Abstract
The Korean water deer (Hydropotes inermis argyropus) is considered a vermin in Korea because it damages crops, but also listed as a vulnerable species on the IUCN’s red list. Therefore, it is indispensable to manage them appropriately by understanding the ecology such as food habits. Here, we aimed to apply high-throughput sequencing (HTS), a sensitive and objective method, to investigate the dietary composition of the Korean water deer inhabiting the lowland and forest areas in summer and winter. We targeted the internal transcribed spacer 2 (ITS2) region for plant identification. From a total of 40 fecal samples analyzed, 63 plant genera were identified, with Morus being the most abundant, and some of the plant taxa identified by HTS were detected for the first time as the diets of Korean water deer. By type, woody plants (68.6%) were the most predominant, followed by forbs (7.0%) and graminoids (0.7%). We found that the deer in the forest area ate more woody plants (84.6%) than those in the lowland area (52.7%). It was also found that the type of woody plants that the deer ate changed by season. Overall, our results indicate that the Korean water deer is a browser that is seasonally adaptable and feeds on a wide variety of woody plants. We expect that the results and genetics methods reported here, by parallelly investigating their habitat range and reproductive behavior in the future, will help the management and conservation of the Korean water deer, which is in contradictory situations.
Similar content being viewed by others
Introduction
The water deer (Hydropotes inermis) is a small bodied species in the family Cervidae, and tends to take solitary behaviors1. Unlike other Cervidae, the deer lacks antlers and they are characterized by their long canines2. The water deer consists of two subspecies inhabiting China (Hydropotes inermis inermis) and the Korean peninsula (Hydropotes inermis argyropus)2,3. Some of the populations were introduced to England and France for hunting game and exhibition purposes4 and the arrival of the deer has been reported in Russia recently5. The population of the Chinese water deer is declining due to habitat fragmentation and illegal hunting6, and the water deer is listed as a vulnerable species on the International Union for Conservation of Nature (IUCN) Red List6. However, the Korean water deer inhabits high density from lowlands to forest areas throughout the Korean Peninsula4, and the Ministry of Environment of Korea designated it as a vermin because it damages crops7. Nonetheless, it is also thought to face serious threats such as habitat changes and roadkills4. Therefore, more research (e.g., diet, reproduction, behavior) is needed to take appropriate management and conservation strategies in Korea.
Ruminants are grouped into browsers feeding on plant leaves, grazers feeding on grasses, and the intermediate feeders8. The water deer is an intermediate feeder close to a browser8. For instance, the Chinese water deer in Zhoushan Archipelago are found to feed on forbs as a main item followed by woods, ferns, and grasses as secondary items9. The Korean water deer in Daebu Island are found to mainly feed on forbs or woods10. Analysis of the gut contents in the Korean water deer, which died of traffic accidents or natural causes, revealed that forbs were main diets (70%)11. Another study conducted in Janghang wetland reported that the Korean water deer fed on forbs (49%), woods (36%), and graminoids (15%)12. However, studies conducted in England reported that the captive water deer fed mainly on graminoids and exhibited intermediate characteristics which might be nutritionally suboptimal13. Additionally, the deer is more likely to feed on barks or twigs of woody plants in winter season when fresh leaves are not available and increased consumption of woody plants in winter than in summer is observed9,12. Thus, habitat vegetation and seasonal differences may have influenced their dietary habits and types10,12.
Dietary analysis of wildlife has been traditionally done by examining the feeding traces and/or observing the morphology of gut or fecal contents14. However, the problem lies in the need of taxonomic expertise for identifying dietary contents15. Recently, high-throughput sequencing (HTS) technologies have been introduced to molecular ecology research16,17. These technologies have enabled higher taxonomic resolution18 and accurate dietary analysis without the need of special taxonomic expertise14,19. Due to its objectivity in species identification and the ability to read a large number of DNA sequences (and therefore high detection sensitivity), the technologies have become widely used in dietary analysis of wildlife, including endangered species20,21,22,23,24.
Here, we aimed to apply HTS to investigate the differences in the diet of the Korean water deer by habitats and seasons. Specifically, we collected fecal samples of the water deer in two seasons (summer and winter) in two distinctive habitats (lowland and forest) in Korea, from which DNA was extracted to sequence the internal transcription spacer 2 (ITS2) region of plants. So far, the diet of the water deer has been investigated by feeding sign observation in the field9,25, microhistological analysis of the feces9, and molecular analyses of the feces, such as polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE) of the rbcL gene26 and Sanger sequencing (after cloning) of the rbcL gene10, ITS2 and rbcL gene12. In this study, HTS was used to sequence the ITS2 region, which is a primary DNA barcode marker for plants27,28. The use of HTS is expected to provide better understandings of feeding habits of the water deer and their seasonal and regional characteristics. We expect that the results of this study will help provide basic information for proper management and conservation strategies for the water deer in Korea.
Materials and methods
Study sites and feces sampling
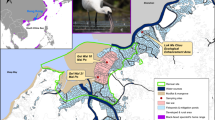
As the Korean water deer inhabit various areas from lowlands to mountainous areas in South Korea3,10, we conducted sampling in two different sites to test the diet differences between the habitat types. We selected the Taehwa Research Forest (TRF) as a representative forest for forest habitats and the Civilian Control Zone (CCZ) as a lowland shrub and meadow habitat representative. The elevation of TRF and CCZ is 150–641 m and 0–116 m above sea level respectively. Both sites are in Gyeonggi Province in South Korea (Fig. 1). TRF is an experimental forest administered by Seoul National University, located in Gwangju City, Gyeonggi Province (37°18′34″ N, 127°18′07″ E). TRF consists of both deciduous trees (e.g., Quercus mongolica and Quercus variabilis) and coniferous trees (e.g., Pinus koraiensis and Larix kaempferi) artificially planted in the 1970s. The other site CCZ (37°54′56″ N, 126°44′58″E) consists of forest vegetation dominated by Acer ginnala, Alnus japonica, and Quercus acutissima29, arable land, and artificial buildings, all of which are typical lowland landscapes of South Korea. As TRF is an experimental forest administered by Seoul National University and the CCZ has restricted access, these areas made for ideal sites for collecting the fecal samples with the fewest amount of human disturbance. The mean annual temperature in 2017 was 10.5 °C for CCZ and 11.7 °C for TRF according to the nearest meteorological observation centers of the Korean Meteorological Administration. Similarly, the total annual precipitation in 2017 was 948 mm in CCZ and 1,020 mm in TRF. The fecal samples were collected in June, July, and August 2017 in the summer, and in December 2017, January and February in 2018 in the winter. In each site in each season, 10 samples were collected, resulting in a total of 40 samples. Each sample consists of several fecal pellets collected from the same pile of fecal pellets. We collected fresh fecal samples that were wet, sticky on the outside. We also avoided sampling after rain and snow. Each of the collected samples was individually placed in a zipper bag, air-dried at room temperature, and stored at − 20 °C10.
DNA extraction
PowerMax Soil DNA extraction kit (Mobio Laboratory, Inc., Carlsbad, CA, USA) was used. From each sample, four fecal pellets were transferred into a conical tube and 4 mL of ultra-pure water was added. Using a sterile spatula, preliminary homogenization was done manually. Then, approximately 0.1 g of each homogenized sample was added to the 2 mL tube of the kit, with the addition of beads (300 mg of 0.1 mm diameter glass beads and 100 mg of 0.5 mm diameter glass beads). The homogenization was performed using a bead beater (BioSpec Products, Inc., Bartlesville, OK, USA) for 3 min at 3,000 g. After that, DNA was extracted as per the manufacturer’s instruction and eluted into 50 µL of TE (10 mM Tris–HCl, 1 mM EDTA, pH = 8.0). The eluted DNA extracts were kept at − 80 °C until the subsequent procedure.
DNA sequencing
Forty libraries were prepared with plant-specific universal primers of ITS-p3 (YGACTCTCGGCAACGGATA) and ITS-u4 (RGTTTCTTTTCCTCCGCTTA)30 to amplify its internal transcribed spacer 2 (ITS2) region. The adapter sequences were attached to universal primers for Illumina MiSeq (Illumina, Inc., San Diego, CA, USA). Each PCR mixture was comprised of a 25 µL of 2 × PCR Solution Premix Taq™ (Takara Bio Inc., Otsu, Shiga, Japan), 1 µL of the DNA extract, and 5 µL of each primer, and 14 µL of PCR-graded water. PCR amplifications were conducted with a T100™ thermal cycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA) and the thermal cycle was as follows: 10 min at 94 °C for initial denaturation, 35 cycles of 30 s at 94 °C, 40 s at 55 °C, 60 s at 72 °C, and 10 min at 72 °C for final extension. The PCR products were purified by AMPure XP beads (Beckman Coulter, Inc., Brea, CA, USA). Then, the purified PCR products were indexed with a Nextera XT Index kit (Illumina) and the thermal cycle was as follows: 3 min at 95 °C for initial denaturation 8 cycles of 30 s at 95 °C, 30 s at 55 °C, 30 s at 72 °C, and 5 min at 72 °C for final extension. The indexed libraries were purified by AMPure XP beads. The indexed and purified libraries were normalized by Quant-iT PicoGreen dsDNA reagent kit (Life Technologies, Carlsbad, CA, USA). The normalized libraries were pooled and loaded to a v3 600 cycle-kit reagent cartridge (Illumina) for 2 × 300 bp paired-end sequencing on an Illumina MiSeq system.
DNA sequence processing
The raw sequence reads were demultiplexed, and the reads with quality scores of less than 20 were removed by the MiSeq reporter v2.5 (Illumina). The subsequent procedure of sequence assembly, quality check, and assignment of taxonomy was performed using USEARCH v.11.0.66731. Reads with more than 1.0 expected errors were removed. The joined reads less than 200 bp were also removed. The default USEARCH settings were used for the identification of unique sequences. Using the UNOISE algorithm, chimeric reads were removed, and the sequence reads were clustered into zero-radius operational taxonomic units (ZOTUs). Each ZOTU was taxonomically assigned against the ITS2 database32,33 using the SINTAX algorithm with an 80% confidence level34. The ZOTUs with the same taxonomy identification were combined for further taxonomic analysis.
Data analyses
Prior to the diversity analyses, using the rrarefy function in vegan version 2.5–735, the number of sequence reads in each sample was normalized to the number of sequence reads of the sample from which the lowest number of sequence reads was obtained (i.e., 18,697 reads).
To see whether the Korean water deer has diet selectivity, α diversity analyses were conducted. For α diversity analyses, the rarefaction curves of observed ZOTU richness against the number of sequence reads were created using the iNEXT package version 2.0.2036. Alpha diversity indicators such as ZOTU richness, Shannon diversity index, and inverse Simpson index were calculated in vegan. To test for differences in the α diversity indicators between the habitats (forest vs. lowlands) and between the seasons (summer vs. winter), we used a two-way analysis of variance (ANOVA) with habitat and season as fixed factors. Prior to ANOVA, the normality and homoscedasticity of the data were checked and the data were logarithmically transformed if necessary.
To understand the diet composition differences between seasons and sites, we conducted β diversity analyses. For β diversity analyses, the compositional differences between the samples were visualized using nonmetric multidimensional scaling (NMDS) plotting based on the Bray–Curtis dissimilarity using the metaMDS function in vegan. The statistical differences in composition between sites and between seasons were tested with a permutational multivariate analysis of variance (PERMANOVA) with 999 permutations37 using the adonis function of vegan. The homogeneity of the compositional dispersion within the same site or within the same season was calculated using the betadisper function of vegan. A variation partitioning analysis was conducted to test the effect size of site and season on variation using the varpart function of vegan. The indicspecies package version 1.7.938 was used to perform the indicator species analysis with 999 permutations to identify plants with different abundances depending on the site or season. To visualize and compare the number of ZOTUs shared between sites and seasons, the Venn diagram illustrating the intersection of the sets of ZOTUs observed at each site and each season was created using the ggvenn package version 0.1.839.
For the diet type analyses, we classified detected plant genera into woody, forb, graminoid, or algae with reference to local flora data (Table S1). Some plants are classified as “not applicable” if the detected genus contains species with different growth forms, or if the taxonomic identification was ambiguous below the 80% confidence level. The local flora data for TRF were adopted from Choi, et al.40 and Ko and Shin41, Lee, et al.42 and those for CCZ were adopted from Kim and Kang43 and Gyeonggi Tourism Organization44. The differences in the diet type between the sites and between the seasons was statistically tested using the Wilcoxon rank sum test.
All graphs were created using the ggplot2 package version 3.3.545. All analyses were performed on R versions 3.5.3 to 4.0.346. The differences were considered significant if p < 0.05.
Results
Sequencing results
From a total of 40 libraries, 1,298,966 sequence reads and 1,544 ZOTUs were observed (Table S2). Each library contained 18,697 to 46,475 reads with an average of 32,474 reads. The number of ZOTUs ranged from 55 to 271 per library. The rarefaction curves reached asymptotes for all libraries (Fig. S1), with the sample coverage ranging from 0.9984 to 0.9999 (Table S2). The sampling of sequence reads to the lowest number per library resulted in a total of 747,880 reads and 1544 ZOTUs. After rarefaction, 50 to 264 ZOTUs were identified per library. These ZOTUs were taxonomically classified into 63 genera.
Taxonomic composition
From a total of 1,544 ZOTUs detected, 42 families and 63 genera of plants were identified. The dominant families and their mean relative abundances were Rosaceae (22.0%), Moraceae (19.0%), Betulaceae (10.0%), Fabaceae (8.7%), Fagaceae (2.9%), Asteraceae (2.5%), Amaranthaceae (2.0%), Pinaceae (1.6%), Cannabaceae (1.6%), and Aceraceae (1.0%) (Fig. 2a). At the genus rank, Morus (18.9%) was the most dominant, followed by Rubus (9.6%), Prunus (8.8%), Corylus (8.1%), Robinia (3.2%), Rosa (3.1%), Quercus (2.7%), Pueraria (2.6%), Humulus (1.6%), Pinus (1.1%), Carpinus (1.0%), Acer (1.0%), and Trifolium (1.0%) (Figs. 2b and 3). It can be seen that most of the detected genera are present in the local flora (Table S1).
Indicator species analysis
The indicator species analysis revealed eight indicator genera including Prunus, Quercus, Actinidia, and Corylus in TRF, and nine indicator genera including Humulus, Robinia, Ambrosia, and Glycine in CCZ (Table 1). All of the eight indicator genera in TRF were woody plants. The indicator genera found in CCZ consisted of forbs, woods, and graminoids. The seasonal analysis revealed two indicator genera of Rubus and Rosa in summer, and seven indicator genera including Prunus, Amorpha, Rhododendron, and Hovenia in winter (Table 2). In general, woody plants were the main indicator genera in both seasons. However, in winter, forbs such as Trifolium and Amaranthus, as well as graminoids such as Avena were also identified as indicator taxa.
Alpha diversity
ZOTU richness was not significantly different between the sites (p > 0.05; two-way ANOVA) (Fig. 4 and Table S3). However, ZOTU richness was significantly higher in winter than in summer with no interaction effect by site (p < 0.05; two-way ANOVA). Conversely, the sample-based rarefaction curves demonstrated that ZOTU richness was higher in summer than in winter (Fig. S2). No statistically significant difference in the Shannon diversity index was observed between the sites and between the seasons (p > 0.05; two-way ANOVA). Similarly, no statistically significant difference in the inverse Simpson index was observed between the sites and between the seasons (p > 0.05; two-way ANOVA).
Alpha diversity indices. (a) ZOTU richness by site. (b) Shannon diversity index by site. (c) Inverse Simpson index by site. (d) ZOTU richness by season. (e) Shannon diversity index by season. (f) Inverse Simpson index by season. There was no interaction between sites and seasons (p > 0.05; two-way ANOVA). n.s. denotes non-significant and * denotes significant at p < 0.05. The rarefied data were used.
Beta diversity
The distinct differences in plant assemblage structures were observed between the sites and between the seasons (Fig. 5). PERMANOVA test also demonstrated that the seasons explained about 12% of the compositional variance (pseudo-F = 5.78; p < 0.05, R2 = 0.120) and that the sites explained about 9% of the compositional variance (pseudo-F = 4.23; p < 0.05, R2 = 0.088) (Table S4). The statistically significant interaction between site and season was observed (pseudo-F = 2.18; p < 0.05, R2 = 0.045) although it explained only about 5% of the variance.
The variation partitioning analysis shows that the variation by season (R2adj = 0.13) was larger than the variation by site (R2adj = 0.03) with no interaction between season and site, suggesting season had greater explanatory power for compositional differences (Fig. S3). Moreover, the beta dispersion analysis shows that the dispersion from the centroid position was significantly higher in summer than in winter (p < 0.05; ANOVA), suggesting more diverse diet composition can be seen in summer than in winter (Fig. S4). However, the difference was insignificant between the sites (p > 0.05; ANOVA).
The shared ZOTU analysis using the Venn diagram shows that 446 ZOTUs (29%) were shared between the two sites while 507 ZOTUs (33%) and 592 ZOTUs (38%) were found only in the TRF and CCZ samples, respectively (Fig. S5). It also shows that 631 ZOTUs (41%) were shared between the two seasons while 572 ZOTUs (37%) and 342 ZOTUs (22%) were found only in summer and winter, respectively. Seventy-nine ZOTUs (5%) were found in all sites and all seasons. The largest number of site- and season-specific ZOTUs (274 ZOTUs) was observed in CCZ summer, while 177, 135, and 218 ZOTUs were observed uniquely in TRF summer, TRF winter, and CCZ winter, respectively.
Plant type
When plants were classified by growth morphology, the composition was 68.6%, 7.0%, and 0.7% for woods, forbs, and graminoids, respectively (Fig. 6). This trend was consistent across all sites and seasons. However, the relative abundance of each plant type differed between the sites and between the seasons. Specifically, TRF had more woody plants (84.6% of mean relative abundance) than CCZ (52.7%) (p < 0.05; Wilcoxon rank-sum test) (Fig. S6). Conversely, CCZ had more forb plants (12.4%) than TRF (1.5%) (p < 0.05; Wilcoxon rank-sum test). Similarly, CCZ had more graminoid plants (1.3%) than TRF (0.05%) (p < 0.05; Wilcoxon rank sum-test). The comparison between the seasons showed that the only woody plants significantly differed between winter (76.8%) and summer (60.5%) (p < 0.05; Wilcoxon rank-sum test).
Discussion
Understanding the diets of wildlife allows understanding of their ecological role and environmental interactions. In this study, we used HTS to investigate the diet of the Korean water deer, a vulnerable species on the IUCN’s Red List. From a total 40 fecal samples analyzed, we have detected 42 plant families and 63 plant genera. Our HTS-based study detected more plant taxa compared to the results of traditional dietary surveys reported so far for the Korean water deer (Table S5). This is consistent with the results of previous wildlife dietary studies comparing HTS with morphological analysis e.g.,18,47. The detected plants included those belonging to the families Rosaceae, Moraceae, Betulaceae, Fabaceae, Fagaceae, Asteraceae, and Amaranthaceae (Fig. 2a). These plants have also been reported as dietary items of the Korean water deer by previous studies using traditional methods10,11,12,26,48. Additionally, we identified, for the first time, plants belonging to the Pinaceae, Araliaceae, Potamogetonaceae, and Apiaceae families in the feces of the Korean water deer.
The seasonal effects are greater than the habitat effects
Using HTS, this study investigated the seasonal and site differences in the diet composition of the Korean water deer. Overall, we found that the seasons have greater impacts on diet than the sites, in both α (Table S3) and β diversity (Table S4). According to α diversity analysis, the deer seems to have selectivity on favored plant species. The large herbivores generally selectively feed on high-quality forage49,50. However, in winter, they consumed more diverse forage than in summer. In winter season, low-quality and availability of forage constrain forage consumption51; in response to this, the deer expand their diet range being less selective52,53. Meanwhile, as more diverse plant species are available in summer, the sample-based rarefaction curve showed the opposite trend (Fig. S2).
Likewise, the diet composition is mainly affected by season than habitat. The observed tendency appears to be the opposite of that reported in studies investigating other deer species. For example, it has been reported that altitude is the primary factor that affects the dietary differences, and season is secondary for the red deer (Cervus elaphus) in a mountainous landscape in New Zealand54. Similarly, a review study on the red deer in Europe reports that their diet composition is mainly influenced by habitat rather than season55. For the roe deer (Capreolus capreolus) in Europe, a review study reports that their diet is more habitat-influenced than seasonal56. One explanation for the opposite tendency observed in this study is that the habitat effects relative to the seasonal effects may have been smaller in this study than in other studies. Although we have selected two sampling sites with different landscapes, and the deer have a reported inhabitation range of approximately 2.77 km257 and are unlikely to migrate between the two sites, the two sites are relatively close (approximately 83 km in a straight line distance) (Fig. 1). This may have obscured the habitat effects compared to the seasonal effects. Consequently, the differences in the local flora, and thus the feeding habits of the water deer, may have become obscured. Despite the relatively weak habitat effect, it should be noted that this study, as in previous studies, confirmed the distinct difference in the diet composition between the two sites (lowland vs. forest) (Fig. 5 and Table S4).
The deer in forest areas eat more woody plants than those in lowland areas
The indicator species analysis revealed the plants specifically eaten by the Korean water deer in each site (Table 1). For example, the deer in the forest area (TRF) was found to specifically eat plants belonging to the genera Prunus, Quercus, Actinidia, Corylus, and Pinus. All of these genera are woody plants and the main constituents in this forest site. Meanwhile, the deer in the lowland area (CCZ) was found to specifically eat plants belonging to the genera Humulus, Robinia, Ambrosia, Glycine, Salix, Amorpha, Trifolium, Digitaria, and Acalypha. Of them, Humulus, Ambrosia, Glycine, Trifolium, and Acalypha are forbs, and Digitaria is a graminoid. These plants are known to be abundant in lowlands and open areas such as the lowland area of CCZ43. The analysis of plants by growth morphology also supports the observed tendency. For instance, we found higher proportion of woods in the forest area (TRF) than in the lowland area (CCZ) and higher proportion of forbs and graminoids in the lowland area (CCZ) than in the forest area (TRF) (Figs. 6 and S6). Our observations are consistent with those by Kim et al.10 who reported that more forbs (95%) were eaten by the Korean water deer in a lowland area and more woody plants (58%) than forbs (31%) were eaten by those in a mountainous area. These findings make sense because the water deer is known to opportunistically feed on readily available plants8.
The Korean water deer is a seasonally adaptable browser eating a wide variety of woody plants
In addition to the habitat differences, the seasonal differences were also observed. For instance, the proportion of woody plants was higher in winter (76.8%) than in summer (60.5%) in both sampling sites (Figs. 6 and S6). This was also in line with a previous investigation conducted in the Janghang wetland in Korea, by which the increased proportion of woody plants in winter (80%) than in summer (28%) was reported12. The increased proportion of woody plants in winter has also been reported in the Chinese water deer9. The reported tendency is reasonable since it is well known that the water deer feed on fresh leaves and sprouts in spring and summer, fruits in fall, and twigs, barks and buds of shrubs or trees in winter when fresh leaves are not available10,12. It is expected that the consumption of barks and/or twigs in winter greatly contributed to the increase in the proportion of woody plants.
Nevertheless, it was found that woody plants are consumed greatly not only in winter but also in summer (Fig. 6), suggesting that the water deer is a browser by its nature. This is supported by a morphophysiological study, too. For instance, the Chinese water deer introduced to the Whipsnade Zoo Park in England was reported to be predominantly grass-eating due to the lack of woody plants; however, a subsequent investigation suggested that the deer were undernourished due to the unbalanced diets13. In addition, although woody plants were predominant in both summer and winter, seasonal differences were observed in the types of woody plants consumed by the deer. For instance, Rubus and Rosa are found to be preferentially consumed in summer, and Prunus, Amorpha, Rhododendron and Hovenia are found to be preferentially consumed in winter (Table 2). From these, the Korean water deer can be regarded as a browser that is seasonally highly adaptable and feeds on a wide variety of woody plants.
Limitations and future prospects
In this study, we investigated the seasonal and habitat differences of diet composition of the Korean water deer. However, the dietary composition of deer is known to be influenced not only by season and site, but also by other factors, such as gender. For example, female red deer and white-tailed deer (Odocoileus virginianus) are known to eat higher quality food than males58,59. Conversely, male sika deer (Cervus nippon) are known to eat higher-protein diets than females in winter60. However, few studies have reported on sex differences in eating habits of the Korean water deer. In future research, in addition to dietary survey, it is desirous to perform sex determination, for example, by analyzing the sex-determining region Y (SRY) gene61.
Additionally, the gut microbiota of wild animals is related to host species, their diet, and environmental changes such as seasons62, and it is also crucial for maintaining health condition of its host63. Therefore, analyzing the gut microbiota of wild animals has been proposed as a conservation tool for wildlife, including endangered species64,65,66. In addition to the dietary analysis we have investigated in this study, we propose to analyze the microbiota to contribute to our understanding of the health status of Korean water deer and their protection and management in the future.
In this study, we used DNA barcode sequencing to investigate the dietary composition of the Korean water deer. Though this method is objective and convenient, it also has some drawbacks, as pointed out by previous research20,67,68,69. Most importantly, the parts of plants eaten (e.g., leaves, twigs, bark, and fruits) cannot be differentiated by DNA-based methods. Each part of the plant may have different nutritional values, so an accurate assessment of them is essential to know their ecology. In addition, each part of the plant may have different digestibility70. The differences in digestibility cause biases in the proportion of dietary content measured71,72,73. In addition to digestion biases, DNA barcoding can be biased for a variety of reasons, such as variations in the number of cells per plant tissue and the number of copies of the target DNA marker per cell69. Recognizing these uncertainties, the results obtained should be used for relative comparisons rather than absolute quantification19,67. Depending on the purpose, DNA barcoding may be better to be combined with traditional methods (e.g., morphological observation)47.
Conclusion
In this study, by analyzing fecal samples of the Korean water deer by DNA barcoding, we succeeded in elucidating their dietary tendencies by habitat and season. By using HTS, more plant taxa were detected compared to previous studies, and some taxa were identified for the first time as the diets of Korean water deer. Overall, we found that the Korean water deer is a browser that is seasonally highly adaptable and feeds on a wide variety of woody plants. Diet is associated with animal health, growth, survival, and reproduction15. Moreover, understanding their eating habits helps to elucidate their ecological roles, such as their interaction with the environment and their role in the local food web14,74, and to formulate conservation strategies such as habitat restoration19,75. We expect that our findings and methodologies reported here will be useful for future ecological surveys and for the formulation of their conservation and management strategies. For example, when planning habitat restoration for water deer in the future, it is necessary to restore vegetation with food items that they prefer because restoring vegetation with the target animal’s preferred diet item is suggested76,77. Furthermore, for the purpose of reproduction enhancement and population growth, supplementary feeding can be carried out78. At the same time, supplementary feeding can be conducted to reduce the damage to arable crops78,79. The effectiveness of supplementary feeding seems to relate to the feeding type of the target species80, therefore supplying favored forage would be effective. However, supplementary feeding may cause other consequences other than intended management goals79,81. Thus, beforehand, it is necessary to assess whether intended consequences can be obtained and consider the comprehensive impacts following supplementary feeding. In addition, in future research, in parallel with the food habits of the wildlife as investigated in this study, it is expected that more accurate conservation and management strategies will be established by investigating the ecology of the target wildlife, such as their habitat range and reproductive behavior.
Data availability
Raw sequence data are available at NCBI under the accession number PRJNA791828.
References
Schilling, A.-M. & Rössner, G. E. The (sleeping) beauty in the beast—a review on the water deer, Hydropotes inermis. Hystrix Ital. J. Mammal. 28, 121–133 (2017).
Geist, V. Deer of the World: Their Evolution, Behaviour and Ecology (Stackpole Books, Pennsylvania, 1998).
Cooke, A. Muntjac and Water Deer: Natural History, Environmental Impact and Management (Pelagic Publishing Ltd, UK, 2019).
Kim, B. J., Lee, B. K. & Kim, Y. J. Korean water deer (National Institute of Ecology, South Korea, 2016).
Belyaev, D. A. & Jo, Y.-S. Northernmost finding and further information on water deer Hydropotes inermis in Primorskiy Krai, Russia. Mammalia 85, 71–73 (2021).
Harris, R. B. & Duckworth, J. W. Hydropotes inermis. The IUCN Red List of Threatened Species, e.T10329A22163569 (2015).
National Institute of Biological Resources. Harmful wildlife. https://species.nibr.go.kr/home/mainHome.do?cont_link=011&subMenu=011016&contCd=011016001 (2015).
Hofmann, R. R. Evolutionary steps of ecophysiological adaptation and diversification of ruminants: a comparative view of their digestive system. Oecologia 78, 443–457 (1989).
Guo, G. & Zhang, E. Diet of the Chinese water deer (Hydropotes inermis) in Zhoushan Archipelago, China. Acta Theriol. Sin. 25, 122–130 (2005).
Kim, B. J., Lee, N. S. & Lee, S. D. Feeding diets of the Korean water deer (Hydropotes inermis argyropus) based on a 202 bp rbcL sequence analysis. Conserv. Genet. 12, 851–856 (2011).
Park, J.-E., Kim, B.-J., Oh, D.-H., Lee, H. & Lee, S.-D. Feeding habit analysis of the Korean water deer. Korean J. Environ. Ecol. 25, 836–845 (2011).
Kim, J., Joo, S. & Park, S. Diet composition of Korean water deer (Hydropotes inermis argyropus) from the Han River Estuary Wetland in Korea using fecal DNA. Mammalia 85, 487–493 (2021).
Hofmann, R., Kock, R. A., Ludwig, J. & Axmacher, H. Seasonal changes in rumen papillary development and body condition in free ranging Chinese water deer (Hydropotes inermis). J. Zool. 216, 103–117 (1988).
Nielsen, J. M., Clare, E. L., Hayden, B., Brett, M. T. & Kratina, P. Diet tracing in ecology: Method comparison and selection. Methods Ecol. Evol. 9, 278–291 (2018).
Birnie-Gauvin, K., Peiman, K. S., Raubenheimer, D. & Cooke, S. J. Nutritional physiology and ecology of wildlife in a changing world. Conserv. Physiol. 5, cox030 (2017).
Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C. & Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 21, 2045–2050 (2012).
Glenn, T. C. Field guide to next-generation DNA sequencers. Mol. Ecol. Resour. 11, 759–769 (2011).
Nichols, R. V., Åkesson, M. & Kjellander, P. Diet assessment based on rumen contents: A comparison between DNA metabarcoding and macroscopy. PLoS ONE 11, e0157977 (2016).
Pompanon, F. et al. Who is eating what: diet assessment using next generation sequencing. Mol. Ecol. 21, 1931–1950 (2012).
Kumari, P. et al. DNA metabarcoding-based diet survey for the Eurasian otter (Lutra lutra): Development of a Eurasian otter-specific blocking oligonucleotide for 12S rRNA gene sequencing for vertebrates. PLoS ONE 14, e0226253 (2019).
Iwanowicz, D. D. et al. Metabarcoding of fecal samples to determine herbivore diets: A case study of the endangered Pacific pocket mouse. PLoS ONE 11, e0165366 (2016).
Berry, T. E. et al. DNA metabarcoding for diet analysis and biodiversity: A case study using the endangered Australian sea lion (Neophoca cinerea). Ecol. Evol. 7, 5435–5453 (2017).
Ford, M. J. et al. Estimation of a killer whale (Orcinus orca) population’s diet using sequencing analysis of DNA from feces. PLoS ONE 11, e0144956 (2016).
Ando, H. et al. Diet analysis by next-generation sequencing indicates the frequent consumption of introduced plants by the critically endangered red-headed wood pigeon (Columba janthina nitens) in oceanic island habitats. Ecol. Evol. 3, 4057–4069 (2013).
Kim, E.-K. Behavioral ecology, habitat evaluation and genetic characteristics of water deer (Hydropotes inermis) in Korea. Ph.D. thesis. Kangwon National University (2011).
Park, J.-E., Kim, B.-J. & Lee, S.-D. A study of potential of diet analysis in the Korean water deer (Hydropotes inermis argyropus) using polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE). Korean J. Environ. Ecol. 24, 318–324 (2010).
Hollingsworth, P. M. Refining the DNA barcode for land plants. Proc. Natl. Acad. Sci. USA 108, 19451–19452 (2011).
Li, D.-Z. et al. Comparative analysis of a large dataset indicates that internal transcribed spacer (ITS) should be incorporated into the core barcode for seed plants. Proc. Natl. Acad. Sci. USA 108, 19641–19646 (2011).
Park, E. & Nam, M. Changes in land cover and the cultivation area of ginseng in the Civilian Control Zone -Paju City and Yeoncheon County-. Korean J. Environ. Ecol. 27, 507–515 (2013).
Cheng, T. et al. Barcoding the kingdom Plantae: new PCR primers for ITS regions of plants with improved universality and specificity. Mol. Ecol. Resour. 16, 138–149 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Ankenbrand, M. J., Keller, A., Wolf, M., Schultz, J. & Förster, F. ITS2 database V: Twice as much. Mol. Biol. Evol. 32, 3030–3032 (2015).
Sickel, W. et al. Increased efficiency in identifying mixed pollen samples by meta-barcoding with a dual-indexing approach. BMC Ecol. 15, 20 (2015).
Edgar, R. C. Accuracy of taxonomy prediction for 16S rRNA and fungal ITS sequences. PeerJ 6, e4652 (2018).
Oksanen, J. et al. vegan: Community ecology package v 2.5–7 (R Foundation, Vienna, Austria, 2020).
Hsieh, T., Ma, K. & Chao, A. iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol. Evol. 7, 1451–1456 (2016).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
De Cáceres, M. & Legendre, P. Associations between species and groups of sites: indices and statistical inference. Ecology 90, 3566–3574 (2009).
Yan, L. ggvenn: Draw venn diagram by 'ggplot2' v. 0.1.8 (R Foundation, Vienna, Austria, 2021).
Choi, D.-Y. et al. Flora of province Gyonggi-do. Bull. Seoul Nat’l Univ. Arbor. 21, 25–76 (2001).
Ko, S. & Shin, Y. Flora of middle part in Gyeonggi Province. Korean J. Plant Res. 22, 49–70 (2009).
Lee, S.-K., Ryu, Y. & Lee, E. J. Endozoochorous seed dispersal by Korean water deer (Hydropotes inermis argyropus) in Taehwa Research Forest, South Korea. Glob. Ecol. Conserv. 40, e02325 (2022).
Kim, K.-H. & Kang, S.-H. Flora of western civilian control zone (CCZ) in Korea. Korean J. Plant Res. 32, 565–588 (2019).
Gyeonggi Tourism Organization. Pyeonghwa-Nuri Trail ecological resource survey. (Paju City, Gyeonggi Province, Korea, 2018).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis 2nd edn. (Springer, New York, 2016).
R Core Team. R: A language and environment for statistical computing (R Foundation, Vienna, Austria, 2020).
Pertoldi, C. et al. Comparing DNA metabarcoding with faecal analysis for diet determination of the Eurasian otter (Lutra lutra) in Vejlerne. Denmark. Mammal. Res. 66, 115–122 (2021).
Lee, B. Morphological, ecological and DNA taxonomic characteristics of Chinese water deer (Hydropotes inermis Swinhoe). Ph.D. thesis. Chungbuk National University (2003).
Wilmshurst, J. F., Fryxell, J. M. & Hudsonb, R. J. Forage quality and patch choice by wapiti (Cervus elaphus). Behav. Ecol. 6, 209–217 (1995).
Langvatn, R. & Hanley, T. A. Feeding-patch choice by red deer in relation to foraging efficiency. Oecologia 95, 164–170 (1993).
Gray, P. B. & Servello, F. A. Energy intake relationships for white-tailed deer on winter browse diets. J. Wildl. Manag. 59, 147–152 (1995).
Brown, D. T. & Doucet, G. J. Temporal changes in winter diet selection by white-tailed deer in a northern deer yard. J. Wildl. Manag. 55, 361–376 (1991).
Takahashi, H. & Kaji, K. Fallen leaves and unpalatable plants as alternative foods for sika deer under food limitation. Ecol. Res. 16, 257–262 (2001).
Bee, J. N. et al. Spatio-temporal feeding selection of red deer in a mountainous landscape. Austral Ecol. 35, 752–764 (2010).
Gebert, C. & Verheyden-Tixier, H. Variations of diet composition of red deer (Cervus elaphus L.) in Europe. Mammal. Rev. 31, 189–201 (2001).
Cornelis, J., Casaer, J. & Hermy, M. Impact of season, habitat and research techniques on diet composition of roe deer (Capreolus capreolus): a review. J. Zool. 248, 195–207 (1999).
Kim, B. J. & Lee, S.-D. Home range study of the Korean water deer (Hydropotes inermis agyropus) using radio and GPS tracking in South Korea: Comparison of daily and seasonal habitat use pattern. J. Ecol. Field Biol. 34, 365–370 (2011).
Beier, P. Sex differences in quality of white-tailed deer diets. J. Mammal. 68, 323–329 (1987).
Staines, B. W., Crisp, J. M. & Parish, T. Differences in the quality of food eaten by red deer (Cervus elaphus) stags and hinds in winter. J. Appl. Ecol. 19, 65–77 (1982).
Koga, T. & Ono, Y. Sexual differences in foraging behavior of sika deer, Cervus nippon. J. Mammal. 75, 129–135 (1994).
Han, S.-H., Lee, S.-S., Cho, I.-C., Oh, M.-Y. & Oh, H.-S. Species identification and sex determination of Korean water deer (Hydropotes inermis argyropus) by duplex PCR. J. Appl. Anim. Res. 35, 61–66 (2009).
You, Z. et al. Seasonal variations in the composition and diversity of gut microbiota in white-lipped deer (Cervus albirostris). PeerJ 10, e13753 (2022).
Zhao, W. et al. Metagenomics analysis of the gut microbiome in healthy and bacterial pneumonia forest musk deer. Gene Genom. 43, 43–53 (2021).
Amato, K. R. et al. Gut microbiome, diet, and conservation of endangered langurs in Sri Lanka. Biotropica 52, 981–990 (2020).
Stumpf, R. M. et al. Microbiomes, metagenomics, and primate conservation: New strategies, tools, and applications. Biol. Conserv. 199, 56–66 (2016).
Redford, K. H., Segre, J. A., Salafsky, N., del Rio, C. M. & McAloose, D. Conservation and the microbiome. Conserv. Biol. 26, 195–197 (2012).
Deagle, B. E. et al. Counting with DNA in metabarcoding studies: How should we convert sequence reads to dietary data?. Mol. Ecol. 28, 391–406 (2019).
Corse, E. et al. A from-benchtop-to-desktop workflow for validating HTS data and for taxonomic identification in diet metabarcoding studies. Mol. Ecol. Resour. 17, e146–e159 (2017).
Alberdi, A. et al. Promises and pitfalls of using high-throughput sequencing for diet analysis. Mol. Ecol. Resour. 19, 327–348 (2019).
Nakahara, F. et al. The applicability of DNA barcoding for dietary analysis of sika deer. DNA Barcodes 3, 200–206 (2015).
Thomas, A. C., Jarman, S. N., Haman, K. H., Trites, A. W. & Deagle, B. E. Improving accuracy of DNA diet estimates using food tissue control materials and an evaluation of proxies for digestion bias. Mol. Ecol. 23, 3706–3718 (2014).
Deagle, B. E., Eveson, J. P. & Jarman, S. N. Quantification of damage in DNA recovered from highly degraded samples–a case study on DNA in faeces. Front. Zool. 3, 11 (2006).
Coissac, E., Riaz, T. & Puillandre, N. Bioinformatic challenges for DNA metabarcoding of plants and animals. Mol. Ecol. 21, 1834–1847 (2012).
Estes, J. A. et al. Trophic downgrading of planet Earth. Science 333, 301–306 (2011).
Clare, E. L. Molecular detection of trophic interactions: emerging trends, distinct advantages, significant considerations and conservation applications. Evol. Appl. 7, 1144–1157 (2014).
Ramirez, R., Quintanilla, J. & Aranda, J. White-tailed deer food habits in northeastern Mexico. Small Rumin. Res. 25, 141–146 (1997).
Anouk Simard, M., Côté, S. D., Weladji, R. B. & Huot, J. Feedback effects of chronic browsing on life-history traits of a large herbivore. J. Anim. Ecol. 77, 678–686 (2008).
Putman, R. J. & Staines, B. W. Supplementary winter feeding of wild red deer Cervus elaphus in Europe and North America: justifications, feeding practice and effectiveness. Mammal Rev. 34, 285–306 (2004).
Milner, J. M., Van Beest, F. M., Schmidt, K. T., Brook, R. K. & Storaas, T. To feed or not to feed? Evidence of the intended and unintended effects of feeding wild ungulates. J. Wildl. Manag. 78, 1322–1334 (2014).
Carpio, A. J., Apollonio, M. & Acevedo, P. Wild ungulate overabundance in Europe: contexts, causes, monitoring and management recommendations. Mammal Rev. 51, 95–108 (2021).
Cappa, F., Lombardini, M. & Meriggi, A. Influence of seasonality, environmental and anthropic factors on crop damage by wild boar Sus scrofa. Folia Zool. 68, 261–268 (2019).
Funding
This research was partly supported by Seoul National University Research Grant in 2018 (NY).
Author information
Authors and Affiliations
Contributions
S.K.L., E.J.L., and N.Y. conceived the research. S.K.L. and N.Y. designed the research. S.K.L. collected the samples. CW performed the laboratory analysis and bioinformatics. S.K.L. analyzed the data. S.K.L. and N.Y. wrote the first draft of the manuscript, and C.W. and E.J.L. commented on and edited the manuscript. All authors read, revised, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, SK., Woo, C., Lee, E.J. et al. Using high-throughput sequencing to investigate the dietary composition of the Korean water deer (Hydropotes inermis argyropus): a spatiotemporal comparison. Sci Rep 12, 22271 (2022). https://doi.org/10.1038/s41598-022-26862-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26862-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.