Abstract
The global prevalence of colistin-resistant Klebsiella pneumoniae (ColRkp) facilitated by chromosomal and plasmid-mediated Ara4N or PEtN-remodeled LPS alterations has steadily increased with increased colistin usage for treating carbapenem-resistant K. pneumoniae (CRkp). Our study demonstrated the rising trend of ColRkp showing extensively and pandrug-resistant characteristics among CRkp, with a prevalence of 28.5%, which was mediated by chromosomal mgrB, pmrB, or phoQ mutations (91.5%), and plasmid-mediated mcr-1.1, mcr-8.1, mcr-8.2 alone or in conjunction with R256G PmrB (8.5%). Several genetic alterations in mgrB (85.1%) with increased expressions of Ara4N-related phoPQ and pmrK were critical for establishing colistin resistance in our isolates. In this study, we discovered the significant associations between extensively drug-resistant bacteria (XDR) and pandrug-resistant bacteria (PDR) ColRkp in terms of moderate, weak or no biofilm-producing abilities, and altered expressions of virulence factors. These ColRkp would therefore be very challenging to treat, emphasizing for innovative therapy to combat these infections. Regardless of the underlying colistin-resistant mechanisms, colistin-EDTA combination therapy in this study produced potent synergistic effects in both in vitro and in vivo murine bacteremia, with no ColRkp regrowth and improved animal survival, implying the significance of colistin-EDTA combination therapy as systemic therapy for unlocking colistin resistance in ColRkp-associated bacteremia.
Similar content being viewed by others
Introduction
Carbapenem resistant Klebsiella pneumoniae (CRkp) is frequently encountered in clinical settings1. Faced with resistance to all standard therapeutic options, clinicians are being urged to reconsider colistin as a viable treatment option in response to rising global CRkp prevalence, few effective therapeutic alternatives and limitations in novel antibiotic development2,3. Colistin, a bactericidal polycationic peptide, attaches to anionic lipid A of lipopolysaccharide (LPS), displacing Ca++ and Mg++ which form bridges between LPS, causing bacterial membrane destabilization. Colistin also triggers bacterial death through hydroxy radical-induced oxidative stress and impairment of bacterial respiratory chains3,4,5. Nevertheless, as colistin use has increased, worldwide prevalence of ColRkp has risen3,4,5,6,7.
Colistin resistance develops in K. pneumoniae due to a reduction in net negative charge of LPS generated by amino-4-deoxy-l-arabinose (Ara4N) and phosphoethanolamine (pEtN)-mediated LPS alterations, causing diminished electrostatic affinity between colistin and LPS3,4. The pmrHFIJKLM operon facilitates Ara4N-remodeled LPS alteration, whereas the pmrCAB operon supports pEtN-integrated LPS modification3,4. As chromosomal-mediated mechanisms, mgrB, phoPQ, and pmrAB mutations influence pmrHFIJKLM and pmrCAB expressions to trigger LPS alterations3,4. Plasmid-mediated colistin-resistant mechanisms include presence of mcr with different alleles (mcr 1–9) that encode phosphoethanolamine transferase causing PEtN-modified LPS3,4.
Furthermore, several studies observed establishment of hypervirulent ColRkp with diverse virulence characteristics8,9,10,11. In K. pneumoniae, biofilm development is a crucial virulence characteristic, and managing biofilms is extremely challenging as they confer significant tolerance to host defense responses and antibiotic effects12,13,14. Type 3 adhesin, mrkD, is implicated in bacterial adherence and biofilm development12,13,14. Hypervirulent K. pneumoniae displays iron-scavenging siderophores—ybtS and kfu that facilitate systemic survival by regulating immunological responses12,15. OmpK35 and OmpK36 are outer membrane porins that help bacteria survive by maintaining membrane integrity and delivering important nutrients14,16. LPS genes—uge and wabG are implicated in promoting pathogenicity by protecting bacteria against host humoral defenses12,14,17. Type 2 quorum-sensing regulatory system-luxS facilitates biofilm development by encouraging cell-to-cell communication12,18. It has been reported that not only PmrAB and PhoPQ support bacterial virulence by regulating virulence gene transcripts, but in addition mgrB-related LPS alterations also augment virulence by suppressing early host defense19,20,21, highlighting the importance of exploring the association between colistin resistance and other virulence factors that influence bacterial pathogenicity. Due to converging colistin resistance and hypervirulence, clinically untreatable K. pneumoniae superbugs may evolve, emphasizing the urgent need to develop viable therapeutic strategy to minimize mortality, morbidity and health-care expenses associated with these infections8,12,22.
Colistin combination therapy, when used to treat drug-resistant bacteria, has been shown to significantly lower treatment failure rates and enhance patient survival23. Ethylenediamine tetra-acetic acid (EDTA) is an anti-virulence drug that disrupts permeability-associated resistance mechanisms and restores antibiotic potency against resistant bacteria through metal ion chelation24, suggesting that it could be used as adjuvant in colistin combination therapy to counteract ColRkp expressing different virulence characteristics. Thus, the aims of the present study were to determine the underlying chromosomal and plasmid-mediated resistance mechanisms encoded in ColRkp, identify any association between colistin resistance and diverse virulence characteristics, and to assess the effectiveness of colistin-EDTA combination in unlocking colistin resistance in ColRkp with different colistin-resistance mechanisms.
Results
Prevalence of colistin resistance among CRkp clinical isolates was on the rising trend
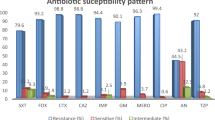
Between 2016 and 2021, 165 CRkp retrieved from hospitalized patients exhibited different antibiotic resistance profiles, with highest resistance to ceftazidime and ciprofloxacin (100%), followed by imipenem (94.5%), meropenem (90.3%), fosfomycin (31.5%) and amikacin (23.6%). Of these, 47 isolates (28.5%) showed colistin resistance, with their incidence growing rapidly over time from 14.9% in 2016 to 36.2% in 2021 (Table 1) (Supplementary Fig. 1). The majority of ColRkp isolates (91.5%) exhibited extensively drug-resistance with the remaining 8.5% pan-drug-resistant (Table 2). Furthermore, when compared to their corresponding planktonic MIC, biofilms of these isolates showed antibiotic tolerance with 1000 times higher MBEC to tested antibiotics (Table 1).
Both chromosomal-mediated and plasmid-mediated mechanisms were responsible for establishing colistin resistance
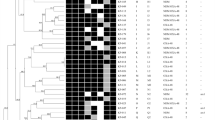
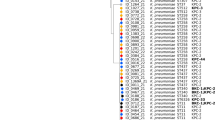
Among chromosomal-mediated colistin resistance (91.5%), mgrB was the most inactivated chromosomal gene (85.1%) primarily by insertion sequences (IS) (61.7%) including IS1-like, ISkpn14-like, IS3-like, IS5-like and IS1380-like elements (ISEcp1-like) (Table 2, Fig. 1a). These were found to insert with different orientations using their inverted repeats within the coding region and the upstream between mgrB start codon and putative promoter region (Table 2, Fig. 1a). Furthermore, different genetic alterations in the mgrB coding sequence (12.8%) were detected, including point mutations in the initial codon (G3A, GTG > GTA), as well as those generating an internal stop codon (A7T-AAA > TAA, G60A-TGG > TGA). We also observed deletion of mgrB since no mgrB PCR amplicons could be detected despite utilizing various primers (10.6%) in both XDR and PDR ColRkp isolates (Table 2, Fig. 1b). Point mutations in pmrB causing the amino acid substitutions—T157P (A469C, ACC > CCC) and phoP-E82K (G244A, GAA > AAA) were detected in 4.25% and 2.1% respectively (Table 2, Fig. 1c,d). Interestingly, all 47 ColRkp isolates used in this study harbored wild type pmrA and phoQ genes. The presence of plasmid-mediated mcr 1.1 or 8.2 alone with low level colistin resistance (8 mg/L), and the combined presence of mcr 1.1 or 8.1 with PmrB—R256G (C766G, CGC > GGC) demonstrating increased colistin resistance (16–32 mg/L) were investigated in 4 ColRkp isolates (8.5%) (Table 2, Fig. 2a–c). Statistically non-significant distributions of ESBLs and carbapenemases, apart from KPC and IMP, were observed in ColRkp with different colistin-resistant mechanisms (Table 2) (Supplementary results Table 2). The most common ESBLs and carbapenemase profiles were TEM-CTXM (n = 20, 46.5%) and OXA-48 (n = 15, 34.8%) in ColRkp with chromosomal-mediated colistin resistance, as well as SHV,TEM, CTX-M (n = 3, 75%) and NDM (n = 1, 25%) in ColRkp with plasmid-mediated colistin resistance, respectively (Supplementary results Table 2).
Chromosomal-mediated colistin-resistant mechanisms including (a) mgrB inactivation by different IS within the coding region and the upstream between mgrB start codon and putative promoter region of ColRkp (n = 29), (b) MgrB inactivation by point mutations—A7T MgrB, G3A MgrB, G60A MgrB in ColRkp (n = 6), (c) T157P PmrB in ColRkp (n = 2), (d) E82K PhoP in ColRkp (n = 1), observed in this study.
Expressions of LPS modification genes were significantly upregulated in ColRkp isolates
In ColRkp isolates with inactivated mgrB by IS1-like and G60A point mutation, transcriptional levels of Ara4N-related phoPQ and pmrK were significantly increased as compared to the levels in colistin-susceptible clinical strains (ColSkp) with wild type mgrB (Fig. 3a,b; p < 0.0001). The ColRkp with deleted mgrB showed significantly upregulated Ara4N-related phoPQ, connector-pmrD, and PEtN-related pmrCAB transcripts (Fig. 3c; p < 0.05). In ColRkp with T157P PmrB, PEtN-related pmrCAB was significantly overexpressed (Fig. 3d; p < 0.001). The ColRkp with E82K PhoP had considerably upregulated phoPQ, pmrD, and pmrK transcriptions (Fig. 3e; p < 0.0001). In ColRkp with combined presence of mcr-8.1 and R256G PmrB, significantly overexpressed pmrCAB were observed (Fig. 3f; p < 0.0001). Additionally, IS1-like integration in mgrB promoter region and G3A mgrB mutation significantly reduced mgrB expressions (p < 0.05) (Supplementary results Fig. 2). Because the majority of ColRkp had IS-mediated or point mutations in mgrB with significant overexpression of Ara4N-related LPS modification genes, several genetic alterations in mgrB with increased expressions of Ara4N-related phoPQ and pmrK were considered to be important for establishing colistin resistance in our isolates (Fig. 3a–f).
The abilities to produce biofilms varied between XDR and PDR ColRkp isolates
The majority of ColRkp isolates (95.7%) produced biofilms, with 76.7% of XDR ColRkp produced strong biofilms, 11.6% had moderate biofilms, 9.3% formed weak biofilms and 2.3% developed no biofilms (Fig. 4a,b). Meanwhile, 75% of PDR ColRkp developed strong biofilms and 25% had no biofilms (Fig. 4b). Significant associations existed between XDR and PDR ColRkp isolates in terms of moderate or weak abilities to produce biofilms, which were higher in XDR ColRkp than in PDR isolates. Interestingly, no biofilm-producing ability was higher in PDR than XDR ColRkp (Fig. 4b; p < 0.0001). It was also observed that all five colistin-susceptible K. pneumoniae (ColSkp) strains developed significantly stronger biofilms (100%) when compared to XDR and PDR ColRkp (Supplementary results Table 1; p < 0.0001).
Coexistence and altered expressions of virulence factors were observed in ColRkp
The existence of all tested bacterial virulence genes as mrkD-kfu-ybtS-ompK35-ompK36-uge-wabG-luxS combination was detected in 12.8% of XDR ColRkp isolates (Fig. 5a). Meanwhile, the most prevalent virulence gene combination was mrkD-ybtS-ompK35-ompK36-uge-wabG-luxS combination, which was observed in 70.2% of XDR strains and 8.5% of PDR strains (Fig. 5a). The combination of mrkD-kfu-ompK35-ompK36-uge-wabG-luxS was identified in 4.3% of XDR strains (Fig. 5a). The least common virulence gene combinations were mrkD-ompK35-ompK36-uge-wabG-luxS and mrkD-ybtS-ompK35-ompK36-wabG-luxS combinations which were observed in 2.1% each of XDR ColRkp isolates (Fig. 5a). In comparison to ColSkp clinical isolates, XDR ColRkp displayed altered expressions of virulence factors, showing significantly higher expressions of ompK35, ompK36, kfu, uge, and luxS (p < 0.0001), as well as significantly lower expression of wabG (Fig. 5b; p < 0.001).
Colistin-EDTA combination showed remarkable synergistic effects against ColRkp in vitro
The adjuvant-EDTA was discovered to have inhibitory effects on ColRkp at concentrations of 3-24 mg/mL (Table 2). When we performed checkerboard assays on ColRkp isolates (n = 45), colistin MIC was lowered to 0.25 mg/L when given in conjunction with 12 mg/mL EDTA, and this colistin-EDTA combination displayed substantial synergistic effects (FICI ≤ 0.5) on all tested 45 ColRkp isolates (Table 2). Using time-kill synergy confirmation assays, synergistic effects of colistin-EDTA combination were confirmed in 11 representative ColRkp with different colistin-resistant mechanisms, including 3 PDR ColRkp with mgrB loss, E82K PhoP, and mcr-1.1, and 8 XDR ColRkp with mgrB inactivation by IS1-like, IS3-like, IS5-like, IS1380-like elements, point mutations- G60A, A7T mgrB, T157P PmrB and combined mcr-8.1 with R256G PmrB (Fig. 6a–k). Despite bacterial regrowth after 6 h of colistin and EDTA monotherapy, colistin-EDTA combination exhibited remarkable synergistic activities in reducing > 3log10 of bacteria starting 2 h after treatment and produced prolonged bactericidal effects with no regrowth until 24 h, in all tested XDR and PDR ColRkp isolates (n = 11), regardless of their underlying colistin resistance mechanisms (Fig. 6a–k).
Administration of colistin-EDTA combination showed potent synergistic activities in murine ColRkp-associated bacteraemia
When compared to colistin and EDTA monotherapy, single intraperitoneal administration of colistin-EDTA combination significantly reduced bacterial burden in murine ColRkp-associated bacteraemia induced by strong biofilm-producing XDR ColRkp with inactivated mgrB by IS1-like element with the presence of evaluated virulence factors (Fig. 7a; p < 0.01). Intraperitoneal colistin-EDTA combination therapy once a day significantly improved the survival of treated mice with ColRkp-associated bacteraemia as compared to their monotherapy (Fig. 7b; p < 0.0001).
Discussion
Klebsiella pneumoniae is a pathogen causing severe untreatable hospital-acquired infections in immunocompromised patients owing to increasing rates of antimicrobial resistance1. With the rising prevalence of CRkp, colistin has emerged as a feasible therapeutic option due to paucity of effective therapeutic alternatives and restrictions in novel antibiotic development3. Consequently, the global prevalence of ColRkp has steadily increased as a result of expanded usage of colistin, revealing the significant threat for the emergence and spread of extensively and pandrug-resistant strains around the world3. In our study, a total of 165 CRkp isolates were observed to exhibit different antibiotic susceptibilities in both planktonic and biofilm environments. Among CRkp, we discovered a 28.5% prevalence of XDR and PDR ColRkp (n = 47), which increased over time from 14.9% in 2016 to 36.2% in 2021.The colistin-resistant rate in our clinical setting is comparable to India (30%)25 and Italy (22.4%)26, but higher than other clinical settings in Thailand (6.6%)27, Nigeria (9.1%)28 and other regions of the world29. Despite the lack of clinical data on colistin use in our hospital, the rising trend of ColRkp is almost certainly due to selective pressure from increased colistin use in clinical settings for increasing CRkp burden, and in poultry industry setting as a short-term colistin preventive strategy for Gram-negative bacterial infections30,31. This could lead to the emergence and colonization of ColRkp among patients, healthy adults, and food animals27,30,31, causing further circulation of colistin resistance with a higher regional colistin-resistant prevalence in Thailand.
Because colistin resistance is continuously growing and varying between countries over time3,31, addressing the underlying colistin-resistant mechanisms has become critical to deduce. Among ESBL and carbapenemase-producing ColRkp in this study, mgrB alteration played a significant role (85.1%), with inactivation by IS1-like, ISkpn14-like, IS3-like, IS5-like and IS1380-like elements (61.7%), point mutation (12.8%) and deletion (10.6%), which is consistent with several studies around the world32,33,34. Downregulated expressions of mgrB were observed in ColRkp with IS integration in promoter region and G3A mgrB, as proposed previously32,35. Moreover, as evidenced in previous studies, expressions of Ara4N-related and PEtN-related LPS modification genes were significantly upregulated in ColRkp isolates with different underlying colistin resistance mechanisms35,36,37. Because the majority of ColRkp (85.1%) had inactivated mgrB, several genetic alterations in mgrB with upregulated Ara4N-related LPS modification genes were considered to be crucial in establishing colistin resistance in our isolates, as previously demonstrated32,35,36. In this study, 4.25% of isolates exhibited the T157P PmrB, along with overexpression of PEtN-related pmrCAB operons, which has previously been proved to cause colistin resistance in K. pneumoniae38. Amino acid substitution—R256G PmrB with significant pmrCAB transcription, was revealed as a combined colistin-resistant mechanism in 4.25% of our isolates (n = 2) harboring mcr-1.1 or mcr-8.2 genes. Although the PROVEAN bioinformatic tool anticipated a deleterious effect of R256G PmrB on its protein function, this substitution was discovered as lineage-specific mutations in both polymyxin-susceptible as well as resistant K. pneumoniae, and it has been confirmed by others to be unrelated to colistin resistance in K. pneumoniae39,40,41,42. Furthermore, PhoQ E82K substitution with significant expression of phoQ was identified in 2.1% of ColRkp isolates, which is consistent with prior research43.
This study showed an 8.5% prevalence of plasmid-mediated mcr genes, which is higher than reports described in earlier studies in Thailand (< 1 to 3.2%)41,42,43,44,45. Until recently, K. pneumoniae of livestock origins from different regions of Thailand have been documented to harbor plasmid-mediated mcr-8 phosphoethanolamine transferase27,41. Our study is the first to show the presence of mcr-8.1 and mcr-8.2 in ESBL and carbapenemase-producing ColRkp isolates from human clinical samples in Thailand. These findings revealed that mcr-8 and its variants have been existing for a period and are widely disseminated among K. pneumoniae of both human and animal origins in this region, suggesting a growing threat of antibiotic resistance in the years46. The judicious use of colistin, as well as continuous monitoring of mcr genes transferability and stability, will tremendously help in the prevention and control of antimicrobial resistance47.
Previous studies reported the establishment of hypervirulent ColRkp with diverse virulence characteristics8,9,10,11 and the potential roles of mgrB, pmrAB and phoPQ in supporting bacterial virulence19,20,21, highlighting the importance of exploring the association between colistin resistance and other virulence factors that influence bacterial pathogenicity. Biofilm formation and diverse virulent factors including mrkD, kfu, ybtS, ompK35, ompK36, uge, wabG and luxS have been implicated in bacterial colonization, invasion, and pathogenicity within the host12,13,14,15,16,17,18. In this study, we discovered that majority of ColRkp produced biofilms, whereas significant associations existed between XDR and PDR ColRkp in terms of moderate, or weak biofilm-producing abilities, which were higher in XDR than in PDR isolates. Interestingly, no biofilm-producing ability was higher in PDR than XDR ColRkp. These findings suggest that not only PDR ColRkp, but XDR ColRkp may also have increased biofilm-mediated antibiotic tolerance which could enhance their abilities to resist the antibiotics effects in order to produce untreatable infections48. Additionally, all of the evaluated virulence genes were encoded in 12.8% of XDR ColRkp stains, and other virulence gene combinations were encoded in varying frequencies in other XDR and PDR ColRkp isolates. There were significant overexpression of ompK35, ompK36, kfu, uge, and luxS as well as lower expression of wabG in ColRkp compared to ColSkp, demonstrating the coexistence and altered expression of bacterial virulence factors in ColRkp. Due to their resistance to last resort colistin therapy, increased biofilm-producing abilities, coexistence and altered expression of virulence factors, these isolates would be very challenging to treat and it emphasizes the critical requirement for innovative therapy to combat these infections in healthcare settings8,11,22.
Consistent with earlier findings, a colistin-EDTA combination demonstrated in vitro potent synergistic effects at significantly lower colistin concentrations with no bacterial regrowth, implying lower probabilities of developing colistin toxicities under this combination therapy, and inferring fewer resistance concerns after its prolonged therapy6. Despite previous studies highlighting colistin-EDTA combination as lock therapy in localized catheter-associated infections6,49, administration of this combination in our murine bacteremia model with ColRkp resulted in a reduction of bacterial burden and increased animal survival, indicating their potent in vivo efficacy as systemic therapy in overcoming ColRkp-associated bacteremia. The potent synergistic effects of colistin-EDTA against XDR and PDR ColRkp isolates could be attributed to EDTA ions sequestration activities, which increase bacterial outer membrane permeabilities and then sensitize as well as synergize with colistin to regain colistin efficacy of increased permeabilizations, leading to enhanced intracellular content release and bacterial death3,50. EDTA chelation could augment colistin's entry into bacteria to exert bactericidal effects by blocking intracellular targets of colistin—essential respiratory enzymes, thereby unlocking colistin resistance regardless of underlying colistin-resistant mechanisms51.
In conclusion, these data revealed a rising trend of both chromosomal and plasmid-mediated colistin resistance in K. pneumoniae isolated from Chulalongkorn Memorial Hospital, Thailand between 2016 to 2021, which was also linked to altered bacterial virulence factors. Potent synergistic effects of colistin-EDTA combination against ColRkp-associated bacteremia suggest their promising application as systemic therapy in unlocking colistin resistance of untreatable superbugs, regardless of underlying colistin-resistant mechanisms, but more clinical trials are necessary to further evaluate their clinical efficacy, tolerance, and safety. The effects of altered virulence gene expression in ColRkp will need to be investigated further to learn more about how colistin-resistant bacteria modulate their pathogenicity inside the host, which will support the implementation of more effective targeted strategies to overcome and mitigate their infectivity.
Materials and methods
Bacterial isolates and antimicrobial susceptibility testing
A total of 165 CRkp clinical isolates which showed resistance to either imipenem or meropenem or both, were obtained from Chulalongkorn Memorial Hospital, Thailand during 2016 to 2021 after approved by the Institutional Review Board (IRB) of the Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand. All isolates are identified by 16srRNA sequencing. To determine planktonic susceptibilities (MIC) to colistin and EDTA, drugs were serially diluted two-fold in 96-well microtiter plates using standard broth microdilution according to criteria in EUCAST (criteria for Enterobacteriaceae for colistin only)52 and CLSI53. Planktonic susceptibilities to other antibiotics including imipenem, meropenem, ceftazidime, ciprofloxacin, amikacin and fosfomycin supplemented with glucose-6-phosphate were determined by agar dilution53. MIC was determined as the lowest concentration that inhibited the visible growth of the bacteria. According to antibiotic susceptibilities, ColRkp were categorized to XDR (non-susceptibility to at least one agent in all but two or fewer antimicrobial categories) and PDR (non-susceptibility to all agents in all antimicrobial categories)54.
Characterization of colistin resistance mechanisms
Chromosomal-mediated colistin resistance mechanisms were analyzed by targeted amplification and sequencing of mgrB, pmrA, pmrB, phoP and phoQ using previously described primers35,38 (Supplementary Methods). The resulting nucleotide and amino acid sequences were analyzed by Basic Local Alignment Search Tool (BLAST) and multiple sequence alignment by Florence Corpet (http://multalin.toulhouse.inra.fr/mutalin/multalin.html) was used to compare mutations observed in ColRkp nucleotide and amino acid sequences to reference sequences of K. pneumoniae subsp. pneumoniae MGH 78578 (GenBank accession number. CP_000647.1), Insertion Sequences (ISs) were analyzed using the IS finder web site (www-is.biotoul.fr). For determination of plasmid-mediated colistin resistance, mcr-1-9 genes were screened by PCR using primers as established previously and confirmed by subsequent sequence analysis using the primers that target for amplification of mcr gene of interest55,56. The PROVEAN tool v.1.1.5 (http://provean.jcvi.org/index.php) was used to predict the effect of amino acid substitutions on protein function57. PROVEAN score ≤ − 2.5 was deleterious for protein function, and a score > − 2.5 was considered to have a neutral effect on protein function.
Determination of extended spectrum β-lactamase (ESBL) and carbapenemase genes
Presence of ESBL (CTXM,TEM,OXA,SHV) and carbapenemase genes (KPC, NDM, OXA-48, IMP, VIM) conferring resistance to broad range of β-lactam and carbapenem antibiotics were analyzed among ColRkp isolates using primers as previously reported58,59.
Determination of the expressions of LPS modification genes associated with ColRkp
By using specific primers as previously described60,61, Quantitative RT-PCR (qRT-PCR) was used to assess the expression levels of LPS modification genes among ColRkp isolates with various colistin-resistant mechanisms. These genes include Ara4N-related pmrK and phoPQ, connector pmrD, and PEtN-related pmrCAB that are known to be involved in establishing Ara4N-related and PEtN-related LPS modification for colistin resistance. The tested isolates were ColRkp isolates with IS1-like, G60A and deleted mgrB, T157P PmrB, E82K PhoP and combined presence of mcr-8.1 and R256G PmrB. Briefly, Monarch Total RNA Miniprep Kit (Biolabs, New England) was used to extract total RNA from tested bacterial cultures grown in Luria–Bertani broth (Merck, Darmstadt, Germany) during the mid-logarithmic growth phase. These DNase-treated purified RNA was subsequently reverse-transcribed into cDNA and qRT-PCR expression assays were performed using cDNA of tested ColRkp and ColSkp clinical isolates.
Determination of in vitro biofilm-mediated colistin tolerance
Using crystal violet assay, the amounts of biofilms produced by ColRkp were determined for assessing their biofilm-producing abilities and biofilm-mediated antibiotic tolerance62. Briefly overnight cultures of bacteria were standardized with an OD600 of 0.02 at 600 nm (5 × 107 CFU mL−1) and 100 μL aliquots are added in triplicate to flat-bottomed 96-well polystyrene microtiter plates (SPL Life Sciences). Plates were then incubated at 37 °C for 24 h. Adherent biofilms were fixed with crystal violet (0.1%) and stained biofilms were solubilized with 30% acetic acid. Absorbance (OD) at 560 nm was then determined using a microtiter-plate-reading fluorimeter (Varioskan Flash Multimode Reader; Thermo Fisher Scientific). All experiments were performed in triplicate and repeated three times. Minimal biofilm eradication concentration (MBEC) is the lowest concentration of antimicrobial agent that eradicates all mature biofilm biovolume and biofilm-embedded bacteria reducing bacterial viability by ≤ 10% as compared to growth controls63,64. Crystal violet assay was used to determine the drugs MBECs on mature ColRkp biofilms biovolume by measuring the percent eradication of biofilm biovolume6,62,63,64. Prestoblue assay was used to evaluate drugs MBECs by measuring the percent reduction of biofilm-embedded viable bacteria. In this assay, cell viability indicator—Prestoblue (Invitrogen) which turns fluorescent red in environment of viable cell was used. The fluorescence intensity of drug-treated biofilm was measured with excitation 535 nm and emission 590 nm under a microtiter-plate-reading fluorimeter6,62,63,64,65,66.
Determination of the coexistence and expressions of virulence factors in ColRkp
The presence of virulence factors (kfu, luxS, mrkD, ompK35, ompK36, uge, wabG and ybtS) were firstly identified to determine their coexistence in ColRkp, by utilizing PCR with bacterial DNA extracted by Purelink genomic DNA micro kit (Invitrogen, USA). Using previously reported specific primers and bacterial cDNA produced from mRNA isolated from bacterial cultures grown in Luria–Bertani broth during mid-logarithmic phase, qRT-PCR was then used to assess the relative expression of these 8 different virulence factors in ColRkp and ColSkp clinical isolates67.
Quantitative RT-PCR (qRT-PCR)
Using the 2∆∆CT method, the relative expressions of 6 different LPS-modification genes and 8 different virulence factors were computed after normalization with control housekeeping gene—rpoD followed by subsequent normalization against the value obtained for ColSkp isolate to evaluate and compare the fold change differences.
Synergy assays for colistin-EDTA combination against ColRkp isolates in vitro
The synergistic activities of colistin-EDTA combination against ColRkp clinical isolates were first determined by checkerboard assay68. The synergistic activities were interpreted according to fractional inhibitory concentration index (FICI): Synergy: FIC index ≤ 0.5 and Additive: 0.5 > FIC index ≤ 1. The synergistic activities were then confirmed using time-kill assays against 11 representative PDR and XDR ColRkp isolates with various underlying colistin-resistant mechanisms68. Briefly, adjusted bacterial suspension of each ColRkp isolate was added to into each flask of 9 different growth conditions, including no drug (growth control), 1× and 0.5×MIC of colistin and EDTA as monotherapy and combination therapy, and incubated for 24 h at 37 °C. The viable cell counts (CFU/mL) under different growth conditions were then determined at 0, 2, 4, 6, 8, 12, and 24 h by plate counting. The synergistic activities are interpreted when there is ≥ 2 log10 (CFU/mL)-fold decrease in combination compared with the single antibiotic; Bactericidal activity is defined as a ≥ 3 log10 (CFU/mL)-fold decrease when compared to the number of viable cells at initial time point68.
Animal study
6–8-week-old C57BL/6 male mice were purchased from Nomura Siam International (Pathumwan, Bangkok, Thailand) and were used in all experiments. Animals were at rest for 1 week in the animal facility before use. Animals received food and water ad libitum and were housed at a maximum of 2 mice per cage, weighed and closely monitored for any signs of distress throughout experimental periods. The animal study was conducted according to guidelines and protocols approved by the Institutional Animal Care and Use Committee of the Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand, based on the National Institutes of Health (NIH), USA.
In vivo murine bacteraemia model
To establish ColRkp-associated bacteremia in vivo, the previously published murine bacteremia model through intraperitoneal inoculation was performed69. We used a clinical XDR ColRkp isolate with the presence of evaluated virulence factors collected from the blood of bacteremia patients. This ESBL and carbapenemase-producing ColRkp also had inactivated mgrB from IS1-like insertion between nucleotides + 71 and + 72, which was one of the most common colistin-resistant mechanisms observed in this study. Immunocompetent male C57BL/6 mice were inoculated intraperitoneally with 1 × 106 CFU of bacterial suspension with 5% porcine mucin (Sigma-Aldrich) and murine ColRkp-associated bacteremia was allowed to develop for 1 h as previously reported69. To evaluate in vivo effects of colistin, EDTA and colistin-EDTA combination, the animals with ColRkp-associated bacteremia were given a single dose of PBS (control), colistin (20 mg/kg), EDTA (40 mg/kg) and colistin-EDTA (20 mg/kg + 40 mg/kg) intraperitoneally, for a total of 4 groups with 10 animals in each group. All mice were then euthanized at 14 h post-infection. Peritoneal fluid was collected by injecting 2 mL sterile saline solution into the peritoneum, followed by gentle massage and aspiration. Peritoneal fluid samples were then serially diluted and plated on nutrient agar for counting of bacterial load. For analysis of survival in mice, the same treatment procedure was repeated once daily, and survival of mice were monitored until clinical endpoint or experimental endpoint was reached. Clinical endpoint was determined using a five-point body condition score analysing weight loss, decrease in body temperature, respiratory distress, hampered mobility, and hunched posture. Experimental endpoint was defined as 10 days post infection for mice not reaching clinical endpoint.
Data analysis
All statistical analysis was conducted using R statistic package70. Data were compared by either unpaired two-tailed Student’s t-test or unpaired two-tailed Mann–Whitney’s U test. Statistical significance was accepted at p < 0.05, p < 0.01, p < 0.001, and p < 0.0001.
Ethics approval
The study protocol was approved by the Institutional Review Board (IRB) of the Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand (COA No. 045/2020, IRB No. 774/63) was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments and comparable ethical standards. Animal care and use protocol are based upon the National Institutes of Health (NIH), USA. The protocol was approved by the Institutional Animal Care and Use Committee of the Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand (Certificate No- 033/2563, Research Project No—020/2563). The study was carried out in compliance with the ARRIVE guidelines (Animal Research: Reporting of In Vivo Experiments).
Informed consent
For this retrospective study of anonymous clinical isolates, the requirement for informed consent from patients was waived by Institutional Review Board (IRB) of the Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand (COA No. 045/2020, IRB No. 774/63).
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its additional information.
References
Sharma, L., Cruz, D. & Zhang, D. Clinical epidemiology, risk factors, and control strategies of Klebsiella pneumoniae infection. Front. Microbiol. 12, 750662–750662 (2021).
Navon-Venezia, S., Kondratyeva, K. & Carattoli, A. Klebsiella pneumoniae: A major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol. Rev. 41, 252–275 (2017).
Gogry, F. A., Siddiqui, M. T., Sultan, I. & Haq, Q. M. R. Current update on intrinsic and acquired colistin resistance mechanisms in bacteria. Front. Med. 8, 677720. https://doi.org/10.3389/fmed.2021.677720 (2021).
El-Sayed Ahmed, M.A.E.-G. et al. Colistin and its role in the Era of antibiotic resistance: An extended review (2000–2019). Emerg. Microbes Infect. 9, 868–885 (2020).
Srisakul, S. et al. Overcoming addition of phosphoethanolamine to lipid A mediated colistin resistance in Acinetobacter baumannii clinical isolates with colistin–sulbactam combination therapy. Sci. Rep. 12, 11390. https://doi.org/10.1038/s41598-022-15386-1 (2022).
Shein, A. M. S. et al. Novel colistin-EDTA combination for successful eradication of colistin-resistant Klebsiella pneumoniae catheter-related biofilm infections. Sci. Rep. 11, 1–13 (2021).
Luk-in, S. et al. Occurrence of mcr-mediated colistin resistance in Salmonella clinical isolates in Thailand. Sci. Rep. 11, 14170. https://doi.org/10.1038/s41598-021-93529-6 (2021).
Lu, Y., Feng, Y., McNally, A. & Zong, Z. The occurence of colistin-resistant hypervirulent Klebsiella pneumoniae in China. Front. Microbiol. 9, 2568 (2018).
Esposito, E. P. et al. Molecular epidemiology and virulence profiles of colistin-resistant Klebsiella pneumoniae blood isolates from the hospital agency “Ospedale dei Colli”, Naples, Italy. Front. Microbiol. 9, 1463 (2018).
Dogan, O. et al. Virulence determinants of colistin-resistant K. pneumoniae high-risk clones. Biology 10, 436 (2021).
Liu, X. et al. Emergence of colistin-resistant hypervirulent Klebsiella pneumoniae (CoR-HvKp) in China. Emerg. Microbes Infect. 11, 1–36 (2022).
Zhu, J., Wang, T., Chen, L. & Du, H. Virulence factors in hypervirulent Klebsiella pneumoniae. Front. Microbiol. 12, 734 (2021).
Jagnow, J. & Clegg, S. Klebsiella pneumoniae MrkD-mediated biofilm formation on extracellular matrix-and collagen-coated surfaces. Microbiology 149, 2397–2405 (2003).
Paczosa, M. K. & Mecsas, J. Klebsiella pneumoniae: Going on the offense with a strong defense. Microbiol. Mol. Biol. Rev. 80, 629–661 (2016).
Holden, V. I., Breen, P., Houle, S., Dozois, C. M. & Bachman, M. A. Klebsiella pneumoniae siderophores induce inflammation, bacterial dissemination, and HIF-1α stabilization during pneumonia. MBio 7, e01397-e11316 (2016).
Tsai, Y.-K. et al. Klebsiella pneumoniae outer membrane porins OmpK35 and OmpK36 play roles in both antimicrobial resistance and virulence. Antimicrob. Agents Chemother. 55, 1485–1493 (2011).
Cortés, G. et al. Molecular analysis of the contribution of the capsular polysaccharide and the lipopolysaccharide O side chain to the virulence of Klebsiella pneumoniae in a murine model of pneumonia. Infect. Immun. 70, 2583–2590 (2002).
Wang, Y., Liu, B., Grenier, D. & Yi, L. Regulatory mechanisms of the LuxS/AI-2 system and bacterial resistance. Antimicrob. Agents Chemother. 63, e01186-e11119 (2019).
García-Calderón, C. B., Casadesús, J. & Ramos-Morales, F. Rcs and PhoPQ regulatory overlap in the control of Salmonella enterica virulence. J. Bacteriol. 189, 6635–6644 (2007).
Warner, D. M., Duval, V. & Levy, S. B. The contribution of PmrAB to the virulence of a clinical isolate of Escherichia coli. Virulence 4, 634–637 (2013).
Kidd, T. J. et al. A Klebsiella pneumoniae antibiotic resistance mechanism that subdues host defences and promotes virulence. EMBO Mol. Med. 9, 430–447 (2017).
Shein, A. M. S. et al. Will there ever be cure for chronic, life-changing colistin-resistant Klebsiella pneumoniae in urinary tract infection? Front. Medicine 8 (2021).
Qureshi, Z. A. et al. Treatment outcome of bacteremia due to KPC-producing Klebsiella pneumoniae: Superiority of combination antimicrobial regimens. Antimicrob. Agents Chemother. 56, 2108–2113 (2012).
Gill, E. E., Franco, O. L. & Hancock, R. E. Antibiotic adjuvants: Diverse strategies for controlling drug-resistant pathogens. Chem. Biol. Drug Des. 85, 56–78 (2015).
Manohar, P. et al. The distribution of carbapenem-and colistin-resistance in Gram-negative bacteria from the Tamil Nadu region in India. J. Med. Microbiol. 66, 874–883 (2017).
Giani, T. et al. Epidemic diffusion of KPC carbapenemase-producing Klebsiella pneumoniae in Italy: Results of the first countrywide survey, 15 May to 30 June 2011. Eurosurveillance 18, 20489 (2013).
Olaitan, A. O. et al. Worldwide emergence of colistin resistance in Klebsiella pneumoniae from healthy humans and patients in Lao PDR, Thailand, Israel, Nigeria and France owing to inactivation of the PhoP/PhoQ regulator mgrB: An epidemiological and molecular study. Int. J. Antimicrob. Agents 44, 500–507 (2014).
Ngbede, E. O. et al. Concurrent resistance to carbapenem and colistin among Enterobacteriaceae recovered from human and animal sources in Nigeria is associated with multiple genetic mechanisms. Front. Microbiol. 12 (2021).
Bradford, P. A. et al. Correlation of β-lactamase production and colistin resistance among Enterobacteriaceae isolates from a global surveillance program. Antimicrob. Agents Chemother. 60, 1385–1392 (2015).
Phetburom, N. et al. Klebsiella pneumoniae complex harboring mcr-1, mcr-7, and mcr-8 isolates from slaughtered pigs in Thailand. Microorganisms 9, 2436 (2021).
Binsker, U., Käsbohrer, A. & Hammerl, J. A. Global colistin use: A review of the emergence of resistant Enterobacterales and the impact on their genetic basis. FEMS Microbiol. Rev. 46, fuab049 (2022).
Poirel, L. et al. The mgrB gene as a key target for acquired resistance to colistin in Klebsiella pneumoniae. J. Antimicrob. Chemother. 70, 75–80 (2015).
Haeili, M. et al. MgrB alterations mediate colistin resistance in Klebsiella pneumoniae isolates from Iran. Front. Microbiol. 8, 2470 (2017).
Kumar, A. et al. Colistin resistance due to insertional inactivation of the mgrB in Klebsiella pneumoniae of clinical origin: First report from India. Rev. Esp. Quimioter. 31, 406 (2018).
Cannatelli, A. et al. MgrB inactivation is a common mechanism of colistin resistance in KPC-producing Klebsiella pneumoniae of clinical origin. Antimicrob. Agents Chemother. 58, 5696–5703 (2014).
Cannatelli, A. et al. In vivo emergence of colistin resistance in Klebsiella pneumoniae producing KPC-type carbapenemases mediated by insertional inactivation of the PhoQ/PhoP mgrB regulator. Antimicrob. Agents Chemother. 57, 5521–5526 (2013).
Kong, Y. et al. In vivo emergence of colistin resistance in carbapenem-resistant Klebsiella pneumoniae mediated by premature termination of the mgrB gene regulator. Front. Microbiol. 12, 1643 (2021).
Jayol, A. et al. Resistance to colistin associated with a single amino acid change in protein PmrB among Klebsiella pneumoniae isolates of worldwide origin. Antimicrob. Agents Chemother. 58, 4762–4766 (2014).
Pitt, M. E. et al. Multifactorial chromosomal variants regulate polymyxin resistance in extensively drug-resistant Klebsiella pneumoniae. Microb. Genom. 4 (2018).
Cheng, Y. H. et al. Colistin resistance mechanisms in Klebsiella pneumoniae strains from Taiwan. Antimicrob. Agents Chemother. 59, 2909–2913. https://doi.org/10.1128/aac.04763-14 (2015).
Azam, M. et al. Colistin resistance among multiple sequence types of Klebsiella pneumoniae is associated with diverse resistance mechanisms: A report from India. Front. Microbiol. 12, 215 (2021).
Pragasam, A. K. et al. Molecular mechanisms of colistin resistance in Klebsiella pneumoniae causing bacteremia from India—A first report. Front. Microbiol. 7, 2135 (2017).
Wand, M. E., Bock, L. J., Bonney, L. C. & Sutton, J. M. Mechanisms of increased resistance to chlorhexidine and cross-resistance to colistin following exposure of Klebsiella pneumoniae clinical isolates to chlorhexidine. Antimicrob. Agents Chemother. 61, e01162-e11116 (2017).
Eiamphungporn, W. et al. Prevalence of the colistin resistance gene mcr-1 in colistin-resistant Escherichia coli and Klebsiella pneumoniae isolated from humans in Thailand. J. Glob. Antimicrob. Resist. 15, 32–35 (2018).
Malchione, M. D., Torres, L. M., Hartley, D. M., Koch, M. & Goodman, J. Carbapenem and colistin resistance in Enterobacteriaceae in Southeast Asia: Review and mapping of emerging and overlapping challenges. Int. J. Antimicrob. Agents 54, 381–399 (2019).
Luo, Q., Wang, Y. & Xiao, Y. Prevalence and transmission of mobilized colistin resistance (mcr) gene in bacteria common to animals and humans. Biosaf. Health 2, 71–78 (2020).
World Health Organization. Global Antimicrobial Resistance Surveillance System (GLASS): The Detection and Reporting of Colistin Resistance. (World Health Organization, 2018).
Olivares, E. et al. Clinical impact of antibiotics for the treatment of Pseudomonas aeruginosa biofilm infections. Front. Microbiol. 10, 2894 (2020).
Chauhan, A., Lebeaux, D., Ghigo, J.-M. & Beloin, C. Full and broad-spectrum in vivo eradication of catheter-associated biofilms using gentamicin-EDTA antibiotic lock therapy. Antimicrob. Agents Chemother. 56, 6310–6318 (2012).
Finnegan, S. & Percival, S. L. EDTA: An antimicrobial and antibiofilm agent for use in wound care. Adv. Wound Care 4, 415–421. https://doi.org/10.1089/wound.2014.0577 (2015).
Deris, Z. Z. et al. A secondary mode of action of polymyxins against Gram-negative bacteria involves the inhibition of NADH-quinone oxidoreductase activity. J. Antibiot. 67, 147–151 (2014).
European Committee on Antimicrobial Susceptibility Testing (EUCAST). EUCAST Reading Guide for Broth Microdilution,Version 2.0. 2020. https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Disk_test_documents/2020_manuals/Reading_guide_BMD_v_2.0_2020.pdf (accessed 7 Jan 2022).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 31st ed. CLSI supplement M100. (Clinical ans Laboratory Standard Institute, 2021).
Magiorakos, A.-P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281 (2012).
Borowiak, M. et al. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J. Antimicrob. Chemother. 72, 3317–3324 (2017).
Tolosi, R. et al. Rapid detection and quantification of plasmid-mediated colistin resistance genes (mcr-1 to mcr-5) by real-time PCR in bacterial and environmental samples. J. Appl. Microbiol. 129, 1523–1529 (2020).
Choi, Y. & Chan, A. P. PROVEAN web server: A tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31, 2745–2747 (2015).
Poirel, L., Walsh, T. R., Cuvillier, V. & Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 70, 119–123 (2011).
Ellington, M. J., Kistler, J., Livermore, D. M. & Woodford, N. Multiplex PCR for rapid detection of genes encoding acquired metallo-β-lactamases. J. Antimicrob. Chemother. 59, 321–322 (2007).
Gomes, A. É. I. et al. Selection and validation of reference genes for gene expression studies in Klebsiella pneumoniae using reverse transcription quantitative real-time PCR. Sci. Rep. 8, 1–14 (2018).
Jayol, A., Nordmann, P., Brink, A. & Poirel, L. Heteroresistance to colistin in Klebsiella pneumoniae associated with alterations in the PhoPQ regulatory system. Antimicrob. Agents Chemother. 59, 2780–2784 (2015).
Wannigama, D. L. et al. Simple fluorometric-based assay of antibiotic effectiveness for Acinetobacter baumannii biofilms. Sci. Rep. 9, 1–14 (2019).
Kifer, D., Mužinić, V. & Klarić, M. Š. Antimicrobial potency of single and combined mupirocin and monoterpenes, thymol, menthol and 1, 8-cineole against Staphylococcus aureus planktonic and biofilm growth. J. Antibiot. 69, 689–696 (2016).
Tang, H.-J. et al. In vitro efficacy of fosfomycin-containing regimens against methicillin-resistant Staphylococcus aureus in biofilms. J. Antimicrob. Chemother. 67, 944–950 (2012).
Wannigama, D. L. et al. A rapid and simple method for routine determination of antibiotic sensitivity to biofilm populations of Pseudomonas aeruginosa. Ann. Clin. Microbiol. Antimicrob. 19, 8. https://doi.org/10.1186/s12941-020-00350-6 (2020).
Phuengmaung, P. et al. Coexistence of Pseudomonas aeruginosa with Candida albicans enhances biofilm thickness through alginate-related extracellular matrix but is attenuated by N-acetyl-l-cysteine. Front. Cell Infect. Microbiol. 10, 594336–594336. https://doi.org/10.3389/fcimb.2020.594336 (2020).
Compain, F. et al. Multiplex PCR for detection of seven virulence factors and K1/K2 capsular serotypes of Klebsiella pneumoniae. J. Clin. Microbiol. 52, 4377–4380 (2014).
Vidaillac, C., Benichou, L. & Duval, R. E. In vitro synergy of colistin combinations against colistin-resistant Acinetobacter baumannii, Pseudomonas aeruginosa, and Klebsiella pneumoniae isolates. Antimicrob. Agents Chemother. 56, 4856–4861 (2012).
MacNair, C. R. et al. Overcoming mcr-1 mediated colistin resistance with colistin in combination with other antibiotics. Nat. Commun. 9, 1–8 (2018).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Acknowledgements
We thank the staff of the bacteriology division, Department of Microbiology at King Chulalongkorn Memorial Hospital, for providing the K. pneumoniae clinical isolates.
Funding
This work was supported by a grant from the 90th Year Anniversary Ratchadapiseksompotch Endowment Fund from the Faculty of Medicine and Graduate School, Chulalongkorn University, Bangkok, Thailand (batch No. 51 (1/65)). Aye Mya Sithu Shein was supported under the Chulalongkorn University Graduate Scholarship Program for ASEAN Countries. Dhammika Leshan Wannigama was supported by Chulalongkorn University Second Century Fund-C2F Fellowship, and the University of Western Australia (Overseas Research Experience Fellowship). For this project Sirirat Luk-in is also funded and supported by National Research Council of Thailand. The sponsor(s) had no role in study design; in the collection, analysis, and interpretation of data; in the writing of the report; or in the decision to submit the article for publication.
Author information
Authors and Affiliations
Contributions
A.M.S.S.: investigation, data curation, formal analysis, writing the original draft of the manuscript and contributed equally to this work as first authors. D.L.W.: conception, investigation, funding acquisition, data curation, formal analysis, supervision, editing the original draft of the manuscript. P.G.H.: supervision, critical review, editing of the manuscript and contributed equally to this work as first authors. C.H.: formal analysis, supervision, methodology, validation, critical review, editing of the manuscript and contributed equally to this work as first authors. S.A.: formal analysis, supervision, methodology, validation, critical review, editing of the manuscript and contributed equally to this work as first authors. P.H.: formal analysis, supervision, methodology, validation, critical review, editing the original draft of the manuscript and contributed equally to this work as first authors. N.C.: confocal laser scanning microscopy data acquisition, curation, methodology, validation. T.S.: Bioinformatics analysis, methodology, validation. S.L.: data acquisition, curation, formal analysis, methodology, validation, critical review and editing of the manuscript. T.L.: conception for mouse model, formal analysis, supervision, critical review and editing of the manuscript. S.N.: bacteria identification and clinical collection. U.R.: bacteria identification and clinical collection. N.K.: bacteria identification and clinical collection. S.S.: data curation, formal analysis, methodology, validation. A.A.: bacteria identification and clinical collection. M.L.: bacteria identification and clinical collection. L.Y.H.: bacteria identification and clinical collection. M.Y.: bacteria identification and clinical collection. N.K.D.R.: supervision, critical review and editing of the manuscript. T.T.: formal analysis of real-time PCR data, critical review and editing of the manuscript. N.N.: supervision, critical review and editing of the manuscript. M.A.: supervision, critical review and editing of the manuscript. P.O.: supervision, critical review and editing of the manuscript. R.K.: conception for mouse model, formal analysis, supervision, critical review and editing of the manuscript. P.P.: supervision, critical review and editing of the manuscript. V.N.B.: formal analysis, supervision, methodology, validation, critical review and editing of the manuscript. A.L.: supervision, methodology, validation, critical review and editing of the manuscript. A.K.: supervision, critical review and editing of the manuscript. T.C.: conception, funding acquisition, supervision, critical review and editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shein, A.M.S., Wannigama, D.L., Higgins, P.G. et al. High prevalence of mgrB-mediated colistin resistance among carbapenem-resistant Klebsiella pneumoniae is associated with biofilm formation, and can be overcome by colistin-EDTA combination therapy. Sci Rep 12, 12939 (2022). https://doi.org/10.1038/s41598-022-17083-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-17083-5
This article is cited by
-
Respiratory carriage of hypervirulent Klebsiella pneumoniae by indigenous populations of Malaysia
BMC Genomics (2024)
-
The effect of combinations of a glyphosate-based herbicide with various clinically used antibiotics on phenotypic traits of Gram-negative species from the ESKAPEE group
Scientific Reports (2024)
-
Enhancing colistin efficacy against Salmonella infections with a quinazoline-based dual therapeutic strategy
Scientific Reports (2024)
-
Streptomyces pratensis-Mediated Fabrication of Silver Nanoparticles and Its Applications as Antimicrobial and Anticancer
BioNanoScience (2024)
-
Solithromycin in Combination with Other Antimicrobial Agents Against the Carbapenem Resistant Klebsiella pneumoniae (CRKP)
Indian Journal of Microbiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.