Abstract
At the group level, antidepressant efficacy of rTMS targets is inversely related to their normative connectivity with subgenual anterior cingulate cortex (sgACC). Individualized connectivity may yield better targets, particularly in patients with neuropsychiatric disorders who may have aberrant connectivity. However, sgACC connectivity shows poor test–retest reliability at the individual level. Individualized resting-state network mapping (RSNM) can reliably map inter-individual variability in brain network organization. Thus, we sought to identify individualized RSNM-based rTMS targets that reliably target the sgACC connectivity profile. We used RSNM to identify network-based rTMS targets in 10 healthy controls and 13 individuals with traumatic brain injury-associated depression (TBI-D). These “RSNM targets” were compared with consensus structural targets and targets based on individualized anti-correlation with a group-mean-derived sgACC region (“sgACC-derived targets”). The TBI-D cohort was also randomized to receive active (n = 9) or sham (n = 4) rTMS to RSNM targets with 20 daily sessions of sequential high-frequency left-sided stimulation and low-frequency right-sided stimulation. We found that the group-mean sgACC connectivity profile was reliably estimated by individualized correlation with default mode network (DMN) and anti-correlation with dorsal attention network (DAN). Individualized RSNM targets were thus identified based on DAN anti-correlation and DMN correlation. These RSNM targets showed greater test–retest reliability than sgACC-derived targets. Counterintuitively, anti-correlation with the group-mean sgACC connectivity profile was also stronger and more reliable for RSNM-derived targets than for sgACC-derived targets. Improvement in depression after RSNM-targeted rTMS was predicted by target anti-correlation with the portions of sgACC. Active treatment also led to increased connectivity within and between the stimulation sites, the sgACC, and the DMN. Overall, these results suggest that RSNM may enable reliable individualized rTMS targeting, although further research is needed to determine whether this personalized approach can improve clinical outcomes.
Similar content being viewed by others
Introduction
The antidepressant efficacy of repetitive transcranial magnetic stimulation (rTMS) may be related to the connectivity of the stimulation target1,2. Scalp measurements or structural MRI are commonly used to identify a target in the dorsolateral prefrontal cortex (DLPFC)3, but recent studies have attempted to personalize rTMS targets based on functional connectivity (FC) with “seed” regions deeper in the brain4. At the group level, antidepressant efficacy of rTMS is related to normative anti-correlation between the stimulation site and the subgenual anterior cingulate cortex (sgACC), suggesting that treatment may be suppressing activity in sgACC and the limbic system5,6,7,8. Antidepressant efficacy is even more strongly related to normative connectivity between the stimulation site and a recently-derived “depression circuit” that closely overlaps with the dorsal attention network (DAN). To improve upon these group-level targets, individualized connectivity measurements have also been used to identify patient-specific stimulation sites based on connectivity to the sgACC9,10,11 or to the DAN12,13. These individualized targets may be more effective than the optimal normative targets7,8,14,15.
While the sgACC approach has been employed most commonly7,8,10,11,14,15,16,17, this approach is limited by the fact that sgACC connectivity is unreliable at the individual level9,11,15,18. Reliability assessments have shown weak test–retest correlation for sgACC connectivity to the DLPFC (spatial r < 0.5)9,15 and marked variability in DLPFC targets identified based on sgACC connectivity (mean test–retest variability of 25 mm)11. Targets can be identified more reliably based on connectivity to the “network” of regions most correlated with the sgACC at the group level9. This network may be personalized using individualized resting-state network mapping (RSNM), which can reliably map brain networks based on resting-state functional MRI (rsfMRI)19,20,21,22,23. RSNM has been successfully used for neurosurgical pre-operative mapping24 and has recently been evaluated as a method for mapping prefrontal topography to identify rTMS targets13.
RSNM enables precise individualized mapping of the DMN25, which is highly correlated with sgACC and the limbic system26,27. Individualized DMN mapping may thus serve as a more reliable proxy for sgACC connectivity. DMN is strongly anti-correlated with the DAN28,29,30, so individualized DAN mapping may yield a TMS target that is reliably anti-correlated to sgACC. Indeed, RSNM studies have found that DAN usually includes a node in the DLPFC. While the precise location of this node varies greatly between individuals31,32, rTMS targets have been identified at this node are reliable within individuals13. In a pilot clinical trial, stimulation of this node led to changes in sgACC connectivity with the DAN stimulation sites and with the DMN13. Furthermore, TMS sites, deep brain stimulation sites, and brain lesions connected to DAN have been causally linked to changes in depression severity33.
Of note, much of the existing knowledge about individualized brain mapping has been based on studies of healthy individuals. It remains unclear whether this generalizes to patients with neuropsychiatric illnesses and brain injuries. Inter-individual variability may be particularly prominent in traumatic brain injury-associated depression (TBI-D), which is associated with altered FC in the DLPFC, sgACC, DAN, and DMN34,35,36,37. This raises additional questions regarding the appropriateness of group-mean rTMS targets or individualized targets derived from seed-based connectivity.
As a first step to addressing these questions, we explored the differences between potential target sites generated using individualized RSNM (RSNM-based targets), standard anatomical methods (structural targets), and the point of maximal anti-correlation with the group-mean location of the sgACC (sgACC-derived targets). We also explored the connectivity changes induced by stimulation of RSNM-based targets in a recent pilot clinical trial13. We hypothesized that RSNM-based targets would approximate the sgACC connectivity profile more reliably than a group-based sgACC seed, which has previously been proposed for rsfMRI-based rTMS targeting, in both healthy controls and TBI-D patients7,9,11. After stimulation of these RSNM-based targets in TBI-D patients, we hypothesized that connectivity changes would be observed in the targeted networks, that these connectivity changes would covary with antidepressant response, and that antidepressant response would be predicted by baseline sgACC connectivity to the stimulation site.
Methods
Full methodological details are presented in the supplement.
Standard protocol approvals and participants
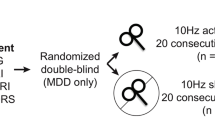
Data were collected as part of a pilot randomized clinical trial of rTMS for TBI-D13. Methods and hypotheses were pre-registered with the Open Science Foundation (osf.io/vjddq)38 and ClinicalTrials.gov (NCT02980484, posted 2 December 2016). The protocol was approved by the Human Research Protection Office at Washington University School of Medicine in St. Louis. All participants provided written informed consent and all procedures were carried out in accordance with relevant regulations. rTMS treatment parameters were within FDA guidelines. Participants were allocated via simple randomization to receive 20 daily sessions of active or sham rTMS with a 1:1 ratio using a random number generator, as described in our prior work. All randomization and enrollment procedures were conducted by the principal investigator (SHS). Participants and raters were blinded to the group allocation. Sham stimulation was applied using a device that produces a similar sound, but does not deliver active stimulation. Further details about the clinical trial are reported in Siddiqi et al.13.
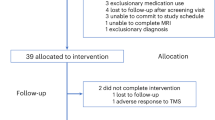
32 potential participants were screened for the study. 15 participants (11 males, ages 19–64) were included if they scored at least 10 on the Montgomery-Asberg Depression Rating Scale (MADRS) and had a history of at least one concussive or moderate TBI. One participant in the sham group was lost to follow-up before starting treatment. The remaining 14 participants completed the intent-to-treat endpoint (MADRS after 10 treatments). Two participants (one active, one sham) missed multiple treatments due to headaches, and were thus unable to complete the full treatment course within the pre-specified 5-weeek timeframe; however, both completed a MADRS and MRI for intent-to-treat analysis. One participant in the sham group chose not to complete a post-treatment MRI scan. This analysis was limited to 13 subjects (10 males) who completed both the pre-treatment and post-treatment scan sessions. Of note, the clinical trial was terminated prior to the first interim analysis (planned at n = 20) because the investigators left the institution for unrelated reasons.
10 healthy control subjects (3 males, ages 22–35) with no history of neuropsychiatric disease were chosen randomly from the Human Connectome Project (HCP) database39. Only 10 subjects were chosen in order to approximately match the number of TBI-D patients and to confirm that utility of individualized RSNM can be demonstrated with small sample sizes; clinical practicality of personalized medicine approaches may be questionable if large sample sizes are required to demonstrate their utility. We did not attempt to match these participants to the TBI-D sample because (a) HCP participants are all under age 35, and (b) no statistical comparisons were made between the two samples, since any matching attempts would be limited by the fact that the datasets were collected in different settings.
MRI acquisition and pre-processing
For TBI-D subjects, a 3 T Siemens Magnetom Prisma magnetic resonance scanner was used to acquire 16.5 min of resting-state blood oxygen-level dependent (BOLD) data in three runs. The baseline MRI scan was acquired at least one day before treatment to provide time for preprocessing and target selection. The post-treatment MRI scan was acquired at least one day after the conclusion of treatment to avoid bias from acute effects of treatment. For HCP subjects, a 3 T Siemens Connectome Skyra was used to acquire 58 min of resting-state BOLD data in four runs. Preprocessing was conducted using in-house scripts described in Power et al.40. For each subject, BOLD time courses were used to construct seven individual-level RSN maps via a multilayer perceptron (MLP)-based machine learning classifier as described in Hacker et al.21. To create individualized regions of interest for further analyses, a winner-take-all map was created by assigning each voxel to the network with maximum likelihood of membership. Further details are described in the supplement.
Confirmation of candidate network targets
Normative connectivity data
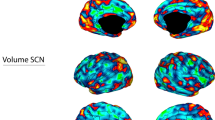
Data from the HCP 800-subject release39 were used to construct normative maps of resting-state functional connectivity with the sgACC (Fig. 1a), as described in Weigand et al.6. We hypothesized that the individualized map of the DAN would best resemble the normative map of sgACC anti-correlations. We also hypothesized that the individualized map of the DMN would best resemble the normative map of sgACC positive correlations. To initially evaluate these hypotheses, we quantified the degree of overlap between our group-based and the individualized RSN maps for each subject (Fig. 1b). All subsequent analyses were conducted using subject-specific connectivity data rather than group connectome data.
(a) Normative map of sgACC functional connectivity, indicating strongest areas of correlation (orange-yellow), and anti-correlation (blue). Strong normative sgACC anti-correlation is prominent at a DLPFC site (red arrow) which has previously been shown to be an effective rTMS target in major depression5,6. (b) Individualized winner-take-all maps of DAN (blue) and DMN (yellow) for two representative example subjects. Red arrows depict the group-mean stimulation site, which shows differing spatial relationships with DAN in the two patients (c) Example of overlap between individualized RSNM DAN/DMN maps (from patient 2) and normative sgACC seed map.
Comparison of individualized RSN maps to sgACC seed maps
For each subject, we quantified the degree to which each of the seven individualized RSN maps overlapped with the normative sgACC seed map. The sgACC seed map was masked with each individual RSN map to identify its overlap with that network (Fig. 1c). This yielded a map of normative sgACC connectivity values at each voxel within each RSN. Using this map, the mean normative sgACC connectivity value of overlapping voxels was calculated for each RSN and each subject. This yielded a single metric of the degree of overlap between the continuous normative sgACC seed map and each binary RSN map. This value was compared between DAN/DMN and the remaining RSNs by calculating Fisher’s least significant difference via one-way ANOVA.
rTMS target selection and comparison
Specific analytical procedures/tools are described in the supplement.
Target selection
Three approaches were used to identify potential rTMS targets:
-
1.
Individualized RSNM-based targeting—The individualized DMN map was subtracted from the individualized DAN map for each subject. The peak DLPFC cluster was identified in this map following the methods described in Siddiqi et al. (Fig. S1)12.
-
2.
Structural MRI-based targeting—Targets were chosen at DLPFC coordinates (± 38, 44, 26), which were originally derived based on normative sgACC connectivity5 and have been used for targeting at the world’s current largest neuronavigated rTMS clinic41.
-
3.
Individualized sgACC-derived target—This method relies on an individual subject’s anti-correlation with group-mean sgACC coordinates, as described in Fox et al.9 and implemented in Cash et al., 202014 and Siddiqi et al.15. The sgACC ROI was defined at the group level as a sphere centered at the coordinates (6, 16, − 10)9.
Comparison of resting state functional connectivity of the potential targets
For each potential stimulation site, resting-state functional connectivity was calculated with a population-derived definition of DAN and DMN42. To confirm that effects were not biased by the fact that RSNM-based DAN/DMN parcels would be expected to show high correlation to the consensus group-mean DAN/DMN parcels, connectivity was also calculated with the normative sgACC seed map. If effects were driven by this source of bias, then the normative sgACC seed map would be most anti-correlated with the sgACC-derived targets.
Potential target correlations with the DAN, DMN, and the sgACC seed map were compared between the different targeting methods across all subjects via within-subjects two-way ANOVA. Results from the two groups of subjects (TBI-D and healthy controls) were not compared with one another due to potential influence of methodological variability and demographic differences.
Comparison of spatial locations of the potential targets
For each subject, the three potential targets were also compared in terms of spatial distance between one another. The mean distances of RSNM targets and sgACC-derived targets from the structural target were compared using paired t-tests. Inter-individual variances for RSNM targets and sgACC-derived targets were determined using F-tests based on the distance of each target from the mean of all coordinates generated by that method.
rTMS treatment
To explore the effects of stimulating our proposed targets, TBI-D subjects were randomized to receive 20 daily sessions of active or sham rTMS using the RSNM targets. The clinical trial protocol, results, and CONSORT checklist are detailed in Siddiqi et al.13. Briefly, clinical treatment included 4000 pulses of high-frequency (10 Hz) left-sided stimulation, followed by 1000 pulses of low-frequency (1 Hz) right-sided stimulation. Using a Brainsight neuronavigation device, target coordinates were plotted on a surface reconstruction of each subject’s brain. No stimulation at other targets was performed in the current study.
Treatment-induced changes
Detailed analysis parameters are described in the supplement.
Target stability over time
Nine TBI-D subjects were randomized to active treatment and four were randomized to sham. For each of the three targeting methods, connectivity with the normative sgACC seed map was calculated for pre-treatment and post-treatment scans. Two-way ANOVA was used to compare the three targeting methods in terms of difference in connectivity between the two time points. Again, it should be noted that only individualized RSNM-based targeting was performed.
Euclidean distances between pre-treatment and post-treatment targets were calculated for RSNM and sgACC-derived targets to assess the stability of target location. Due to non-normal distribution of these distances, a Wilcoxon matched-pairs signed-rank test was used to compare the two targeting methods in terms of stability over time.
Exploratory evaluation of treatment-induced connectivity changes
To explore how the target RSNs were affected by rTMS, active and sham groups were compared in terms of treatment-induced change in connectivity. Connectivity was calculated between five a priori ROIs defined in the original clinical trial, including left/right stimulation sites, DAN, DMN, and sgACC. This analysis was conducted using covariance rather than correlation, since covariance is less sensitive to the potential influence of changing amplitudes of BOLD fluctuations between different time points43.
To explore how the rest of the brain was affected by rTMS, ROI-ROI connectivity was calculated with each of the 17 Yeo networks and voxel-wise connectivity was calculated with the whole brain. Active and sham groups were compared using a general linear model (GLM) with group assignment as the primary predictor, post-treatment connectivity as the outcome, and pre-treatment connectivity as a covariate. Except where required for voxel-wise multiple comparisons correction, statistical hypothesis testing was not conducted for active-sham comparisons because the trial did not reach its original target sample size13.
Prediction of antidepressant response
To examine connectivity-based predictors of response in the active treatment group, whole-brain connectivity of each stimulation site was compared with antidepressant response. For each voxel, a least squares regression model was constructed using baseline target-voxel connectivity and baseline MADRS as predictors of post-treatment MADRS. Because antidepressant response could not be assumed to be normally distributed in this small sample, all data were rank-transformed, which is consistent with prior methods described in Weigand et al.6.
Results
Confirmation of candidate network targets
In both groups, the positive correlations in the normative sgACC seed map (Fig. 1a, yellow/orange regions) showed stronger overlap with the individual DMN map than with any other individualized network map (Fig. 2a), as quantified by the mean voxel value of the sgACC seed map that fell within the boundaries of each network. The anti-correlations in the sgACC seed map (Fig. 1a, blue regions) showed stronger overlap with the individualized DAN map than with any other network map (Fig. 2a). In comparison with the other individualized networks in the TBI-D group, DAN showed significantly stronger overlap with the negative component of the sgACC seed map, while DMN showed significantly stronger overlap with the positive component of the sgACC seed map (Fig. 2b). The same trend was evident in the HCP group, except that the DAN-ventral attention network (VAN) difference and the DAN-frontoparietal control network (FPC) difference did not reach significance (Fig. 2b). Overall, DAN anti-correlation and DMN correlation provided the best individualized approximation of the sgACC seed map.
(a) Individualized similarity between each RSN and the normative sgACC seed map at baseline. The group-mean sgACC seed map positive correlations overlapped more with DMN and the anti-correlations overlapped more with DAN than any other individualized RSN map for the majority of individual subjects. (b) Mean similarity between individualized RSNs and the normative sgACC seed map. DMN was the only network showing strong overlap with the positive correlations in the sgACC seed map. Several RSNs showed notable overlap with the anti-correlations in the sgACC seed map.
Evaluation of expected stimulation profile for each potential target
Nearly all potential targets showed positive correlation with the DAN, anti-correlation with the DMN, and anti-correlations with the group mean sgACC seed map (Fig. 3a). Within-subjects two-way ANOVA revealed a significant effect of potential targeting method on left- and right-sided target connectivity with each of these RSNs in each of the two datasets (Table S1). The magnitude of these differences was similar but not identical between the two datasets (Fig. 3b and Table S1). As expected, in comparison with each of the other two targets, the RSNM-based target showed stronger connectivity to group-mean DAN and anti-correlation to group-mean DMN in 11/13 TBI-D patients and 8/10 healthy controls (p = 0.002, single-proportion z-test with expected proportion of 50%). Overall, RSNM-based targets appeared to provide a better individualized approximation of the desired networks than the other two potential targeting methods.
Functional connectivity of targets yielded by the three approaches. (a) Functional connectivity of RSNM, sgACC-derived, and structural targets with DAN, DMN, and the normative sgACC connectivity map in each group. (b) Differences in connectivity profiles between the three potential targeting methods. On most metrics, RSNM targets showed significantly stronger connectivity with all three regions of interest.
Spatial distribution of derived targets
In both groups, RSNM-based target coordinates were spatially distinct from both comparator targets (described in Section “Target selection”) with 95% confidence intervals that were greater than zero (Fig. 4a,b). The structural target was significantly closer to the RSNM target than to the sgACC-derived target in both the TBI-D (p = 0.006) and HCP (p = 4 × 10–5) groups. The sgACC-derived targets also showed wider variance between subjects than the RSNM targets for both the TBI-D (F = 0.4, p = 0.01) and HCP (F = 0.4, p = 0.03) groups (Table S2). The anatomical locations of targets generated by the different methods are depicted in Fig. 4b along with an example of approximate predicted stimulation volumes for each target in one representative subject. Thus, RSNM targets were less variable anatomically than sgACC-derived targets.
Anatomical distributions of targets yielded by the three approaches. Targets are displayed on a 3D surface rendering to facilitate visual comparison; the analyses were conducted in volume space. (a) Mean and 95% confidence interval for the Euclidean distance between target coordinates generated by the different methods. (b) 3D scatter plot of the target sites generated by the different methods to illustrate approximate spatial distribution of targets. Background is a representative example of a single-subject surface reconstruction with approximate predicted stimulation volumes in that subject (in the TBI depression group). These approximate stimulation volumes are cortical surface projections of the estimated 15-mm sphere centered at the stimulation site for that subject; shapes are asymmetric and irregular due to normal variation in cortical surface anatomy. Across all subjects and in this representative example, RSNM-based actual stimulation sites were different from the structural group-mean site (green dot).
Stability of connectivity and target location before vs. after RSNM targeted rTMS treatment in TBI-D patients
13 TBI depression patients were scanned again after a full course of active rTMS (n = 9) or sham (n = 4), which was applied to the RSNM target derived from the pre-TMS scan. Connectivity with the normative sgACC seed map remained relatively stable for the RSNM target and the structural target (Fig. 5a). sgACC-derived targets, by contrast, showed significantly different connectivity profiles between pre-treatment and post-treatment scans (p = 0.03). These results were unchanged when repeating the analysis after controlling for active versus sham stimulation (p = 0.03), and there was no significant effect of treatment group (p = 0.59).
(a) Change in connectivity profile of left- and right-sided potential targets identified based on pre-treatment and post-treatment scans. Group mean sgACC seed map connectivity remained relatively stable for RSNM targets and structural targets but was more unstable for the potential sgACC-derived targets. Each symbol represents 1 target; there were 2 targets per subject (right and left) × 13 subjects who underwent either active or sham rTMS treatment. (b) Spatial change in target coordinates between pre-treatment and post-treatment scans. After a course of RSNM-targeted treatment, the location of the potential sgACC-derived target sites changed significantly more than the location of RSNM-based target sites (p < 0.0001, Wilcoxon matched-pairs signed-rank test). (c) Treatment-induced change in connectivity within and between a priori ROIs. Active treatment was associated with connectivity changes within and between stimulation sites, sgACC, DAN, and DMN. Magnitude of change is quantified using Cohen’s d (adapted with permission from Siddiqi et al., J Neurotrauma 2019).
Between the two scan sessions (pre- and post-treatment), the mean absolute Euclidean distance change in target coordinates was 6.6 mm for RSNM targets and 17.7 mm for sgACC-derived targets (Wilcoxon matched-pairs signed rank test p < 10–4, Fig. 5b). These results did not differ when repeating the analysis after controlling for active versus sham stimulation (p < 10–3), and there was no significant effect of treatment group (p = 0.14).
Thus, after active or sham rTMS at the RSNM target, the location of the RSNM targets remained more stable than the sgACC-derived targets. Consistency of connectivity was similar between RSNM targets and structural targets. Of note, we did not directly assess stability of the location of the sgACC-derived targets before and after stimulation at these targets because no stimulation was performed at sgACC-derived targets.
Target engagement: treatment-induced change in connectivity
In comparison with sham, active rTMS was associated with large connectivity changes in several of the a priori ROI pairs, including DMN to sgACC, Left to Right stimulation site, and Left stimulation site to sgACC. There were also large changes in within-ROI connectivity in both stimulation sites and sgACC (Fig. 5c). Pre-treatment and post-treatment connectivity of each stimulation site with each of the 17 Yeo networks is depicted in Fig. S2.
Results and statistical methods for exploratory analyses are detailed in the supplement. First, treatment-induced connectivity change was compared between active and sham groups. Partial Spearman correlation was computed between group and post-treatment connectivity after controlling for pre-treatment connectivity (Fig. S3). For the right stimulation site, there was a decrease in FC with the cingulo-opercular network parcel (rho = − 0.56) and the parieto-occipital DAN parcel (rho = − 0.55), increase in FC with the parahippocampal/retrosplenial DMN parcel (rho = 0.65) (Fig. S3a), and increase in FC with a voxel cluster in the left ventral hippocampus (r > 0.7, corrected p < 0.05) (Fig. S3b). For the left stimulation site, active versus sham treatment led to a trend towards decreased parieto-occipital DAN connectivity and increased prefrontal/parietal DMN connectivity (Fig. S3c), as well as a decrease in FC with a voxel cluster in the dorsomedial prefrontal cortex (r > 0.7, corrected p < 0.05) (Fig. S3d).
Baseline predictors of clinical efficacy
For both stimulation sites, rTMS treatment efficacy was related to baseline connectivity of the stimulation site. For the right stimulation site, antidepressant response was significantly predicted by baseline anti-correlation with bilateral sgACC, anti-correlation with motor cortex, and positive correlation with dorsal ACC (corrected p < 0.05) (Fig. 6a). For the left stimulation site, antidepressant response to rTMS was predicted by baseline correlation with right precuneus and anti-correlation with right sgACC, bilateral lateral parietal lobe, and bilateral dorsomedial prefrontal regions traditionally associated with the resting-state salience network (corrected p < 0.05) (Fig. 6b). Permutation testing confirmed that this whole-brain map was stronger than expected by chance for the right stimulation site (p = 0.04), but not the left stimulation site (p = 0.27).
Connectivity profile associated with increased antidepressant efficacy of stimulation sites. White regions depict the a priori subgenual ROI. Clusters detected with threshold of r > 0.8 (uncorrected p < 0.001), minimum extent of 729 mm3, and cluster significance defined at p < 0.05. (a) Antidepressant response was significantly predicted by right stimulation site anti-correlation with bilateral sgACC, anti-correlation with motor cortex, and positive correlation with dorsal ACC (corrected p < 0.05). (b) Antidepressant response was significantly predicted by left stimulation site correlation with right precuneus and anti-correlation with right sgACC, bilateral lateral parietal lobe, and bilateral dorsomedial prefrontal cortex (corrected p < 0.05). (c, d) Antidepressant response was inversely related to seed-based connectivity of both stimulation sites with contralateral subgenual regions defined by a more recent cortical parcellation. Left panels depict the relationship between predicted and actual MADRS improvement, while right panels depict the overlap between the corresponding ROI (red) and the voxels whose stimulation site anti-correlation predicts MADRS improvement (blue). MNI coordinates of the center of each ROI are reported in maroon. (c) Right stimulation site connectivity with the a priori subgenual ROI (predicted MADRS change) was significantly predictive of antidepressant response (rho = 0.70, p = 0.035). This relationship was stronger when using exploratory subgenual ROIs based on a more recent cortical parcellation (rho = 0.90 and 0.93, p = 0.0009 and 0.0003). (d) Left stimulation site connectivity with the a priori subgenual ROI (predicted MADRS change) appeared to predict antidepressant response, but this relationship did not reach significance (rho = 0.54, p = 0.13). One of the two exploratory subgenual ROIs was significantly predictive of antidepressant response (rho = 0.90, p = 0.0009).
Antidepressant response was inversely correlated with the right-sided stimulation site’s FC with the a priori subgenual ROI (Fig. 6c, top panel; Spearman rho = 0.70, p = 0.03). The left-sided stimulation site showed a trend in the same direction, but did not reach significance (Fig. 6d, top panel).
The voxel-wise maps of connections that predicted antidepressant response (Fig. 6a,b) suggested that this effect was more prominent for specific subgenual regions that were only partially overlapping with the predefined sgACC ROI (Fig. 6c,d, bottom two panels). To explore this further, the ROI-based analysis was repeated post hoc using recent sub-classified sgACC parcels44. Stimulation site connectivity with contralateral sgACC regions revealed strong predictive value for post-treatment MADRS (right stimulation site: rho = 0.90 and 0.93, p = 0.001 and 0.0003; left stimulation site: rho = 0.90 and 0.54, p = 0.001 and 0.13, respectively). Treatment efficacy was thus predictable using baseline stimulation site connectivity.
Discussion
Our findings suggest that individualized RSNM may be used to reliably identify rTMS targets based on their connectivity profile. We identified subject-specific rTMS targets at the networks that are likely being approximated by sgACC connectivity maps, which have previously been shown to predict efficacy of rTMS for major depression6. These target coordinates were stable and spatially distinct from prior approaches. Furthermore, these individualized RSNM-based targets showed stronger functional connectivity with the intended network targets than other candidate rTMS targets, even when these networks are defined conservatively based on consensus group-mean maps. Furthermore, the RSNM-based targets approximated the sgACC connectivity map more effectively than individualized targets generated using the widespread sgACC-derived approach. While it appears counter-intuitive that sgACC-based targets were less connected with a map generated using sgACC as a seed, this may be because the sgACC-derived approach is unreliable. This is consistent with our hypothesis that our RSNM-based approach would identify a target that approximates the sgACC connectivity profile more effectively than a group-based sgACC seed.
Our proposed targeting approach was based on individualized mapping of DAN and DMN. The involvement of these networks in depression treatment may be related to dysfunctional interactions between externally-oriented attention-switching, which involves the DAN, and internally-oriented emotion engagement, which involves the DMN45. Such interactions appear to be affected in major depression46 and are modulated by deep brain stimulation of the sgACC47. This is consistent with our finding that antidepressant response was predicted by stimulation site connectivity with a large subgenual region. Treatment was also associated with changes in subgenual connectivity to itself, to the left DAN stimulation site, and to the DMN. This further suggests that our targeting approach may identify a network that modulates subgenual connectivity, even though it is not directly based on subgenual connectivity.
Nevertheless, our choice of DAN and anti-DMN targeting remains speculative in the absence of a head-to-head trial of antidepressant efficacy in comparison to rTMS applied to other targets. In addition, there are several existing approaches to individualized RSN mapping6,16,17 and we did not assess which approach (including group-level mapping) best predicts neurophysiological and clinical response. There are also several approaches to resting-state fMRI pre-processing; for instance, our use of global signal regression may affect the identification of anti-correlated networks48. Similarly, there are several approaches to TMS-induced electric field modeling, but we chose not to use individualized finite element modeling because this method has not yet been validated for functional connectivity analyses. Careful validation of these techniques may help to further optimize our methods. Finally, there are some emerging approaches to improve reliability of sgACC-derived targets, such as a “searchlight approach,” which simulates multiple potential targets and chooses the one whose connectivity profile is closest to the subgenual seed map49. Our approach is compatible with these types of approaches, as it would simply require substitution of the subgenual seed map with the individualized DAN-DMN map.
Our interpretations of treatment-induced changes are limited by small sample size. This may increase the risk of a false negative result due to lack of power or false positive results due to chance. Furthermore, we only assessed bilateral stimulation, and the interactions between the two stimulation sites are uncertain. This does not affect our reliability assessments, but does limit our ability to confirm whether the neurophysiological and clinical effects are consistent with our hypotheses. Prospective studies comparing unilateral stimulation vs. bilateral stimulation will be required to disentangle the neurophysiological effects of the bilateral stimulation employed in this study. It is not known whether the approach to selecting a left excitatory stimulation site should be the same as the approach used to selecting a right inhibitory stimulation site, since stimulation of the two hemispheres may have different effects50.
Further research will be required before these findings can be considered generalizable. This study was conducted using cutting-edge MRI scanners and recently-optimized scan protocols, so it remains unclear whether similar results can be achieved using more readily-available equipment. The patient population was also carefully selected as patients with relatively mild TBI and clear major depressive symptoms; its applicability to primary major depression or moderate/severe TBI requires further investigation.
Despite these limitations, our results support the emerging notion that variability in effects of rTMS may be related to inter-individual variability in functional topography of the DLPFC1,51,52. While the clinical implications of individualized RSN-based targeting are not yet clear, this method yields targets that are consistently connected to regions that have been implicated in antidepressant response to rTMS, including the sgACC. Stimulation of these targets also appears to modulate these key regions in a manner that is related to antidepressant response. This should help to inform an alternative and possibly more rational approach to prospective individualized target selection in future rTMS studies as well as retrospective analysis of results from existing studies.
In conclusion, the use of individualized RSN mapping for identification of distinct patient-specific rTMS targets may represent a promising method for reducing variability in targeting rTMS. This lays the foundation for development of more robust approaches for personalized medicine in neuromodulation.
References
Luber, B. M. et al. Using neuroimaging to individualize TMS treatment for depression: Toward a new paradigm for imaging-guided intervention. Neuroimage 148, 1–7. https://doi.org/10.1016/j.neuroimage.2016.12.0831 (2017).
Siddiqi, S. H., Kording, K. P., Parvizi, J. & Fox, M. D. Causal mapping of human brain function. Nat. Rev. Neurosci. 23, 361–375 (2022).
Fitzgerald, P. B. et al. A randomized trial of rTMS targeted with MRI based neuro-navigation in treatment-resistant depression. Neuropsychopharmacology 34(5), 1255–1262. https://doi.org/10.1038/npp.2008.233 (2009).
Fox, M. D. et al. Resting-state networks link invasive and noninvasive brain stimulation across diverse psychiatric and neurological diseases. Proc. Natl. Acad. Sci. USA. 111(41), E4367–E4375. https://doi.org/10.1073/pnas.1405003111 (2014).
Fox, M. D., Buckner, R. L., White, M. P., Greicius, M. D. & Pascual-Leone, A. Efficacy of transcranial magnetic stimulation targets for depression is related to intrinsic functional connectivity with the subgenual cingulate. Biol. Psychiatry. 72(7), 595–603. https://doi.org/10.1016/j.biopsych.2012.04.028 (2012).
Weigand, A. et al. Prospective validation that subgenual connectivity predicts antidepressant efficacy of transcranial magnetic stimulation sites. Biol. Psychiatry. 84(1), 28–37. https://doi.org/10.1016/j.biopsych.2017.10.028 (2018).
Cash, R. F. H. et al. Subgenual functional connectivity predicts antidepressant treatment response to transcranial magnetic stimulation: Independent validation and evaluation of personalization. Biol. Psychiatry. 86(2), e5–e7. https://doi.org/10.1016/j.biopsych.2018.12.002 (2019).
Cash, R. F. H. et al. Using brain imaging to improve spatial targeting of transcranial magnetic stimulation for depression. Biol. Psychiatry https://doi.org/10.1016/j.biopsych.2020.05.033 (2020).
Fox, M. D., Liu, H. & Pascual-Leone, A. Identification of reproducible individualized targets for treatment of depression with TMS based on intrinsic connectivity. Neuroimage 66, 151–160. https://doi.org/10.1016/j.neuroimage.2012.10.082 (2013).
Williams, N. R. et al. High-dose spaced theta-burst TMS as a rapid-acting antidepressant in highly refractory depression. Brain 141(3), e18. https://doi.org/10.1093/brain/awx379 (2018).
Ning, L., Makris, N., Camprodon, J. A. & Rathi, Y. Limits and reproducibility of resting-state functional MRI definition of DLPFC targets for neuromodulation. Brain Stimul. 12(1), 129–138. https://doi.org/10.1016/j.brs.2018.10.004 (2019).
Siddiqi, S. H. et al. Individualized connectome-targeted transcranial magnetic stimulation for neuropsychiatric sequelae of repetitive traumatic brain injury in a retired NFL player. J. Neuropsychiatry Clin. Neurosci. 31(3), 254–263. https://doi.org/10.1176/appi.neuropsych.18100230 (2019).
Siddiqi, S. H. et al. Repetitive transcranial magnetic stimulation with resting-state network targeting for treatment-resistant depression in traumatic brain injury: A randomized, controlled double-blinded pilot study. J. Neurotrauma. 36(8), 1361–1374. https://doi.org/10.1089/neu.2018.5889 (2019).
Cash, R. F. H., Cocchi, L., Lv, J., Fitzgerald, P. B. & Zalesky, A. Functional magnetic resonance imaging-guided personalization of transcranial magnetic stimulation treatment for depression. JAMA Psychiat. https://doi.org/10.1001/jamapsychiatry.2020.3794 (2020).
Siddiqi, S. H., Weigand, A., Pascual-Leone, A. & Fox, M. D. Identification of personalized TMS targets based on subgenual cingulate connectivity: An independent replication. Biol. Psychiatry. 90, e55–e56 (2021).
Cole, E. J. et al. Stanford accelerated intelligent neuromodulation therapy for treatment-resistant depression. Am. J. Psychiatry 177(8), 716–726. https://doi.org/10.1176/appi.ajp.2019.19070720 (2020).
Cole, E. J. et al. Stanford neuromodulation therapy (SNT): A double-blind randomized controlled trial. Am. J. Psychiatry 179(2), 132–141. https://doi.org/10.1176/appi.ajp.2021.20101429 (2022).
Mueller, S. et al. Reliability correction for functional connectivity: Theory and implementation. Hum. Brain Mapp. 36(11), 4664–4680. https://doi.org/10.1002/hbm.22947 (2015).
Laumann, T. O. et al. Functional system and areal organization of a highly sampled individual human brain. Neuron 87(3), 657–670. https://doi.org/10.1016/j.neuron.2015.06.037 (2015).
Glasser, M. F. et al. A multi-modal parcellation of human cerebral cortex. Nature 536(7615), 171–178. https://doi.org/10.1038/nature18933 (2016).
Hacker, C. D. et al. Resting state network estimation in individual subjects. Neuroimage 82, 616–633. https://doi.org/10.1016/j.neuroimage.2013.05.108 (2013).
Wang, D. et al. Parcellating cortical functional networks in individuals. Nat. Neurosci. 18(12), 1853–1860. https://doi.org/10.1038/nn.4164 (2015).
Birn, R. M. et al. The effect of scan length on the reliability of resting-state fMRI connectivity estimates. Neuroimage 83, 550–558. https://doi.org/10.1016/j.neuroimage.2013.05.099 (2013).
Lee, M. H. et al. Clinical resting-state fMRI in the preoperative setting: Are we ready for prime time?. Top. Magn. Reson. Imaging. 25(1), 11–18. https://doi.org/10.1097/RMR.0000000000000075 (2016).
Gordon, E. M. et al. Precision functional mapping of individual human brains. Neuron 95(4), 791–807. https://doi.org/10.1016/j.neuron.2017.07.011 (2017).
Greicius, M. D. et al. Resting-state functional connectivity in major depression: Abnormally increased contributions from subgenual cingulate cortex and thalamus. Biol. Psychiatry. 62(5), 429–437. https://doi.org/10.1016/j.biopsych.2006.09.020 (2007).
Liston, C. et al. Default mode network mechanisms of transcranial magnetic stimulation in depression. Biol. Psychiatry. 76(7), 517–526. https://doi.org/10.1016/j.biopsych.2014.01.023 (2014).
Fox, M. D. et al. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl. Acad. Sci. USA. 102(27), 9673–9678. https://doi.org/10.1073/pnas.0504136102 (2005).
Chai, X. J., Castanon, A. N., Ongur, D. & Whitfield-Gabrieli, S. Anticorrelations in resting state networks without global signal regression. Neuroimage 59(2), 1420–1428. https://doi.org/10.1016/j.neuroimage.2011.08.048 (2012).
Wong, C. W., Olafsson, V., Tal, O. & Liu, T. T. Anti-correlated networks, global signal regression, and the effects of caffeine in resting-state functional MRI. Neuroimage 63(1), 356–364. https://doi.org/10.1016/j.neuroimage.2012.06.035 (2012).
Gordon, E. M. et al. Individual-specific features of brain systems identified with resting state functional correlations. Neuroimage 146(918–939), 918–939. https://doi.org/10.1016/j.neuroimage.2016.08.032 (2017).
Gordon, E. M., Laumann, T. O., Adeyemo, B. & Petersen, S. E. Individual variability of the system-level organization of the human brain. Cereb. Cortex. 27(1), 386–399. https://doi.org/10.1093/cercor/bhv239 (2017).
Siddiqi, S. H. et al. Brain stimulation and brain lesions converge on common causal circuits in neuropsychiatric disease. Nat. Hum. Behav. 5, 1707 (2021).
Han, K. et al. Disrupted modular organization of resting-state cortical functional connectivity in U.S. military personnel following concussive “mild” blast-related traumatic brain injury. Neuroimage 84, 76–96. https://doi.org/10.1016/j.neuroimage.2013.08.017 (2014).
Han, K., Chapman, S. B. & Krawczyk, D. C. Disrupted intrinsic connectivity among default, dorsal attention, and frontoparietal control networks in individuals with chronic traumatic brain injury. J. Int. Neuropsychol. Soc. 22(2), 263–279. https://doi.org/10.1017/S1355617715001393 (2016).
van der Horn, H. J. et al. Graph analysis of functional brain networks in patients with mild traumatic brain injury. PLoS ONE 12(1), e0171031. https://doi.org/10.1371/journal.pone.0171031 (2017).
Caeyenberghs, K., Verhelst, H., Clemente, A. & Wilson, P. H. Mapping the functional connectome in traumatic brain injury: What can graph metrics tell us?. Neuroimage 160, 113–123. https://doi.org/10.1016/j.neuroimage.2016.12.003 (2017).
Siddiqi, S. H. rTMS for major depression associated with TBI. Open Science Framework. osf.io/vjddq
Van Essen, D. C. et al. The human connectome project: A data acquisition perspective. Neuroimage 62(4), 2222–2231. https://doi.org/10.1016/j.neuroimage.2012.02.018 (2012).
Power, J. D. et al. Methods to detect, characterize, and remove motion artifact in resting state fMRI. Neuroimage 84, 320–341. https://doi.org/10.1016/j.neuroimage.2013.08.048 (2014).
Blumberger, D. M. et al. Unilateral and bilateral MRI-targeted repetitive transcranial magnetic stimulation for treatment-resistant depression: A randomized controlled study. J. Psychiatry Neurosci. 41(4), E58-66. https://doi.org/10.1503/jpn.150265 (2016).
Yeo, B. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106(3), 1125–1165. https://doi.org/10.1152/jn.00338.201110.1152/jn.00338.2011 (2011).
Cole, M. W., Yang, G. J., Murray, J. D., Repovš, G. & Anticevic, A. Functional connectivity change as shared signal dynamics. J. Neurosci. Methods 259, 22–39. https://doi.org/10.1016/j.jneumeth.2015.11.011 (2016).
Schaefer, A. et al. Local-global parcellation of the human cerebral cortex from intrinsic functional connectivity MRI. Cereb. Cortex. 28(9), 3095–3114. https://doi.org/10.1093/cercor/bhx179 (2018).
Hamilton, J. P. et al. Default-mode and task-positive network activity in major depressive disorder: Implications for adaptive and maladaptive rumination. Biol. Psychiatry. 70(4), 327–333. https://doi.org/10.1016/j.biopsych.2011.02.003 (2011).
Kaiser, R. H., Andrews-Hanna, J. R., Wager, T. D. & Pizzagalli, D. A. Large-scale network dysfunction in major depressive disorder: A meta-analysis of resting-state functional connectivity. JAMA Psychiat. 72(6), 603–611. https://doi.org/10.1001/jamapsychiatry.2015.0071 (2015).
Choi, K. S., Riva-Posse, P., Gross, R. E. & Mayberg, H. S. Mapping the “depression switch” during intraoperative testing of subcallosal cingulate deep brain stimulation. JAMA Neurol. 72(11), 1252–1260. https://doi.org/10.1001/jamaneurol.2015.2564 (2015).
Murphy, K. & Fox, M. D. Towards a consensus regarding global signal regression for resting state functional connectivity MRI. Neuroimage 154, 169–173. https://doi.org/10.1016/j.neuroimage.2016.11.052 (2017).
Cash, R. F. H. et al. Personalized connectivity-guided DLPFC-TMS for depression: Advancing computational feasibility, precision and reproducibility. Hum. Brain Mapp. 42(13), 4155–4172. https://doi.org/10.1002/hbm.25330 (2021).
Chen, J. et al. Left versus right repetitive transcranial magnetic stimulation in treating major depression: A meta-analysis of randomised controlled trials. Psychiatry Res. 210(3), 1260–1264. https://doi.org/10.1016/j.psychres.2013.09.007 (2013).
Sale, M. V., Mattingley, J. B., Zalesky, A. & Cocchi, L. Imaging human brain networks to improve the clinical efficacy of non-invasive brain stimulation. Neurosci. Biobehav. Rev. 57, 187–198. https://doi.org/10.1016/j.neubiorev.2015.09.010 (2015).
Opitz, A., Fox, M. D., Craddock, R. C., Colcombe, S. & Milham, M. P. An integrated framework for targeting functional networks via transcranial magnetic stimulation. Neuroimage 127, 86–96. https://doi.org/10.1016/j.neuroimage.2015.11.040 (2016).
Acknowledgements
We would especially like to thank the experimental participants. We thank Dr. Abraham Snyder for logistical support with implementation of individualized RSN mapping. We thank Dr. Benjamin Srivastava for assisting with clinical assessments. We thank Linda Hood for technical support with MRI equipment. We thank Drs. Sindhu Jacob and Martin Wice for referring participants. We additionally acknowledge valuable discussions with Drs. Michael Fox, Steven Petersen, Charles Conway, Eric Wasserman, Sarah Lisanby, Bruce Luber, Andrew Drysdale, Irving Reti, Vani Rao, Maurizio Corbetta, Gordon Shulman, and Abraham Snyder which led to the conception and refinement of the study design.
Funding
The study protocol was funded by the McDonnell Center for Systems Neuroscience (New Resource Proposal) and the Mallinckrodt Institute of Radiology (funding for scans). SHS received fellowship support from the Sidney R. Baer Foundation. SHS received salary support from the Center for Neuroscience and Regenerative Medicine at the Uniformed Services University for Health Sciences. DLB was an employee of Washington University at the time the study was conducted, and is now an employee of the Uniformed Services University of the Health Science, Department of Defense.
Author information
Authors and Affiliations
Contributions
S.H.S., D.L.B., A.R.C., and N.T.T. designed the clinical protocol. A.R.C. provided rTMS expertise and equipment as well as functional connectivity expertise. C.D.H. provided expertise and novel analytical scripts for individualized R.S.N. mapping and functional connectivity analysis. E.C.L. provided expertise for clinical implementation of RSN mapping techniques. S.K. provided expertise and developed scripts for functional connectivity processing. S.H.S. coordinated the study, recruited subjects, administered treatments, processed MRI data, conducted functional connectivity analyses, and conducted all statistical analyses. N.T.T. and C.D.H. also administered study treatments. S.H.S., and N.T.T. acquired MRI scans and conducted clinical assessments. S.H.S., D.L.B., and A.R.C. wrote the manuscript with intellectual contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
SHS serves as a scientific consultant for Magnus Medical and a clinical consultant for Kaizen Brain Center and Acacia Mental Health. SHS has received research support from Neuronetics Inc and Brainsway Ltd. SHS owns shares in Brainsway Ltd (publicly-traded) and Magnus Medical (not publicly traded). SHS has served as an speaker on behalf of Otsuka (unbranded educational material) and Brainsway (unbranded scientific material). None of these entities were involved in the present work. DLB has served as a consultant for Pfizer Inc, Intellectual Ventures, Signum Nutralogix, Kypha Inc, Sage Therapeutics, iPerian Inc, Navigant, Avid Radiopharmaceuticals (Eli Lilly & Co), the St Louis County Public Defender, the United States Attorney’s Office, the St Louis County Medical Examiner, GLG, Stemedica, Luna Innovations, and QualWorld. DLB holds equity in the company Inner Cosmos. DLB receives royalties from sales of Concussion Care Manual (Oxford University Press) and an honorarium from Mary Ann Liebert Inc for services as Editor-in-Chief of Journal of Neurotrauma. No conflicts of interest with the presented work. DLB is an employee of the Department of Defense; the views expressed here do not reflect those of the Uniformed Services University of the Health Sciences, the US Department of Defense, or the US Government. ECL holds equity in the companies Neurolutions and Inner Cosmos. CDH, ECL, and SHS hold intellectual property related to the use of RSNM to target TMS. The remaining authors report no conflicts of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Siddiqi, S.H., Kandala, S., Hacker, C.D. et al. Individualized precision targeting of dorsal attention and default mode networks with rTMS in traumatic brain injury-associated depression. Sci Rep 13, 4052 (2023). https://doi.org/10.1038/s41598-022-21905-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-21905-x
This article is cited by
-
Prospects for Nerve Regeneration and Gene Therapy in the Treatment of Traumatic Brain Injury
Journal of Molecular Neuroscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.