Abstract
Malaria, caused by Plasmodium parasites, is a major global health challenge. Whole genome sequencing (WGS) of Plasmodium falciparum and Plasmodium vivax genomes is providing insights into parasite genetic diversity, transmission patterns, and can inform decision making for clinical and surveillance purposes. Advances in sequencing technologies are helping to generate timely and big genomic datasets, with the prospect of applying Artificial Intelligence analytical techniques (e.g., machine learning) to support programmatic malaria control and elimination. Here, we assess the potential of applying deep learning convolutional neural network approaches to predict the geographic origin of infections (continents, countries, GPS locations) using WGS data of P. falciparum (n = 5957; 27 countries) and P. vivax (n = 659; 13 countries) isolates. Using identified high-quality genome-wide single nucleotide polymorphisms (SNPs) (P. falciparum: 750 k, P. vivax: 588 k), an analysis of population structure and ancestry revealed clustering at the country-level. When predicting locations for both species, classification (compared to regression) methods had the lowest distance errors, and > 90% accuracy at a country level. Our work demonstrates the utility of machine learning approaches for geo-classification of malaria parasites. With timelier WGS data generation across more malaria-affected regions, the performance of machine learning approaches for geo-classification will improve, thereby supporting disease control activities.
Similar content being viewed by others
Introduction
Malaria, caused by Plasmodium parasites and transmitted by Anopheles mosquitoes, remains a pressing global health problem, with a mortality and morbidity burden heavily concentrated among children less than five years old. The morbidity and mortality impacts of Plasmodium falciparum malaria are predominantly concentrated in Sub-Saharan Africa, whereas the burdens of Plasmodium vivax are most heavily felt in Asia and South America1. The complex co-evolutionary history between Plasmodium parasites, humans, and Anopheles mosquitoes is contained within the genome of each organism, and genomic tools and data are of key importance for understanding the fundamental genetic underpinning of malaria, its geo-spatial distribution and control strategies to eliminate it. There is a rapidly growing number of P. falciparum and P. vivax isolate DNA that have undergone whole genome sequencing (WGS), with continued advances in genomic technologies likely to accelerate the timely generation of datasets from clinical and surveillance blood samples to inform disease epidemiology and control.
The rich information contained in WGS data can be used to infer transmission patterns, detect drug resistance, and support wider malaria control initiatives and elimination strategies2,3. WGS data in combination with population genomic methods can detect selective sweeps associated with drug resistance and infer the geographic origin of infections, including if infections are found to be imported or drug resistant and whether treatment should be adapted accordingly. It is known that malaria parasites have a population structure primarily based on geography4,5. Several informative molecular barcodes for speciation and geography have been developed2,3, but typically these barcodes have not used the whole genome due to the high-dimensionality of the data and the associated computational cost3. However, machine learning (a subfield of Artificial Intelligence) with its ability to incorporate and analyse very large and high-dimensional datasets in an efficient manner, seems potentially well suited for geo-predicting using WGS data. Machine learning can be applied for classification, which concerns predicting a label (e.g., country, continental region), and regression, which involves predicting a quantity (e.g., longitude or latitude).
Machine learning has been applied effectively across a variety of problems in malaria research, including the detection of evolutionary selection associated with drug resistance6,7, the classification and detection of parasites in red blood cells8,9,10,11, and antimalarial drug discovery12. Deep learning is a subset of machine learning where algorithms aim to extract and learn series of hierarchical representations, often leveraging large amounts of data. The application of deep learning, and especially neural networks, has been explored within population genetics13,14, including for other pathogens15,16. Pioneering work has also shown that machine learning, including deep learning convolutional neural networks (CNNs), can be used to predict geographic locations from human, mosquito and P. falciparum genetic variation17, building on methods and the use of large genotyping chips or WGS for population structure assessment18,19. Here, we aim to further expand on the application of geo-prediction for malaria parasites by using a very large dataset of isolates sourced globally, (P. falciparum, n = 5957, 27 countries; P. vivax, n = 659, 13 countries) across 11 regions (South East Asia (SEA), Southern SEA (SSEA), South Asia, South America, West Africa, Central Africa, South Central Africa, East Africa, Horn of Africa, Southern Africa, Oceania). We explore the potential of both regular machine learning approaches that aim to learn representations from sequence and geographical data, as well as deep learning approaches that aim to learn and extract layers of hierarchical representations of SNP combinations linked to geography. We compare four commonly applied approaches, including classification methods that predict locations and subsequently interpolate to specific coordinates, as well as compare the performance across geographies (countries) both including the observations within those and excluding them from the training sets used to develop the models.
Materials and methods
Processing of raw sequencing data
Publicly available raw Illumina (> 150 bp paired end) sequence data from previously published studies of P. falciparum and P. vivax was downloaded from the ENA repository (see S1 Table and S2 Table for accession numbers), and accompanied by meta-data including locations of sampling (see S1 Table and S2 Table for latitude and longitude coordinates). The data included public raw sequence and GPS data from MalariaGEN projects (www.malariagen.net). Raw WGS data for P. falciparum (n = 5957) and P. vivax (n = 659) were aligned with the Pf3D7 (v3) and PvP01 (v1) reference genomes, respectively, using bwa-mem software (v0.7.12) using default parameter settings (e.g., concerning mismatch and sequence read clipping penalties; see http://bio-bwa.sourceforge.net/bwa.shtml). The samtools (v1.9) functions fixmate and markdup were applied to the resulting BAM files to call a set of potential variants20. For variant quality control, calibration assessments were performed using the GATK’s BaseRecalibrator and ApplyBQSR functions, benchmarking off known high quality variants from genetic crosses for P. falciparum5,21 and previously curated datasets for P. vivax20. A revised set of SNPs and insertions/deletions (indels) was called with GATK’s HaplotypeCaller (version 4.1.4.1) using the option -ERC GVCF5,22. Variants were then assigned a quality score using GATK’s Variant Quality Score Recalibration (VQSR), and those with a VQSLOD score < 0, representing variants more likely to be false than true, were filtered out7,22. Additionally, SNPs were removed if they had more than 10% missing alleles7,22.The resulting dataset comprised of parasite genomes of P. falciparum (5,957 isolates, 750 k SNPs) and of P. vivax (659 isolates, 588 k SNPs). The population structure was assessed using a principal component analysis (PCA) of between isolate SNP differences. In parallel, ADMIXTURE analysis23 was performed to understand the composition of ancestral groups across geography, where the optimal number of groups (K) was established using cross validation with values ranging between 1 and 20. This cross validation analysis led to 10 ancestral groups for both P. falciparum and P. vivax (K = 10).
Statistical models and performance
Using machine learning (ML) and deep learning (DL) statistical models, the goal was to use SNPs to predict geographical source at a location (GPS), country, and regional resolution. We applied two standard models for classification at a country and region level: (1) penalized multinomial logistic regression classifier (LOG-C; ML); (2) CNN (CNN-C; DL). Subsequently, we used the predictive probabilities placed on different locations to perform a weighted interpolation between these locations and make predictions at the GPS coordinate level.
In particular, the final prediction location (longitude and latitude) was determined by a weighted average of classifier predictions, where weights are the probabilities placed by the model on each location.
We also applied two regression models for GPS coordinate prediction: (iii) penalised linear regression model (LIN-R; ML); (iv) CNN (CNN-R; DL). The LOG-C and LIN-R models were tuned on the regularization strength C for the L1 penalty (LASSO) and implemented in the sklearn Python package (https://scikit-learn.org). The penalty parameters were tuned using cross-validation (see below, S3 Table). The deep learning CNN architecture was implemented using the Keras library (version 2.2.4)24 in Python. Our CNN models had an architecture with a soft-max prediction layer and regularization through dropout25 to prevent overfitting and support transferability. The main model had one convolutional layer with 4 filters, with respective filter size of (40, 9) followed by two drop-out and dense layers with ReLu activation (similar to17), and applied the Stochastic Gradient Descent algorithm for optimisation. We trained and validated the models for 1000 epochs. The parameterisation of the models is summarised (S3 Table). We created a stratified three-fold split in the dataset (80% training, 10% validation, 10% test) for all models, and used the validation dataset to cross-validate parameters (S3 Table). The LOG-C and LIN-R models were cross-validated (stratified, four-fold) on the regularization strength C for the L1 penalty. The reported scores (accuracy, mean weighted distance error) were calculated by making predictions on the hold-out test set (see S3 Table for the final parameter set). In addition, we conducted a “leave-one-geography-out”, where each single geography in the training dataset was omitted in turn, with the model trained on the remaining geographies, to understand generalizability towards previously unseen locations26.
Classification accuracy was determined after assigning predicted latitude and longitude pairs to individual countries. For the classification models, a mean (weighted) distance error was calculated using the Haversine method to allow for (angular) distance calculations along a sphere, based on the difference of the actual and estimated location. The latter was determined by a weighted average of classifier predictions, where weights are the probabilities placed by the model on each location. The accuracy was calculated based on the labels of the prediction versus the test data. In particular, the baseline accuracy using a naive prediction based on the most common country would be 18.8% for P. falciparum (Cambodia) and 24.3% for P. vivax (Thailand). For the regression models, the error was calculated using the Haversine method based on the difference between the predicted and actual latitude and longitude using angular distance.
Results
Malaria isolate sequence data and population structure
Raw WGS data with accompanying geographic origin information was available in the public domain for P. falciparum (n = 5957, 27 countries) and P. vivax (n = 659, 13 countries) (Table 1), which represent the global distributions for each parasite. Most P. falciparum isolates were sourced from SEA (2,648, 44.5%) followed by West Africa (2,042, 34.3%) and East Africa (451, 7.6%). Whilst, for P. vivax, most isolates were sourced from SEA (282, 42.9%) followed by South America (220, 33.4%) and SSEA (48) (Table 1). By analysing each species separately, high quality genome-wide SNPs were identified across the isolates (P. falciparum 750 k SNPs, P. vivax 588 k SNPs). Most SNPs have low minor allele frequencies (SNPs with MAF < 1%: P. falciparum 94.6%, P. vivax 77.6%) (S1 Figure). Most SNPs were in genic regions (P. falciparum 76.5%, P. vivax 54.3%), with a high proportion of non-synonymous (NS) amino acid changes (P. falciparum 63.0%, P. vivax 42.5%). The genetic diversity amongst P. falciparum isolates was relatively homogeneous across the 27 countries (SNP π: median 0.037, range 0.027–0.053), and lower in magnitude than P. vivax, whose data was sourced from 13 countries (SNP π: median 0.056, range 0.037–0.066) (Table 1).
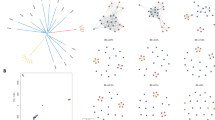
Unsupervised clustering methods were applied to the genome-wide SNPs of each species to reveal the extent of their population structure and linked (pseudo-)ancestral patterns. Principal component analysis (PCA) of P. falciparum and P. vivax isolates revealed the expected separation by continent, and clear evidence of population structure at both the regional and country level (Fig. 1). An analysis of population structure and ancestry using ADMIXTURE software23 determined the number of ancestral groups (P. falciparum K = 10, P. vivax K = 10), and their relative abundance for each isolate was estimated (Fig. 2). For P. falciparum, there were dominant ancestral groups across region and continent (Africa 4, SEA 4, Oceania 1, South America 1), with some evidence of mixture of ancestries (e.g., SEA isolates with 3 ancestral populations), but a general consistency within country. For P. vivax, the numbers of dominant ancestral groups by region differed from P. falciparum (South America 4, SEA 2, SSEA 2, East Africa 1, South Asia 1), due to sampling and Plasmodium species endemicity differences, such as the near absence of P. vivax in Africa. Overall, there was more homogeneity of ancestral groups within P. vivax isolates, with some groups broadly linked to neighbouring countries (comparison with Fig. 1). These analyses confirmed that spatial-genomic clustering and classification is possible using WGS data.
Application of geo-classification models
For P. falciparum, the predictive performance of the classification methods (LOG-C, CNN-C) was stronger than for the regression models (LIN-R, CNN-R) in regional (Table 2) and country-wide (Table 3) analyses (mean distance error (km): LIN-R 470, LOG-C 93, CNN-R 245, CNN-C 77). For locations included in the training dataset, the performance of the classification models was close to 100% at the regional level, and close to 90% at the country level (S4 Table, S5 Table). The poorest performance of the models was for African populations, for example, the mean distance error for CNN-C was high in West African (267 km) and East African countries (117 km, especially Kenya and Uganda), as well as Malawi (530 km) (Table 3), compared to other regions. This observation is consistent with the complex ancestries in African populations (Fig. 2), as well as another deep learning analysis17. As expected, where we predicted countries absent in data used by the training models, the distance errors (km) were at least ~ five-fold larger (LIN-R 2246, LOG-C 1848, CNN-R 1983, CNN-C 1540), with the poorest predictions for Peru (Table 4). The best performing model in this setting was the CNN-C classifier (Fig. 3).
Maps with predicted vs. actual locations for the best predictive models. Blue points are the actual locations in the dataset, red points are the predicted locations (where different to actual), with red lines link the actual and the predicted locations. CNN-C deep learning Convolutional Neural Network classifier. LOG-C penalised multinomial logistic regression classifier.
For P. vivax, the predictive performance of the classification methods (LOG-C, CNN-C) was also superior compared to regression models (LIN-R, CNN-R) across regional (Table 2) and country-wide (Table 3) analyses (mean distance error (km): LIN-R 890, LOG-C 33, CNN-R 819, CNN-C 36) (Table 3). For locations included in the training dataset, the performance of the classification models was close to 100% at both the regional and country level, with the poorest performance in neighbouring China and Myanmar (S4 Table, S5 Table). The (mean) distance error for the countries not used in the development of the model is distinctively larger (km: LIN-R 1481, LOG-C 2508, CNN-R 2512, CNN-C 2405), with the poorest predictions for Ethiopia and Peru (Table 4). The best performing model in this setting was a LIN-R regression (Fig. 3).
Discussion
WGS data of Plasmodium parasites can detect imported infections, drug resistance, and transmission patterns, thereby assisting decision making in clinical and malaria control settings. With the implementation of WGS gaining traction across health systems, there is an opportunity to implement statistical learning methodologies to assist surveillance activities. A clear use-case includes the determination of the geographical origin of isolates, building on insights from previous work which shows that genomic data can be used to cluster parasites by geography2,3,4,5. Our work reveals that machine learning approaches, particularly those focusing on classification (e.g., deep learning CNNs), have the potential to accurately predict geographic locations at a GPS and country-level resolution. As expected, the performance was much stronger for isolates of which the geographic origin was already represented at the country level in the dataset, demonstrating the need for WGS to be implemented more widely to fill country gaps in genetic diversity. The weakest predictions were for P. falciparum in West and East Africa, where common ancestries, mixed infections, movement of people, drug resistance and malaria endemicities can complicate genetic diversity analysis. The distance errors are similar to a previous machine learning analysis of P. falciparum (median < 20 km), which implemented a single deep learning approach on a smaller dataset17. Our CNN for classification approach appeared to perform well across parasite species, was implemented with measures to minimise the effects of over-fitting, and its performance is likely to improve with greater isolate sampling and WGS data.
Whilst we have implemented a limited set of machine learning methods, there is scope to test alternative approaches (e.g., gradient boosted trees, support vector machines)16 or further optimise our model parametrisations (beyond the default settings) to improve performance. For example, while L1-penalized regression approaches are generally quite competitive, stability selection on top of the LASSO leads generally to improvements27. Moreover, the resulting model is white box and leads to a set of interpretable SNPs. CNNs are the most utilised deep learning network type, and known to outperform alternative approaches28. However, one limitation of CNN models is their “black box” nature, with a complex architecture consisting of several layers, and in our context (and others17) making it difficult to establish which (combinations of) SNPs are informative for the geographical profiling. Other studies have used population genomic approaches to determine informative SNPs, with a focus on applying genotyping assays or amplicon sequencing for resource poor settings2,3. We provide computer code to implement the models, to assist future assessments in simulation or empirical studies. Future work should focus on the development of an online “geo-locator” tool that reveals a prediction of location, which can be assessed for its plausibility against the actual position, if known, and feedback into the model building and learning process. Such a framework could also be extended to integrate explicit drug resistance markers29, as well as genomic data for malaria vectors17, and use sequences generated on portable and field deployable sequencing platforms (e.g., Oxford Nanopore Technology MinION). Such tools would be of immediate value to malaria control programs in endemic countries, including those that are implementing elimination activities who wish to differentiate between locally acquired or imported infections. It would also assist those countries with low malaria burden, including through the detection of imported parasites that could threaten malaria elimination targets.
In summary, our study has demonstrated that machine learning methods can play an informative role in determining the geographic origin of WGS isolates, thereby providing important insights for both control and surveillance activities. Further, such approaches will be scalable when WGS becomes routine and cost effective, resulting in a setting with increasingly “big data” being available for decision making. The utility of this “learning” system will improve with time, as underlying methodologies and model performances improve with more data becoming available, and they are implemented within informatic tools to assist surveillance and clinical decision making. This utility underscores the benefit of making sequencing data and linked geographical information publicly available to global databases in a more-timely fashion to understand infection dynamics, the advantages of which have also been demonstrated by the COVID-19 crisis.
Conclusion
Advances in sequencing technologies are making real time genomics-informed surveillance and clinical management a reality. With the resulting big genomic datasets, our study has shown that machine learning methods, a subset of Artificial Intelligence, can accurately predict the geographical source of malaria parasites from sequence data. With greater geographical coverage and informatics infrastructure, such approaches will improve in performance and assist malaria control and elimination activities.
Data availability
The raw WGS data is available from the European Nucleotide Archive (ENA) (see S1 Table and S2 Table for project accession numbers). Computing code and machine learning models are available from https://github.com/WDee/GeoComparison.
References
World Health Organization. World Malaria Report (2020).
Preston, M. D. et al. A barcode of organellar genome polymorphisms identifies the geographic origin of Plasmodium falciparum strains. Nat. Commun. 5, 1–7 (2014).
DiezBenavente, E. et al. A molecular barcode to inform the geographical origin and transmission dynamics of Plasmodium vivax malaria. PloS Genet. 16, e1008576 (2020).
Diez Benavente, E. et al. Distinctive genetic structure and selection patterns in Plasmodium vivax from South Asia and East Africa. Nat. Commun. 12, 1–11 (2021).
Samad, H. et al. Imputation-based population genetics analysis of plasmodium falciparum malaria parasites. PLOS Genet. 11, e1005131 (2015).
Pybus, M. et al. Hierarchical boosting: A machine-learning framework to detect and classify hard selective sweeps in human populations. Bioinformatics 31, 493 (2015).
Deelder, W. et al. Using deep learning to identify recent positive selection in malaria parasite sequence data. Malar. J. 20, 1–9 (2021).
Quan, Q., Wang, J. & Liu, L. An effective convolutional neural network for classifying red blood cells in malaria diseases. Interdiscip. Sci. Comput. Life Sci. 12, 217–225 (2020).
Liang, Z. et al. CNN-based image analysis for malaria diagnosis. In: Proc. - 2016 IEEE Int. Conf. Bioinforma. Biomed. BIBM 2016 493–496 (2017). https://doi.org/10.1109/BIBM.2016.7822567.
Poostchi, M., Silamut, K., Maude, R. J., Jaeger, S. & Thoma, G. Image analysis and machine learning for detecting malaria. Transl. Res. 194, 36–55 (2018).
Fuhad, K. M. F. et al. Deep learning based automatic malaria parasite detection from blood smear and its smartphone based application. Diagnostics 10, 329 (2020).
Neves, B. J. et al. Deep Learning-driven research for drug discovery: Tackling malaria. PLoS Comput. Biol. 16, e1007025 (2020).
Flagel, L., Brandvain, Y. & Schrider, D. R. The unreasonable effectiveness of convolutional neural networks in population genetic inference. Mol. Biol. Evol. 36, 220–238 (2019).
Sanchez, T., Cury, J., Charpiat, G. & Jay, F. Deep learning for population size history inference: design, comparison and combination with approximate Bayesian computation. bioRxiv https://doi.org/10.1101/2020.01.20.910539 (2020).
Deelder, W. et al. Machine learning predicts accurately mycobacterium tuberculosis drug resistance from whole genome sequencing data. Front. Genet. 10, 922 (2019).
Libiseller-Egger, J., Phelan, J., Campino, S., Mohareb, F. & Clark, T. G. Robust detection of point mutations involved in multidrug-resistant mycobacterium tuberculosis in the presence of co-occurrent resistance markers. PLoS Comput. Biol. 16, e1008518 (2020).
Battey, C. J., Ralph, P. L. & Kern, A. D. Predicting geographic location from genetic variation with deep neural networks. Elife 9, 1–22 (2020).
Guillot, G., Jónsson, H., Hinge, A., Manchih, N. & Orlando, L. Accurate continuous geographic assignment from low- to high-density SNP data. Bioinformatics 32, 1106–1108 (2016).
Bhaskar, A., Javanmard, A., Courtade, T. A., Tse, D. & Valencia, A. Novel probabilistic models of spatial genetic ancestry with applications to stratification correction in genome-wide association studies. Bioinformatics 33, 879–885 (2017).
Li, H. Improving SNP discovery by base alignment quality. Bioinformatics 27, 1157–1158 (2011).
Miles, A. et al. Indels, structural variation, and recombination drive genomic diversity in Plasmodium falciparum. Genome Res. 26, 1288–1299 (2016).
Benavente, E. D. et al. Genomic variation in Plasmodium vivax malaria reveals regions under selective pressure. PLoS ONE 12, e0177134 (2017).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Chollet, F., & others. Keras. GitHub. Retrieved from https://github.com/fchollet/keras (2015).
Srivastava, N., Hinton, G., Krizhevsky, A. & Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Mordelet, F. & Vert, J. P. ProDiGe: Prioritization of disease genes with multitask machine learning from positive and unlabeled examples. BMC Bioinform. 12, 1–15 (2011).
Mahé, P. & Tournoud, M. Predicting bacterial resistance from whole-genome sequences using k-mers and stability selection. BMC Bioinform. 19, 1–11 (2018).
Alzubaidi, L. et al. Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J. Big Data 8, 1–74 (2021).
Turkiewicz, A. et al. Genetic diversity of the Plasmodium falciparum GTP-cyclohydrolase 1, dihydrofolate reductase and dihydropteroate synthetase genes reveals new insights into sulfadoxine-pyrimethamine antimalarial drug resistance. PLoS Genet. 16, e1009268 (2020).
Acknowledgements
TGC was funded by Medical Research Council UK (Grant no. MR/M01360X/1, MR/N010469/1, MR/R025576/1, MR/R020973/1 and MR/X005895/1) grants. SC was funded by BloomsburySET and Medical Research Council UK grants (MR/M01360X/1, MR/R025576/1, MR/R020973/1 and MR/X005895/1). We thank Aleksei Ponomarev for providing support on Python coding.
Author information
Authors and Affiliations
Contributions
W.D., S.C., L.P., and T.G.C. conceived and designed the study. E.M. and J.E.P. performed the bioinformatic processing of the raw sequencing data. W.D. and E.M. performed the population genetic and statistical analysis, under the supervision of S.C., L.P. and T.G.C. W.D. wrote the first draft of the manuscript. All authors commented on and edited the manuscript and approved the final version. W.D. and T.G.C. compiled the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Deelder, W., Manko, E., Phelan, J.E. et al. Geographical classification of malaria parasites through applying machine learning to whole genome sequence data. Sci Rep 12, 21150 (2022). https://doi.org/10.1038/s41598-022-25568-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25568-6
This article is cited by
-
Risk assessment of imported malaria in China: a machine learning perspective
BMC Public Health (2024)
-
Rapid profiling of Plasmodium parasites from genome sequences to assist malaria control
Genome Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.