Abstract
The prion-like transsynaptic propagation of misfolded tau along the brain’s connectome has previously been modeled using connectome-based network diffusion models. In addition to the connectome, interactions between the general neurological “milieu” in the neurodegenerative brain and proteinopathic species can also contribute to pathology propagation. Such a molecular nexopathy framework posits that the distinct characteristics of neurodegenerative disorders stem from interactions between the network and surrounding molecular players. However, the effects of these modulators remain unquantified. Here, we present Nexopathy in silico (“Nexis”), a quantitative model of tau progression augmenting earlier models by including parameters of pathology propagation defined by the molecular modulators of connectome-based spread. Our Nexis:microglia model provides the first quantitative characterization of this effect on the whole brain by expanding previous models of neuropathology progression by incorporating microglial influence. We show that Trem2, but not microglial homeostasis genes, significantly improved the model’s predictive power. Trem2 appears to reduce tau accumulation rate while increasing its interregional spread from the hippocampal seed area, causing higher tau burden in the striatum, pallidum, and contralateral hippocampus. Nexis provides an improved understanding and quantification of microglial contribution to tau propagation and can be flexibly modified to include other modulators of progressive neurodegeneration.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder that exhibits extracellular amyloid-\(\beta\) plaques and intracellular aggregates of hyperphosphorylated tau as its main neuropathological hallmarks. The regional location and density of tau tangles correlate more with the degree of clinical symptoms than those of amyloid-\(\beta\) plaques1,2. The stronger association between tau and regional atrophy, hypometabolism, and cognitive decline in AD3,4 makes it imperative to shift the long-standing focus from amyloid-\(\beta\) to tau and to further investigate modulators of tau accumulation and spread.
Although tau mainly spreads via the brain’s anatomic connectome, the complete characterization of tauopathies should be based on the notion of Molecular Nexopathy5, which states that divergent spatiotemporal patterns of different tauopathies may result from biochemical differences between tau species and the neural cells in the brain that interact with tau as well as the connectome-based mechanism of transsynaptic transmission5. To unpack this, even though the key mediatory role in tau spread is played by the connectome, there are many other interactions between cells, circuits, proteopathic species, and the general neurological milieu in the neurodegenerative brain that remains outside of the connectome-centric view of these diseases. Evidence for this notion has become stronger over years suggesting the existence of important condition-altering differences in pathological tau species between degenerative conditions. For instance, tau displays differential 3R and 4R isoforms6 and hyperphosphorylation sites7 in different tauopathies. Gross tangle structure also differs across diseases, with AD tau forming neurofibrillary tangles and pre-helical formations, Pick bodies in Pick’s disease, and progressive supranuclear palsy, corticobasal degeneration, and frontotemporal dementia presenting more abundant tangles8. Indeed, pathological tau transits with a different mechanism in a disorder-dependent manner5,9.
Tau propagates in a stereotypical pattern in AD, starting from the entorhinal cortex to the hippocampus followed by the neocortex as described by Braak staging10. Experiments involving the injection of synthetic tau fibrils in the mouse brain have demonstrated that tau can propagate from the injection site to distant but connected regions11. Tauopathy progression is prion-like - templated and self-propagating12, with pre-existing aggregates acting as seeds to induce further misfolding. Since tau primarily migrates along the white matter tracts of the brain, it is reasonable to model its spatiotemporal dynamics in a first-order approximation as a diffusive process between connected brain regions. Prior work by our group has quantified pathology progression in humans and animals using such an approach via a graph-theory based system of differential equations called the Network Diffusion Model (NDM)13,14,15. However, although the NDM framework considers the connectome graph as the central (sole) mediator of pathology spread along a networked representation of the brain, it fails to consider the impact of molecular nexopathic species on tau spread.
Modeling of tau propagation and the hypothesized influence of microglia. Panel (a): Representation of the brain as a network, with each ball indicating the center of mass of a region and the line between any pair of regions indicating a connection between them. Panel (b): Connectivity matrix of unilateral 213 cortical regions of the adult mouse brain. Panel (c): Graphic of the whole brain connectivity network with nodes (i,j,k) representing brain regions and connection strengths (Cij, Cjk) representing mesoscale connectivity. Panel (d): Illustration of Nexis-modeled progression of tau across time with each ball representing a region and ball-size indicating the degree of pathology burden. Panel (e): Per-region baseline microglial density is depicted in blue on the graphic of the sagittal mouse brain. Tau pathology at different time points post tau seeding is depicted in red on the mouse brains. The blue arrow on the glass brain at time point 1 indicates the regional injection site (hippocampus in this case). Panels (a), (c) and (d) are adapted from Mezias et al.15. AMY, amygdala; CRB, cerebellar region; CORT, cortical subplate; HPC, hippocampal formation; HYPO, hypothalamus; ISOCTX, isocortex; MED, medulla; MIDBR, midbrain; OLF, olfactory areas; PAL, pallidum; PONS, pons; STR, striatum; THAL, thalamic areas.
Here, we attempt to capture this nexopathy paradigm using mathematical in silico modeling, which we refer to as Nexopathy in silico or “Nexis”. Nexis models the following biological processes: trans-neuronal transmission along the connectome (modeled using the NDM); prion-like amplification of tau over time; and the nexopathic effects of extra-connectomic molecular players that modulate these processes. We formulate from first principles the process by which neuronal and other cells, genes, and other modulators might interact with misfolded tau during its accumulation and transmission along neural connections. Although these principles are qualitatively well-established from preclinical in vivo and in vitro studies, the unique benefit of formulating them in a mathematical model is that the model then becomes a means for testing interesting hypotheses and becomes a testbed for novel hypothesis generation. The overall design of Nexis and its typical workflow is depicted in Fig. 1a–d.
Important nexopathic effects are wielded by microglia16, the brain’s resident immune cells, and tau dynamics are well known to be influenced by the action of microglia often found in the vicinity of pathological aggregates. Recent studies of postmortem AD brains have revealed that microglia co-localize with tau and activate themselves after internalizing tau aggregates17. Tau-containing neurons can also activate adjacent microglia causing them to form junctions with other neurons as conduits to transfer tau and propagate pathology18. Using mouse models of rapid tau propagation, Asai et al.19 and others have demonstrated that microglia markedly contribute to the clearance, recruitment, and spread of pathology. Microglial activation and neuroinflammation accompany tau tangle formation and importantly, parallel AD clinical symptoms20. Specifically, activated microglia express the triggering receptor expressed on myeloid cells 2 (TREM2). Trem2 is a well-known AD risk-gene21, with loss-of-function mutations (such as R47H and R62H) increasing the susceptibility to late-onset AD22,23 and being related to the rate of cognitive decline24. In vivo, Trem2 over-expression in pathological conditions is related to the recruitment of microglia to pathological aggregates. Although, studies assessing the relationship between Trem2 deletion and tau pathology and neurodegeneration have yielded variable results21, empirical clinical and preclinical data suggest that the modulation of tau propagation by the action of microglia19,25,26 is germane to the study of progressive tauopathies.
Owing to the microglial contributions, quantitative estimates based solely on connectivity and regional pathology levels, e.g. those from our prior NDM13, cannot recapitulate all aspects of pathology spread patterns in AD. Therefore here, we apply our novel Nexis framework to interrogate the modulatory effect of microglia on tauopathy progression, by introducing microglial signatures within the Nexis model (Fig. 1e). We explore microglial influence under two potential modes, whereby microglia alter: (1) the rate of intra-regional tau accumulation and (2) the rate of inter-regional tau spread. These effects are not mutually exclusive but have different biological and mathematical implications. We first apply the non-microglia-influenced Nexis model of tau aggregation and spread (i.e. Nexis:global) characterized solely by tau accumulation/clearance rate in addition to its diffusion along the connectome. This model is further extended into a microglia-based Nexis:microglia model to incorporate their effect on tauopathy progression. We apply these Nexis frameworks to data from mouse models of tauopathy27 using a 426-region mouse connectome28. By utilizing microglial homeostasis genes29 (first principal component of P2ry12, Cx3cr1, Fcrls, Olfml3, Hexb, Siglech, Sox5, and Jun) and an activated microglial signature (Trem2) gene30, we show that the inclusion of microglia in the model quantitatively improves its predictive power, with Trem2 inclusion largely outperforming that of microglial homeostasis genes. We developed an optimization algorithm to infer model parameters. Results confirm and quantify reduction of tau burden (accumulation) by Trem2 and increase in tau transmissibility to regions ipsi- and contralateral to the hippocampal seed area. Given the strain-specific nature of tau pathology, we verified that the Nexis:Trem2 model parameters for datasets DS6 and DS9 also fit the pathology from the corresponding 1:10 dilution studies from Kaufman et al.27. We then explored Apoe, another putative modulator of tau propagation, and found that it does not contribute significantly to tau progression in the current datasets used. Nexis:microglia was also applied to some other microglial genes (Sorl1, Cd33, Clu, Sirpa, and Aif1) regulating phagocytosis and neuroinflammation in AD progression31,32.
Overall, the Nexis:microglia framework provides a more complete model of tau propagation. This study brings together empirical mouse tauopathy models27, the mouse mesoscale connectome28, and gene expression atlas30 to achieve the first quantitative study of microglial involvement in tauopathy propagation developed and validated at the whole-brain level.
Results
Association between spatiotemporal tau pathology and microglial spatial distribution
Correlations of tau pathology spread in sample dataset DS9 (Fig. 2a) with microglial homeostasis genes (Fig. 2b, c) as well as with Trem2 (Fig. 2b, d) were significant, although modest. These data form a solid basis on which to undertake a more detailed and model-based exploration of the effects of microglia on tauopathy progression.
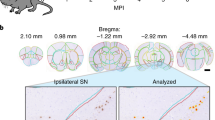
Correlation between spatiotemporal empirical tau pathology at 4, 8, and 12 weeks post hippocampal tau injection and baseline microglial gene spatial distribution for DS9. Panel (a): Empirical tau pathology in DS9 dataset post hippocampal seeding of exogenous tau27. Panel (b): Baseline spatial distribution of first principal component of microglial homeostasis genes and Trem2 from a healthy adult mouse30. Panels (c) and (d): Correlation between spatiotemporal tau pathology progression at 4, 8, and 12 weeks post hippocampal seeding, and baseline expression levels of microglial homeostasis genes as well as Trem2. Sphere sizes in panel a correspond to the degree of tau burden in specific brain regions. HIP, hippocampus; EC, entorhinal cortex; PAL, pallidum; STR, striatum.
Workflow of model implementation and validation
To implement and validate the Nexis models we chose whole-brain analysis approach with both the structural connectome and regional tau defined at the same parcellation scheme consisting of 426 regions (213 each hemisphere), each of which serves as a node of the connectome graph. Fig. 1a–c schematizes the network representation of the mouse brain for which we extract the connectivity matrix C. This connectome is the central (sole) mediator of tau spread in NDM (Fig. 1d). For Nexis, the analytical framework is augmented to account for the influence of tau accumulation/clearance rate (Nexis:global) followed by the influence of microglia (Nexis:microglia) (Fig. 1e). Since homeostasis genes and Trem2 have distinct functional roles and display distinct baseline distributions (Fig. 2b), we chose to model them separately with two models following the Nexis:microglia framework: Nexis:homeostasis and Nexis:Trem2.
We validated the models at the whole-brain level on mouse tauopathy datasets DS4, DS6, and DS9 from Kaufman et al.27. Instead of pre-selecting certain brain regions and their tau levels as is usually done in such studies, we used all regional tau data, wherever available, in a completely unbiased fashion.
Model parameterization depends on tau strain
The Nexis:microglia models include the following parameters: global accumulation/clearance rate of pathology, \(\alpha\); global diffusivity rate constant denoting the diffusion of pathology between regions, \(\beta\); accumulation/clearance of pathology as modulated by microglia, p; and pathology diffusion rate as modulated by microglia, b. Nexis:global is the non-microglial model with \(p = b = 0\). The tau strains of the selected datasets were derived from mutually distinct sources: DS4 mice were injected with human AD-derived tau isolates, whereas DS6 and DS9 mice had tau derived from a P301S mouse and recombinant tau fibrils, respectively27. These pre-characterized datasets were made available by the Kaufman et al. study27 where they demonstrated distinct tau bio-activity marked by varying capacities to form and spread tau aggregates (seeding and spreading potential), widespread induction of pathology, and capacity to induce microgliosis. Figure S1 illustrates the distinct spatiotemporal patterns of tau pathology in DS4, DS6, and DS9 captured in silico.
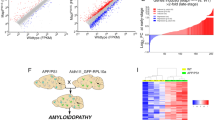
Mean and 95% confidence intervals for the global and microglial parameters as well as model fit for datasets DS4, DS6, and DS9. Panels (a) and (b): 100-iteration 80% resampled bootstrapped values of the Nexis:global and Nexis:Trem2 model parameters. Panel (c): Comparison between model fits of the 80% resampled bootstrapped dataset with the full dataset (magenta dotted line). Global parameters: \(\gamma\) (seed rescale), \(\alpha\) (intra-regional tau accumulation rate), and \(\beta\) (inter-regional tau spread rate). Microglial parameters: p (microglia-mediated intra-regional tau accumulation rate) and b (microglia-mediated inter-regional tau spread rate). Model fit is depicted by \(R^2\).
We implemented a robust strategy for parameter estimation using a 100 iteration bootstrapping approach on 80% resampled data, allowing us to acquire 95% confidence intervals around the parameter means and to assess reproducibility. Table 1 summarizes the inferred parameters and quality-of-fit metrics. The results of parameter estimation using the bootstrapping approach (Fig. 3) include the mean and 95% confidence intervals for the model fit, global, and microglial parameters for Nexis:global and Nexis:Trem2 of DS4, DS6, and DS9. Non-zero p and b values demonstrate microglial influence on tau accumulation and spread above and beyond Nexis:global. When Trem2 microglial effects were included, DS6 and DS9 indicated enhanced clearance (p < 0) but increased inter-regional spread of tau (b > 0). Homeostatic microglial effects, however, led to lower inter-regional spread in DS6 and DS9 (b < 0 for DS6 and DS9; Table 1). DS4 demonstrated increased tau accumulation and spread due to Trem2 compared to homeostatic microglia (Table 1).
Inclusion of microglia improves the predictive power of Nexis
The NDM performed weakly compared to Nexis models for all datasets (Table 1). Nexis:global performed weakly for DS4 (\(R^2\) = 0.17) but well for DS6 and DS9 (\(R^2\) = 0.46 and 0.44, respectively), successfully recapitulating the spatiotemporal profile of empirical tau pathology. This suggests that passive network diffusion is insufficient and additional aggregation processes enhance the model’s predictive power.
The inclusion of microglial gene expression demonstrated higher predictive power (\(R^2\)) compared to Nexis:global. The improvement was statistically significant (per Fisher’s-R-to-Z transform p value) for DS6 and DS9. Trem2 imparted significantly higher predictive power compared to microglial homeostasis genes. It is important to note that across all datasets, AIC and BIC decreased upon the inclusion of microglia, indicating that the better fits achieved by Nexis:microglia were not simply due to the introduction of additional model parameters. Instead, these results reflect reduction in uncertainty in the global parameters, \(\alpha\) and \(\beta\), as indicated by their narrower 95% confidence intervals. The eight chosen microglial homeostasis genes are used in the model as their first principal component. The accumulation and spread parameters for these Nexis:microglia models were either negative, non-significant, or much lower than those obtained for Nexis:Trem2 (Table S1).
Further, we consider the datasets DS6_110 and DS9_110, which use the same injected tau strains as DS6 and DS9, respectively, but at 1:10 dilution27. We expected that the tau-strain-specific parameters \(\alpha\), \(\beta\), b, and p should be consistent between these experiments, but the seed rescale parameter, \(\gamma\), should be lower. We found that by applying Nexis:Trem2 on DS6_110 and DS9_110 after fixing the former parameters to their respective values for DS6 and DS9 and fitting only for \(\gamma\), the model provides strong fits to the data with lower \(\gamma\) values, as anticipated (Table 2).
Effect of Trem2 on tau burden and spread
NDM, Nexis:global, and Nexis:Trem2 model fits for dataset DS9. Panel (a): Empirical tau pathology observed in DS9 post hippocampal seeding of exogenous tau27. Panels (b)–(d): NDM, Nexis:global, and Nexis:Trem2 modeled tau pathology progression, respectively, in DS9 dataset at 4, 8, and 12 weeks post hippocampal injection. Panels (e)−(g): Scatter-plots depicting the correlation between empirical and ND-modeled (e), Nexis:global-modeled (f), as well as Nexis:Trem2-modeled (g) tau pathology, respectively, at 4, 8, and 12 weeks post hippocampal tau seeding. Pearson r values are provided for each scatter plot. HIP, hippocampal area; EC, entorhinal cortex; PAL, pallidum; STR, striatum.
Since the fitting results highlight Trem2 as the more significant modulator compared to homeostatic genes, the remainder of our results will focus on it. Figure 4 shows the results for DS9 in detail; they are very similar for DS6 (Fig. S3). For dataset DS4 (Fig. S2), while the Nexis models were significantly better than the passive NDM, improvement from Nexis:global to Nexis:microglia was not statistically significant (Table 1). Both Nexis-modeled distributions differ significantly from the NDM-modeled distribution. While both Nexis-modeled distributions look similar visually, there are clear differences between them that begin to become apparent at later time-points, especially in striatal and entorhinal areas. While both produce excellent recapitulation of empirical tau burden at all time points, the Nexis:Trem2 model produces by far the better fits (Fig. 4f, g). Model fits in all cases improve over time as may be expected from a ramifying pathological process. The Nexis:Trem2 fits incurred an increase in the microglia-dependent spread parameter, b (Table 1); hence Trem2 appears to allow more pathology to efflux from the hippocampal seed regions at early time-points (Fig. 4d).
Flow-map representations of tau accumulation and spread in DS9 dataset post hippocampal seeding. Tau burden and spread differences were evaluated between Nexis:global and Nexis:Trem2 modeled tau pathology. Blue arrows of tau spread (a) depict Nexis:global-modeled tau flow without the influence of Trem2 and indicate a positive difference (tau spread: Nexis:global > Nexis:Trem2). Magenta arrows of tau spread (b) depict tau-flow mediated by Trem2, above and beyond that modeled by Nexis:global and indicate a negative difference (tau spread: Nexis:global < Nexis:Trem2). Red and green spheres represent higher tau burden as modeled by Nexis:global and Nexis:Trem2, respectively. Sphere sizes correspond to the degree of tau burden. HIP: hippocampal area, contra. HIP, contralateral hippocampal area; lateral EC, lateral entorhinal cortex; PAL, pallidal area; PAS, parasubiculum; STR, striatal area.
We next assessed, visually, the effect that microglia are imparting on the spread of tau within the model. Flow-maps depicting major pathology spreading directions were constructed using Brainframe, our in-house Matlab-based function. Figure 5 highlights differences in tau burden and spread as modeled by Nexis:global and Nexis:Trem2 for DS9. A prevalence of magenta arrows (Fig. 5b) indicates that Trem2 allowed more widespread propagation of tau (also reflected in the higher b values in Table 1 for DS9). Although modeled tau propagated from the hippocampal seed area to connected regions, Trem2 appeared to mediate more spread of tau to the striatal, entorhinal, pallidal, and contralateral hippocampal regions. Similarly, a prevalence of green spheres in the striatal, pallidal, and hippocampal areas indicates that these regions evidence higher tau burden due to Trem2 than would otherwise be expected. Tau burden was unaffected by the presence of Trem2 in the parasubiculum (red sphere at all time points).
Changes in Trem2 expression and the subsequent response of tau
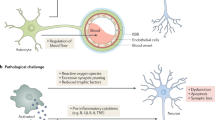
Summary of the relationship between Trem2 and tau. Panel (a): End time-point (12 weeks here) tau levels in response to simulated scaling of Trem2 expression levels in selected regions of the mouse brain. The inset depicts the baseline expression levels of Trem2 in the selected brain regions. Panel (b): Summary of the effect of Trem2 on tau accumulation and spread as modeled by Nexis:Trem2. The rate of tau accumulation (parameter p) was lower and the rate of tau spread (parameter b) was higher in the presence of Trem2. Nexis:global, which does not consider the effect of microglia, failed to capture the entire dynamics of tau, leading to poor model performance. Note that the illustrated relationship depends on the strain of tau being modeled. Note also that our speculation of exosomes contributing to trans-neuronal and microglia-mediated spread of tau has not been modeled using Nexis.
Given the effect of Trem2 on model performance and its quantified effects on tau transmissibility (b), we hypothesized that there would be a dose-dependent effect of Trem2 on tau, where higher Trem2 expression would lead to lower tau levels. In silico simulation of changes in baseline Trem2 levels at the global brain level, followed by application of the Nexis:Trem2 model, indicated a unidirectional trend in tau. Higher Trem2 expression resulted in a reduction of tau in five sample regions of the mouse brain (Fig. 6a). Specifically, there appeared to be a preponderance of an early effect of Trem2 in the hippocampus and the entorhinal cortex.
Nexis:microglia models applied to additional candidate genes implicated in tauopathies
We investigated Apoe, a strongly implicated AD-risk gene33,34,35 expressed both in microglia and astrocytes36. Nexis:Apoe did not outperform Nexis:global in any of the datasets (Table S2, Fig. S4). Since Trem2 and Apoe also work in concert in AD37, we conducted a combined Nexis:microglia analysis including them both, but results for all three datasets matched those produced by Nexis:Trem2 alone (Table S2). We then applied Nexis:microglia to some candidate genes involved in phagocytosis and inflammation (Sorl1, Cd33, Clu, Sirpa, and Aif1). The accumulation (p) and spread (b) parameters obtained for these genes were either negative or insignificant, agreeing with their role in containing and eliminating pathological entities.
Discussion
The spread of hyperphosphorylated tau in AD occurs along the brain’s connectome and is also influenced by the surrounding molecular players such as certain cell types. The presented modeling effort was inspired by and expands upon the “molecular nexopathy” paradigm5 in that a central role is noted for network-mediated spread, but also includes other sources of protein aggregation and mediators of trans-neuronal spread, especially microglia. Microglia co-localize with tau inclusions, taking up and releasing bioactive seed-competent tau, and thus emerging as key mediators of AD pathology propagation along with connectome-mediated spread18,38. Phagocytic microglia, instead of degrading tau, may package it into exosomes19 and form aberrant junctions with surrounding neurons through which tau might be transmitted between cells18,26. While prior studies13,14,15 have successfully quantified key aspects of pathology progression along the connectome, they do not constitute a true Nexopathy model, since they did not account for the influence of non-neuronal modulators.
Thus, here we presented a mathematical model called Nexis, that attempts to incorporate diverse effects including misfolded protein seeding, aggregation, connectome-mediated spread, as well as the modulation of all the above processes by extra-connectomic agents residing within the brain’s neurological milieu. Nexis augments prior quantitative models (NDM)13 by including microglial modulation via two key processes: local tau accumulation and its enhanced trans-neuronal transmission. Our results quantify how the inclusion of microglia improves the predictive power of the model and reduces the uncertainty in the accumulation and diffusion parameter estimates (see AIC and narrower confidence intervals around model parameters, Table 1). The full model recapitulates empirical data with \(R^2\) values up to 0.60. The NDM, on the other hand, cannot capture the extent of the spatiotemporal pathology because it fails to capture scale. Thus even though the individual Pearson r values per time-point are significant, the overall model fit (\(R^2\)) is very poor with the NDM.
The Nexis-modeled spatiotemporal progression of tau closely matches the Braak staging scheme10, with pathology progressing from the hippocampal injection site to the contralateral hippocampus and entorhinal cortex, followed by the neocortical, striatal, and thalamic areas. We acknowledge that the data used here were from a seeded mouse model, which may have biased the starting point of pathology to be the hippocampus. Future models applied to unseeded mouse models of AD would help since they do demonstrate tau spatiotemporal behavior as dictated by Braak staging39. In addition, the data are from a model of primary tauopathy, and hence cannot necessarily be characterized in the same way as AD tauopathy. This is because there is insufficient information on staging of primary tauopathies40, and the classical pattern of tau spread to the frontal regions might be due to the influence of amyloid-\(\beta\) in AD.
Microglial modulation was strongly dependent on the tau strain used in the specific dataset. Datasets DS6 and DS9 have medium to high seeding capacity and are characterized by tau from a P301S tauopathy mouse and recombinant fibrils, respectively. For these datasets, Trem2 was associated with decreased tau accumulation rate (p) and increased spread (b) to regions ipsi- and contralateral to the hippocampal seed area (Table 1). Moreover, Trem2 distribution (Fig. 2b) did not spatially correlate with regions with higher Trem2-mediated tau burden (Fig. 4d), suggesting that microglial effects are not strictly local. Instead, as shown by differences in fitted model parameters, Trem2 appears to mobilize tau increasing its network transmissibility. We hypothesize that Trem2 renders tau more mobile, thus increasing its brain-wide spread but reducing its accumulation rate. Although not possible to model here, the effect of Trem2 on tau mobilization may be occurring via the action of exosomes19. Trem2-mediated high tau efflux with slower accumulation may reflect clearance and possible mitigation of pathology. Mouse studies have indeed demonstrated Trem2-mediated decrease in tau accumulation in early disease stages41,42, Trem2-mediated neuroprotection43, as well as Trem2 loss resulting in enhanced tau seeding and exacerbated pathology24. Microglia can internalize and release seed-competent tau, but also potentially degrade its seeding activity38. Our model parameters here suggest that the decreased accumulation rate (p) may support the empirically observed reduced seeding activity38, whilst still maintaining tau spreading potential (b). Intriguingly, our results posit this effect of microglia to be specific to Trem2, and not to homeostatic microglial genes.We schematically summarize the relationship between Trem2 and tau as modeled by Nexis and as observed in these datasets, in Fig. 6.
Neither Nexis framework recapitulates one of the datasets, DS4, particularly well (Table 1). DS4, characterized by the human AD-brain derived tau isolate, demonstrated elevated rates of both tau accumulation (p) and spread (b) in the presence of either Trem2 or homeostatic microglia. However, neither Nexis:homeostasis nor Nexis:Trem2 was significantly better than Nexis:global. As observed by Kaufman et al.27, DS4 was a low-tangle tau strain with lower seeding capacity than DS6 and DS9. This may explain its comparatively weaker Nexis:global model fit, with a failure of any additional improvements with the inclusion of microglial genes. We also speculate that since DS4 was characterized by amyloid-comorbid tau from a human-AD brain it evinced higher values of accumulation and spread rates. AD derived amyloid-comorbid tau can acquire more pathogenic conformations leading to its aggregation and accumulation at a faster rate44. Our results thus point to the importance of quantifying the heterogeneous effects that tau has on neuropathology progression, importantly because tau strain diversity based on differences in bioactivity (seeding and spreading) may explain the myriad of AD clinical outcomes45.
Trem2 is upregulated in microglia during disease progression, and in fact a lack of Trem2 worsens tau pathology46. Similar observations have been made by Zhu et al.47 where Trem2 deletion caused enhanced spread of tau pathology from the injection site to other connected regions of a Trem2 knock-out mouse model. We showed here that Nexis is capable of confirming the histopathologically observed mechanistic relationship between Trem2 and tau (Fig. 6a). Trem2 expression levels lower and higher than baseline simulated its deletion (or knock-out) and over-expression, respectively. Nexis:Trem2 set up using varying scales of Trem2 expression could provide quantifiable trends of its inverse relationship with tau as quantified in five sample regions of the mouse brain. The baseline expression levels of Trem2 (Fig. 6a inset) in these regions is in agreement with the pattern observed in murine and human studies48, with the hippocampus and entorhinal cortex evidencing the highest levels followed by the neocortical and striatal regions. Apoe is a canonical risk-gene implicated in AD, and along with being a strong influencer of amyloid pathology, it is also suggested to influence microglial function and tau pathology49. We thus explored the effects of Apoe via Nexis:microglia model alone and in conjunction with Trem2 as these two genes appear to act in concert37. Compared to Nexis:Trem2, the lack of improvement in Nexis:Apoe and comparable performance by Nexis:Trem2:Apoe can be related to the fact that Apoe acts downstream of Trem237 (in disease conditions). Therefore, the key mediatory role manifested by Trem2 can be expected to encompass the contribution of Apoe.
In conclusion, our in silico Nexis:microglia model quantifies the contribution of microglia to tau neuropathology progression. It recapitulates the characteristics of observed tau pathology progression, providing distinct results specific to the tau species employed in the mouse models we studied. Although prior bench studies have suggested a modulatory role for microglia, the present study is the first to demonstrate this quantitatively on the whole mouse brain, and for specific microglial genes and specific tau conformations. In particular, our results provide quantitative support to the observations made by Asai et al.19 in that, microglia aid increased propagation of tau across brain regions. The Nexis:microglia framework can greatly aid experimentalists in further hypothesis generation, validation, and quantification. It may be readily adapted to interrogate the effects of other putative modulators of pathology progression such as astrocytes and oligodendrocytes.
Limitations and future directions
The datasets used monitored the levels of exogenous tau at 4, 8, and 12 weeks post hippocampal seeding. Additional time-points may enable a more thorough understanding of the spatiotemporal behavior of tau progression. The current monitoring over time was also not longitudinal in the real sense since measurements did not track the same animal. Instead, tau progression was observed in groups of tau-injected mice sacrificed at different time-points post-injection. Further, the gene expression data used as microglial signatures were from a healthy mouse brain and thus can infer only about the effects of baseline microglial density on pathology. However, interestingly, a recent study indeed supports the effect of baseline microglia, rather than the brain’s anatomic network, on tau propagation26. We were also restricted to using the coronal series from the AGEA due to its superior spatial coverage relative to the sagittal series, but with significantly fewer quantified genes. As a result, not all microglial genes of interest could be assessed with Nexis.
Next, Trem2 assumes opposing roles (protective vs. detrimental) in AD pathology depending on disease-stage. Such disease-stage dependent dual roles of Trem2 have been observed with respect to amyloid \(\beta\) plaque pathology50. Our current in silico investigation of the role of Trem2 in neuropathology progression is restricted to tau and is conducted on data acquired within three months of exogenous tau seeding in mice. Future goals would be to apply this approach to data collected at extended time-points post tau-seeding (depending on data availability), as well as to include amyloid pathology in the model since amyloid \(\beta\) has been speculated to potentiate microglial influence on tauopathy propagation26. A possibility that the seed region where tau was initially injected may confer specific molecular vulnerability to tau, remains outside the scope of the current study design. Our Nexis approach itself is fully capable of testing for site-specific effects but paucity of experimental data to date makes this challenging.
We also note that the nexopathy paradigm enumerates a far more diverse set of potential mechanisms by which molecular dysfunction might interact with the neural architecture than is possible to explore in a single study. Epigenetic51 or post-translational modifications52 due to variable causes will produce gene expression in diseased groups that is different from healthy gene expression used here. However, the primary goal of uncovering the molecular versus network correlates of AD topography may be largely unaffected by these additional factors, since they are either unlikely to have spatial gradients (e.g. age, environment) or have non-AD-like spatial patterning (oxidative/vascular stress).
Finally, the current modeling attempts were restricted to data from mouse models of primary tauopathy. There are two main reasons why we decided to start with mouse models: (1) the datasets compared here are from mice having the same genetic background thus minimizing any effects of genetic heterogeneity on model results, and (2) since mice are exogenously injected with pathology in specific brain regions, pathology progression from a common seed location can be modeled and compared. Pathology seeding data from humans are not possible, and despite the stereotyped progression of AD tau pathology, there is evidence that significant heterogeneity exists between patients at disease inception53. Another advantage of seeded mouse models is that the diverse effects of different strains of the pathological species can be compared. That being said, the true translational aspect of Nexis will be appreciated by its application to human data. This will also be important for inter-species comparison of the glial response to pathology, since transcriptional signatures of glia in AD pathology can be markedly different between mouse AD models and specimens from human AD patients54. Nevertheless, despite this difference, the fact that Trem2 impacts microglia function is a common theme in mouse and human AD54. We are currently in the process of curating and inspecting human data to which we can apply Nexis.
Methods
Study design
Empirical spatiotemporal tau pathology data were obtained from a study on PS19 mice expressing 4R1N P301S with human tau, exogenously injected with distinct strains of pathological tau27. Regional levels of tau burden were extracted from heat maps displaying semi-quantified pathology anatomically following the approach from Mezias et al.55. The Kaufman datasets used in this study are DS4 (containing tau isolate derived from the human AD-brain), DS6 (contained tau injectate from a P301S tauopathy mouse), and DS9 (characterized by recombinant tau fibrils). Our 426-region mouse connectome is from the Allen Brain Atlas (ABA)28, with 213 region bi-hemispheric symmetry. The microglial baseline spatial gene expression maps employed as signatures of baseline homeostatic and phagocytic microglia are from the ABA gene expression atlas (AGEA) of healthy 56-day old male C57BL/6J mice30. Note that these microglial signatures represent quiescent rather than activated states and may not be considered disease-activated microglia (DAM). Moreover we selectively use only the genes from the coronal AGEA and exclude those from the sagittal atlas because the spatial coverage of gene expression is higher in the former. Thus, the final gene list that we referred to has a subset containing 3855 genes from a total of approximately 26000 genes in the AGEA (sagittal + coronal).
Regional homeostatic microglial abundance was defined as the first principal component of the expression levels of P2ry12, Cx3cr1, Fcrls, Olfml3, Hexb, Siglech, Sox5, and Jun29,56. We separately tested the influence of Trem2, Cd33, and Apoe, activated microglial surface-markers and regulators of microglia-mediated phagocytosis strongly implicated in AD pathogenesis57. Cd33’s influence (Table S3) was negligible compared to Trem2 given its smaller effect size compared to Trem258.
Nexopathy in silico Model (Nexis)
The Nexis model is derived from the Network Diffusion Model (NDM)13—a mathematical model that describes pathology spread as a diffusive process between connected brain regions. Let \(c_{12}\) denote the inter-region connection strength between two brain regions (1,2) as inferred from tractography data, let and \(x_{1}(t)\) and \(x_{2}(t)\) be the pathology concentration at two brain regions at time t as inferred from semi-quantitative immunohistochemistry analysis from the specific reference study. Following the NDM13, we impose first-order diffusive dynamics:
where \(\beta\) is the global diffusivity rate that controls how fast pathology can flow from the more concentrated region to the less concentrated one independent of any other modulators.
Nexis:global adds another parameter to the NDM, \(\alpha x_{1}\), a linear accumulation or clearance term depending on its sign.
This pair-wise relationship can be extended to the entire brain connectivity network using vector \(\vec{x} \in {\mathbb {R}}^{n}\) over all n ROIs, to give \(\frac{d\vec{x}}{dt} = -\beta L \vec{x}\), where L is the graph Laplacian matrix (when \(\alpha = 0\))13.
Nexis:microglia
The Nexis:microglia framework incorporates the potential effects that different modulators, for instance cell-types such as microglia, may have on neuropathology progression, particularly in the context of AD and other tauopathies. It has two additional cell-type specific parameters p and b, which indicate the effect of the chosen cell type, u, on pathology accumulation/clearance and spread, respectively. Letting \(u_{1}\) and \(u_{2}\) indicate the local gene expression in regions 1 and 2, the extended pairwise relationship in (2) becomes
Equation (3) reduces to NDM equation (1) if \(b = p = \alpha = 0\), and can similarly be extended brain-wide via pathology \(\vec{x}\) and gene \(\vec{u}\) vectors to \(\frac{d \vec{x}}{dt} = [\Lambda ( \vec{u} ) - \beta {\tilde{L}}( \vec{u} )] \vec{x}\), where \(\Lambda\) is a diagonal matrix and \({\tilde{L}}\) is a new Laplacian, both now depending on gene expression u. See the Supplementary Information for a detailed explanation.
For convenience of reading, in the remainder of the document the model name ’Nexis:microglia’ will be replaced by Nexis:homeostasis in the case of homeostatic microglia or Nexis:(microglial gene name) in the case of a particular microglial gene being investigated.
Seed scaling and model initialization
Both NDM and Nexis require a seed-rescale parameter \(\gamma\), which denotes the scaling factor for initial pathology at the site of injection of pathology (tau in the current case). This parameter needs to be quantified since, although we know the site of the injection (hippocampus for instance), we do not know the amount of pathology being injected. Thus, the pathology at t = 0 is represented as a binary vector multiplied by \(\gamma\) and is used to initialize the model.
Robust strategy for parameter estimation
We implemented a two-step, robust strategy for estimating parameters for the NDM and Nexis models. First, we fit the spread parameter (\(\beta\)) for the NDM, followed by the accumulation and spread parameters (\(\alpha\) and \(\beta\)) for Nexis:global via grid search and random initialization. This was then followed by 100 iterations of bootstrapping with 80\(\%\) re-sampling. Next, for the Nexis:microglia model, we initialized the values of \(\alpha\) and \(\beta\) to those obtained from Nexis:global while also fitting (without initialization) the microglia-mediated accumulation and spread parameters (p and b). Anchoring the parameters shared by both Nexis frameworks within a reasonable range from each in the optimization routines prevents the algorithm from reaching local minima that yield better fits but have overall biologically-implausible parameter values. This also allowed us to assess reproducibility by acquiring the 95% confidence intervals around the mean estimates of the model fit as well as global and microglial parameters (simulations in Fig. 3 for Nexis:global and Nexis:Trem2). Importantly, model parameters were fit using all available quantified regions and all time points; prior formulations only allowed fitting to single time points14,15.
Statistics and reproducibility
The statistical significance of the differences in model fits between the NDM and the Nexis frameworks was determined by carrying out Fisher’s-R-to-Z transformation. The p value of significance was obtained by the Student’s t test on the Fisher-transformed Z scores. The model fits presented in Table 1 were determined by averaging the parameters across 100 iterations of bootstrapping with 80% re-sampling of the regions quantified by Kaufman et al.27. When fitting the Nexis:microglia models we used the mean and 95% confidence intervals determined for the Nexis:global parameters \(\alpha\), \(\beta\), and \(\gamma\) for each dataset as the initial guesses and bounds for those parameters, respectively; this helps to minimize the risk of finding degenerate solutions. Please see Supplementary Information for details on parameter fitting. The \(R^2\) values of the full model are within the range of the bootstrapped iterations of the 80% re-sampled data (Fig. 3c), thus confirming reproducibility. All analyses were conducted in Matlab R2020b.
Simulation of changes in Trem2 expression to assess response of tau
Significant improvement in model fit upon inclusion of Trem2 warranted further in silico quantification of tau response to simulated changes in Trem2 expression levels. Baseline Trem2 levels were scaled by factors of two above and below the baseline level and tau was quantified in the hippocampus, entorhinal cortex, neocortex, striatum, and amygdala. The scaling was performed by multiplying the microglial gene (here, Trem2) vector by powers of 2 (from \(2^{-3}\) to \(2^4\)) before entering it into the model.
Ethics approval and consent to participate
All data were obtained from publicly available datasets and published papers. No human subjects, live animals, patient or animal derived tissue, cell lines, or biological material were directly used in this research and thus, no consent or ethics approval was required.
Data availability
The source data on the mouse models, gene expression, and mesoscale mouse connectome as well as the computer code to run the models are available at our repository at https://github.com/Raj-Lab-UCSF/Nexis.
References
Ossenkoppele, R. et al. Tau pet patterns mirror clinical and neuroanatomical variability in alzheimer’s disease. Brain 139, 1551–67 (2016).
Arriagada, P. V., Growdon, J. H., Hedley-Whyte, E. T. & Hyman, B. T. Neurofibrillary tangles but not senile plaques parallel duration and severity of alzheimer’s disease. Neurology 42, 9 (1992).
Forman, M. S. et al. Signature tau neuropathology in gray and white matter of corticobasal degeneration. Am. J. Pathol. 160, 2045–53 (2002).
Reitz, C., Honig, L., Vonsattel, J., Tang, M.-X. & Mayeux, R. Memory performance is related to amyloid and tau pathology in the hippocampus. J. Neurol. Neurosurg. Psychiatry 80, 7 (2009).
Warren, J. D. et al. Molecular nexopathies: A new paradigm of neurodegenerative disease. Trends Neurosci. 36, 561–9 (2013).
Higuchi, M., Trojanowski, J. Q. & Lee, V. M. Tau protein and tauopathies. Neuropsychopharmacol. Fifth Gener. Prog. 16, 564 (2002).
Hanger, D. P., Anderton, B. H. & Noble, W. Tau phosphorylation: The therapeutic challenge for neurodegenerative disease. Trends Mol. Med. 15, 112–9 (2009).
Clavaguera, F. et al. Brain homogenates from human tauopathies induce tau inclusions in mouse brain. Proc. Natl. Acad. Sci. USA 110, 9535–40 (2013).
Hardy, J. & Revesz, T. The spread of neurodegenerative disease. N. Engl. J. Med. 366, 2126–8 (2012).
Braak, H. & Braak, E. Neuropathological stageing of alzheimer-related changes. Acta Neuropathol. 82, 239–59 (1991).
Iba, M. et al. Synthetic tau fibrils mediate transmission of neurofibrillary tangles in a transgenic mouse model of alzheimer’s-like tauopathy. J. Neurosci. 33, 1024–37 (2013).
Frost, B. & Diamond, M. I. Prion-like mechanisms in neurodegenerative diseases. Nat. Rev. Neurosci. 11, 155–9 (2010).
Raj, A., Kuceyeski, A. & Weiner, M. A network diffusion model of disease progression in dementia. Neuron 73, 1204–15 (2012).
Raj, A. et al. Network diffusion model of progression predicts longitudinal patterns of atrophy and metabolism in alzheimer’s disease. Cell Rep. 10, 359–369 (2015).
Mezias, C., LoCastro, E., Xia, C. & Raj, A. Connectivity, not region-intrinsic properties, predicts regional vulnerability to progressive tau pathology in mouse models of disease. Acta Neuropathol. Commun. 5, 61 (2017).
Hammond, T. R., Robinton, D. & Stevens, B. Microglia and the brain: Complementary partners in development and disease. Annu. Rev. Cell Dev. Biol. 34, 523–544. https://doi.org/10.1146/annurev-cellbio-100616-060509 (2018).
Bolos, M. et al. Direct evidence of internalization of tau by microglia in vitro and in vivo. J. Alzheimers Dis. 50, 77–87 (2016).
Busche, M. & Hyman, B. Synergy between amyloid-beta and tau in alzheimer’s disease. Nat. Neurosci. 23, 11 (2020).
Asai, H. et al. Depletion of microglia and inhibition of exosome synthesis halt tau propagation. Nat. Neurosci. 18, 1584–93 (2015).
Ingelsson, M. et al. Early abeta accumulation and progressive synaptic loss, gliosis, and tangle formation in ad brain. Neurology 62, 925–31 (2004).
McQuade, A. et al. Gene expression and functional deficits underlie trem2-knockout microglia responses in human models of alzheimer’s disease. Nat. Commun. 11, 5370. https://doi.org/10.1038/s41467-020-19227-5 (2020).
Ulrich, J. D., Ulland, T. K., Colonna, M. & Holtzman, D. M. Elucidating the role of trem2 in alzheimer’s disease. Neuron 94, 237–248 (2017).
Parhizkar, S. et al. Loss of trem2 function increases amyloid seeding but reduces plaque-associated apoe. Nat. Neurosci. 22, 191–204. https://doi.org/10.1038/s41593-018-0296-9 (2019).
Leyns, C. E. G. et al. Trem2 function impedes tau seeding in neuritic plaques. Nat. Neurosci. 22, 1217–1222 (2019).
Maphis, N. et al. Reactive microglia drive tau pathology and contribute to the spreading of pathological tau in the brain. Brain 138, 1738–55 (2015).
Pascoal, T. A. et al. Microglial activation and tau propagate jointly across braak stages. Nat. Med. 2021, 117 (2021).
Kaufman, S. K. et al. Tau prion strains dictate patterns of cell pathology, progression rate, and regional vulnerability in vivo. Neuron 92, 796–812 (2016).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–14 (2014).
Eme-Scolan, E. & Dando, S. J. Tools and approaches for studying microglia in vivo. Front. Immunol. 11, 583647 (2020).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–76 (2007).
Villegas-Llerena, C., Phillips, A., Garcia-Reitboeck, P., Hardy, J. & Pocock, J. M. Microglial genes regulating neuroinflammation in the progression of alzheimer’s disease. Curr. Opin. Neurobiol. 36, 74–81. https://doi.org/10.1016/j.conb.2015.10.004 (2016).
Nuvolone, M. et al. Prion pathogenesis is unaltered in the absence of sirpalpha-mediated “don’t-eat-me’’ signaling. PLoS ONE 12, e0177876. https://doi.org/10.1371/journal.pone.0177876 (2017).
Strittmatter, W. J. et al. Apolipoprotein e: High-avidity binding to beta-amyloid and increased frequency of type 4 allele in late-onset familial alzheimer disease. Proc. Natl. Acad. Sci. USA 90, 1977–81. https://doi.org/10.1073/pnas.90.5.1977 (1993).
Shi, Y. et al. Apoe4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549, 523–527. https://doi.org/10.1038/nature24016 (2017).
Deming, Y. et al. Genome-wide association study identifies four novel loci associated with alzheimer’s endophenotypes and disease modifiers. Acta Neuropathol. 133, 839–856. https://doi.org/10.1007/s00401-017-1685-y (2017).
Fernandez, C. G., Hamby, M. E., McReynolds, M. L. & Ray, W. J. The role of apoe4 in disrupting the homeostatic functions of astrocytes and microglia in aging and alzheimer’s disease. Front. Aging Neurosci. 11, 14. https://doi.org/10.3389/fnagi.2019.00014 (2019).
Krasemann, S. et al. The trem2-apoe pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Hopp, S. C. et al. The role of microglia in processing and spreading of bioactive tau seeds in alzheimer’s disease. J. Neuroinflam. 15, 269 (2018).
Hurtado, D. E. et al. Abeta accelerates the spatiotemporal progression of tau pathology and augments tau amyloidosis in an alzheimer mouse model. Am. J. Pathol. 177, 1977–88. https://doi.org/10.2353/ajpath.2010.100346 (2010).
Chung, D. C., Roemer, S., Petrucelli, L. & Dickson, D. W. Cellular and pathological heterogeneity of primary tauopathies. Mol. Neurodegener. 16, 57. https://doi.org/10.1186/s13024-021-00476-x (2021).
Sayed, F. A. et al. Differential effects of partial and complete loss of trem2 on microglial injury response and tauopathy. Proc. Natl. Acad. Sci. USA 115, 10172–10177 (2018).
Lee, S. H. et al. Trem2 restrains the enhancement of tau accumulation and neurodegeneration by beta-amyloid pathology. Neuron 2021, 65 (2021).
Condello, C., Yuan, P. & Grutzendler, J. Microglia-mediated neuroprotection, trem2, and alzheimer’s disease: Evidence from optical imaging. Biol. Psychiatry 83, 377–387. https://doi.org/10.1016/j.biopsych.2017.10.007 (2018).
Stancu, I. C., Vasconcelos, B., Terwel, D. & Dewachter, I. Models of beta-amyloid induced tau-pathology: The long and folded road to understand the mechanism. Mol. Neurodegener. 9, 51 (2014).
Dujardin, S. et al. Tau molecular diversity contributes to clinical heterogeneity in alzheimer’s disease. Nat. Med. 26, 1256–1263 (2020).
Jiang, T. et al. Silencing of trem2 exacerbates tau pathology, neurodegenerative changes, and spatial learning deficits in p301s tau transgenic mice. Neurobiol. Aging 36, 3176–3186. https://doi.org/10.1016/j.neurobiolaging.2015.08.019 (2015).
Zhu, B. et al. Trem2 deletion enhances tau dispersion and pathology through microglia exosomes. Mol. Neurodegener. 17, 58. https://doi.org/10.1186/s13024-022-00562-8 (2022).
Guilarte, T. R., Rodichkin, A. N., McGlothan, J. L., Acanda-Rocha, A. M. & Azzam, D. J. Imaging neuroinflammation with tspo: A new perspective on the cellular sources and subcellular localization. Pharmacol. Ther. 234, 108048. https://doi.org/10.1016/j.pharmthera.2021.108048 (2022).
Shi, Y. et al. Apoe4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549, 523–527 (2017).
Jay, T. R. et al. Disease progression-dependent effects of trem2 deficiency in a mouse model of alzheimer’s disease. J. Neurosci. 37, 637–647 (2017).
Razin, A. & Cedar, H. Dna methylation and gene expression. Microbiol. Rev. 55, 451–8 (1991).
Saxena, S. & Caroni, P. Selective neuronal vulnerability in neurodegenerative diseases: From stressor thresholds to degeneration. Neuron 71, 35–48 (2011).
Torok, J., Maia, P. D., Powell, F., Pandya, S. & Raj, A. A method for inferring regional origins of neurodegeneration. Brain141, 863–876 (2018). https://doi.org/10.1093/brain/awx371, https://academic.oup.com/brain/article-pdf/141/3/863/25046306/awx371.pdf.
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal trem2-dependent and trem2-independent cellular responses in alzheimer’s disease. Nat. Med. 26, 131–142. https://doi.org/10.1038/s41591-019-0695-9 (2020).
Mezias, C. & Raj, A. The role of directionality in determining spatiotemporal tau pathology differs between ad-like and non-ad-like mouse models. BioRxiv 2020, 63 (2020).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–28. https://doi.org/10.1038/ni.2419 (2012).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Zhao, L. Cd33 in alzheimer’s disease—biology, pathogenesis, and therapeutics: A mini-review. Gerontology 65, 323–331 (2019).
Acknowledgements
We thank the NIH for their generous support. This work was funded by grants RF1AG062196 and R01AG072753 from the NIH NIA to Dr. Ashish Raj, and R01NS092802/183412 from the NIH NINDS to Dr. Ashish Raj.
Author information
Authors and Affiliations
Contributions
A.R. conceptualized the study and developed the original network diffusion model (Raj et al.13). C.A. performed the literature search and helped in the conceptualization of the study. P.M., J.T., and C.M. expanded the NDM and optimized the Nexis framework. J.T. validated the reproducibility of the model by bootstrapping. C.M. curated and processed the tauopathy data. J.T. and C.M developed a Matlab-based 'Brainframe' toolbox for pathology depiction on glass brains. C.A. conducted the study, analyzed the results, and generated the figures. C.A. and P.M. co-wrote the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Anand, C., Maia, P.D., Torok, J. et al. The effects of microglia on tauopathy progression can be quantified using Nexopathy in silico (Nexis) models. Sci Rep 12, 21170 (2022). https://doi.org/10.1038/s41598-022-25131-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25131-3
This article is cited by
-
The anti-Alzheimer potential of novel spiroindolin-1,2-diazepine derivatives as targeted cholinesterase inhibitors with modified substituents
Scientific Reports (2023)
-
Network models to enhance the translational impact of cross-species studies
Nature Reviews Neuroscience (2023)
-
Connectome-based modelling of neurodegenerative diseases: towards precision medicine and mechanistic insight
Nature Reviews Neuroscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.