkage disequilibrium (LD) based on 999 permutations using R-package “poppr” version 2.8.2 testing if populations are clonal (where significant disequilibrium is expected due to linkage among loci) or sexual (where linkage among loci is not expected), expressed by the index of association (IA) proposed by Brown et al.90. Fixation index (FST) matrices based on pairwise FST values were calculated for all pairs of populations and regions using R-package “hierfstat” version 0.04–2291.
In addition, population structure was assessed using STRUCTURE 2.3.492. The admixture model and correlated allele frequencies between populations were selected as specified by Falush et al.55 to determine the number of genetic clusters (K) best fitting the data93. The length of the burn-in period was 100,000 runs followed by 500,000 Markov Chain Monte Carlo permutations. 20 iterations were performed for each value of K ranging from 1 to 15. Optimal values of K were determined based on log likelihood values as described by Pritchard et al.92. Principal Coordinate Analysis was calculated to show the interference of all individuals based on their genotypic similarity using Nei’s genetic distances94,95.
In the next step, spatial autocorrelation analysis was performed using GenAlEx version 6.50389 to describe the degree of similarity among regions and among populations within the regions. In addition, isolation by distance (IBD) was analyzed using R-packages “adegenet” version 2.1.196 and “MASS”97 based on Mantel test with 999 replicates between a matrix of Euclidean distances and a matrix of geographic distances. Spatial autocorrelation test in GenAlEx was based on the distribution of permuted (rp) values under the assumption of no spatial structure and 999 random shuffling of individuals among distance classes.
All genetic diversity calculations including null allele frequency analysis, HWE, IA, and FST-matrices were performed for two data sets, one containing all individuals, including multiples (duplicates) of the same MLGs (“ori”), and a second data set including only unique MLGs, subsequently called “clone corrected” data set (“cc”). Allelic richness per population was estimated using a rarefaction approach to correct for differences in populational sample size using R-package “diveRsity” version 1.9.9098. In addition, we estimated observed heterozygosity (Ho), expected heterozygosity (He), and the inbreeding coefficient (Fis) for both, the ori and the cc data.
Confidence intervals for Fis were obtained using 999 bootstrap permutations. Moreover, a hierarchical Analysis of Molecular Variance (AMOVA99) for populations and regions was performed using the R-package “adegenet” version 2.1.196 with statistical significance testing based on 999 permutations.
To analyze the genetic structure among different groupings, each genetic parameter was at first tested by Kolmogorov–Smirnov for normality. In consequence, Ho (ori and cc) and Fis (ori and cc) were treated by ANOVA and consecutive Tukey HSD for testing significant group median differences, and all other parameters were tested by non-parametric ranked Kruskal–Wallis-Test followed by a post-hoc Mann–Whitney-U-Test for significant group median differences. We tested for differences by six regions, four population age classes and five population size classes applying a significance level of p < 0.05 after adjusting by the Bonferroni correction for multiple tests (cf. Table 1). Population age was estimated from documented first occurrences until 2019 in the respective neighborhood of the 60 populations (see Supplementary Table S3) and classified into age class 1 (5–20 years), 2 (21–60), 3 (61–99), and 4 (≥ 100). Population size was classified into 1 (40–99 ramets), 2 (100–999), 3 (1,000–9,999), 4 (10,000–99,999), and 5 (100,000–500,000).
Data availability
Source data on the distribution of Ambrosia psilostachya in Europe and the full FST-matrix can be provided on a reasonable request by the first author. All other data generated or analyzed for this study are included in this published article.
References
Van Kleunen, M. et al. Global exchange and accumulation of non-native plants. Nature 525, 100–101 (2015).
Simberloff, D. et al. Impacts of biological invasions: What’s what and the way forward. Trends Ecol. Evol. 28, 58–66 (2013).
Fried, G., Chauvel, B., Reynaud, P. & Sache, I. Decreases in crop production by non-native weeds, pests, and pathogens. In Impact of Biological Invasions on Ecosystem Services (ed. Vilà, M.) 83–101 (Springer, 2017).
Nentwig, W., Mebs, D. & Vilà, M. Impact of non-native animals and plants on human health. In Impact of Biological Invasions on Ecosystem Services (ed. Vilà, M.) 277–293 (Springer, 2017).
Smith, M., Cecchi, L., Skjøth, C. A., Karrer, G. & Šikoparija, B. Common ragweed: A threat to environmental health in Europe. Environ. Int. 61, 115–126 (2013).
Strother, J. L. Ambrosia L. in Flora of North America, Vol. 21 efloras.org. http://www.efloras.org/florataxon.aspx?flora_id=1&taxon_id=101325 (2007). Accessed 10 August 2022.
Oswalt, M. L. & Marshall, G. D. Ragweed as an example of worldwide allergen expansion. All. Asth. Clin. Immun. 4, 130–135 (2008).
Payne, W. W. Biosystematic studies of four widespread weedy species of ragweeds, Ambrosia: Compositae. PhD Thesis, University of Michigan (1962).
Burbach, G. J. et al. Ragweed sensitization in Europe—GA(2)LEN study suggests increasing prevalence. Allergy 64, 664–665 (2009).
Ghosh, B. et al. Immunological and molecular characterization of Amb P V allergens from Ambrosia psilostachya (western ragweed) pollen. J. Immunol. 152, 2882–2889 (1994).
Karrer, G. et al. Ambrosia in Europe. Habitus, Leaves, Seeds, 6 European Ragweed Species. Comparison of traits. EU-COST-Action FA-1203 ‘Sustainable management of Ambrosia artemisiifolia in Europe’. http://internationalragweedsociety.org/smarter/wp-content/uploads/6AmbrosiaSpecies.pdf (2016). Accessed 10 August 2022.
Essl, F. et al. Biological flora of the British Isles: Ambrosia artemisiifolia L.. J. Ecol. 103, 1069–1098 (2015).
Payne, W. W. A re-evaluation of the genus Ambrosia (Compositae). J. Arnold Arbor. 45, 401–438 (1964).
Müller-Schärer, H. et al. Cross-fertilizing weed science and plant invasion science. Basic Appl. Ecol. 33, 1–13 (2018).
Chapman, D. S. et al. Modelling the introduction and spread of non-native species: International trade and climate change drive ragweed invasion. Glob. Change Biol. 22, 3067–3079 (2016).
Mang, T., Essl, F., Moser, D. & Dullinger, S. Climate warming drives invasion history of Ambrosia artemisiifolia in central Europe. Preslia 90, 59–81 (2018).
Liu, X.-L. et al. The current and future potential geographical distribution of common ragweed, Ambrosia artemisiifolia in China. Pak. J. Bot. 53, 167–172 (2021).
Allard, H. A. The North American ragweeds and their occurrence in other parts of the world. Science 98, 292–293 (1943).
Greuter, W. Compositae (pro parte majore) in Compositae. Euro+Med Plantbase - the information resource for Euro-Mediterranean plant diversity (ed. Greuter, W. & Raab-Straube, E. von) https://europlusmed.org/cdm_dataportal/taxon/76610e67-b2d4-4aef-a785-c4555af5b150 (Accessed 22 August 2022).
Abramova, L. M. Expansion of invasive alien plant species in the Republic of Bashkortostan, the Southern Urals: Analysis of causes and ecological consequences. Russ. J. Ecol. 43, 352–357 (2012).
Montagnani, C., Gentili, R., Smith, M., Guarino, M. F. & Citterio, S. The worldwide spread, success, and impact of ragweed (Ambrosia spp.). Crit. Rev. Plant. Sci. 36, 139–178 (2017).
Vermeire, L. T. & Gillen, R. L. Western ragweed effects on herbaceous standing crop in Great Plains grasslands. J. Range Manag. 53, 335–341 (2000).
Reece, P. E., Brummer, J. E., Northup, B. K., Koehler, A. E. & Moser, L. E. Interactions among western ragweed and other sandhills species after drought. J. Range Manag. 57, 583–589 (2000).
Wagner, W. H. & Beals, T. F. Perennial ragweeds (Ambrosia) in Michigan, with description of a new, intermediate Taxon. Rhodora 60, 177–204 (1958).
Hansen, A. Ambrosia L. In Flora Europaea Vol. 4 (eds Tutin, T. G. et al.) (Cambridge University Press, 1976).
Sell, P. & Murrell, G. Flora of Great Britain and Ireland, Campanulaceae–Asteraceae Vol. 4, 513–514 (Cambridge University Press, 2006).
Pignatti, S. Flora d’Italia Vol. 3 (Edagricola, 1982).
Amor Morales, À., Navarro Andrés, F. & Sánchez Anta, M. Datos corológicos y morfológicos de las especies del género Ambrosia L. (Compositae) presentes en la Península Ibérica. Bot. Complut. 36, 85–96 (2012).
Karrer, G. Ambrosia. In Flora d’Italia 2nd edn, Vol. 3 (eds Guarino, R. & La Rosa, M.) 808–810 (Edagricola, 2018).
Rich, T. C. G. Ragweeds (Ambrosia L.) in Britain. Grana 33, 38–43 (1994).
Chauvel, B., Fried, G., Monty, A., Rossi, J. P. & Le Bourgeois, T. Analyse de Risques Relative à L’ambroisie à Épis Lisses (Ambrosia Psilostachya DC.) et Élaboration de Recommandation De gestion (ANSES, 2017).
Lawalreé, A. Les Ambrosia adventices en Europe occidentale. Bull. Jard. Botan. l’Etat Bruxelles 18, 305–315 (1947).
Karrer, G. Interessante Gefäßpflanzenfunde aus Österreich, 1. Neilreichia 12, 183–187 (2021).
Bassett, I. J. & Crompton, C. W. The biology of Canadian weeds. 11. Ambrosia artemisiifolia L. and A. psilostachya DC. Can. J. Plant Sci. 55, 463–476 (1975).
Djemaa, S. Caractérisation de la banque de graines de l'Ambroisie à épis lisses Ambrosia psilostachya DC (Asteraceae) et moyens de contrôle de cette espèce envahissante et allergène (Rapport de stage de Master 1 – Université de Montpellier 2 – Master IEGB, 2014).
Chun, Y. J., Le Corre, V. & Bretagnolle, F. Adaptive divergence for a fitness-related trait among invasive Ambrosia artemisiifolia populations in France. Mol. Ecol. 20, 1378–1388 (2011).
Genton, B. J. et al. Isolation of five polymorphic microsatellite loci in the invasive weed Ambrosia artemisiifolia (Asteraceae) using an enrichment protocol. Mol. Ecol. Notes 5, 381–383. https://doi.org/10.1111/j.1365-294X.2005.02750.x (2005).
Genton, B. J., Shykoff, J. A. & Giraud, T. High genetic diversity in French invasive populations of common ragweed, Ambrosia artemisiifolia, as a result of multiple sources of introduction. Mol. Ecol. 14, 4275–4285 (2005).
Gaudeul, M., Giraud, T., Kiss, L. & Shykoff, J. A. Nuclear and chloroplast microsatellites show multiple introductions in the worldwide invasion history of common Ragweed Ambrosia artemisiifolia. PLoS One 6, e17658. https://doi.org/10.1371/journal.pone.0017658 (2011).
Chun, Y. J., Fumanal, B., Laitung, B. & Bretagnolle, F. Gene flow and population admixture as the primary post-invasion processes in common ragweed (Ambrosia artemisiifolia) populations in France. New Phytol. 185, 1100–1107 (2010).
Gladieux, P. et al. Distinct invasion sources of common ragweed (Ambrosia artemisiifolia) in Eastern and Western Europe. Biol. Invasions 13, 933–944 (2010).
Li, X.-M., Liao, W.-J., Wolfe, L. M. & Zhang, D.-Y. No evolutionary shift in the mating system of North American Ambrosia artemisiifolia (Asteraceae) following its introduction to China. PLoS One 7(2), e31935. https://doi.org/10.1371/journal.pone.0031935 (2012).
Kočiš Tubić, N., Djan, M., Veličković, N., Anačkov, G. & Obreht, D. Microsatellite DNA variation within and among invasive populations of Ambrosia artemisiifolia from the southern Pannonian Plain. Weed Res. 55, 268–277 (2015).
Ciappetta, S. et al. Invasion of Ambrosia artemisiifolia in Italy: Assessment via analysis of genetic variability and herbarium data. Flora 223, 106–113 (2016).
Meyer, L. et al. New gSSr and EST-SSR markers reveal high genetic diversity in the invasive plant Ambrosia artemisiifolia L. and can be transferred to other invasive Ambrosia species. PLoS One 12(5), e0176197. https://doi.org/10.1371/journal.pone.0176197 (2017).
Van Boheemen, L. A. et al. Multiple introductions, admixture and bridgehead invasion characterize the introduction history of Ambrosia artemisiifolia in Europe and Australia. Mol. Ecol. 26, 5421–5434 (2017).
Kropf, M., Huppenberger, A. S. & Karrer, G. Genetic structuring and diversity patterns along rivers—Local invasion history of Ambrosia artemisiifolia (Asteraceae) along the Danube River in Vienna (Austria) shows non-linear pattern. Weed Res. 58, 131–140 (2018).
Sun, Y. & Roderick, G. K. Rapid evolution of invasive traits facilitates the invasion of common ragweed Ambrosia artemisiifolia. J. Ecol. 107, 2673–2687 (2019).
Li, F. et al. Patterns of genetic variation reflect multiple introductions and pre-admixture sources of common ragweed (Ambrosia artemisiifolia) in China. Biol. Invasions 21, 2191–2209 (2019).
Payne, W. W., Raven, P. H. & Kyhos, D. W. Chromosome numbers in Compositae. IV. Ambrosieae. Am. J. Bot. 51, 419–424 (1964).
Miller, H. E., Mabry, T. J., Turner, B. L. & Payne, W. W. Infraspecific variation of sesquiterpene lactones in Ambrosia psilostachya (Compositae). Am. J. Bot. 55, 316–324 (1968).
Del Amo Rodriguez, S. & Gomez-Pompa, A. Variability in Ambrosia cumanensis (Compositae). Syst. Bot. 1, 363–372 (1976).
Grünwald, N. J., Everhart, S. E., Knaus, B. J. & Kamvar, Z. N. Best practices for population genetic analyses. Phytopathology 107, 1000–1010 (2017).
Arnaud-Haond, S., Stoeckel, S. & Bailleul, D. New insights into the population genetics of partially clonal organisms: When seagrass data meet theoretical expectations. Mol. Ecol. 29, 3248–3260 (2020).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Watkinson, A. & Powell, J. Seedling recruitment and the maintenance of clonal diversity in plant populations—A computer simulation of Ranunculus repens. J. Ecol. 81, 707–717 (1993).
Balloux, F., Lehmann, L. & de Meeus, T. The population genetics of clonal and partially clonal diploids. Genetics 164, 1635–1644 (2003).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: A r package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281. https://doi.org/10.7717/peerj.281 (2014).
Bonin, A. et al. How to track and assess genotyping errors in population genetics studies. Mol. Ecol. 13, 3261–3273 (2004).
Guretzky, J., Anderson, A. & Fehmi, J. Grazing and military vehicle effects on grassland soils and vegetation. Great Plains Res. 16, 51–61 (2006).
Vitalos, M. & Karrer, G. Dispersal of Ambrosia artemisiifolia seeds along roads: the contribution of traffic and mowing machines. NeoBiota 8, 53–60 (2009).
Karrer, G. Das österreichische Ragweed Projekt—übertragbare Erfahrungen. The Austrian Ragweed Project—Experiences and Generalisations. Julius-Kühn-Archiv 445, 27–33 (2014).
Lemke, A., Buchholz, S., Kowarik, I., Starfinger, U. & von der Lippe, M. Interaction of traffic intensity and habitat features shape invasion dynamics of an invasive alien species (Ambrosia artemisiifolia) in a regional road network. NeoBiota 64, 155–175 (2021).
Orlić, M., Gačić, M. & La Violette, P. E. The currents and circulation of the Adriatic Sea. Oceanol. Acta 15, 109–124 (1992).
Fumanal, B., Chauvel, B., Sabatier, A. & Bretagnolle, F. Variability and cryptic heteromorphism of Ambrosia artemisiifolia seeds: What consequences for its invasion in France?. Ann. Bot. 100, 305–313 (2007).
González, L. et al. An Atlantic Odissey: The fate of invading propagules across the coastline of the Iberian Peninsula. In 15th Ecology and Management of Alien Plant Invasions (EMAPi) Book of Abstracts: Integrating Research, Management and Policy (eds Pyšek, P. et al.) 24 (Institute of Botany, Czech Academy of Sciences, 2019).
Ward, S. Genetic analysis of invasive plant populations at different spatial scales. Biol. Invasions 8, 541–552 (2006).
Halkett, F., Simon, J.-C. & Balloux, F. Tackling the population genetics of clonal and partially clonal organisms. Trends Ecol. Evol. 20, 194–201 (2005).
Kočiš Tubić, N., Djan, M., Veličković, N., Anačkov, G. & Obreht, D. Gradual loss of genetic diversity of Ambrosia artemisiifolia L. populations in the invaded range of central Serbia. Genetika 46, 255–268 (2014).
Suehs, C. M., Affre, L. & Médail, F. Invasion dynamics of two alien Carpobrotus (Aizoaceae) taxa on a Mediterranean island: I. Genetic diversity and introgression. Heredity 92, 31–40 (2004).
Stoeckel, S. et al. Heterozygote excess in a self-incompatible and partially clonal forest tree species—Prunus avium L. Mol. Ecol. 15, 2109–2118 (2005).
Balloux, F. Heterozygote excess in small populations and the heterozygote-excess effective population size. Evolution 58, 1891–1900 (2004).
Hansson, B. & Westerberg, L. On the correlation between heterozygosity and fitness in natural populations. Mol. Ecol. 11, 2467–2474 (2002).
Hewitt, A., Rymer, P., Holford, P., Morris, E. C. & Renshaw, A. Evidence for clonality, breeding system, genetic diversity and genetic structure in large and small populations of Melaleuca deanei (Myrtaceae). Aust. J. Bot. 67, 36–45 (2019).
Dlugosch, K. M. & Parker, I. M. Founding events in species invasions: Genetic variation, adaptive evolution, and the role of multiple introductions. Mol. Ecol. 17, 431–449 (2008).
Novak, S. J. & Mack, R. N. Genetic bottlenecks in alien plant species: influences of mating systems and introduction dynamics. In Species Invasions: Insights into Ecology, Evolution, and Biogeography (eds Sax, D. F. et al.) 201–228 (Sinauer Associates, 2005).
Karnkowski, W. Pest Risk Analysis and Pest Risk Assessment for the territory of the Republic of Poland (as PRA area) on Ambrosia spp., updated version. (Torun, 2001).
Karrer, G. et al. Ausbreitungsbiologie und Management einer extrem allergenen, eingeschleppten Pflanze – Wege und Ursachen der Ausbreitung von Ragweed (Ambrosia artemisiifolia) sowie Möglichkeiten seiner Bekämpfung. (Final Report, BMLFUW, Vienna, Austria). https://dafne.at/projekte/ragweed (2011). Accessed 10 August 2022.
Honnay, O. & Jacquemyn, H. A meta-analysis of the relation between mating system, growth form and genotypic diversity in clonal plant species. Evol. Ecol. 22, 299–312 (2008).
Vallejo-Marín, M., Dorken, M. E. & Barrett, S. C. H. The ecological and evolutionary consequences of clonality for plants mating. Annu. Rev. Ecol. Syst. 41, 193–213 (2010).
McKey, D., Elias, M., Pujol, B. & Duputiè, A. The evolutionary ecology of clonally propagated domesticated plants. New Phytol. 186, 318–332 (2010).
WFO Ambrosia psilostachya DC. http://www.worldfloraonline.org/taxon/wfo-0000137200 (accessed 21 July 2022).
Tomasello, S., Stuessy, T. F., Oberprieler, C. & Heubl, G. Ragweeds and relatives: Molecular phylogenetics of Ambrosiinae (Asteraceae). Mol. Phylogenet. Evol. 130, 104–114 (2019).
Délye, C., Matéjicek, A. & Gasquez, J. PCR-based detection of resistance to Acetyl-CoA carboxylase-inhibiting herbicides in black-grass (Alopecurus myosuroides Huds) and ryegrass (Lolium rigidum Gaud). Pest Manag. Sci. 58, 474–478 (2002).
Adamack, A. T. & Gruber, B. PopGenReport: Simplifying basic population genetic analyses in R. Methods Ecol. Evol. 5, 384–387 (2014).
Brookfield, J. F. Y. A simple new method for estimating null allele frequency from heterozygote deficiency. Mol. Ecol. 5, 453–455 (1996).
Harper, J. L. Population Biology of Plants (Academic Press, 1977).
Lambertini, C. et al. Genetic diversity in three invasive clonal aquatic species in New Zealand. BMC Genet. 11(52), 1–18. https://doi.org/10.1186/1471-2156-11-52 (2010).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: Genetic analysis in excel. Population genetic software for teaching and research—An update. Bioinformatics 28, 2537–2539 (2012).
Brown, A. H. D., Feldman, M. W. & Nevo, E. Multilocus structure of natural populations of Hordeum spontaneum. Genetics 96, 523–536 (1980).
Goudet, J. Hierfstat, a package for R to compute and test hierarchical F-statistics. Mol. Ecol. Notes 5, 184–186 (2005).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Kropf, M., Comes, H. P. & Kadereit, J. W. An AFLP clock for the absolute dating of shallow-time evolutionary history based on the intraspecific divergence of southwestern European alpine plant species. Mol. Ecol. 18, 697–708 (2009).
Nei, M. Genetic distance between populations. Am. Nat. 106, 283–292 (1972).
Jombart, T. adegenet: A r package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S 4th edn. (Springer, 2002).
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. diversity: An R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Acknowledgements
Financial support was given from the EU COST-Action 1235 SMARTER to R.H. for lab work, and to G.K. for travelling to herbaria and for sampling. Furthermore, G.K. was also aided from the BOKU-project NEOPHYTA. We appreciate very much samples provided by C. Montagnani, H. Karrer, E. Tordoni, F. Petruzellis, and I. Karrer.
Funding
The funding was provided by European Cooperation in Science and Technology, FA1203.
Author information
Authors and Affiliations
Contributions
The study was conceptualized by G.K.; G.K. coordinated and executed field sampling together with acknowledged colleagues; lab work was done by R.H. and V.L.; G.K., R.H. and M.K. were engaged in data analysis; G.K. wrote the first draft of the paper, R.H. and G.K. created the figures; all authors contributed to writing of subsequent versions and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karrer, G., Hall, R.M., Le Corre, V. et al. Genetic structuring and invasion status of the perennial Ambrosia psilostachya (Asteraceae) in Europe. Sci Rep 13, 3736 (2023). https://doi.org/10.1038/s41598-023-30377-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30377-6
This article is cited by
-
Environmental impacts on intraspecific variation in Ambrosia artemisiifolia genome size in Slovakia, Central Europe
Environmental Science and Pollution Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.