Abstract
Antimicrobial resistance, especially carbapenem resistance, poses a serious threat to global public health. Here, a carbapenem-resistant Comamonas aquatica isolate SCLZS63 was recovered from hospital sewage. Whole-genome sequencing showed that SCLZS63 has a 4,048,791-bp circular chromosome and three plasmids. The carbapenemase gene blaAFM-1 is located on the 143,067-bp untypable plasmid p1_SCLZS63, which is a novel type of plasmid with two multidrug-resistant (MDR) regions. Notably, a novel class A serine β-lactamase gene, blaCAE-1, coexists with blaAFM-1 in the mosaic MDR2 region. Cloning assay showed that CAE-1 confers resistance to ampicillin, piperacillin, cefazolin, cefuroxime, and ceftriaxone, and elevates the MIC of ampicillin-sulbactam two-fold in Escherichia coli DH5α, suggesting that CAE-1 functions as a broad-spectrum β-lactamase. Amino acid sequences analysis suggested that blaCAE-1 may originate from Comamonadaceae. The blaAFM-1 in p1_SCLZS63 is located in a conserved structure of ISCR29-ΔgroL-blaAFM-1-ble-ΔtrpF-ΔISCR27-msrB-msrA-yfcG-corA. Comprehensive analysis of the blaAFM-bearing sequences revealed important roles of ISCR29 and ΔISCR27 in the mobilization and truncation of the core module of blaAFM alleles, respectively. The diverse passenger contents of class 1 integrons flanking the blaAFM core module make the complexity of genetic contexts for blaAFM. In conclusion, this study reveals that Comamonas may act as an important reservoir for antibiotics-resistance genes and plasmids in the environment. Continuous monitoring for the environmental emergence of antimicrobial-resistant bacteria is needed to control the spread of antimicrobial resistance.
Similar content being viewed by others
Introduction
Comamonas spp. are a group of Gram-negative, nonfermenting and rod-shaped bacteria belonging to the Comamonadaceae family of the phylum Proteobacteria1. This organism frequently grows in a wide range of habitats, such as wastewater, wetlands, soil, and hospital environments2, and has been reported as one of the major members of microbial communities in wastewater bioaugmentation and bioremediation3. In spite of its uncommon pathogenesis, some Comamonas species have been increasingly reported to be closely associated with invasive infections in humans, such as bacteremia4,5, urinary tract infection6, intra-abdominal infection7, and meningitis8.
β-lactams are the most commonly used antibiotics in clinical settings due to their safety, efficacy, and broad-spectrum of activity9. The biggest challenge to the use of β-lactams is the production of β-lactamases, which is the most common and important resistance mechanism for β-lactamases in Gram-negative bacteria10. Based on the amino acid sequence identity, β-lactamases are divided into four molecular classes (A-D)11. Classes A, C, and D β-lactamases hydrolyze their substrates through the formation of an acyl enzyme with an active-site serine, while the hydrolytic reaction of class B β-lactamases requires one or two essential zinc ions11. At present, the widespread of extended-spectrum β-lactamase (ESBL)- and carbapenemase-producing organisms pose a serious challenge for clinical management and public health for their hydrolytic activity against expanded-spectrum of β-lactam antibiotics9,12,13. Hospital wastewater is a complex matrix containing a high abundance of bacteria combined with sublethal antibiotic concentrations from clinical settings, which serves as a hot spot for the evolution and dissemination of antimicrobial resistance (AMR), and also a reservoir of novel resistance genes14. Marathe et al.15 demonstrated that hospital sewage effluent creates a niche where pathogens acquire novel antibiotic resistance genes (ARGs), including carbapenemases genes. Hem et al.16 isolated a subset of carbapenem-resistant Comamonas strains from wastewater, of which most possess novel unknown resistance mechanisms for carbapenems. Therefore, wastewater is an ideal place to identify novel ARGs.
To date, carbapenem resistance genes blaNDM17, blaIMP-818, and blaGES-516 have been reported in Comamonas isolates. AFM-1 is a newly emergent carbapenemase that was first identified in a clinical Alcaligenes faecalis strain AN-70 in China and was later reported in Pseudomonas aeruginosa19 and Aeromonas hydrophila20, in which ISCR elements are involved in the mobilization of blaAFM-1. In silico analysis with the GenBank database indicated four variants of AFM (AFM-1 to -4) that were also present in Stenotrophomonas maltophilia, Bordetella trematum, as well as Comamonas isolates. In this study, we described a novel type of multidrug-resistance plasmid carrying the carbapenemase gene blaAFM-1 in a Comamonas aquatica strain from hospital sewage. Of note, we also characterized a novel class A serine β-lactamase gene, blaCAE-1, which coexists with blaAFM-1 on the plasmid. In addition, we performed a comprehensive genomic comparison of blaAFM-bearing sequences to gain a better understanding of the dissemination patterns of this novel carbapenemase gene.
Materials and methods
Bacterial isolation and in vitro susceptibility testing
C. aquatica SCLZS63 was recovered from the sewage outlet of the affiliated hospital of Southwest Medical University, Sichuan Province, China, in November 2019. As previously described21, 5 ml of water sample was concentrated by centrifugation for 5 min at 5000 g, and the sediment was resuspended in sterile 0.9% NaCl solution, then, the bacterial suspension was plated onto MacConkey agar containing meropenem (2 μg/ml) and incubated for 24 h at 37 °C. A single colony was picked, and initial species identification was performed by sequence analysis of 16S rRNA gene after PCR amplifying and Sanger sequencing22. The susceptibility to ceftazidime, cefotaxime, aztreonam, meropenem, and imipenem for SCLZS63 was examined by using the broth microdilution method and interpreted according to the Clinical and Laboratory Standards Institute (CLSI) guidelines for other non-enterobacterales bacteria23.
Genome sequencing and analysis
Genomic DNA of SCLZS63 was obtained using the QIAamp DNA Mini Kit (Qiagen), and the purified DNA was subjected to whole-genome sequencing on a HiSeq 2000 platform (Illumina, San Diego, CA, USA) using a paired-end library with an insert size of 150 bp, followed by the long-read MinION Sequencer (Nanopore, Oxford, UK). The de novo hybrid assembly of the Illumina reads and MinION reads were carried out by using Unicycler v0.4.324. Gene annotation for the assembled genomes was performed with the RAST tools25 and BLASTp/BLASTn searches against the UniProtKB/SwissProt database26. Bacterial precise species were identified using pairwise ANI analysis between strain SCLZS63 and reference genomes of Comamonas with the online software JSpeciesWS (https://jspecies.ribohost.com/jspeciesws/). A > 96% ANI cut-off was used for species circumscription27. Plasmid incompatibility types, antibiotic resistance genes, and insertion elements were predicted using PlasmidFinder 2.1 (95%, minimum threshold for identity; 60%, minimum coverage)28, ResFinder (90%, minimum threshold for identity; 60%, minimum coverage)29, and ISfinder30.
Phylogenetic analysis
Amino acid sequences of 17 class A β-lactamases were retrieved from the Beta-Lactamase DataBase (BLDB, http://bldb.eu/), and were utilized for the phylogenetic analysis. Sequences were aligned using the program Clustal W31. A maximum-likelihood tree was generated by MEGA 6 software32, with 1000 bootstrap replicates, and was then annotated using iTOL33.
Gene cloning
PCR amplification of the complete coding sequence of blaCAE-1 and its promoter region from C. aquatica SCLZS63 was performed using the primers blaCAE-F: 5′-cagcaaatgggtcgcggatccGCTTACTTTCACTCATGACGTCACC-3′and blaCAE-R: 5′-gtggtggtggtggtgctcgagGGATGTTGGAAGACCCGACC-3′. The resulting PCR fragment was then ligated into an expression vector pET28a using a ClonExpress® II One Step Cloning Kit (Vazyme, China) to construct pET28a-blaCAE-1, which was introduced into Escherichia coli DH5α by chemical transformation. Potential transformants containing the recombinant plasmid were selected on Luria–Bertani (LB) agar plates containing 50 mg/L kanamycin and verified by PCR assays. The susceptibility to antimicrobial agents (ampicillin, piperacillin/tazobactam, ampicillin/sulbactam, cefazolin, ceftriaxone, ceftazidime, cefotaxime, cefepime, aztreonam, ertapenem, meropenem, and imipenem) for the recombination strain was performed using the broth microdilution method according to the CLSI guidelines with E. coli strain ATCC 25922 as the quality control strain. E. coli DH5α containing the empty pET28a served as a negative control.
Conjugation and electroporation experiments
Conjugation experiments were performed using both broth- and filter-based methods, with the azide-resistant E. coli J53 as the recipient, as described previously34. Equal amounts of donor and recipient cells at the exponential stage (the optical density at 600 nm reaches ~ 0.5) were mixed and incubated at 37 °C in LB broth or on the filter that was placed on an LB agar plate overnight. Subsequently, cells were resuspended and diluted in 0.9% NaCl, and potential transconjugants were selected on LB agar plates containing 150 µg/ml sodium azide and 4 µg/ml cefotaxime. Conjugation assays were repeated with different donor/recipient ratios.
Electroporation was carried out with E. coli DH5α as the recipient. Plasmids of C. aquatica SCLZS63 were extracted using the E.Z.N.A. plasmid Mini Kit I (OMEGA, Bio-Tek, USA), verified by agarose gel electrophoresis, and then transferred by electroporation (Micro-Pulser electroporator; Bio-Rad, USA) into DH5α competent cells. Transformants were selected on LB agar plates containing 4 µg/ml cefotaxime. The presence of blaCAE-1 in the transformant was examined by PCR assays with primers blaCAE-F/R.
Comparative analysis of bla AFM-bearing sequences
To obtain the blaAFM-bearing sequences, a BLASTn with standard options was performed with the nucleotide sequences of blaAFM-1 (GenBank accession no. NG_063835) as a query in the NCBI GenBank database. Plasmids and chromosomes with a full-length hit to blaAFM-1 (100% query coverage and ≥ 99.88% identity) were selected. Alignments of the blaAFM-bearing sequence were performed using BLASTn and visualized with Easyfig v 2.2.3.
Results
Genome feature of C. aquatica strain SCLZS63
Strain SCLZS63 was initially identified as a Comamonas strain, which was resistant to ceftazidime (MIC, 128 μg/ml), cefotaxime (MIC, 128 μg/ml), aztreonam (MIC, 256 μg/ml), meropenem (MIC, 16 μg/ml), and imipenem (MIC, 16 μg/ml), respectively. Whole-genome sequencing (WGS) revealed that SCLZS63 contains one single circular chromosome with a size of 4,048,791 bp and an average GC content of 64.53%, and three circular plasmids, p1_SCLZS63, p2_SCLZS63, and p3_SCLZS63 (Table 1). SCLZS63 belongs to C. aquatica as it had 96.55% identity (81.16% coverage) to the C. aquatica reference strain NEB418 by average nucleotide identity (ANI) analysis. It had 9 known ARGs, mediating resistance to aminoglycosides [aac(6')-IIa, aph(6)-Id, and aph(3'')-Ib], β-lactams (blaAFM-1), macrolides [mph(E) and msr(E)], sulfonamides (sul1), trimethoprim (dfrA5), and amphenicols (cmx). Among them, aph(6)-Id and aph(3'')-Ib are located on the chromosome, while the remainder ARGs are carried by p1_SCLZS63.
CAE-1 confers resistance to several β-lactams
Notably, a 909-bp open reading frame (ORF) encoding a putative 302-amino-acid class A β-lactamase was identified on p1_SCLZS63. The novel class A β-lactamase is most closely related to CzoA-135 (Accession no. EFI60385), with only 52.7% amino acid identity, followed by PAU-136 (48.8%, APC57487), and AXC-1 (48.3%, ATG32091) (Fig. 1A). Therefore, it was given a new family name CAE-1 (for C. aquatica enzyme). The novel CAE-1 contains the four conserved motifs of class A β-lactamases, namely, 70SXXK73, 130SDN132, 166EXXXN170, and 234KTG23637,38 (Fig. 1B). Only one amino acid residue 69C that is associated with carbapenemase activity was identified in CAE-137 (Fig. 1B).
Comparison of CAE-1 with other known class A beta-lactamase. (a) An unrooted phylogenetic tree of CAE-1 and its close homologs and representative class A beta-lactamase. Proteins with carbapenemase activity are highlighted in blue and CAE-1 is in red. The tree scale indicates substitutions per site. The β-lactamases (GenBank accession numbers) are CTX-M-1(DQ915955), KPC-2 (LDDY01000008), SFC-1(AY354402), FRI-1(KT192551), SME-1 (Z28968), IMI-1 (U50278), MM3-1 (MK831000), PAD-1(LNTU01000040), GPC-1 (MH211206), XCC-16 (QUZR01000001), L2-28 (CP011010), HBL-1 (CP012077), BOR-1 (BX640443), PSV-1 (KU926347), PAU-1 (KU881625), AXC-1 (MF767301), and CzoA-1 (ADVQ01000062). (b) Amino acid sequence alignment of CAE-1 with CzoA-1, PAU-1, FRI-1, IMI-1, KPC-2 and SME-1. The residue numbers are positioned above the sequences according to the standard numbering scheme for the class A beta-lactamases39. The four conserved motifs of class A β-lactamases are outlined in black and the red triangle indicates the conserved residue 69C among class A carbapenemases.
To determine whether CAE-1 mediates resistance to β-lactams or not, blaCAE-1 was cloned into the pET28a vector and transformed into E. coli DH5α. Compared with the control strain DH5α carrying the empty pET28a, the recombinant strain DH5α/pET28a-blaCAE-1 exhibited resistance to some β-lactams tested, including ampicillin, piperacillin, cefazolin, cefuroxime, and ceftriaxone (Table 2), and the MIC for ampicillin-sulbactam increased for two-fold. The acquisition of blaCAE-1 had no effect on the MICs of carbapenems and aztreonam. This finding implies that CAE-1 has broad-spectrum activity profiles, and functions as broad-spectrum β-lactamase.
The mobilization and possible origin of bla CAE-1
We screened the presence of blaCAE-1 in GenBank database by BLASTn (Accessed by 20 August 2022), and four matches were found, including one plasmid from C. aquatica (CP079746, China, water, 2019), and three chromosomes of P. aeruginosa (CP042967, patient, Thailand, 2016), C. aquatica (LR813086, Spain, Water, 2020) and Comamonas thiooxydans (CP063057, China, patient, 2019). Analysis of the adjacent genetic elements of blaCAE-1 in p1_SCLZS63 and the above four blaCAE-1-carrying sequences showed that blaCAE-1 is always reversely located at the immediate upstream of the LysR family transcriptional activator-encoding gene ampR (Fig. 2), which consists with the commonly seen lysR-accompanied class A β-lactamases as described previously36. In all sequences except pB1A, the blaCAE-1-ampR element is bounded by an intact IScaq2 (upstream of blaCAE-1) and a ΔIScaq2 (immediately downstream of blaCAE-1) that is truncated by a parallel IScaq1 (Fig. 2). IScaq1/IScaq2 (Accession no. LR813086/CP063057) are IS5 family transposases that are initially identified in C. aquatica in the GenBank database. However, the flanking IScaq2 elements are in opposite orientations, which denies the assumption of an IScaq2-based composite transposon. In pB1A, the blaCAE-1-ampR element is bracketed by two IS110 family transposases, which are in the reversed orientation as well (Fig. 2). The exact mechanism of the mobilization of blaCAE-1 remains unclear.
Genetic contexts of blaCAE-1. The position and transcriptional direction of ORFs are indicated by arrows. blaCAE-1 and mobile genetic elements are highlighted in red and yellow, respectively. Regions of > 90% nucleotide sequence identity are marked by grey shading. Δ represents truncated insertion sequences.
To investigate the possible origin of blaCAE-1, BLASTp search of CAE-1 against non-redundant protein database was performed. We found that CAE-1 shows significant amino acid identity with chromosomally encoded class A β-lactamase of bacteria mainly from Comamonas and Acidovorax sp., which both belong to the family Comamonadaceae. Additionally, we found that the average GC-content for the immediate genetic background of the blaCAE-1 gene (namely, the blaCAE-1-ampR element) is 63.81%, which is close to the chromosomal GC-content of Comamonas (64.53% for SCLZS63). These findings suggest that Comamonadaceae may be an ancestral source of the blaCAE-1 gene.
bla CAE-1 and bla AFM-1 coexist on a novel type of plasmid p1_SCLZS63
The resistant plasmid p1_SCLZS63 is 143,067 bp in size, with GC content of 56.68%, and contains 141 ORFs. The backbone of p1_SCLZS63 includes regions responsible for plasmid replication (repA), maintenance (parAB, umuCD), and conjugative transfer (tra genes), and it could not be assigned into any known incompatibility group. No similar plasmids were found in the GenBank database using the backbone sequences of p1_SCLZS63 as a query, suggesting that p1_SCLZS63 is a novel type of plasmid. Within the backbone, p1_SCLZS63 harbors two multidrug-resistant (MDR) regions and an arsenate resistance operon (arsCDABCH) (Fig. 3A). In the 7.4-kb MDR1 region, the class 1 integron intI1-dfrA5-aac(6')-IIa-qacEΔ1-sul1 is bounded by an IS5 element (upstream) and an IS6100 (downstream).
Genetic characterization of p1_SCLZS63. (a) Circular organization of p1_SCLZS63. Arrows on the outer ring indicate deduced ORFs and their orientations. Backbone genes for replication, conjugal transfer, and plasmid maintenance are highlighted in green, brown, and light blue, respectively. blaCAE-1 and blaAFM are colored red, and other resistance genes are in fuchsia. Insertion elements, integrase genes, and genes for heavy metal resistance (ars and mer gene clusters) are indicated in yellow, dark blue, and olive, respectively. Two multidrug-resistant regions (MDR1 and MDR2) are indicated. Δ represents truncated genes. (b) Linear comparison of the MDR2 region in p1_SCLZS63 with related regions. Arrows indicate deduced ORFs and their orientations. Regions of > 90% nucleotide sequence identity are indicated by grey shading. The arrow colors are the same as in (a).
The 32-kb MDR2 region shows a complex structure and is a mosaic with areas of diverse origin (Fig. 3B). The Tn3-borne defective mercury resistance operon (mer) and the neighboring blaCAE-1 region were remarkably similar (99.9% identity, 100% coverage) to a region on the chromosome of P. aeruginosa strain PA99 (Accession no. CP042967). The msr(E)-mph(E) unit flanked by IS26 is universally found in many bacterial families, such as Enterobacteriaceae and Moraxellaceae. The blaAFM-1 region resembled that on the plasmid pSS332-218k (Accession no. CP071152) from a clinical Aeromonas caviae in Zhejiang, China. In this region, two copies of intI1 (one complete and one truncated)-sul1 element in parallel orientation bracketed the core blaAFM-1 platform, wherein the blaAFM-1 is located at a conserved fragment ISCR29-ΔgroL-blaAFM-1-ble-ΔtrpF-ΔISCR27-msrB-msrA-yfcG-corA, as had been reported previously in other plasmids19,20.
To determine the transfer ability of p1_SCLZS63, conjugation experiments were carried out with E. coli J53 as the recipient. Despite repeated attempts, no transconjugants containing p1_SCLZS63 were obtained. In addition, the transfer of p1_SCLZS63 into E. coli DH5α by electroporation was also unsuccessful after several attempts.
Genetic contexts of bla AFM alleles
A total of 14 blaAFM-bearing plasmids (n = 11) and chromosomes (n = 2) were retrieved from the GenBank database (Accessed on 19 October 2022). blaAFM was identified on the chromosomes of S. maltophilia, B. trematum, and P. aeruginosa, and it was also carried by different types of plasmids with various sizes (61,915–495,621 bp) from Comamonas testosterone, A. caviae, P. aeruginosa, Pseudomonas asiatica, A. faecalis, and C. aquatica from China. These blaAFM-bearing plasmids are generally untypable, except that some IncP-2 type plasmids (pAR19640, pNDTH9845, and pWTJH17) carry blaAFM-2 in P. aeruginosa, and an IncW plasmid pAN70-1 carries blaAFM-1 in A. faecalis.
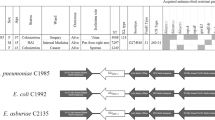
Generally, two kinds of blaAFM-bearing core modules were identified, namely ISCR29-ΔgroL-ΔfloR-blaAFM-ble-ΔtrpF-ΔISCR27-msrB-msrA-yfcG-corA (designated type A) and its truncated version ISCR29-ΔgroL-ΔfloR-blaAFM-ble-ΔtrpF-ΔISCR27 (type B). The truncation of type B seems to have resulted from the recombination event of ΔISCR27 (Fig. 4). Almost all the blaAFM-1 and blaAFM-4 genes are found in the type A module, except that blaAFM-1 in pMD9A is in the type B module, in which form blaAFM-2 and blaAFM-3 genes are embedded, and that the blaAFM-1-bearing type A module in pAN70-1 is disrupted by a Tn6346-like transposon19. In addition, we found that almost all the blaAFM-bearing core modules are always flanked by class 1 integrons (Fig. 4). The intI1-sul1-blaAFM-1 core module-ΔintI1-sul1 in p1_SCLZS63 in this study seems to be an ancestor structure, from which class 1 integrons with different cassette arrays surrounding the blaAFM core module have arisen.
The genetic contexts and mobilization mechanisms of blaAFM genes. The position and transcriptional direction of ORFs are indicated by arrows. The blaAFM genes are highlighted in red and other resistance genes are in fuchsia. Insertion elements and integrase genes are colored yellow and dark blue, respectively. Genes in the blaAFM-bearing core module are indicated in green. The strain or plasmid names, species, isolation sources, and GenBank accession numbers are shown. Δ represents truncated genes or elements.
Discussion
Antimicrobial resistance represents a growing threat to medical care. Infections caused by carbapenem-resistant bacteria are usually associated with poor prognoses and increased morbidity and mortality rates40. Close monitoring of carbapenem-resistant bacteria in the hospital sewage is essential. Comamonas, especially, carbapenem-resistant Comamonas, are abundant in wastewater16, while their genome characteristics of antibiotic resistance are poorly characterized. In this work, we isolated a carbapenem-resistant C. aquatica from hospital sewage, and its genetic information of drug resistance was characterized by high-resolution WGS. C. aquatica has been isolated as the causative agent of bacteremia and septic shock41. The prevalence of multidrug-resistance, especially carbapenem resistance in C. aquatica warrants further public health surveillance.
Ambler classe A β-lactamases are prevalent and diverse considering other molecular classes, and they represent the most important enzymatic source of both natural and acquired resistance to β-lactams in Gram-negative bacilli38. In this study, a new enzyme CAE-1 has been added to the list of class A serine β-lactamase. It exhibits resistance to ampicillin, piperacillin, cefazolin, cefuroxime, and ceftriaxone, as revealed by the in vitro susceptibility analysis. To confirm this, kinetics analysis on pure CAE-1 enzyme would have been informative. With this goal in mind, the blaCAE-1 gene without its promoter region was cloned in a pET28a expression vector and introduced in an E. coli expression strain. However, repeated attempts at expressing and purifying a N-terminus his-tagged version of the CAE-1 enzyme (expected molecular mass of ~ 34 kDa) proved unsuccessful. Possible reasons for why the heterologous expression did not work include low expression under detection limit and an inappropriate host. Further work will be needed to understand the enzyme kinetics of CAE-1.
Plasmids play a vital role in the dissemination of ARGs via horizontal gene transfer. Antibiotic resistance plasmids are rarely reported in Comamonas. Two IncP-1 plasmids were previously reported in Comamonas from aquatic environments, one of which was associated with the degradation of dyes42, and the other one was involved in resistance to heavy metal and oxidative stress16. None of the two IncP-1 plasmids carry any ARGs. In the strain SCLZS63, most ARGs are located on the untypable plasmid p1_SCLZS63, including the carbapenem resistance gene blaAFM-1. The previously reported carbapenemase genes blaGES-5, and blaIMP-8 in Comamonas are both chromosomally located16,18. The emergence of the carbapenemase gene on the plasmid in Comamonas has important public health implications, which demands more attention. The conjugation experiments indicated that the blaAFM-1-harboring p1_SCLZS63 is non-transmissible in this case. While the genomic sequences presented here infer the transfer potential of p1_SCLZS63, the unsuccessful conjugation might result from the exceptionally low transferability that is below detectable limits, or the non-replication of this plasmid due to the unsuitability of E. coli J53 recipient strain used in this study for p1_SCLZS63 from C. aquatica. Electroporation of p1_SCLZS63 into E. coli DH5α was also not successful, which again implies that p1_SCLZS63 may not be readily maintained in a different host of E. coli cells.
AFM is a newly identified subclass B1b metallo-β-lactamase, which shows partial identity to the widespread NDM19. blaAFM has been found in different species of non-fermenting Gram-negative bacteria and is spreading in clinical P. aeruginosa isolates at a low rate in China43,44. ISCR elements are known to move and pick up adjacent genetic components via a rolling-circle mechanism45. In the cases of blaAFM alleles, ISCR29 may have initially acquired the ΔgroL-blaAFM-ble-ΔtrpF-ΔISCR27-msrBA-yfcG-corA-ΔISCR27 region and mobilized it upstream of an intI1-sul1 genetic segment, meanwhile truncating the intI1 gene, followed by a second insertion downstream of the second intI1-sul1 segment. The intI1-sul1-blaAFM-1 core module-ΔintI1-sul1 structure in plasmid like p1_SCLZS63 serves as the intermediate source for the dissemination of blaAFM, most likely by recombination events. The flanking class 1 integrons have acquired a variety of passenger genes in the subsequent genetic actions, generating complex genetic contexts for blaAFM.
Conclusions
In the present study, we identified and characterized a novel class A β-lactamase gene blaCAE-1 conferring resistance to broad-spectrum cephalosporin, from an environmental C. aquatica isolate, where blaCAE-1 coexists with the carbapenemase gene blaAFM-1 in a mosaic MDR region on a novel type of plasmid. This finding highlights the importance of the environment as a reservoir of novel antibiotic resistance plasmids and resistance determinants. Effective surveillance is required for understanding the transmission and ongoing evolution of this multidrug-resistance plasmid in clinical settings.
Data availability
The complete sequences of the C. aquatica SCLZS63 chromosome and plasmids p1_SCLZS63, p2_SCLZS63, and p3_SCLZS63 have been submitted to GenBank with accession numbers CP104279 to CP104282, respectively.
References
Wu, Y., Zaiden, N. & Cao, B. The core- and pan-genomic analyses of the genus Comamonas: From environmental adaptation to potential virulence. Front. Microbiol. 9, 3096. https://doi.org/10.3389/fmicb.2018.03096 (2018).
Ma, Y. F. et al. The complete genome of Comamonas testosteroni reveals its genetic adaptations to changing environments. Appl. Environ. Microbiol. 75, 6812–6819. https://doi.org/10.1128/AEM.00933-09 (2009).
Zhu, G. et al. How bioaugmentation with Comamonas testosteroni accelerates pyridine mono-oxygenation and mineralization. Environ. Res. 193, 110553 (2021).
Tiwari, S. & Nanda, M. Bacteremia caused by Comamonas testosteroni an unusual pathogen. J. Lab. Phys. 11, 87–90. https://doi.org/10.4103/JLP.JLP_116_18 (2019).
Sammoni, A., Abdalah, A. & Al-Aissami, M. Comamonas testosteroni bacteremia: A rare unusual pathogen detected in a burned patient: Case report and literature review. Ann. Med. Surg. (Lond.) 75, 103371. https://doi.org/10.1016/j.amsu.2022.103371 (2022).
Almuzara, M., Cittadini, R., Estraviz, M. L., Ellis, A. & Vay, C. First report of Comamonas kerstersii causing urinary tract infection. New Microbes New Infect. 24, 4–7. https://doi.org/10.1016/j.nmni.2018.03.003 (2018).
Almuzara, M. N. et al. Intra-abdominal infections due to Comamonas kerstersii. J. Clin. Microbiol. 51, 1998–2000. https://doi.org/10.1128/JCM.00659-13 (2013).
Arda, B. et al. Comamonas testosteroni meningitis in a patient with recurrent cholesteatoma. APMIS 111, 474–476. https://doi.org/10.1034/j.1600-0463.2003.1110404.x (2003).
Bush, K. & Bradford, P. A. Interplay between beta-lactamases and new beta-lactamase inhibitors. Nat. Rev. Microbiol. 17, 295–306. https://doi.org/10.1038/s41579-019-0159-8 (2019).
Bonomo, R. A. Beta-lactamases: A focus on current challenges. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a025239 (2017).
Bush, K. Past and present perspectives on beta-lactamases. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.01076-18 (2018).
Hernando-Amado, S., Coque, T. M., Baquero, F. & Martinez, J. L. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat. Microbiol. 4, 1432–1442. https://doi.org/10.1038/s41564-019-0503-9 (2019).
Castanheira, M., Simner, P. J. & Bradford, P. A. Extended-spectrum beta-lactamases: An update on their characteristics, epidemiology and detection. JAC Antimicrob. Resist. 3, dlab092. https://doi.org/10.1093/jacamr/dlab092 (2021).
Karkman, A., Do, T. T., Walsh, F. & Virta, M. P. J. Antibiotic-resistance genes in waste water. Trends Microbiol. 26, 220–228. https://doi.org/10.1016/j.tim.2017.09.005 (2018).
Marathe, N. P. et al. Sewage effluent from an Indian hospital harbors novel carbapenemases and integron-borne antibiotic resistance genes. Microbiome 7, 97. https://doi.org/10.1186/s40168-019-0710-x (2019).
Hem, S. et al. Genomic analysis of carbapenem-resistant Comamonas in water matrices: Implications for public health and wastewater treatments. Appl. Environ. Microbiol. 88, e0064622. https://doi.org/10.1128/aem.00646-22 (2022).
Le, T. H. et al. Occurrences and characterization of antibiotic-resistant bacteria and genetic determinants of hospital wastewater in a tropical country. Antimicrob. Agents Chemother. 60, 7449–7456. https://doi.org/10.1128/AAC.01556-16 (2016).
Guo, X. et al. Emergence of IMP-8-producing Comamonas thiooxydans causing urinary tract infection in China. Front. Microbiol. 12, 585716. https://doi.org/10.3389/fmicb.2021.585716 (2021).
Zhang, X. et al. Characterization of the novel plasmid-encoded MBL gene blaAFM-1, integrated into a blaIMP-45-bearing transposon Tn6485e in a carbapenem-resistant Pseudomonas aeruginosa clinical isolate. J. Antimicrob. Chemother. 77, 83–88. https://doi.org/10.1093/jac/dkab342 (2021).
Lin, X. et al. Molecular and functional characterization of a novel plasmid-borne bla NDM-like gene, blaAFM-1, in a clinical strain of Aeromonas hydrophila. Infect. Drug Resist. 14, 1613–1622. https://doi.org/10.2147/IDR.S297419 (2021).
Li, Y. et al. Genetic and virulence characteristics of a Raoultella planticola isolate resistant to carbapenem and tigecycline. Sci Rep 12, 3858. https://doi.org/10.1038/s41598-022-07778-0 (2022).
Lane, D. J. 16S/23S rRNA sequencing. Nucleic Acid Techniques in Bacterial Systematics 115–175 (1991).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 29th Edition. CLSI supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute (2019).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595. https://doi.org/10.1371/journal.pcbi.1005595 (2017).
Aziz, R. K. et al. The RAST server: Rapid annotations using subsystems technology. BMC Genomics 9, 75. https://doi.org/10.1186/1471-2164-9-75 (2008).
Boutet, E. et al. UniProtKB/Swiss-Prot, the manually annotated section of the UniProt KnowledgeBase: How to use the entry view. Methods Mol. Biol. 1374, 23–54. https://doi.org/10.1007/978-1-4939-3167-5_2 (2016).
Rossello-Mora, R. & Amann, R. Past and future species definitions for Bacteria and Archaea. Syst. Appl. Microbiol. 38, 209–216. https://doi.org/10.1016/j.syapm.2015.02.001 (2015).
Carattoli, A. & Hasman, H. PlasmidFinder and in silico pMLST: Identification and typing of plasmid replicons in whole-genome sequencing (WGS). Methods Mol. Biol. 2075, 285–294. https://doi.org/10.1007/978-1-4939-9877-7_20 (2020).
Bortolaia, V. et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 75, 3491–3500. https://doi.org/10.1093/jac/dkaa345 (2020).
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 34, D32-36. https://doi.org/10.1093/nar/gkj014 (2006).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948. https://doi.org/10.1093/bioinformatics/btm404 (2007).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 47, W256–W259. https://doi.org/10.1093/nar/gkz239 (2019).
Li, Y. et al. Whole-genomic analysis of NDM-5-producing Enterobacteriaceae recovered from an urban river in China. Infect. Drug Resist. 14, 4427 (2021).
Zhuang, W., Liu, H., Li, J., Chen, L. & Wang, G. Regulation of class A beta-lactamase CzoA by CzoR and IscR in Comamonas testosteroni S44. Front. Microbiol. 8, 2573. https://doi.org/10.3389/fmicb.2017.02573 (2017).
Wang, J. et al. PAU-1, a novel plasmid-encoded ambler class A beta-lactamase identified in a clinical Pseudomonas aeruginosa isolate. Infect. Drug Resist. 12, 3827–3834. https://doi.org/10.2147/IDR.S225288 (2019).
Majiduddin, F. K. & Palzkill, T. Amino acid residues that contribute to substrate specificity of class A beta-lactamase SME-1. Antimicrob. Agents Chemother. 49, 3421–3427. https://doi.org/10.1128/AAC.49.8.3421-3427.2005 (2005).
Philippon, A., Jacquier, H., Ruppe, E. & Labia, R. Structure-based classification of class A beta-lactamases, an update. Curr. Res. Transl. Med. 67, 115–122. https://doi.org/10.1016/j.retram.2019.05.003 (2019).
Ambler, R. P. et al. A standard numbering scheme for the class A beta-lactamases. Biochem. J. 276(Pt 1), 269–270. https://doi.org/10.1042/bj2760269 (1991).
Perez, F. & Bonomo, R. A. Carbapenem-resistant Enterobacteriaceae: Global action required. Lancet Infect. Dis https://doi.org/10.1016/s1473-3099(19)30210-5 (2019).
Kaeuffer, C. et al. First case of Comamonas aquatica bacteremia complicated by septic shock. Med. Mal Infect. 48, 540–542. https://doi.org/10.1016/j.medmal.2018.08.004 (2018).
Stolze, Y. et al. IncP-1beta plasmids of Comamonas sp. and Delftia sp. strains isolated from a wastewater treatment plant mediate resistance to and decolorization of the triphenylmethane dye crystal violet. Microbiology (Reading) 158, 2060–2072. https://doi.org/10.1099/mic.0.059220-0 (2012).
Chen, M. et al. Plasmid-borne AFM alleles in Pseudomonas aeruginosa clinical isolates from China. Microbiol. Spectr. https://doi.org/10.1128/spectrum.02035-22 (2022).
Li, Y. et al. Alcaligenes faecalis metallo-beta-lactamase in extensively drug-resistant Pseudomonas aeruginosa isolates. Clin. Microbiol. Infect. 28, 880 e881-880 e888. https://doi.org/10.1016/j.cmi.2021.11.012 (2022).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. https://doi.org/10.1128/CMR.00088-17 (2018).
Funding
This work was supported by National Natural Science Foundation of China (31900125), Scientific and technological project in Sichuan Province (2022JDRC0144), the Joint Funds of the Luzhou and Southwest Medical University Natural Science Foundation (2020LZXNYDJ34).
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.L. and L.Z.; methodology, C.F. and Q.L.; software, X.D.; formal analysis, X.D.; resources, Y.Q.; writing-original draft preparation, Y.L.; writing-review and editing, L.Z. and X.W.; funding acquisition, Y.L. and L.Z. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Fang, C., Wang, X. et al. A new class A beta-lactamase gene blaCAE-1 coexists with blaAFM-1 in a novel untypable plasmid in Comamonas aquatica. Sci Rep 13, 3634 (2023). https://doi.org/10.1038/s41598-023-28312-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-28312-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.