Abstract
Globally, since the outbreak of the Omicron variant in November 2021, the number of confirmed cases of COVID-19 has continued to increase, posing a tremendous challenge to the prevention and control of this infectious disease in many countries. The global daily confirmed cases of COVID-19 between November 1, 2021, and February 17, 2022, were used as a database for modeling, and the ARIMA, MLR, and Prophet models were developed and compared. The prediction performance was evaluated using mean absolute error (MAE), mean absolute percentage error (MAPE), and root mean square error (RMSE). The study showed that ARIMA (7, 1, 0) was the optimum model, and the MAE, MAPE, and RMSE values were lower than those of the MLR and Prophet models in terms of fitting performance and forecasting performance. The ARIMA model had superior prediction performance compared to the MLR and Prophet models. In real-world research, an appropriate prediction model should be selected based on the characteristics of the data and the sample size, which is essential for obtaining more accurate predictions of infectious disease incidence.
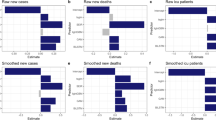
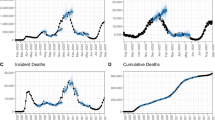
Similar content being viewed by others
Introduction
Since November 2021, the Omicron variant has rapidly spread worldwide. The B.1.1.529 variant was first reported by WHO in South Africa on November 24, 20211. The World Health Organization (WHO) announced the SARS-CoV-2 variant Omicron (B.1.1.529) on November 26, 20212,3. Consequently, many countries have enacted various restrictions to prevent the spread of Omicron variants.
Globally, as of February 14, 2022, a total of 416,614,051 confirmed cases of COVID-19 comprising 5,844,097 deaths were reported by the WHO4. It was estimated that the R0 of the Omicron variants may be as high as 105. Therefore, it is crucial that prediction models are used to forecast the COVID-19 epidemic trend, which can help the government and relevant authorities take effective measures to respond in advance6. Time series forecasting models play an important role in disease surveillance7. Accurate prediction results are required for the prevention and control of COVID-19 to provide early warning information to government officials.
Numerous mathematical models, including traditional time series and machine learning models, have been applied to predict the incidence of COVID-19. In particular, in the traditional time series model, the ARIMA time series model is the most widely used for COVID-19 incidence prediction. Ceylan et al.8 used the ARIMA model to estimate the overall prevalence of COVID-19 in three European countries, and the results can help politics and health authorities allocate medical resources reasonably. Sun et al.9 used a modified ARIMA model to forecast the COVID-19 pandemic in Alberta, Canada. Roy et al.10 analyzed the effectiveness of COVID-19 epidemiological surveillance using ARIMA models. Malki et al.11 applied the ARIMA model to predict the spread of COVID-19 worldwide. James et al.12 adopted the ARIMA model to forecast the short-term trajectory of the acceleration of fatalities caused by COVID-19. Dawoud et al.13 utilized the ARIMA model to estimate COVID-19 cumulative confirmed cases. Alzahrani et al.14 used the autoregressive model (AR), moving average (MA), a combination of both (ARMA), and integrated ARMA (ARIMA) to forecast the COVID-19 pandemic and found that the performance of the ARIMA model outperformed the other models.
In addition, the ARIMA model is used not only in the estimation of the number of COVID-19 pandemics, but also in the estimation of the number of fully vaccinated people or in the estimation of electricity consumption and natural gas amounts. Cihan et al.15 developed the ARIMA model to predict electricity and natural gas consumption in an industrial zone in Turkey. Cihan et al.16 used the ARIMA model to determine the number of people fully vaccinated against COVID-19.
However, some of the research has focused on the use of machine learning models to predict COVID-19 incidence, such as LSTM, GRU, SVR, XGBoost, RNNs, etc. Shahid et al.17 constructed the ARIMA, SVR, LSTM, and Bi-LSTM models to forecast COVID-19 confirmed cases, deaths, and recoveries in ten major countries, and stated that Bi-LSTM achieved much better prediction results than other models. Luo et al.18 established and compared the prediction performance of the LSTM and XGBoost algorithms. ArunKumar et al.19 developed GRU, LSTM, and RNN models to forecast future trends of the cumulative COVID-19 confirmed cases for the top-10 countries.
However, to date, no studies have compared global COVID-19 incidence predictions using ARIMA, MLR, and Prophet models since the outbreak of Omicron variants. In this study, the global daily confirmed cases of COVID-19 between November 1, 2021, and February 17, 2022, were obtained from the WHO website. Based on the sample size and data characteristics, ARIMA, MLR, and Prophet models were constructed and compared, and the optimum model was selected to predict the global daily confirmed cases of COVID-19 from February 18 to March 18, 2022. To the best of our knowledge, this is the first study to explore in detail the construction and comparison of the ARIMA, MLR, and Prophet models for predicting daily confirmed cases of COVID-19 worldwide. We hope that the prediction results of this study will serve as a reference for COVID-19 prevention and control worldwide.
Materials and methods
Materials
Data source
We collected daily confirmed cases of COVID-19 globally between November 1, 2021, and February 17, 2022, from the website of the World Health Organization (https://covid19.who.int/). Microsoft Excel was used to create the time series database. All data were updated daily. In this study, 109 observations were divided into training and validation sets, 80% of which was the training set, and the rest (20%) was the test set. The datasets for November 1, 2021, and January 27, 2022, were considered as the training set, and data from January 28, 2022, to February 17, 2022, were considered as the validation set.
Methods
ARIMA model
The autoregressive integrated moving average (ARIMA) model, a classic time series prediction technique, was proposed by Box and Jenkins in the early 1970s, and has been extensively applied to the prediction of infectious diseases20. ARIMA is a mathematical model that uses historical values to forecast future values of a variable21. The basic equation for ARIMA is as follows22:
In this equation, yt is the predictive value, B is the backward shift operator, εt is the residuals from time series23,\(\Theta_{P}\) and \(\theta_{p}\), \(\Phi_{Q}\), and \(\varphi_{q}\) represent the four parameters in the ARIMA model p, q, P, and Q, respectively. Here, d and D represent the degrees of the seasonal and trend differences, respectively. The ARIMA model parameters p, P, q, Q, and s represent the order of auto-regression, seasonal auto-regression lag, order of moving average, seasonal moving average, and seasonal periodicity, respectively24.
In general, the ARIMA model is defined as ARIMA(p, d, q) (P, D, Q)s. In this study, however, the ARIMA model was expressed as ARIMA(p, d, q) because the daily confirmed COVID-19 cases in the time series were non-seasonal data, and its equation can be written as follows23:
The construction process of the ARIMA model includes several steps25,26,27,28. First, the daily confirmed COVID-19 case sequence was plotted to determine whether the time series was stationary. Sequences with non-stationary time series were transformed into stationary sequences using difference and log transformations. Second, the parameters of the ARIMA model were estimated by analyzing auto-correlation and partial auto-correlation function graphs. The parameters p, P, q, and Q were determined using auto-correlation function (ACF) and partial auto-correlation function (PACF) graphs after difference and log transformations. The candidate ARIMA model was determined initially. Third, the ARIMA model diagnosis and evaluation were determined using the Ljung-Box (Q) test and the t-test, respectively. The Ljung-Box (Q) test required that residuals of the daily COVID-19 case time series were white noise (significant level, p > 0.05). A t-test was used to determine whether the parameters of each candidate ARIMA model were significant. The optimum model depends on the maximum R-square value, minimum normalized BIC, and RMSE values, and the residuals are white noise sequences. Bayesian information criterion (BIC) is commonly used for model selection in time series forecasting29. It was developed by Schwarz and is defined as29,30:
where L is the maximized value of the likelihood function of the model, n is the sample size, and k is the number of parameters estimated by the model. The normalized Bayesian information criterion (BIC) was used to confirm the adequacy of the model30. The smaller the value of the normalized BIC, the more adequate the model fits30.
MLR model
Multiple linear regression model(MLR), an extension of simple linear regression, is used to describe the a linear relationship between multiple independent variables and a single dependent variable31. The formula for the MLR model is as given below32.
where Y is the dependent variable; X1, X2, … are the independent variables; β is the Y-intercept; β0, β1,… βk are the regression coefficients; and ε is the random error term.
Prophet model
The Prophet model, an open-source time-series forecasting algorithm, was created by Facebook in 2017, and can be run using R or Python33. The basic formula for the Prophet model is as follows34,35:
Here, y(t) is the predictive value, g(t) is the trend function that models non-periodic changes in the time series of daily confirmed COVID-19 cases, s (t) signifies periodic changes(weekly characteristics of confirmed COVID-19 cases time series), and h (t) signifies the effects of holidays on potentially irregular schedules. For example, Christmas Day. \(\varepsilon_{t}\) signifies idiosyncratic changes that are not accommodated by the model36.
In trend model g (t), there are two types of models: a saturating growth model and a one-piece linear model that covers numerous Facebook applications. The formula for the nonlinear saturation growth model is as follows37:
where C is the carrying capacity, k is the growth rate, and m is the offset parameter.
The formula for the piecewise logistic growth model is as follows36:
where \(\delta\) is a vector of rate adjustments and \(\gamma\) is the correct adjustment at the change point.
The seasonality s(t) depends on the Fourier series to provide a viable model for periodic effects. This formula is expressed as follows34.
where a is standard Fourier series, P is the periodic changes.
Holidays and events h(t) have a greater influence on predicting time-series performance because they do not follow a periodic pattern37.
where t is during holiday i and ki is the holiday parameter and a prior k ~ normal (0, ν2).
Evaluation of the prediction performance
In this study, the mean absolute error (MAE), mean absolute percentage error (MAPE), and root mean square error (RMSE) were used to evaluate the prediction performances of the ARIMA, MLR, and Prophet models. The smaller the values of MAE, MAPE, and RMSE, the better is the prediction performance of the model. These evaluation indices are expressed as38:
where \({\hat{\text{X}}}_{t}\) is the predicted value, \(X_{t}\) is the observed value, and n is the sequence sample size.
Statistical software
SPSS (version 24.0; IBM Corp., Armonk, NY, USA, URL: https://www.ibm.com/support/pages/node/724325?mhsrc=ibmsearch_a&mhq=statistics%2024) and EView (version10.0; IHS Global Inc., Irvine, CA, USA, URL:https://eviews.com/download/ev10download.shtml) were used to create the ARIMA model. SPSS version 24.0 (version 24.0; IBM Corp., Armonk, NY, USA, URL: https://www.ibm.com/support/pages/node/724325?mhsrc=ibmsearch_a&mhq=statistics%2024) was used to create the MLR model. R software (version 4.1.1, URL:https://stat.ethz.ch/pipermail/r-announce/2021/000672.html) was used to construct the Prophet model. Among which, “Prophet” package of R software was used in construction of the Prophet model. The level of significance was set at p < 0.05.
Ethical approval
Data were obtained from publicly accessible sources. Formal ethical approval was not required for this study.
Results
General analysis
A total of 167,658,527 confirmed cases of COVID-19 were reported worldwide between November 1, 2021, and February 27, 2022. Descriptive Statistics of the daily confirmed cases of COVID-19 are shown in Table 1. The histogram chart of the daily confirmed cases of COVID-19 is shown in Fig. 1. As shown in Fig. 2, there was a rising periodicity trend characteristic of the daily confirmed cases of the COVID-19 time series. The growth rate of new confirmed coronary cases was 1.92% per day during this period. In addition, the confirmed cases occurred at a minimum peak on the first day and then reached a high peak two days later every other week with a cycle of 7 days (Fig. 2).
ARIMA model
The original sequence of the daily confirmed cases of the COVID-19 time series fluctuated greatly and presented an upward and periodic trend, indicating that this was a non-stationary time series (Fig. 2). Therefore, we used the first-order difference and natural logarithm transformation to convert the original sequence into a stationary time series; thus, parameter d was 1. The transformed time series presented random and stable characteristics (Fig. 3) and was a stationary time series. The ADF test also showed that the transformed time series was stationary ( t = − 9.247, p < 0.001).
The autocorrelation function (ACF) and partial autocorrelation function (PACF) graphs can help identify the p, q, P, and Q parameters of the ARIMA model. The candidate ARIMA models were constructed by combining the parameters p, q, P, and Q. From the analysis in Figs. 4, 5, we found that after a first-order difference and natural logarithm transformed time series displayed trailing and slower decaying convergence, the maximum was on the order of 7, which was significantly higher than orders 1 to 6; therefore, the parameter of p was 7, and q was in the range of 0 to 7. Therefore, the candidate ARIMA models are as follows: ARIMA (7,1,0), ARIMA (7,1,1), ARIMA (7,1,2), ARIMA (7,1,3), ARIMA (7,1,4), ARIMA (7,1,5), ARIMA (7,1,6), and ARIMA (7,1,7).
In addition, all candidate ARIMA models were tested using Ljung-Box Q for white noise. The results show that only three models passed the Ljung-Box Q test(p > 0.05): ARIMA(7,1,0), ARIMA(7,1,1), and ARIMA(7,1,2) (Table 2). The larger the R-squared value, the better is the fit of the ARIMA model. As shown in Table 2, the difference between the R squared values of the three models was not significant, indicating that the degree of the fitting effect was not different. Furthermore, we found that ARIMA(7,1,0) had the lowest RMSE and normalized BIC values and passed the t-test(p < 0.001), indicating that it was the optimum model (Table 3). Figure 6 shows that the residual ACF and PACF charts of ARIMA(7,1,0) are stationary time series, which also demonstrates that ARIMA(7,1,0) is the optimum model.
MLR model
The confirmed cases occurred at a minimum peak on the first day and then reached a high peak two days later every other week with a cycle of 7 days. Therefore, the every6th moment(day) might have affected the values at the latter moment(day). We used the data sliding method to set the input variables (X1-X6) and the independent variable(Y) and then constructed a multiple linear regression model. The R value of the MLR model was 0.949, indicating that the model fit well. The results of the F-test showed that the linear regression equation was significant (F = 144.08, p < 0.05). The MLR model equation that we fitted was \({\text{Y}} = 90416.43 + 0.4{\text{X}}_{1} - 0.1{\text{X}}_{2} + 0.02{\text{X}}_{3} + 0.01{\text{X}}_{4} - 0.16{\text{X}}_{5} + 0.71{\text{X}}_{6}\). The results are shown in Table 4.
Prophet model
A total of 109 observations are included in this section. In this study, the Prophet model that we constructed excluded holidays because of the differences in holidays between countries and the rapid spread of Omicron variants worldwide. As shown in Fig. 7, the global daily confirmed cases of the COVID-19 time series showed a fast-growing upward trend between November 2021 and February 2022. The day of the week curve shows that the global daily confirmed cases of the COVID-19 time series dropped to their lowest point on Tuesdays, quickly reached their highest point on Wednesdays, and then gradually increased.
Figure 8 showed the prediction performance of the Prophet model. In Fig. 8, the black dots represent the observed values, the blue line represents the predicted values, and the light-blue areas represent the 95% confidence intervals of the predicted values. As shown in Fig. 8, the predicted values were relatively stable within the 95% confidence interval from November 2021 to January 2022; however, after that, most of the predicted values fell outside the 95% confidence interval. This is due to the fact that the data this time is relatively stable and there are fewer outliers observations.
Comparison of ARIMA and Prophet models
As a first-order difference was conducted in the process of constructing the ARIMA (7,1,0) model and the data sliding method was carried out, only 102 observations were available to compare the predicted performances of the ARIMA, MLR, and Prophet models. The predicted and observed values fitted by the three models were used to calculate the MAE, MAPE, and RMSE, which were used to compare the predicted performances in this study. As shown in Table 5, the MAE, MAPE, and RMSE values of the ARIMA(7,1,0) model were lower than those of the MLR and Prophet models both in fitting performance and forecasting performance parts, indicating that the ARIMA(7,1,0) model has superior prediction performance and can be applied for the prediction of daily confirmed COVID-19 cases. The predicted value curve fitted by ARIMA(7,1,0) overlapped with the actual trend of COVID-19 incidence, indicating that ARIMA(7,1,0) was able to simulate the COVID-19 incidence well, and the prediction results were more accurate than those of the MLR and Prophet models (Fig. 9). Therefore, the ARIMA (7,1,0) model was used to perform an extrapolation to predict global daily confirmed COVID-19 cases from February 18 to March 18, 2022 (Table 6).
Discussion
Globally, with the rapid spread of the Omicron variant, the number of confirmed COVID-19 cases has continued to increase. Many countries are facing severe epidemic trends for this infectious disease. In this study, the global daily confirmed cases of COVID-19 between November 1, 2021, and February 17, 2022, were obtained from the World Health Organization website. The ARIMA, MLR,and Prophet models were applied to forecast the COVID-19 epidemic trends. Our findings showed that the ARIMA, MLR, and Prophet models could be applied to forecast daily confirmed COVID-19 cases; however, the ARIMA model had a superior prediction performance compared to the MLR and Prophet models.
According to the characteristics of the data, distribution, and sample size38, choosing a suitable model for daily confirmed COVID-19 cases is a prerequisite for obtaining more accurate prediction results. There were 109 observations in this study, and the sample sizes and data characteristics met the requirements for constructing the ARIMA, MLR, and Prophet models. Moreover, the ARIMA model is a classical time series prediction approach with several advantages for predicting the incidence of infectious diseases38. The major advantage of the ARIMA model is that it addresses linear problems that can reveal the dynamic laws between historical and predicted data20,39. The ARIMA model considers the trend, periodicity, and randomness of the time series, which can also quantify the expression by virtue of the model parameters40. Multiple linear regression models are widely used to predict the incidence of infectious diseases, and have the advantages of simple and fast modeling31,32. Rath et al.32 used a multiple linear regression model to forecast new active cases of the COVID-19 pandemic, and the model achieved remarkable accuracy in COVID-19 recognition.
The Prophet model, developed by Facebook in 2017, has been widely used in medicine41, environment35, and biology42 in recent years. Compared with traditional time-series forecasting models, the Prophet model has many advantages, such as its ability to consider trends, periodicity, special events, and outlier factors in the modelling process35, and its flexibility and simplicity of construction. In addition, the Prophet model has strong generalization capability and performs better in predicting the incidence of infectious diseases33. Xie et al.33 used the ARIMA and Prophet models to predict the incidence of HFMD, and the results showed that the prediction performance of the Prophet model was better than that of the ARIMA model. Tulshyan et al.41 used the Prophet models to forecast COVID-19 positive cases and fatalities in India over a 30-Day, the study showed that the Prophet model performs better in terms of accuracy with real data.
However, in this study, the MAE, MAPE, and RMSE values of the ARIMA(7,1,0) model were lower than those of the MLR and Prophet models both in fitting performance and forecasting performance parts. Our findings proved that the ARIMA model had superior prediction performance compared to the Prophet model, which was the opposite of their findings33,41. There are several possible explanations for this finding. First, the sample size of this study was 109, which met the requirements for modeling the ARIMA, MLR, and Prophet models. However, the prophet model is based on time series decomposition and machine learning fitting, which is more suitable for the long-term prediction of large samples and stabilized data34,35. Second, the MLR model has some disadvantages concerning its practical application43. For example, it tends to over fit when noisy data are used43. When outliers and influential observations are used to build MLR models, the accuracy of their predictions decreases. The prophet method was initially developed to address business-related issues44. Third, the time span of the data was from November 1, 2021, to February 17, 2022, which was the period of an outbreak of the Omicron variant. Therefore, the MLR and Prophet models were inferior to the ARIMA model in capturing short-term dramatic changes in the daily confirmed COVID-19 case sequences.
Therefore, we cannot apply predictive techniques blindly to real-world research. In general, data on the incidence of infectious diseases are characterized by linearity, seasonality, periodicity, and randomness40. Once the research data were obtained, the characteristics of the data and sample size were observed, and an appropriate predictive model was selected for the prediction. The traditional ARIMA time series forecasting model is well suited, particularly when the information on the research data is insufficient. It can rapidly predict infectious disease epidemics.
Our study has several limitations. First, the ARIMA model specializes in addressing linear problems12. However, the nonlinear part of a time series cannot be handled well27. Second, there may be the possibility of under-reporting of cases and deaths or delays in notifications, which may also lead to biased results. Third, the predicted values of the global daily confirmed COVID-19 cases from February 18 to March 18, 2022, all fell within the 95% confidence interval, indicating that there were no unexpected outbreaks of the Omicron variant during this period. However, the prevalence of COVID-19 is closely related to meteorological factors45, health care factors, and human mobility35. Therefore, in future studies, we need to consider the influential factors that affect the occurrence of COVID-19 in the modelling procedures and update the data continuously to obtain more accurate predictions.
Conclusions
In our study, we collected data on global daily confirmed cases of COVID-19 between November 1, 2021, and February 17, 2022, from the World Health Organization website. ARIMA, MLR, and Prophet models were constructed and compared. The study showed that the ARIMA model had superior prediction performance compared to the MLR and Prophet models. These prediction results can provide reference information for COVID-19 prevention and control worldwide.
Data availability
The data used or analyzed during the current study are available from the website of the World Health Organization(https://covid19.who.int/).
References
Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of concern. World health organization.https://www.who.int/news/item/26-11-2021-classification-of-omicron-(b.1.1.529)-sars-cov-2-variant-of-concern(2022).
Chen, J., Wang, R., Gilby, N. B. & Wei, G. W. Omicron (B.1.1.529): Infectivity, vaccine breakthrough, and antibody resistance. J. Chem. Inf. Model 62, 412–422. https://doi.org/10.1021/acs.jcim.1c01451 (2022).
He, X., Hong, W., Pan, X., Lu, G. & Wei, X. SARS-CoV-2 Omicron variant: Characteristics and prevention. MedComm. 2, 838–845. https://doi.org/10.1002/mco2.110 (2021).
WHO Coronavirus (COVID-19) Dashboard. World health organization. https://covid19.who.int/(2022).
Burki, T. K. Omicron variant and booster COVID-19 vaccines. Lancet Respir. Med. 10, 17. https://doi.org/10.1016/S2213-2600(21)00559-2 (2022).
Wang, G. et al. Comparison of ARIMA, ES, GRNN and ARIMA–GRNN hybrid models to forecast the second wave of COVID-19 in India and the United States. Epidemiol. Infect. 149, 240. https://doi.org/10.1017/S0950268821002375 (2021).
Wang, Y. W., Shen, Z. Z. & Jiang, Y. Comparison of autoregressive integrated moving average model and generalised regression neural network model for prediction of haemorrhagic fever with renal syndrome in China: A time-series study. BMJ open. 9, 025773. https://doi.org/10.1136/bmjopen-2018-025773 (2019).
Ceylan, Z. Estimation of COVID-19 prevalence in Italy, Spain, and France. Sci. Total Environ. 729, 38817. https://doi.org/10.1016/j.scitotenv.2020.138817 (2020).
Sun, Jian. Forecasting COVID-19 pandemic in Alberta, Canada using modified ARIMA models. Comput. Methods Programs Biomed. Update 1, 100029. https://doi.org/10.1016/j.cmpbup.2021.100029 (2021).
Roy, S., Bhunia, G. S. & Shit, P. K. Spatial prediction of COVID-19 epidemic using ARIMA techniques in India. Model Earth Syst. Environ. 7, 1385–1391. https://doi.org/10.1007/s40808-020-00890-y (2021).
Malki, Z. et al. ARIMA models for predicting the end of COVID-19 pandemic and the risk of second rebound. Neural Comput. Appl. 33, 2929–2948. https://doi.org/10.1007/s00521-020-05434-0 (2021).
James, A. & Tripathi, V. Time series data analysis and ARIMA modelling to forecast the short-term trajectory of the acceleration of fatalities in Brazil caused by the corona virus (COVID 19). PeerJ 9, 11748. https://doi.org/10.7717/peerj.11748 (2021).
Dawoud, I. Modelling Palestinian COVID-19 cumulative confirmed cases: A comparative study. Infect Dis. Model. 5, 748–754. https://doi.org/10.1016/j.idm.2020.09.001 (2020).
Alzahrani, S. I., Aljamaan, I. A. & Al-Fakih, E. A. Forecasting the spread of the COVID-19 pandemic in Saudi Arabia using ARIMA prediction model under current public health interventions. J. Infect. Public Health 13, 914–919. https://doi.org/10.1016/j.jiph.2020.06.001 (2020).
Cihan, P. Impact of the COVID-19 lockdowns on electricity and natural gas consumption in the different industrial zones and forecasting consumption amounts: Turkey case study. Int. J. Electr. Power Energy Syst. 134, 107369. https://doi.org/10.1016/j.ijepes.2021.107369 (2022).
Cihan, P. Forecasting fully vaccinated people against COVID-19 and examining future vaccination rate for herd immunity in the US, Asia, Europe, Africa, South America, and the World. Appl. Soft Comput. 111, 107708. https://doi.org/10.1016/j.asoc.2021.107708 (2021).
Shahid, F., Zameer, A. & Muneeb, M. Predictions for COVID-19 with deep learning models of LSTM GRU and Bi-LSTM. Chaos Solitons Fractals 140, 110212. https://doi.org/10.1016/j.chaos.2020.110212 (2020).
Luo, J., Zhang, Z., Fu, Y. & Rao, F. Time series prediction of COVID-19 transmission in America using LSTM and XGBoost algorithms. Results Phys. 27, 104462. https://doi.org/10.1016/j.rinp.2021.104462 (2021).
ArunKumar, K. E., Kalaga, D. V., Kumar, C., Kawaji, M. & Brenza, T. M. Forecasting of COVID-19 using deep layer recurrent neural networks (RNNs) with gated recurrent units (GRUs) and long short-term memory (LSTM) cells. Chaos Solitons Fractals 146, 110861. https://doi.org/10.1016/j.chaos.2021.110861 (2021).
Alim, M. et al. Comparison of ARIMA model and XGBoost model for prediction of human brucellosis in mainland China: A time-series study. BMJ open. 10, 039676. https://doi.org/10.1136/bmjopen-2020-039676 (2020).
Alabdulrazzaq, H. et al. On the accuracy of ARIMA based prediction of COVID-19 spread. Results Phys. 27, 104509. https://doi.org/10.1016/j.rinp.2021.104509 (2021).
Zhai, M. et al. Research on the predictive effect of a combined model of ARIMA and neural networks on human brucellosis in Shanxi Province, China: A time series predictive analysis. BMC Infect. Dis. 21, 280. https://doi.org/10.1186/s12879-021-05973-4 (2021).
Wang, Y., Xu, C., Wang, Z. & Yuan, J. Seasonality and trend prediction of scarlet fever incidence in mainland China from 2004 to 2018 using a hybrid SARIMA-NARX model. PeerJ 7, 6165. https://doi.org/10.7717/peerj.6165 (2019).
Wei, W. et al. Application of a combined model with autoregressive integrated moving average (ARIMA) and generalized regression neural network (GRNN) in forecasting hepatitis incidence in Heng county China. Plos One 11, e0156768. https://doi.org/10.1371/journal.pone.0156768 (2016).
Wang, H., Tian, C. W., Wang, W. M. & Luo, X. M. Time-series analysis of tuberculosis from 2005 to 2017 in China. Epidemiol. infect. 146, 935–939. https://doi.org/10.1017/S0950268818001115 (2018).
Ye, G. H. et al. Improving the precision of modelling the incidence of hemorrhagic fever with renal syndrome in mainland China with an ensemble machine learning approach. Plos One 16, e0248597. https://doi.org/10.1371/journal.pone.0248597 (2021).
Wu, W. et al. Comparison of two hybrid models for forecasting the incidence of hemorrhagic fever with renal syndrome in Jiangsu province China. Plos One 10, e0135492. https://doi.org/10.1371/journal.pone.0135492 (2015).
Tsan, Y. T. et al. The prediction of influenza-like illness and respiratory disease using LSTM and ARIMA. Int. J. Environ. Res. Public Health 19, 1858. https://doi.org/10.3390/ijerph19031858 (2022).
SchwarzG, E. Estimating the dimension of a model. Ann. Stat. 2, 461–464 (1978).
Clement, E. P. Using normalized bayesian information criterion (BIC) to improve box-jenkins model building. Am. J. Math. Stat. 5, 214–221 (2014).
Patil, S. & Pandya, S. Forecasting dengue hotspots associated with variation in meteorological parameters using regression and time series models. Front. Public Health 9, 798034. https://doi.org/10.3389/fpubh.2021.798034 (2021).
Rath, S., Tripathy, A. & Tripathy, A. R. Prediction of new active cases of coronavirus disease (COVID-19) pandemic using multiple linear regression model. Diabetes Metab. Syndr. 5, 1467–1474. https://doi.org/10.1016/j.dsx.2020.07.045 (2020).
Xie, C. et al. Trend analysis and forecast of daily reported incidence of hand, foot and mouth disease in Hubei China by Prophet model. Sci. Rep. 11, 1445. https://doi.org/10.1038/s41598-021-81100-2 (2021).
Liao, S., Yang, C. & Li, D. Improving precise point positioning performance based on prophet model. Plos One 16, 0245561. https://doi.org/10.1371/journal.pone.0245561 (2021).
Shen, J., Valagolam, D. & McCalla, S. Prophet forecasting model: A machine learning approach to predict the concentration of air pollutants (PM2.5, PM10, O3, NO2, SO2, CO) in Seoul South Korea. PeerJ. 8, 9961. https://doi.org/10.7717/peerj.9961 (2020).
Bashir, Tasarruf, Chen, H. Y., Tahir, Muhammad Faizan & Zhu, L. Q. Short term electricity load forecasting using hybrid prophet-LSTM model optimized by BPNN. Energy Rep. 8, 1678–1686. https://doi.org/10.1016/j.egyr.2021.12.067 (2022).
Taylor, S. J. & Letham, B. Forecasting at scale. Am. Stat. https://doi.org/10.7287/peerj.preprints.3190v (2017).
Zheng, Y. et al. Predictive analysis of the number of human brucellosis cases in Xinjiang. China. Sci. Rep. 11, 11513. https://doi.org/10.1038/s41598-021-91176-5 (2021).
Zhang, R. et al. Comparison of ARIMA and LSTM for prediction of hemorrhagic fever at different time scales in China. PloS One 17, e0262009. https://doi.org/10.1371/journal.pone.0262009 (2022).
Wang, L. et al. Epidemic Situation of brucellosis in Jinzhou city of China and prediction using the ARIMA Model. Can. J. Infect. Dis. Med. Microbiol. 2019, 1429462. https://doi.org/10.1155/2019/1429462 (2019).
Tulshyan, V., Sharma, D. & Mittal, M. An eye on the future of COVID-19: Prediction of likely positive cases and fatality in India over a 30-day horizon using the prophet model. Disaster Med. Public Health Prep. 18, 1–7 (2022).
Clark, N. J., Proboste, T., Weerasinghe, G. & Soares Magalhães, R. J. Nearterm forecasting of companion animal tick paralysis incidence: An iterative ensemble model. Plos Comput. Biol. 18, e1009874. https://doi.org/10.1371/journal.pcbi.1009874 (2022).
Salleh, F. H., Zainudin, S. & Arif, S. M. Multiple linear regression for reconstruction of gene regulatory networks in solving cascade error problems. Adv. Bioinform. 2017, 4827171. https://doi.org/10.1155/2017/4827171 (2017).
Hasan, M. N. et al. The global case-fatality rate of COVID-19 has been declining since may 2020. Am J Trop Med Hyg. 104, 2176–2184. https://doi.org/10.4269/ajtmh.20-1496 (2017).
Damette, O., Mathonnat, C. & Goutte, S. Meteorological factors against COVID-19 and the role of human mobility. Plos One 16, e0252405. https://doi.org/10.1371/journal.pone.0252405 (2021).
Acknowledgements
We thank the Sichuan Provincial Primary Health Service Development Research Center (grant no. SWFZ21-Q-59), Sichuan Provincial Orthopedics Hospital (grant no. 2021GL01), and Sichuan Provincial Hospital Management and Development Research Center (grant no. SCYG 2020-24) for funding this study.
Funding
This study was supported by the Sichuan Provincial Primary Health Service Development Research Center (Grant No. SWFZ21-Q-59), Sichuan Provincial Orthopedics Hospital (grant no. 2021GL01), and Sichuan Provincial Hospital Management and Development Research Center (Grant No. SCYG 2020–24).
Author information
Authors and Affiliations
Contributions
Z., D., Z., H., and Z., R. designed the study. Z., D., H., S. collected and analyzed the data. Z., D., H., S. wrote the first draft. Z., D., Z., H., and Z., R. reviewed and edited this manuscript. All authors contributed to the final draft and have approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, D., Zhang, R., Zhang, H. et al. Prediction of global omicron pandemic using ARIMA, MLR, and Prophet models. Sci Rep 12, 18138 (2022). https://doi.org/10.1038/s41598-022-23154-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23154-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.