Abstract
In the development of polymer materials, it is an important issue to explore the complex relationships between domain structure and physical properties. In the domain structure analysis of polymer materials, 1H-static solid-state NMR (ssNMR) spectra can provide information on mobile, rigid, and intermediate domains. But estimation of domain structure from its analysis is difficult due to the wide overlap of spectra from multiple domains. Therefore, we have developed a materials informatics approach that combines the domain modeling (http://dmar.riken.jp/matrigica/) and the integrated analysis of meta-information (the elements, functional groups, additives, and physical properties) in polymer materials. Firstly, the 1H-static ssNMR data of 120 polymer materials were subjected to a short-time Fourier transform to obtain frequency, intensity, and T2 relaxation time for domains with different mobility. The average T2 relaxation time of each domain is 0.96 ms for Mobile, 0.55 ms for Intermediate (Mobile), 0.32 ms for Intermediate (Rigid), and 0.11 ms for Rigid. Secondly, the estimated domain proportions were integrated with meta-information such as elements, functional group and thermophysical properties and was analyzed using a self-organization map and market basket analysis. This proposed method can contribute to explore structure–property relationships of polymer materials with multiple domains.
Similar content being viewed by others
Introduction
Traditional design approaches for materials are experimentally driven, facing significant challenges due to the vast design space of materials. Experimental science can be supported by materials informatics1,2,3 that makes full use of theoretical and computational science such as density functional theory (DFT)4,5 and molecular dynamics (MD)6 and data science using computers (Artificial Intelligence; AI)7,8. Computational science solves equations numerically based on theory and physical models. On the other hand, data science explores candidate materials with a certain function from a large quantity of material data. Its physical meaning needs to be verified by experimental, theoretical and computational science. The meta-information involved in the production process of the material also plays an important role in desired materials development9,10.
In recent years, the development of sustainable polymers that meet the needs of consumers without destroying the environment has become an important issue due to global problems such as marine pollution, waste disposal, and global warming caused by plastics11,12. Furthermore, “carbon–neutral” bio-based polymers, such as polylactic acid (PLA), polybutylene succinate (PBS), poly(3-hydroxybutyrate-co-3-hydroxyhexanoate) (PHBH), and poly-ε-caprolactone (PCL), have become a focus in the era of biorefinery materials as an alternative to oil-based materials13,14. Polymers such as PLA15, PCL16, are multiple domain systems, are often employed as high-performance materials which can display various properties.
Solid-state nuclear magnetic resonance (ssNMR) spectroscopy is a powerful tool that is used to characterize the native structure, components and dynamics of solid-state samples at the atomic level, and has been increasingly applied in material sciences17,18. In addition, NMR measurements, especially low magnetic field NMR, is a method for routine material evaluations, which has produced a lot of NMR datasets19. Typical ssNMR methods are cross-polarization (CP)/magic-angle spinning (MAS) methods with elimination of linewidth broadening due to chemical shift anisotropy for high resolution. On the other hand, there are different complementary approaches to tackle complexity of polymer domain structure. 1H-static ssNMR can be applied to quantify domain mobility in terms of dynamic heterogeneity20. In addition, the use of magic-and-polarization echo (MAPE)21 and double-quantum (DQ)22 filters can determine the spectral parameters for the mobile amorphous domains with the long-time decay and the strongly dipole–dipole-coupled crystalline domains with the quickly decay, respectively.
In the case of characterization of a solid-state sample with domains of rigid, intermediate and mobile types, the 1H-static ssNMR measurement is useful as a measure of the kinetic nature of higher order structures, although its analysis is difficult because the spectrum is broadened and overlapped23. Therefore, application of signal deconvolution is needed to characterize structure and property of the sample. Several methods for spectral separation19, fitting and numerical simulation24 such as SIMPSON25, SPINEVOLUTION26, dmfit27, EASY-GOING deconvolution28, INFOS29, Fityk30, ssNake31, and a noise reduction method based on principal component analysis32 have been developed. So far, in NMR data analysis, signal simulation and fitting have targeted only the frequency domain or the time domain. In our previous study, we proposed signal deconvolution methods that combines short-time Fourier transform (STFT; a time–frequency analytical method) and probabilistic sparse matrix factorization33, and non-negative tensor/matrix factorization34. In our method using STFT, by simulating the signal for both the frequency and time domain, it was possible to separate the signal related to the motility characteristics of the domain structure based on the indicators of chemical shift and T2 relaxation time. The NMR signal can be calculated by functions such as Lorentzian35, Gaussian36, and Voigt37 in the frequency domain, and by the T2 relaxation equation38 in the time domain. In addition, the difference in T2 relaxation times can be adjusted by the Weibull coefficient39. Analysis of the relaxation time of a sample's free-induction decay (FID) provides important insights into the chemical composition, structure, and mobility of the sample38,40.
The polymer domain structure has a significant influence on their macroscopic properties41. Materials informatics, which is the emerging field, support analysis of relationships of structure and property from materials data sets2,3,7,8,42. NMR signal has potential for use as a descriptor having the structural features of the molecules contributing to their physical/chemical/biological properties43. In previous studies, a self-organization map (SOM) has been applied to tool wear monitoring44. Market basket analysis (MBA) has been applied to predict drug-drug interactions45. Bayesian optimization has been applied in real and virtual degradable experiments of bioplastics46. Generative topographic mapping regression (GTMR) has also been applied to the analysis of CP-MAS spectra to predict 13C NMR spectrum of the material in its solid-state based on its thermophysical properties34. Machine learning methods have applied for various material studies such as cloud-point engineering of polymers47, prediction of drug-polymer amorphous solid dispersion miscibility and stability48, atomic/inter-atomic properties prediction49, solubility prediction50, descriptor selection for investigating physical properties of biopolymers in hairs51, classification of the membrane materials52, prediction of crystallization tendency53, prediction of density, glass transition temperature, melting temperature, and dielectric constants of polymer9, macromolecular modeling54.
In this study, we propose a materials informatics approach to explore the structure–property relationships of polymers that combines the polymer domain modeling and the integrated analysis of polymer materials meta-information. For polymer domain modeling, 1H-static ssNMR spectral parameters obtained using STFT were utilized, including T2 relaxation time, frequency, and intensity. The domain structure with different mobility in the polymeric material was estimated by fitting the physical indices such as T2 relaxation time, frequency, and linewidth. In addition, using a SOM and MBA, the relationships between the estimated domain structure and the meta-information such as elements, functional group, and thermophysical property were explored.
Results and discussion
A materials informatics approach to exploring structure–property relationships using domain modeling
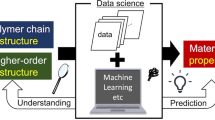
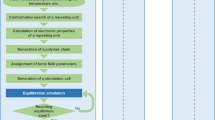
The conceptual diagram of materials informatics approach to exploring structure–property relationships using domain modeling is shown in the Fig. 1. The detailed analytical flow of this method is shown in Supporting Information Fig. S1. We have utilized the input polymer information that are 1H-static ssNMR (1H-static ssNMR) data, primary structure of the polymer, and thermophysical property data (TG/DTA/DTG). Then, frequency and time information are obtained by STFT against FIDs obtained by 1H-static ssNMR. The domain modeling method is the following. The domain components are firstly separated by fitting the obtained frequency and time information (Fig. S2). Secondly, the domain component ratio is calculated by 3D modeling (Fig. S3). To reduce the error between total of the simulated values of domain components using our signal processing method and the original static data, Bayesian optimization as one of the optimization methods was performed using Eq. 1 (see in the Materials Methods section) in the text to search for the ratio between the mobile and rigid components according to Eqs. (2–5). After that, we performed statistical analysis, MBA, and SOM, which are materials informatics methods, on the obtained domain information, primary structure information, and thermophysical property information to associate the structure and physical properties. The detailed results are shown in the following sections.
Conceptual diagram of materials informatics approach to exploring structure–property relationships using domain modeling. (a) To exploring relationships between structure and property of polymer materials, 1H-static solid-state NMR (ssNMR) spectrum, primary structure information and physical property data obtained from TG, DTA and DTG were used. (b) Domain modeling was applied to the input 1H-static ssNMR spectrum of polymer materials to obtain domain structural information. (c) The domain proportion, primary structure data and thermophysical property data were integrated for data analysis such as statistical analysis, SOM and MBA. 1H-static ssNMR, Proton static solid-state nuclear magnetic resonance; TG, Thermogravimetry; DTA, Differential thermal analysis; DTG, Derivative thermogravimetry; Tm, Melting temperature; Td, Thermal decomposition temperature; Tg, Glass transition temperature; STFT, Short-time Fourier transform; MBA, Market basket analysis; SOM, Self-organization map.
Results of domain modeling of polymer materials using time–frequency simulation of 1H-static ssNMR spectra
The ratios of the domain components were calculated from the volume ratios obtained from domain modeling of 71 samples of the polymer materials (Fig. 2). Here, the domain component is defined as a region of higher-order structure distinguished by molecular mobility in polymers. NMR characterizes this domain component by analyzing the spectral widths or T2 relaxation time. The volumes of four domain component (Mobile, Intermediate (Mobile), Intermediate (Rigid) and Rigid) are calculated from Eqs. 2–5 (\(M_{Mobile}\), \(M_{IM}\), \(M_{IR}\) and \(M_{Rigid}\)). The volume ratio is defined as the ratio of the volume of each domain component estimated by 3D simulation based on those equations to total volume. We have classified analyzed data in this study into four domain components: Mobile, Intermediate (Mobile), Intermediate (Rigid), and Rigid domain components, those we regard to mobile, slightly mobile, slightly rigid, and rigid components in their material states, respectively. As a result, the domain component ratios indicated differences among not only different polymer materials but also similar ones that composed of the same monomers, which can be attributed to the molding conditions and molecular weight. In the case of PCL (Fig. 2, upper right), which is the sample with the most Mobile domain component: the Mobile domain component ratio was 37.7%; the Intermediate (Mobile) domain component ratio was 11.2%; the Intermediate (Rigid) domain component ratio was 18.1%; and the Rigid domain component ratio was 33.0%. In the case of PHBH sample (Fig. 2, upper left), which has the highest Rigid domain component: the Mobile domain component ratio was 10.4%; the Intermediate (Mobile) domain component ratio was 3.1%; the Intermediate (Rigid) domain component ratio was 36.5%; and the Rigid domain component ratio was 50.5%. From this result, we were able to calculate the difference in domain ratios caused by the different monomers used to synthesize the polymer.
Results of domain proportion calculation of polymer materials. The 1H-static ssNMR spectra of 71 polymer materials were separated by domain modeling. Based on the results, the domain models are shown. (a) An example of PHBH, (b) an example of PCL, and (c) the domain ratios of 71 polymer materials. Polymer sample ID numbers in the figure refer to Table S2. Red: Mobile, Magenta: Intermediate (Mobile), Violet: Intermediate (Rigid), Blue: Rigid.
The calculated distributions of T2 relaxation time and thermophysical properties of the domain components of each polymer are shown (Fig. 3). The box-and-whisker plot shows that the average T2 relaxation time information is 0.96 ms for Mobile, 0.55 ms for Intermediate (Mobile), 0.32 ms for Intermediate (Rigid), and 0.11 ms for Rigid. The weighted average (WA) of the polymer materials showed a distribution among the samples, among which PCL (Fig. 3, red squares) was high and PHBH (Fig. 3, yellow circles) was low. The same was true for the results of thermal analysis spectral data (Table S3). Based on the domain component ratios, the estimated domain ratio diagrams were inserted for the highest PCL and lowest PHBH of WA.
Diagram of polymer properties with T2 relaxation time among four domain components and melting temperature. M: Red box plot; Mobile, I(M): Pink box plot; Intermediate (Mobile), I(R): Violet box plot; Intermediate (Rigid), R: Blue box plot; Rigid, WA: Weighted average, Tm: Melting temperature. x: Average T2 relaxation time. The box and whisker plot displays the minimum, first quartile, median, third quartile, maximum, and outliers. The estimated domain proportion models were inserted for the highest PCL and lowest PHBH of WA.
Self-organizing map analysis integrating domain proportions and quantitative spectral data in polymer materials
In order to evaluate the relationships between domain structure and thermophysical properties of polymer materials, we integrated the domain proportion described in the previous section, 13C-CP/MAS spectra55, which easily reflect primary chemical structure, and quantitative thermophysical data (thermogravimetry (TG), differential thermal analysis (DTA), derivative thermogravimetry (DTG), differential scanning calorimetry (DSC))56. To capture the characteristics of the integrated data, clustering by SOM was performed (Fig. 4). For input data of SOM, the domain component information calculated by the component separation method and meta-information of thermophysical properties, elements, and functional groups used is listed in Tables S1 and S2. These materials clustered in the following way i.e., navy blue circle symbols are polyethylene terephthalate (PET), light blue circle symbols are polyethylene (PE), and the clusters of these polymer materials are on the top, blue triangle symbols are poly(butylene adipate-co-terephthalate) (PBAT), light pink circle symbols are polylactic acid (PLA), light green circle symbols are poly(hexano-6-lactam) (nylon), and the clusters of these polymer materials are on the bottom left, black cross symbols are polybutylene succinate (PBS), orange diamond symbols are poly(butylene succinate-co-butylene adipate) (PBSA), and red square symbols are PCL, and the clusters of these polymer materials are found to exist solidly in the lower right corner, respectively. These clusters formed two groups: high heat resistant polymers with many rigid domain components, and low heat resistant polymers with many mobile domain components57,58. The results of thermal analysis for PET, PE, PBAT, PLA, nylon, PBS, PBSA, and PCL were consistent with their characteristics shown above (Fig. S4, Table S3).
Result of SOM. Visualization of 3D SOM compressed into 2D SOM with 4 × 6 segments, red: positive, blue: negative. red square: PCL, orange: PBSA, black cross: PBS, blue triangle: PBAT, yellow circle: PHBH, light pink: PLA, light green: nylon, white: PES, light blue: PE, navy blue: PET, black bar: other polymers. Polymer numbers in the figure refer to Table S1.
Market basket analysis integrating quantitative domain proportion and qualitative meta-information in polymer materials
The MBA was performed to evaluate the relationships between the quantitative domain proportion as well as qualitative meta-information such as elements (linearly connected methylenes > 4 carbons), functional groups (aromaticity), and thermophysical properties. For input data of MBA, the domain component information calculated by the component separation method and meta-information of thermophysical properties, elements, and functional groups used is listed in Tables S1, S2 and S3. Using the transaction data based on the MBA, a network diagram is shown in Fig. 5, where the T2 information of the four domain components (Mobile, Intermediate (Mobile), Intermediate (Rigid), and Rigid) shows a high lift value with the primary structure information (aromaticity, linearly connected methylenes > 4 carbons). While the thermophysical properties (melting temperature, Tm; thermal decomposition temperature, Td; glass transition temperature, Tg) show a dominant lift value with their structure information. The lift value here is one of the indicators for correlation analysis in MBA. Figure S5 shows the MBA network diagram using the temperature information from thermal analysis, where the Mobile domain ratio correlated with the lower temperature (< 100 °C) of thermophysical properties. The Intermediate (Mobile) domain ratio from the MAPE Filter and the Intermediate (Rigid) domain ratio from the DQ Filter showed intermediate thermophysical properties (100 to 300 °C). The rigid domain ratio correlated with the lower temperature (< 100 °C) and of the higher temperature (> 300 °C) thermophysical properties. In common, PCL is a thermoplastic biodegradable polyester with good thermal processability and low melting point57. In PCL, there is a melting temperature at 66.5 °C of DTG (Fig. S4, Table S3). While the DTG peak of lower temperature is correlated with the Rigid domain (Fig. S5a).
Selected MBA network for T2 relaxation time data. From the results of the market basket analysis, only the relationships between the simulated domain structure information and T2 relaxation time data, experimental melting temperature (Tm), thermal decomposition temperature (Td), glass transition temperature (Tg), and data on primary structure were extracted. M: Mobile domain, Im: MAPE Filter-derived Intermediate domain (Intermediate (Mobile)), Ir: DQ Filter-derived Intermediate domain (Intermediate (Rigid)), R: Rigid domain, T2(M): T2 relaxation time information for mobile domain, T2(Im): T2 relaxation time information for Intermediate (Mobile) domain, T2(R): T2 relaxation time information for rigid domain, T2(Ir): T2 relaxation time information for Intermediate (Rigid) domain, > 4 carbon: linearly connected methylenes > 4 carbons, Aroma: aromaticity, Amounts: sample amounts.
Conclusion
In the development of materials, in addition to the chemical structure from the primary to the higher-order, meta-information such as molding process and additives are important factors because these have great influences on the final material properties. We have developed a materials informatics approach that combines the domain modeling and the integrated analysis of materials meta-information. To estimate the domain structure information, we have introduced a time–frequency simulation method for calculating multiple domain components from 1H-static ssNMR spectra. In our integrated analysis of domain proportions and meta-information, SOM was a useful tool for capturing trends across polymer material data. On the other hand, MBA was able to investigate the strong relationships between structure and meta-information in individual materials, including qualitative data as well as quantitative data. The relationships between mobility of domain structure and melting temperature were similar to the results shown by SOM and MBA (Figs. 3, 4, 5)57,59. This materials informatics approach is expected to efficiently explore relationships between structure and properties of high-performance and low environmental impact polymer materials.
Materials and methods
Materials
Polymer materials (Table S1) were prepared using a press molding machine (H300-01, AS ONE Corp., Osaka, Japan) and molding methods reported in a previous study46.
Time–frequency simulation of 1H-static ssNMR spectra
The time–frequency simulation method was developed in Python 3, by using the packages of nmrglue60 for processing of NMR data, Scipy.signal for the Fourier transform, STFT, and mathematical processing, the curve-fit function of scipy.optimize and BayesianOptimization for the fitting process, and mpl_toolkits for visualization of 3D (time, frequency, and intensity) simulation model. Before applying this method, the FID data was phase-corrected, baseline-corrected, and inverse Fourier transformed using TopSpin (Bruker-BioSpin, MA, USA). In order to calculate the ratio between the Mobile and Intermediate (Mobile) domain components obtained from the MAPE filtered spectra, and the Rigid and Intermediate (Rigid) domain components obtained from the DQ filtered spectra, the calculation errors between the four domain components and the STFT 1H-static spectra (Static) were calculated using the following equation (Eq. 1).
After finding the α and β parameters that minimize the error, we created a 3D model of the four domain components based on the frequency and T2 relaxation time information.
The domain component ratios contained in the polymer material were calculated using the following equation (Eq. 2–5).
A detailed description of the time–frequency simulation method is given in Supporting Information Figs. S1 and S2.
The weighted average (WA) of the T2 relaxation times for a single polymer material was calculated using the following equation (Eq. 6).
Self-organizing maps of domain proportions and thermophysical data in polymer materials
In the integrated analysis of domain proportions and meta-information of polymer materials, a SOM was produced using the R package kohonen61. In order to evaluate the relationships between domain structure and thermophysical properties of polymer materials, we integrated the domain component information calculated by the component separation method, 13C-CP/MAS spectra, which easily reflect primary chemical structure, and quantitative thermophysical data (TG, DTA, DTG, DSC). To capture the characteristics of the integrated data, clustering by SOM was performed (Tables S1, S2). A basic explanation of SOM is presented in the SOM section of the Supporting Information.
Market basket analysis of domain proportions and meta-information in polymer materials
MBA was performed using the R package arules62,63. Linearly connected methylenes > 4 carbons, oxygen containing and aromaticity was set to 1 if true and 0 if false. The numeric data of domain component information calculated by the component separation method and meta-information of thermophysical properties, elements, and functional groups listed in Tables S1, S2 and S3, were converted to “high” and “low” ranked data. The “high” or “low” ranked data were defined as the top 25% or the bottom 25% of all values. Association rules were determined using criterion values of support, confidence, and lift. Since lift values < 1 do not independent relationship as association rules, this study adopted a cutoff value of 1 as a lift value threshold for association rules. In addition to this, the probabilities of random occurrences are 6.25% for support and 25% for confidence because each variable was ranked by using the top or bottom 25% of all values. The maxlen (maximum size of mined frequent item sets) were set to 2. A basic explanation of MBA is presented in the MBA section of the Supporting Information. The association network was visualized using the Cytoscape program.
Tool development for automated spectral simulation
We have created a Bayesian optimization-based50 spectral simulation tool that automates the T2 relaxation time domain fitting, frequency fitting, and 3D domain modeling of our domain component separation method. The details of the Python program including T2 relaxation time information, frequency information, and 3D domain modeling of the present domain component separation method can be obtained at https://github.com/riken-emar/matrigica. For improving the level of accuracy of the prediction, intermediate regression models were employed when performing in-phase machine learning. In addition, we developed a website dedicated to the established domain component ratio calculation, which is freely available at http://dmar.riken.jp/matrigica/.
Data availability
The analytical tool and numerical data for association analysis used in this study were deposited on the following websites: http://dmar.riken.jp/NMRinformatics/MatRigiCa.zip and http://dmar.riken.jp/NMRinformatics/DatasetForMatRigiCa.zip.
References
Ben-Sasson, A. J. et al. Design of biologically active binary protein 2D materials. Nature 589, 468–473. https://doi.org/10.1038/s41586-020-03120-8 (2021).
Jha, D. et al. Enabling deeper learning on big data for materials informatics applications. Sci. Rep. 11, 4244. https://doi.org/10.1038/s41598-021-83193-1 (2021).
Ramprasad, R., Batra, R., Pilania, G., Mannodi-Kanakkithodi, A. & Kim, C. Machine learning in materials informatics: recent applications and prospects. npj Comput. Mater. 3, 54. https://doi.org/10.1038/s41524-017-0056-5 (2017).
Ito, K., Xu, X. & Kikuchi, J. Improved prediction of carbonless NMR spectra by the machine learning of theoretical and fragment descriptors for environmental mixture analysis. Anal. Chem. 93, 6901–6906. https://doi.org/10.1021/acs.analchem.1c00756 (2021).
Ito, K., Obuchi, Y., Chikayama, E., Date, Y. & Kikuchi, J. Exploratory machine-learned theoretical chemical shifts can closely predict metabolic mixture signals. Chem. Sci. 9, 8213–8220. https://doi.org/10.1039/c8sc03628d (2018).
Mori, T. et al. Exploring the conformational space of amorphous cellulose using NMR chemical shifts. Carbohydr. Polym. 90, 1197–1203. https://doi.org/10.1016/j.carbpol.2012.06.027 (2012).
Himanen, L., Geurts, A., Foster, A. S. & Rinke, P. Erratum: Data-driven materials science: Status, challenges, and perspectives. Adv. Sci. 7, 1903667. https://doi.org/10.1002/advs.201903667 (2020).
Chen, G. et al. Machine-learning-assisted de novo design of organic molecules and polymers: Opportunities and challenges. Polymers 12, 163. https://doi.org/10.3390/polym12010163 (2020).
Ma, R. & Luo, T. PI1M: A benchmark database for polymer informatics. J. Chem. Inf. Model. 60, 4684–4690. https://doi.org/10.1021/acs.jcim.0c00726 (2020).
Granda, J. M., Donina, L., Dragone, V., Long, D. L. & Cronin, L. Controlling an organic synthesis robot with machine learning to search for new reactivity. Nature 559, 377–381. https://doi.org/10.1038/s41586-018-0307-8 (2018).
Borrelle, S. B. et al. Predicted growth in plastic waste exceeds efforts to mitigate plastic pollution. Science 369, 1515–1518. https://doi.org/10.1126/science.aba3656 (2020).
Kubowicz, S. & Booth, A. M. Biodegradability of plastics: Challenges and misconceptions. Environ. Sci. Technol. 51, 12058–12060. https://doi.org/10.1021/acs.est.7b04051 (2017).
Mohanty, A. K., Vivekanandhan, S., Pin, J. M. & Misra, M. Composites from renewable and sustainable resources: Challenges and innovations. Science 362, 536–542. https://doi.org/10.1126/science.aat9072 (2018).
Ragauskas, A. J. et al. The path forward for biofuels and biomaterials. Science 311, 484–489. https://doi.org/10.1126/science.1114736 (2006).
Inkinen, S., Hakkarainen, M., Albertsson, A. & Sodergard, A. From lactic acid to poly(lactic acid) (PLA): Characterization and analysis of PLA and its precursors. Biomacromolecules 12, 523–532. https://doi.org/10.1021/bm101302t (2011).
Schaler, K., Achilles, A., Barenwald, R., Hackel, C. & Saalwachter, K. Dynamics in crystallites of poly(epsilon-caprolactone) as investigated by solid-state NMR. Macromolecules 46, 7818–7825. https://doi.org/10.1021/ma401532v (2013).
Eden, M. Editorial for the special issue on solid-state NMR spectroscopy in materials chemistry. Molecules 25, 2720. https://doi.org/10.3390/molecules25122720 (2020).
Kikuchi, J., Ito, K. & Date, Y. Environmental metabolomics with data science for investigating ecosystem homeostasis. Prog. Nucl. Magn. Reson. Spectrosc. 104, 56–88. https://doi.org/10.1016/j.pnmrs.2017.11.003 (2018).
Yamada, S. et al. InterSpin: Integrated supportive webtools for low- and high-field NMR analyses toward molecular complexity. ACS Omega 4, 3361–3369. https://doi.org/10.1021/acsomega.8b02714 (2019).
Barenwald, R., Achilles, A., Lange, F., Ferreira, T. M. & Saalwachter, K. Applications of solid-state NMR spectroscopy for the study of lipid membranes with polyphilic guest (macro)molecules. Polymers 8, 439. https://doi.org/10.3390/polym8120439 (2016).
Demco, D. E., Johansson, A. & Tegenfeldt, J. Proton spin diffusion for spatial heterogeneity and morphology investigations of polymers. Solid State Nucl. Magn. Reson. 4, 13–38. https://doi.org/10.1016/0926-2040(94)00036-c (1995).
Buda, A. et al. Domain sizes in heterogeneous polymers by spin diffusion using single-quantum and double-quantum dipolar filters. Solid State Nucl. Magn. Reson. 24, 39–67. https://doi.org/10.1016/S0926-2040(03)00020-1 (2003).
Schaler, K. et al. Basic principles of static proton low-resolution spin diffusion NMR in nanophase-separated materials with mobility contrast. Solid State Nucl. Magn. Reson. 72, 50–63. https://doi.org/10.1016/j.ssnmr.2015.09.001 (2015).
Schneider, H., Saalwachter, K. & Roos, M. Complex morphology of the intermediate phase in block copolymers and semicrystalline polymers as revealed by 1H NMR spin diffusion experiments. Macromolecules 50, 8598–8610. https://doi.org/10.1021/acs.macromol.7b00703 (2017).
Bak, M., Rasmussen, J. & Nielsen, N. SIMPSON: A general simulation program for solid-state NMR spectroscopy. J. Magn. Reson. 147, 296–330. https://doi.org/10.1006/jmre.2000.2179 (2000).
Veshtort, M. & Griffin, R. SPINEVOLUTION: A powerful tool for the simulation of solid and liquid state NMR experiments. J. Magn. Reson. 178, 248–282. https://doi.org/10.1016/j.jmr.2005.07.018 (2006).
Massiot, D. et al. Modelling one- and two-dimensional solid-state NMR spectra. Magn. Reson. Chem. 40, 70–76. https://doi.org/10.1002/mrc.984 (2002).
Grimminck, D. et al. EASY-GOING deconvolution: Automated MQMAS NMR spectrum on a model with analytical crystallite excitation efficiencies. J. Magn. Reson. 228, 116–124. https://doi.org/10.1016/j.jmr.2012.12.012 (2013).
Smith, A. INFOS: Spectrum fitting software for NMR analysis. J. Biomol. NMR 67, 77–94. https://doi.org/10.1007/s10858-016-0085-2 (2017).
Wojdyr, M. Fityk: A general-purpose peak fitting program. J. Appl. Crystallogr. 43, 1126–1128. https://doi.org/10.1107/S0021889810030499 (2010).
van Meerten, S., Franssen, W. & Kentgens, A. ssNake: A cross-platform open-source NMR data processing and fitting application. J. Magn. Reson. 301, 56–66. https://doi.org/10.1016/j.jmr.2019.02.006 (2019).
Kusaka, Y., Hasegawa, T. & Kaji, H. Noise reduction in solid-state NMR spectra using principal component analysis. J. Phys. Chem. A 123, 10333–10338. https://doi.org/10.1021/acs.jpca.9b04437 (2019).
Yamada, S., Kurotani, A., Chikayama, E. & Kikuchi, J. Signal deconvolution and noise factor analysis based on a combination of time-frequency analysis and probabilistic sparse matrix factorization. Int. J. Mol. Sci. 21, 2978. https://doi.org/10.3390/ijms21082978 (2020).
Yamada, S., Chikayama, E. & Kikuchi, J. Signal deconvolution and generative topographic mapping regression for solid-state NMR of multi-component materials. Int. J. Mol. Sci. 22, 1086. https://doi.org/10.3390/ijms22031086 (2021).
Sun, Y. C. & Xin, J. Lorentzian peak sharpening and sparse blind source separation for NMR spectroscopy. SIViP 16, 633–641. https://doi.org/10.1007/s11760-021-02002-4 (2022).
Wang, F., Deng, Z., Yang, Z. & Sun, P. Heterogeneous dynamics and microdomain structure of high-performance chitosan film as revealed by solid-state NMR. J. Phys. Chem. C 125, 13572–13580. https://doi.org/10.1021/acs.jpcc.1c01801 (2021).
Bruce, S. D., Higinbotham, J., Marshall, I. & Beswick, P. H. An analytical derivation of a popular approximation of the Voigt function for quantification of NMR spectra. J. Magn. Reson. 142, 57–63. https://doi.org/10.1006/jmre.1999.1911 (2000).
Besghini, D., Mauri, M. & Simonutti, R. Time domain NMR in polymer science: From the laboratory to the industry. Appl. Sci. 9, 1801. https://doi.org/10.3390/app9091801 (2019).
Rossini, A. J. et al. Dynamic nuclear polarization enhanced NMR spectroscopy for pharmaceutical formulations. J. Am. Chem. Soc. 136, 2324–2334. https://doi.org/10.1021/ja4092038 (2014).
Borsacchi, S. et al. Rubber-filler interactions in polyisoprene filled with in situ generated silica: A solid state NMR study. Polymers 10, 822. https://doi.org/10.3390/polym10080822 (2018).
Schlagnitweit, J. et al. A solid-state NMR method to determine domain sizes in multi-component polymer formulations. J. Magn. Reson. 261, 43–48. https://doi.org/10.1016/j.jmr.2015.09.014 (2015).
Audus, D. & de Pablo, J. Polymer informatics: Opportunities and challenges. ACS Macro Lett. 6, 1078–1082. https://doi.org/10.1021/acsmacrolett.7b00228 (2017).
Verma, R. P. & Hansch, C. Use of 13C NMR chemical shift as QSAR/QSPR descriptor. Chem. Rev. 111, 2865–2899. https://doi.org/10.1021/cr100125d (2011).
Yen, C., Lu, M. & Chen, J. Applying the self-organization feature map (SOM) algorithm to AE-based tool wear monitoring in micro-cutting. Mech. Syst. Signal Process. 34, 353–366. https://doi.org/10.1016/j.ymssp.2012.05.001 (2013).
Ferdousi, R., Jamali, A. & Safdari, R. Identification and ranking of important bio-elements in drug-drug interaction by Market Basket Analysis. Bioimpacts 10, 97–104. https://doi.org/10.34172/bi.2020.12 (2020).
Yamawaki, R., Tei, A., Ito, K. & Kikuchi, J. Decomposition factor analysis based on virtual experiments throughout Bayesian optimization for compost-degradable polymers. Appl. Sci. 11, 2820. https://doi.org/10.3390/app11062820 (2021).
Kumar, J. et al. Machine learning enables polymer cloud-point engineering via inverse design. npj Comput. Mater. 5, 73. https://doi.org/10.1038/s41524-019-0209-9 (2019).
Walden, D. M. et al. Molecular simulation and statistical learning methods toward predicting drug-polymer amorphous solid dispersion miscibility, stability, and formulation design. Molecules 26, 182. https://doi.org/10.3390/molecules26010182 (2021).
Gao, P., Zhang, J., Qiu, H. & Zhao, S. A general QSPR protocol for the prediction of atomic/inter-atomic properties: A fragment based graph convolutional neural network (F-GCN). Phys. Chem. Chem. Phys. 23, 13242–13249. https://doi.org/10.1039/d1cp00677k (2021).
Kurotani, A., Kakiuchi, T. & Kikuchi, J. Solubility prediction from molecular properties and analytical data using an in-phase deep neural network (Ip-DNN). ACS Omega 6, 14278–14287. https://doi.org/10.1021/acsomega.1c01035 (2021).
Takamura, A., Tsukamoto, K., Sakata, K. & Kikuchi, J. Integrative measurement analysis via machine learning descriptor selection for investigating physical properties of biopolymers in hairs. Sci. Rep. 11, 24359. https://doi.org/10.1038/s41598-021-03793-9 (2021).
Yokoyama, D., Suzuki, S., Asakura, T. & Kikuchi, J. Chemometric analysis of NMR spectra and machine learning to investigate membrane fouling. ACS Omega 7, 12654–12660. https://doi.org/10.1021/acsomega.1c06891 (2022).
Venkatram, S. et al. Predicting crystallization tendency of polymers using multifidelity information fusion and machine learning. J. Phys. Chem. B 124, 6046–6054. https://doi.org/10.1021/acs.jpcb.0c01865 (2020).
Fleishman, S. J. et al. RosettaScripts: A scripting language interface to the Rosetta macromolecular modeling suite. PLoS ONE 6, e20161. https://doi.org/10.1371/journal.pone.0020161 (2011).
Spaccini, R., Todisco, D., Drosos, M., Nebbioso, A. & Piccolo, A. Decomposition of bio-degradable plastic polymer in a real on-farm composting process. Chem. Biol. Technol. Agric. 3, 4. https://doi.org/10.1186/s40538-016-0053-9 (2016).
Budrugeac, P. et al. The use of thermal analysis methods for predicting the thermal endurance of an epoxy resin used as electrical insulator. J. Therm. Anal. Calorim. 146, 1791–1801. https://doi.org/10.1007/s10973-020-10156-5 (2021).
Zhong, Y., Godwin, P., Jin, Y. & Xiao, H. Biodegradable polymers and green-based antimicrobial packaging materials: A mini-review. Adv. Ind. Eng. Polym. Res. 3, 27–35. https://doi.org/10.1016/j.aiepr.2019.11.002 (2020).
Post, W., Kuijpers, L., Zijlstra, M., van der Zee, M. & Molenveld, K. Effect of mineral fillers on the mechanical properties of commercially available biodegradable polymers. Polymers 13, 394. https://doi.org/10.3390/polym13030394 (2021).
Oyama, T. et al. Biodegradable compatibilizers for poly(hydroxyalkanoate)/poly(epsilon-caprolactone) blends through click reactions with end-functionalized microbial poly(hydroxyalkanoate)s. ACS Sustain. Chem. Eng. 7, 7969–7978. https://doi.org/10.1021/acssuschemeng.9b00897 (2019).
Helmus, J. J. & Jaroniec, C. P. Nmrglue: An open source Python package for the analysis of multidimensional NMR data. J. Biomol. NMR 55, 355–367. https://doi.org/10.1007/s10858-013-9718-x (2013).
Ito, K., Sakata, K., Date, Y. & Kikuchi, J. Integrated analysis of seaweed components during seasonal fluctuation by data mining across heterogeneous chemical measurements with network visualization. Anal. Chem. 86, 1098–1105. https://doi.org/10.1021/ac402869b (2014).
Wei, F., Sakata, K., Asakura, T., Date, Y. & Kikuchi, J. Systemic homeostasis in metabolome, ionome, and microbiome of wild yellowfin goby in estuarine ecosystem. Sci. Rep. 8, 3478. https://doi.org/10.1038/s41598-018-20120-x (2018).
Shiokawa, Y., Misawa, T., Date, Y. & Kikuchi, J. Application of market basket analysis for the visualization of transaction data based on human lifestyle and spectroscopic measurements. Anal. Chem. 88, 2714–2719. https://doi.org/10.1021/acs.analchem.5b04182 (2016).
Acknowledgements
The authors thank Yuuri Tsuboi, Tomoko Matsumoto, Akiyo Tei (RIKEN) for their support with NMR data acquisition and handling. S.Y. was supported by the RIKEN Junior Research Associate Program and the promotion program for informatics and data science of RIKEN Center for Sustainable Resource Science during the part of a period of this research. This work was partially supported by a Grant from the Agriculture, Forestry, and Fisheries Research Council, as well as Strategic Innovation Program (SIP) from CAO of Japan (to J.K.).
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript. K.H., S.Y., E.C., and J.K. designed this study. Analytical flow and python tools were developed by K.H., S.Y. and A.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hara, K., Yamada, S., Kurotani, A. et al. Materials informatics approach using domain modelling for exploring structure–property relationships of polymers. Sci Rep 12, 10558 (2022). https://doi.org/10.1038/s41598-022-14394-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14394-5
This article is cited by
-
Review of Material Modeling and Digitalization in Industry: Barriers and Perspectives
Integrating Materials and Manufacturing Innovation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.