Abstract
Spiders have distinct predatory behaviours selected along Araneae’s evolutionary history but are mainly based on the use of venom for prey paralysis. Uloboridae spiders have lost their venom glands secondarily during evolution. Because of this, they immobilise their prey by extensively wrapping, and digestion starts with the addition of digestive fluid. During the extra-oral digestion, the digestive fluid liquefies both the prey and the AcSp2 spidroins from the web fibres. Despite the efficiency of this process, the cocktail of enzymes involved in digestion in Uloboridae spiders remains unknown. In this study, the protein content in the midgut of Uloborus sp. was evaluated through enzymatic, proteomic, and phylogenetic analysis. Hydrolases such as peptidases (endo and exopeptidases: cysteine, serine, and metallopeptidases), carbohydrases (alpha-amylase, chitinase, and alpha-mannosidase), and lipases were biochemically assayed, and 50 proteins (annotated as enzymes, structural proteins, and toxins) were identified, evidencing the identity between the digestive enzymes present in venomous and non-venomous spiders. Even enzymes thought to be unique to venom, including enzymes such as sphingomyelinase D, were found in the digestive system of non-venomous spiders, suggesting a common origin between digestive enzymes and enzymes present in venoms. This is the first characterization of the molecules involved in the digestive process and the midgut protein content of a non-venomous spider.
Similar content being viewed by others
Introduction
The success of spiders during evolution is frequently associated with venom1,2, web for prey capture and immobilisation3, and digestive enzymes for prey degradation. Spiders digest a large amount of food by regurgitating the digestive fluid from their midgut in a process known as extra-oral digestion (EOD)4,5. EOD is complemented by intracellular digestion at the digestive cells. Insects, which are the main prey of spiders, are principally composed of proteins, carbohydrates, and lipids. Thus, spider digestive fluid hydrolases are expected to be capable of catalysing the breakdown of these nutrients. Historically, digestive fluid has been studied for some decades in order to comprehend the enzymes involved in this process. Perret6 described the presence of proteolytic activity in venom contaminated with “saliva”, a consequence of digestive fluid extraction by electro stimulation. Mommsen7,8,9,10,11 assayed carbohydrases, such as amylase, chitinases, and hexosaminidases as well as peptidases and carboxyl esterases in the digestive juice (term replaced by digestive fluid) of Tegenaria atrica. Other studies on digestion in spiders reported the isolation of peptidase from the digestive fluid of Argiope aurantia12,13,14,15 and identification of metallopeptidases in spider digestion and serine peptidase inhibitors in the digestive fluid of this spider15. The ultrastructure of the midgut of Coelotes terrestris has been described in detail16. Joo et al.17 isolated a serine peptidase from the whole body of Nephila clavata with a pro-thrombin-like specificity. Furthermore, our group described the proteome/transcriptome analysis of spider midgut and its digestive fluid in Nephilingis cruentata18, elucidating the types and action mechanism of enzymes secreted in EOD and their action role in final intracellular digestion. Walter et al.19 corroborated these data through proteomic analyses of the digestive fluid of Stegodyphus mimosarum and Acanthoscurria geniculata.
Although the importance of digestive enzymes has already been demonstrated in different spider species, many spider venoms also contain different types of enzymes2,20,21, such as hyaluronidases, astacins, and serine peptidases. The overlapping of enzymes raises the possibility of contamination of the digestive fluid by spider venom and vice-versa, and the discussion of the importance and balance of enzymes in the digestive fluid and venom for prey digestion is still unclear.
Therefore, spiders without venom would make better models for studying enzymes exclusively involved in prey degradation.
Uloboridae, commonly known as hackled orbweavers, is a spider family composed of 19 genera and 291 species, and is the only spider family that has lost its venom glands secondarily during evolution22. Uloboridae spiders are cribellate orb-weavers that exhibit particular prey capture behaviour3. Their prey are immobilised by strong and extensive wrapping (up to 7000 wrapping movements), as observed in Philoponella vicina23, forming a package. Although other spiders only wet the portion close to their chelicerae with the digestive fluid24, Uloboridae spiders regurgitate their digestive fluid all over the package surface25. Thus, Uloboridae spiders are useful models for understanding the single role of the digestive system in prey degradation.
Based on previous proteomic data of the digestive fluid and the diet of Uloboridae spiders, enzymes such as peptidases, astacin-like metallopeptidases, cysteine peptidases and, carboxypeptidases; carbohydrases, amylase, and chitinases; and lipases were expected to be identified in the digestive tract of Uloboridae spiders. However, some studies have suggested that several enzymes, usually involved in the envenoming process, can also be expressed even in non-venomous animals26, highlighting the necessity and importance of comparative studies between the enzymes of the venom and the digestive system. One such enzyme is sphingomyelinase D (SMaseD). SMaseD (EC 3.1.4.41) is a hydrolase which catalyses the sphingomyelin cleavage. Structurally, these enzymes can be divided into two groups: classes I and II. The distinction between class I and II is defined by the presence of a single disulphide bond in class I and two disulphide bonds in class II. Class II is further divided into two subclasses: classes IIa and IIb27. Classes IIa and IIb of SMaseD are differentiated based on the dual substitutions at positions 95 (Gly → Asn) and 134 (Pro → Glu) in class IIb27. The SMaseD class I acts more specifically to hydrolyse sphingomyelin, class IIa acts on sphingomyelin and more substrates, and Class IIb has non-catalytic or reduced activity against sphingomyelin27,28,29, because of the hydrophilic environment at the entrance to the active site, due to the high polarity of Asn and Glu substitutions27.
In the current study, the digestive tract was isolated from Uloborus sp., homogenized and, was characterised by electrophoresis separation, enzymatic assays, and proteomic analyses. These data were compared with transcriptomic data from other Uloboridae species and proteomic, transcriptomic, and genomic data from other venomous spider families to understand the enzyme cocktail involved in digestion and envenoming, in order to comprehend the possible shared origin and evolution of enzymatic components in spider digestion and venom.
Results
Protein profile of the midgut of Uloborus sp
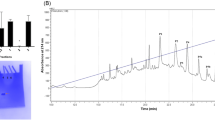
The SDS-PAGE profile in Fig. 1 shows proteins along the gel but is mainly distributed at 66–21 kDa. This protein molecular mass range is compatible with the molecular masses of other digestive enzyme profiles in spiders18,19 (Fig. 1).
SDS PAGE profile of midgut samples from Uloborus sp. About 10 μg of proteins from the midgut of Uloborus sp. was submitted to electrophoresis in a 12% polyacrylamide gel at 150 V. MWS—Molecular Weight Standard, 97 kDa—Phosphorylase B, 66 kDa—Albumin, 45 kDa—Albumin from egg, 30 kDa—Carbonic anhydrase, 20.1 kDa—Trypsin inhibitor and 14.4 kDa—Lysozyme. The gel was silver stained according to76.
Enzymes activities
According to Fuzita et al.18 and Walter et al.19, many classes of enzymes have been described in the spider midgut. In this study, 11 different enzymes within three major classes were evaluated: carbohydrases, peptidases, and lipases; the values of enzyme activities are shown in Supplementary Table S1 and Fig. 2.
Values of endopeptidases, exopeptidases, carbohydrases and lipase specific activities from the midgut diverticula (MD) of Uloborus sp. Enzyme assays were performed as shown in Table 1. N = 5 to endopeptidases (2A); N = 6 to exopeptidases (2B); N = 8 to carbohydrases (2C), except to alpha-amylase (N = 4) and N = 8 to lipase (2D). Colored bars represent specific activity, mean and ± standard deviation of biological replicates for a specific enzyme according to color legend. * Means no activity detected.
Cysteine peptidases and astacin showed the major activities of endopeptidases (Fig. 2A), mainly cysteine peptidases (21,400 ± 6700 mU.mg−1). No serine endopeptidase activity, such as trypsin or chymotrypsin, was detected despite the different substrates and pH conditions tested, namely: Z-Phe-Arg-MCA, Z-Gly-Gly-Arg-MCA, benzoyl-Arg-pNa, N-succinyl-Ala-Pro-Phe-pNa, and H–D-Phe-Pip-Arg-pNA (thrombin substrate) in a range of pH 3–8 (Table 1) with 10 mM CaCl2. Exopeptidases, represented by aminopeptidase and carboxypeptidase, presented activities of 0.9 ± 0.2 mU.mg−1 and 4.8 ± 0.7 mU.mg−1, respectively, (Fig. 2B).
Carbohydrate digestion involves enzymes such as alpha-amylase, chitinase, beta-N-acetyl-glucosaminidase (hexosaminidase), alpha-L–fucosidase, and alpha-mannosidase (Fig. 2, panel C). Alpha-amylase displayed the highest activity (200.2 ± 81.4 mU.mg−1) followed by mannosidase (4.4 ± 1.6 mU.mg−1), chitinase (4.3 ± 1.2 mU.mg−1), hexosaminidase (1.2 ± 0.2 mU.mg−1), and α-L-fucosidase (0.8 ± 0.2 mU.mg−1). All carbohydrases present an acidic pH optimum (4.5–5.5).
Lipase activity was measured using 2,3-Dimercapto-1-propanol tributyrate (DMPTB) as substrate. This is a generic lipase substrate, which might be hydrolysed by triacylglycerol lipases and allows the hydrolysis by other enzymes such as other esterases, including phospholipases A2 and B. Thus, the measured values of lipase activity (9.2 ± 1.8 mU.mg−1) (Fig. 2D) are a sum of these enzymes.
Proteomic identification
In total, 523 tryptic peptides were identified, of which, 50 proteins were matched with different databases. Among the 50 identified proteins (Fig. 3; Supplementary Table S2), 18 were structural proteins such as actin, calmodulin, and alpha-tubulin, or enzymes involved in cell metabolism such as aldehyde dehydrogenase, thioredoxin, and proteasome subunits. The other 33 identified proteins were correlated with proteins already associated with digestion by proteomic analysis of the digestive fluid of N. cruentata, S. mimosarum, and A. geniculata18,19, as shown in Supplementary Table S3.
The amount and protein categories by Uloborus sp. midgut proteome. Proteins were set apart into six different categories and their amount in Uloborus sp. midgut proteome: structural (7), carbohydrases (3), peptidases (21), phospholipases (1), toxins (7) and metabolism (11), summing a total of 50 proteins. Plot was generated in Excel.
Peptides mainly related to endopeptidases and exopeptidases were most abundant. Cathepsin L, cathepsin O (both cysteine endopeptidases), carboxypeptidase (metallo and serine carboxypeptidases), astacin-like metalloendopeptidases, and serine endopeptidases were the peptidases identified in Uloborus sp. (Supplementary Table S2). Regarding carbohydrate digestion, 23 peptides related to alpha-mannosidase, chitinase, and alpha-amylase activity were identified. No enzymes belonging to triacylglycerol lipase family were detected. However, an enzyme typical of venom, phospholipase D (or sphingomyelinase, SMaseD), was detected. In addition to phospholipase, U21, U24, and loxtox proteins were also found30.
Sphingomyelinase classification and evolution
Proteomic analysis of Uloborus sp. digestive system samples showed the presence of SMaseD (Supplementary Table S2). The alignment of spider SMaseD (Figs. 4 and 5) indicate that venom SMaseD belonged to class I and digestive SMase belonged to class IIb.
Spider venom and midgut SMaseD multiple sequence alignment and their classification by cysteine conservation. SMaseD of class I, class IIa and class IIb from L. venom gland were aligned against SMaseD from abdomen A. geniculata, midgut N. cruentata, digestive fluid S. mimosarum, and whole body S. dumicola and O. yesoensis in multiple sequence alignment (MUSCLE software), to classify the SMaseD found in the midgut/digestive fluid of spiders. Sequences with only one disulfide bond between Cys97 and Cys103 are classified as “Class I SMaseD” and sequences with an additional disulfide bond between Cys99 and Cys 250 are classified as “Class II SMaseD''. Red box: 1º disulfide bond cysteine conservation; Purple box: 2º disulfide bond cysteine conservation; Blue line/arrow; columns of amino acids positions hidden (95–108 and 240–253). Midgut/digestive fluid SMaseD sequences only conserve the four cysteine residues (Cys97, Cys99, Cys 103, and Cys 250). Alignment was generated in Jalview (https://www.jalview.org)81.
Spider venom and midgut SMaseD multiple sequence alignment, their classification by glycine and proline substitutions, and conservation of residues involved in catalysis. SMaseD of class I, class IIa and class IIb from L. venom gland were aligned against SMaseD from abdomen A. geniculata, midgut N. cruentata, digestive fluid S. mimosarum, and whole body S. dumicola and O. yesoensis in multiple sequence alignment (MUSCLE software) 80, to classify the SMaseD found in midgut/digestive fluid of spiders. Most of the class I and class IIa SMaseD conserve the glycine and proline (purple highlighted and red arrowed indicated) residues, and class IIb SMaseD has a substitution in glycine/proline residues. As well, the conservation of histidines (His57 and His93), glutamic acid (Glu77), aspartic acid (Asp79), lysine (Lys139), tyrosine (Tyr278), and tryptophan (Trp280), involved in ion magnesium coordination and substrate recognition. Purple box: conserved histidine, glutamic acid, aspartic acid, lysine, glycine, and proline; Green box: conserved tyrosine; Blue box: conserved tryptophan. Midgut/digestive fluid SMaseD sequences only substitute the glycine residue (Gly141). Alignment was generated in Jalview (https://www.jalview.org81.
A phylogenetic tree of Chelicerata (Supplementary Fig. S4) has three main groups of SMases represented by distinct Chelicerata species; however, were supported by low bootstrap values. Therefore, a maximum likelihood analysis of SMaseD exclusive from spider SMaseD sequences (midgut and venom) was performed to enhance the statistical bootstrap support (Fig. 6). The tree presents three main groups: (a) a group containing the proteome confirmed digestive SMaseD, classified as SMaseD IIb; (b) a group containing other spider venom and some Loxosceles laeta and L. intermedia SMaseD II; and (c) a group containing majorly L. laeta, L. intermedia and L. reclusa venom SMaseD class I (Fig. 6).
Maximum Likelihood phylogenetic tree from SMaseD sequences. A Maximum Likelihood phylogenetic tree analysis by IQ Tree software82 and coloured by ITOL (https://itol.embl.de)83, composed of midgut, venom gland sequences from spiders. Bootstrap test of phylogeny was applied with 1000 resampling. All sequences are listed in Supplementary Table S5.
Discussion
Due to lack of venom, Uloboridae spiders have developed unique strategies for prey capture, prey handling, and digestion. These strategies include behavioural changes such as resting and feeding positions. Several Uloboridae species rest on their webs in a cryptic position to hide their anterior legs for protection against predators31. However, members of the genus Uloborus, for example, change these postures during feeding. Esquivel et al.24, showed that the digestive fluid of these spiders damages their legs, indicating that the leg position changing behaviour during feeding is to avoid leg damage, suggesting that digestive fluid is rich in hydrolytic enzymes. However, to date, specific digestive enzymes involved in prey liquefaction are not known in this group of spiders. Insects, the main Uloborid prey, have a higher percentage of proteins than carbohydrates and lipids32 in their body composition. Therefore, high activities or many digestive peptidases are expected in Uloboridae because total protein ingestion is the sum of protein content in prey and protein present in silk consumption from the wrapping envelope33.
Biochemical assays of soluble fraction of the digestive tract of Uloborus sp. revealed endopeptidase and exopeptidase activity (Supplementary Table S1 and Fig. 2A,B). Regarding exopeptidase activities, Uloborus sp. presented similar enzyme activities and peptides as other spider species, corroborating the data that carboxypeptidases are the predominant exopeptidase enzymes involved in protein degradation. The identified carboxypeptidases belonged to the serine carboxypeptidase (EC 3.4.16.6) and metallocarboxypeptidases (EC 3.4.17.17; EC 3.4.17.24) families. Similar results were obtained for previously characterised spiders, indicating that exopeptidases are important components secreted during the first phase of protein digestion.
The endopeptidases, astacin and cysteine peptidases, were identified by enzyme activity and peptide identification by mass spectrometry, as expected based on the literature data of spiders’ digestive fluid. The involvement of astacin in Uloboridae digestion was expected because of its specificity and the composition of the web used for prey immobilisation34,35. Astacin preferentially hydrolyses peptide bonds formed by Ala, Gly, and Ser residues at P1 and P1’36.
In contrast, the Uloboridae catching web comprises different types of spidroins, including aciniform spidroins (AcSp). The AcSp rich in alanine (Ala), glycine (Gly), and serine (Ser) forms aciniform spidroins type 1 (AcSp1)37,38, which is suitable for astacin hydrolysis. The importance of the spider digestive fluid has been demonstrated in the degradation of the thinner type-A aciniform spidroin lines during Uloboridae feeding.
However, contrary to the expected, the specific activity of Uloborus sp. astacin was lower than that of other spiders astacins. Proteomes from N. cruentata digestive tract and also from the digestive fluids from N. cruentata, S. mimosarum, and A. geniculata show a higher proportion of astacins than other endopeptidases, mainly, by an abundant presence of astacins isoforms18,19, respectively 27, 34 and 11 astacins were identified in the digestive fluids of spiders. However, Uloborus sp. midgut proteomic data revealed only two sequences of metallopeptidases. The lower astacin content identified in Uloborus sp. might be related to the distinct mass spectrometry methods applied to sample analysis. However, proteomic data from other spider samples using the same methodology allowed the identification of more astacin peptides than those identified in Uloborus sp. Another limitation in the identification of astacin peptides may be related to the absence of Uloborus sp. transcriptome data. The proteomic analysis was done using a database composed mainly of Araneus ventricosus and S. mimosarum sequences which are the best databases available of spiders and from the transcriptome of the Uloboridae spider Octonoba yesoensis39. The transcriptome of O. yesoensis has 50 distinct transcripts of astacins, which is a compatible number of astacins in spiders18,19, possibly indicating that the low activity and peptides of Uloborus sp. astacin could still be related to other factors such as astacin specificity/assay substrate and feeding conditions.
Walter et al.19 suggested that the difference between astacin numbers could be related to web degradation and the evolutionary scale of different spiders. To understand this, other specific transcriptome and proteome data are required.
Although astacins and cysteine peptidases are the main peptidases in spiders, another family of endopeptidases, namely serine peptidase, might be involved in protein degradation. Many peptides from the S1 (serine peptidase) family have been identified by proteomic analysis in all spiders in which digestive enzymes were studied18,19,40. However, identification of serine peptidase activity was not possible even with different substrates and assay conditions. The only measured serine peptidase activity was observed in Trichonephila clavipes17, indicating enzyme specificity similar to that of thrombin hydrolysis. Serine peptidases are also present in the EOD enzyme cocktail, although the expressed trypsins in the midgut of spiders seem to be different from classical trypsins, such as bovine and human trypsins, owing to the presence of accessory domains40. Our research group identified accessory domains analysing the data obtained from the midgut proteome and transcriptome of N. cruentata18, digestive fluid of A. geniculata, and S. mimosarum19, and the whole body transcriptome of O. yesoensis39.
Acidic protein degradation catalysed by cysteine peptidase is also important, as previously reported18,19,41 and is involved in both, the extracellular digestion during prey liquefaction, and intracellular digestion. Our results also demonstrate the participation of cathepsin L-like enzymes in the digestive process of Uloborus sp. (biochemically and by proteomic analysis) and from the transcriptome from O. yesoensis. Thus, the presence of acidic peptidases is mandatory in spider digestion, even in the digestive fluid and midgut digestive cells.
As described by18,41, enzymes that are mainly involved in intracellular digestion, such as cysteine peptidases, alpha-L-fucosidase, and alpha-mannosidase, perform a more acid hydrolysis, and secreted enzymes, such as chitinases, astacins, alpha-amylase, aminopeptidases, and carboxypeptidases conduct a less acidic EOD. The pattern of enzymes pH optimum, which correlates with enzymes location/phase of digestion, is the same in Uloborus sp. as that described for other spiders18,19,41.
Carbohydrates digestion
The main enzymes involved in carbohydrate digestion in spiders are chitinases and hexosaminidases (β-N-acetylglucosaminidase), which act synergistically to catalyse the cleavage of chitin polymers, and amylase. Chitinase and β-N-acetylglucosaminidase were firstly isolated from Cuppienius salei11 and their activities are related to insect carcass degradation, while promoting the accessibility of the venom and the digestive fluid to prey tissues, as well as a defensive role against microorganisms.
Amylase catalyses the cleavage of the α-1,4-glycosidic bonds in starch and glycogen. In insects, glycogen is abundantly stored in the fat body42. However, starch is also an available substrate for spider amylase. Starch can be present in the pollen grain, adhered to the spider’s web, and ingested during web recycling5; it also could be attached with a prey that is a pollinator, or even in the midgut content at digestive tract of herbivorous insects ingested as prey43,44. Furthermore, actively feeding on pollen is important for orb-weaver spider nourishment, particularly for juvenile orb-weaver’s spiders45. Uloborus sp. presented both alpha amylase activity, identified biochemically, and alpha amylase peptides in the proteomic analysis. Measurements of enzymatic activity using glycogen and starch as substrates indicated that Uloborus amylase evenly hydrolysed both substrates.
Proteomic and biochemical data also identified enzymes involved in the removal of carbohydrates from glycoconjugates, including alpha-L-fucosidase and alpha-mannosidase which are related to the intracellular digestive process in Uloborus sp. The first characterisation of a spider alpha-L-fucosidase was obtained from the midgut of N. cruentata46. In insects, the N-glycosylation pathways involve a high mannosylation and two fucosylations (Man3GlcNAc(± α3/6Fuc) GlcNAc) of glycoproteins, forming a structure known as paucimannosidic glycans47. Thus, alpha-mannosidase and alpha-L-fucosidase of spiders participate in the removal of these glycan residues during prey digestion. Furthermore, diverse glycans repertoires participate in host–parasite relationship48, in glycoproteins and glycolipids so-called ‘host-like’ glycans49. In this manner, alpha-mannosidase and alpha-L-fucosidase could also play defensive roles50.
Lipid digestion
Different types of triacylglycerol, diacylglycerol, and monoacylglycerol lipases, and phospholipases are involved in lipid and phospholipids hydrolysis.
Lipid storage (lipid droplets) in the midgut of spiders is essential for nourishment, during extensive periods of starvation51, and for delivering lipid pathways to other tissues52. Lipids are one of the major macronutrients in insect53, thus, lipases are crucial for lipid digestion in spiders.
The midgut proteome of Uloborus sp. identified only phospholipases, such as phospholipases C and D. Lipase activity was measured using DMPTB. This activity may be related to hydrolysis by non-specific lipases54. Therefore, triacylglycerol, diacylglycerol, and monoacylglycerol lipases could hydrolyse DMPTB, endorsing the presence of lipases in Uloborus sp. midgut; as well as phospholipases C release diacylglycerol during phospholipids hydrolysis, making it a suitable product for lipases.
The midgut of Uloborus sp., used as a source of digestive enzymes in the present study, has limited information on the feeding conditions of the individuals studied, but they were at least 72 h fasting. As observed in Pardosa prativaga, a higher metabolism rate was observed in the first 3 h of digestion55, and in N. cruentata, there are more lipases in animals under fasting conditions18. Thus, feeding conditions could be the reason that we did not identify lipases in Uloborus sp. midgut proteome. In addition, as the midgut diverticula are one of the tissues with higher volumes in a spider, the whole-body transcriptome of O. yesoensis, an Uloboridae spider, revealed 21 lipase transcripts, some of which might belong to the midgut diverticula.
SMaseD phylogenetics
Some peptides identified in the proteomic analysis of Uloborus midgut are orthologues of peptides identified in the venom gland of other non-Uloboridae spiders, corroborating previous data reported in the literature that some peptides with toxin function are expressed in tissues distinct from the venom gland. Previously, the identification of some proteins homologous to venom toxins has been attributed to digestive fluid contamination with venom. Foradori et al.15 were the first to isolate several peptidase inhibitors similar to those identified in spider venom in the digestive fluid of A. aurantia. The transcriptome of N. cruentata (GenBank: GEWZ00000000.1)18 allowed the identification of toxin and enzyme orthologues transcripts of spider midgut to those found in venoms.
These toxin-like peptides have already been identified in other non-venomous Arachnida, such as mites and ticks26. Similar to peptide toxins, enzymes involved in the envenomation process, which are exclusive to venom, are also shared between the venom gland and other organs and have been described in mites and ticks. SMaseD is an example of such enzyme.
Although SMaseD in Arachnida is a group of orthologous genes, it can have distinct disulphide binding patterns and activities. SMaseD is expressed in the salivary glands of ticks and modulates the host immune system, facilitating pathogen transmission26. Some tick genera, such as Rhipicephalus and Amblyomma56, cause inflammation and necrosis in the host. However, recombinant SMaseD from Ixodes scapularis did not show necrotic effects. Differences in SMaseD catalytic efficiency and specificity have also been observed between Sicaridae species57.
Our data on proteomic analysis of the digestive tract of Uloborus sp. and the analysis of the transcriptome of O. yesoensis suggested the presence of SMaseD in Uloboridae spiders. The identification of toxins including SMaseD, in the midgut of Uloborus sp. is a complete proof of the involvement of this enzyme in digestion with potential insecticidal activity, facilitating prey paralysis and capture even in a non-venomous spider.
Analysis of the midgut transcriptome and proteome from N. cruentata, of the opisthosoma (abdomen) of A. geniculata19, and the midgut proteome of Loxosceles gaucho58 also identified SMaseD involved in the digestive tract. The analysis of S. mimosarum digestive fluid proteome identified peptides corresponding to SMaseD, but in low concentration. In contrast, analysis of the proteome of the digestive fluid from N. cruentata and A. geniculata did not detect SMaseD peptides. These data suggest the involvement of SMaseD mainly in the intracellular phase of digestion. This is in accordance with the SMase identified in tick genomes, which are predicted to be acidic and neutral SMases, localised in the lysosome.
SMase multiple alignment allowed the maximum likelihood algorithm to expand the history of the SMaseD gene in some Chelicerata species (Supplementary Fig. S4). Orthologues of SMaseD genes were identified even in Limulus transcriptome data (Supplementary Table S5), a basal group of the Chelicerata subphylum, suggesting that the association of SMaseD with digestion is older than its relation to envenomation in spiders. Another aspect observed in this analysis is an unusual distribution of ticks and mites SMaseD, although both groups possess class IIb SMaseD, mite SMaseD has sequences represented in the three groups distinguished in the tree; however, most sequences are more similar to SMasesD from spiders midgut than from ticks SMaseD. Usually, regarding other digestive enzymes, mites and ticks present sequences with high identity values and are frequently grouped together18,41 indicating a distinct selective pressure to the evolution of SMaseD.
Although this tree is informative, it has low bootstrap values support. In order to enhance the statistical bootstrap support, and evidence the grouping of the distinct classes of SMase in spiders, an exclusive spider maximum likelihood tree was performed.
The exclusive spider analysis and sequence alignment indicated that the midgut SMaseD sequences belong to the class II SMaseD subfamily, conserving the two disulphide bridges (Fig. 4) between Cys97 and Cys103 (corresponding to Cys51 and Cys57, according to PDB entry)27,59 and an additional disulphide bridge between Cys99 and Cys250 (corresponding to Cys53 and Cys201, according to PDB entry)27,59. Moreover, alignment analysis suggested that digestive SMaseD belongs to class IIb, due to substitution at Gly141 (corresponding to Gly95, according to PDB entry)27 by an asparagine, lysine, or histidine residue (Fig. 5); and conservation of proline residue Pro180 (corresponding to Pro134, according to PDB entry)27. Furthermore, midgut SMaseD conserves the histidine, glutamic acid, and aspartic acid residues: His57, His93, Glu77, and Asp79 (Fig. 5), according to LiRecDT160, involved in the catalytic process and coordination of magnesium ion at the active site28, as well as, conserves lysine, tyrosine, and tryptophan residues: Lys139, Tyr278, and Trp280 (Fig. 5), according to LiRecDT1, involved in substrate recognition and stabilization in the cleft of the active site28.
Previous analyses27,59, comparing the relationship and adaptive evolution of SMaseD to glycerophosphodiester phosphodiesterase were so far made, exhibiting a positive selection on spider venom SMaseD, and an increase of necrotic activity.
Although SMaseD has been widely described for their toxicity in mammals, the main prey of these spiders, even of the family Sicaridae, are other arthropods. Loxosceles laeta, Loxosceles intermedia and Loxosceles arizonica for example have, as their main prey, other spiders, ants and scorpions. On the other hand, little is known about lipidic constituents in arthropods. In Diptera the main complex lipids are ceramide phosphoethanolamines. However, in other groups, such as hemipterans the main lipids are sphingomyelin61. Diets composed of prey rich in sphingomyelin constitute an important selective pressure on more effective SMaseD, culminating in the selection of insecticide molecules.
Another aspect that should be emphasised is the common enzyme between the salivary glands of ticks and the venom gland of spiders. One of the hypotheses of their appearance in the venom gland in spiders is that they originate from the salivary glands2,62, concomitantly with the gene duplication events in the Arachnida and Chelicerata groups63,64,65,66. Our results suggest that the spider venom gland origin is closely related to the salivary gland in ticks and is derived from a gland associated with digestion because of the presence of class IIb SMaseD and other shared components between these tissues.
Conclusion
Here, we summarise for the first time the profile of the midgut enzymes of the non-venomous spider Uloborus sp., using proteomic and enzymological approaches. The results indicated that Uloborus sp. shares the main digestive enzymes, such as metallopeptidases and cysteine peptidases, carboxypeptidases, chitinases and amylases, with the families of venomous spiders. In addition to typical digestive enzymes, the midgut of Uloboridae also contains enzymes such as SMaseD and other toxin peptides usually found in venom glands, suggesting an insecticide activity of digestive fluid in Uloboridae and indicating that the origin of spider toxins during evolution is possibly the digestive system.
Material and methods
Animals and sample preparation
Adult females of Uloborus sp. were collected in Fazenda Nova Monte Carmelo, located between municipalities of Araguari, Romaria, Nova Ponte, Estrela do Sul and Monte Carmelo, in Minas Gerais, Brazil (18°49′27″S, 47°51′47″W). The area is an Eucalyptus crop interspersed with native Cerrado vegetation (i.e. Brazilian.
Savanna). As all of the Brazilian Uloborus species have been described based only on color patterns, species identification still requires a taxonomic revision of South American species. The species object of this study in Fig. 7 is the same previously studied by Nascimento67 and voucher specimens were deposited in the collection of Universidade Federal de Minas Gerais (Reference number: 16685).
Sampled individuals were immediately sent to Instituto Butantan (São Paulo, SP) and were kept in 15 mL tubes, at room temperature, for at least 72 h of fasting. The spiders were then anesthetized in CO2 for about 10–15 min and subsequently dissected, using a stereomicroscope and isotonic saline solution (NaCl 300 mM)5 for the isolation of the midgut, silk gland, egg sac, and carcass. The different tissues were collected and stored at −20ºC.
Midguts were individually homogenized with a Potter–Elvehjem to fragment the internal membrane of enterocytes and release content, using cold Ultrapure Water (Milli-Q). Thereafter, the homogenate was centrifuged at 16,000 × g for 30 min at 4ºC. The soluble fraction was used as an enzyme source, and it was stored at −20ºC.
Protein determination and enzymatic assays
The protein content from Uloborus sp midgut samples was estimated by the bicinchoninic acid method68 using albumin from the chicken egg as standard protein. Enzymatic assays were performed at 30 °C using a series of chromogenic and fluorescent substrates. Product reaction was measured using a spectrophotometer (SpectraMax M2—Molecular Devices) and spectrofluorometer (SpectraMax Gemini XPS—Molecular Devices), through at least four different intervals of time to ensure enzyme initial velocities. Blanks of the enzyme (enzyme plus buffer at the same assay condition) and the substrate (substrate plus enzyme solvent) were used as negative controls. Enzymes, buffers, substrates, and their final concentrations at the well plate are listed below (Table 1). At least, 5 different biological samples were used to each enzyme assay.
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE)
SDS-PAGE was performed with one individualized midgut homogenized in 500 μl of ultra-filtered cold water. The homogenate sample was submitted to VivaSpin 20 centrifugal concentrator MWCO 10 kDa using ultrafiltered water as diluent was concentrated 500 μl of homogenate sample, which corresponds to, was resuspended in 20 μl of sample buffer (approximately, a total of 10 μg). Samples from Uloborus sp. midgut (approximately 10 μg of protein) were submitted to VivaSpin 20 centrifugal concentrator MWCO 10 kDa using ultrafiltered water as diluent. After that, samples were concentrated in vacuum centrifugation (speed vac), and resuspended (total of 10 μg of protein) in 20 μl of sample buffer, containing: 60 mM Tris–HCl buffer (pH 6.8), 2.5% SDS, 0.36 mM b-mercaptoethanol, 10% (v/v) glycerol, and 0.005% (w/v) bromophenol blue, thereafter, the sample was heated at 100 ºC for 5 min. Samples were applied onto a 12% polyacrylamide gel according to75. The electrophoresis was performed at 150 V, and the gel was silver stained76.
Bottom-up proteomics
An aliquot of Uloborus sp. midgut was concentrated in vacuum centrifugation (speed vac), to a total of 10 μg of protein. Afterward, the procedure of digestion in solution was: 20 μl of 8 M Urea, 10 min at 22 °C, 0.5 μl of 1 M dithiothreitol (DTT) for 30 min and 30 °C, 5 μl of 200 mM iodoacetamide (IAA) in the dark for 30 min at 22 °C, 90 μl of digestion buffer [50 mM NH4HCO3 + 10% acetonitrile (ACN)] to dilute urea concentration and addition of trypsin to protein ratio of 1:100. The digestion was incubated for 18 h at 30 °C and stopped by the addition of 0.1% formic acid (FA). After that, ACN was removed by vacuum centrifugation.
The sample concentrated by vacuum centrifugation was resuspended in 50 μl of acidic solution (0.1% FA) to mass spectrometry procedure. Tandem mass spectrometry analysis of tryptic peptides was performed by an LC–MS/MS Q-TOF (Shimadzu), samples (50 µL aliquot) were loaded into a C18 column (Kinetex C18, 5 μm; 50 × 2.1 mm) and peptides eluted by binary gradient of 5% to 40%, solvent A—Water:FA (999: 1) and solvent B—ACN:Water:FA (900: 99: 1), at a constant flow of 0.2 mL.min-1 for 40 min.
Raw LCD, LCMSolution—Shimadzu, data were converted into MGF by the LCMSolution tool and then loaded into Peaks Studio V7.0 (BSI, Canada). Data were processed according to the following parameters: MS and MS/MS error mass were: 0.1 Da; methionine oxidation and carbamidomethylation as variable and fixed modification, respectively; trypsin as cleaving enzyme; maximum missed cleavages (3), maximum variable PTMs per peptide (3) and non-specific cleavage (both); only proteins with score ≥ 40 and containing at least 1 unique peptide were considered in this study.
Bioinformatic analysis/databases
Different databases from spiders employed these searches, such as N. cruentata midgut proteome18, S. mimosarum, and A. geniculata genome77, and their digestive fluid proteome19. Furthermore, we employed Trans Decoder78 (Galaxy Australia), finding coding regions software, to the raw database from O. yesoensis whole body transcriptome (GenBank: IAQA00000000.1), afterwards, the translation and annotation of the enzyme's nucleotide sequences were obtained by blastp, using A. ventricosus and S. mimosarum amino acid enzyme sequences as queries.
Protein sequences of SMaseD Class I, IIa, and IIb were obtained at: Uniprot reviewed database (venom gland Loxosceles genera);18 SMaseD from the midgut of N. cruentata;39, from the whole body of O. yesoensis;77 transcriptomic data from S. mimosarum and A. geniculata corroborated by proteomic data from the spider´s digestive fluid and/or abdomen19, and submitted to multiple sequence alignment analysis and classification of midgut SMaseD sequences. SMaseD sequences from O. yesoensis were acquired by Trans Decoder78 (Galaxy Australia) annotation of the transcriptome (GenBank: IAQA00000000.1), filtered SMaseD nucleotide sequences using N. cruentata as Blastn query (NCBI), and translated to amino acid sequence by Translate Tool (Expasy)79. Protein alignment of SMaseD sequences was built by MUSCLE software80 and analyzed in Jalview (2.11.2.2 version) software81, in order to compare the cysteine residues conservation and pattern and glycine/proline residue substitutions at specific positions to enzyme classification.
For the construction of an SMaseD phylogenetic tree of Arachnida and Chelicerata species we collected nucleotide SMaseD sequences from midgut Tityus serrulatus, Ixodes ricinus, Rhipicephalus microplus, and Amblyomma aureolatum, from the salivary gland of Ixodes scapularis, from whole body Dermacentor silvarum, Rhipicephalus sanguineus, Tetranychus urticae, Metaseiulus occidentalis and Limulus polyphemus, venom gland Hemiscorpius lepturus; sequences were collected by keywords (Sphingomyelinase D, Phospholipase D and Dermonecrotic toxin) search in Nucleotide Advanced Search Builder (NCBI), filtered to Arachnida group and only mRNA and EST sequences were selected. Furthermore, midgut sequences were also added to the construction at the phylogenetic tree. The sequences and access codes are listed in Supplementary Table S5.
The Nucleotide sequence alignment was performed by MUSCLE software80. The phylogenetic tree was constructed using by IQ Tree (Galaxy Version 2.1.2 + galaxy2) software82 by a “Maximum Likelihood Tree”, with a bootstrap method as a test of phylogeny with 1000 bootstrap replications. The trees were coloured using iTol (Interactive tree of life) v6 software83.
Data availability
The datasets generated and/or analyzed during the current study are available in: 1- Proteomic data: the PRIDE repository, Data are available via ProteomeXchange with identifier PXD037345 Project https://doi.org/10.6019/PXD037345. Reviewer account details Username: reviewer_pxd037345@ebi.ac.uk Password: OafQcthc. (https://www.ebi.ac.uk/pride/login). 2- The translated sequences from Octonoba yesoensis 39—https://repositorio.butantan.gov.br/handle/butantan/4540.
References
Pineda, S. S. et al. Structural venomics reveals evolution of a complex venom by duplication and diversification of an ancient peptide-encoding gene. Proc. Natl. Acad. Sci. 117, 11399–11408 (2020).
Lüddecke, T., Herzig, V., Reumont, B. M. & Vilcinskas, A. The biology and evolution of spider venoms. Biol. Rev. 97, 163–178 (2022).
Correa-Garhwal, S. M., Baker, R. H., Clarke, T. H., Ayoub, N. A. & Hayashi, C. Y. The evolutionary history of cribellate orb-weaver capture thread spidroins. BMC Ecol Evol 22, 89 (2022).
Cohen, A. C. Extra-Oral Digestion in Predaceous Terrestrial Arthropoda. Annu Rev Entomol 40, 85–103 (1995).
Foelix, R. Biology of spiders. (OUP USA, 2011).
Perret, B. A. Proteolytic activity of tarantula venoms due to contamination with saliva. Toxicon 15, 505–510 (1977).
Mommsen, T. P. Comparison of digestive?-amylases from two species of spiders (Tegenaria atrica andCupiennius salei). J. Compar. Physiol. B 127, 355–361 (1978).
Mommsen, T. P. Digestive enzymes of a spider (Tegenaria atrica koch)—III. Esterases, phosphatases, nucleases. Comp. Biochem. Physiol. A Physiol. 60, 377–382 (1978).
Mommsen, T. P. Digestive enzymes of a spider (Tegenaria atrica koch)—II. Carbohydrases. Comp. Biochem. Physiol. A Physiol. 60, 371–375 (1978).
Mommsen, T. P. Digestive enzymes of a spider (Tegenaria at rica Koch)—I. General remarks, digestion of proteins. Comp. Biochem. Physiol. A Physiol. 60, 365–370 (1978).
Mommsen, T. P. Chitinase and from the digestive fluid of the spider. Biochimica et Biophysica Acta (BBA) - Enzymology 612, 361–372 (1980).
Tillinghast, E. K. & Kavanagh, E. J. The alkaline proteases of Argiope and their possible role in web digestion. J. Exp. Zool. 202, 213–222 (1977).
Kavanagh, E. J. & Tillinghast, E. K. The alkaline proteases of Argiope—II. Fractionation of protease activity and isolation of a silk fibroin digesting protease. Compar. Biochem. Physiol. Part B: Compar. Biochem. 74, 365–372 (1983).
Tugmon, C. R. & Tillinghast, E. K. Proteases and protease inhibitors of the spiderArgiope aurantia (Araneae, Araneidae). Naturwissenschaften 82, 195–197 (1995).
Foradori, M. J., Tillinghast, E. K., Smith, J. S., Townley, M. A. & Mooney, R. E. Astacin family metallopeptidases and serine peptidase inhibitors in spider digestive fluid. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 143, 257–268 (2006).
Ludwig, M. & Alberti, G. Mineral congregations, ?spherites? in the midgut gland ofCoelotes terrestris (Araneae): Structure, composition and function. Protoplasma 143, 43–50 (1988).
Joo, H.-S. et al. Purification and characterization of a Prothrombin-activating protease from Nephila clavata. Toxicon 40, 289–296 (2002).
Fuzita, F. J., Pinkse, M. W. H., Patane, J. S. L., Verhaert, P. D. E. M. & Lopes, A. R. High throughput techniques to reveal the molecular physiology and evolution of digestion in spiders. BMC Genomics 17, 716 (2016).
Walter, A. et al. Characterisation of protein families in spider digestive fluids and their role in extra-oral digestion. BMC Genomics 18, 600 (2017).
Kuhn-Nentwig, L., Schaller, J. & Nentwig, W. Purification of toxic peptides and the amino acid sequence of CSTX-1 from the multicomponent venom of Cupiennius salei (Araneae:Ctenidae). Toxicon 32, 287–302 (1994).
Langenegger, N., Nentwig, W. & Kuhn-Nentwig, L. Spider Venom: Components, Modes of Action, and Novel Strategies in Transcriptomic and Proteomic Analyses. Toxins (Basel) 11, 611 (2019).
Opell, B. D. American Arachnological Society Prey Handling and Food Extraction by the Triangle-Web Spider Hyptiotes cavatus (Uloboridae). Source: The Journal of Arachnology vol. 16. http://www.jstor.org/stable/3705766 (1988).
Eberhard, W. G., Barrantes, G. & Weng, J.-L. The mystery of how spiders extract food without masticating prey. Bull. Br. arachnol. Soc. (2006).
Esquivel, C., Escalante, I. & Eberhard, W. G. The effect of regurgitated digestive fluid on the spider’s own legs in Philoponella vicina (Araneae: Uloboridae). Sour.: J. Arachnol. 40 (2012).
Eberhard, W. G., Barrantes, G. & Weng, J.-L. Tie them up tight: wrapping by Philoponella vicina spiders breaks, compresses and sometimes kills their prey. Naturwissenschaften 93, 251–254 (2006).
Rajendran, K. v., Neelakanta, G. & Sultana, H. Sphingomyelinases in a journey to combat arthropod‐borne pathogen transmission. FEBS Lett 595, 1622–1638 (2021).
Murakami, M. T. et al. Structural insights into the catalytic mechanism of sphingomyelinases D and evolutionary relationship to glycerophosphodiester phosphodiesterases. Biochem. Biophys. Res. Commun. 342, 323–329 (2006).
Vuitika, L. et al. Active site mapping of Loxosceles phospholipases D: Biochemical and biological features. Biochimica et Biophysica Acta (BBA) – Mol. Cell Biol. Lipids 1861, 970–979 (2016).
Lajoie, D. M. et al. Variable substrate preference among phospholipase D toxins from sicariid spiders. J. Biol. Chem. 290, 10994–11007 (2015).
Dantas, A. E. et al. Description of Loxtox protein family and identification of a new group of Phospholipases D from Loxosceles similis venom gland. Toxicon 120, 97–106 (2016).
Opell, B. D. & Eberhard, W. G. Resting postures of orb-weaving uloborid spiders (Araneae, Uloboridae). J Arachnol 11, 369–376 (1983).
Paul, A. et al. Grasshoppers as a food source? A review. BASE 337–352 (2016). https://doi.org/10.25518/1780-4507.12974.
Weng, J.-L., Barrantes, G. & Eberhard, W. G. Feeding by Philoponella vicina (Araneae, Uloboridae) and how uloborid spiders lost their venom glands. Can. J. Zool. 84, 1752–1762 (2006).
Tremblay, M.-L. et al. Spider wrapping silk fibre architecture arising from its modular soluble protein precursor. Sci. Rep. 5, 11502 (2015).
Babb, P. L. et al. The Nephila clavipes genome highlights the diversity of spider silk genes and their complex expression. Nat. Genet. 49, 895–903 (2017).
Rawlings, N. D. et al. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 46, D624–D632 (2018).
Chaw, R. et al. Intragenic homogenization and multiple copies of prey-wrapping silk genes in Argiope garden spiders. BMC Evol. Biol. 14, 31 (2014).
Wen, R. et al. Molecular cloning and analysis of the full-length aciniform spidroin gene from Araneus ventricosus. Int. J. Biol. Macromol. 117, 1352–1360 (2018).
Kono, N., Nakamura, H., Mori, M., Tomita, M. & Arakawa, K. Spidroin profiling of cribellate spiders provides insight into the evolution of spider prey capture strategies. Sci. Rep. 10, 15721 (2020).
Passos, R. A. S. Avaliação preditiva estrutural das serino peptidases de aranhas: as relações evolutivas e de estrutura/função. (Insituto Butantan, 2022).
Fuzita, F. J., Pinkse, M. W. H., Verhaert, P. D. E. M. & Lopes, A. R. Cysteine cathepsins as digestive enzymes in the spider Nephilengys cruentata. Insect Biochem. Mol. Biol. 60, 47–58 (2015).
Arrese, E. L. & Soulages, J. L. Insect fat body: Energy, Metabolism, and regulation. Annu. Rev. Entomol. 55, 207–225 (2010).
Wyatt, G. R. The Biochemistry of Sugars and Polysaccharides in Insects. In Advances in insect physiology 287–360 (1967). https://doi.org/10.1016/S0065-2806(08)60210-6.
Clarke, D., Morley, E. & Robert, D. The bee, the flower, and the electric field: electric ecology and aerial electroreception. J. Comp. Physiol. A. 203, 737–748 (2017).
Eggs, B. & Sanders, D. Herbivory in spiders: the importance of pollen for orb-weavers. PLoS ONE 8, e82637 (2013).
Perrella, N. N., Fuzita, F. J., Moreti, R., Verhaert, P. D. E. M. & Lopes, A. R. First characterization of fucosidases in spiders. Arch. Insect Biochem. Physiol. 98, e21462 (2018).
Walski, T., de Schutter, K., Cappelle, K., van Damme, E. J. M. & Smagghe, G. Distribution of glycan motifs at the surface of midgut cells in the cotton leafworm (Spodoptera littoralis) Demonstrated by Lectin Binding. Front. Physiol. 8, (2017).
Veríssimo, C. D. M., Graeff-Teixeira, C., Jones, M. K. & Morassutti, A. L. Glycans in the roles of parasitological diagnosis and host–parasite interplay. Parasitology 146, 1217–1232 (2019).
van Die, I. & Cummings, R. D. Glycan gimmickry by parasitic helminths: A strategy for modulating the host immune response?. Glycobiology 20, 2–12 (2010).
Moreti, R., Perrella, N. N. & Lopes, A. R. Carbohydrate digestion in ticks and a digestive α-l-fucosidase. J. Insect Physiol. 59, 1069–1075 (2013).
Lipovšek, S., Novak, T., Janžekovič, F., Brdelak, N. & Leitinger, G. Changes in the midgut diverticula epithelial cells of the European cave spider, Meta menardi, under controlled winter starvation. Sci. Rep. 8, 13645 (2018).
Laino, A., Cunningham, M. L., García, F. & Heras, H. First insight into the lipid uptake, storage and mobilization in arachnids: Role of midgut diverticula and lipoproteins. J. Insect Physiol. 55, 1118–1124 (2009).
Jensen, K., Mayntz, D., Toft, S., Raubenheimer, D. & Simpson, S. J. Nutrient regulation in a predator, the wolf spider Pardosa prativaga. Anim. Behav. 81, 993–999 (2011).
Choi, S.-J., Hwang, J.-M. & Kim, S.-I. A colorimetric microplate assay method for high throughput analysis of lipase activity. BMB Rep. 36, 417–420 (2003).
Jensen, K., Mayntz, D., Wang, T., Simpson, S. J. & Overgaard, J. Metabolic consequences of feeding and fasting on nutritionally different diets in the wolf spider Pardosa prativaga. J. Insect Physiol. 56, 1095–1100 (2010).
Latif, A. A., Maina, J. N., Dhadialla, T. S. & Nokoe, S. Histological Reactions to Bites of Amblyomma variagatum and Rhipicephalus appendiculatus (Acari: Ixodidae) Fed Simultaneously on Naive or Sensitized Rabbits. J. Med. Entomol. 27, 316–323 (1990).
Zobel-Thropp, P. A., Kerins, A. E. & Binford, G. J. Sphingomyelinase D in sicariid spider venom is a potent insecticidal toxin. Toxicon 60, 265–271 (2012).
Valladão, R. Comparative molecular physiology of spider’s digestion: Transcriptomic, proteomic and enzimologic analysis physiological and molecular aspects. (University of São Paulo , 2022).
Pedroso, A., Matioli, S. R., Murakami, M. T., Pidde-Queiroz, G. & Tambourgi, D. v. Adaptive evolution in the toxicity of a spider’s venom enzymes. BMC Evol. Biol. 15, 290 (2015).
Chaim, O. M. et al. Phospholipase-D activity and inflammatory response induced by brown spider dermonecrotic toxin: Endothelial cell membrane phospholipids as targets for toxicity. Biochimica et Biophysica Acta (BBA) – Mol. Cell Biol. Lipids 1811, 84–96 (2011).
Shi, X.-X. et al. Two sphingomyelin synthase homologues regulate body weight and sphingomyelin synthesis in female brown planthopper, N. lugens (Stål). Insect Mol. Biol. 28, 253–263 (2019).
Coons, L. B. & Roshdy, M. A. Fine structure of the salivary glands of unfed male Dermacentor variabilis (Say) (Ixodoidea: Ixodidae). J. Parasitol. 59, 900–912 (1973).
Sharma, P. P., Santiago, M. A., González-Santillán, E., Monod, L. & Wheeler, W. C. Evidence of duplicated Hox genes in the most recent common ancestor of extant scorpions. Evol. Dev. 17, 347–355 (2015).
Kenny, N. J. et al. Ancestral whole-genome duplication in the marine chelicerate horseshoe crabs. Heredity (Edinb) 116, 190–199 (2016).
Schwager, E. E. et al. The house spider genome reveals an ancient whole-genome duplication during arachnid evolution. BMC Biol. 15, 62 (2017).
Turetzek, N., Pechmann, M., Schomburg, C., Schneider, J. & Prpic, N.-M. Neofunctionalization of a Duplicate dachshund gene underlies the evolution of a novel leg segment in Arachnids. Mol. Biol. Evol. 33, 109–121 (2016).
Nascimento, A. L. & Gonzaga, M. O. Maternal defensive behaviors of Uloborus sp. (Araneae, Uloboridae): behavioral repertoire and influence of clutch size and female size on female aggressiveness. Acta Ethol. 19, 33–41 (2016).
Smith, P. K. et al. Measurement of protein using bicinchoninic acid. Anal. Biochem. 150, 76–85 (1985).
Baker, J. E. & Woo, S. M. ß-glucosidases in the rice weevil, Sitophilus oryzae:Purification, properties, and activity levels in wheat- and legume-feeding strains. Insect Biochem. Mol. Biol. 22, 495–504 (1992).
Alves, L. C., Almeida, P. C., Franzoni, L., Juliano, L. & Juliano, M. A. Synthesis of N alpha-protected aminoacyl 7-amino-4-methyl-coumarin amide by phosphorous oxychloride and preparation of specific fluorogenic substrates for papain. Pept. Res. 9, 92–6.
Noelting, G. & Bernfeld, P. Sur les enzymes amylolytiqucs III. La β-amylase: dosage d’activité et contrôle de l’absence d’α-amylase. Helv. Chim. Acta 31, 286–290 (1948).
Twining, S. S. Fluorescein isothiocyanate-labeled casein assay for proteolytic enzymes. Anal. Biochem. 143, 30–34 (1984).
Nicholson, J. A. & Kim, Y. S. A one-step l-amino acid oxidase assay for intestinal peptide hydrolase activity. Anal. Biochem. 63, 110–117 (1975).
Erlanger, B. F., Kokowsky, N. & Cohen, W. The preparation and properties of two new chromogenic substrates of trypsin. Arch. Biochem. Biophys. 95, 271–278 (1961).
LAEMMLI, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Blum, H., Beier, H. & Gross, H. J. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 8, 93–99 (1987).
Sanggaard, K. W. et al. Spider genomes provide insight into composition and evolution of venom and silk. Nat. Commun. 5, 3765 (2014).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Gasteiger, E. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31, 3784–3788 (2003).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Acknowledgements
The Authors sincerely acknowledge Shimadzu for sample analysis by QTOF equipment. We would like to thank Editage (www.editage.com) for English language editing.This work was supported by the Brazilian research agencies FAPESP 2015/23745-7 and Fapemig. Rodrigo is a graduate fellow from CNPq (133680/2020-7). A. R. Lopes (310868/2018-1) and D.C Pimenta (301975/2019-5) are CNPq research fellow.
Author information
Authors and Affiliations
Contributions
R.V.: experimental design and execution, data analysis and manuscript composition; O.B.S.N., experimental execution, data analysis and critical reading, M.O.G. Uloborus sp supply and critical reading; D.C.P., data analysis and critical reading; A.R.L.: experimental design, data analysis and manuscript composition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Valladão, R., Neto, O.B.S., de Oliveira Gonzaga, M. et al. Digestive enzymes and sphingomyelinase D in spiders without venom (Uloboridae). Sci Rep 13, 2661 (2023). https://doi.org/10.1038/s41598-023-29828-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-29828-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.