Abstract
The integration of smartphones and microfluidics is nowadays the best possible route to achieve effective point-of-need testing (PONT), a concept increasingly demanded in the fields of human health, agriculture, food safety, and environmental monitoring. Nevertheless, efforts are still required to integrally seize all the advantages of smartphones, as well as to share the developments in easily adoptable formats. For this purpose, here we present the free platform appuente that was designed for the easy integration of microfluidic chips, smartphones, and the cloud. It includes a mobile app for end users, which provides chip identification and tracking, guidance and control, processing, smart-imaging, result reporting and cloud and Internet of Things (IoT) integration. The platform also includes a web app for PONT developers, to easily customize their mobile apps and manage the data of administered tests. Three application examples were used to validate appuente: a dummy grayscale detector that mimics quantitative colorimetric tests, a root elongation assay for pesticide toxicity assessment, and a lateral flow immunoassay for leptospirosis detection. The platform openly offers fast prototyping of smartphone apps to the wide community of lab-on-a-chip developers, and also serves as a friendly framework for new techniques, IoT integration and further capabilities. Exploiting these advantages will certainly help to enlarge the use of PONT with real-time connectivity in the near future.
Similar content being viewed by others
Introduction
Internet-of-Things (IoT) emerged in the last few years and rapidly grew to become one of the most important high-tech trends1. Although there is no clear consensus on IoT definition, it may be thought as the interconnection of people, objects and systems for smart data collection, processing and reaction2. IoT opens limitless possibilities in applications including agriculture, smart cities, commerce and industry3. Most authors identify three main layers within IoT: a first layer for sensing purposes; a second layer in charge of communication, storage, and processing; and a third layer to interface the user, providing information management and data analytics4. The second and third layers are transverse to all application fields, and have shown great advances in the last years2. In contrast, the first layer is application-specific and its readiness level is then heterogeneous. A relevant case is the detection of biomolecules in real time, which is connected to the concept of point-of-need testing (PONT), i.e. in-field, real-time chemical determinations. When the test is related to human health, this concept is referred to as point-of-care testing (POCT).
PONT is demanded in the fields of human and animal health, food safety, environmental monitoring and surveillance5. In 2003, the World Health Organization defined the ASSURED criteria to define the ideal POCT, which should be affordable, sensitive, specific, user-friendly, rapid and robust, equipment-free and deliverable to end users6. This concept was recently reaffirmed and renewed, with the REASSURED acronym, now including real-time connectivity and easy specimen collection7. Even though many efforts have been conducted from academic and industrial sectors, the PONT necessity is still unmet8, excluding some exceptional cases9. The progress achieved on PONT is based on microfluidic technology, in any of its forms: glass chips10 lateral flow immunoassays (LFIA)11, paper-based analytical devices (µPAD)12, polymeric microfluidic devices13, or hybrid devices14. Microfluidics presents inherent advantageous characteristics like requiring small amounts of sample and reagents, decreased reaction times15, and the potential to implement chemical operations on a reduced chip16. However, most of the approaches fail to accomplish all REASSURED criteria simultaneously, mainly because additional hardware is needed for the purpose of providing off-chip operations like heating, pumping, valve actuation and reading. In the best case, all the operations needed are centralized in a single piece of equipment17,18. Usually, sensitivity, specificity and speed are addressed but neglecting autonomy, portability, deliverability and sometimes cost.
In the last decade, the integration of smartphones and microfluidic devices has been proposed19 and explored in many directions, as described in a series of recently published review articles20,21,22. Smartphones offer many complementary advantages, like computing power, data storage, power sourcing, connectivity, a diversity of sensors, traceability, geolocation, the possibility of guiding the user throughout the test23,24, as well as control of the validity and quality of measurements, using specific apps (hereafter app will be used to mean software application, while application will be used to denote a specific field of use). Even more, smartphones are a natural gateway for PONT to give place to complex IoT applications involving sensing, smart decision making and even actuation. The majority of the smartphone-based approaches also rely on additional hardware. According to the application, this hardware can be: (i) a potentiostat25,26, (ii) a dark-box for digital imaging, meant to provide proper geometry and light parameters26,27,28, (iii) a heating module29,30, (iv) a fluid handling module or (v) combinations of the previous ones17,31,32,33,34. The accessories create additional costs, may present hygiene issues, and usually are phone-model specific35. These approaches are also detrimental to portability, autonomy and deliverability, deviating from the ASSURED criteria.
Interestingly, some PONT microfluidic devices use smartphones without additional hardware. This approach mainly relies on digital imaging using the integrated camera, with rare exceptions36. Nowadays smartphones are ubiquitous37 and PONT approaches could include them without loss of portability, deliverability or autonomy. Moreover, smartphones can be used to improve other characteristics of ASSURED, like sensibility, specificity, usability and robustness. Even more, smartphones guarantee the real time connectivity deemed necessary in REASSURED. Advantages of only using smartphones compared to the approaches using additional hardware are: (i) more autonomy and portability, (ii) lower cost, (iii) easier handling, (iv) fewer cross-contamination and hygiene issues26. When solely using a smartphone, different reproducibility issues emerge due to: (i) ambient light intensity and color variability38,39,40,41, (ii) non-uniform illumination41, (iii) variability in geometric parameters like capture distance and angles41, (iv) variability in camera parameters (focus mode, exposure time, ISO, resolution)41,42, (v) phone-specific pre-processing algorithms41,43 and (vi) brand/model-specific variability40. Various strategies have been developed to circumvent these issues. For example, chromatic and light source effects can be separated by using different color spaces35,39,40,42,43 or advanced processing methods41. The built-in LED can be used as the main light source to address light uniformity35,42. Also, on-site or on-chip calibration can be performed. The former40 requires additional time, testing, and training, while the latter involves additional features in the chip, such as color35,39 or chart references41,43, controls44,45, or standard addition assays38. Also, alignment marks41,42,46 or frames47 can be included to control the variability of geometric parameters.
Irrespective of the inclusion of additional hardware, most of the works exploit a limited set of features offered by phones, without taking advantage of capabilities such as image processing, timers, alarms, and traceability controls48, or the aforementioned ability to guide and control the user actions through step-by-step multimedia instructions49. Moreover, previous works are neither customizable, free or open-source, meaning that developed tools cannot be easily adopted by others, hindering fast prototyping and technology accessibility. The few exceptions to some of these issues are an open-source potentiostat for electrochemistry-related applications26, a free stand-alone software for colorimetry analysis for test developers50, and a paid software product where customization is undertaken by a team of programmers51. Thus, to leverage IoT-PONT applications, a platform enabling easy integration of smartphones to microfluidic-based projects is needed.
This paper reports the development of a free platform called appuente, for the easy integration of microfluidic chips, smartphones, and the cloud, which can be further exploited for IoT. This platform includes a web app for PONT developers, to easily customize and manage the data of the administered tests. It also includes a smartphone app, to guide and control the end-user actions throughout the test, by using tools and peripherals provided by smartphones. The platform was conceived from the beginning to be customizable and ideally serve any application, being also flexible to add further features. Moreover, appuente is meant to help test developers to take advantage of all the potential offered by smartphones, like (i) step-by-step multimedia guidance, (ii) control mechanisms like timers, alarms and tracking, (iii) compatibility with all the mentioned strategies to improve reproducibility and (iv) cloud connectivity and IoT integration, among others. Although appuente is mainly thought for the integration of microfluidics and smartphones without additional hardware, the platform is also compatible with several miniaturized laboratory assays (see for example52), as well as with applications that involve additional hardware (see for example53,54). In order to illustrate and validate the performance of the platform, three application cases are presented. The first one is a dummy application to detect levels of gray, (mimicking e.g. the output of quantitative colorimetric test), which enables full characterization of the system. The second application case is a milli-fluidic root elongation assay for pesticide toxicity assessment52. The last case is a LFIA for leptospirosis detection55. The potential of the platform to produce equipment-free, robust, qualitative and quantitative assays is achieved in this article by including proper calibration features within the chip. Finally, it is worth to remark that we are presenting a mixed platform that gathers all the advantages that mobile devices and the cloud offer to microfluidic devices for PONT. Some of these advantages have been previously reported, but they were given separately and some of them are inaccessible. Here, we integrate all the improvements on appuente and provide them in an easily accessible format.
Appuente
Before going through a detailed description of the platform, it is relevant to introduce some declarations. Firstly, it is worth mentioning that puente means bridge in Spanish, hence the name appuente sets the mission of making accessible the large number of smartphones capabilities to the community of microfluidic developers. Secondly, it should be noted that there are three key roles in appuente: developers, users and patients. The first one is the person/team developing the test, who will mainly use the web app. The user is the person administering the test, like a health-care worker, in the case of POCT requiring trained personnel, or the interested itself, in the cases of elementary tests. The user will handle the mobile app. The patient role, which may exist or not depending on the application, is the person to whom the test is administered. This role might only provide the sample to be analyzed or, in the case of completely autonomous PONTs, can be simultaneously the user. Finally, it must be stated that appuente, in its present form, is intended for research, education, and product prototyping. The platform should not be straightforwardly used in operational environments that require compliance with safety policies or other regulation. In what follows, we describe the main components of the developed platform: microfluidic chip, mobile app, and web app.
Microfluidic chip
As represented in Fig. 1a, appuente requires microfluidic chips to include alignment marks and an ID code. The marks are meant to avoid the geometric variability from assay to assay, due to different distances and angles when the image is taken. The ID code is meant to identify the test type and the singular chip, in order to apply the controls described in the next subsection. Also, the features that determine the assay result must be defined. The number, size, position, and meaning of such features depend on the application, and will be usually associated with a calibration procedure and/or result reading (for example, test and control lines in LFIA).
The ID codes and alignment marks are automatically generated by the platform and provided in a vector graphics file (SVG or PDF format) together with the mentioned features. The developer can use such file as a starting point to add the application-specific microfluidics, and then to materialize the chip (via wax printing, 3D printing, laser engraving). Note that appuente does not require a particular material for the chip, which depends entirely on the design purposes of the developer.
Elementary parts of appuente. (a) Microfluidic chip structures required by appuente. Besides alignment marks and ID code, the developer must define the features that will be used to obtain the test result. In this example, N features (F1, F2,...FN) were defined. (b) Inputs and outputs for the custom algorithm. The developer must provide a .m file with the desired algorithm. This algorithm will receive a data structure with the available information about the project and the particular assay, and must return another structure to be used in the final report. (c) The information returned is shown in the smartphone, at the end of the process, in a customized report.
Mobile app
As represented in Fig. 2, the operation of the mobile app is as follows. First, the user scans the ID code of the chip using a scanner built-in the app. Using this ID code the mobile app downloads the test instructions from the server. At this point, the server runs several controls to check for expiration date, no-reuse (for disposable tests), and batch validity (for possible device recalls). If the chip passes such controls, the app guides the user throughout the test-specific steps using the multimedia resources provided by the developer, such as text and images. During this stage, user actions can be controlled by the app with alarms and timers, e.g. by setting a window of opportunity for reading results. In the last step, the mobile app helps the user to take a picture with standard characteristics (regarding geometry and ambient light conditions), in order to guarantee result reproducibility within a range. After this operation, the app sends the cropped features inside an assay data structure to the server, together with other relevant information about the test. The server runs a custom algorithm in GNU Octave and sends back a report structure (refer to the Supplementary Information for detail about the named structures) to the app. This process is schematized in Fig. 1b. Finally, the mobile app shows the results to the user (Fig. 1c), along with other information deemed necessary by the developer (e.g. error ranges and recommendations). A video showing the operation of the mobile app is provided with the Supplementary Information.
Representation of the typical flow diagram followed by appuente developers and users and interactions through the database (DB). (*) Note that after the ID code scan, the specific steps to be followed depend on the project; thus, the “Follow steps” box enumerates a subset of the offered possibilities.
Web app
The web app (appuente.com) is meant for test developers and allows guided customization of both the mobile app and the microfluidic chip, to cover the application needs. The developer can follow two workflows, as represented in Fig. 2: one for project creation and the other for data management. In the former, the developer firstly logs in and creates a new project, then fills a form with general information about the test (e.g. name, description, icon) and specific information that will be part of the mobile app: e.g test summary and number, type and details of steps to be followed. Note that the last step will always be a smart picture, i.e., a picture where geometric and ambient-light parameters are controlled. Then, the developer loads geometric information about the chip and its features. Next, uploads the algorithm that will be applied to the information collected in the smart picture step. Such algorithm has to be uploaded in a .m file, and will be run in GNU Octave, a free, open-source alternative to Matlab and compatible with it. This algorithm has to be created and debugged by the developer, and the only requirement is that the input (assay) and output (report) data structures are used for communication with the platform. Some of the algorithms used in this work are given as examples in the Supplementary Information. Next, the developer has to design the report that will be seen by the user. Finally, the developer can download the vector geometry of the chip to add the microfluidics and materialize (e.g. by printing or engraving) the actual chips. The developer is expected to iterate this workflow until the mobile application behaves as desired. Note that, if the developer wishes further customization (e.g. to serve specific normative or to manage the database in another server), the project can be ejected from the platform. With this action, the source code of the mobile app is provided under a BSD-3-Clause License and can be downloaded to be freely managed elsewhere.
In the second workflow, where the test is already designed and available to users, the developer can access the database and manage the collected information. In the current version, appuente only allows data inspection and downloading; on-site analytics and integration to external databases will be included in the near future.
Results
Application case #1: Grayscale detector
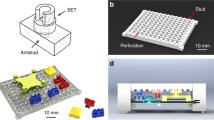
The developed platform was tested with a simple application case meant to measure the level of gray of a region within the chip, mimicking tests based on colorimetric reading, where the amount of analyte can be correlated to the intensity of the reflected light. On the other hand, this application case also allows thorough characterization of the appuente performance, and serves as a tutorial from which new developers can start their projects. The chips used are depicted in Fig. 3a,b, where the alignment marks, the ID code and the features (color references and the unknown level of gray) can be observed. Following the flow diagram for developers in Fig. 2, a new project was created and general information about the test was filled in, as well as geometric information about the chip, feature sizes and positions. Next, an Octave algorithm (described in Methods) was selected after being coded and debugged in the desktop version of GNU Octave. The result returned by this algorithm is informed on a custom report shown on the smartphone screen (Fig. 3g). Next, the chips were printed on regular copy paper and cut. Proper functioning of the whole project was achieved after a couple of iterations of the previously described workflow.
In order to quantify the performance of the platform, several studies were carried out, whose results are collected in Table 1. The first five rows measure the error in determining a known level of gray under different situations. In particular, the fist two rows compare the performance of two different ways of providing the color references: the simplest one, where the extreme white and black references are provided (Fig. 3a) and an enhanced version, where the complete gray gradient is printed on the chip (Fig. 3b). In the third row, results obtained under ambient light intensity varying from 150 to 10,000 lx are compared. The fourth and fifth rows assess the variations obtained using different phone models and by different operators. The last two rows measure the ability to find the actual feature within the chip, as a function of the feature size. Again, two alternatives are compared: the simplest one, using the native capabilities of the platform, and an enhanced version using image processing techniques to find the actual feature within the image, as shown in Fig. 3d,e. Finally, results obtained using Octave where compared with Matlab (vR2017a, mathworks.com). For this purpose, a subset of the images from the experiment in the second row of Table 1, were analyzed with the same algorithm used for Octave, with minimum differences due to sintaxis variation between both programs. Results obtained were practically equivalent (differences lower than 0.1%).
Application case #1: grayscale detector. The chips used include two kinds of features: the unknown and the color references, provided as (a) the extreme gray values or (b) a gradient with the full palette. (c) A screenshot of the mobile app showing the auto-shooting step. (d) The cropped features are passed to the algorithm for processing. (e) The algorithm segments the analyzed features in order to recognize the pixels containing useful information. (f) Plot of the measured reference gradient and the absolute error (between such gradient and the mean unknown value) as a function of position. The position where the error reaches a minimum is proportional to the unknown gray value. (g) Screenshot of the final report showing the obtained result.
Application case #2: Device for root elongation assays
As mentioned in the Introduction, appuente was applied to a bioassay for measuring root elongation, which is referred to as Plant chip. Root elongation assays are commonly used to assess the phytotoxicity of chemical compounds. Seeds of certain species are grown and, after a specified time period, the root lengths of test and control groups are compared. This procedure is traditionally carried out in Petri dishes, which requires several time-consuming tasks. Plant chip was recently presented as a surpassing alternative, where roots are grown in an array of 20 milli-channels, allowing the operator to measure root lengths directly from a digital picture with the aid of image processing software52. The combination of Plant chip and appuente is proposed here to further improve and simplify the measurement procedure, as well as to evaluate root elongation as a function of time, which is a current need in the field52. Table 2 summarizes the different steps necessary for root elongation assays in different formats: the conventional assay on Petri dishes, Plant chip, and the combination of Plant chip and appuente.
For this application case, the original Plant chip devices were modified to enhance contrast, using 3D-printed black PLA milli-channel array (Fig. 4a). A detailed description about plant chip fabrication can be found in the Methods section, as well as in the previous publication52. The appuente platform was used to produce a mobile app in which the user is prompted to (i) scan the ID code placed at the top of the chip, (ii) draw a red mark over each channel at the end position of every root (Fig. 4b), and (iii) align the chip for the app to automatically take a picture (Fig. 4c). Then, the cropped channel area is fed to the custom algorithm, the result of which is returned and shown in a final report that indicates the mean measured length and the standard deviation. An alternative procedure avoids the red mark step by using a more complex algorithm, which is capable of detecting roots and performing measurements directly. Such an approach makes operation even simpler, at the expense of additional processing time and the requirement of higher resolution images. To overcome these issues, a third alternative that makes use of machine learning techniques is currently being explored (not shown here). Figure 4d shows the results obtained using the simpler approach, in which the average root length obtained in four different experiments (n = 20, seeds per experiment) is shown as a function of time. In the same figure, the average length of the 80 analyzed roots is compared with the measurements performed by an expert, using the original Plant chip approach.
Application case #2. (a) Exploded-view drawing of the Plant chip device. (b) Photograph of the device showing an assay in progress and the red marks made during the reading process as prompted by the mobile app. (c) Automatic imaging of the chip by using the appuente app. (d) Root lengths as a function of time. Symbols (\(+\) \(\times \) \(\bigtriangleup \) \(\bigtriangledown \)) represent the results returned by the app in four different experiments. In the same plot, the average values of the analyzed roots at different times (solid line-circle) are compared to the measurements made by an expert using the original Plant chip method (solid line-squares).
Application case #3: LFIA chips for leptospirosis detection
Leptospirosis is a zoonotic infectious disease caused by pathogenic bacteria of the genus leptospira. Although leptospirosis is spread worldwide, it constitutes a special health problem in humid areas, such as tropical and subtropical regions, where most low-resource populations are found. Leptospirosis can be fatal, but effective and specific treatment is available, making early diagnostic valuable56. Recently, a LFIA for leptospirosis detection was developed55, which captures specific human IgM using a bacterial lysate as the antigen. The test showed high diagnostic performance, constituting a potential candidate for the implementation of early and accurate detection of leptospirosis. For this application case, the described LFIA strips were placed in custom 3D-printed plastic cassettes, with embedded alignment marks on top and individual ID codes at the bottom. The parts of the LFIA strip and the cassette are shown in Fig. 5a. The appuente platform was used to produce the corresponding mobile app, capable of guiding and prompting the user through the required steps (Fig. 5b). The app also allows online reading of the assay results, by using a custom made algorithm capable of classifying test as “Positive”, “Negative” or “Invalid”. The result is indicated in a final report on the phone screen, avoiding user subjectivity, specially relevant in cases where the intensity of test lines is weak. The performance of the appuente enhanced LFIA for leptospirosis was evaluated with four different analyte dilutions (n = 5). Results were read by an experienced operator using the naked eye and, immediately after, by using the mobile app running on two different smartphones. The classification effectiveness was 100%, i.e., the expert and the mobile app on each cellphone correctly identified all positive and negative cases. Moreover, the images produced by the mobile app were saved and shown to a panel of ten non-expert operators, who were asked to classify the test results. The average classification effectiveness was 79.5% (min 55% , max 90%). Also, the inter-observer agreement, measured with the Fleiss’s \(\kappa \) coefficient57 was 0.63, indicating intermediate agreement among the members of the panel. Examples of prototypes used for the experimental runs are shown in Fig. 5c.
Application case #3. (a) A 3D render of the LFIA parts (top panel) and its cassette (bottom panel). (b) Screenshots of the custom mobile app created using appuente, at six running steps: (1) Start new test, (2) scan ID code, (3) add sample, (4) chronometer, (5) automatic imaging and (6) result report. (c) Real examples of LFIA chips for positive (top) and negative (bottom) samples.
Discussion
The developed platform was successfully implemented and validated with three application cases, which in turn produced valuable information about the platform performance. Besides, it was evident how appuente improved several aspects of the assays, as described next. The grayscale detector showed a good performance in the color determination accuracy (percent error below 10%), with a simple algorithm that assumes a linear trend between black and white color references. As expected, the alternative that includes the full gray intensity palette for direct comparison with the unknown showed better performance. Even better performance should be possible after solving some remaining issues (regarding perspective and accurate indication of the beginning and end positions of the gray gradient) and allowing for a non-linear gradient to provide extra sensitivity. Thus, the grayscale detector showed that the color determination percent error can be very low (below 5% in all cases and around 1% in most cases), provided that proper algorithms and color references for calibration are used. Moreover, outstanding reproducibility was obtained, with error ranges below 5% in every case, and below 2% in most cases. Results obtained with this configuration were found to be independent from the operator, phone model, algorithm codification language, and light intensity. In the latter case, it should be noted that auto-shooting was not possible below 150 lx. Concerning cropping accuracy, native capabilities admit improvement, but they can also be significantly enhanced using image segmentation algorithms (Fig. 3e), as reported in Table 1.
Regarding the root elongation assay, it is readily observed in Table 2 how appuente reduces the number of experimental steps. Correspondingly, there is a clear decrease in assay complexity, time consumption, and the possibility of operator errors. As it can be seen in Fig. 4, the method showed very good agreement with the original Plant chip method considering both averages and standard deviations, with the absolute relative error (\(<{10}\)% in every case) being significantly smaller than the typical standard deviation of the assay. Thus, the approach proposed here is as accurate an reproducible as the validated assay formats, although significantly simpler and less time demanding. Furthermore, the combination of Plant chip and appuente allows for running time-dependent assays (Fig. 4d), which are time-consuming with Plant chip alone, and impossible to implement on traditional Petri dishes (where root elongation measurements can only be made at the end of the incubation). Live imaging provides rich information about the biological events that occur during plant development, particularly germination and root elongation, which can now be successfully followed without interrupting the experiment.
The platform was also tested with a POCT application, by implementing a novel LFIA for leptospirosis detection. Most of the platform capabilities were used: from the “no re-use” control, through user guidance, to smart-imaging of the chips. The performance of the mobile app in correctly classifying all the samples was compared against an expert, having both 100% effectiveness. When compared to users with no specific knowledge on LFIA tests, the average effectiveness was < 80% and the inter-operator agreement \(\kappa \) was 0.63, indicating intermediate agreement among the operators. This means that the user-independent reading method presented here, clearly contributes improving result interpretation, even when naked-eye reading is possible, given that the user subjectivity plays a significant role for the assay outcome.
It should be noted that not every qualitative LFIA test might need integration with appuente or mobile devices in general. Examples are test not prone to weak positives, or those where false negatives do not constitute a risk. In such cases, naked-eye qualitative reading might be enough. It should be stated though, that the other advantages listed in the introduction can still be found of value, and justify integration with appuente.
The developed platform allows for fast prototyping of mobile apps for the integration with IoT and microfluidic devices. It was conceived to provide all the advantages offered by mobile phones: user guidance, chip identification and tracking, user control, processing, report, cloud and IoT integration and imaging. In particular, appuente provides smart-imaging capabilities, in which the geometric variability is minimized. The platform is also prepared to deal with other sources of variability, like camera model and settings and ambient light intensity and color. Specifically, it can be achieved by providing proper references in the chip, as shown in Fig. 3a,b. The aforementioned elements allow not only qualitative determinations, like in application case #3, but also quantitative assessments, as demonstrated in application cases #1 and #2.
Future work will focus on adding iPhone compatibility to the platform, data analytics, and offline operation. Native cropping capabilities will also be improved (a workaround was presented in this article). Some challenging issues still remain within the field, namely: hygiene and cross-contamination (particularly when human samples are involved), phone anchoring, and the handling of light non-uniformity, especially for applications using smartphones without additional hardware. Concerning hygiene, it will depend on the specific biological marker under detection, as well as if the user is also the patient, so that several ad hoc solutions could be explored. Regarding light non-uniformity handling, video based approaches have been proposed58. Also, using the phone LED light has been proposed, although such approach fails in situations where ambient light sources are prevalent. Another alternative requires locating the chip between the dominant light source and the user41, although it is highly user-dependent. The last two approaches can be addressed by appuente, by automatically turning on the LED light when shooting and/or by providing additional guidance to the user. Moreover, specific features can be positioned within the chip to measure light gradients and correct or discard the result if uniformity is not proper. Finally, even if several aspect are to be improved, the here presented free, flexible and customizable tool for the integration of microfluidics and smartphones, will hopefully allow innovations and spreading of PONT-IoT applications.
Methods
Web and mobile apps
The web server runs on a LNMP stack (i.e. Linux, Nginx, MySQL, PHP), using an Ubuntu virtual machine (v20.04 LTS, ubuntu.com), a Nginx web server (v1.18.0, nginx.org), a MySQL database (v8, mysql.com), and PHP programming language (v8.0). GNU Octave (v5.2.0, gnu.org/software/octave) installed on the server also interacts with the rest of the elements. The website was developed using the Laravel framework (v8.12, laravel.com) for PHP, providing web services to connect with the mobile app using the JSON (json.org) format. The TCPDF library (tcpdf.org) is used to generate the ID codes. The web app requires no installation since it runs on a web browser, and can be accessed from appuente.com. Although this app runs ideally on any browser, it was mainly tested on Firefox (v88.0 mozilla.org) Chrome (v88.0.4324.96, google.com/chrome) and Safari (v15.0 apple.com/safari/). The mobile app was developed using the React Native framework (reactnative.dev), using JavaScript. An Android native module was developed in Java for image processing using the library openCV (v3.10, opencv.org). The web services send to and receive from the server the required information at every step. The app (appuente 1.0.0) can be downloaded from the Google Play Android store and from appuente.com. For both the web and mobile apps, GitLab (gitlab.com) was used for version control.
Grayscale detection chips and project setting
For the grayscale detector application example, two appuente projects were created, one for the chip shown in Fig. 3a, and the other for the chip in Fig. 3b. In both cases, a single smart-imaging step (Fig. 3c) was used. The “geometry and features” sections were filled with the desired physical dimensions of the chips, and proper algorithms were chosen. In the first case, containing three features, the algorithm (grayScaleDetect3.m, see Supplementary Information) averages the RGB pixels at each one of the features, converts it to the grayscale, and interpolates between the white reference (considered as 255) and the black reference (considered as 0) to find the level of gray of the unknown. In the second case, including two features, the algorithm (grayScaleDetect4.m, see Supplementary Information) converts the RGB values to grayscale, and finds the best match of the unknown within the reference gradient, containing all the possible gray values (Fig. 3f). In both cases, the features as defined in appuente were considerably larger than their physical counterparts, to make sure that no information is lost during the cropping procedure. Therefore, as a first step, the mentioned algorithms need to search for the physical feature, which is achieved using image segmentation. The chip geometry, with the embedded align marks and ID codes, was downloaded from the “Batch management” tab of the project and edited in Inkscape (v1.1, inkscape.org). Then, the chips were wax-printed (Xerox ColorQube 8580, Norwal, USA) on regular copy paper.
Grayscale detection experiments
The described grayscale detector was used in different sets of experiments keeping the following conditions constant, unless noted: ambient light intensity = 300 lx, feature side = 10 mm, unknown level of gray = 127, smartphone brand-model = TCL L10+, algorithm = grayScaleDetect4.m. In the two first sets of experiments (Table 1), the accuracy achieved when measuring different levels of gray was studied when grayScaleDetect3.m and grayScaleDetect4.m were used, respectively. Next, the accuracy obtained under variable ambient light intensity (measured with the ambient light sensor embedded in the same phone) was determined. After that, the accuracy obtained using different smartphone models and operators was assessed. Finally, the feature cropping accuracy was evaluated when the native capabilities of the platform were used, and when those capabilities were enhanced using the image segmentation algorithms previously mentioned. In every case, 5 repetitions were performed for each experiment (n = 5). In all the experiments, the only measure taken for light uniformity, was to place the chips between the light source and the operator to avoid shadowing41.
Root elongation chip preparation and project setting
For this application example, laser-cut PMMA top and bottom layers were used, as in the original device52. These transparent layers serve as the top and bottom walls of the channel where the roots grow, while allowing direct optic measurements. The middle layer was replaced by a black 3D-printed channel array, in order to enhance contrast and improve the performance of automatic identification and subsequent data analysis from (possibly low-resolution) images. The ID codes, obtained from the appuente environment, were printed on copy paper and glued to the chip. Germination paper was used as a substrate for the seeds (Lactuca sativa), the growing roots, and for water supply52.
After creating a new appuente project, the general data and chip geometry were set. Then, a custom algorithm was created and validated in a desktop version of GNU Octave. The algorithm (plantchip.m, see Supplementary Information) starts by identifying the locations of each of the 20 channels in the device, after which it looks for the relative position of the mark (or lack thereof) on each channel to output the corresponding physical root lengths. Algorithmic identification of the channels allows proper data analysis even in the presence of minor distortions in the image (as may be introduced by the smartphone’s camera lens), which can be expected due to the wide device compatibility of the appuente platform.
Root elongation chip experiments
Germination paper was first moistened with DI water (2.55 mL) and a seed was placed in every channel. Incubation temperature was kept at 24 ± \({2}^\circ {\hbox {C}}\). Assays were performed in fourfold (n = 80) for 120 h each. When possible, images were taken every 8 h using the mobile app. Illumination was achieved by using a 12 W LED panel as a backlight, attenuated with a thin, light blue cardboard sheet. The appuente platform determined the root lengths as described above. For comparison, the images captured by appuente were also used by an expert to measure the root lengths with the straight line tool from ImageJ (v1.53k, imagej.nih.gov/ij/). The global average and standard deviation were computed from both the app’s and the expert’s measurements. In all cases, channels with no detectable (\(< {5}\,{\hbox {mm}}\)) or inconsistent growth were removed from the analysis.
Leptospirosis LFIA chip preparation and appuente project setting
Plastic cassettes were 3D printed on PLA and the alignment marks were cut in self-adhesive vinyl. ID codes, obtained from the appuente environment, were printed on copy paper and glued to the case bottom. LFIA pads were assembled manually into the plastic cassettes, using the guides 3D printed at the bottom. Pads were mounted on two-sided tape in the following order: reaction pad (FF120HP, GE Life Sciences), conjugate pad (Fusion 5, GE Life Sciences) containing a solution of gold nanoparticles conjugated to goat anti-human IgM, sample and absorbent pads (CF3, Whatman, GE Life Sciences). Then, test and control lines were drawn onto the reaction pad by dipping silicone seals, respectively, into the leptospiral antigen preparation or a rabbit anti-goat IgG solution (Sigma-Aldrich) and stamping on the nitrocellulose membrane according to demarcated positions on the cassettes.
An appuente project was created and properly configured to guide the user through the required steps: sample load, 15-min incubation, and reading (see Fig. 5b). Geometric properties were also loaded on the project to match the cassette marks. A single feature was defined coincident with the black rectangle around the result window. An Octave algorithm (lfiaRead3.m, see Supplementary Information) was created, which (i) automatically finds the result window within the feature area, (ii) converts the RGB to a gray scale signal and takes average values in the longitudinal direction of the window, (iii) obtains a binary signal using an experimentally predefined threshold, (iv) looks for coincidences between the binary signal and the expected positions for the test and control lines, (v) uses this information to classify the test as positive, negative, or invalid.
Leptospirosis LFIA chip experiments
The chips for leptospirosis detection were tested with two real serum samples corresponding to a negative and a positive case of leptospirosis, classified at Laboratorio Nacional de Referencia de Leptospirosis of the Instituto Nacional de Enfermedades Respiratorias (INER) “Dr E. Coni”, Santa Fe, Argentina. Four different analyte concentrations were assessed in quintuplicate (n = 5): 1:100 dilution of the negative sample, 1:100, 1:200 and 1:500 dilutions of the positive sample. One hundred microliters of each were dispensed into the sample window of the chip and after 15 min, results were read and classified in two ways: naked eye reading by an expert and running the mobile app on different smartphones (TCL L10+ and Xiaomi Redmi 8). In addition, a non-expert panel was assembled to classify each sample using the images taken by appuente. The panel was composed of ten adults (ages: 27–39 y.o.) with degrees in natural sciences, who received a brief explanation of the task. The classification effectiveness of the expert, the app and the panel was calculated as the percentage of samples correctly classified (as negative, positive or invalid), compared to the known sample composition. The inter-operator agreement was measured using the Fleiss’s \(\kappa \) coefficient57, as a measure of the degree of concord among the members of the panel, accounting for chance. Complete agreement among the members of the panel means \(\kappa =1\) while agreement equal to the expected by chance, means \(\kappa =0\). The LFIA assays were evaluated and approved by the Bioethics Committee of FBCB, UNL (CE2019-37, Acta 05/19). All the experiments were performed in accordance with relevant national and international guidelines59,60. Informed consent was obtained from all participants and/or their legal guardians to use their samples for the research.
Data availability
The datasets analyzed in this article are available from the corresponding author on reasonable request. The GNU Octave algorithms used for this work are available in the Supplementary Information.
References
Nord, J. H., Koohang, A. & Paliszkiewicz, J. The internet of things: Review and theoretical framework. Expert. Syst. with Appl. 133, 97–108 (2019).
Čolaković, A. & Hadžialić, M. Internet of things (IoI): A review of enabling technologies, challenges, and open research issues. Comput. Netw. 144, 17–39 (2018).
Asghari, P., Rahmani, A. M. & Javadi, H. H. S. Internet of things applications: A systematic review. Comput. Netw. 148, 241–261 (2019).
Nauman, A. et al. Multimedia internet of things: A comprehensive survey. IEEE Access 8, 8202–8250 (2020).
Yetisen, A. K., Akram, M. S. & Lowe, C. R. based microfluidic point-of-care diagnostic devices. Lab on a Chip 13, 2210–2251 (2013).
Mabey, D., Peeling, R. W., Ustianowski, A. & Perkins, M. D. Diagnostics for the developing world. Nat. Rev. Microbiol. 2, 231–240 (2004).
Land, K. J., Boeras, D. I., Chen, X.-S., Ramsay, A. R. & Peeling, R. W. Reassured diagnostics to inform disease control strategies, strengthen health systems and improve patient outcomes. Nat. Microbiol. 4, 46–54 (2019).
Tseng, C.-C. et al. Recent advances in microfluidic paper-based assay devices for diagnosis of human diseases using saliva, tears and sweat samples. Sens. Actuators B Chem., 130078 (2021).
Noviana, E. et al. Microfluidic paper-based analytical devices: From design to applications. Chem. Rev. (2021).
Floris, A. et al. A prefilled, ready-to-use electrophoresis based lab-on-a-chip device for monitoring lithium in blood. Lab on a Chip 10, 1799–1806 (2010).
Wong, R. & Tse, H. Lateral Flow Immunoassay (Springer Science & Business Media, 2008).
Martinez, A. W., Phillips, S. T., Whitesides, G. M. & Carrilho, E. Diagnostics for the developing world: Microfluidic paper-based analytical devices (2010).
Tsao, C.-W. Polymer microfluidics: Simple, low-cost fabrication process bridging academic lab research to commercialized production. Micromachines 7, 225 (2016).
Sanjay, S. T., Dou, M., Sun, J. & Li, X. A paper/polymer hybrid microfluidic microplate for rapid quantitative detection of multiple disease biomarkers. Sci. Rep. 6, 1–10 (2016).
Bruus, H. Theoretical Microfluidics Vol. 18 (Oxford University Press, 2008).
Salentijn, G. I., Grajewski, M. & Verpoorte, E. Reinventing (bio) chemical analysis with paper. Anal. Chem. 90, 13815–13825 (2018).
Zhu, X. et al. A novel microfluidic device integrated with chitosan-modified capillaries for rapid Zikv detection. Micromachines 11, 186 (2020).
Wu, X. et al. Ms 2 device: Smartphone-facilitated mobile nucleic acid analysis on microfluidic device. Analyst (2021).
Martinez, A. W. et al. Simple telemedicine for developing regions: Camera phones and paper-based microfluidic devices for real-time, off-site diagnosis. Anal. Chem. 80, 3699–3707 (2008).
Vashist, S. K., Mudanyali, O., Schneider, E. M., Zengerle, R. & Ozcan, A. Cellphone-based devices for bioanalytical sciences. Anal. Bioanal. Chem. 406, 3263–3277 (2014).
Quesada-González, D. & Merkoçi, A. Mobile phone-based biosensing: An emerging “diagnostic and communication” technology. Biosens. Bioelectron 92, 549–562 (2017).
Kim, D. W., Jeong, K. Y. & Yoon, H. C. Smartphone-based medical diagnostics with microfluidic devices. In Smartphone Based Medical Diagnostics, 103–128 (Elsevier, 2020).
Schaumburg, F., Kler, P. A., Carrell, C. S., Berli, C. L. & Henry, C. S. USB powered microfluidic paper-based analytical devices. Electrophoresis 41, 562–569 (2020).
Lee, S. et al. Nutriphone: A mobile platform for low-cost point-of-care quantification of vitamin B12 concentrations. Sci. Rep. 6, 1–8 (2016).
Lillehoj, P. B., Huang, M.-C., Truong, N. & Ho, C.-M. Rapid electrochemical detection on a mobile phone. Lab on a Chip 13, 2950–2955 (2013).
Ainla, A. et al. Open-source potentiostat for wireless electrochemical detection with smartphones. Anal. Chem. 90, 6240–6246 (2018).
Barbosa, A. I., Gehlot, P., Sidapra, K., Edwards, A. D. & Reis, N. M. Portable smartphone quantitation of prostate specific antigen (PSA) in a fluoropolymer microfluidic device. Biosens. Bioelectron. 70, 5–14 (2015).
Halvorsen, C. P. et al. A rapid smartphone-based lactate dehydrogenase test for neonatal diagnostics at the point of care. Sci. Rep. 9, 1–8 (2019).
Choi, J. R. et al. An integrated paper-based sample-to-answer biosensor for nucleic acid testing at the point of care. Lab on a Chip 16, 611–621 (2016).
Kaarj, K., Akarapipad, P. & Yoon, J.-Y. Simpler, faster, and sensitive zika virus assay using smartphone detection of loop-mediated isothermal amplification on paper microfluidic chips. Sci. Rep. 8, 1–11 (2018).
Smith, G. T. et al. Robust dipstick urinalysis using a low-cost, micro-volume slipping manifold and mobile phone platform. Lab on a Chip 16, 2069–2078 (2016).
Wu, D. et al. A paper-based microfluidic dot-ELISA system with smartphone for the detection of influenza A. Microfluid. Nanofluidics 21, 43 (2017).
Ganguli, A. et al. Hands-free smartphone-based diagnostics for simultaneous detection of zika, chikungunya, and dengue at point-of-care. Biomed. Microdevices 19, 1–13 (2017).
Rajendran, V. K., Bakthavathsalam, P., Bergquist, P. L. & Sunna, A. Smartphone detection of antibiotic resistance using convective PCR and a lateral flow assay. Sens. Actuators B Chem. 298, 126849 (2019).
Kong, T. et al. Accessory-free quantitative smartphone imaging of colorimetric paper-based assays. Lab on a Chip 19, 1991–1999 (2019).
Lee, S. et al. Flexible opto-electronics enabled microfluidics systems with cloud connectivity for point-of-care micronutrient analysis. Biosens. Bioelectron. 78, 290–299 (2016).
Statista. Number of smartphone users worldwide. https://www.statista.com/statistics/330695/number-of-smartphone-users-worldwide/ (2021).
Chaplan, C. A., Mitchell, H. T. & Martinez, A. W. based standard addition assays. Anal. Methods 6, 1296–1300 (2014).
Hong, J. I. & Chang, B.-Y. Development of the smartphone-based colorimetry for multi-analyte sensing arrays. Lab on a Chip 14, 1725–1732 (2014).
Yetisen, A. K., Martinez-Hurtado, J., Garcia-Melendrez, A., da Cruz Vasconcellos, F. & Lowe, C. R. A smartphone algorithm with inter-phone repeatability for the analysis of colorimetric tests. Sens. Actuators B Chem. 196, 156–160 (2014).
Karlsen, H. & Dong, T. Smartphone-based rapid screening of urinary biomarkers. IEEE Trans. Biomed. Circuits Syst. 11, 455–463 (2017).
Lopez-Ruiz, N. et al. Smartphone-based simultaneous pH and nitrite colorimetric determination for paper microfluidic devices. Anal. Chem. 86, 9554–9562 (2014).
Baker, L. B. et al. Skin-interfaced microfluidic system with personalized sweating rate and sweat chloride analytics for sports science applications. Sci. Adv. 6, eabe3929 (2020).
Ho, N. R. et al. Visual and modular detection of pathogen nucleic acids with enzyme-DNA molecular complexes. Nat. Commun. 9, 1–11 (2018).
Mahato, K. & Chandra, P. based miniaturized immunosensor for naked eye alp detection based on digital image colorimetry integrated with smartphone. Biosens. Bioelectron. 128, 9–16 (2019).
Brangel, P. et al. A serological point-of-care test for the detection of IgG antibodies against Ebola virus in human survivors. ACS Nano 12, 63–73 (2018).
San Park, T., Li, W., McCracken, K. E. & Yoon, J.-Y. Smartphone quantifies salmonella from paper microfluidics. Lab on a Chip 13, 4832–4840 (2013).
Otal, E. H. et al. Open-source portable device for the determination of fluoride in drinking water. ACS Sensors 6, 259–266 (2021).
Tsagkaris, A. et al. A microfluidic paper-based analytical device (\(\mu \)pad) with smartphone readout for chlorpyrifos-oxon screening in human serum. Talanta 222, 121535 (2021).
Parker, R. W., Wilson, D. J. & Mace, C. R. Open software platform for automated analysis of paper-based microfluidic devices. Sci. Rep. 10, 1–10 (2020).
BBI Solutions. Novarum. https://bbisolutions.com/en/services/novarum-smartphone-reader.html/ (2021).
Macagno, J., Lescano, M. R. & Berli, C. L. Milli-channel array for direct and quick reading of root elongation bioassays. Ecotoxicol. Environ. Saf. 178, 51–57 (2019).
Wang, X. et al. Microfluidic-based fluorescent electronic eye with CdTe/CdS core-shell quantum dots for trace detection of cadmium ions. Anal. Chim. Acta 1131, 126–135 (2020).
Sun, F. et al. Smartphone-based multiplex 30-minute nucleic acid test of live virus from nasal swab extract. Lab on a Chip 20, 1621–1627 (2020).
Pujato, N. et al. Development of an accurate lateral flow immunoassay for enhanced diagnosis of human leptospirosis. Manuscr. progress (2021).
World Health Organization. Human leptospirosis: Guidance for diagnosis, surveillance and control. WHO/CDS/CSR/EPH 2002.23 (World Health Organization, 2003).
Fleiss, J. L. Measuring nominal scale agreement among many raters. Psychol. Bull. 76, 378 (1971).
Coleman, B., Coarsey, C., Kabir, M. A. & Asghar, W. Point-of-care colorimetric analysis through smartphone video. Sens. Actuators B Chem. 282, 225–231 (2019).
Guía para Investigaciones con Seres Humanos. Guidelines, Ministerio de Salud de la Repúbilca Argentina (2011).
UNESCO. International declaration on human genetic data. https://en.unesco.org/themes/ethics-science-and-technology/human-genetic-data (2003).
Acknowledgements
The authors gratefully acknowledge Fundación Bunge y Born, Argentina, for funding this research program through their Grants for Research of Infectious Diseases in Vulnerable Social Contexts. The authors also acknowledge the people who took part in the inter-operator study for the grayscale detector and the non-expert panel for the leptospirosis LFIA. Finally, the authors acknowledge Leandro M. Suero for his support with both the web and mobile apps.
Author information
Authors and Affiliations
Contributions
F.S. conceived the platform, carried out microfabrication, and wrote most of the manuscript. J.P.V. implemented both the web and mobile apps. G.S.G. participated in developing and testing the automatic identification and data capture algorithms for Plant chip assays in appuente. N.P. prepared the LFIA strips for leptospirosis detection, selected the suitable serum samples for appuente assessment, conducted the experiments and classified the results. J.M. adapted the Plant chip platform for use with appuente; directed, undertook experimental aspects and data processing of the Plant chip assays. P.A.K. took part in the conception of the platform and developed Plant chip support jointly with G.S.G and J.M. C.L.A.B. took part in the conception of the platform, supervised the whole process, and edited the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video 1.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schaumburg, F., Vidocevich, J.P., Gerlero, G.S. et al. A free customizable tool for easy integration of microfluidics and smartphones. Sci Rep 12, 8969 (2022). https://doi.org/10.1038/s41598-022-13099-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13099-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.