Abstract
To predict disease outcome in muscle-invasive bladder cancer (MIBC), we constructed a prognostic autophagy-related (PAR) lncRNA signature. Comprehensive bioinformatics analyses were performed using data from TCGA and GTEx databases. Univariate Cox, and least absolute shrinkage and selection operator regression analyses were also performed, based on differentially expressed genes, to identify PAR-related lncRNAs to establish the signature. Furthermore, the Kaplan–Meier OS curve and receiver operating characteristic curve analyses were performed and a nomogram was constructed, all of which together confirmed the strong predictive ability of the constructed signature. Patients with MIBC were then divided into high- and low-risk groups. Gene enrichment and immune infiltration analyses revealed the potential mechanisms in MIBC. We also further evaluated the signature of molecules related to immune checkpoints and the sensitivity toward chemotherapeutic agents and antitumor-targeted drugs to find better treatment prescriptions. We identified a number of PAR-related lncRNA signatures, including HCP5, AC024060.1, NEAT1, AC105942.1, XIST, MAFG-DT, and NR2F1-AS1, which could be valuable prognostic tools to develop more efficient, individualized drug therapies for MIBC patients.
Similar content being viewed by others
Introduction
Urothelial carcinoma, especially bladder cancer (BCa), is a common malignant tumor of the urinary system. Data from the Global Cancer Observatory (GCO, https://gco.iarc.fr/) shows that, 573,278 individuals were diagnosed with BCa and 212,536 individuals died of BCa in 2020, making BCa the fourth most common and tenth most fatal tumor worldwide1. It has also been reported that men are more susceptible to BCa (possibly due to smoking), making BCa the sixth most common carcinoma and ninth most fatal carcinoma in men. Most importantly, approximately 30% of BCa patients were diagnosed with muscular invasive bladder cancer (MIBC)2. Due to the high recurrence rate and poor prognosis of MIBC cases, patients usually experience great physical and mental stress and economic burden, which underscores the need to investigate new prognostic biomarkers in order to improve disease outcomes in patients through early detection and early treatment of high-risk patients.
Autophagy, as its name suggests, is a kind of cell “self-eating” and is prevalent as a self-protection mechanism of eukaryotic cells3. Autophagy leads to cell renewal and maintains homeostasis by degrading damaged organelles and macromolecules4. In cancer, autophagy can both promote and inhibit tumor progression, although the specific mechanism remains unclear.
Long non-coding RNAs (lncRNAs) are a type of ncRNA with transcripts > 200 nucleotides in length but with no protein-coding capacity5. lncRNAs participate in the occurrence and development of cancer through various biological processes, including the cell cycle, cell growth, cell death, drug resistance, and epigenetic regulation5.
Recently, the prevalence of autophagy-related lncRNAs, such as BLACAT16, MALAT17, XIST8, SNHGs9, HULC10, CASC211, and GAS512, was found to be associated with early diagnosis and prognosis of BCa, indicating that autophagy-related lncRNAs may be used as prognostic biomarkers and potential therapeutic targets. Therefore, we used autophagy-related lncRNAs to establish a prognostic autophagy-related (PAR) lncRNA signature for predicting disease outcome in BCa.
Materials and methods
Data acquisition
We acquired the RNA sequencing data of MIBC patients from the Cancer Genome Atlas (TCGA, https://cancergenome.nih.gov) and the Genotype-Tissue Expression Project (GTEx, https://www.genome.gov). Relevant clinical variables, such as age, sex, survival data, grade, and TNM stage, were also obtained from TCGA. We extracted 232 autophagy-related genes from the human autophagy database (HADb, http://autophagy.lu/clustering/index.html).
Establishment and testing of the Risk Score Model
We randomly divided the MIBC patient samples from TCGA into training (n = 184) and testing (n = 180) groups (Table 1). The training cohort was used to establish lncRNA features and the testing cohort was used to verify the model. By applying differential expression analysis, univariate Cox regression analysis, least absolute shrinkage and selection operator (LASSO) regression analysis, Venn analysis, and multivariate Cox regression analysis, we identified seven PAR-lncRNAs which can be viewed as a signature to predict the disease outcomes of patients with MIBC. We established a lncRNA-mRNA co-expression network and Sankey diagram with the seven PAR-lncRNAs and 26 co-expressive mRNAs to visualize the potential relationship of the lncRNAs and mRNA.
Subsequently, we divided the training group into high- and low-risk groups using the median risk score as the cut-off point. Kaplan–Meier survival curve analysis was performed to show the difference in OS between the two groups. We also plotted the time-dependent receiver operating characteristic (ROC) curve and risk nomogram to evaluate the prediction accuracy of the model.
Furthermore, univariate and multivariate Cox analyses were performed to confirm that the seven PAR-lncRNAs signature was an independent prognostic factor for MIBC compared to other clinical characteristics—such as age, sex, clinical stage, TNM, and risk score. We analyzed the correlation between the risk scores and clinical parameters to investigate whether there was any difference in risk scores among different clinical parameter stratifications.
Gene enrichment analysis
Gene ontology (GO) term enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were performed to investigate potential signaling pathways and functions related to the seven lncRNAs included in the model13,14,15.
Immune infiltration analysis and prediction of the sensitivity toward chemotherapeutic agents, antitumor targeted drugs, and immune checkpoint inhibitors
We used the R package “GSEABase” to investigate the difference in the expression and function of 16 infiltrating immune cells between the high- and low-risk groups. We also used the R package “pRRophetic” to analyze drug sensitivity in the high- and low-risk groups. In addition, we used the R package “ggpubr” to explore the sensitivity toward immune checkpoint inhibitors (ICIs) in the high- and low-risk groups.
Statistical analysis
Cytoscape software (version 3.9.0) was used to construct an autophagy-related lncRNA co-expression network. R software (version x64 4.0.5) was used for statistical analysis, including Cox regression analysis, ROC curve analysis, gene enrichment analysis, and immune infiltration analysis.
Results
Data acquisition
The RNA sequencing and clinical data from the TCGA and GTEx databases included 375 MIBC patients and 28 para-cancerous tissues (19 from TCGA and 9 from GTEx). Differential expression analysis (|log2FC|> 2) revealed 175 differentially expressed lncRNAs in tumors compared with the corresponding expression in para-cancerous tissues.
We also extracted 232 autophagy-related genes from the human autophagy database (HADb, http://autophagy.lu/clustering/index.html), of which 112 were identified by the “limma” R package and Pearson correlation analysis.
Risk Score Model
The MIBC samples from TCGA were randomly assigned to a training group (n = 184) and a testing group (n = 180). The training cohort was used to establish lncRNA features, and the testing cohort was used to verify the model. We then selected 10 prognosis-related lncRNAs from the training group for univariate Cox regression analysis (Fig. 1A). The least absolute shrinkage and selection operator (LASSO) regression analysis was then applied to further screen the 10 lncRNAs (Fig. 1B,C). Furthermore, we applied a Venn analysis to screen for nine possible PAR-lncRNAs (Fig. 1D). Multivariate Cox regression analysis showed that seven of the nine lncRNAs were suitable for the construction of a prognostic signature (Fig. 1E). Among the seven PAR-related lncRNAs included in the prognostic signature, HCP5 and AC024060.1 were considered protective factors (HR values less than 1), and NEAT1, AC105942.1, XIST, MAFG-DT, and NR2F1-AS1 were considered risk factors (HR values greater than 1).
Selection of prognostic autophagy-related lncRNAs(PAR lncRNAs) with prognostic value. (A) Risk ratio forest plot shows that 10 autophagy-related lncRNAs, NR2F1-AS1, MALAT1, AC105942.1, MAFG-DT, HCP5, NEAT1, AC005261.1, U62317.2, AC024060.1, XIST, were significantly related to OS of MIBC patients from training group. (B) Adjusted parameters of LASSO regression model. (C) Figure for LASSO coefficient spectrum of prognostic lncRNAs. (D)The Venn analysis between DRGs and Prognostic genes. DRGs, differentially expressed genes. (E) Forest plot of multivariate Cox regression analysis shows the prognostic value of the 7 prognostic autophagy-related lncRNAs(PAR lncRNAs).
In addition, we used the “survival” and “survminer” R packages to calculate the correlation coefficients. The risk score for the prognostic lncRNA model was calculated using the following formula:
The expression and coefficients of each lncRNA in the signature are shown in Supplementary Table 1.
lncRNA-mRNA co-expression network
Once the seven PAR-lncRNAs were identified, we established a lncRNA-mRNA co-expression network to investigate the potential functions of the seven PAR-lncRNAs. The network was visualized using Cytoscape (Fig. 2A). The graph consisted of seven PAR-lncRNAs and 26 mRNAs (|coefficient value|> 0.3 and P-value < 0.001). Moreover, we drew a Sankey diagram to further display the risk/protective relationship between the seven PAR-lncRNAs and the 26 mRNAs (Fig. 2B).
The establishment of lncRNA-mRNA co-expression network. (A) The PAR lncRNAs-mRNA co-expression network contains 7 PAR lncRNAs and 26 mRNAs with | coefficient value |> 0.3 and P value < 0.001; (B) the Sankey diagram shows the connection degree between the 7 PAR lncRNAs and 26 mRNAs (risk/protective). PAR lncRNAs, prognostic autophagy-related lncRNAs.
Testing of the Risk Score Model and independent prognostic analysis.
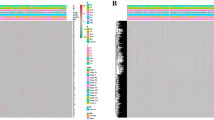
Patients in the training group were divided into high- and low-risk groups using the median risk score as the cut-off point. The results of the Kaplan–Meier survival curve analysis showed that the OS in the high-risk group was much lower than that in the low-risk group (Fig. 3A,B). For the training group, the expression heatmap of these seven PAR-related lncRNAs, the risk score of the signature in the low- and high-risk groups, and the survival time of MIBC patients are displayed in Fig. 3C,E,G. The same assessments were applied to the test group (Fig. 3D,F,H). We also drew a time-dependent ROC curve for the patients in the two groups (Fig. 3I,J). The area under the curve (AUC) values were used to evaluate the prediction accuracy of the model. In addition, a risk nomogram was established to further determine the predictive ability of the model including seven-PAR-lncRNA signature and univariate significant features, such as gender, age, T, M, and risk score (Fig. 4A). Univariate and multivariate Cox analyses showed that the seven-PAR-lncRNA signature may be a reliable predictor of OS in patients with MIBC. Univariate Cox analysis revealed that clinical characteristics—such as age, sex, clinical stage, TNM, and risk score—were related to OS (Fig. 4B). Multivariate Cox analysis of these clinical characteristics showed that the seven-PAR-lncRNA signature was an independent prognostic factor for MIBC (P < 0.05; Fig. 4C). The ROC curve showed that the seven-PAR-lncRNA signature is an excellent predictive indicator of prognosis compared to any other clinical characteristics, with an AUC value of 0.766 (Fig. 4D).
(A) The overall survival rate of the low-risk group was better than the high-risk group in the training group (P < 0.01); (B) the overall survival rate of the low-risk group was better than the high-risk group in the test group (P < 0.01); (C) the heatmap showed the differential expression of 7 PAR lncRNAs between the high- and low-risk groups in the training group; (D) the heatmap showed the differential expression of 7 PAR lncRNAs between the high- and low-risk groups in the test group; (E) the distribution of the risk score for every MIBC patient in the training group; (F) the distribution of the risk score for every MIBC patient in the test group; (G) the survival status of every MIBC patient with different risk scores in the training group; (H) the survival status of every MIBC patient with different risk scores in the test group; (I) the AUC of 1-year OS was 0.747 in the training group; (J) the AUC of 1-year OS was 0.674 in the test group; PAR lncRNAs, prognostic autophagy-related lncRNAs; MIBC, muscle-invasive bladder cancer; AUC, Area Under Curve; OS, Overall Survival.
(A) Nomogram for predicting the 1- 3- and 5-year survival rates of MIBC patients; (B) Univariate Cox analysis reveals that clinical characteristics, like age, gender, clinical stage, TNM and risk score, were related to OS; (C) Multivariate Cox analysis of these clinical characteristics shows that the 7-PAR lncRNAs signature is an independent prognostic factor for MIBC; (D) The 7-PAR lncRNAs signature is an excellent predictive indicator of prognosis than any other clinical characteristic, with an area under the curve (AUC) of 0.766.
Gene enrichment analysis
To investigate the potential signaling pathways and functions related to the seven genes included in the model, we performed GO enrichment analysis (Fig. 5A,B) and KEGG pathway analysis (Fig. 5C,D). The GO enrichment analysis consists of three terms: biological process (BP), cellular component (CC), and molecular function (MF). In terms of BP, the seven ARGs with prognostic features were highly associated with muscle system processes, muscle contraction, extracellular matrix organization, and extracellular structure organization. In terms of CC, these genes were related to the collagen-containing extracellular matrix, contractile fiber, myofibril, and sarcomere. Furthermore, the major functions of these genes were revealed to be actin binding, extracellular matrix structural constituent, glycosaminoglycan binding, and sulfur compound binding. KEGG analysis showed that the seven PAR-related lncRNAs were mostly enriched for cell adhesion molecules, followed by dilated cardiomyopathy, hypertrophic cardiomyopathy, and vascular smooth muscle contraction.
Gene enrichment analysis (A,B) Gene Ontology (GO) analysis results show the enriched biological processes, cell components and molecular functions associated with the mRNAs that co-express with the 7 prognostic autophagy-related lncRNAs. (C,D) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis results shows the enriched signaling pathways associated with the mRNAs that co-express with the 7 prognostic autophagy-related lncRNAs.
Clinical correlation analysis
We then analyzed the correlation between the risk scores and other clinical parameters in the case of the training group (having exact clinicopathological characteristics). The results showed that patients aged > 70 years were at a higher risk than those aged ≤ 70 years. A similar result was observed between females and males. The risk score had no statistical differences between stages T, N, and M (Fig. 6).
Correlation analyses of the prognostic autophagy-related lncRNAs signature with various clinicopathological characteristics of the MIBC patients. (A) patients aged > 70 have higher risk than those aged ≤ 70; (B) female have higher risk than male; (C) risk score has no statistical differences among stage II, III and IV; (D) risk score has no statistical differences among T2, T3 and T4; (E) risk score has no statistical differences among N0, N1, N2; (F) risk score has no statistical differences between M0 and M1.
Immune infiltration analysis and prediction of the sensitivity toward chemotherapeutic agents, antitumor targeted drugs, and immune checkpoint inhibitors
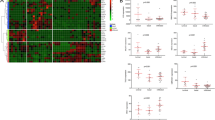
We used the R package “GSEABase” to investigate the difference in the expression and function of 16 infiltrating immune cells between the high- and low-risk groups. We found statistically significant differences in the expression of CD8+ T cells, mast cells, and Th2 cells (Fig. 7A). Furthermore, our findings also showed statistical differences in cytolytic activity, HLA, inflammation-promoting, MHC class I, and type I IFN Response (Fig. 7B).
Immune infiltration analysis and prediction of the sensitivity of chemotherapeutic agents. (A) The expression of infiltrating immune cells has statistical difference in CD8+ T cells, Mast cells, Th2 cells between the high- and low-risk group; (B) The function of infiltrating immune cells has statistical difference in Cytolytic activity, HLA, Inflammation-promoting, MHC class I, Type I IFN Reponse between the high- and low-risk group; (C,D) In terms of drug sensitivity to cisplatin and gemcitabine, there is no statistical difference between high-risk group and low-risk group; (E–G) The sensitivity to Doxorubicin, Methotrexate, Vinblastine are much higher in high-risk group than in low-risk group. (H–J) The sensitivity to Sunitinib, Sorafenib, and Axitinib has no statistical difference between the high- and low- risk groups; (K) The sensitivity to Pazopanib is much higher in high-risk group than in low-risk group; (L) The expression level of PDCD1(PD-1) was remarkably higher in the low-risk group with P < 0.05; (M) The expression level of LGALS9 (GAL9)were remarkably higher in the low-risk group with P < 0.01; (N–T) The expression levels of PDCD1LG2 (PD-1LG2), TIGHT, LAG-3, CD274 (PD-L1), CTLA-4, HAVCR2 (TIM-3) and TNFRSF9 had no statistical difference between the high- and low- risk groups with the P > 0.05.
To identify a more effective approach to treating MIBC, we analyzed the sensitivity of the high- and low-risk groups toward chemotherapeutic agents using the R package “pRRophetic”. From the boxplots, we found that there was no difference in sensitivity toward gemcitabine and cisplatin between the two groups (Fig. 7C,D), whereas higher sensitivity was observed for doxorubicin, methotrexate, and vinblastine in the high-risk group than in the low-risk group (Fig. 7E–G). This indicates that patients in the high-risk group may be more likely to gain better efficacy with the MAVC drug regimen (consisting of cisplatin, doxorubicin, methotrexate, and vinblastine) regimen rather than with the GC regimen (gemcitabine and cisplatin).
Antitumor drugs, such as sorafenib and sunitinib, have been shown to have an excellent effect on renal cell carcinoma. However, in BCa, the effect of these antitumor-targeted drugs, excluding erdafitinib, is still unclear. Accordingly, we investigated the sensitivity of these cells to common antitumor-targeted drugs. The data from these studies showed that there was no statistically significant difference between the high- and low-risk groups in terms of their sensitivity to sunitinib, sorafenib, and axitinib (Fig. 7H–J) and that the sensitivity to pazopanib was much higher in the high-risk than in the low-risk group (Fig. 7K).
We also compared the high- and low-risk groups for differences in the sensitivity toward immune checkpoint inhibitors (ICIs), such as pembrolizumab and atezolizumab, which have been reported to show excellent durable responses and disease control in patients with BCa16,17. We utilized the "limma" and "gpubr" packages to generate violin plots. As can be seen in Fig. 7M, the expression levels of PDCD1(PD-1) (P < 0.05, Fig. 7L) and LGALS9 (GAL9) (P < 0.01) were remarkably higher in the low-risk group. The expression levels of PDCD1LG2 (PD-1LG2), TIGHT, LAG-3, CD274 (PD-L1), CTLA-4, HAVCR2 (TIM-3), and TNFRSF9 were not significantly different between the high- and low-risk groups (P > 0.05, Fig. 7N–T).
Altogether, these results seem to indicate that there is significant potential for improving therapeutic regimens for MIBC.
Discussion
BCa is one of the most common tumors of the urinary system, and data from the Global Cancer Observatory shows that the number of new BCa patients in 2020 is ranked fourth among that of all malignancies1. Among all patients diagnosed with BCa, approximately 30% have MIBC2, which has a poor prognosis. Although various therapeutic regimens ameliorate the effects of the disease, there is room for improvement in terms of categorizing patients for more individualized treatment and a better prognosis. That is why we carry on this study. By this signature, patients will be divided into high- and low-risk groups, and then they can get individualized treatment plans, considering the result of drug sensitivity, so that they can gain a better prognosis.
Recently, researchers have found that autophagy and lncRNAs play important roles in the occurrence and development of cancer18,19,20,21,22,23. In the present study, we established a seven-PAR-lncRNA signature, based on the data of MIBC patients from TCGA, to predict the prognosis of MIBC patients. We evaluated the prediction accuracy of the signature using Kaplan–Meier survival curve analysis, time-dependent ROC curve, risk nomogram, univariate and multivariate Cox analyses, and other analyses. In examining the drug sensitivity, we found that patients in the high-risk group may be more likely to benefit from the MAVC regimen than by the GC regimen, which suggests that MAVC should be given higher priority over GC when choosing a chemotherapy regimen for high-risk patients.
In our study, HCP5 and AC024060.1 were considered protective factors, with HR values less than 1. NEAT1, AC105942.1, XIST, MAFG-DT, and NR2F1-AS1 were considered as risk factors with HR values greater than 1. Therefore, we searched the published literature for these genes.
HCP5 can negatively regulate the expression of miR-29b-3p to promote BCa cell viability, migration, and invasion24. HCP5 is a part of an immune-related lncRNA signature, which is related to the prognosis of BCa25. AC024060.1 could be seen in the other two prognosis-related signatures of BCa; however, research on the specific mechanism is still lacking26,27.
NEAT1, located on chromosome 11, is a structural component of nuclear paraspeckles, which plays a significant oncogenic role in proliferation and cell migration28. It has been shown that the overexpression of NEAT1 is associated with poor prognosis of many cancers29. Shan et al. reported that NEAT1 promotes bladder progression through the NEAT1/miR-410/HMGB1 axis30. NEAT1 has also been shown to play an important role in the treatment of BCa. Zhao et al. found that the suppression of cell growth, invasion, and apoptosis of BCa cells under cisplatin chemotherapy could be enhanced by silencing NEAT131. Similarly, by negatively regulating miR-214-3p expression, NEAT1 promotes doxorubicin resistance in BCa via the Wnt/β-catenin pathway32.
XIST, located in the X chromosome inactivation center of Xq13.233, has been shown to play a wide range of roles in the occurrence and progression of therioma34,35. The results of previous studies have indicated that XIST can participate in the occurrence, progression, and metastasis of therioma via the XIST-TET1-p53 regulatory network36, miR-133a37, XIST/miR-200c38, miR-124 dependent androgen receptor regulation39, and the miR-139-5p mediated Wnt/β-catenin signaling pathway40.
In another two prognosis-related signatures of BCa, AC105942.1 was also revealed to be a part of the signatures, but the specific mechanism of action of AC105942.1 in cancer is still unclear41,42. A similar conclusion was also found for MAFG-DT42,43,44 and NR2F1-AS145,46.
In recent years, the treatment of BCa has become increasingly diversified, including surgery, chemotherapy, immunotherapy, and targeted therapy. Here, we explored alternative treatments for MIBC that could be more effective than the existing ones.
Chemotherapy plays an important role in MIBC treatment. GC and MVAC are the most commonly used chemotherapeutic regimens. Through sensitivity tests of these drugs, we found that the high-risk MIBC patients had a much higher sensitivity toward MVAC than that toward GC, which indicates that patients in the high-risk group may be more likely to benefit from the MAVC regimen rather than the GC regimen. Similar tests were performed for antitumor-targeted drugs for urothelial carcinoma and immune checkpoint inhibitors. We hope that these results will help clinicians choose more effective, treatment options for individual patients.
However, our study has some limitations. First, external validation of the signature in large-scale multicenter cohorts is necessary before the signature is widely adopted, owing to the limitations of the data source and sample size of our study. In addition, although we determined that these seven lncRNAs play an important role in BCa, the precise mechanism of action remains unclear. Further functional experiments are required to elucidate the mechanisms of these genes. In summary, further studies are needed to elucidate the mechanisms and effects of MIBC.
Conclusion
Utilizing MIBC patient data obtained from TCGA, we established a seven-PAR-lncRNA signature to predict the disease outcome of patients with MIBC. Furthermore, we explored alternative pharmaceutical treatments that could be individualized for MIBC patients based on disease risk. Our results indicate that patients in the high-risk group may benefit more from the MAVC drug regimen than the GC regimen, pazopanib may be more effective in high-risk MIBC patients, and drugs acting on PD-1 and LGALS9 gene loci may be more effective in low-risk patients.
Data availability
All the data in this study can be found in The Cancer Genome Atlas(TCGA, https: //cancergenome.nih.gov)and Genotype-Tissue Expression Project (GTEx, https://www.genome.gov).
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249. https://doi.org/10.3322/caac.21660 (2021).
Sun, M. & Trinh, Q. D. Diagnosis and staging of bladder cancer. Hematol. Oncol. Clin. North Am. 29(2), 205–vii. https://doi.org/10.1016/j.hoc.2014.10.013
Masclaux-Daubresse, C., d’Andrea, S., Bouchez, I. & Cacas, J. L. Reserve lipids and plant autophagy. J. Exp. Bot. 71(10), 2854–2861. https://doi.org/10.1093/jxb/eraa082 (2020).
Kimmelman, A. C. & White, E. Autophagy and tumor metabolism. Cell Metab. 25(5), 1037–1043. https://doi.org/10.1016/j.cmet.2017.04.004 (2017).
Evans, J. R., Feng, F. Y. & Chinnaiyan, A. M. The bright side of dark matter: lncRNAs in cancer. J. Clin. Investig. 126(8), 2775–2782. https://doi.org/10.1172/JCI84421 (2016).
He, W. et al. linc-UBC1 physically associates with polycomb repressive complex 2 (PRC2) and acts as a negative prognostic factor for lymph node metastasis and survival in bladder cancer. Biochem. Biophys. Acta. 1832(10), 1528–1537. https://doi.org/10.1016/j.bbadis.2013.05.010 (2013).
Cao, Y. et al. Long non-coding RNA in bladder cancer. Clin. Chim. Acta 503, 113–121. https://doi.org/10.1016/j.cca.2020.01.008 (2020).
Su, G., He, Q. & Wang, J. Clinical values of long non-coding RNAs in bladder cancer: A systematic review. Front. Physiol. 9, 652. https://doi.org/10.3389/fphys.2018.00652 (2018).
Yang, Z., Li, Q., Zheng, X. & Xie, L. Long noncoding RNA small nucleolar host gene: A potential therapeutic target in urological cancers. Front. Oncol. 11, 638721. https://doi.org/10.3389/fonc.2021.638721 (2021).
Wang, J., Ma, W. & Liu, Y. Long non-coding RNA HULC promotes bladder cancer cells proliferation but inhibits apoptosis via regulation of ZIC2 and PI3K/AKT signaling pathway. Cancer Biomark 20(4), 425–434. https://doi.org/10.3233/CBM-170188 (2017).
Pei, Z. et al. Down-regulation of lncRNA CASC2 promotes cell proliferation and metastasis of bladder cancer by activation of the Wnt/β-catenin signaling pathway. Oncotarget 8(11), 18145–18153. https://doi.org/10.18632/oncotarget.15210 (2017).
Liu, Z. et al. Downregulation of GAS5 promotes bladder cancer cell proliferation, partly by regulating CDK6. PLoS ONE 8(9), e73991. https://doi.org/10.1371/journal.pone.0073991 (2013).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 49(D1), D545–D551. https://doi.org/10.1093/nar/gkaa970 (2021).
Necchi, A. et al. Updated results of PURE-01 with preliminary activity of neoadjuvant Pembrolizumab in patients with muscle-invasive bladder carcinoma with variant histologies. Eur. Urol. 77(4), 439–446. https://doi.org/10.1016/j.eururo.2019.10.026 (2020).
Marcq, G. et al. Phase 1 trial of Atezolizumab Plus trimodal therapy in patients with localized muscle-invasive bladder cancer. Int. J. Radiat. Oncol. Biol. Phys. 110(3), 738–741. https://doi.org/10.1016/j.ijrobp.2020.12.033 (2021).
Li, X., He, S. & Ma, B. Autophagy and autophagy-related proteins in cancer. Mol. Cancer 19(1), 12. https://doi.org/10.1186/s12943-020-1138-4 (2020).
Levy, J., Towers, C. G. & Thorburn, A. Targeting autophagy in cancer. Nat. Rev. Cancer 17(9), 528–542. https://doi.org/10.1038/nrc.2017.53 (2017).
Amaravadi, R. K., Kimmelman, A. C. & Debnath, J. Targeting autophagy in cancer: Recent advances and future directions. Cancer Discov. 9(9), 1167–1181. https://doi.org/10.1158/2159-8290.CD-19-0292 (2019).
Statello, L., Guo, C. J., Chen, L. L. & Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 22(2), 96–118. https://doi.org/10.1038/s41580-020-00315-9 (2021).
McCabe, E. M. & Rasmussen, T. P. lncRNA involvement in cancer stem cell function and epithelial-mesenchymal transitions. Semin. Cancer Biol. 75, 38–48. https://doi.org/10.1016/j.semcancer.2020.12.012 (2021).
Saw, P. E., Xu, X., Chen, J. & Song, E. W. Non-coding RNAs: The new central dogma of cancer biology. Science China. Life sciences 64(1), 22–50. https://doi.org/10.1007/s11427-020-1700-9 (2021).
Zhao, C. et al. LncRNA HCP5 promotes cell invasion and migration by sponging miR-29b-3p in human bladder cancer. OncoTargets Ther. 13, 11827–11838. https://doi.org/10.2147/OTT.S249770 (2020).
Zhang, L. et al. Identification of immune-related lncRNA signature to predict prognosis and immunotherapeutic efficiency in bladder cancer. Front. Oncol. 10, 542140. https://doi.org/10.3389/fonc.2020.542140 (2021).
Wang, J. et al. Identification and verification of an immune-related lncRNA signature for predicting the prognosis of patients with bladder cancer. Int. Immunopharmacol. 90, 107146. https://doi.org/10.1016/j.intimp.2020.107146 (2021).
Wan, J. et al. Autophagy-related long non-coding RNA is a prognostic indicator for bladder cancer. Front. Oncol. 11, 647236. https://doi.org/10.3389/fonc.2021.647236 (2021).
Taheri, M., Omrani, M. D. & Ghafouri-Fard, S. Long non-coding RNA expression in bladder cancer. Biophys. Rev. 10(4), 1205–1213. https://doi.org/10.1007/s12551-017-0379-y (2018).
Yang, C. et al. Long non-coding RNA NEAT1 overexpression is associated with poor prognosis in cancer patients: A systematic review and meta-analysis. Oncotarget 8(2), 2672–2680. https://doi.org/10.18632/oncotarget.13737 (2017).
Shan, G., Tang, T., Xia, Y. & Qian, H. J. Long non-coding RNA NEAT1 promotes bladder progression through regulating miR-410 mediated HMGB1. Biomed. Pharmacother. 2020(121), 109248. https://doi.org/10.1016/j.biopha.2019.109248 (2020).
Zhao, W. et al. Silencing long non-coding RNA NEAT1 enhances the suppression of cell growth, invasion, and apoptosis of bladder cancer cells under cisplatin chemotherapy. Int. J. Clin. Exp. Pathol. 12(2), 549–558 (2019).
Guo, Y. et al. Non-coding RNA NEAT1/miR-214-3p contribute to doxorubicin resistance of urothelial bladder cancer preliminary through the Wnt/β-catenin pathway. Cancer Manag. Res. 10, 4371–4380. https://doi.org/10.2147/CMAR.S171126 (2018).
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71(3), 527–542. https://doi.org/10.1016/0092-8674(92)90520-m (1992).
Liu, X. et al. Long non-coding RNA XIST predicts worse prognosis in digestive system tumors: a systemic review and meta-analysis. Biosci. Rep. 38(3), BSR20180169. https://doi.org/10.1042/BSR20180169 (2018).
Wang, D. et al. Abnormal X chromosome inactivation and tumor development. Cell Mol. Life Sci. 77(15), 2949–2958. https://doi.org/10.1007/s00018-020-03469-z (2020).
Hu, B., Shi, G., Li, Q., Li, W. & Zhou, H. Long noncoding RNA XIST participates in bladder cancer by downregulating p53 via binding to TET1. J. Cell. Biochem. 120(4), 6330–6338. https://doi.org/10.1002/jcb.27920 (2019).
Zhou, K., Yang, J., Li, X. & Chen, W. Long non-coding RNA XIST promotes cell proliferation and migration through targeting miR-133a in bladder cancer. Exp. Ther. Med. 18(5), 3475–3483. https://doi.org/10.3892/etm.2019.7960 (2019).
Xu, R. et al. LncRNA XIST/miR-200c regulates the stemness properties and tumourigenicity of human bladder cancer stem cell-like cells. Cancer Cell Int. 18, 41. https://doi.org/10.1186/s12935-018-0540-0 (2018).
Xiong, Y. et al. The long non-coding RNA XIST interacted with MiR-124 to modulate bladder cancer growth, invasion and migration by targeting androgen receptor (AR). Cell Physiol. Biochem. 43(1), 405–418. https://doi.org/10.1159/000480419 (2017).
Hu, Y. et al. Long non-coding RNA XIST promotes cell growth and metastasis through regulating miR-139-5p mediated Wnt/β-catenin signaling pathway in bladder cancer. Oncotarget 8(55), 94554–94568. https://doi.org/10.18632/oncotarget.21791 (2017).
Tong, H. et al. An epithelial-mesenchymal transition-related long noncoding RNA signature correlates with the prognosis and progression in patients with bladder cancer. Biosci. Rep. 41(1), BSR20203944. https://doi.org/10.1042/BSR20203944 (2021).
Zheng, Z. et al. Identification of a novel glycolysis-related LncRNA signature for predicting overall survival in patients with bladder cancer. Front. Genet. 12, 720421. https://doi.org/10.3389/fgene.2021.720421 (2021).
Cui, Y., Zhou, Z., Chai, Y., Che, X. & Zhang, Y. Identification of a nomogram from ferroptosis-related long noncoding RNAs signature to analyze overall survival in patients with bladder cancer. J. Oncol. 2021, 8533464. https://doi.org/10.1155/2021/8533464 (2021).
Zhou, M. et al. Computational recognition of lncRNA signature of tumor-infiltrating B lymphocytes with potential implications in prognosis and immunotherapy of bladder cancer. Brief. Bioinform. 22(3), bbaa047. https://doi.org/10.1093/bib/bbaa047 (2021).
Peng, M., Cheng, X., Xiong, W., Yi, L. & Wang, Y. Integrated analysis of a competing endogenous RNA network reveals a prognostic lncRNA signature in bladder cancer. Front. Oncol. 11, 684242. https://doi.org/10.3389/fonc.2021.684242 (2021).
Zhong, W. et al. Integrative analysis of prognostic long non-coding RNAs with copy number variation in bladder cancer. J. Zhejiang Univ. Sci. B 22(8), 664–681. https://doi.org/10.1631/jzus.B2000494 (2021).
Acknowledgements
Great thanks to the assistance of all the people who contributed in this study. We particularly thank Dr. Shengjie Yu for the discussion and critical reading of the manuscript and fund support.
Funding
This study was supported by the Chongqing Science and Technology Benefiting project [cstc2016kjhmpt1001-5].
Author information
Authors and Affiliations
Contributions
W.T. and Y.Y. conceived of the whole study and participated in design. Wei Tan and Hao Huang drafted the manuscript. W.T., J.M., Y.L., Y.G. assisted in bioinformatics analysis of the data. Z.H. and H.W. revised the manuscript, and they contributed equally. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tan, W., Yuan, Y., Huang, H. et al. Comprehensive analysis of autophagy related long non-coding RNAs in prognosis, immunity, and treatment of muscular invasive bladder cancer. Sci Rep 12, 11242 (2022). https://doi.org/10.1038/s41598-022-13952-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13952-1
This article is cited by
-
Carcinogenic roles of MAFG-AS1 in human cancers
Clinical and Translational Oncology (2023)
-
Identification of necroptosis-related long non-coding RNAs prognostic signature and the crucial lncRNA in bladder cancer
Journal of Cancer Research and Clinical Oncology (2023)
-
Roles of non-coding RNAs in the metabolism and pathogenesis of bladder cancer
Human Cell (2023)
-
Potential roles of lncRNA-XIST/miRNAs/mRNAs in human cancer cells
Clinical and Translational Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.