Abstract
Livestock abortion is an important cause of productivity losses worldwide and many infectious causes of abortion are zoonotic pathogens that impact on human health. Little is known about the relative importance of infectious causes of livestock abortion in Africa, including in subsistence farming communities that are critically dependent on livestock for food, income, and wellbeing. We conducted a prospective cohort study of livestock abortion, supported by cross-sectional serosurveillance, to determine aetiologies of livestock abortions in livestock in Tanzania. This approach generated several important findings including detection of a Rift Valley fever virus outbreak in cattle; high prevalence of C. burnetii infection in livestock; and the first report of Neospora caninum, Toxoplasma gondii, and pestiviruses associated with livestock abortion in Tanzania. Our approach provides a model for abortion surveillance in resource-limited settings. Our findings add substantially to current knowledge in sub-Saharan Africa, providing important evidence from which to prioritise disease interventions.
Similar content being viewed by others
Introduction
Infectious causes of abortion, including early foetal loss and stillbirth, affect livestock production worldwide with the potential to cause major economic losses1. Many abortions are caused by zoonotic pathogens that also pose a risk of infection to people2. Although the economic losses can be substantial in high-production, intensive agricultural systems1, abortion-associated livestock production losses have marked negative impacts for the poorest communities in the world, many millions of whom are critically dependent on livestock for economic livelihoods, food security, health, and wellbeing3,4.
Livestock abortions can be caused by a wide range of pathogens as well as non-infectious causes such as nutritional deficiencies and trauma2. While many infectious causes of abortions occur worldwide, aetiological data based on robust laboratory-confirmed diagnoses are dominated by studies from farming systems in high- and middle-income countries5,6,7 with data lacking from low-income settings. Comparison of aetiologies across geographic areas is challenging as studies vary in methodologies and in the specific pathogens that are investigated. Furthermore, the ability to identify a specific aetiology of a livestock abortion is often relatively low, with less than 35% cases attributed to specific diagnoses, regardless of the income status of the country5,7,8. Factors that can affect the ability to establish the aetiology of livestock abortion include timing of investigations post-abortion, the sample types available and degree of autolysis of tissue samples, the range of aetiological agents selected for investigation, and diagnostic approach used.
Considerable surveillance and diagnostic infrastructure is required for robust case attribution of livestock abortions5,9, which typically limits the data available from low resource settings. For countries where data do exist, geographic patterns appear to vary quite widely. For example, Coxiella burnetii has been determined to be a common cause of abortion in sheep and goats in Canada10, and in goats in the Netherlands11, but only rarely in sheep in the United States of America (USA)8,12 or the United Kingdom (UK)9, despite evidence of widespread seropositivity in both countries13,14. Chlamydia abortus is one of the most common causes of infectious ovine abortion worldwide, including the UK, the USA and most of northern Europe but has not been found to be an abortion issue in sheep in Australia or New Zealand15 and there is little available information from the African continent, other than in a few select countries (e.g. Tunisia, Algeria, Namibia, Zimbabwe, and Morocco16,17,18,19. In tropical areas, epidemic-prone pathogens, such as Rift Valley fever virus (RVFV) which poses a severe health threat to both animals and people, are also a concern20. No less important are the endemic zoonoses such as Brucella abortus and B. melitensis, and zoonotic protozoal agents such as Toxoplasma gondii, which are of substantial concern in subsistence farming settings where people live closely with their animals3,21 or in communities where HIV predisposes the human population to severe infection e.g., with toxoplasmosis22.
In Africa, inferences about the potential role and relative importance of abortigenic agents are often made from serological studies of livestock23,24,25. This is due to a number of factors including a lack of national-level surveillance for syndromes such as abortions in livestock; restricted access to veterinary care for many subsistence farmers, who are therefore unable to investigate individual abortion events; and limitations in diagnostic laboratory capacity (e.g., little to no access to histopathology services or molecular diagnostic tests that could help to generate data on different aetiologies of abortion). While serological data are valuable to identify the presence of abortigenic agents circulating within a livestock population, there is also a need to recognise the limitations of such data and the inferences that can be made based on it26,27. Livestock seropositivity indicates only that an animal has been exposed to a pathogen and does not necessarily indicate current disease status or causality of an abortion event. Additionally, chronic low-level carriage of some abortigenic agents (e.g., C. burnetii) in healthy animals complicates attribution of abortion. To date, in sub-Saharan Africa, there remains an almost complete absence of published data on livestock abortion aetiologies determined through confirmatory diagnostic testing, resulting in a very weak evidence base on which to prioritise animal health interventions.
In Tanzania, livestock diseases are prioritised for prevention and control largely on the basis of the potential for transboundary spread and zoonotic impacts28. A recent exercise to prioritise zoonotic diseases ranked two abortigenic agents, Brucella and RVFV amongst the top six zoonoses for prevention and control29. While there is clear evidence that brucellosis and Rift Valley fever (RVF) are both diseases of public health concern in Tanzania30,31,32, no data currently exist on the relative importance of these or other zoonotic pathogens, such as C. burnetii, Leptospira spp., C. abortus or T. gondii, as causes of livestock abortion. Nor are there any data available on the contribution of other abortigenic pathogens of global importance, including N. caninum, bluetongue virus and pestiviruses such as bovine viral diarrhoea virus (BVDV) and border disease virus (BDV). Without robust aetiological data, it is unlikely that current approaches to livestock disease prioritisation will reflect the true impact of abortigenic pathogens, with the risk that scarce resources for disease control and prevention will be targeted ineffectively and achieve sub-optimal productivity and public health outcomes.
As 50% of Tanzanian households keep livestock33 and livestock in Tanzania are concentrated in the north, we sought to address knowledge gaps around the aetiology of livestock abortion in northern Tanzania through a prospective cohort study conducted in the Arusha, Kilimanjaro, and Manyara Regions that span a range of agro-ecological settings34. We used a combination of field investigation and laboratory diagnosis of abortion events in cattle, goats, and sheep to determine aetiologies of livestock abortions for a targeted selection of known abortigenic agents. For these selected agents, we developed a set of case definitions to attribute abortion events to a specific aetiological agent based on molecular diagnostic assays in the absence of comprehensive histopathology data. In the same study area, we also carried out serological analyses of a large sample of livestock sera collected by cross-sectional studies in the same area to determine seroprevalence to the same range of abortigenic pathogens at a regional level.
Results
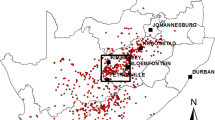
Figure 1 shows the location of sampling for the serological cross-sectional and abortion cohort studies.
Distribution of villages in cross-sectional studies for serologic baseline testing (2013–2016; red dots) and abortion cohort sampling wards (2017–2019; yellow shading) with number of abortion events investigated, Arusha, Kilimanjaro, and Manyara Regions, northern Tanzania. Map created using QGIS version 3.22.1 (https://www.qgis.org/en/site/). Shapefiles were downloaded from GADM (https://gadm.org/) and ICPAC (geoportal.icpac.net).
Cross-sectional sample serology
Samples were available from 3629 cattle, 3344 goats, and 2628 sheep from 567 households in 43 villages. Figure 2 summarises the cross-sectional seroprevalence of pathogens in ruminant livestock sampled in northern Tanzania. Seroprevalence at individual level for cross sectional livestock with Clopper Pearson 95% confidence intervals is detailed in supplementary materials (S1, A).
Abortion cohort study
During the prospective cohort study period, a total of 215 abortion events were investigated, 71 (33%) in cattle, 100 (46.5%) in goats, and 44 (20.5%) in sheep. The numbers of each sample type collected for abortion events are detailed in Table 1.
Abortion cohort acute serology
The acute seroprevalence results by pathogen in the abortion cohort was broadly similar to the cross-sectional survey. A table showing these results is available in the supplementary materials (S1; A).
Diagnosis of abortion aetiology
We attributed 42 (19.5%) of 215 abortion events to one or more of the abortigenic pathogens included in our diagnostic panel. Table 2 summarises the number of abortion events with positive diagnostic test results that met the study case definitions. The strength of evidence for attributing the cause of abortion for each pathogen is detailed in the methods section (Table 5). Histopathology and immunohistochemistry (IHC) results are shown in supplementary materials S1, B.
For some bacterial pathogens, despite evidence of infection in the dam, laboratory data was not considered sufficient to support a diagnosis of abortion associated with the pathogen. This was particularly true for C. burnetii, which was detected by qPCR with a threshold cycle (Ct) value < 40 in 16 (22.5%) of 71 cattle, 24 (24.5%) of 98 goats, and 12 (27.3%) of 44 sheep, however only six (11.5%) of these 52 animals met the study case definition (Ct ≤ 27) for C. burnetii as a presumptive aetiology of abortion. For C. abortus, although seroconversion between acute and convalescent samples to C. abortus was detected in one (1.8%) of 56 goats with paired sera available for testing, this animal had a negative vaginal swab qPCR result and therefore did not meet our case definition for C. abortus-associated abortion.
Of the protozoal pathogens, Neospora DNA was diagnosed by nested conventional polymerase chain reaction (PCR) in nine (12.7%) of 71 cattle and one (1.0%) of 98 goat abortion events. In one cattle case, a co-infection with BHV-1 was also detected suggesting a multi-factorial aetiology. Toxoplasma DNA was detected by nested conventional PCR in one (2.3%) of 44 sheep abortion events. The single T. gondii PCR positive sheep sample was genotyped at 7/10 marker and was found to have type II alleles at all of them, indicative of a type II clonal genotype infection.
Of the viral pathogens, RVF was most commonly diagnosed, with RVFV RNA detected by reverse transcriptase qPCR (RT-qPCR) in 14 (19.7%) of 71 cattle abortion events. Evidence of seroconversion was detected in one animal that was positive by RT-qPCR. The other 13 RT-qPCR positive animals were positive in both acute and convalescent sera, meeting World Organisation for Animal Health (OIE) guidelines for definitive diagnosis of infection. Pestiviruses were also commonly detected with two (2.8%) of 71 cattle, six (13.6%) of 44 sheep, and one (1.0%) of 100 goat abortion events positive by RT-qPCR.
Pestivirus speciation results
Among the nine animals that tested positive for pestiviruses by RT-qPCR, NPro coding sequence was obtained from six samples for phylogenetic analysis (Fig. 3). BVDV infection was confirmed in two animals (one cow and one sheep) with a distinct genotype detected in each case. The cattle sample (SF1-190) was most similar to reference sequences from BVDV genotype 1d whereas the sheep sample (SD1-147) was most similar to reference sequences from BVDV genotype 1b. Sequences obtained from the remaining four samples (SD1-144, SD1-145, SD1-155, SD1-160), all obtained from sheep, were similar to each other but appeared to be distinct from all of the reference sequences. These sequences represent a potentially novel, previously unreported pestivirus species and formed a separate clade on phylogenetic analysis. The sequences were assigned accession numbers OU548650-OU548655 by the European Nucleotide Archive (https://www.ebi.ac.uk/ena/browser/home) and, for the purposes of that submission, this novel clade was named Pestivirus abortion/sheep/SD1/2018/Tanzania.

taken from aborting dam; SF denotes swab taken from aborted foetus. APPV atypical porcine pestivirus, BDV border disease virus, BVDV bovine viral diarrhoea virus, CSFV classical swine fever virus.
Results of evolutionary analysis of the NPro coding region from pestivirus sequences obtained from cattle and sheep sampled in the abortion cohort study, northern Tanzania. The evolutionary history was inferred using the Maximum Likelihood method and Tamura-Nei model of nucleotide substitution35. The tree with the highest log likelihood is shown. The percentage of trees from 500 Bootstrap repeats in which the associated taxa clustered together is shown next to the branches. Evolutionary analyses were conducted in MEGA X36. Reference sequences representing nine pestivirus species were also included37 and are labelled with virus species (A–H,K), genotype and strain name and Genbank accession numbers. Pestivirus species are also shown by bracket labels on the right of the figure. Samples analysed in this study are shown in bold and labelled with unique identifiers (SD/F1-XXX), where SD denotes vaginal swab
Discussion
To our knowledge, this is the first multi-pathogen study of the aetiologies of livestock abortion in Tanzania. We developed and applied an approach designed to optimise abortion investigations in resource limited settings, complemented by traditional cross-sectional surveillance to generate a more comprehensive understanding of the distribution and impact of several important abortigenic pathogens. This approach generated several new insights for Tanzania including: (a) detection of an RVF outbreak in cattle during an inter-epidemic period; (b) relatively high prevalence of C. burnetii infection in cattle, goats and sheep; (c) the first demonstration of N. caninum, T. gondii, and pestiviruses, including novel pestivirus species, associated with livestock abortion in Tanzania, including detection in less typical hosts (for example, N. caninum in goats); but, contrary to prior expectations, (d) limited or no detection of Brucella spp., C. abortus, and Leptospira spp. in cases of livestock abortion. Our findings provide important information on the health and production impacts of a range of livestock pathogens in East Africa with relevance for prioritising disease interventions in the region.
Our study demonstrates the importance of zoonotic pathogens as causes of abortion in ruminants in Tanzania, with implications for human as well as animal health. Two important zoonotic pathogens were diagnosed in multiple abortion events by our study: RVFV, a virus that can cause devastating outbreaks of human and animal disease38,39, and C. burnetii, an important bacterial cause of human febrile illness in Tanzania40. RVFV is a high priority zoonosis in East Africa and outbreaks are frequently associated with livestock abortion storms20. Direct animal to human transmission of infection can occur following inhalation of aerosolized virus particles or during close contact with tissues and bodily fluids of infected animals, for example, when assisting with parturition or handling aborted foetuses41,42. Although RVF usually occurs as large outbreaks occurring every 10–20 years, there is increasing serological evidence of transmission in the period between these epidemics43. This study detected the occurrence of an apparent outbreak in cattle that was previously unreported in Tanzania, despite RVF outbreaks being detected in Kenya, Uganda, and Rwanda at the same time41,44. Although an alert of RVF was issued in Tanzania in June 2018 at the time of this study45, to our knowledge, no human cases were reported. Our finding highlights the value of livestock abortion surveillance in early detection of this epidemic-prone pathogen and also demonstrates the value of molecular diagnostic capacity for detection of RVF livestock cases.
Coxiella burnetii is a pathogen of concern in Europe, where outbreaks of infection in small ruminant livestock have also led to outbreaks of human disease46. C. burnetii has also been detected in humans and in a wide range of animal species across Africa47,48, and accounts for notable proportions of severe human febrile illness in Africa (Vanderburg et al., 2014), including in northern Tanzania40,49. Our results provide important evidence of the impact of C. burnetii as a cause of livestock reproductive losses, particularly in small ruminants, revealing a dual human and animal health burden of this zoonotic pathogen. In our abortion cohort study, we applied a stringent case definition that exceeded the minimum guidelines stated by the OIE for diagnosis of C. burnetii-associated abortion. Using this case definition, we diagnosed six cases of C. burnetii-associated abortion, but it is also worth noting that C. burnetii DNA was detected in a total of 52 (24.4%) abortion events indicating that shedding of this zoonotic bacterium is even more prevalent than the number of attributed cases would suggest. Based on our findings, we suggest that the development of preventive and control measures against C. burnetii should be a priority for future work that could have substantial benefits for human and animal health.
Our study confirmed T. gondii, another zoonotic pathogen, is associated with livestock abortion in Tanzania. Toxoplasmosis is also an important, sometimes fatal, zoonosis in Tanzania, particularly among immunocompromised people22. There are limited data on genotypes present in Africa but the Type II clonal lineage found in the single PCR positive sheep abortion event is known to be dominant in North and East Africa so aligns with previous studies50. The cross-sectional serological data showing widespread exposure of T. gondii in sheep and goats are consistent with previous published findings of high seroprevalence in people and animals, including livestock51,52 and wild felid species in East Africa53. Given that T. gondii is one of the most important abortigenic agents of sheep worldwide, it is perhaps surprising that we did not detect more cases of T. gondii as a cause of sheep abortion in this study. However, high flock seroprevalences are typically associated with low rates of abortion due to the development of protective immunity in ewes infected with T. gondii prior to mating54.
Other zoonotic pathogens were detected infrequently in the abortion cohort study, with only a single case attributed to Brucella spp. in a cow, a single detection of seroconversion to C. abortus in a goat, and no evidence of abortions associated with Leptospira spp. in any livestock species. Although C. abortus has been reported as a cause of abortion in sheep elsewhere in northern and southern Africa55,56,57, our study generated little evidence that C. abortus was contributing to the burden of production losses in sheep in northern Tanzania at the time of our study. Our cross-sectional serological findings also support the conclusion that levels of exposure in small ruminants are very low across northern Tanzania. However, these findings do raise concerns about the potential impact should infection be introduced to this predominantly naïve population in the future.
Both Brucella spp. and Leptospira spp. are recognised as important zoonoses in northern Tanzania, causing large proportions of severe human febrile illness21,49,58,59 and are known to be circulating in livestock in these areas of Tanzania. Seroprevalence of Brucella spp. was low in all three livestock species (Fig. 2), which is consistent with the low frequency of detection in abortion cases. In contrast, although Leptospira was not detected by qPCR in any abortion event investigated, evidence of exposure to Leptospira borgpetersenii serovar Hardjo, and Leptospira interrogans serovar Hardjo (subsequently referred to as Leptospira Hardjo serovars) was relatively high in cattle in the cross-sectional study. Several possible explanations may explain this discrepancy. Similar to T. gondii, prior exposure to Leptospira serovars may lead to protective immunity prior to animals becoming pregnant. Additionally, livestock abortions associated with Leptospira are often seen in outbreaks or ‘abortion storms’ following either a new introduction into a herd or a seasonal outbreak where flooding or wet weather conditions support transmission of novel serovars60. The timespan of our project meant that we may have missed intermittent or infrequent abortion storms associated with Leptospira in livestock. However, even accounting for these limitations, our study suggests that in these agricultural settings, the proportion of abortions attributable to Brucella spp. and Leptospira spp. may be lower than is widely perceived. Alongside other constraints to the uptake and use of livestock brucellosis vaccines these data raise questions about the likely effectiveness and sustainability of livestock vaccination-based control strategies in this context if based on a farmer pays model only. To effectively mitigate both the animal and human impacts of brucellosis a diversity of strategies may be needed to achieve the vaccination coverage required to reduce animal production impacts and also to reduce the substantial human health burden from this disease.
In addition to zoonotic pathogens, our study also provides valuable data on several production-limiting infections of ruminants that are often over-looked in sub-Saharan Africa. In particular, while N. caninum and pestiviruses are considered important causes of abortions and productivity loss in Europe and the USA, there are few data available describing their impact in African livestock. Our results provide the first direct evidence of livestock abortion associated with N. caninum infection in Tanzania, a conclusion which is supported by a positive relationship between N caninum seroprevalence and in-herd abortion rates in cattle detected by earlier work in our study area61. In this study, N. caninum was predominately diagnosed among cattle abortions, but notably also in one goat abortion event. From the abortion cohort we have evidence that N. caninum was a likely cause of abortions in this cohort. From the larger cross-sectional serological studies, the exposure to N. caninum is very low, particularly in goats (1.7%) and sheep (1.5%). Although abortion and foetal pathology associated with N. caninum has been described in goats and sheep62,63,64, natural infection of N. caninum in small ruminants is not widely reported. Given the importance of small ruminants within livestock economies and for food security in Africa65,66, further investigation of the patterns and impact of N. caninum infection in small ruminants are warranted.
Pestiviruses were the most commonly diagnosed pathogen associated with abortion in sheep in this study, but pestiviruses are rarely discussed as priority production-limiting pathogens in livestock in Africa. Analysis of the six pestivirus sequences obtained in this study identified at least three different viral lineages associated with abortion in livestock in northern Tanzania including a novel lineage in sheep. Based on our analysis of the NPro coding region, this novel lineage appears to be distinct from other pestiviruses and is likely to be a new, previously unreported pestivirus species. Notably, several of the most similar pestivirus species were detected in sympatric wildlife species including giraffe in Kenya (Genbank accession numbers U80907 and AY163647). As wildlife interactions with livestock are common in many areas of Tanzania, we hypothesise that spill-over from wildlife may also be involved in the epidemiology of this novel pestivirus lineage, which appears to be a contributing cause of ovine reproductive loss in northern Tanzania.
Serological data from the cross-sectional study confirmed widespread exposure to BHV-1 in the study area (Fig. 2), and previous studies in this area have demonstrated an association between seropositivity and cattle abortion67. However, the evidence for BHV-1 as a cause of abortion in this study is relatively weak. As latent BHV-1 infection may lead to false positive results by molecular detection68, we relied on serology to diagnose infection. Two cattle showed evidence of BHV-1 seroconversion suggesting recent infection which may be implicated in the aetiology of abortion in these cases but further data are needed to evaluate the impact of BHV-1 pathogen on livestock reproduction in Tanzania and our findings highlight the challenges of attribution in settings where endemic pathogens may co-exist in cattle herds69.
BTV, which causes production losses in livestock in Europe70,71, was not detected by RT-qPCR in any samples from livestock abortion events, despite high BTV seroprevalences detected in all species in the cross-sectional study population. Serological evidence of infection is widespread East Africa72,73,74 and several studies have also demonstrated viraemia using RT-qPCR assays, reporting infection prevalence as high as 88.9% in cattle in Kenya74 and 56.0% in goats in Uganda73. Collectively, these studies suggest that BTV infection is endemic in livestock in East Africa. However, research indicates that African breeds of livestock may be relatively resistant to clinical disease associated with BTV. Studies from southern Africa have indicated that indigenous African breeds of livestock are less susceptible to clinical disease from BTV infection than European breeds75. Furthermore, transplacental foetal infection is only associated with certain BTV strains, especially attenuated vaccine strains76. Our findings supported by these trends in BTV epidemiology and pathogenesis suggest that BTV is not a major cause of livestock abortions in northern Tanzania and underscores the need for caution in making inferences about livestock diseases and causes of abortion in particular based solely on seroprevalence data.
The approach described in our abortion cohort study enabled us to attribute approximately 20% of reported abortion events to a specific aetiology. Our success rate was not dissimilar to the quoted rate of aetiological abortion diagnosis in high income settings (35%)5,7,8 despite working in a challenging and resource-limited environment. Our approach was specifically designed to optimise abortion investigations in a resource-limited setting, where veterinary surveillance may be limited. Our focus on the use of molecular diagnostics to detect a range of selected pathogens proved to be valuable, particularly for high priority pathogens such as RVFV. Although 215 livestock abortions were investigated, the precision of subgroup analyses, such as by animal species or geographic area becomes, by necessity, diminished in power. The serological cross-sectional study sampled a much greater number of livestock enabling us to have a robust baseline of exposure to the pathogens we were investigating. While molecular diagnostic capability was established for several known abortigenic agents, it was not possible for our diagnostic panel to be fully comprehensive of all possible aetiologies of abortion and did not include agents such as Campylobacter spp., Listeria spp., and Salmonella enterica serovars that are known to be important causes of livestock abortion worldwide. Non-infectious causes were also not considered for investigation and these two factors may account for some of the undiagnosed cases.
Although histopathology is conventionally considered the most robust approach for attributing causality in livestock abortion for some pathogens, histopathology was of limited value in this study due to the limited availability and poor quality of tissues samples obtained through field investigations that were often conducted in remote settings. Additional challenges to inference of causality were encountered due to the nature of our study site. In areas such as rural Tanzania, where remote locations, high ambient temperatures and scavenging animals mean that the type and quality of diagnostic samples is often sub-optimal, inference about the cause can be particularly challenging. For example, placental tissues were not available for sampling from many small ruminants, and even where placental tissues were available for collection, most tissue samples showed evidence of significant autolysis, which precluded a definitive diagnosis in some cases. However, despite these sampling limitations, molecular diagnostics provided a wealth of information regarding the presence of abortigenic agents in swab and tissue samples, and therefore we recommend that molecular diagnostic capacity be invested in and included as a core element of abortion surveillance studies to infer aetiology in resource-limited settings.
In considering appropriate criteria for attributing the aetiology of abortion events reported to this study, we became aware of variations in case definitions and the limitations of the evidence base underpinning attribution criteria for livestock abortions. While methods for assigning attribution in situations of imperfect diagnostic testing are well-developed and widely adopted in aetiological studies of human diseases77, these questions are rarely addressed in veterinary investigations. Attribution was particularly problematic for C. burnetii, as the pathogen is frequently detected in vaginal fluids at parturition in clinically healthy animals78 and environmental contamination of placental material or foetal surfaces may also lead to false positives in high risk environments79. In this study, we developed a case definition for C. burnetii based on OIE guidelines80,81 but also driven by population-level data generated by this study. Further development of this case definition would be beneficial to understand how clinical history, gross pathology (where available), measured bacterial load and other adjunct diagnostics could be used to more confidently attribute abortion cases both within and outside the Tanzanian context.
Substantial differences in reporting rates between wards further limits the representativeness of our sample. Important differences in livestock seroprevalence between production systems have been observed in northern Tanzania82 as well as in other areas of East Africa83. Spatial heterogeneity in livestock exposure risk, with clustering in seropositivity at the village-level, have also been described for pathogens such as Brucella84. It is therefore possible that sample collection in more administrative areas, with improved representation of pastoral and agropastoral livestock keepers, would reveal different frequencies of pathogen detection than described here. Our data should therefore be considered as only indicative of the relative importance of different pathogens as causes of livestock abortion in northern Tanzania as a whole.
A key insight from our study is that, apart from RVF, none of the abortigenic pathogens identified by this study have been considered important pathogens in Tanzania, or prioritised for interventions. Gaps remain in our understanding of the causes of livestock abortion in Tanzania. However, even with testing a limited range of pathogens, our study has been able to generate data that adds substantially to the existing evidence base for the development of interventions to reduce the burden of production losses associated with livestock abortions. Our study further highlights the value and importance of establishing laboratory-based molecular diagnostics and developing clear case definitions and diagnostic criteria for use in resource-limited settings. With laboratory capacity and technical capability expanding rapidly in Africa, there is enormous potential and an urgent need for the veterinary sector to shift towards more evidence-based policy and decision-making, with the potential for this transition to lead to substantial improvements in animal and human health.
Methods
Research ethics
Ethics approval for this research was granted by Kilimanjaro Christian Medical Centre (KCMC) Ethics Committee (No. 535 & No. 832); National Institute of Medical Research (NIMR), Tanzania (NIMR/HQ/R.8a/Vol.IX/1522 & NIMR/HQ/R.8a/Vol.IX/2028); Research Ethics Coordinating Committee and the Institutional Review Board for Clinical Investigations of Duke University Health System in the United States (Pro00037356), University of Otago Ethics Committee (H15/069 & H17/069); and University of Glasgow College of Medical Veterinary and Life Sciences Ethics Committee (200140152 & 200170006). Sample shipments between the UK and Tanzania were performed in accordance with the Nagoya Protocol on Access to Genetic Resources under Material Transfer Agreements between KCMC and University of Glasgow, and between KCMC and Moredun Research Institute (MRI) with authorisation for export and import, respectively, from the Tanzanian Ministry of Livestock and Fisheries (permit number VIC/ZIS/8582), and The Scottish Government’s Agriculture, Food and Rural Communities Directorate (TARP(S) 2019/07). All experiments were performed in accordance with relevant guidelines and regulations.
Cross-sectional and prospective cohort studies
Both cross-sectional and prospective cohort studies were used to collect the data reported. The cross-sectional livestock serological studies were undertaken to provide a baseline for exposure to a range of abortigenic pathogens in the study area. The prospective cohort study was undertaken on livestock that had aborted that were reported to the project field team. The details of both these study types are described below.
Cross-sectional serological studies
Two cross-sectional livestock serological studies were performed from 2013 through 2016. The full details of the study methods, including site selection and sampling strategy, have been extensively described elsewhere82,85. In brief, blood samples were collected from clinically healthy cattle, goats, and sheep at sampled households across six districts in Arusha Region, three districts in Kilimanjaro Region, and four districts in Manyara Region. Multi-stage sampling was used to select villages, households, and individual animals. From each randomly selected household, whole blood samples were collected from a maximum of 15 animals per livestock species. Serum was separated from coagulated whole blood samples by centrifugation (≤ 1300×g for 10 min at room temperature) and aliquoted to create replicate sets. Samples were transported to the Zoonoses Laboratory at Kilimanjaro Clinical Research Institute (KCRI), Moshi, Tanzania, for archiving at − 80 °C prior to testing. Cattle, goat, and sheep sera were tested for exposure to Brucella spp., C. burnetii, and RVFV using commercial ELISA, as per manufacturer’s instructions (Table 3). Cattle sera were additionally tested for exposure to Leptospira Hardjo serovars, BHV-1, and BVDV (Table 3). Absorbance was measured using a MultiSkan FC plate reader (Fisher Scientific, Leicestershire, UK). For each sample, one of the duplicate aliquots was heat treated at 56 °C for two hours and archived at − 80 °C until it was shipped, on dry ice, to MRI or University of Glasgow, UK, for testing. All or a random selection of available samples were tested by indirect ELISA for N. caninum and T. gondii, and C. abortus (goats and sheep only) as described previously86,87,88. A random selection of cross-sectional sera was also tested for exposure BTV (Table 3). Clopper-Pearson 95% confidence intervals were calculated using the binom package89 in R statistical software (version 3.6.0).
Abortion cohort study
The prospective abortion cohort study was undertaken from October 2017 through September 2019 in 13 wards in Arusha, Kilimanjaro, and Manyara Regions of northern Tanzania (Fig. 1). Study wards were selected from those included in the cross-sectional exposure studies. Two wards were selected purposively from among those included in these cross-sectional studies because of existing relationships with the livestock-keeping community that were expected to promote participation. The remaining eleven wards were selected randomly from the wards targeted in the cross-sectional studies.
Livestock field officers (LFOs), animal health professionals working for the Tanzanian Ministry of Livestock and Fisheries, were recruited to participate in the study with one LFO responsible for a single ward. At the start of the study the LFOs attended a training course on causes of livestock abortion, as well as the practical investigation and management of abortion events. Training included the safe collection, storage, and transport of biological samples from the aborting dam, foetus, and placental material, the use of personal protective equipment for the safe handling and disposal of aborted material and methods for data collection. Following the training course, LFOs were asked to disseminate information about the study to the community living within their respective wards, requesting livestock owners to report any instances of livestock abortion to them so that an investigation could be carried out. Following receipt of a report, the LFO was asked to pass on the information to the research coordinator, a veterinarian, for investigation. For the purposes of this study, an abortion event was defined as: farmer reported visual evidence of premature foetal loss or stillbirth in cattle, goats, or sheep. The abortion was considered eligible for inclusion in this study if the research field team or LFO could attend within 72 h of the event occurring. Two investigation options were available. Where LFOs carried out the investigation, vaginal swabs and 10 ml blood samples were collected in red-top vacutainer tubes (Becton, Dickinson and Company, New Jersey, USA) from the dam(s) that had aborted, as identified by the livestock keeper. Where the veterinarian-led research team was able to join the LFO’s within 72 h, additional samples were collected, including swabs from the surface of the foetus exterior and placental (cotyledon) tissues, when available. Duplicate swab samples (breakpoint shaft, viscose tip; Technical Service Consultants Ltd, Lancashire, UK) and duplicate tissue samples were stored in 1 ml DNA/RNA shield (Zymo Research, California, USA) to inactivate infectious agents and preserve nucleic acid integrity prior to molecular testing. Due to practical limitations, samples comprehensive sets of tissues including foetal fluids for bacterial culture were not collected in this study. Tissue samples were also stored in duplicate in 10% buffered formal saline to preserve tissue structure for histopathology and IHC. Samples were transported to the Zoonoses Laboratory at KCRI, for archiving at − 80 °C prior to testing against a panel of abortigenic agents.
Abortion cohort serological testing
Sera from dams that aborted were tested using the range of host and pathogen specific assays described in Table 3. For each sample, one of these duplicate tubes was heat treated at 56 °C for two hours prior to export to the UK for testing. The remaining tube was retained for testing at KCRI and stored at − 80 °C. Households were re-visited 4–6 weeks later to collect a convalescent blood sample from the same dam(s). Seroconversion was defined as a negative acute serum with a positive convalescent serum.
DNA and RNA extraction
RNA was extracted using a RNeasy® Mini kit (QIAGEN, Hilden, Germany) following manufacturer’s instructions. For RNA extraction from the placental tissues, a sterile scalpel was used to add approximately 25 mg of tissue to 600 µl RLT buffer in a sterile bead beating tube containing ceramic beads (CK28, Precellys, Bertin Instruments, France). Bead beating was performed for one minute at 3450 rpm using a MiniBeadBeater-16 (BioSpec Products, Inc., Oklahoma, USA). The liquid was aliquoted into a microcentrifuge tube with 590 µl nuclease-free water and 10 µl Proteinase K (QIAGEN, Hilden, Germany). For RNA extraction from swabs, 200 µl sample liquid was added to 600 µl RLT buffer, and 10 µl Proteinase K. All tubes were then incubated at 55 °C for 10 min. Once cooled to room temperature, tubes were centrifuged at 10,000×g for three minutes. The supernatant was transferred to a new microcentrifuge tube where half the volume of ethanol (96–99%) was added to the tube (e.g., 450 µl ethanol to 900 µl supernatant) and mixed by pipetting up and down gently. RNA was extracted from 700 µl of the lysate/ethanol mix using a final elution volume of 50 µl.
DNA was extracted using a DNeasy® blood and tissue kit (QIAGEN), following manufacturer’s instructions. For DNA extraction from placental tissues, a sterile scalpel was used to add approximately 25 mg of placenta into a microcentrifuge tube with 180 µl ATL buffer and 20 µl Proteinase K. The tubes were incubated at 56 °C, vortexing occasionally, until the tissue was lysed, after which, 200 µl AL buffer was added, and the tube was vortexed briefly. For DNA extraction from swabs, 200 µl sample liquid was added to 200 µl AL buffer with 20 µl Proteinase K and vortexed. All tubes were incubated at 56 °C for 10 min. When the buffer/sample mix had cooled to room temperature, 200 µl ethanol (96–99%) was added, and the tube was vortexed. DNA was extracted from the entire volume of the lysate/ethanol mix. Final elution volumes for swabs and placental tissue samples were 100 µl and 200 µl, respectively.
All batches of nucleic acid extractions included a negative extraction control where sample was replaced by nuclease-free water and the run alongside the nucleic acid extraction process.
Molecular testing
The molecular targets for all pathogens are listed in Table 4. Primer and probe sequences and cycling conditions for each PCR, qPCR, and RT-qPCR assay, are detailed in supplementary materials (S1; Table C). DNA or RNA extracts from positive controls listed in Table 4 were used for individual PCR, qPCR, and RT-qPCR assays. All available vaginal swab, foetal swab, and placental tissue samples for all three livestock species were for all pathogens included in this study. Samples were tested in duplicate except for pathogens tested using commercial kits (BTV and pestiviruses) where single samples were tested. Nuclease-free water and extraction controls were included in each PCR, qPCR, and RT-PCR run as negative controls. Samples were considered positive only if negative extraction controls and negative controls showed no evidence of amplification and the internal and external positive controls also amplified. Where single wells amplified for samples tested in duplicate, the assay was repeated. Unless specified, testing was performed at KCRI.
Previously published qPCR assays designed and validated for the diagnosis of infection with the respective pathogens were used to detect the presence of Brucella spp. (two gene targets91,92), C. burnetii46, C. abortus93, and pathogenic Leptospira spp.94. These qPCRs were performed in single well reactions using a QuantiNova® Probe kit (QIAGEN) with 5 μl of DNA template (or 2.5 μl per target for Brucella spp.).
Nested conventional PCR assays for the presence of Neospora spp. and Toxoplasma gondii were performed targeting the internal transcribed spacer (ITS1) region between the 18S and 5.8S rRNA genes98. The primary (1°) nested PCR was performed using 2 μl of DNA template. The secondary (2°) nested PCR was performed using with 2 μl of a 1:5 diluted primary PCR product. Both 1° and 2° assays were performed with 2 × PCR master mix (Promega Corporation, Wisconsin, USA). The final PCR product from the 2° nested PCR was checked by electrophoresis in a 1.5% agarose gel. Samples were considered positive if the size of bands visible under UV light were congruous with those expected of the Neospora (249 bp) and Toxoplasma (227 bp) targets.
The presence of BTV and pestivirus RNA in samples was determined by RT-qPCR using commercial kit-based assays validated for the detection of the respective pathogens (Table 4). Single samples were tested, as per kit instructions, using 5 μl of RNA template. Finally, the presence of RVFV RNA was assessed using the QuantiNova® RT-PCR probe kit (QIAGEN). Samples were tested in duplicate using 5 μl of RNA template per well.
Pestivirus speciation
As commercial kit-based RT-qPCR assays for pestiviruses do not distinguish between the different pestivirus species, genotyping was performed at MRI to determine the infecting pestivirus species. The RNA extracts that were positive in pan-pestivirus RT-PCR screening assays were subjected to RT-PCR and then nested PCR using four primer pairs developed for genotyping of livestock pestiviruses: 324–326 (5’UTR); BD1-BD3/BD4 (Npro); BD1-BD2 (Npro-C); and 17f-1400r (UTR-Erns) to determine the infecting virus type99,98,101. RT-PCR products of the expected sizes were purified from agarose gels and submitted for sequencing using the relevant PCR primers (Eurofins-MWG). Sequence traces for each sample were assembled using SeqMan Pro (DNASTAR Lasergene) and consensus sequences that excluded the terminal primer sites were generated. The 504 nt Npro coding sequences were aligned with a set of NPro reference sequences that included representatives of pestivirus species A-H plus K, as described previously37. Alignment was performed using MAFFT (https://mafft.cbrc.jp/alignment/). Maximum likelihood (ML) phylogenetic analysis was performed using MEGA version X. Model selection was used to identify the best evolutionary model for the dataset. The ML analysis was bootstrapped 500 times to estimate the reliability of the resulting phylogeny.
Additional pathogen genotyping
T. gondii genotyping was also performed to determine the infecting sub-type. The methods of this genotyping work are detailed in the supplementary materials S1; D.
Pathology and immunohistochemistry (IHC)
Histopathology was performed on placental tissue samples where available. For samples that tested positive for C. burnetii by qPCR, C. burnetii IHC was also performed. The methods for the histopathology and IHC are given in supplementary materials S1; B.
Diagnosis of abortion aetiology
For the purposes of our study, we determined a set of case definitions that we used to attribute livestock abortion events to a specific aetiology. These case definitions are shown in Table 5. Case definitions were based on OIE guidelines for the diagnosis of each pathogen. Due to the challenges in systematically collecting tissue samples for histopathology from abortion cases (e.g., lack of available tissue in many cases), we designed these diagnostic criteria (Table 5) to be independent of histopathological examination but used evidence from histopathological examination to support our diagnosis where samples existed to support this diagnostic approach. For N. caninum, for which no OIE diagnostic guidelines exist, the case definition for was based on criteria used by MRI, UK102, a specialist diagnostic veterinary laboratory. For C. burnetii, OIE guidelines indicate that definitive diagnosis of infectious abortions resulting from C. burnetii infection can only be robustly achieved at the flock or herd level, but also that bacterial load is predictive of causality in individual animals80,81. Therefore, to determine the C. burnetii case definitions, we used a two-step approach. First, we used the minimum bacterial load associated with abortions as stated by the OIE (> 104 bacteria per swab or per gram of tissue, equivalent to Ct values of 37.5 and 42.7 respectively on our PCR platform). Secondly, using a histogram of the Ct results at the study population-level, we identified a bimodal distribution in the results, suggesting two distinct biological processes within our sampled population (see supplementary material S1; E). We used this frequency distribution, supported by histopathology and IHC results to select a conservative threshold value using a cut-off Ct value of ≤ 27 for attributing abortion events to C. burnetii in this study.
Confidence in this study’s diagnostic case definition was determined to be either ‘definitive’, ‘presumptive’ or ‘suggestive’, depending on how closely our diagnostic approach aligned with the OIE guidelines for diagnosis of abortions caused by each pathogen. Where our study diagnostic approach met OIE definitions for a definitive diagnosis, we classified our diagnosis as ‘definitive’ and had the highest confidence in our aetiologic diagnosis. Where our study diagnostic approach met OIE definitions for a presumptive diagnosis or for N. caninum where no OIE guidelines exist, we classified our diagnosis as ‘presumptive’. Where our study showed evidence of recent dam infection by serology, but did not detect the presence of the pathogen directly in any samples (e.g., for BHV-1 in cattle, or C. abortus in small ruminants), we classified the diagnosis as ‘suggestive’ indicating a lower level of confidence in these aetiologic diagnoses.
Communication of results
For every livestock abortion case notified to our team, a preliminary laboratory report reported the PCR results. The report, identifiable to ward level, was supplied to the Tanzania Veterinary Laboratory Agency, Zonal Veterinary Investigation Centre, the Livestock Field Officer, and the farmer. If a pathogen was detected, an interpretation of what that aetiological agent was, and what could be done to prevent future infections, was included in the report. A translation into Kiswahili was available and the veterinarians in our field team were contactable for further consultation. The preliminary laboratory report template is available in the supplementary materials (S1; F).
References
De Vries, A. Economic value of pregnancy in dairy cattle. J. Dairy Sci. 89, 3876–3885 (2006).
Givens, M. D. & Marley, M. S. D. Infectious causes of embryonic and fetal mortality. Theriogenol 70, 270–285 (2008).
FAO. World Livestock. Livestock in food security 2011 (FAO, 2011).
FAO. World Livestock: Transforming the livestock sector through the Sustainable Development Goals. Rome: Food and Agriculture Organization of the United Nations; 2018. Report No.: 222 pp. Licence: CC BY-NC-SA 3.0 IGO.
Cabell, E. Bovine abortion: Aetiology and investigations. In Pract. 29, 455–463 (2007).
Campero, C. M., Moore, D. P., Odeón, A. C., Cipolla, A. L. & Odriozola, E. Aetiology of bovine abortion in Argentina. Vet. Res. Commun. 27, 359–369 (2003).
Wolf-Jäckel, G. A. et al. Diagnostic studies of abortion in Danish cattle 2015–2017. Acta Vet. Scand. 62, 1 (2020).
Kirkbride, C. A. Bacterial agents detected in a 10-year study of bovine abortions and stillbirths. J. Vet. Diagn. Invest. 5, 64–68 (1993).
Buxton, D. & Henderson, D. Infectious abortion in sheep. In Pract. 21, 360–368 (1999).
Hazlett, M. J. et al. A prospective study of sheep and goat abortion using real-time polymerase chain reaction and cut point estimation shows Coxiella burnetii and Chlamydophila abortus infection concurrently with other major pathogens. J. Vet. Diagn. Invest. 25, 359–368 (2013).
van den Brom, R. et al. Abortion in small ruminants in the Netherlands between 2006 and 2011. Tijdschrift Diergeneesk 137, 450–457 (2012).
Oliveira, R. D. et al. Domestic sheep show average Coxiella burnetii seropositivity generations after a sheep-associated human Q fever outbreak and lack detectable shedding by placental, vaginal, and fecal routes. PLoS ONE 12, e0188054 (2017).
Loftis, A. D., Reeves, W. K., Miller, M. M. & Massung, R. F. Coxiella burnetii, the agent of Q fever, in domestic sheep flocks from Wyoming United States. Vector Borne Zoon. Dis 12, 189–191 (2012).
Velasova, M. et al. Herd-level prevalence of selected endemic infectious diseases of dairy cows in Great Britain. J. Dairy Sci. 100, 9215–9233 (2017).
Essig, A. & Longbottom, D. Chlamydia abortus: New aspects of infectious abortion in sheep and potential risk for pregnant women. Curr. Clin. Microbiol. Rep. 2, 22–34 (2015).
Benkirane, A., Jabli, N. & Rodolakis, A. Frequency of abortion and seroprevalence of the principal diseases causing ovine infectious abortion in the area of Rabat (Morocco). Ann. Rech. Vet. 21, 267–273 (1990).
Hireche, S., Ababneh, M. M., Bouaziz, O. & Boussena, S. Seroprevalence and molecular characterization of Chlamydia abortus in frozen fetal and placental tissues of aborting ewes in northeastern Algeria. Trop. Anim. Health Prod. 48, 255–262 (2016).
Rekiki, A. et al. Isolation and characterisation of local strains of Chlamydophila abortus (Chlamydia psittaci serotype 1) from Tunisia. Vet. Res. 33, 215–222 (2002).
Seth-Smith, H. M. B. et al. European Chlamydia abortus livestock isolate genomes reveal unusual stability and limited diversity, reflected in geographical signatures. BMC Genomics 18, 344 (2017).
Budasha, N. H., Gonzalez, J. P., Sebhatu, T. T. & Arnold, E. Rift Valley fever seroprevalence and abortion frequency among livestock of Kisoro district, South Western Uganda (2016): A prerequisite for zoonotic infection. BMC Vet. Res. 14, 271 (2018).
Bodenham, R. F. et al. Prevalence and speciation of brucellosis in febrile patients from a pastoralist community of Tanzania. Sci. Rep. 10, 7081 (2020).
Mboera, L. E. G., Kishamawe, C., Kimario, E. & Rumisha, S. F. Mortality patterns of toxoplasmosis and its comorbidities in Tanzania: A 10-year retrospective hospital-based survey. Front. Public Health 7, 25 (2019).
Alemayehu, G., Mamo, G., Alemu, B., Desta, H., Tadesse, B., Benti, T., & Wieland, B. Causes and flock level risk factors of sheep and goat abortion in three agroecology zones in Ethiopia. Front. Vet. Sci. 8 (2021).
Okumu, T. A., John, N. M., Wabacha, J. K., Tsuma, V. & VanLeeuwen, J. Seroprevalence of antibodies for bovine viral diarrhoea virus, Brucella abortus and Neospora caninum, and their roles in the incidence of abortion/foetal loss in dairy cattle herds in Nakuru District Kenya. BMC Vet. Res. 15, 95 (2019).
Tolosa, T., Bezabih, D. & Regassa, F. Study on seroprevalence of bovine brucellosis and abortion and associated risk factor. Bull. Anim. Health Prod. Afr. 58 (2010).
Reichel, M.P., Wahl, L.C. & Hill, F.I. Review of diagnostic procedures and approaches to infectious causes of reproductive failures of cattle in Australia and New Zealand. Front. Vet. Sci. 5 (2018).
Vidal, S. et al. Neglected zoonotic agents in cattle abortion: tackling the difficult to grow bacteria. BMC Vet. Res. 13, 373 (2017).
Maziku, M., Gebru, G. & Stapleton, J. Livestock health priorities in the Tanzania livestock master plan. Tanzania Livestock Master Plan Brief 4. Nairobi, Kenya: IRLI (2017).
CDC. Workshop summary. One health zoonotic disease prioritization for multisectoral engagement in Tanzania. In: Centers for Disease Control and Prevention (2017).
Crump, J. A. et al. Invasive bacterial and fungal infections among hospitalized HIV-infected and HIV-uninfected children and infants in northern Tanzania. Trop. Med. Int. Health 16, 830–837 (2011).
Nyarobi, J. The epidemiology of Rift Valley fever in northern Tanzania. PhD thesis, University of Glasgow (2020).
Sindato, C. et al. A spatial analysis of Rift Valley fever virus seropositivity in domestic ruminants in Tanzania. PLoS ONE 10, e0131873–e0131873 (2015).
Covarrubias, K., Nsiima, L. & Zezza, A. Livestock and livelihoods in rural Tanzania: a descriptive analysis of the 2009 National Panel Survey. Joint paper of the World Bank, FAO, AU-IBAR, ILRI and the Tanzanian Ministry of Livestock and Fisheries Development. (2012).
de Glanville, W. A. et al. Classification and characterisation of livestock production systems in northern Tanzania. PLoS ONE 15, e0229478 (2020).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10, 512–526 (1993).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Braun, U., Hilbe, M., Peterhans, E. & Schweizer, M. Border disease in cattle. Vet J 246, 12–20 (2019).
Munyua, P. et al. Rift Valley fever outbreak in livestock in Kenya, 2006–2007. Am. J. Trop. Med. Hyg. 83, 58–64 (2010).
Woods, C. W. et al. An outbreak of Rift Valley fever in Northeastern Kenya, 1997–98. Emerg. Infect. Dis. 8, 138–144 (2002).
Prabhu, M. et al. Q fever, spotted fever group, and typhus group rickettsioses among hospitalized febrile patients in northern Tanzania. Clin. Infect. Dis. 53, e8–e15 (2011).
Anyamba, A., Soebiyanto, R.P., Small, J.L., Linthicum, K.J., Forshey, B., Toolin, C.F., & Tucker, C.J. Rift Valley fever outbreak in East Africa: signature of climate extremes. American Geophysical Union, Fall Meeting 2018, abstract #GH14A-07; 2018. pp. GH14A-07.
Mohamed, M. et al. Epidemiologic and clinical aspects of a Rift Valley fever outbreak in humans in Tanzania, 2007. Am. J. Trop. Med. Hyg. 83, 22–27 (2010).
Mbotha, D. et al. Inter-epidemic Rift Valley fever virus seroconversions in an irrigation scheme in Bura, South-East Kenya. Transbound. Emerg. Dis. 65, e55–e62 (2018).
Hassan, A. et al. Epidemiological investigation of a Rift Valley fever outbreak in humans and livestock in Kenya, 2018. Am. J. Trop. Med. Hyg. 103, 1649–1655 (2020).
ProMED-mail. Rift Valley fever - Eastern Africa: (Tanzania) alert. ProMED-mail. Archive Number: 20180619.5863493 2018 [cited 2021 March 19] Available from: http://www.promedmail.org.
Roest, H. I. et al. Molecular epidemiology of Coxiella burnetii from ruminants in Q fever outbreak, the Netherlands. Emerg. Infect. Dis. 17, 668–675 (2011).
Salifu, S. P., Bukari, A. A., Frangoulidis, D. & Wheelhouse, N. Current perspectives on the transmission of Q fever: Highlighting the need for a systematic molecular approach for a neglected disease in Africa. Acta Trop. 193, 99–105 (2019).
Vanderburg, S. et al. Epidemiology of Coxiella burnetii infection in Africa: A OneHealth systematic review. PLoS Negl. Trop. Dis. 8, e2787–e2787 (2014).
Crump, J. A. et al. Etiology of severe non-malaria febrile illness in Northern Tanzania: A prospective cohort study. PLoS Negl. Trop. Dis. 7, e2324–e2324 (2013).
Galal, L. et al. Toxoplasma and Africa: One parasite, two opposite population structures. Trends Parasitol. 34, 140–154 (2018).
Swai, E. S. & Kaaya, J. E. A survey of Toxoplasma gondii antibodies by latex agglutination assay in dairy goats in northern Tanzania. Trop. Anim. Health Prod. 45, 211–217 (2013).
Tonouhewa, A. B. et al. Toxoplasma gondii infection in meat animals from Africa: systematic review and meta-analysis of sero-epidemiological studies. Vet. World 10, 194–208 (2017).
Ferreira, S. C. M. et al. Evidence of high exposure to Toxoplasma gondii in free-ranging and captive African carnivores. Int. J. Parasitol. Parasites Wildl. 8, 111–117 (2019).
Innes, E. A., Bartley, P. M., Buxton, D. & Katzer, F. Ovine toxoplasmosis. Parasitol 136, 1887–1894 (2009).
Barkallah, M. et al. Molecular prevalence of Chlamydia and Chlamydia-like bacteria in Tunisian domestic ruminant farms and their influencing risk factors. Transbound. Emerg. Dis. 65, e329–e338 (2018).
Bhandi, S. et al. Brucellosis and chlamydiosis seroprevalence in goats at livestock-wildlife interface areas of Zimbabwe. Onderstepoort. J. Vet. Res. 86, e1–e9 (2019).
Samkange, A., Katsande, T. C., Tjipura-Zaire, G. & Crafford, J. E. Seroprevalence survey of Chlamydophila abortus infection in breeding goats on commercial farms in the Otavi Veterinary District, northern Namibia. Onderstepoort. J. Vet. Res. 77, E1-5 (2010).
Allan, K. J. et al. Assessment of animal hosts of pathogenic Leptospira in northern Tanzania. PLoS Negl. Trop. Dis. 12, e0006444–e0006444 (2018).
Bouley, A. J. et al. Brucellosis among hospitalized febrile patients in northern Tanzania. Am. J. Trop. Med. Hyg. 87, 1105–1111 (2012).
Ellis, W.A. Animal Leptospirosis. In: Adler, B. (ed). Leptospira and Leptospirosis, vol. 387. Springer: Berlin (2015).
Semango, G., Hamilton, C.M., Kreppel, K., Katzer, F., Kibona, T., Lankester, F., & de Glanville, W.A. The sero-epidemiology of Neospora caninum in cattle in northern Tanzania. Front. Vet. Sci. 6 (2019).
Eleni, C. et al. Detection of Neospora caninum in an aborted goat foetus. Vet. Parasitol. 123, 271–274 (2004).
Varaschin, M. S. et al. Congenital neosporosis in goats from the State of Minas Gerais Brazil. Korean J. Parasitol. 50, 63–67 (2012).
González-Warleta, M. et al. Endogenous transplacental transmission of Neospora caninum during successive pregnancies across three generations of naturally infected sheep. Vet. Res. 49, 106 (2018).
Wodajo, H. D. et al. Contribution of small ruminants to food security for Ethiopian smallholder farmers. Small Rumin. Res. 184, 106064 (2020).
Smith, T., Godfrey, S. H., Buttery, P. J. The contribution of sheep and goats in alleviating poverty: communicating messages from research: Proceedings of the third DFID Livestock Production Programme link project (R7798) workshop for small ruminant keepers. Izaak Walton Inn, Embu, Kenya, 4–7 February 2003.
Main, K. Seroprevalence and risk factors associated with bovine herpesvirus-1 infection in cattle in northern Tanzania. MSc thesis, University of Glasgow (2019).
Newcomer, B. W. & Givens, D. Diagnosis and control of viral diseases of reproductive importance: Infectious bovine rhinotracheitis and bovine viral diarrhea. Vet. Clin. N. Am. Food Anim. Pract. 32, 425–441 (2016).
Barkallah, M. et al. Survey of infectious etiologies of bovine abortion during mid- to late gestation in dairy herds. PLoS ONE 9, e91549 (2014).
Gethmann, J., Probst , C. & Conraths, F.J. Economic impact of a bluetongue serotype 8 epidemic in Germany. Front. Vet. Sci. 7 (2020).
Nusinovici, S., Souty, C., Seegers, H., Beaudeau, F. & Fourichon, C. Decrease in milk yield associated with exposure to bluetongue virus serotype 8 in cattle herds. J. Dairy Sci. 96, 877–888 (2013).
Abera, T. et al. Bluetongue disease in small ruminants in south western Ethiopia: Cross-sectional sero-epidemiological study. BMC Res. Notes 11, 112 (2018).
Mulabbi, E. N., Ayebazibwe, C., Majalija, S., Batten, C. A. & Oura, C. A. Circulation of bluetongue virus in goats in the Karamoja region of Uganda. J. S. Afr. Vet. Assoc. 84, E1-3 (2013).
Toye, P. G. et al. Bluetongue and epizootic haemorrhagic disease virus in local breeds of cattle in Kenya. Res. Vet. Sci. 94, 769–773 (2013).
Coetzee, P., Stokstad, M., Venter, E. H., Myrmel, M. & Van Vuuren, M. Bluetongue: A historical and epidemiological perspective with the emphasis on South Africa. Virol. J. 9, 198 (2012).
Maclachlan, N. J., Drew, C. P., Darpel, K. E. & Worwa, G. The pathology and pathogenesis of bluetongue. J. Comp. Pathol. 141, 1–16 (2009).
Knoll, M. D. et al. Bayesian estimation of pneumonia etiology: Epidemiologic considerations and applications to the pneumonia etiology research for child health study. Clin. Infect. Dis. 64, S213-s227 (2017).
Hilbert, A. et al. Prevalence of Coxiella burnetii in clinically healthy German sheep flocks. BMC Res. Notes 5, 152 (2012).
Borel, N. et al. Laboratory diagnosis of ruminant abortion in Europe. Vet. J. 200, 218–229 (2014).
Sidi-Boumedine, K., Rousset, E., Henning, K., Ziller, M., Niemczuck, K., Roest, H.I. & Thiéry, R. Development of harmonised schemes for the monitoring and reporting of Q‐fever in animals in the European Union. EFSA Supporting Publication 7 (2010).
World Organisation for Animal Health (OIE). Q Fever. In: OIE, editor. OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals (2018).
Herzog, C. M. et al. Pastoral production is associated with increased peste des petits ruminants seroprevalence in northern Tanzania across sheep, goats and cattle. Epidemiol. Infect. 147, e242 (2019).
Kadohira, M., McDermott, J. J., Shoukri, M. M. & Kyule, M. N. Variations in the prevalence of antibody to Brucella infection in cattle by farm, area and district in Kenya. Epidemiol. Infect. 118, 35–41 (1997).
Kairu-Wanyoike, S., Nyamwaya, D., Wainaina, M., Lindahl, J., Ontiri, E., Bukachi, S., & Bett, B. Positive association between Brucella spp. seroprevalences in livestock and humans from a cross-sectional study in Garissa and Tana River Counties, Kenya. PLoS Negl. Trop. Dis. 13, e0007506 (2019).
Bodenham, R.F., Mazeri, S., Cleaveland, S., Crump, J.A., Fasina, F.O., de Glanville, W.A., & Halliday, J.E.B. Latent class evaluation of the performance of serological tests for exposure to Brucella spp. in cattle, sheep, and goats in Tanzania. PLoS Negl. Trop. Dis. 15, e0009630-e0009630 (2021).
Helmick, B., Otter, A., McGarry, J. & Buxton, D. Serological investigation of aborted sheep and pigs for infection by Neospora caninum. Res. Vet. Sci. 73(2), 187-189. https://doi.org/10.1016/s0034-5288(02)00093-0 (2002).
Marques, P. X. et al. Detection of Toxoplasma gondii antigens reactive with antibodies from serum, amniotic, and allantoic fluids from experimentally infected pregnant ewes. Vet Parasitol. 185(2–4), 91–100. https://doi.org/10.1016/j.vetpar.2011.10.028 (2012).
Wilson, K., Livingstone, M. & Longbottom, D. Comparative evaluation of eight serological assays for diagnosing Chlamydophila abortus infection in sheep. Vet. Microbiol. 135, 38–45 (2009).
Dorai-Raj, S. binom: binomial confidence intervals for several parameterizations. R package version 1.1–1; 2014.
Hamilton, C.M., Kelly, P.J., Bartley, P.M., Burrells, A., Porco, A., Metzler, D., & Katzer, F. Toxoplasma gondii in livestock in St. Kitts and Nevis, West Indies. Parasit. Vectors 8, 166 (2015).
Matero, P., Hemmilä, H., Tomaso, H., Piiparinen, H., Rantakokko-Jalava, K., Nuotio, L. & Nikkari, S. Rapid field detection assays for Bacillus anthracis, Brucella spp., Francisella tularensis and Yersinia pestis. Clin. Microbiol. Infect. 17, 34–43 (2011).
Probert, W.S., Schrader, K.N., Khuong, N.Y., Bystrom, S.L. & Graves, M.H. Real-time multiplex PCR assay for detection of Brucella spp., B. abortus, and B. melitensis. J. Clin. Microbiol. 42, 1290–1293 (2004).
Livingstone, M., Wheelhouse, N., Maley, S. W. & Longbottom, D. Molecular detection of Chlamydophila abortus in post-abortion sheep at oestrus and subsequent lambing. Vet. Microbiol. 135, 134–141 (2009).
Stoddard, R.A., Gee, J.E., Wilkins, P.P., McCaustland, K. & Hoffmaster, A.R. Detection of pathogenic Leptospira spp. through TaqMan polymerase chain reaction targeting the LipL32 gene. Diagn. Microbiol. Infect. Dis. 64, 247–255 (2009).
Buxton, D. et al. The pathogenesis of experimental neosporosis in pregnant sheep. J. Comp. Pathol. 118, 267–279 (1998).
Hurtado, A., Aduriz, G., Moreno, B., Barandika, J. & García-Pérez, A. L. Single tube nested PCR for the detection of Toxoplasma gondii in fetal tissues from naturally aborted ewes. Vet. Parasitol. 102, 17–27 (2001).
Drosten, C. et al. Rapid detection and quantification of RNA of Ebola and Marburg viruses, Lassa virus, Crimean-Congo hemorrhagic fever virus, Rift Valley fever virus, dengue virus, and yellow fever virus by real-time reverse transcription-PCR. J. Clin. Microbiol. 40, 2323–2330 (2002).
Burrells, A. et al. Evidence of the three main clonal Toxoplasma gondii lineages from wild mammalian carnivores in the UK. Parasitol 140, 1768–1776 (2013).
Becher, P. et al. Phylogenetic analysis of pestiviruses from domestic and wild ruminants. J. Gen. Virol. 78(Pt 6), 1357–1366 (1997).
Vilcek, S., Nettleton, P. F., Paton, D. J. & Belák, S. Molecular characterization of ovine pestiviruses. J. Gen. Virol. 78, 725–735 (1997).
Vilcek, S. et al. Bovine viral diarrhoea virus genotype 1 can be separated into at least eleven genetic groups. Arch. Virol. 146, 99–115 (2001).
Dubey, J. P. & Schares, G. Neosporosis in animals—the last five years. Vet. Parasitol. 180, 90–108 (2011).
World Organisation for Animal Health (OIE). Brucellosis (Brucella abortus, B. melitensis and B. suis) (infection with B. abortus, B. melitensis and B. suis). In: OIE, editor. Manual of diagnostic tests and vaccines for terrestrial animals (2018).
World Organisation for Animal Health (OIE). Enzootic Abortion of Ewes (Ovine Chlamydiosis) (Infection with Chlamydia abortus). In: OIE, editor. Manual of diagnostic tests and vaccines for terrestrial animals (2018).
Touratier, A., Baurier, F., Beaudeau, F., Bendali, F., Buret, Y., DeCremoux, R., & Dufour, A. Comment faire le diagno stic d'un élevage cliniquement atteint de fièvre Q? Recueil des Journées Nationales des GTV, Nantes, 147–155 (2007).
World Organisation for Animal Health (OIE). Leptospirosis In: OIE, editor. OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals (2018).
Dubey, J. P. & Schares, G. Diagnosis of bovine neosporosis. Vet Parasitol 140, 1–34 (2006).
World Organisation for Animal Health (OIE). Toxoplasmosis. In: OIE, editor. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals (2018).
World Organisation for Animal Health (OIE). Infectious Bovine Rhinotracheitis/Infectious Pustular Vulvovaginits. In: OIE, editor. Manual of diagnostic tests and vaccines for terrestrial animals (2018).
World Organisation for Animal Health (OIE). Bluetongue (infection with bluetongue virus). In: OIE, editor. Manual of diagnostic tests and vaccines for terrestrial animals (2018).
World Organisation for Animal Health (OIE). Bluetongue. Aetiology, epidemiology, diagnosis, prevention and control references (2021).
World Organisation for Animal Health (OIE). Bovine Viral Diarrhoea. In: OIE, editor. Manual of diagnostic tests and vaccines for terrestrial animals (2018).
World Organisation for Animal Health (OIE). Rift Valley Fever (infection with Rift Valley fever virus). In: OIE, editor. OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals (2018).
Acknowledgements
The abortion cohort study was supported by the Supporting Evidence Based Interventions project, University of Edinburgh (grant number R83537 CH). The cross-sectional studies were supported by US National Institutes of Health (NIH)—National Science Foundation (NSF) Ecology and Evolution of Infectious Disease program (R01 TW009237) and the UK Biotechnology and Biological Sciences Research Council (BBSRC) (grant numbers: BB/J010367/1, BB/L018926/1). MD was funded by the Scottish Government. CH, CH, EI, FK, ML, GR and DL are supported by funding from the Scottish Government Rural and Environment Science and Analytical Services Division (RESAS). BW is funded by the BBSRC. We would like to thank Rosanne de Jong, Divine Ekwem, Erin Hodgkinson, Karen Main and Jen Jen Yu for assistance with serological testing at the University of Glasgow and Clare Underwood (MRI) for expert immunohistological preparations. We appreciate the huge efforts of our field team, Rigobert Tarimo, Fadhili Mshana, and Hassan Hussein as well as the assistance from District Veterinary Officers and LFOs. Thanks to Paul Johnson (University of Glasgow) for support with data management and curation, and Dassa Nkini (NM-AIST) and Elizabeth Kussaga (KCRI) for administrative support.
Author information
Authors and Affiliations
Contributions
K.T., J.R.C., Wd.G., F.L., J.B., J.A.C., M.D., J.H., C.H., E.A.I., F.K., D.L., B.M., O.N., N.W., S.C., K.A. developed the study concept and design of the work. T.K., Wd.G., F.L., G.S. collected samples and metadata. K.T., N.A., R.C., G.C., M.D., C.H., E.I., F.K., M.L., C.M., V.M., J.N., G.R., N.W., B.W., S.C., K.A. processed samples. K.T., J.R.C., Wd.G., F.L., N.A., R.C., G.C., J.A.C., M.D., J.H., C.H., E.I., F.K., M.L., D.L., C.M., V.M., J.N., G.R., N.W., B.W., S.C., K.A. analysed and interpreted data. K.T., Wd.G., F.L., J.H., S.C., K.A. drafted the article. All authors provided critical revision of the article and provided approval for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thomas, K.M., Kibona, T., Claxton, J.R. et al. Prospective cohort study reveals unexpected aetiologies of livestock abortion in northern Tanzania. Sci Rep 12, 11669 (2022). https://doi.org/10.1038/s41598-022-15517-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-15517-8
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.