Abstract
Archaeological and genomic evidence suggest that modern Homo sapiens have roamed the planet for some 300–500 thousand years. In contrast, global human mitochondrial (mtDNA) diversity coalesces to one African female ancestor (“Mitochondrial Eve”) some 145 thousand years ago, owing to the ¼ gene pool size of our matrilineally inherited haploid genome. Therefore, most of human prehistory was spent in Africa where early ancestors of Southern African Khoisan and Central African rainforest hunter-gatherers (RFHGs) segregated into smaller groups. Their subdivisions followed climatic oscillations, new modes of subsistence, local adaptations, and cultural-linguistic differences, all prior to their exodus out of Africa. Seven African mtDNA haplogroups (L0–L6) traditionally captured this ancient structure—these L haplogroups have formed the backbone of the mtDNA tree for nearly two decades. Here we describe L7, an eighth haplogroup that we estimate to be ~ 100 thousand years old and which has been previously misclassified in the literature. In addition, L7 has a phylogenetic sublineage L7a*, the oldest singleton branch in the human mtDNA tree (~ 80 thousand years). We found that L7 and its sister group L5 are both low-frequency relics centered around East Africa, but in different populations (L7: Sandawe; L5: Mbuti). Although three small subclades of African foragers hint at the population origins of L5'7, the majority of subclades are divided into Afro-Asiatic and eastern Bantu groups, indicative of more recent admixture. A regular re-estimation of the entire mtDNA haplotype tree is needed to ensure correct cladistic placement of new samples in the future.
Similar content being viewed by others
Introduction
Descent with modification1 is perhaps the greatest insight in all biology. By inferring common ancestry between orthologous DNA sequences, the field of molecular phylogenetics has revolutionized how we design cancer therapies2, trace infectious diseases3, unlock the secrets of aging4, and ultimately study human history5. Mitochondrial DNA (mtDNA) is the oldest and most ubiquitous genetic tool for revealing the shared origins and migration patterns of humankind6,7,8. Its uniparental inheritance and unique properties make it ideal for reconstructing the human matrilineal phylogeny: abundant cytoplasmic copies make it easy to sequence, rapidly accumulating mutations can resolve recent events, and non-recombining transmission allows for ancient relationships to be finely estimated.
Since the original Cambridge Reference Sequence was published 40 years ago9, the field of human mtDNA phylogenetics has increasingly shown that non-African lineages are a subset, or small clade nested within a much older and hyper-diverse African phylogeny6,8,10,11. That same pattern of ancient African origins and subsequent Out-of-Africa (OOA) expansion some 70–50 thousand years ago (kya) has since been corroborated by Y-DNA12,13, autosomal DNA14,15,16, patterns of archaic introgression17,18,19, and African fossils20,21,22. Although the uniparental mtDNA and Y-DNA loci have a coalescence time of 250–150 kya, archaeological, nuclear, and genome-wide datasets suggest the possibility of a much older (500–300 kya) origin for modern humans23,24,25. The true age may rest somewhere among those two ranges.
Genome-wide coalescent analyses suggest that ancient populations began to take structure 200 kya, which led to a rift between Khoisan and non-hunter-gatherer groups (i.e., Niger-Congo, Nilo-Saharan, Afro-Asiatic) by 160 kya, followed shortly by a split between Khoisan and RFHG groups 120–100 kya26,27. Mitochondrial studies have reinforced this pattern of Stone Age divergences and subsequent admixture amongst rainforest28, and Khoisan29,30 hunter-gatherer groups. Much of the past 200 thousand years of human evolution has therefore been a story of population structuring and diffusion within the continent of Africa.
Our prevailing understanding of human matrilineal ancestry for nearly two decades has defined seven major African lineages or “L haplogroups.” Macrohaplogroup L is a paraphyletic group that contains all modern humans in haplogroups L0–L631, except for the OOA clades M and N. We consider an L haplogroup to be any African lineage (pre-OOA) that is not nested within an existing named L haplogroup. The oldest lineage “L1” (sensu auct.) was initially rooted with an Asian sequence10, and was later reclassified due to paraphyly into L0, L1, and L532,33,34. Subclades of these three oldest haplogroups reach peak frequencies in human populations thought to be outgroups to the rest of modern humans (L0d and L0k in Khoisan; L1c in Baka; L0a2b in Mbuti; L5 in Mbuti, Tshwa, Sandawe). The most widespread haplogroup in sub-Saharan Africans is L2, likely owing to the Bantu expansion, but also to a 70–50 kya climatic oscillation that drove a mass migration35. The remaining lineages fall into L3'4'6; i.e., L3 and its two low-frequency outgroups L4 and L636. These haplogroups have an East African center of diversity, and L3 contains the only two OOA clades M and N.
We recently discovered several sequences in the private FamilyTreeDNA (FTDNA) database of full mtDNA sequences (mtFull; > 200,000 records) that cannot be classified as descendants of any of the previously described L haplogroups (see “Materials and methods”). These sequences ostensibly formed a sister clade to the ancient L5, which would be the eighth L haplogroup. This novel L7 lineage would be a major discovery, because it would be the oldest mitochondrial lineage described in two decades. Moreover, if other L7 sequences have already been mistaken as L5, this would underscore the urgency of calls to update the mtDNA tree37,38. The de facto resource for defining the human mtDNA topological structure is PhyloTree31, however it has not been updated since 2016, and its phylogenetic reconstruction methods have always been unclear. If new sequences or subclades are simply grafted onto an existing structure, then the space of possible trees is not explored, and the new context of synapomorphies is not fully considered. With just n = 50 sequences, there are (2n – 3)!! bifurcating rooted trees, or approximately the number of atoms in the known universe (~ 1080). Given that Phylotree v17 has > 5,000 haplogroups with many homoplastic variants, there is the possibility that a lineage as significant as L7 could go unnoticed.
In this study, we had three primary goals: (1) define the structure of L7 and its subclades; (2) estimate the timing of their origin; and (3) infer any likely population origins or migration events that led to its current distribution. Our work will shed light on the deep coalescent structure of humankind in the cradle of Africa and offer a new perspective on the most ubiquitous tool in human phylogenetics.
Materials and methods
Sample selection
In order to estimate the placement and age of the putative L7 clade, we selected complete mtDNA genomes to represent the backbone of the haplotype tree. These include the major subclades for L0 (L0a1, L0a2, L0d1, L0d2, L0d3, L0f., L0k), L1 (L1b, L1c1, L1c2, L1c3, L1c4), L2 (L2a, L2b, L2c, L2d, L2e), L6, L4, and L3. Inclusion of backbone sequences is important to give phylogenetic context and polarity of mutations while reconstructing L5, L7, and their relationships. To reconstruct each of these backbone branches, we selected pairs of sequences (Supplementary Table S1) representing the maximum topological distance according to the latest version of PhyloTree (v1739), and used HaploGrep240 to infer haplogroups. We verified that publicly chosen samples did not influence the structure of L5, L7, or other parts of the tree as a consequence of any private or artifact mutations by re-running the analyses several times with multiple backbone sample sets. For L5'7, we searched extensively for any complete sequences with L5 or likely L7 mutations that could resolve this portion of the tree; we found a total of 49 L5 and 18 L7 unique sequences (Supplementary Table S2). Data sources included FTDNA private collections, Genographic Project participants opting into scientific research, the 1000 Genomes Project41, and the academic literature28,30,42,43,44,45,46,47,48,49,50,51,52.
Next-generation DNA sequencing
Private collections were sequenced using the mtFull Sequence procedure in the Gene by Gene, Ltd. lab (Houston, TX). Briefly, tissue from cheek swabs were extracted using a magnetic bead procedure. Samples were LR-PCR amplified using a KAPA HiFi Hotstart PCR kit (Roche Sequencing) with the profile: 95 °C for 5 min, [98 °C for 20 s, 65 °C for 15 s, 72 °C for 3 min] × 35, 72 °C for 5 min, 4 °C hold. Amplicons were fragmented enzymatically using a QIAseq FX DNA Library kit (Qiagen), sequencing adapters were ligated, and libraries were pooled on an Illumina MiSeq lane and sequenced with 2 × 150 bp reads to 1000 × depth. We used NextGENe v2.3.1 to align raw reads and call variants. Alignment was done to the rCRS reference9 with a matching requirement of ≥ 12 bases and ≥ 85%, and variant calling was done according to the manufacturer's protocols. This included a 20% threshold of original reads for heteroplasmies, which were scored with IUPAC ambiguity codes.
Sanger sequencing
We used Sanger sequencing to validate that the L7 haplogroup-defining mutations were not artifactual. Although we only needed one sequence for validation purposes, we chose three of the novel L7 sequences, one from each subclade, for robustness (see “Results”). For the sequencing reaction, proprietary primers were used with a combination of BigDye Terminator v3.1 (Applied Biosystems by Life Technologies) and custom in-house chemistries. Samples were run through capillary electrophoresis in a 3730 Genetic Analyzer (Applied Biosystems by Life Technologies), and the resulting electropherograms were scored with Sequencher 5.0 software (GeneCodes).
Phylogenetic reconstruction and divergence dating
Each mtDNA genome was aligned to the Reconstructed Sapiens Reference Sequence53 using MUSCLE54. We removed the four most recurrent positions (152, 195, 310, 16519) to reduce the effect of homoplasy. To account for site mutation rate variation, we partitioned the data into six bins: codon position 1, 2, and 3; the hypervariable segments (HVS; HVS-I, HVS-II, and HVS-III combined); the more conserved transfer RNA (tRNA) genes; and other non-coding regions. Phylogenetic inference and divergence dating were performed with BEAST 2.5.255,56. Although only SNP markers were used due to their simple model of evolution, INDELs were identified, annotated according to PhyloTree v1739 conventions, and used later for annotation purposes. INDELs with high recurrence or alignment ambiguity at positions 309–315, 515–524, 3105–3107, 16183–16184, and 16193 were ignored. All heteroplasmies were treated as uncertain bases. All data conversion, marker selection, and file formatting steps (e.g., Nexus) were performed in R 3.5.157. PartitionFinder v1.1.158 was used to select the best site model for each partition, which ranged from simple (HKY + Γ) to complex (GTR + Γ + I). After preliminary analyses using these partition-specific models, the complexity of up to 36 substitution parameters (six rates across six partitions) resulted in poor convergence. Therefore, we simplified this into an HKY + Γ model, with empirical nucleotide frequencies used for the equilibrium states. Exponential priors were used for the gamma shape parameters, and log normal priors were used for the kappa parameters of HKY. We initially compared strict and relaxed log-normal clock models, but observed no appreciable substitution rate variation across branches, based on near-zero standard deviation of the uncorrelated log-normal relaxed clock (ucld.sdev << 1; coefficient of variation 95% HPD overlapping zero), justifying the use of a strict clock.
We calibrated the strict clock with a mutation rate (μ) of 2.285 × 10−8 site−1 year−1, an average of mtDNA mutation rates surveyed in the literature12,59,60,61,62,63,64. Distributions were averaged using the distr65 package in R and used to specify a lognormal prior for the strict clock model. It is important to incorporate a consilience of rate estimates into the total uncertainty to offset potential bias from any one estimate66. We allowed the relative rates of the six partitions to be estimated and fixed to the mean overall rate. For the tree prior we selected a non-parametric Bayesian skyline model so as not to assume anything a priori about population size or tree shape through time. We performed two independent analyses of 5 × 107 MCMC steps sampling every 103 and used Tracer to assess a stationary distribution of posterior samples, within and between MCMC chains. We combined posterior tree samples with LogCombiner and summarized the maximum clade credibility (MCC) tree using TreeAnnotator. We repeated this process separately for just the L5 and L7 samples to estimate effective population size (Ne) as a function of number of coalescent events through time (Bayesian Skyline analysis).
The MCC tree was summarized by collapsing any nodes with < 0.5 posterior probabilities into polytomies. We used the R package treeio67 to input the BEAST tree, and custom functions to collapse uncertain clades. Next, we determined which mutations were synapomorphies (i.e., defining variants) for each remaining clade. We used the data.tree package (github.com/gluc/data.tree) to convert the tree into a hierarchical data structure, and ancestral state reconstruction with the ace function in the ape package68. Ancestral variants defining each clade were estimated using a discrete model fitted by maximum likelihood procedure.
Inference of the best tree topology and support values of each clade differ for Bayesian and maximum likelihood inference69,70. Therefore, we complemented the BEAST analysis with a RAxML71 estimation to ensure consistent topologies, and to compare posterior probabilities with bootstrap frequencies. We ran RAxML 8.2.12 with the “-f a” algorithm to perform 103 rapid bootstraps and search for the best-scoring ML tree, under the GTRCAT model. In order to determine the outgroup for RAxML, and confirm the correct root placement by BEAST, we performed a separate BEAST analysis using 28 complete archaic mitogenomes from NCBI GenBank. All parameters were kept constant, except that only two samples from the eight L haplogroups were included, and the tree tip dates on archaic sequences were constrained using the calibrated C14 dates.
Some of the sequences from Gonder et al.52, particularly in L0, were previously identified as erroneously missing some root-defining mutations, and gaining some “phantom” (false novel) ones, due to mysterious data artifacts43,72. Although neither those authors nor we could find any such issues affecting the five L5/L7 sequences used in this study, we omitted them and repeated the phylogenetic analyses to ensure that the primary results were unaffected.
Haplotype frequency maps
We obtained haplotype data from a large and representative set of studies focusing on African mtDNA to summarize frequency patterns across the continent11,28,29,30,32,35,36,41,42,43,44,45,46,47,48,50,51,52,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100. We used HaploGrep240 to infer haplotypes from PhyloTree v1739 for both full sequences and control region data. If any sequence was classified by the original authors as L5 (or L1e), we used the L5/L7 clade-defining variants (see above) to determine whether the sequence was unambiguously L5/L7, or ambiguously placed in L5'7. Definitive placement required 100% of the available SNPs for each branch or sub-branch to be positive, with no such pattern in the sister clade’s SNPs. Pie charts and heatmaps were then plotted in R v3.5.157 for each major haplogroup using the ggplot2 v3.3.5101 and scatterpie v0.1.7 (https://github.com/GuangchuangYu/scatterpie/) packages.
Ancestral state reconstruction of major population
We used ancestral state reconstruction to find major population patterns in the L5'7 tree. We used largest possible population groupings to capture old and distinct genetic structure: Khoisan-speakers, RFHGs, Bantu, Niger-Congo (non-Bantu), Nilo-Saharan, Afro-Asiatic, and Arabic. Several sequences had unknown population labels but did have admixture analysis results from other studies or the FTDNA database, in which case the dominant sub-Saharan population was used. We used the ace function in the ape package68 and a discrete model fitted by maximum likelihood procedure. Only ancestral nodes with high (> 0.9) likelihood of one population were interpreted as biologically informative.
Ethics statement
All experiments in this study were conducted in adherence with the set of ethical principles of the Declaration of Helsinki. Ethical clearance for the study was obtained from the Pearl Institutional Review Board (protocol number: 21-GBYG-101). Informed consent was obtained from all participants.
Results
Phylogenetic reconstruction and divergence dating
Eighteen unique sequences were found to have the unique pattern: derived for the three L5 synapomorphic mutations T3423C, C12432T, and A16166G, yet negative, or ancestral for the other four (459.1C, A7972G, A12950G, C16148T), and further derived for two additional SNPs A6527G and T11809C (Fig. 1). They were also found to share the retromutation at position 195, but we excluded this hypervariable mutation from our analysis (see “Materials and methods”). Our BEAST and RAxML analyses recovered a topology that confidently splits these 18 samples into a new clade (L7) that is reciprocally monophyletic with L5, united by the new parent clade L5'7 (Fig. 2, Supplementary Figs. S1–S4; Supplementary Tables S3, S4). All newly sequenced samples were sequenced to a mean depth of 1167× across all sites, and 1402× at these relevant sites.
Phylogenetic Placement of New L7 Lineage. Major clades are collapsed, showing the existing (red) and newly discovered (blue) clades. Defining mutations are shown for L5'7, L5, L7, and the six major branches in L7 (L7a, L7a*, L7a1, L7b, L7b1, L7b2). Mutations formerly defining L5b2 that actually define L7 are highlighted in red.
Ultrametric Time Tree. Mean divergence times estimated by BEAST with a fixed clock of \(2.285\times {10}^{-8}\) substitutions site−1 year−1 are shown. L5'7 clades are highlighted in blue. Black circles denote clades with \(\ge\) 0.9 posterior probability, open circles denote clades with \(\ge\) 0.5 but \(<\) 0.9 posterior probability, and polytomies include any clades \(<\) 0.5 posterior probability. Previous haplogroup assignments (in square brackets) are denoted as follows: * = L5, A = L5a, B = L5b, C = L5c (which is an undescribed haplogroup).
Three L7 samples in the academic literature have previously been assigned to subclades of L5 based on PhyloTree v17: KM986522 (L5a1b;51), MF696086 (L5b2;45,102), and MF621126 (L5b2a;50). However, Vyas et al.51 indicated an uncertain placement (“L2'3'4'5'6”), and Cabrera et al.50 correctly suggested their Sudanese sequence MF621126 formed a sister group to known L5 which they termed “L5c.” We elevate this clade to “L7” for two reasons. (1) The samples currently assigned to L5a and L5b are not valid (reciprocally monophyletic) groups since samples in L5b2 are actually sister lineages to all of L5 (i.e., they are L7). Hence, the current nomenclature must change (see Supplementary Table S5 for a mapping between new and previous clades). Elevating L7 re-establishes monophyly and integrity for L5, L5a, and L5b. (2) L7 is even older than L5, making it the third oldest haplogroup after L0 and L1 (Fig. 2; Supplementary Table S3). The newly described L5'7 split is the seventh oldest divergence in the tree at 100.0 [111.9–87.7] kya. We note that the name L7 has been proposed once before to clade “L4g”46, however that was never adopted elsewhere in the literature or PhyloTree database.
In addition to the three L7 samples previously assigned to L5 subclades, we found four L7 Tanzanian samples assigned generically to L552 or “L2'3'4'5'6a1a”102, five samples in the 1000 Genomes Project41, and six unique samples from the private FTDNA database and the Genographic Project103. Sanger sequencing independently confirmed the variant calls in samples from three major L7 subclades: FT-L7-002 (L7a*), FT-L7-004 (L7b1), and FT-L7-006 (L7b2). We found no effect on any parent or sister clades after experimentally removing the samples from Gonder et al.52. Preliminary BEAST analyses using a relaxed log-normal clock showed little rate variation across branches (ucld.sdev = 0.208; coefficient of variation = 0.21) justifying the assumption of a strict clock.
One sequence within the L7a clade (L7a*) shares a most recent common ancestor approximately 79 kya with its sister clade L7a1 (Table 1). Two first cousins were found to share this identical sequence: FT-L7-001, and FT-L7-002 (unused in phylogenetic analyses), both individuals of primarily European descent with South African matrilineal ancestry. The L7a* lineage is therefore the oldest autapomorphic sequence currently known in the human mtDNA tree, with 27 derived SNPs (Table 1; Fig. 2).
Three major subclades of L7 were discovered in addition to the L7a* singleton branch: L7a1 dated to 16.3 [23.4–9.7] kya, L7b1 dated to 22.1 [30.9–14.0] kya, and L7b2 dated to 26.4 [36.4–16.9] kya. BEAST and RAxML both yielded consistent topologies for all major clades (Supplementary Figs. S1, S2). The major L5 subclades L5a1, L5a2, and L5b1 were also found to have mean divergence time estimates between ca. 22 and 28 kya, coinciding with an estimated increase in population size (Fig. 3). Our BEAST analysis of ancient mitogenomes from Neanderthals, Denisovans, and Homo heidelbergensis (Sima de los Huesos sample; SDLH) confirmed L0 as the outgroup in modern human mtDNA and estimated a divergence time of ~ 725 kya for their common hominid ancestor (Supplementary Fig. S3; Supplementary Table S6). The inferred introgression event from pre-modern humans into Neanderthals was dated to ~ 380 kya, which is consistent with earlier work on the SDLH sample5,104,105.
Haplotype frequency maps
African frequencies of the eight L haplogroups reach their peak diversity in the eastern Rift Valley countries of Ethiopia, Kenya, and Tanzania (Fig. 4, Supplementary Fig. S5, S6). We accumulated frequency data from 11,089 African samples across 46 studies (Supplementary Table S7). Minority clades such as L3i, L3x, L4, L5, L6, and L7 reach their highest frequencies in these countries, although generally at less than 10% of the local population (except for L4: 35% in Tanzania). After reclassifying all known L5'7 sequences into either L5 or L7 wherever the available markers permitted, we found that L5 reaches peak frequency (9.32%) in the Mbuti population of eastern Democratic Republic of the Congo (D.R.C.), and L7 (1.37%) in the Sandawe population of Tanzania (Fig. 4b,c).
Distribution of Haplotypes by Population. (a) Frequencies of each major haplogroup by African country, with color codes identical to Fig. 2. D.R.C. = Democratic Republic of the Congo, C.A.R. = Central African Republic; (b) Heatmap of L5 frequency by country normalized to maximum; (c) Heatmap of L7 frequency by country normalized to maximum. For b–c, Countries shaded in gray have no data, and an ‘ × ’ marks the locality with highest frequency (Mbuti population in b, Sandawe population in c).
Ancestral state reconstruction of major population
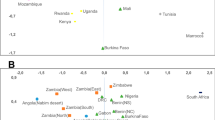
Recent subclades of L5 and L7 tend to share common population groupings, although older subclades have far more uncertain population origins (Fig. 5). We used a population classification scheme that attempts to group the 221 recorded African populations in our dataset into seven major population lineages (Supplementary Table S8). For example, we found that L7a1 is comprised entirely of Bantu (Luhya and Swahili) ethnic groups in Kenya. In contrast, the four L7b1 samples are associated with Afro-Asiatic groups in Ethiopia and outside of Africa in the Middle East: Cushitic and Semitic admixture for two FTDNA customers from Dubai and Ethiopia, Jordanian and Palestinian history in a third. The fourth sample with Sudanese background could not be unambiguously grouped. The multiple occurrences of L7b1 outside of Africa are peculiar and would require further sample discovery on either side of the Red Sea to help establish geographic origin for the subclade. Similarly, L7b2 samples from FTDNA have Ethiopian origins and Semitic-Cushitic admixture in two cases, or Yemeni origins in a third case.
Importantly, the four Khoisan-speaking Sandawe people form their own ~ 4 kya subclade within the ~ 26 kya clade L7b2. An analogous pattern of primarily Bantu or Afro-Asiatic clades is seen in the L5a and L5b topologies, which circumscribe smaller subclades of foraging populations. For example, an ~ 8 kya subclade of Mbuti RFHG people is found in L5a1, and a ~ 1.7 kya subclade of Khoisan Tshwa people is found in L5a2 (Fig. 5). The L5b* lineage is only known from a single sequence shared by two Khoisan Shua individuals in Botswana, whose most recent common ancestor with L5b1 lived ~ 40 kya.
Discussion
Discovery of an ancient lineage
Discovery of an eighth L haplogroup that diverged ~ 100 kya and has a coalescence time of ~ 90 kya is an exciting but unexpectedly late advance in human mtDNA phylogenetics after amassing thousands of sequences for 40 years. The first clades to be assigned “haplogroups” were ordered by their discovery, first A–G in Asia and the Americas106,107, H–K in Europe108, and only one haplogroup L for the most diverse continent of Africa10. Due to incorrect rooting with an Asian sequence, the initial L haplogroups L1 and L2 were rerooted to achieve monophyly in L1, resulting in the first five L haplogroups L0, L1, L5, L2, and L3 containing OOA-subclades M and N32,33,34. Thus, in early papers the L5 lineage was originally described as “L1e”109. Rare eastern African L haplogroups L4 and L6 were the last two to be described36. The ancient divergences in L7, its novel parent group L5'7 (coalesced ~ 100 kya), and the oldest singleton branch L7a* (diverged ~ 80 kya), may shed light on early demographic events of modern humans in East Africa.
Middle stone age expansions in early humans
The earliest modern human population divergences are thought to include: (1) foraging peoples from the ancestors of non-foraging peoples by ~ 160 kya, and (2) Khoisan from other foragers such as RFHGs and Hadza and Sandawe ancestors by ~ 120–100 kya26,27. Subsequent divisions between Niger-Congo, Nilo-Saharan, and Afro-Asiatic lineages and between eastern and western branches of Khoisan and RFHGs occurred less than 55 kya. Modern-day populations such as Hadza and Sandawe illustrate the complexity that ensued: after a ~ 88 kya split from each other, they admixed with Afro-Asiatic groups such as Omotic and Cushitic, and with the Khoisan from whom they likely derive their click-language110. Gradual population growth followed by several major demographic expansions in the Middle Stone Age ~ 75–55 kya35,111,112,113 were the likely catalysts for advancing new terrestrial and marine foraging abilities, microlithic technology, novel pigments in art, and possibly even syntactic language114,115,116,117,118,119. Although most African populations retain a genetic signal of expansion from ~ 70 kya or as early as ~ 110 kya in East African Nilotic groups, serial bottlenecks have largely erased this expansion signal from hunter-gatherer groups120.
Such oscillating periods of population divergence followed by major expansion events with admixture between groups may reflect episodic climatic change across the continent. During glacial maxima, reduced forestation has resulted from increased aridity and desertification, restricting humans to lakeshores, river margins, oceansides, and highlands of Kenya and Ethiopia with persistent canopy cover121. Such oases of habitat may have been important refugia during the arid OIS 6 period from ~ 200–125 kya, if woodland habitat was favored by humans as a source of water, food, and protection from heat and predation. Archaeological and climatic evidence suggests an improvement after this period, however East and tropical Central Africa show inverse population abundances, especially from 130–60 kya, with an increase in Central Africa and a decrease in East Africa122. Changing monsoonal patterns caused wet periods in East Africa during 145–120, 110–95, 80–65, and 55–50 kya, and simultaneous arid periods in Central Africa, including a “megadrought” from 115 to 90 kya122,123,124. Volcanic activity caused by a series of caldera collapses along the Eastern African Rift System also likely made the region uninhabitable for segments of time122. Thus, asynchronous pulses of climatic and tectonic instability125 likely forced human dispersals back and forth across the African continent during much of prehistory.
The L7 and L5 haplogroups diversified ~ 90 kya during this complex period of climatically driven divergence events in East Africa. That coincides with the end of favorably wet conditions in the East African cradle of mtDNA diversity, and the end of a western superdrought, which may have pressured a slight western and southward expansion into the eastern Congo Basin and Great Lakes region of western Tanzania. Other major lineages such as L2 also arose in West Africa at this time before later returning eastward35. Both haplogroups L7 and L5 retain minor subclades that are purely foraging groups: Sandawe people of Tanzania in the case of L7; and both Tshwa/Shua Khoisan of Botswana and east RFHGs of the D.R.C. for L5 (Fig. 5). Each haplogroup also reaches its peak frequency within those populations (Fig. 4b,c). This pattern may hint at the original autochthonous populations that founded these two clades. If Khoisan, RFHGs, and Sandawe foraging populations had relatively small sizes compared to the groups they have recently intermixed with, genetic drift would be expected to drive higher rates of lineage extinction. Previous studies have surmised this process to explain high haplotype frequencies in many foraging groups: L0d and L0k across Khoisan groups29,43; L5 in Tshwa and Shua Khoisan30; L3d in Damara Khoisan30; L1c1a and L1c4 in Baka RFHGs126; L0a2b, L5, L2a2, L2a4 in Mbuti RFHGs28; L4 in Sandawe and Hadza78; and now L7 (formerly “L5”) in Sandawe52.
Population history is rarely represented by any single genetic locus such as mtDNA, for many reasons: (1) phylogenies are diverging (bifurcating or multifurcating) whereas populations are usually reticulating since most populations are products of admixture between ancestral populations; (2) genetic drift causes lineages within smaller populations to be lost faster; and (3) incomplete lineage sorting (fueled by drift) can prevent any single-locus phylogeny from tracking the correct branching order of populations. However, even if a large clade has mixed population identity due to drift or gene flow, a younger subclade may still provide useful population ancestry information if most subclade members are from one population. For example, Khoisan ancestors are thought to be the outgroup to other modern humans, yet L0 (the mtDNA outgroup) is found among many African populations; however, the two subclades L0d and L0k are comprised almost entirely (82% and 83% respectively) of Khoisan-speakers. Typically, this is interpreted to reinforce a correspondence between Khoisan and L030,43,52. The high population diversity of other L0 subclades may represent ancient admixture with those groups, and the specificity of Khoisan-speakers in L0d/L0k may represent the drift within this shrinking population.
Shrinking populations of autochthonous foragers during the past 20 kya may explain why the majority of L5'7 descendants today are primarily Afro-Asiatic (e.g., Cushitic Ethiopian, Ethiopian Jewish, or Yemenite), or eastern Bantu (e.g., Luhya). For example, eastern RFHGs who retain the highest L5 percentage gradually experienced a 40% reduction in effective population size (Ne) starting 20 kya28. A simultaneous seven-fold increase in pre-farming populations suggests that non-foraging populations expanded and possibly subsumed many rare haplogroups by female gene flow. This would explain the long stem lengths for L5a, L5b, L7a, and L7b, with more recent coalescence times of ~ 20 kya (Fig. 2), coinciding with a higher growth rate (Fig. 3). The mtDNA and ethno-linguistic diversity in East Africa is unparalleled in the continent, largely due to the coexistence of ancient linguistic and genetic groups such as Omotic, Cushitic, Semitic, Nilotic, and Chadic73,88. Admixture with these expanding groups may have preserved the exceedingly rare L7 lineage (maximum frequency 1.37%; Fig. 4c). West African Bantu farmers expanded eastward and southward 5–2 kya and assimilated with most local genetic and linguistic groups, leading to their having 15–25% ancestry from west RFHGs, Khoisan, and Afro-Asiatic farmers47,127,128. This second wave of admixture may be the source of the Bantu ancestry in L7a1 and many subclades of L5. The ~ 80 kya relict sequence L7a* from a primarily European woman of South African matrilineal descent is very intriguing. It exposes the process of lineage extirpation, either due to tens of thousands of years of low Ne, or a recent bottleneck (or both). Given that there is < 1% detectable sub-Saharan ancestry in the individual, and the next closest sequence is 80 kya removed, it is imprudent to speculate about the history of this lineage. With time and intensive sampling in southeastern Africa, it is possible more L7a* sequences will be discovered and will help resolve the mystery.
Need to revise the human mtDNA tree
The L7 sequence remained undiscovered partly due to its rarity (< 2% of any population). Only 18 unique full sequences are currently known, and the two synapomorphic SNPs for L7 are in the coding region, thus making them invisible across numerous control region datasets. However, 12 of the 18 full sequences were already publicly available. PhyloTree39 and other semi-static reference trees are also in need of continual re-estimation to avoid overlooking important topological changes such as the L7 clade. So called “phylogenetic placement” methods that use a fixed tree topology as prior information to assign unknown sequences have their place129,130,131, particularly when the sequence does not differ significantly from known sequences, or in metagenomics. However, any new sequence added to a tree alters the context of ancestral and derived states, which is why heuristic algorithms such as TBR or NNI are normally employed in modern phylogenetics to explore optimal tree space132.
Often in human mtDNA phylogenetic studies, maximum parsimony (MP) methods are used34,42,53,79,88, which do indeed explore tree space heuristically. However, MP uses the optimality criterion of fewest changes (minimum homoplasy), which is not often true with increasing timespans and faster mutation rates of hypervariable base positions. A common issue with MP is long branch attraction, whereby unrelated branches that experienced large amounts of evolutionary change are erroneously placed together due to shared mutations133,134. Independently derived mutations can be shared for many reasons in mtDNA, including rapid mutation rate causing convergence by chance, but also selection for mito-nuclear compatibility, which often sorts heteroplasmies according to the nuclear genetic background135,136. Maximum likelihood and Bayesian methods are more ideal for phylogenetic reconstruction of nucleotide sequences because they explicitly model site rate variation and between-lineage heterotachy, although in theory all methods should converge on the same answer if given informative data137.
What is needed now is a periodic re-estimation of the mtDNA tree that incorporates all complete sequences worldwide, and then accurately classifies unknown sequences using the novel reference tree. Such a project has been undertaken recently with a total of 2,243 sequences and MP37, but this falls short of the hundreds of thousands of existing haplotypes that would undoubtedly form myriad new haplogroups. The recently initiated Million Mito Project138 combines data from multiple sources to re-estimate the phylogenetic Tree of Womankind, restructuring, redefining and annotating the branches using more than a quarter of a million current samples, with the goal of incorporating one million full mtDNA sequences.
Conclusions
We have discovered the third oldest L haplogroup fully 40 years after Anderson9 unveiled the first human mtDNA sequence. Hidden within L7 is the oldest known singleton branch of the tree. This exciting twist to the matrilineal story of humanity is undoubtedly not the last, as many unique haplotypes lie hidden and waiting for discovery. Future work should corroborate our preliminary inference that the Sandawe were an autochthonous source of L7 ancestry by resampling that and related populations in the Dodoma region of Tanzania. Although we attempted to identify control region sequences that are L7 using downstream L7a and L7b SNPs, many possible L7 may still elude us, and therefore future studies should fully sequence any such L5'7 samples. We encourage the human mtDNA research community to seek out other ambiguously placed haplotypes based on PhyloTree as potentially new L haplogroups, or other rare or misplaced haplogroups. A global initiative to catalogue and re-estimate the phylogeny for all full mtDNA sequences, such as the Million Mito Project138, should aim to more efficiently track the evolving shape of the Tree of Womankind.
Data availability
The seven novel L7 complete mitochondrial sequences produced in this study are available in the NCBI GenBank repository, under accession numbers ON156774–ON156780. The ancillary L5 sequences produced during the current study are not publicly available due to FamilyTreeDNA privacy terms, but upon reasonable request to the corresponding author, permission can be sought from the participants.
References
Darwin, C. On the Origin of Species by Means of Natural Selection, or the Preservation of Favoured Races in the Struggle for Life. (John Murray, 1859).
Schwartz, R. & Schäffer, A. A. The evolution of tumour phylogenetics: Principles and practice. Nat. Rev. Genet. 18, 213–229 (2017).
Kühnert, D., Wu, C. H. & Drummond, A. J. Phylogenetic and epidemic modeling of rapidly evolving infectious diseases. Infect. Genet. Evol. 11, 1825–1841 (2011).
Smith, S. D., Pennell, M. W., Dunn, C. W. & Edwards, S. V. Phylogenetics is the new genetics (for most of biodiversity). Trends Ecol. Evol. 35, 415–425 (2020).
Petr, M. et al. The evolutionary history of Neanderthal and Denisovan Y chromosomes. Science 369, 1653–1656 (2020).
Cann, R. L., Stoneking, M. & Wilson, A. C. Mitochondrial DNA and human evolution. Nature 325, 31–36 (1987).
Brown, W. M. Polymorphism in mitochondrial DNA of humans as revealed by restriction endonuclease analysis. Proc. Natl. Acad. Sci. USA 77, 3605–3609 (1980).
Vigilant, L., Stoneking, M., Harpending, H., Hawkes, K. & Wilson, A. C. African populations and the evolution of human mitochondrial DNA. Science 253, 1503–1507 (1991).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981).
Chen, Y. S., Torroni, A., Excoffier, L., Santachiara-Benerecetti, A. S. & Wallace, D. C. Analysis of mtDNA variation in African populations reveals the most ancient of all human continent-specific haplogroups. Am. J. Hum. Genet. 57, 133–149 (1995).
Watson, E., Forster, P., Richards, M. & Bandelt, H. J. Mitochondrial footprints of human expansions in africa. Am. J. Hum. Genet. 61, 691–704 (1997).
Poznik, G. D. et al. Sequencing Y chromosomes resolves discrepancy in time to common ancestor of males versus females. Science 341, 562–565 (2013).
Hammer, M. F. et al. The geographic distribution of human Y chromosome variation. Genetics 145, 787–805 (1997).
Malaspinas, A. S. et al. A genomic history of Aboriginal Australia. Nature 538, 207–214 (2016).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Takahata, N., Lee, S. H. & Satta, Y. Testing multiregionality of modern human origins. Mol. Biol. Evol. 18, 172–183 (2001).
Bergström, A., Stringer, C., Hajdinjak, M., Scerri, E. M. L. & Skoglund, P. Origins of modern human ancestry. Nature 590, 40–43 (2021).
Reich, D. et al. Genetic history of an archaic hominin group from Denisova cave in Siberia. Nature 468, 1053–1060 (2010).
Chen, L., Wolf, A. B., Fu, W., Li, L. & Akey, J. M. Identifying and interpreting apparent Neanderthal ancestry in African individuals. Cell 180, 1–11 (2020).
Stringer, C. The origin and evolution of Homo sapiens. Philos. Trans. R. Soc. B Biol. Sci. 371, 20150237 (2016).
Hublin, J. J. et al. New fossils from Jebel Irhoud, Morocco and the pan-African origin of Homo sapiens. Nature 546, 289–292 (2017).
White, T. D. et al. Pleistocene Homo sapiens from Middle Awash, Ethiopia. Nature 423, 742–747 (2003).
Schlebusch, C. M. et al. Southern African ancient genomes estimate modern human divergence to 350,000 to 260,000 years ago. 655, 652–655 (2017).
Garrigan, D. & Hammer, M. F. Reconstructing human origins in the genomic era. Nat. Rev. Genet. 7, 669–680 (2006).
Scerri, E. M. L. et al. Did our species evolve in subdivided populations across Africa, and why does It matter?. Trends Ecol. Evol. 33, 582–594 (2018).
Fan, S. et al. African evolutionary history inferred from whole genome sequence data of 44 indigenous African populations. Genome Biol. 20, 1–14 (2019).
Pereira, L., Mutesa, L., Tindana, P. & Ramsay, M. African genetic diversity and adaptation inform a precision medicine agenda. Nat. Rev. Genet. 22, 284–306 (2021).
Batini, C. et al. Insights into the demographic history of African Pygmies from complete mitochondrial genomes. Mol. Biol. Evol. 28, 1099–1110 (2011).
Barbieri, C. et al. Ancient substructure in early mtDNA lineages of Southern Africa. Am. J. Hum. Genet. 92, 285–292 (2013).
Barbieri, C. et al. Unraveling the complex maternal history of Southern African Khoisan populations. Am. J. Phys. Anthropol. 153, 435–448 (2014).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, 386–394 (2009).
Mishmar, D. et al. Natural selection shaped regional mtDNA variation in humans. Proc. Natl. Acad. Sci. USA 100, 171–176 (2003).
Salas, A. et al. The African diaspora: Mitochondrial DNA and the Atlantic slave trade. Am. J. Hum. Genet. 74, 454–465 (2004).
Shen, P. et al. Reconstruction of patrilineages and matrilineages of Samaritans and other Israeli populations from Y-chromosome and mitochondrial DNA sequence variation. Hum. Mutat. 24, 248–260 (2004).
Silva, M. et al. 60,000 years of interactions between Central and Eastern Africa documented by major African mitochondrial haplogroup L2. Sci. Rep. 5, 1–13 (2015).
Kivisild, T. et al. Ethiopian mitochondrial DNA heritage: tracking gene flow across and around the gate of tears. Am. J. Hum. Genet. 75, 752–770 (2004).
Heinz, T., Pala, M., Gómez-Carballa, A., Richards, M. B. & Salas, A. Updating the African human mitochondrial DNA tree: Relevance to forensic and population genetics. Forensic Sci. Int. Genet. 27, 156–159 (2017).
Blanco, R., Mayordomo, E., Montoya, J. & Ruiz-Pesini, E. Rebooting the human mitochondrial phylogeny: An automated and scalable methodology with expert knowledge. BMC Bioinf. 12, 1–13 (2011).
van Oven, M. PhyloTree Build 17: Growing the human mitochondrial DNA tree. Forensic Sci. Int. Genet. Suppl. Ser. 5, e392–e394 (2015).
Weissensteiner, H. et al. HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res. 44, W58–W63 (2016).
Siva, N. 1000 Genomes project. Nat. Biotechnol. 26, 256 (2008).
Derenko, M. et al. Complete mitochondrial DNA diversity in Iranians. PLoS ONE 8, e80673 (2013).
Behar, D. M. et al. The dawn of human matrilineal diversity. Am. J. Hum. Genet. 82, 1130–1140 (2008).
Behar, D. M. et al. Counting the founders: the matrilineal genetic ancestry of the Jewish Diaspora. PLoS ONE 3, e2062 (2008).
Brucato, N. et al. The Comoros show the earliest Austronesian gene flow into the Swahili Corridor. Am. J. Hum. Genet. 102, 58–68 (2018).
Torroni, A., Achilli, A., Macaulay, V., Richards, M. & Bandelt, H. J. Harvesting the fruit of the human mtDNA tree. Trends Genet. 22, 339–345 (2006).
Barbieri, C. et al. Migration and interaction in a contact zone: mtDNA variation among Bantu-speakers in Southern Africa. PLoS ONE 9, e99117 (2014).
Barbieri, C., Butthof, A., Bostoen, K. & Pakendorf, B. Genetic perspectives on the origin of clicks in Bantu languages from southwestern Zambia. Eur. J. Hum. Genet. 21, 430–436 (2013).
Lippold, S. et al. Human paternal and maternal demographic histories: insights from high-resolution Y chromosome and mtDNA sequences. Investig. Genet. 5, 1–17 (2014).
Cabrera, V. M., Marrero, P., Abu-Amero, K. K. & Larruga, J. M. Carriers of mitochondrial DNA macrohaplogroup L3 basal lineages migrated back to Africa from Asia around 70,000 years ago. BMC Evol. Biol. 18, 1–16 (2018).
Vyas, D. N. et al. Bayesian analyses of Yemeni mitochondrial genomes suggest multiple migration events with Africa and Western Eurasia. Am. J. Phys. Anthropol. 159, 382–393 (2016).
Gonder, M. K., Mortensen, H. M., Reed, F. A., De Sousa, A. & Tishkoff, S. A. Whole-mtDNA genome sequence analysis of ancient African lineages. Mol. Biol. Evol. 24, 757–768 (2007).
Behar, D. M. et al. A ‘Copernican’ reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012).
Edgar, R. C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinf. 5, 1–119 (2004).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 1–8 (2007).
Bouckaert, R. et al. BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
R Core Team. R: A language and environment for statistical computing. (2022).
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Soares, P. et al. Correcting for purifying selection: an improved human mitochondrial molecular clock. Am. J. Hum. Genet. 84, 740–759 (2009).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Brotherton, P. et al. Neolithic mitochondrial haplogroup H genomes and the genetic origins of Europeans. Nat. Commun. 4, 1–11 (2013).
Rieux, A. et al. Improved calibration of the human mitochondrial clock using ancient genomes. Mol. Biol. Evol. 31, 2780–2792 (2014).
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–44913 (2014).
Kivisild, T. Maternal ancestry and population history from whole mitochondrial genomes. Investig. Genet. 6, 1–10 (2015).
Camphausen, A. F., Kohl, M., Ruckdeschel, P., Stabla, T. & Ruckdeschel, P. The distr Package. October 1–208 (2019).
Ho, S. Y. W. & Endicott, P. The crucial role of calibration in molecular date estimates for the peopling of the Americas. Am. J. Hum. Genet. 83, 142–146 (2008).
Wang, L. et al. Treeio: an R package for phylogenetic tree input and output with richly annotated and associated data. Mol. Biol. Evol. 37, 599–603 (2020).
Paradis, E. & Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Svennblad, B., Erixon, P. E. R., Oxelman, B. & Britton, T. O. M. Fundamental differences between the methods of maximum likelihood and maximum posterior probability in phylogenetics. Syst. Biol. 55, 116–121 (2006).
Douady, C. J., Boucher, Y., Doolittle, W. F. & Douzery, E. J. P. Comparison of Bayesian and maximum likelihood bootstrap measures of phylogenetic reliability. Mol. Biol. Evol. 20, 248–254 (2003).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Yao, Y. G., Salas, A., Logan, I. & Bandelt, H. J. mtDNA data mining in GenBank needs surveying. Am. J. Hum. Genet. 85, 929–933 (2009).
Boattini, A. et al. MtDNA variation in East Africa unravels the history of Afro-Asiatic groups. Am. J. Phys. Anthropol. 150, 375–385 (2013).
Barbieri, C. et al. Contrasting maternal and paternal histories in the linguistic context of Burkina Faso. Mol. Biol. Evol. 29, 1213–1223 (2012).
Castrì, L. et al. mtDNA variability in two bantu-speaking populations (Shona and Hutu) from Eastern Africa: Implications for peopling and migration patterns in sub-Saharan Africa. Am. J. Phys. Anthropol. 140, 302–311 (2009).
Poloni, E. S. et al. Genetic evidence for complexity in ethnic differentiation and history in East Africa. Ann. Hum. Genet. 73, 582–600 (2009).
Quintana-Murci, L. et al. Maternal traces of deep common ancestry and asymmetric gene flow between Pygmy hunter-gatherers and Bantu-speaking farmers. Proc. Natl. Acad. Sci. USA 105, 1596–1601 (2008).
Tishkoff, S. A. et al. History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation. Mol. Biol. Evol. 24, 2180–2195 (2007).
Kivisild, T. et al. The role of selection in the evolution of human mitochondrial genomes. Genetics 172, 373–387 (2006).
Howell, N., Elson, J. L., Turnbull, D. M. & Herrnstadt, C. African haplogroup L mtDNA sequences show violations of clock-like evolution. Mol. Biol. Evol. 21, 1843–1854 (2004).
Torroni, A. et al. Do the four clades of the mtDNA haplogroup L2 evolve at different rates?. Trans. Am. Soc. Agric. Eng. 69, 1348–1356 (2001).
Ingman, M., Kaessmann, H., Pääbo, S. & Gyllensten, U. Mitochondrial genome variation and the origin of modern humans. Nature 408, 708–713 (2000).
Li, J. Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Maca-Meyer, N., González, A. M., Larruga, J. M., Flores, C. & Cabrera, V. M. Major genomic mitochondrial lineages delineate early human expansions. BMC Genet. 2, 13 (2001).
Maca-Meyer, N. et al. Mitochondrial DNA transit between West Asia and North Africa inferred from U6 phylogeography. BMC Genet. 4, 1–11 (2003).
Soares, P. et al. The expansion of mtDNA haplogroup L3 within and out of Africa. Mol. Biol. Evol. 29, 915–927 (2012).
Gomes, V. et al. Mosaic maternal ancestry in the Great Lakes region of East Africa. Hum. Genet. 134, 1013–1027 (2015).
Non, A. L., Al-Meeri, A., Raaum, R. L., Sanchez, L. F. & Mulligan, C. J. Mitochondrial DNA reveals distinct evolutionary histories for Jewish populations in Yemen and Ethiopia. Am. J. Phys. Anthropol. 144, 1–10 (2011).
Kampira, E., Kumwenda, J., Van Oosterhout, J. J. & Dandara, C. Mitochondrial DNA subhaplogroups L0a2 and L2a modify susceptibility to peripheral neuropathy in Malawian adults on stavudine containing highly active antiretroviral therapy. J. Acquir. Immune Defic. Syndr. 63, 647–652 (2013).
Pierron, D. et al. Genomic landscape of human diversity across Madagascar. Proc. Natl. Acad. Sci. USA 114, E6498–E6506 (2017).
Göbel, T. M. K. et al. Mitochondrial DNA variation in sub-Saharan Africa: forensic data from a mixed West African sample, Côte d’Ivoire (Ivory Coast), and Rwanda. Forensic Sci. Int. Genet. 44, 1–6 (2020).
Brucato, N. et al. The imprint of the slave trade in an African American population: mitochondrial DNA, Y chromosome and HTLV-1 analysis in the Noir Marron of French Guiana. BMC Evol. Biol. 10, 1–19 (2010).
Ottoni, C. et al. First genetic insight into Libyan Tuaregs: A maternal perspective. Ann. Hum. Genet. 73, 438–448 (2009).
Bekada, A. et al. Introducing the Algerian mitochondrial DNA and Y-chromosome profiles into the North African landscape. PLoS ONE 8, e56775 (2013).
Coudray, C. et al. The complex and diversified mitochondrial gene pool of Berber populations. Ann. Hum. Genet. 73, 196–214 (2009).
Rando, J. C. et al. Mitochondrial DNA analysis of Northwest African populations reveals genetic exchanges with European, Near-Eastern, and sub-Saharan populations. Ann. Hum. Genet. 62, 531–550 (1998).
Cherni, L. et al. Post-last glacial maximum expansion from Iberia to North Africa revealed by fine characterization of mtDNA H haplogroup in Tunisia. Am. J. Phys. Anthropol. 139, 253–260 (2009).
Ely, B., Wilson, J. L., Jackson, F. & Jackson, B. A. African-American mitochondrial DNAs often match mtDNAs found in multiple African ethnic groups. BMC Biol. 4, 1–14 (2006).
González, A. M. et al. Mitochondrial DNA variation in Mauritania and Mali and their genetic relationship to other Western Africa populations. Ann. Hum. Genet. 70, 631–657 (2006).
Wickham, H. ggplot2. Wiley Interdiscip. Rev. Comput. Stat. 3, 180–185 (2011).
MTree. YFull MTree 1.02.8880. YFull (2021). Available at: https://www.yfull.com/mtree/L2'3'4'5'6a/. (Accessed: 22nd April 2021).
Behar, D. M. et al. The Genographic Project public participation mitochondrial DNA database. PLoS Genet. 3, 1083–1095 (2007).
Posth, C. et al. Deeply divergent archaic mitochondrial genome provides lower time boundary for African gene flow into Neanderthals. Nat. Commun. 8, 1–9 (2017).
Meyer, M. et al. A mitochondrial genome sequence of a hominin from Sima de los Huesos. Nature 505, 403–406 (2014).
Torroni, A. et al. Asian affinities and continental radiation of the four founding Native American mtDNAs. Am. J. Hum. Genet. 53, 563–590 (1993).
Torroni, A. et al. Mitochondrial DNA analysis in Tibet: implications for the origin of the Tibetan population and its adaptation to high altitude. Am. J. Phys. Anthropol. 93, 189–199 (1994).
Torroni, A. et al. mtDNA and the origin of Caucasians: identification of ancient Caucasian-specific haplogroups, one of which is prone to a recurrent somatic duplication in the D-loop region. Am. J. Hum. Genet. 55, 760–776 (1994).
Salas, A. et al. The making of the African mtDNA landscape. Am. J. Hum. Genet. 71, 1082–1111 (2002).
Shriner, D., Tekola-Ayele, F., Adeyemo, A. & Rotimi, C. N. Genetic ancestry of Hadza and Sandawe peoples reveals ancient population structure in Africa. Genome Biol. Evol. 10, 875–882 (2018).
Atkinson, Q. D., Gray, R. D. & Drummond, A. J. mtDNA variation predicts population size in humans and reveals a major Southern Asian chapter in human prehistory. Mol. Biol. Evol. 25, 468–474 (2008).
Kim, H. L. et al. Khoisan hunter-gatherers have been the largest population throughout most of modern-human demographic history. Nat. Commun. 5, 1–8 (2014).
Henn, B. M., Cavalli-Sforza, L. L. & Feldman, M. W. The great human expansion. Proc. Natl. Acad. Sci. USA 109, 17758–17764 (2012).
Henshilwood, C. S., D’errico, F., Marean, C. W., Milo, R. G. & Yates, R. An early bone tool industry from the Middle Stone Age at Blombos Cave, South Africa: implications for the origins of modern human behaviour, symbolism and language. J. Hum. Evol. 41, 631–678 (2001).
Henshilwood, C. S. & Dubreuil, B. Reading the artifacts: gleaning language skills from the Middle Stone Age in southern Africa. in The Cradle of Language (eds. Botha, R. & Knight, C.) 41–61 (Oxford University Press, 2009).
Henshilwood, C. S. et al. Emergence of modern human behavior: Middle Stone Age engravings from South Africa. Science 295, 1278–1280 (2002).
Mellars, P. Why did modern human populations disperse from Africa ca. 60,000 years ago? A new model. Proc. Natl. Acad. Sci. 103, 9381–9386 (2006).
McBrearty, S. & Brooks, A. S. The revolution that wasn’t: a new interpretation of the origin of modern human behavior. J. Hum. Evol. 39, 453–563 (2000).
Scheinfeldt, L. B., Soi, S. & Tishkoff, S. A. Working toward a synthesis of archaeological, linguistic, and genetic data for inferring African population history. Proc. Natl. Acad. Sci. USA 107, 8931–8938 (2010).
Excoffier, L. & Schneider, S. Why hunter-gatherer populations do not show signs of Pleistocene demographic expansions. Proc. Natl. Acad. Sci. USA 96, 10597–10602 (1999).
Basell, L. S. Middle Stone Age (MSA) site distributions in eastern Africa and their relationship to Quaternary environmental change, refugia and the evolution of Homo sapiens. Quat. Sci. Rev. 27, 2484–2498 (2008).
Blome, M. W., Cohen, A. S., Tryon, C. A., Brooks, A. S. & Russell, J. The environmental context for the origins of modern human diversity: A synthesis of regional variability in African climate 150,000–30,000 years ago. J. Hum. Evol. 62, 563–592 (2012).
Scholz, C. A. et al. East African megadroughts between 135 and 75 thousand years ago and bearing on early-modern human origins. Proc. Natl. Acad. Sci. USA 104, 16416–16421 (2007).
Cohen, A. S. et al. Ecological consequences of early Late Pleistocene megadroughts in tropical Africa. Proc. Natl. Acad. Sci. USA 104, 16422–16427 (2007).
Shultz, S. & Maslin, M. Early human speciation, brain expansion and dispersal influenced by African climate pulses. PLoS ONE 8, e76750 (2013).
Batini, C. et al. Phylogeography of the human mitochondrial L1c haplogroup: Genetic signatures of the prehistory of Central Africa. Mol. Phylogenet. Evol. 43, 635–644 (2007).
Patin, E. et al. Dispersals and genetic adaptation of Bantu-speaking populations in Africa and North America. Science 356, 543–546 (2017).
de Filippo, C., Bostoen, K., Stoneking, M. & Pakendorf, B. Bringing together linguistic and genetic evidence to test the Bantu expansion. Proc. R. Soc. B Biol. Sci. 279, 3256–3263 (2012).
Matsen, F., Kodner, R. & Armbrust, E. pplacer: Linear time maximum-likelihood and Bayesian phylogenetic placement of sequences onto a fixed reference tree. BMC Bioinformatics 11, 538 (2010).
Balaban, M., Sarmashghi, S. & Mirarab, S. APPLES: scalable distance-based phylogenetic placement with or without alignments. Syst. Biol. 69, 566–578 (2020).
Czech, L., Barbera, P. & Stamatakis, A. Methods for automatic reference trees and multilevel phylogenetic placement. Bioinformatics 35, 1151–1158 (2019).
John, K. S. Review paper: the shape of phylogenetic treespace. Syst. Biol. 66, e83–e94 (2017).
Philippe, H., Zhou, Y., Brinkmann, H., Rodrigue, N. & Delsuc, F. Heterotachy and long-branch attraction in phylogenetics. BMC Evol. Biol. 5, 1–8 (2005).
Spencer, M., Susko, E. & Roger, A. J. Likelihood, parsimony, and heterogeneous evolution. Mol. Biol. Evol. 22, 1161–1164 (2005).
Wei, W. et al. Germline selection shapes human mitochondrial DNA diversity. Science 364, eaau6520 (2019).
Blier, P. U. P., Dufresne, F. & Burton, R. R. S. Natural selection and the evolution of mtDNA-encoded peptides: Evidence for intergenomic co-adaptation. Trends Genet. 17, 400–406 (2001).
Brooks, D. R. et al. Quantitative phylogenetic analysis in the 21st century. Rev. Mex. Biodivers. 78, 225–252 (2007).
Estes, R. J. Million Mito Project. DNAeXplained (2021). Available at: https://dna-explained.com/2020/03/17/the-million-mito-project/. (Accessed: 22nd April 2021).
Acknowledgements
We are grateful to all sample donors for participating in this study. We would like to thank Doron M. Behar for his invaluable feedback during initial validation, and Luisa Sanchez for helping to Sanger validate sequences. Katy Rowe, Haydar Alyassery, and Ibrahim A. A. Ibrahim assisted with customer contact and acquiring consent to upgrades and research participation. We are grateful to Toomas Kivisild of Cambridge University and Maere Reidla of University of Tartu who provided original DNA samples from Ethiopia. Gregory Vilshansky (previously of National Geographic Society) helped query existing sequences to find potential donors for the project. We thank Connie Bormans and the Gene by Gene, Ltd. lab for their help sequencing novel samples. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. We would like to thank Bennett Greenspan (recently retired after founding FamilyTreeDNA) for his two decades of dedication to scientific discovery and genealogy, and for his commitment to scientific research and support for the ambitious Million Mito Project, without which this ongoing research would not be possible.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the conception and initial design of the study. G.R. discovered the first L7 sequence in the FamilyTreeDNA database. G.R. and M.G.V. helped to acquire full sequences for the study. P.A.M. performed phylogenetic analyses, wrote R scripts, summarized public sequences from NCBI, and wrote the manuscript. M.G.V., R.J.E., and G.R. provided important feedback in editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.A.M. and G.R. are employees of and R.J.E and M.G.V. are contractors for FamilyTreeDNA.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maier, P.A., Runfeldt, G., Estes, R.J. et al. African mitochondrial haplogroup L7: a 100,000-year-old maternal human lineage discovered through reassessment and new sequencing. Sci Rep 12, 10747 (2022). https://doi.org/10.1038/s41598-022-13856-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13856-0
This article is cited by
-
mtDNA “nomenclutter” and its consequences on the interpretation of genetic data
BMC Ecology and Evolution (2024)
-
Unraveling the mitochondrial phylogenetic landscape of Thailand reveals complex admixture and demographic dynamics
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.