Abstract
Unnatural substituted amino acids play an important role as chiral building blocks, especially for pharmaceutical industry, where the synthesis of chiral biologically active molecules still represents an open challenge. Recently, modification of the hydrophobic binding pocket of phenylalanine ammonia-lyase from Petroselinum crispum (PcPAL) resulted in specifically tailored PcPAL variants, contributing to a rational design template for PAL-activity enhancements towards the differently substituted substrate analogues. Within this study we tested the general applicability of this rational design model in case of PALs, of different sources, such as from Arabidopsis thaliana (AtPAL) and Rhodosporidium toruloides (RtPAL). With some exceptions, the results support that the positions of substrate specificity modulating residues are conserved among PALs, thus the mutation with beneficial effect for PAL-activity enhancement can be predicted using the established rational design model. Accordingly, the study supports that tailoring PALs of different origins and different substrate scope, can be performed through a general method. Moreover, the fact that AtPAL variants I461V, L133A and L257V, all outperformed in terms of catalytic efficiency the corresponding, previously reported, highly efficient PcPAL variants, of identical catalytic site, suggests that not only catalytic site differences influence the PAL-activity, thus for the selection of the optimal PAL-biocatalysts for a targeted process, screening of PALs from different origins, should be included.
Similar content being viewed by others
Introduction
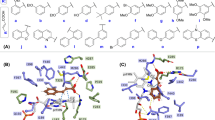
The current state of art of the PAL mediated biotransformations revealed several synthetically useful PALs of eukaryotic (plant and yeast) or bacterial origins1,2,3,4. Their substrate scope has been intensively studied within the last decade, with several PALs, such as those originary from Petroselinum crispum (PcPAL)2,5,6, Anabaena variabilis (AvPAL)7,8, Rhodotorula glutinis (RgPAL)9,10, Arabidopsis thaliana (AtPAL)11, Planctomyces brasiliensis (PbPAL)12, Kangiella koreensis (KkPAL)13, Pseudozyma antarctica (PzaPAL)14, shown to possess broad substrate scope. However, all these studies also revealed significant differences in their catalytic efficiencies towards specific substrates2,7,11,12,14. As example, PbPAL and the recently explored AL-11 PAL15 transformed substrates with electron-donor substituents, previously shown to be poor substrates for other PALs, such as PcPAL, AtPAL and AvPAL. Comparison of the catalytic sites of PALs of different origins shows a highly conserved polar substrate binding region responsible for the fixation of substrate’s carboxyl- and NH2- group (Fig. 1, Fig. S1), which also embeds the catalytically essential 3,5-dihydro-5-methylene-4H-imidazol-4-one MIO-group16,17. Besides, the residues of the polar binding region form an essential H-bond network18 (Fig. S2). The differences within the hydrophobic substrate-binding region of PALs (Fig. 1, Fig. S1), responsible for the facile active site accommodation of the substrate’s aromatic ring, supposedly contribute to the different substrate specificities observed among aromatic ammonia lyases2,4,19. Interestingly, several hydrophobic active site residues, such as those corresponding to I460 and L256 of PcPAL are highly conserved, while large diversification can be observed at positions homologue with 137 and 138 of PcPAL (Fig. 1, Fig. S1). These later residues are well-known for their substrate-specificity modulator effect, specific polar residues (e.g. histidine) at position corresponding to 137 of PcPAL provide also tyrosine ammonia-lyase (TAL) activity, such as in case of RgPAL and RtPAL from Rhodotorula sp., with reported TAL/PAL activities20. Histidine ammonia-lyases (HALs) show a characteristic His residue at position corresponding to 138 of PcPAL19,21. Exploration and characterization of novel PALs from different origins is continuously expanding12,13,14,15 driven by the aim to find PALs of increased operational and/or thermostability, or of activity towards substrates hardly transformed by existing PALs.
Protein engineering efforts on PALs of different origins, such as PcPAL2,6,18,22, AvPAL7,23, RgPAL24, PbPAL12, AL-1115, also focused to provide variants of expanded substrate scope or increased operational and thermal stability. Diversification at positions 137 and 138 has been performed at several PALs, such as PcPAL2,6,18, AvPAL7,8,23, RtPAL24 and PbPAL12, and provided variants of improved catalytic performance, with modifications especially at analogue positions of 137 of PcPAL, highlighting the substrate specificity modulator effect of this residue. Our recent mutational analysis of the hydrophobic binding pocket of PcPAL2, revealed other specificity modulator active site residues and depicted their specific interaction with the differently positioned ortho-, meta-, para- substituents of the non-natural substrates, thus providing excellent tool for the rational protein engineering of PcPAL. Considering the differences in the hydrophobic substrate binding pocket of PALs of different origins, that strongly influence their substrate scope, the general validity of our recently developed rational engineering strategy among PALs of diverse origins, hence of diverse substrate scope, should be assessed. This might provide a desirable general rational design strategy among PALs, allowing facile development of substrate-tailored PALs of various origins. Accordingly, we tested whether the mutational strategy developed for PcPAL applies to other PALs, such as AtPAL and RtPAL, of different sequence similarities to PcPAL, AtPAL possessing high, 81% sequence identity and identical catalytic site with PcPAL, while RtPAL shares lower, 38% sequence identity with PcPAL, and contains specific ‘TAL-activity provider’ His and Gln residues at positions analogue to 137 and 138 of PcPAL (Fig. 1). Notable, that wild-type AtPAL and PcPAL show very similar catalytic efficiencies (kcat), while in comparison RtPAL shows ~ twofold increased kcat values, but lower specificity constants (kcat/KM) in the natural PAL-reaction (deamination of l-Phe) and also within the reverse ammonia addition route of trans-cinnamic acid11.
Catalytic site of PcPAL, with key residues from the hydrophobic substrate-binding region marked based on their proximity2 to the differently positioned (ortho-blue, meta-orange and para-green) aromatic substituents of the substrates and the homologue active site residues in RtPAL (P11544) and AtPAL (P45724) based on sequence alignments with PcPAL (P24481) (right Table) with colour-marked residues subjected to mutagenesis, generating the focused PAL variant-library (bottom Table).
Results and discussion
Mutant library generation and enzyme activity screens
Recent mapping of the hydrophobic binding pocket of PcPAL, revealed mutant variants, obtained by mutagenesis of residues L256, L134, F137 and I460, of enhanced catalytic activity to ortho-, meta-, para- substituted substrates, respectively2. Accordingly, the homologues of all four substrate specificity modulating residues were similarly replaced in AtPAL and RtPAL (Fig. 1), initially obtaining four AtPAL variants (L257V, L133A, F136V and I461V) and four RtPAL variants (L266V, L134A, H137V and I472V). Notable, that residue H137 of RtPAL, homologue of F137 of PcPAL, is most probably involved in H-bonding with Q138, homologue of L138 of PcPAL. Since H137V RtPAL variant showed reduced activity within the activity screens, we presumed that the presence of residue Q138 within a complete hydrophobic active site environment is non-favourable. Thus, the mutational strategy was adapted, and within the ‘PcPAL-like’ RtPAL variants, besides the specific mutation of residues involved in aromatic substituent accommodation, additional mutations H137F and Q138L have also been included. Thus, RtPAL variant H137F/Q138L resembles the catalytic site of wt-PcPAL, variants H137F/Q138L/L266V and H137F/Q138L/I472V being homologues to L256V and I460V PcPAL, respectively, and variant H137V/Q138L RtPAL to F137V PcPAL variant. Despite our efforts, including different PCR protocols for mutagenesis, RtPAL variant L134A/H137F/Q138L resembling the catalytic site of L134A PcPAL could not be obtained, thus only L134A RtPAL was employed within the activity tests. The slight variations in thermal unfolding temperatures (Tm) of the purified enzyme variants indicated that mutations did not affect the protein folding (Figs. S3, S4 and Table S2), the only significant modification being observed in case of variant H137F/Q138L/I472V RtPAL, with Tm value decreased with ~ 6 °C compared to the wild-type RtPAL.
Activity assessments of the PcPAL, AtPAL and RtPAL variant library
The activities of the obtained RtPAL and AtPAL variants were assessed and compared with those of the corresponding PcPAL variants within the deamination and amination reactions of phenylalanines and cinnamic acids, monosubstituted with both electron-donating (− CH3, − OCH3) and electron-withdrawing (− CF3, − Br) groups at all positions (ortho, meta, para) of their aromatic ring (Fig. 2). Both reaction routes have been tested by whole-cell PAL-biocatalysts mediated biotransformations of substrates rac-1a–l and 2a–l, monitoring the conversions, while for the ammonia eliminations enzyme kinetic parameters (KM and kcat), based on initial velocity measurements using purified enzymes, were also assessed. During enzyme kinetic measurements for the ammonia additions, factors such as (i) the high ammonia concentration of the reaction buffer leading to increased background absorbance, (ii) the high extinction coefficient of the cinnamic acid derivatives, hindering the use of large substrate concentrations, (iii) the overlapping absorbance spectra of the cinnamic acid and phenylalanine counterparts, provided high standard deviations, low reproducibility and/or incomplete Michaelis–Menten curves.
Activity assessments for ortho-substituted substrates
Generally, within the whole-cell mediated biotransformations of ortho-substituted substrates 1,2a–d, mutations L257V, L133A in case of AtPAL and mutation L266V in case of RtPAL provided similar enhancement of the conversion-based enzyme activity, relatively to their wild-type variants, as the one reported for L256V PcPAL2 (Table 1). Additional general tendency can be observed among the results obtained with wild-type PALs, AtPAL outperforming in terms of conversion the corresponding PcPAL, while wt-RtPAL provided the lower conversions in both reaction routes (Table 1).
More detailed, in case of o-Br-substituted substrates excellent, equilibrium-approaching conversions are obtained with the wild-type PcPAL (87% for 2a and 42% for 1a after 6 h and 24 h reaction times) and AtPAL (94% for 2a after 1 h reaction time and 48% for 1a after 6 h reaction time), thus the increased catalytic efficiency of variants L256V PcPAL and L257V AtPAL is less reflected within the conversion-based enzyme activities. However, the 2.3- and 2.1-fold increased kcat values, comparatively to the wt-variant’s (Table 1), support the beneficial effect of the mutations. Similar behaviour can be observed in case of substrates 1b and 2b, with high conversions of similar range being registered for both wt- or mutant variants of Pc/AtPAL, but 3.5- and 5.5-fold increased catalytic efficiencies (kcat) of the corresponding L256V and L257V variants. AtPALs provided stationary conversions of ~ 86% for 2b and ~ 50% for rac-1b within significantly shorter reaction times of 3 h and 30 min, respectively, in comparison with similar conversions obtained only after 24 h reaction times using PcPALs. Interestingly, while in case of 1a the KM value was not significantly altered upon mutations analogue to L256V, in case of 1b the mutation resulted in highly decreased substrate affinity (increased KM values) for all three PALs of different origin, supporting a more relaxed accommodation of 1b within the modified active site. Wild-type RtPAL in case of both substrates 2a, 2b provided only moderate conversion of 30% (6 h) and 35% (24 h), respectively, thus the beneficial effect of mutation L266V was clearly visible also within the conversion-based enzyme activity, with 94% (6 h) and 82% (24 h) conversion for 2a and 2b, respectively. In case of o-OCH3- substituted substrates 1c, 2c, the beneficial effect of mutations analogue with L134A from PcPAL was also observed in case of AtPAL, where the corresponding L133A variant provided high conversions of 95% and 3.1-fold increased kcat values. In case of L134A, but also wild-type RtPAL, very low/no conversions of < 1–3% were detected, while enzyme kinetics also revealed low initial velocities and substrate affinities. Interestingly, in this case the ‘PcPAL-like’ RtPAL L266V variant, with mutations H137F/Q138L/L266V, provided increased conversions of 19% for 1c and 17% for 2c after 16 h reaction time. Indeed, in this particular case, due to the bend caused by the oxygen atom of the o-OCH3 substituent, the methyl group positions between residues 134 and 266 (Fig. 3A,B), while the increased hydrophobicity induced by mutation H137F and Q138L most probably facilitates the accommodation of the substrate’s aromatic moiety. In accordance with the experimental results, both flexible and rigid docking of 2c within the active sites of wild-type, L134A and H137F/Q138L/L266V RtPAL revealed substrate orientations of significantly lower energy for both mutant variants in comparison with those obtained for the wild-type RtPAL (Fig. 3A). In case of o-CH3-substituted substrates 1d, 2d the wild-type variants of all three PALs provided high conversion in both reaction routes, the best performer AtPAL reaching in shortest reaction time of 3 h 83% conversion of 2a. Similarly to the case of the o-Br-substituted substrates 1a, 2a, the increased catalytic efficiency of the variants bearing mutations analogue to L256V of PcPAL is supported by their increased kcat values in comparison to their wild-type variants. The less significant, only 1.1–1.3-fold increase in kcat values, than in case of 1a–c, is expectable based on the smallest sterical requirement of the methyl group, which seemingly, when ortho-positioned on the substrate, is favourably accommodated within the active site of all PAL variants.
2-Methoxycinnamic acid 2c docked into the active site of: (A) wt-RtPAL (green, − 4.2 kcal/mol), L134A RtPAL (purple, − 6.9 kcal/mol), and H137F/Q138L/L266V RtPAL (grey, − 8 kcal/mol); (B) wt-PcPAL (green, − 7.3 kcal/mol) and L134A PcPAL (purple, − 7.7 kcal/mol). Steric clashes between the ortho-methoxy group and side chains of residues L266 and L134, respectively, are highlighted with red dashed lines. The modified residues and the active site orientation of 2c within the corresponding PAL variant is marked with similar colour.
Activity assessments for meta-substituted substrates
Interestingly, in case of phenylalanine/cinnamic acid analogues with substituents in meta-position 1e, 1f, 1h and 2e, 2f, 2h, the wild-type variant RtPAL showed superior catalytic efficiency in comparison to wild-type PcPAL and AtPAL, supported by its higher kcat values within the ammonia eliminations of 1e, 1f, 1h or conversion values in both reaction routes of m-CF3- and m-Me-substituted substrates (Table 2). The case of m-methoxy-substituted substrates 1g, 2g acts again as an exception, where RtPAL shows significantly lower, ~ 18% conversion within the ammonia addition, while 52% and 32% conversion are registered with AtPAL and PcPAL, respectively. Within the two presumed active orientations of the meta-substituents (Fig. 1), in case of RtPAL besides the conserved L134 and I472 residues, polar residues Q138, H137 also appear, suggesting their favourable interaction with the polar CF3- and Br- substituents of 1e, 2e and 1f, 2f, that most probably contributes towards the superior activity of RtPAL. Computational results revealed that the substrate orientations exposing the meta-substituent towards residue I460 are energetically favoured in case of wild-type PcPAL, while for wild-type RtPAL the presence of polar residues Q138, H137 shifts the active site orientation of the meta-substituent from those observed for PcPAL, the orientations towards residues L134 showing close or even lower energies than those pointing towards I472 of RtPAL (Fig. 4, Fig. S5, Table S7).
The beneficial effect of the mutational strategy explored at PcPAL for the increased enzyme activity towards m-substituted substrates, was perfectly retained in case of AtPAL, where mutations L133A and I461V provided significantly increased kcat and conversion values in case of all substrates. While the results were expected in the frame of identical architecture of the two catalytic sites of AtPAL and PcPAL, in most of the cases AtPAL variants I461V and L133A showed superior catalytic properties (higher kcat values and higher conversions in shorter reaction time), than their corresponding PcPAL homologues (Table 2). In case of RtPAL, deciphering the beneficial effect of homologue mutations L134A and I472V was hindered by the low activity of I472V RtPAL and its “PcPAL-like” homologue, H137F/Q138L/I472V variant, while the above discussed high activity of the wild-type RtPAL supports that it also represents an optimized variant for meta-substituted substrates, substrate orientations exposing the aromatic substituents towards residue L134 being favoured. Accordingly, variant L134A provided conversion and kinetic data close to those observed for wt-RtPAL for all tested substrates, while the unsuccessful mutagenesis in case of its PcPAL-like L134A/H137F/Q138L variant didn’t allow testing the combined effect of replacing polar residues H137, Q138 and meta-substituted substrate-modulator residue L134. Moreover, variants including mutation I472V used as purified proteins were completely inactive within kinetic measurements, also providing very low conversion when used as whole-cell biocatalysts in biotransformations of 1e, 1f and 2e, 2f. Their thermal denaturing profile (Fig. S4) reveals their lowered thermal stability, similar to those reported for I460A PcPAL variant, for which we supposed that the mutation-induced, non-favourable water-accessibility of the catalytic site is responsible for the activity loss6. Notable, that variant I472V RtPAL and its ‘PcPAL-like’ homologue H137F/Q138L/I472V were also inactive within the biotransformations of p-substituted substrates (Table 3).
Activity assessments for para-substituted substrates
In case of para-substituted phenylalanines 1j, 1k,1l and cinnamic acids 2i–2l very low (< 10%) or no conversion was detected when using wild-type PcPAL and RtPAL variants, in accordance with the reported steric clashes between the p-substituent and active site residues2,6. Interestingly, wt-AtPAL afforded close to maximum conversion of all para-substituted phenylalanines, except for p-OCH3-phenylalanine, where similarly to Pc/RtPALs, low conversion of 14% and kcat value of 0.007 s−1 were obtained within the ammonia elimination of rac-1k and no conversion within the ammonia addition to 2k.
Related to the effect of the mutational strategy, we observed that in case of AtPAL variants I461V and F136V, similarly as in case of PcPAL, provided important conversion and activity enhancements for all substrates 1i–2l and 2i–l. Accordingly, while in case of p-Br- and p-CF3-substituted substrates the mutation-induced increase in the conversions is less significant, due to the well-performing wild-type variant, the 2.9-fold and 3.4-fold increased kcat values of variant I461V for 1i and 1j support the beneficial effect of the mutation. In case of substrates 1k, 1l and 2k, 2l, p-substituted with the electron-donating -OCH3 and -CH3 groups, the superior catalytic efficiency of I461V variant to wt-AtPAL is also resembled within the highly increased conversion values. While mutation F136V of AtPAL also induced significant increase in the conversions of all substrates, in case of substrates 1i, 1j and 2i, 2j even surpassing the conversions registered with I461V variant, however the enantiomeric excess (ee) of the l-phenylalanines 1i, 1j and 1 k produced within the ammonia additions, were of lower value (ee of 92%, 83% and 97%, respectively) in comparison with the highly enantiopure forms (ee > 99%) produced by wt- and I461V variant. This is in accordance with the results from PcPAL, where mutation F137V also decreased the enantioselectivity of the enzyme with ee values of 97% and 82% being obtained for l-1i and l-1j, respectively.
RtPAL, in general, proved to be inefficient for the transformation of para-substituted amino acids, while the destabilization effect of mutation residue I472V, as described in case of meta-substituted substrates, resulted in no detectable activity. Instead, the mutation H137V, provided minor to moderate conversion increase of 7.9–23.1%, in case of p-Br- and p-CF3-substituted substrates 1i, 1j and 2i, 2j, where the beneficial effect of the mutation is also supported by the significantly increased kcat values. The lower catalytic efficiency of H137V RtPAL, reflected in significantly lower conversion values, in comparison to its homologue variants F136V AtPAL and F137V PcPAL, might result from the presence of polar Q138 residue in the proximity of the hydrophobic, mutated V137 residue (Fig. 5), supported by the increased conversions provided by the ‘PcPAL-like’ H137V/Q138L RtPAL, approximating the conversions registered with the homologue At/Pc-PAL variants.
Active orientations of 4-(trifluoromethyl) cinnamic acid 2j within (A) wt-RtPAL (green, − 5.5 kcal/mol), H137V RtPAL (indigo, − 6.9 kcal/mol), H137V/Q138L RtPAL (grey, − 7.5 kcal/mol), and I472V RtPAL (purple, − 5.5 kcal/mol) and (B) wt-PcPAL (green, − 4.8 kcal/mol), F137V PcPAL (grey, − 8.9 kcal/mol), and I460V PcPAL (purple, − 6.4 kcal/mol). The modified residues and the active site orientation of 2j within the corresponding variant is marked with similar colour.
Considering the above described conservation of the catalytic efficiency-enhancing effect (Fig. 6) of the mutational strategy developed for PcPAL, in case of AtPAL (81% sequence identity and identical catalytic site with PcPAL) and RtPAL (38% sequence identity and TAL-activity providing catalytic site, containing H137, Q138 residues at positions analogue to F137, L138 of PcPAL) the general applicability of the rational design strategy among PALs is supported. Notable, that in case of RtPAL, besides the modification of the substrate specificity-modulator residues, replacement of residue Q138, in proximity of position 137, to hydrophobic residues, further enhanced the catalytic properties of H137V RtPAL, supporting that the mutational strategy is adaptable for further additional mutations based on simple rational considerations, allowing facile development of substrate-tailored PALs of various origins. Despite the identical catalytic site residues of AtPAL and PcPAL, in several cases AtPAL variants in comparison with the corresponding PcPAL variants, showed higher catalytic efficiencies/conversions (Fig. 6), highlighting that besides active site residues, other structural elements also determine the different enzyme activities/substrate specificities of PALs of different origins. Besides, the mutational approach revealed several PAL variants, such as L133A AtPAL, I461V AtPAL, L266V and H137V/Q138L RtPAL, which in comparison with their previously reported2 PcPAL homologues, possess enhanced catalytic efficiency within the various ammonia additions producing valuable l-phenylalanines (Fig. 6).
Conversion into l-phenylalanines l-1a–l obtained within the ammonia additions reactions catalyzed by the wild-type PcPAL (orange), AtPAL (blue) and RtPAL (grey) overlayed with the conversions provided by their best performing mutants (marked with non-filled boxes overlayed with the coloured lanes, representing the conversions of wild-type variants), evidencing the conversion-based activity increase provided by the mutations (white zone of each lane-box).
Experimental part
Site-directed mutagenesis
The codon optimized genes encoding PALs from Arabidopsis thaliana and Rhodosporidium toruloides were obtained through the synthesis services of GenScript, followed by their cloning into pET19b vector (using XhoI and Bpu1102I cloning sites for RtPAL and XhoI and NdeI cloning sites for AtPAL). The site-directed mutagenesis was performed following the protocol described by Naismith and Liu25, using as template the pET-19b vector harbouring the gene encoding PALs from Arabidopsis thaliana and Rhodosporidium toruloides, respectively. Using as homology model the active site of PcPAL6 (PDB ID 1W2726), several residues from the hydrophobic binding pocket were selected for point mutations, namely L133/L134 (to A), F136/H137 (to V), L257/L266 (to V) and I461/I472 (to V). New MIO-enzyme libraries were created at these positions and screened for activity towards the substrates. The primers used within the mutagenesis are listed in Table S1.
Protein expression, purification
Expression, isolation and purification of wild-type RtPAL and its mutant variants (L134A, H137V, L266V, I472V, H137F/Q138L/L266V, H137F/Q138L/I472V, H137F/Q138L, H137V/Q138L) was performed according to our optimized protocol via immobilized affinity chromatography (IMAC)27. In case of AtPAL (wt- and L133A, F136V, L257V, I461V mutants), precultures were prepared at 37 °C, 200 rpm, overnight in 50 mL LB (Luria Bertani) medium supplemented with carbenicillin (50 μg/mL) and chloramphenicol (30 μg/mL) from glycerol stocks of E.coli Rosetta (DE3)plysS cells harbouring the pET-19b vector carrying the wt- or mutant atpal gene. 2% (v/v) from the starter culture was used to inoculate 2 × 500 mL LB medium in 2 L flasks. The OD600 was monitored and when a value of 0.45 was reached, the temperature was lowered from 37 to 25 °C, and the shaking continued till an OD600 value of 0.6–0.8, when PAL expression was induced via IPTG (0.5 mM final concentration). The cell growth continued at 25 °C, 200 rpm for another 6 h, when cells were harvested by centrifugation at 4000 rpm (1751×g), 4 °C for 20 min. The supernatant was discarded and the cell pellet was stored at − 20 °C until further use or processed immediately using the optimized protein isolation protocol as described for PcPAL27.
Thermal unfolding profile of purified proteins
The thermal unfolding of all PALs was determined by nanoscale differential scanning fluorimetry measurements, using Prometheus NT.48 nanoDSF instrument (NanoTemper Technologies, München, Germany). PAL variants were diluted with 20 mM Tris, 120 mM NaCl pH 8.8 buffer to a final concentration of 1 mg/mL. 10 μL of each sample were loaded into UV capillaries (NanoTemper Technologies) and unfolding of PAL enzymes was detected during heating in a linear thermal ramp of 1.5 °C/min between 20 and 95 °C, with an excitation power of 70%. Data analysis was performed using NT Melting Control software and melting temperature (Tm) was determined by fitting the experimental data using a polynomial function, in which the maximum slope is indicated by the peak of its first derivative (F350/F330). All measurements were performed in triplicate (Figs. S3, S4 and Table S2).
Preparation of whole-cell PAL biocatalysts
The overnight precultures were prepared in 20 mL LB (Luria Bertani) medium supplemented with carbenicillin (50 μg/mL) and chloramphenicol (30 μg/mL) in 100 mL Erlenmeyer flasks, being inoculated with glycerol stocks of E. coli Rosetta (DE3) pLysS cells harboring the pET-19b vector carrying the wt or mutant atpal or rtpal gene, followed by incubation at 37 °C and shaking at 200 rpm. 2% (v/v) of the overnight culture was used to inoculate 50 mL LB medium. Cultures were grown at 37 °C, 200 rpm until OD600 reached 0.6–0.8 (approx. 3 h), when enzyme production was induced via the addition of 0.5 mM IPTG (final concentration), and the cell growth was maintained at 20 °C, 200 rpm, overnight (approx. 17 h). The final OD600 was measured for each mutant variant and wild-type PAL. The culture volumes required for the biotransformation screenings were harvested by centrifugation in 1.5 mL polypropylene tubes for 10 min at 13,300 rpm (12,000×g). The required volume of bacterial culture, providing the amount of whole-cell pellet needed was calculated considering the volume of the reactions, the whole-cell biocatalysts concentration (with fixed cell density OD600 of ~ 2)2,22 and the final OD600 value of the induced cells. The harvested cells were washed with 500 μL PBS buffer (20 mM phosphate, 150 mM NaCl, pH 8.0) (13,300 rpm, 12,000×g, 10 min) and stored at − 20 °C until further use.
Analytical scale ammonia addition and elimination reactions
The bacterial pellet of PAL-biocatalysts (prepared as described above, in 1.5 mL polypropylene tubes) was resuspended to an OD600 of ~ 2, in 500 µL substrate solution (2 mM cinnamic acids 2a–l or 2 mM racemic amino acids rac-1a–l) prepared in 6 M NH4OH buffer pH 10 adjusted with CO2 (in case of ammonia addition) or 20 mM Tris.HCl, 120 mM NaCl buffer, pH 8.8 (in case of ammonia elimination). The reaction mixtures were incubated at 30 °C, 250 rpm. Reaction samples were taken after 3, 6, 16, and 24 h and quenched by adding an equal volume of MeOH, vortexed and centrifuged (13,400 rpm, 12,000 g×, 10 min). The supernatant was filtered through a 0.22 μm nylon membrane filter prior to analysis by HPLC. In order to determine the conversions values, a Gemini NX-C18 column (150 × 4.5 mm; 5 µm) was chosen, using as mobile phase: A: NH4OH buffer (0.1 M, pH 9.0)/B: MeOH, with a flow rate of 1.0 mL/min. The enantiomeric excess values were determined by chiral HPLC separations, using Crownpak CR-I (+) chiral column (150 × 3 mm; 5 µm) and HClO4 (pH = 1.5)/acetonitrile as mobile phase at a flow rate of 0.4 mL/min. HPLC methods and response factors used for the conversion value determinations, as well as retention times of the enantiomers of rac-1a–l can be consulted in our previous reports2,6. All analytical scale biotransformations were performed in duplicates, while during the initial activity screens using a significantly sized reaction-subset the HPLC analysis have been performed for all samples within the duplicate set (see details in Supporting information, Chapter 6, Table S3).
Enzyme kinetics
The initial enzyme activities were spectrophotometrically determined, using a Tecan Infinite Spark 10 M microplate reader and Corning 96-well Clear Flat Bottom UV-Transparent microplates. The kinetic measurements were performed in triplicate at 30 °C by monitoring the production of trans-cinnamic acid analogues 2a–l at 290 nm (wavelength where the corresponding amino acids rac-1a–l showed no absorption), using substrate concentrations of 0.1–20 mM of 1a–l, 100 mM Tris.HCl, 120 mM NaCl (pH 8.8) as buffer and purified PAL variants at fixed enzyme concentration of 0.322 μM. Kinetic constants (KM, vmax) were obtained from the Michaelis–Menten curves by non-linear fitting. Standard deviations for the determined kinetic parameters are given within Tables S4–S6 (Supporting information).
Computational studies
The ground state geometries of the monosubstituted cinnamic acid derivatives 2a–l were obtained by calculations based on the density functional theory, performed using the Gaussian 09 software28 by employing the B3LYP density functional and the 6-31G(d,p) basis set. Geometry optimizations were carried out in a water solvated environment using the Polarizable Continuum Model (PCM)29.
The molecular docking calculations were performed with the Autodock Vina software30, using flexible-ligand and rigid-receptor docking. The search space was defined by embedding the binding site residues and the MIO prosthetic group. In both cases the receptor grid was defined as a cubic box with the dimension of 20 Å × 20 Å × 20 Å. The exhaustiveness search parameter of Vina was increased to 100.
The crystal structure of PcPAL was retrieved from Protein Data Bank entry 6F6T31, whereas in case of RtPAL, the AlphaFold32 predicted model was retrieved from the UniProt database (entry P11544)33. The assembled tetrameric structure was submitted for minimization using the YASARA web server34. Although crystal structures of RtPAL are available, PDB entries 1T6J and 1Y2M, both structures present the open conformation of the protein, missing the loop containing the Y110 residue, responsible for the catalytic site closure upon substrate binding.
Conclusions
Within this study we tested the applicability of the mutational strategy developed for PAL from Petroselinum crispum to other PALs with the aim to provide a general rational design strategy, highly desirable for developing substrate-tailored PALs of diverse substrate scope and origins. Accordingly, AtPAL and RtPAL, both well-characterized PAL representatives, that share different sequence identity (high degree of 81%, respectively low degree of 38%) to PcPAL, with RtPAL known to possess dual PAL/TAL-activity, were selected for this purpose. As expected, wild-type RtPAL with low sequence identity to Pc/AtPAL, showed different substrate specificity towards the substrate library, revealing its higher catalytic efficiency towards meta-substituted substrates in both ammonia elimination and ammonia addition reaction routes, while the substrate specificities of wt-Pc/AtPAL have been found very similar. However, the enzyme activity tests of the generated focused AtPAL, RtPAL, PcPAL mutant library towards the mono-substituted substrates revealed that AtPAL variants, with some exceptions, surpassed in terms of conversion and catalytic efficiency the corresponding, previously reported PcPAL homologues (L134A, L256V, F137V and I460V). Since their active sites possess identical residues, the results highlight that besides active site residues, other structural elements also determine the different enzyme activities/substrate specificities of PALs. Furthermore, the activity of PAL variants tailored towards substrates of different (ortho-, meta-, para-) substitution pattern, revealed that the mutational approach is applicable among different PALs, resulting the expected catalytic efficiency increase towards the targeted non-natural substrates, with minor sequence alignment-based rational refinements further improving its efficacy. Accordingly, in case of RtPAL, besides the modification of the substrate specificity modulator residues L266V, L134A, F137V and I472V, replacement of residue Q138, in proximity of mutated position 137, to hydrophobic residues, further enhanced the catalytic properties of RtPAL variants. In this context, the study paves the way and contributes for the development of the general rational design strategy among the PAL (E.C. 4.3.1.24) and PAL/TAL families (E.C. 4.3.1.25).
Data availability
The Uniprot identifiers of all protein sequences used within the alignments and experimental work and the Protein Data Bank (PDB) IDs for the protein structures used within the computational part are described within the manuscript, while other datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Parmeggiani, F., Weise, N. J., Ahmed, S. T. & Turner, N. J. Synthetic and therapeutic applications of ammonia-lyases and aminomutases. Chem. Rev. 118, 73–118. https://doi.org/10.1021/acs.chemrev.6b00824 (2018).
Nagy, E. Z. A. et al. Mapping the hydrophobic substrate binding site of phenylalanine ammonia-lyase from petroselinum crispum. ACS Catal. 9, 8825–8834. https://doi.org/10.1021/acscatal.9b02108 (2019).
Parmeggiani, F., Lovelock, S. L., Weise, N. J., Ahmed, S. T. & Turner, N. J. Synthesis of D- and L-phenylalanine derivatives by phenylalanine ammonia lyases: A multienzymatic cascade process. Angewandte Chemie - International Edition 54, 4608–4611. https://doi.org/10.1002/anie.201410670 (2015).
Ahmed, S. T., Parmeggiani, F., Weise, N. J., Flitsch, S. L. & Turner, N. J. Engineered ammonia lyases for the production of challenging electron-rich l -phenylalanines. ACS Catal. 8, 3129–3132. https://doi.org/10.1021/acscatal.8b00496 (2018).
Gloge, A., Zoń, J., Kövári, A., Poppe, L. & Rétey, J. Phenylalanine ammonia-lyase: The use of its broad substrate specificity for mechanistic investigations and biocatalysis—Synthesis of L-arylalanines. Chem. Eur. J. 6, 3386–3390. https://doi.org/10.1002/1521-3765(20000915)6:18%3c3386::AID-CHEM3386%3e3.0.CO;2-5 (2000).
Filip, A. et al. Tailored mutants of phenylalanine ammonia-lyase from petroselinum crispum for the synthesis of Bulky l- and d-Arylalanines. ChemCatChem 10, 2627–2633. https://doi.org/10.1002/cctc.201800258 (2018).
Lovelock, S. L. & Turner, N. J. Bacterial Anabaena variabilis phenylalanine ammonia lyase: A biocatalyst with broad substrate specificity. Bioorg. Med. Chem. 22, 5555–5557. https://doi.org/10.1016/j.bmc.2014.06.035 (2014).
Moffitt, M. C. et al. Discovery of two cyanobacterial phenylalanine ammonia lyases: Kinetic and structural characterization. Biochemistry 46, 1004–1012. https://doi.org/10.1021/bi061774g (2007).
Renard, G., Guilleux, J. C., Bore, C., Malta-Valette, V. & Lerner, D. A. Synthesis of L-phenylalanine analogs by Rhodotorula glutinis. Bioconversion of cinnamic acids derivatives. Biotechnol. Lett. 14, 673–678. https://doi.org/10.1007/BF01021641 (1992).
Yamada, S., Nabe, K. & Izuo, N. Production of L-phenylalanine from trans-cinnamic acid with Rhodotorula glutinis containing L-phenylalanine ammonia-lyase activity. Appl. Environ. Microbiol. 42, 773–778. https://doi.org/10.1128/aem.42.5.773-778.1981 (1981).
Dreßen, A. et al. Phenylalanine ammonia lyase from Arabidopsis thaliana (AtPAL2): A potent MIO-enzyme for the synthesis of non-canonical aromatic alpha-amino acids: Part I: Comparative characterization to the enzymes from Petroselinum crispum (PcPAL1) and Rhodosporidium toruloides (RtPAL). J. Biotechnol. 258, 148–157. https://doi.org/10.1016/j.jbiotec.2017.04.005 (2017).
Weise, N. J. et al. Zymophore identification enables the discovery of novel phenylalanine ammonia lyase enzymes. Sci. Rep. 7, 1. https://doi.org/10.1038/s41598-017-13990-0 (2017).
Varga, A. et al. A novel phenylalanine ammonia-lyase from Kangiella koreensis. Stud. Univ. Babes-Bolyai, Chem. 62, 293–308. https://doi.org/10.24193/subbchem.2017.3.25 (2017).
Varga, A. et al. A novel phenylalanine ammonia-lyase from Pseudozyma antarctica for stereoselective biotransformations of unnatural amino acids. Catal. Today 366, 185–194. https://doi.org/10.1016/j.cattod.2020.04.002 (2021).
Kempa, E. E. et al. Rapid screening of diverse biotransformations for enzyme evolution. Jacs Au 1, 508–516. https://doi.org/10.1021/jacsau.1c00027 (2021).
Rétey, J. Discovery and role of methylidene imidazolone, a highly electrophilic prosthetic group. Biochim. Biophys. Acta Proteins Proteomics 1647, 179–184. https://doi.org/10.1016/S1570-9639(03)00091-8 (2003).
Poppe, L. & Rétey, J. Friedel-crafts-type mechanism for the enzymatic elimination of ammonia from histidine and phenylalanine. Angewandte Chemie Int. Edition 44, 3668–3688. https://doi.org/10.1002/anie.200461377 (2005).
Bartsch, S. & Bornscheuer, U. T. A single residue influences the reaction mechanism of ammonia lyases and mutases. Angewandte Chem. Int. Edition 48, 3362–3365. https://doi.org/10.1002/anie.200900337 (2009).
Csuka, P. et al. Pseudomonas fluorescens strain R124 Encodes three different MIO enzymes. ChemBioChem 19, 411–418. https://doi.org/10.1002/cbic.201700530 (2018).
Calabrese, J. C., Jordan, D. B., Boodhoo, A., Sariaslani, S. & Vannelli, T. Crystal structure of phenylalanine ammonia lyase: Multiple helix dipoles implicated in catalysis. Biochemistry 43, 11403–11416. https://doi.org/10.1021/bi049053+ (2004).
Baedeker, M. & Schulz, G. E. Structures of two histidine ammonia-lyase modifications and implications for the catalytic mechanism. Eur. J. Biochem. 269, 1790–1797. https://doi.org/10.1046/j.1432-1327.2002.02827.x (2002).
Tork, S. D. et al. The production of l- and d-phenylalanines using engineered phenylalanine ammonia lyases from Petroselinum crispum. Sci. Rep. 9, 1. https://doi.org/10.1038/s41598-019-56554-0 (2019).
Weise, N. J. et al. Intensified biocatalytic production of enantiomerically pure halophenylalanines from acrylic acids using ammonium carbamate as the ammonia source. Catal. Sci. Technol. 6, 4086–4089. https://doi.org/10.1039/c6cy00855k (2016).
Rowles, I. et al. Engineering of phenylalanine ammonia lyase from Rhodotorula graminis for the enhanced synthesis of unnatural L-amino acids. Tetrahedron 72, 7343–7347. https://doi.org/10.1016/j.tet.2016.06.026 (2016).
Liu, H. & Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91. https://doi.org/10.1186/1472-6750-8-91 (2008).
Ritter, H. & Schulz, G. E. Structural basis for the entrance into the phenylpropanoid metabolism catalyzed by phenylalanine ammonia-lyase. Plant Cell 16, 3426–3436. https://doi.org/10.1105/tpc.104.025288 (2004).
Dima, N. A. et al. Expression and purification of recombinant phenylalanine ammonia-lyase from Petroselinum crispum. Stud. Univ. Babes-Bolyai, Chem. 61, 21–34 (2016).
Gaussian 16 Rev. C.01 (Wallingford, CT, 2016).
Tomasi, J., Mennucci, B. & Cammi, R. Quantum mechanical continuum solvation models. Chem. Rev. 105, 2999–3093. https://doi.org/10.1021/cr9904009 (2005).
Trott, O. & Olson, A. J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Bata, Z. et al. Substrate tunnel engineering aided by X-ray crystallography and functional dynamics swaps the function of MIO-enzymes. ACS Catal. 11, 4538–4549. https://doi.org/10.1021/acscatal.1c00266 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589. https://doi.org/10.1038/s41586-021-03819-2 (2021).
Bateman, A. et al. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489. https://doi.org/10.1093/nar/gkaa1100 (2021).
Krieger, E. et al. Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: Four approaches that performed well in CASP8. Proteins Struct. Funct. Bioinf. 77, 114–122. https://doi.org/10.1002/prot.22570 (2009).
Acknowledgements
This work was financed by the Swiss National Science Foundation (SNF) project PROMYS, grant no. IZ11Z0_166543 and by the Romanian Ministry of Education and Research, CNCS–UEFISCDI, project number PN-III-P1-1.1-TE-2019-2118, within PNCDI III. M.E.M. thanks for financial support from the Romanian Ministry of Education and Research, National Research Council-UEFISCDI, project number PN-III-P1-1.1-PD-2019-1188. L.C. thanks for the STAR-Institute of the Babeş-Bolyai University and ELTE Márton Áron Special College for the provided student-research fellowships.
Funding
This article was funded by Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung, PROMYS, grant no. IZ11Z0_166543, Romanian Ministry of Education and Research, CNCS-UEFISCDI, PN-III-P1-1.1-TE-2019-2118, PN-III-P1-1.1-PD-2019-1188.
Author information
Authors and Affiliations
Contributions
S.D.T. and M.E.M. contributed to the work equally. S.D.T. was responsible for the preparation of whole-cell-biocatalysts, analytical scale biotransformations and their HPLC monitoring. M.E.M. was responsible for enzyme kinetic measurements. Isolation and purification of PAL variants, mutant library generation was performed by S.D.T., M.E.M. and A.F, while L.C. was involved in substrate synthesis and biotransformation-monitoring by HPLC. L.C.N. performed the computational studies and was responsible for the graphical artworks. L.C.B. conceived the project and was responsible for funding, supervised all experiments, data and wrote the paper together with F.D.I, S.D.T., M.E.M. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tork, S.D., Moisă, M.E., Cserepes, L. et al. Towards a general approach for tailoring the hydrophobic binding site of phenylalanine ammonia-lyases. Sci Rep 12, 10606 (2022). https://doi.org/10.1038/s41598-022-14585-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14585-0
This article is cited by
-
Phenylalanine ammonia-lyases: combining protein engineering and natural diversity
Applied Microbiology and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.