Abstract
The aim of this study was to assess the dynamics of microbial communities and antimicrobial resistance genes (ARGs) in the chicken gut following amoxicillin and thiamphenicol treatments and potential co-selection of ARGs. To this purpose, the microbial community composition, using 16S rRNA NGS, and the abundance of ARGs conferring resistance to β-lactams and phenicols, using qPCRs, were determined. Results revealed that the administered antimicrobials did not significantly reduce the gut microbiota diversity, but changed its composition, with taxa (e.g. Gallibacterium and Megamonas) being enriched after treatment and replacing other bacteria (e.g. Streptococcus and Bifidobacterium). Positive correlations were found between ARGs (e.g. cmlA, blaCMY-2, and blaSHV) and the relative abundance of specific taxa (e.g. Lactobacillus and Subdoligranulum). The selective pressure exerted by both amoxicillin and thiamphenicol resulted in an increased abundance of ARGs conferring resistance to β-lactams (e.g. blaTEM-1, blaSHV, and blaCTX-M1-like) and phenicols (e.g. floR and cmlA). These findings, together with the co-occurrence of genes conferring resistance to the two antimicrobial classes (e.g. blaTEM-1 and cmlA), suggest a possible interaction among antimicrobials on resistance emergence, possibly due to the presence of mobile genetic elements (MGEs) carrying multiple resistance determinants.
Similar content being viewed by others
Introduction
A growing body of evidence suggests that antimicrobial use (AMU) in conventional animal farming contributes to the emergence and spread of antimicrobial resistance (AMR) in bacterial populations, posing a significant threat to both animal and human health1,2,3. Mass drug administration, including antimicrobial administration through feed and drinking water to large numbers of animals at once, is likely to have contributed to the reduced therapeutic efficacy of some antimicrobials and generation of multidrug-resistant bacteria4. Chickens harbour bacteria, including human pathogens, resistant to several classes of antimicrobials5,6, some of which are listed as critically important antimicrobials by the World Health Organisation (WHO)7. Amoxicillin, which belongs to the class of β-lactams, represents one of the most commonly used antimicrobials in humans8 and broilers9. Previous studies showed that β-lactam antibiotic use can favour the emergence of bacteria resistant to third-generation cephalosporins (3GCs), which represent one of the few available options for treating multidrug-resistant bacterial infections in humans10. Thiamphenicol is an analogue of chloramphenicol and a broad-spectrum bacteriostatic antimicrobial, active against both Gram-positive and Gram-negative bacteria11. Due to the paucity of new drugs to counteract AMR, dismissed antimicrobials like chloramphenicol, which use is not allowed in animals, regained utility for the treatment of multidrug-resistant bacteria in human medicine12. Questions remain as to whether the selective pressure exerted by thiamphenicol might result in the emergence of bacteria resistant to chloramphenicol. ARGs against β-lactams and phenicols can be harboured in mobile genetic elements (MGEs), such as plasmids, and intra- and inter-species (e.g. between Enterobacteriaceae and Clostridiaceae) horizontal gene transfer (HGT) is a common event contributing to the emergence and dissemination of resistance in microbial communities13.
In this study, we assessed whether the prophylactic administration of amoxicillin and thiamphenicol affects the chicken gut microbiota composition, increasing or decreasing the microbial diversity and abundance of specific taxa, as well as whether influences the abundance of ARGs conferring resistance to critically or highly important antimicrobials, such as 3GCs, carbapenems and chloramphenicol. Furthermore, we investigated whether the abundance of specific taxa correlated with ARG occurrence and whether the administration of amoxicillin and thiamphenicol resulted in co-selection of ARGs conferring resistance to the other antimicrobial class.
Results
General description of sequences
After the quality filtering step, removal of chimeric fragments, and read merging, a total of 3,378,323 reads with 3007 different features was obtained, with an average of 27,244 sequences per individual sample. After quality filtering, none of the samples was excluded from the analysis of microbial communities.
Amoxicillin and thiamphenicol treatments influence microbial diversity and the abundance of specific taxa
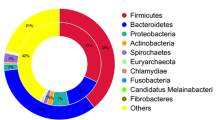
Using 16S rRNA NGS, the gut microbial community composition of the chicks in each group was characterized at different time points. At phylum level, microbiota composition varied with age rather than with treatment (Supplementary Fig. S1). Proteobacteria were the most abundant phyla at 1 day of age (d.o.a.), Firmicutes became dominant at later stages, while Bacteroidota were highly abundant in caecum samples collected at 46 d.o.a. Similar dynamics were observed also at family level, since Enterobacteriaceae and Clostridiaceae were significantly more abundant at 1 d.o.a. in all groups, Lactobacillaceae, Lachnospiraceae, and Ruminococcaceae seemed to bloom at 8 d.o.a., and Rikenellaceae were the dominant family in the caecum samples collected at 46 d.o.a. (Fig. 1; Supplementary Fig. S2).
Heatmap representing the microbial community composition at family level. The heatmap was generated in R (version 4.2.1) (https://www.r-project.org/) using package pheatmap (version 1.0.12).
Early-age administration
In both α-diversity indices (Fig. 2A,B), there was a trend towards increasing diversity from early to late time points in all groups; however, the only significant differences were between the group treated with amoxicillin (AMX1) and the other groups on day 21 post treatment (p.t.), and within AMX1 group between day 21 p.t. and the other time points. PERMANOVA showed that the microbial community was significantly different between the group treated with thiamphenicol (THP1) and the other two groups (i.e. AMX1 and control) on day 1 p.t. (p < 0.001) and on day 12p.t. (p = 0.048), while there were no differences at the last time point. These findings are supported by the NMDS plots (Fig. 2C–E), in which a clear spatial separation between THP1 and the remaining groups at the first two time points was observed.
α-Diversity within groups treated at early-age at different time points using Shannon (A) and Simpson (B) indexes. Boxplots represent 25th to 75th percentiles and whiskers showing a maximum of ×1.5 the interquartile range (IQR), and different letters indicate significant differences within the α-diversity indexes (p < 0.05). β-diversity between treatment groups on day 1 p.t. (C), day 12 p.t. (D) and day 21 p.t. (E). Samples are clustered according to Bray Curtis distances.
LEfSe analysis performed at the genus level at each time point showed an increased abundance of genus cc_115 after thiamphenicol administration (LDA = 4.40), and reduction of Helicobacter (LDA = 4.40) and Candidatus Arthomitus (LDA = 4.49) in the control group on day 1 p.t. (Supplementary Fig. 3A). At the following time point (12 d.p.t.), Bacteroides (LDA = 4.79) were reduced in the AMX1 group, while the genus Streptococcus was enriched in the control group (Supplementary Fig. 3B). On day 21 p.t. (Supplementary Fig. 3C), six taxa were enriched in the AMX1 group, including Sphingomonas (LDA = 4.11), Megamonas (LDA = 4.69) and Bacteroides (LDA = 4.31), one in the THP1 group, i.e. Gallibacterium (LDA = 4.35), and two in the control group, i.e. Streptococcus (LDA = 4.65) and Bifidobacterium (LDA = 4.13).
Middle-age administration
Middle-age administration of amoxicillin and thiamphenicol did not affect the α-diversity of the gut microbiota, as shown by both Shannon’s and Simpson’s indices (Fig. 3A,B). On the contrary, PERMANOVA and NMDS plot (Fig. 3C,D) showed differences in the microbial community composition (β-diversity) between the group treated with thiamphenicol (THP2) and the other two groups (i.e. amoxicillin treated (AMX2) and control), but only on day 1 p.t. (p < 0.001).
α-Diversity within groups treated at middle-age at different time points using Shannon (A) and Simpson (B) indexes. Boxplots represent 25th to 75th percentiles and whiskers showing a maximum of ×1.5 the interquartile range (IQR). β-Diversity between treatment groups on day 1 p.t. (C) and day 9 p.t. (D). Samples are clustered according to Bray Curtis distances.
On day 1 p.t., Megamonas (LDA = 4.63) was enriched in the AMX2 group, Ruminococcus (LDA = 4.61) and cc_115 (LDA = 4.38) in THP2 group, whereas Helicobacter (LDA = 4.69) and Peptoniphilus (LDA = 4.52) were more abundant in the control (Supplementary Fig. 4A). At day 9 p.t. Gallibacterium was more abundant in the thiamphenicol treated group, while Peptoniphilus was less abundant in the control group (Supplementary Fig. 4B).
Amoxicillin and thiamphenicol influence the abundance of β-lactam and phenicol resistance genes
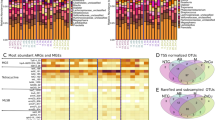
With the exception of blaNDM, all the other ARGs investigated were detected in at least one sample. All of the detected ARGs were also identified in the control group, with the exception of blaVIM-2 and blaOXA-1. Genes conferring resistance to either β-lactams (i.e. blaTEM-1, blaSHV, blaOXA-1 and blaOXA-48) or phenicols (i.e. catA1, catB3, floR, and cmlA) were detected in chicks of 1 d.o.a. (Supplementary Fig. S5). In the caecum samples collected at the slaughter house, seven out of fourteen ARGs were identified (i.e. blaTEM-1, blaCMY-2, catA1, catA2, catB3, floR, and cmlA). All samples were characterized by at least one ARG (min = 1, max = 7, and mean = 3.83), and 89.84% of the samples showed resistance to both antimicrobial classes. Overall, the relative abundance of ARGs conferring resistance to β-lactams and phenicols was significantly increased after treatments; however, genes conferring resistance to β-lactams and phenicols showed different temporal fluctuations in the different groups.
Early-age treatment
The relative abundance of ARGs conferring resistance to phenicols was significantly increased in both AMX1 and THP1 groups compared to the control (Fig. 4A,B); however, while abundance was significantly higher at all time points in the THP1 group, it decreased on day 21 p.t. in the AMX1 group. Compared to the control group, ARGs against β-lactams were more abundant in the THP1 group only on day 1 p.t., while the relative abundance of these ARGs was higher in the AMX1 from day 12 p.t. onwards.
Relative abundance of all genes conferring resistance to β-lactams (A) and phenicols (B) to 16S rRNA copy number per group per time point after early-age administration of amoxicillin and thiamphenicol. Relative abundance of genes floR (C), cmlA (D), blaTEM-1 (E), blaSHV (F), and blaCTX-M1-LIKE (G) to 16S rRNA copy number per group per time point. p < 0.05 shown as *. For easiness of representation, only ARGs showing significant differences between groups at the same time point are reported.
When considering individual ARGs, floR was more abundant in both AMX1 and THP1 groups than in the control group only on day 1 p.t. (Fig. 4C), while cmlA showed higher abundance in both AMX1 and THP1 groups until day 12 p.t. (Fig. 4D). None of the four remaining ARGs conferring resistance to phenicols was enriched after amoxicillin or thiamphenicol administration at any time point. On day 1 p.t., blaTEM-1 was more abundant in both the AMX1 and THP1 groups compared to the control; however, while the abundance in the AMX1 group remained significantly higher at the following time points, it decreased in the THP1 group (Fig. 4E). blaSHV was enriched in the AMX1 group on day 1 p.t. (Fig. 4F), while blaCTX-M1-like was significantly more abundant in the THP1 group compared to the other two groups on day 12 p.t. (Fig. 4G). Of the remaining ARGs conferring resistance to β-lactams, none was enriched after both antimicrobial administrations.
Middle-age administration
The relative abundance of both β-lactam and phenicol ARGs was significantly increased in the treated groups on day 1 p.t.; however, on day 9 p.t. only genes conferring resistance to the class of the antimicrobial administered were enriched (Fig. 5A,B). floR and cmlA were more abundant after the administration of both AMDs but, while cmlA was enriched on day 1 post both treatments, floR was enriched on day 1 p.t. in the AMX2 group and on day 9 p.t. in the THP2 group (Fig. 5C,D). Interestingly, catA1, conferring resistance to phenicols, was enriched only after the administration of amoxicillin (1 d.p.t.), while blaCMY-2, conferring resistance to β -lactams, was enriched in both the AMX2 (1 and 9 d.p.t.) and THP2 (9 d.p.t.) groups (Fig. 5E,F). None of the remaining ARGs was significantly enriched after either treatments at any time-point.
Relative abundance of all genes conferring resistance to β-lactams (A) and phenicols (B) to 16S rRNA copy number per group per time point after middle-age administration of amoxicillin and thiamphenicol. Relative abundance of genes catA1 (C), floR (D), cmlA (E), and blaCMY-2 (F), to 16S rRNA copy number per group per time point. p < 0.05 shown as *. For easiness of representation, only ARGs showing significant differences between groups at the same time point are reported.
Microbiome and ARGs composition of caecum samples
Neither the type of AMDs administered nor the age of treatment seemed to have affected the microbiota diversity of the caecum samples collected at the slaughterhouse, since no significant differences in α- or β-diversity were detected (Fig. 6A–C). Similarly, there were no significant differences in the abundance of ARGs between treated and control groups (Fig. 6D,E).
α-Diversity within caecum samples using Shannon (A) and Simpson (B) indexes. Boxplots represent 25th to 75th percentiles and whiskers showing a maximum of ×1.5 the interquartile range (IQR). β-Diversity between groups (C). Samples are clustered according to Bray Curtis distances. Relative abundance of all genes conferring resistance to β-lactams (D) and phenicols (E) to 16S rRNA copy number per group.
Associations between microbial communities and ARGs and co-occurrence of ARGs
The associations between ARGs occurrence and specific taxa (genus level) are summarized in Table 1. Seventeen taxa were significantly associated with at least one ARG, Lactobacillus showing the highest number of associations (n = 3), while cmlA and blaCMY-2 showed the highest number of associations with specific taxa. The occurrence of blaCMY-2 showed positive associations with five genera (e.g. Subdoligranulum and Butyricicoccus) and negative associations with Streptococcus, Lactobacillus, and Enterococcus, while blaSHV was positively correlated with five taxa, including Lactobacillus and Faecalibacterium. cmlA was positively associated with genera Lactobacillus, Bacteroides, and Subdolingranum, while negative associations were identified with eight taxa, including Escherichia/Shigella and Alistipes. Positive associations were also identified between floR and genera Escherichia/Shigella and Enterococcus. Both catA1 and catB3 were correlated with an increased abundance of Ruminococcus torques, while the latter gene was also associated with Clostridia.
Positive correlations were observed between genes conferring resistance to the same antimicrobial class and also between genes conferring resistance to β-lactams and phenicols. In detail, positive correlations were identified between blaCMY-2 and catA2 (Spearman r = 0.2561, p = 0.003), between blaTEM-1 and floR (Spearman r = 0.3662, p < 0.001) and cmlA (Spearman r = 0.4133, p < 0.001), and between blaCTX-M1-LIKE and cmlA (Spearman r = 0.2286, p = 0.009). Data regarding the co-occurrence of ARGs are reported in Table 2.
Discussion
The present study investigated longitudinally the selective pressure exerted by the administration of amoxicillin and thiamphenicol on the chicken gut microbiota and associated ARGs. Microbial community showed a strong age-dependent dynamic and, in accordance with previous observations14,15, Enterobacteriaceae were the predominant taxa at 1 d.o.a., while Lactobacillaceae, Lachnospiraceae, and Ruminococcaceae dominated the microbiota at later time points. According to previous observations, Rikenellaceae were highly abundant in caecal samples collected at 46 d.o.a.16. Previous studies reported contrasting effects of AMU on chicken gut microbial diversity (α-diversity). Although Le Roy et al. described that AMU significantly decreased the diversity and richness of the chicken gut microbiota17, other studies did not find any significant change, if not a trend toward increased microbial diversity14,18. Indeed, the administration of amoxicillin and thiamphenicol does not seem to reduce microbial diversity and richness, to the point that a significant increase in taxonomic diversity was observed in the chicken gut on day 21 after early-age administration of amoxicillin. However, comparison among bacterial communities (β-diversity) revealed that the gut microbiota of chickens treated with thiamphenicol was significantly different from those of the other groups and that early-age administration caused a longer shift in the microbial community composition (i.e. up to 12 d.p.t. in early-age treated group and only on 1 d.p.t. in middle-age treated group). Overall these findings seem to suggest that resistant bacteria could bloom under the selective pressure of amoxicillin and thiamphenicol, compensating for the loss of non-resistant species and that AMD administration at early-age can cause a more persistent perturbation of the gut microbiota. In addition to the risk of selecting for resistant bacterial populations, even if high genetic diversity of the gut microbiota is generally considered beneficial for chicken health, proliferation of usually lowly abundant bacteria might result in an altered microbial community structure, causing metabolic dysfunction and increasing susceptibility to opportunistic pathogens14,17.
The relative abundance of ARGs conferring resistance to β-lactams and phenicols increased after administration of either amoxicillin or thiamphenicol compared to the control group. Even if the selection of ARGs against the other antimicrobial class seems to be transitory (i.e. up to 12 d.p.t. in the early-age treated groups and only on 1 d.p.t. in the middle-age treated groups), this finding suggests a potential co-selection for resistance to amoxicillin and thiamphenicol, which is consistent with previous studies reporting increased resistance against β-lactams and phenicols after exposure to florfenicol and amoxicillin, respectively9,19. The co-selection and persistence of genes conferring resistance to phenicols (e.g. cmlA and floR) after exposure to amoxicillin might represent a concern for public health, since in recent years chloramphenicol has been used for the treatment of multidrug-resistant bacteria in humans, especially in low income countries where therapeutic alternatives are scarcely available12. Since blaTEM-1, cmlA, and floR were positively correlated and showed similar dynamics in the chicken gut (i.e. increased abundance after both treatments), the observed co-selection of resistance might rely on the presence of resistance determinants to β-lactams and phenicols on the same genetic element, as previously reported20,21. The co-selection for resistance to phenicols and β-lactams seems to involve also AmpC- and extended-spectrum β lactamases (ESBL)-encoding genes, which confer resistance to third-generation cephalosporins (3GCs). Indeed, treatment with thiamphenicol seems to exert a selective pressure on blaCMY-2 and blaCTX-M1-like, which encode for AmpC and ESBL, respectively. Notably, both blaCMY-2 and blaCTX-M1-like co-occurred with genes conferring resistance to phenicols (cmlA and catA2, respectively). Furthermore, cmlA showed positive associations with genera Lactobacillus and Subdoligranulum, which positively correlated with genes conferring resistance to 3GCs (blaCMY-2 and blaSHV, respectively), corroborating the hypothesis of a co-selection for resistance. Although Lactobacillus and Subdoligranulum are beneficial for the chicken gut health, cmlA, blaCMY-2, and blaSHV genes have been previously described on mobile genetic elements (MGEs) carrying multiple resistance determinants20,21,22, thus contributing to the emergence of multidrug-resistant bacteria, including human and animal pathogens, and representing a concern for veterinary and public health. The molecular detection of ARGs was performed on the total DNA, thus the associations inferred via statistical analysis do not necessarily mean that a taxon harboured a specific resistance gene; however, cmlA and bla genes have been previously reported in Lactobacillaceae23,24 and Firmicutes25. Similarly, positively correlations between genes conferring resistance to phenicols (i.e. catA1 and catB3) and genera Ruminococcus and phylum Firmicutes have been described26. The administration of amoxicillin also seems to exert selective pressure on blaSHV and blaCMY-2; even if transitory, it represents another concern for public health, since these genes conferring resistance to 3GCs are able to be transferred to the soil through fertilization with livestock manure, which is an agricultural common practice, and to spread and persist in the environment27,28. Considering the importance regained by chloramphenicol for the treatment of human infections, positive associations between floR and Escherichia/Shigella and Enterococcus represent a threat to human health, because bacteria belonging to these genera can cause severe infections in humans and can harbour multidrug resistance determinants in MEGs, which can be transferred to other bacterial species5,29,30,31.
Caecum samples collected at the slaughterhouse did not show any significant difference in ARGs abundance and microbial community composition between treated and untreated groups. Birds of all groups showed respiratory and intestinal symptoms and therefore were treated with doxycycline for five consecutive days from day 33 onwards. This treatment could be responsible for the flattening of the results, hampering to investigate the long-term effect of the administration of amoxicillin and thiamphenicol on the chicken gut microbiota and associated AMR.
In conclusion, data gathered in the present study suggest that amoxicillin and thiamphenicol treatments might contribute to the co-selection and persistence of resistant bacteria and ARGs against β-lactams (including 3GCs) and phenicols in the chicken gut, posing a health threat. Indeed, conventional poultry farming manure used for soil fertilization can promote the spread and dissemination of AMR in the environment and possibly the transmission to other animals and also humans, via dispersion into waterways or by entering the food chain28,32. Therefore, the use of these antimicrobials for the treatment of bacterial infections in poultry farming should be reconsidered, as a reduction in their use could indirectly contribute to reducing the impact of AMR in a One Health perspective.
Materials and methods
Animals and samples collection
One hundred one-day-old healthy Ross 308 chicks from a single hatchery were randomly allocated in five different pens in the animal facility of the Department of Veterinary Science, University of Turin, and reared until 46 days of age as part of a zootechnical trial. This trial was approved by the Ethics and Animal Welfare Committee of the Department of Veterinary Sciences (protocol n. 2796/2020), University of Turin (Italy). All broilers were vaccinated against Marek’s disease and Infectious Bursal disease at the hatchery. Additionally, one-day old chicks were vaccinated against coccidiosis (coarse spray), Newcastle disease and Infectious Bronchitis disease (fine spray). Each pen was equipped with a bucket-type feeder and a drinker with wood shavings as litter. The environmental conditions (lighting, temperature, relative humidity, and ventilation) were controlled accordingly to the Ross broiler management guidelines. Birds received the following feeding program to meet the standard nutritional requirements: a commercial starter diet (230 g/kg of CP) from day 0 to 12 (starter period) and a commercial grower diet (185 g/kg of CP) from day 13 to 46 (finisher period). Two groups (AMX1 and AMX2) were administered 20 mg of amoxicillin per kg per live weight per day for three consecutive days, once in the morning and once in the evening with an interval of 12 h, while two groups (THP1 and THP2) were administered with 57 mg of thiamphenicol per kg per live weight per day for three consecutive days. Chicks in the AMX1 and THP1 groups were treated from day 5 onwards, while those in the AMX2 and THP2 groups were treated from day 21 onwards. The control group did not receive any antimicrobial. On day 30 of age, birds of all groups started showing respiratory and intestinal symptoms; therefore, the farm veterinarian prescribed and administered 20 mg of doxycycline per kg of live weight a day for five consecutive days from day 33 onwards. All treatments were administered via drinking water. From each group, six birds were randomly selected and cloacal swabs taken on day 1 (T0). Birds in the AMX1, THP1 and control groups were swabbed on 1 d.p.t. (T1), 12 d.p.t. (T2) and 21 d.p.t. (T3). Cloacal swabs were collected from AMX2, THP2 and control groups on days 1 d.p.t. (T1) and 9 d.p.t. (T2). A total of 120 cloacal swabs were collected. Within one hour after sampling, swabs were placed at – 80 °C until DNA extraction. At the end of the rearing cycle (46 days of age) and after the regular withdrawal period and the complete remission of symptoms, birds were regularly slaughtered and caecum contents were aseptically collected and immediately frozen in liquid nitrogen.
DNA extraction
Microbial DNA was extracted from the swabs using a commercial kit (QIAamp UCP Pathogen Mini Kit, Qiagen, Germany), and from each swab, two aliquots were prepared to be used in the downstream analyses. Meanwhile, DNA was extracted from 250 mg of caecal content using the DNeasy PowerSoil DNA Isolation Kit (Qiagen, Hilden, Germany), following manufacturer’s recommendations. DNA quality and quantity were assessed using the Qubit 2.0 Fluorometer (Thermo Fisher Scientific, Monza, Italy).
16S rRNA gene sequencing and data analysis
NGS-based 16S rRNA was performed to identify and compare bacteria present in each sample. Briefly, the V3-V4 region of the 16S rRNA gene was amplified with primers 341F/R806 with overhangs adapters sequences33, and the second stage PCR was performed to add the IDT for Illumina Nextera DNA UD Indexes (IDT). All PCRs were carried out using KAPA HiFi HotStart ReadyMix (Roche) in a 2720 thermal cycler (Applied Biosystems, Waltham, MA), applying the amplification protocol 95 °C for 3 min (min), 25 or 8 cycles at 95 °C for 30 s (s), 55 °C for 30 s and 72 °C for 30 s, for the first and the second stage PCR, respectively. After both PCR stages, amplicons were purified using the SPRIselect purification kit (Beckman Coulter, Brea, CA). Libraries were quantified using Qubit 2.0 Fluorometer (Thermo Fisher Scientific), pooled and sequenced using the Illumina MiSeq sequencing platform (San Diego, California, USA) with a 2 × 300 bp paired-end approach. Within the Quantitative Insights into Microbial Ecology 2 (QIIME2 version 2019.4) software, the DADA2 package was used for 16S rRNA data analysis34,35. Taxa assignment was carried out using SILVA-Naive Bayes sklearn trained database36, while the on-line software Calypso (http://cgenome.net/wiki/index.php/Calypso) and Galaxy (https://galaxyproject.org/)37 were used for the statistical analysis of microbial communities. The microbial community composition was explored using heatmaps, while microbial diversity within each group (α-diversity) was expressed using Shannon and Simpson indexes. Differences in microbial composition among groups (β-diversity) were assessed using permutational multivariable analysis of variance (PERMANOVA) based on the Bray–Curtis dissimilar measure using the Adonis function and visualized with non-metric multidimensional scaling (NMDS). The linear discriminant analysis (LDA) effect size method (LEfSe) was used to identify taxa most likely to explain differences between groups at the same time point.
Quantitative PCR (qPCR) analysis of ARGs
Extracted DNA was screened by qPCR to detect and quantify 14 ARGs conferring resistance to β-lactams (i.e. blaTEM-1, blaSHV, blaCTX-M-1like, blaCMY-2, blaOXA-1, blaOXA-48, blaVIM-2 and blaNDM) and phenicols (i.e. catA1, catA2, catA3, catB Group3, floR and cmlA). Primers’ sequences, optimal concentrations, annealing and melting temperatures, and positive controls used are reported in Supplementary Table 1. All samples were tested for each gene in triplicate using PowerUp™ SYBR Green Master Mix (Thermo Fisher Scientific) with an optimal concentration of each primer in a LightCycler480 Roche (Roche, Basel, Switzerland) real-time platform and the mean values (copy number of each ARG) were estimated. However, since the absolute abundance of ARGs in a given sample is not a significant value, as it is proportional to the total DNA present in the sample, ARGs relative abundance was calculated by normalizing the ARG copies number to 16S rRNA gene copies, and used in the statistical analysis.
Statistical analysis
Differences in α-diversity or ARG abundance among groups (e.g. AMX1, THP1 and control) at the same time points (e.g. T0–T3) were tested for significance using generalized linear models (GLMs) with a log link and Gaussian error distribution (log-transformed Shannon index values) or gamma error distribution (ARG abundances). Only ARGs with ≥ 10% prevalence over all samples were included in the analysis. Hierarchical clustering was used to assess associations between presence/absence of each ARG and groups and time points. Differences in relative abundance of microbial taxa among the three groups were tested for significance using multivariate regression analysis with several dependent variables (i.e. log-transformed relative abundances of the microbial taxa). The same approach was used to assess associations between the relative abundance of the different taxa and ARG abundance. Also for these analyses, only taxa and ARGs present at a confident level of detection of 10% prevalence over samples were included. Furthermore, Spearman’s rank correlation was used to assess the co-occurrence of ARGs conferring resistance to β-lactams and phenicols. All models included cluster-robust (Sandwich) variance estimators to account for clustering of samples originating from the same chicks over time points. Statistical significance was set at p < 0.05. Statistical analysis and data visualization were carried out in R (version 4.2.1) (https://www.r-project.org/) using package Stats (version 4.1.2), Stata 16 (StataCorp, USA) and GraphPad Prism version 9.2.0 (https://www.graphpad.com).
Ethical statement
The zootechnical trial was approved by Ethics and Animal Welfare Committee of the Department of Veterinary Sciences (protocol n. 2796/2020), University of Turin (Italy). (https://www.veterinaria.unito.it/do/organi.pl/Show?_id=twsn). All international, national and/or institutional guidelines for care and use of animals were followed. The authors declare that the animal results of the study are reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Data availability
The datasets generated and/or analysed during the current study are available in the NCBI repository, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA872901, BioProject ID PRJNA872901.
References
Furtula, V. et al. Veterinary pharmaceuticals and antibiotic resistance of Escherichia coli isolates in poultry litter from commercial farms and controlled feeding trials. Poult. Sci. 89, 180–188 (2010).
Diarra, M. S. et al. Impact of feed supplementation with antimicrobial agents on growth performance of broiler chickens, Clostridium perfringens and Enterococcus counts, and antibiotic resistance phenotypes and distribution of antimicrobial resistance determinants in Escheric. Appl. Environ. Microbiol. 73, 6566–6576 (2007).
Gonzalez Ronquillo, M. & Angeles Hernandez, J. C. Antibiotic and synthetic growth promoters in animal diets: Review of impact and analytical methods. Food Control 72, 255–267 (2017).
Diarra, M. S. & Malouin, F. Antibiotics in Canadian poultry productions and anticipated alternatives. Front. Microbiol. 5, 1–15 (2014).
Apostolakos, I. et al. High-resolution characterisation of ESBL/pAmpC-producing Escherichia coli isolated from the broiler production pyramid. Sci. Rep. 10, 1–12 (2020).
Giacomelli, M. et al. Molecular characterization and genotypic antimicrobial resistance analysis of Campylobacter jejuni and Campylobacter coli isolated from broiler flocks in northern Italy. Avian Pathol. 41, 579–588 (2012).
World Health Organization. WHO | WHO list of Critically Important Antimicrobials (CIA). (2019).
Zapata, H. J. & Quagliarello, V. J. The microbiota and microbiomein aging: Potential implications in health and age-related diseases general aspects of thehuman microbiota and microbiome. Geriatr. Biosci. 63, 776–781 (2016).
Burow, E. et al. Antibiotic resistance in Escherichia coli from broiler chickens after amoxicillin treatment in an experimental environment. Microb. Drug Resist. 26, 1098–1107 (2020).
Liu, L. et al. Amoxicillin increased functional pathway genes and beta-lactam resistance genes by pathogens bloomed in intestinal microbiota using a simulator of the human intestinal microbial ecosystem. Front. Microbiol. 11, 1–14 (2020).
Tikhomirov, M., Poźniak, B., Smutkiewicz, A. & Świtała, M. Pharmacokinetics of florfenicol and thiamphenicol in ducks. J. Vet. Pharmacol. Ther. 42, 116–120 (2019).
Williams, C. T., Musicha, P., Feasey, N. A., Adams, E. R. & Edwards, T. ChloS-HRM, a novel assay to identify chloramphenicol-susceptible Escherichia coli and Klebsiella pneumoniae in Malawi. J. Antimicrob. Chemother. 74, 1212–1217 (2019).
Kent, A. G., Vill, A. C., Shi, Q., Satlin, M. J. & Brito, I. L. Genes within individual gut microbiomes revealed through bacterial Hi-C. Nat. Commun. https://doi.org/10.1038/s41467-020-18164-7 (2020).
Schokker, D. et al. Perturbation of microbiota in one-day old broiler chickens with antibiotic for 24 hours negatively affects intestinal immune development. BMC Genom. 18, 1–14 (2017).
Rychlik, I. Composition and function of chicken gut microbiota. Animals 10, 103 (2020).
Clavijo, V., Morales, T., Vives-Flores, M. J. & Reyes Muñoz, A. The gut microbiota of chickens in a commercial farm treated with a Salmonella phage cocktail. Sci. Rep. 12, 1–16 (2022).
Le Roy, C. I., Woodward, M. J., Ellis, R. J., La Ragione, R. M. & Claus, S. P. Antibiotic treatment triggers gut dysbiosis and modulates metabolism in a chicken model of gastro-intestinal infection. BMC Vet. Res. 15, 1–13 (2019).
Cuccato, M. et al. 16S Rrna sequencing analysis of the gut microbiota in broiler chickens prophylactically administered with antimicrobial agents. Antibiotics 10, 1–10 (2021).
Liu, J. et al. On-farm soil resistome is modified after treating dairy calves with the antibiotic florfenicol. Sci. Total Environ. 750, 141694 (2021).
Bischoff, K. M., White, D. G., Hume, M. E., Poole, T. L. & Nisbet, D. J. The chloramphenicol resistance gene cmlA is disseminated on transferable plasmids that confer multiple-drug resistance in swine Escherichia coli. FEMS Microbiol. Lett. 243, 285–291 (2005).
Meunier, D. et al. Plasmid-borne florfenicol and ceftiofur resistance encoded by the floR and blaCMY-2 genes in Escherichia coli isolates from diseased cattle in France. J. Med. Microbiol. 59, 467–471 (2010).
Schwarz, S., Kehrenberg, C., Doublet, B. & Cloeckaert, A. Molecular basis of bacterial resistance to chloramphenicol and florfenicol. FEMS Microbiol. Rev. 28, 519–542 (2004).
Campedelli, I. et al. Genus-wide assessment of antibiotic resistance in Lactobacillus spp. Appl. Environ. Microbiol. 85, 1–21 (2019).
Hummel, A. S., Hertel, C., Holzapfel, W. H. & Franz, C. M. A. P. Antibiotic resistances of starter and probiotic strains of lactic acid bacteria. Appl. Environ. Microbiol. 73, 730–739 (2007).
Su, H. et al. Occurrence and temporal variation of antibiotic resistance genes (ARGs) in shrimp aquaculture: ARGs dissemination from farming source to reared organisms. Sci. Total Environ. 607–608, 357–366 (2017).
Roberts, M. C. & Schwarz, S. Tetracycline and phenicol resistance genes and mechanisms: Importance for agriculture, the environment, and humans. J. Environ. Qual. 45, 576–592 (2016).
Graham, D. W., Knapp, C. W., Christensen, B. T., McCluskey, S. & Dolfing, J. Appearance of β-lactam resistance genes in agricultural soils and clinical isolates over the 20th century. Sci. Rep. 6, 1–8 (2016).
Laconi, A. et al. Microbial community composition and antimicrobial resistance in agricultural soils fertilized with livestock manure from conventional farming in Northern Italy. Sci. Total Environ. 760, 143404 (2021).
Arias, C. A., Contreras, G. A. & Murray, B. E. Management of multidrug-resistant enterococcal infections. Clin. Microbiol. Infect. 16, 555–562 (2010).
Hu, G. Z. et al. Phenotypic and molecular characterization of TEM-116 extended-spectrum β-lactamase produced by a Shigella flexneri clinical isolate from chickens. FEMS Microbiol. Lett. 279, 162–166 (2008).
Worley, J. N. et al. Genomic drivers of multidrug-resistant Shigella affecting. MBio 12, 1–12 (2021).
Marti, R. et al. Safely coupling livestock and crop production systems: How rapidly do antibiotic resistance genes dissipate in soil following a commercial application of swine or dairy manure?. Appl. Environ. Microbiol. 80, 3258–3265 (2014).
Takahashi, S., Tomita, J., Nishioka, K., Hisada, T. & Nishijima, M. Development of a prokaryotic universal primer for simultaneous analysis of Bacteria and Archaea using next-generation sequencing. PLoS ONE 9, e105592 (2014).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Yilmaz, P. et al. The SILVA and ‘all-species Living Tree Project (LTP)’ taxonomic frameworks. Nucleic Acids Res. 42, 643–648 (2014).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Acknowledgements
We would like to thank Dr. Giuditta Tilli and Dr. Lisa Carraro [Department of Comparative Biomedicine and Food Science, University of Padua, Legnaro (PD), Italy] for their support in planning the sampling scheme and extracting the DNA from the faecal samples, respectively. Authors are also grateful to Dr. Giovanni Perona and Dr. Stefano Nurisso [Department of Veterinary Science, University of Turin, Grugliasco (TO), Italy] for providing technical support during the conduction of the zootechnical trial; to Dr. Elena Pagani [Monge & C, Monasterolo di Savigliano (CN), Italy] for providing technical support with sampling procedures and to Dr. Sara Divari, Dr. Paola Pregel, Prof. Frine Eleonora Scaglione and Alessandra Sereno [Department of Veterinary Science, University of Turin, Grugliasco (TO), Italy] for their support in this research.
Author information
Authors and Affiliations
Contributions
A.L., A.P. and F.T.C. designed and conceived the study; A.L. and R.T. acquired and curated the data; A.L. and L.M.G. analysed the data; M.C. managed the animal and collected the samples; A.L., L.M.G. and A.P. wrote the manuscript. All the authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Laconi, A., Tolosi, R., Mughini-Gras, L. et al. Amoxicillin and thiamphenicol treatments may influence the co-selection of resistance genes in the chicken gut microbiota. Sci Rep 12, 20413 (2022). https://doi.org/10.1038/s41598-022-24927-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-24927-7
This article is cited by
-
Antimicrobial residues in meat from chickens in Northeast Vietnam: analytical validation and pilot study for sampling optimisation
Journal of Consumer Protection and Food Safety (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.