Abstract
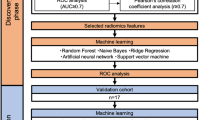
This study aimed to compare the predictive performance of different modeling methods in developing normal tissue complication probability (NTCP) models for predicting radiation-induced esophagitis (RE) in non–small cell lung cancer (NSCLC) patients receiving proton radiotherapy. The dataset was composed of 328 NSCLC patients receiving passive-scattering proton therapy and 41.6% of the patients experienced ≥ grade 2 RE. Five modeling methods were used to build NTCP models: standard Lyman–Kutcher–Burman (sLKB), generalized LKB (gLKB), multivariable logistic regression using two variable selection procedures-stepwise forward selection (Stepwise-MLR), and least absolute shrinkage and selection operator (LASSO-MLR), and support vector machines (SVM). Predictive performance was internally validated by a bootstrap approach for each modeling method. The overall performance, discriminative ability, and calibration were assessed using the Negelkerke R2, area under the receiver operator curve (AUC), and Hosmer–Lemeshow test, respectively. The LASSO-MLR model showed the best discriminative ability with an AUC value of 0.799 (95% confidence interval (CI): 0.763–0.854), and the best overall performance with a Negelkerke R2 value of 0.332 (95% CI: 0.266–0.486). Both of the optimism-corrected Negelkerke R2 values of the SVM and sLKB models were 0.301. The optimism-corrected AUC of the gLKB model (0.796) was higher than that of the SVM model (0.784). The sLKB model had the smallest optimism in the model variation and discriminative ability. In the context of classification and probability estimation for predicting the NTCP for radiation-induced esophagitis, the MLR model developed with LASSO provided the best predictive results. The simplest LKB modeling had similar or even better predictive performance than the most complex SVM modeling, and it was least likely to overfit the training data. The advanced machine learning approach might have limited applicability in clinical settings with a relatively small amount of data.
Similar content being viewed by others
Introduction
Radiotherapy is an essential treatment modality for patients with non-small cell lung cancer (NSCLC). However, radiation-induced toxicity, such as esophagitis, is of great concern to physicians as it lowers patients’ quality of life and may limit the dose-escalation targeted for better tumor control1. Accurate prediction of the normal tissue complication probability (NTCP) offers clinical guidelines for early intervention and facilitates physicians to develop a patient-specific treatment strategy. Moreover, there has been a growing interest in NTCP model-based patient selection for proton therapy2,3,4,5,6. For a given endpoint, modeling of NTCP can be performed using different approaches, ranging from the conventional analytic functions to the latest machine learning techniques7. And yet, there is no consensus on what is the best approach for NTCP modeling in the context of radiotherapy.
Traditional NTCP models were developed using analytic functions based on different assumptions of the dose–response relationship8,9,10,11,12,13. Among the analytic NTCP models, the Lyman–Kutcher–Burman (LKB) is the most well-known and widely used method11,12,13. In addition, it is the only NTCP model available in clinical treatment planning systems for biological evaluation14. In the standard LKB (sLKB) model, only the dosimetric factor was considered and the three-dimensional dose distribution was simplified into a single measure. It has been found the performance of the sLKB model was improved by adding one or more dose-modifying variables that account for the effect of non-dosimetric factors15,16,17. The LKB model with dose-modifying variables was denoted as the generalized LKB (gLKB) model hereafter. Another classic modeling technique to deal with various types of predictors is multivariable logistic regression (MLR), a statistical method to predict a binary outcome based on a set of independent variables18. Compared with the sLKB model which used only one dosimetric measure, the MLR model is capable of handling multiple dose-volume metrics, such as maximum dose, mean dose, Vx values (the percentage volume receiving a dose greater than x Gy), and Dx values (the minimum dose given to x% of the volume)19. The sLKB model showed a slightly lower discriminative ability than the MLR model20, but the difference between the performance of the generalized gLKB model and that of the MLR model has not been reported.
In the era of big data, the application of supervised machine learning in the treatment response modeling for radiotherapy has been rapidly growing. Support vector machine (SVM)21, a commonly used supervised machine learning algorithm, has shown promising accuracy in predicting lung radiation-induced pneumonitis22, local tumor control after stereotactic body radiation therapy for early-stage NSCLC patients23, and other clinical outcomes24,25. With the preset feature selection, the results of the comparison in performance between SVM and MLR were inconsistent26,27. It is of interest to compare their predictive power using a technique-specific feature selection scheme. Moreover, the advantage or the disadvantage of the complex SVM model over the simple LKB model remains unknown.
In this study, we aimed to identify the value of increasing the complexity of the NTCP model and using the advanced modeling technique in clinical practice by comparing the performance of LKB, MLR, and SVM methods for a data set concerning esophagitis.
Materials and methods
Data set
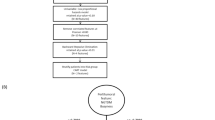
The data set consists of 328 NSCLC patients treated with passive-scattering proton therapy at The University of Texas MD Anderson Cancer Center during April 2006 to February 2012, and 136 (41.5%) patients experienced grade 2 or higher radiation-induced esophagitis20. This retrospective data collection was approved by our institutional review board (The University of Texas MD Anderson Cancer Center) with waivers for the patient’s informed consent. The prescription was 50–82.5 Gy [relative biological effectiveness (RBE)] in 25–37 fractions, with or without concurrent chemotherapy (CCRT). The clinical data were collected from the Epic system (Epic Systems Corporation), and the dosimetric data were extracted from the Eclipse treatment planning system (version 8.9, Varian Medical Systems) and has been converted to equivalent dose in 2-Gy fractions. Details of the patient characteristics, treatment, follow-up schedule, and esophagitis scoring were presented elsewhere20. The clinical characteristics included in the study were age (odds ratio (OR) = 0.99), sex (OR = 0.99), stage (OR = 2.23), and CCRT (OR = 4.76). There was no missing data on the outcome and predictors. All methods were performed in accordance with the relevant guidelines and regulations.
Modeling techniques
Standard Lyman–Kutcher–Burman modeling
In the LKB model, NTCP is calculated using the probit formula coupled with a generalized equivalent uniform dose (EUD) dose-volume histogram-reduction scheme28. The sLKB model described the dose–response relationship characterized by three parameters, in which n represents the volume effect, m denotes the slope of the NTCP curve at TD50, and TD50 is the dose tolerance corresponding to 50% complication risk (Supplementary material 1).
Generalized Lyman–Kutcher–Burman modeling
In the gLKB model, a dose-modifying factor was introduced to account for the effect of clinical features. The use of CCRT has been shown to be associated with a higher probability of developing grade ≥ 2 esophagitis in this cohort of patients20. Therefore, we substituted the parameter TD50 with a group-specific TD50s for the subgroup treated with (TD50y) or without (TD50n) CCRT, while keeping the same n, m for the entire cohort (Supplementary material 1). The dose-modifying factor is the ratio of TD50y and TD50n.
In the sLKB and gLKB modeling, the same methods were used to determine the best fits and the 95% confidence intervals (CI) for the parameters. The maximum likelihood estimation was used to determine the optimal values for the parameters n, m, TD50, TD50y, and TD50n. The 95% CIs for the fitted parameters were obtained using the profile likelihood method29,30.
Multivariable logistic regression
In the multivariable logistic regression model, the NTCP is modeled as a logit transformation of a linear function of several prognostic variables (Supplementary material 1). The candidate prognostic variables for each patient include four clinical and 17 dosimetric parameters. The clinical variables were sex, race, age, stage, and the use of CCRT. The dosimetric variables were EUD, the maximum dose (Dmax), the mean dose (Dmean), and the percentage volume receiving a dose higher than x Gy(RBE) (x ranges from 10 to 75 in increments of 5). The results of the correlation between variables are available elsewhere20. Table 1 listed the encoding values of the clinical variables and the dosimetric values. The EUD was calculated using the parameter n derived for the sLKB model.
For the stepwise feature selection, a forward stepwise algorithm was adopted with the likelihood ratio test. Starting with a null model, the stepwise regression added or removed one variable at each step based on the p value for the likelihood ratio test. The variable selection process stopped if no variable can be added or removed. Stepwise-MLR is referred to as the multivariable logistic regression model using the stepwise feature selection in the following sections.
The LASSO method profiles out the insignificant variables by penalizing the regression coefficients of the variables and shrinks some of them to zero. As such, variables with non-zero coefficients were selected. The degree of penalty is controlled by the regularization parameter λ (Supplementary material 1). A fivefold cross-validation was performed to determine the optimal value of λ. The multivariable logistic regression model using LASSO regulation for feature selection is denoted as LASSO-MLR.
Support vector machine
For binary classification, the SVM searches for a hyperplane that maximizes the margin between the two classes. In this study, a radial kernel function was used to map the data from a low-dimensional space into a high-dimensional feature space where the non-linear boundary became a linear boundary. Two hyperparameters were included in the SVM model. C is the regularization parameter that trades off the margin width against the fitting error, and γ is the parameter in the kernel function that controls the overfitting. The NTCP estimates were then generated from a decision function using a logit transformation (Supplementary material 1).
A grid-research was performed in the space of (log2C = − 5:2:15, log2γ = − 3: − 2: − 15) to identify the optimal parameter pair (C, γ) using fivefold cross validation31. The whole data set was divided into 5 subsets of approximately equal size. In each iteration, a model was trained in 80% of the sample using a pair of (C,γ) and tested in the remaining 20% subsets. The procedure was repeated 5 times. The selection criterion was the averaged value of the area under the receiving operator curve (AUC) computed in the testing samples. All grid points of (C, γ) were tested and the one yielding the maximum AUC was picked. The best parameters of C and γ were then used in the feature selection. The optimal subset of features was determined by sequentially adding features based on the criterion of the fivefold cross-validated AUC until there is no improvement. In the SVM modeling, a total of 24 variables were included, where stage was partitioned as a four-category attribute (1,0,0,0), (0,1,0,0), (0,0,1,0), and (0,0,0,1).
Bootstrapping for feature selection and model validation
The feature selection procedures in Stepwise-MLR, LASSO-MLR, and SVM were repeated in 1000 bootstrap samples drawn from the replacement of the original sample. The features with a selection frequency of greater than 80% were picked for the final models. Another threshold was applied for the selection frequency if the highest selection frequency was lower than 80%. With the selected features, models were built on the original sample and the performance was evaluated (apparent performance). For SVM, the grid-search step was repeated to find the best pair (C, γ) within the selected feature space.
We also used the bootstrap approach to internally validate the predictive performance of different NTCP models. Model fitting including feature selection and coefficient estimation was repeated on each bootstrap sample using the specific modeling technique. From the resulting model, the performance was evaluated in the bootstrap sample (bootstrap performance) and the original sample (test performance). The optimism was calculated as the difference between bootstrap performance and test performance. A lower value of optimism denotes a lower level of overfitting. The optimism should be corrected to reflect the stable and unbiased estimation of the model performance in future data. The optimism-corrected performance was then obtained by subtracting the optimism from the apparent performance.
The performance of the model was assessed using the Negelkerke R2, AUC, and Hosmer–Lemeshow (HL) test. Negelkerke R2 is an overall measure to quantify the variability explained by a model. AUC is used to indicate how well the model classifies patients into different risk groups. HL test assessed the calibration of the model, and a p value of greater than 0.05 indicates good agreement between predicted probability and observed risk.
Software
Model development and data analysis were performed in MATLAB (version R2016b, MathWorks, Inc.). For the SVM modeling, we used the MATLAB version of the freely distributed package LIBSVM32.
Results
In the gLKB model, the optimal values of n and m [n = 0.23 (95% CI: 0.09–0.47), m = 0.54 (95% CI: 0.41–0.74)] are slightly deviated from the values of sLKB model [n = 0.24 (95% CI: 0.10–0.49), m = 0.51 (95% CI: 0.37–0.70)]. The TD50 for patients treated with CCRT (TD50y) was 42.17 Gy (RBE) [95% CI: 31.76–52.53 Gy (RBE)], while it was 57.84 Gy (RBE) [95% CI: 42.65–79.75 Gy (RBE)] for patients treated without CCRT. Therefore, the dose-modifying factor is 1.4. Compared with the sLKB model, the gLKB model showed a better fit of the data, as can be deduced from the the increased value of optimism corrected Negelkerke R2 and AUC.
Figure 1 shows the selection frequency for all the variables in the 1000 bootstrap samples with the modeling techniques Stepwise-MLR, LASSO-MLR, and SVM. For the variables with greater than 80% frequency, stepwise selection and LASSO had two common variables: CCRT (stepwise: 80.8%, LASSO: 99.7%) and EUD (stepwise: 93.6%, LASSO: 96.7%). Compared with stepwise selection, LASSO tended to include more variables and it selected V75 with a frequency of 92.4%. Using 60% as a threshold, the variables selected by SVM were CCRT (66.3%) and EUD (69.2%), matching the variable selection based on the stepwise approach. Table 2 presents the regression coefficients for the two MLR models using the selected variables and the optimal values of C and γ for the SVM model.
A summary of the apparent performance, bootstrap performance, optimism, and the optimism-corrected performance of all the models are listed in Table 3. The optimism-corrected Negelkerke R2 and AUC values of the LASSO-MLR were also higher than the other models. Stepwise-MLR showed a marginal improvement in the optimism-corrected-AUC (0.797) compared with the gLKB model (0.796). A similar predictive performance was found for the SVM and sLKB model with regard to the optimism-corrected Negelkerke R2 and AUC values. Both of the optimism-corrected Negelkerke R2 values of the SVM and sLKB models were 0.301, and the optimism-corrected AUC values were 0.783 and 0.784 for the sLKB and SVM models respectively. Moreover, there were minimal differences between SVM and gLKB, Of note, the advanced SVM modeling technique was more prone to overfitting, as indicated by its highest optimism (0.039 and 0.015 for Negelkerke R2 and AUC respectively). By contrast, the lowest optimism was observed in the conventional sLKB model.
Figure 2 displays the receiving operator curves and the calibration plots of all the models. A comparison of receiving operator curves of all the models (Fig. 2a) showed that sLKB was less accurate in detecting true positive cases at a higher threshold as compared with the other models. The predictions of all the models significantly agreed with the observed outcome, as demonstrated by the non-significant HL tests. The loess smoother of LASSO-MLR was closer to the ideal line, indicating a better calibration performance.
Receiver operator curves (a) and the calibration plot of actual outcome vs. predicted probability for the sLKB (b), gLKB (c), Stepwise-MLR (d), LASSO-MLR (e), and SVM (f) models. Results of the Hosmer–Lemeshow test are displayed in the lower right part of each plot. The open circles are the observed frequency in each group by the deciles of mean predicted probabilities. The dashed line represents the ideal prediction and the solid line is the loess fit for the output of the model.
Discussion
In this study, we investigated and internally validated the predictive performance of NTCP models developed for radiation-induced esophagitis in NSCLC patients receiving proton radiotherapy using different modeling techniques ranging from conventional analytic solution, logistic regression, to advanced machine learning. Of the five NTCP models, LASSO-MLR showed the best fit for the data. The sLKB model had the lowest optimism. SVM modeling resulted in NTCP models with a good apparent performance but with the highest optimism. Our results highlighted the non-inferiority of the conventional LKB modeling to the advanced SVM modeling in the discriminative ability and the accuracy of the predicted probability.
The predictive performance of the NTCP model was improved when more factors were considered. For the LKB modeling, our results of the comparison of the apparent performance were in line with the studies conducted by Peeter et al.15 and Defraene et al.33, who found the modified LKB model with both the dosimetry and clinical factors gave a significantly better fit than the standard LKB model with dosimetry alone for different endpoints. Previous studies also reported similar findings in multivariable logistic regression modeling that the model with more predictors had a higher performance than the one with fewer predictors34,35. However, the comparison of the optimism-corrected performance has not been investigated. Of note is that the optimism-corrected performance was considered a less unbiased assessment of the model in the future data than the apparent performance36. It is more likely to have a higher optimism by including more variables, thus resulting in lower optimism-corrected performance. Xu et al.35 demonstrated that an all-variable logistic regression model was susceptible to overfitting, which suggested a robust variable selection approach. As shown in Table 3, thought the optimisms of gLKB and LASSO-MLR were higher than those of their counterparts using the same modeling technique, the optimism-corrected performance of gLKB and LASSO-MLR were still higher.
The advanced SVM modeling technique did not offer improvement in our prediction of the dose–response relationship over the conventional LKB and logistic regression models. Moreover, the predictive performance of the SVM model was slightly poorer than the Stepwise-MLR that used the same predictors. The results would probably be explained by the γ parameter of the SVM model. In addition to the regulation parameter C, the behavior of the SVM model using the radial basis function kernel is very sensitive to the γ parameter. A large value of the γ parameter means that the radius of the influence area of the support vectors only includes the support vector itself and the regulation from the parameter C is unable to prevent overfitting. A small value of the γ parameter denotes an overly constrained model that cannot capture the complexity of the data, resulting in a model with a similar performance as the model assuming the linearity of the predictors. The γ parameter in the optimized SVM model has a relatively low value of 2−13. Therefore, the SVM model performed comparably to the logistic regression model under the same feature selection and even did not outperform the simplest analytic LKB model. Lynam et al.26 also reported that logistic regression performed as well as the machine learning algorithm in their study population when a small number of predictors were considered. Currently, there is no golden standard for variable selection and fewer significant variables are preferred in clinical practice. The nonlinear machine learning approaches may prove more powerful in the context of a larger dataset with more variables, such as the study of genomics and radiomics. In addition, the signal-to-noise of the clinical data may be lower. Therefore, the advanced machine learning modeling technique may have limited applicability in the current clinical setting.
For clinical practitioners, intuitive interpretability and ease of implementation were two important aspects when choosing the “best” model. The parameter n in the LKB model provides insight into whether the low dose region or the high dose region matters. We can also derive a dose constraint in terms of EUD from the LKB and logistic regression models based on the monotonically increasing functions of EUD, leading toward individualized treatment planning. In contrast, the mapping of input data and the output predictions in the SVM model is considered a “black box”, which prevents practitioners from better understanding the data and quantifying the effect of the dosimetric and non-dosimetric predictors. Incorporating the decision-making platform with a large amount of information given by the treatment planning system and the patient information system such as Epic would facilitate the adoption of a machine learning algorithm in clinical decision-making. In terms of performance and utility, LASSO-MLR was the best modeling technique in the current context.
One limitation of the current study is that only a few clinical predictors had been enrolled in the investigation. A further study would be different NTCP modeling techniques incorporated with functional imaging features and biological markers. In addition, larger and external data sets are needed to verify the findings of this study.
Conclusion
In the context of classification and probability estimation for predicting the NTCP of radiation-induced esophagitis in NSCLC patients receiving proton radiotherapy, a multivariable logistic regression model developed with LASSO provided the best predictive results. The simplest LKB modeling using the analytic function had similar or even better predictive performance than the most complex SVM modeling, and it was least likely to overfit the training data. The advanced machine learning approach might have limited applicability in clinical settings with a relatively small amount of data.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Verma, V., Simone, C. B. 2nd. & Werner-Wasik, M. Acute and late toxicities of concurrent chemoradiotherapy for locally-advanced non-small cell lung cancer. Cancers (Basel) 9, 120 (2017).
McNamara, A. L. et al. Perspectives on the model-based approach to proton therapy trials: A retrospective study of a lung cancer randomized trial. Radiother. Oncol. 147, 8–14 (2020).
Scherman, J. et al. Incorporating NTCP into randomized trials of proton versus photon therapy. Int. J. Part. Ther. 5, 24–32 (2019).
Brodin, N. P. et al. A quantitative clinical decision-support strategy identifying which patients with oropharyngeal head and neck cancer may benefit the most from proton radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 104, 540–552 (2019).
Langendijk, J. A. et al. Selection of patients for radiotherapy with protons aiming at reduction of side effects: The model-based approach. Radiother. Oncol. 107, 267–273 (2013).
Widder, J. et al. The quest for evidence for proton therapy: Model-based approach and precision medicine. Int. J. Radiat. Oncol. Biol. Phys. 95, 30–36 (2016).
El Naqa, I. A Guide to Outcome Modeling in Radiotherapy and Oncology: Listening to the Data (CRC Press, 2018).
Schultheiss, T. E., Orton, C. G. & Peck, R. Models in radiotherapy: Volume effects. Med. Phys. 10, 410–415 (1983).
Källman, P., Ågren, A. & Brahme, A. Tumour and normal tissue responses to fractionated non-uniform dose delivery. Int. J. Radiat. Biol. 62, 249–262 (1992).
Jackson, A., Kutcher, G. & Yorke, E. Probability of radiation-induced complications for normal tissues with parallel architecture subject to non-uniform irradiation. Med. Phys. 20, 613–625 (1993).
Lyman, J. T. Complication probability as assessed from dose-volume histograms. Radiat. Res. Suppl. 104, S13–S19 (1985).
Kutcher, G. J. & Burman, C. Calculation of complication probability factors for non-uniform normal tissue irradiation—The effective volume method. Int. J. Radiat. Oncol. Biol. Phys. 16, 1623–1630 (1989).
Kutcher, G. J. et al. Histogram reduction method for calculating complication probabilities for three-dimensional treatment planning evaluations. Int. J. Radiat. Oncol. Biol. Phys. 21, 137–146 (1991).
Allen Li, X. et al. The use and QA of biologically related models for treatment planning: Short report of the TG-166 of the therapy physics committee of the AAPM. Med. Phys. 39, 1386–1409 (2012).
Peeters, S. T. H. et al. Rectal bleeding, fecal incontinence, and high stool frequency after conformal radiotherapy for prostate cancer: Normal tissue complication probability modeling. Int. J. Radiat. Oncol. Biol. Phys. 66, 11–19 (2006).
Tucker, S. L. et al. Incorporating single-nucleotide polymorphisms into the lyman model to improve prediction of radiation pneumonitis. Int. J. Radiat. Oncol. Biol. Phys. 85, 251–257 (2013).
El Naqa, I. et al. Modeling of normal tissue complications using imaging and biomarkers after radiation therapy for hepatocellular carcinoma. Int. J. Radiat. Oncol. Biol. Phys. 100, 335–343 (2018).
McDonald, J. H. Handbook of Biological Statistics Vol. 2 (Sparky House Publishing, Baltimore, 2009).
El Naqa, I. et al. Multivariable modeling of radiotherapy outcomes, including dose-volume and clinical factors. Int. J. Radiat. Oncol. Biol. Phys. 64, 1275–1286 (2006).
Wang, Z. et al. Lyman–Kutcher–Burman normal tissue complication probability modeling for radiation-induced esophagitis in non-small cell lung cancer patients receiving proton radiotherapy. Radiother. Oncol. 146, 200–204 (2020).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Chen, S. et al. Investigation of the support vector machine algorithm to predict lung radiation-induced pneumonitis. Med. Phys. 34, 3808–3814 (2007).
Klement, R. J. et al. Support vector machine-based prediction of local tumor control after stereotactic body radiation therapy for early-stage non-small cell lung cancer. Int. J. Radiat. Oncol. Biol. Phys. 88, 732–738 (2014).
Pella, A. et al. Use of machine learning methods for prediction of acute toxicity in organs at risk following prostate radiotherapy. Med. Phys. 38, 2859–2867 (2011).
Mizutani, T. et al. Survival time prediction after radiotherapy for malignant glioma patients based on clinical and DVH features using support vector machine. Int. J. Radiat. Oncol. Biol. Phys. 99, E698–E699 (2017).
Lynam, A. L. et al. Logistic regression has similar performance to optimised machine learning algorithms in a clinical setting: Application to the discrimination between type 1 and type 2 diabetes in young adults. Diagn. Progn. Res. 4, 6 (2020).
El Naqa, I. et al. Predicting radiotherapy outcomes using statistical learning techniques. Phys. Med. Biol. 54, S9–S30 (2009).
Niemierko, A. Reporting and analyzing dose distributions: A concept of equivalent uniform dose—Response. Med. Phys. 24, 1325–1327 (1997).
Cox, D.R. & Snell, E.J. Analysis of Binary Data. 2nd Edn (Chapman and Hall/CRC, London, 1989).
Roberts, S. A. & Hendry, J. H. The delay before onset of accelerated tumour cell repopulation during radiotherapy: A direct maximum-likelihood analysis of a collection of worldwide tumour-control data. Radiother. Oncol. 29, 69–74 (1993).
Keerthi, S. S. & Lin, C.-J. Asymptotic behaviors of support vector machines with Gaussian kernel. Neural Comput. 15, 1667–1689 (2003).
Chang, C.-C. & Lin, C.-J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. (TIST) 2, 1–27 (2011).
Defraene, G. et al. The benefits of including clinical factors in rectal normal tissue complication probability modeling after radiotherapy for prostate cancer. Int. J. Radiat. Oncol. Biol. Phys. 82, 1233–1242 (2012).
Cella, L. et al. Development of multivariate NTCP models for radiation-induced hypothyroidism: A comparative analysis. Radiat. Oncol. 7, 224 (2012).
Xu, C.-J. et al. Impact of statistical learning methods on the predictive power of multivariate normal tissue complication probability models. Int. J. Radiat. Oncol. Biol. Phys. 82, e677–e684 (2012).
Steyerberg, E. W. Clinical Prediction Models (Springer, 2019).
Acknowledgements
The University of Texas MD Anderson Cancer Center was supported in part by the National Cancer Institute Cancer Center Support Grant P30 CA016672. This study was supported in part by the National Key Research and Development Program of China (Grant 2016YFC0105409). We acknowledge financial support from the MD Anderson, Texas Advanced Computing Center, Oden Institute for Computational and Engineering Sciences initiative in Oncological Data and Computational Science.
Author information
Authors and Affiliations
Contributions
X.Z. conceptualized and designed the study. M.C. wrote the main manuscript text and prepared all the figures and tables. M.C., Z.W., S.J., and J.S. collected and analyzed the data. Q.N., J.C., and Z.L. reviewed the clinical plan and graded the radiation-induced esophagitis. L.W., N.S., G.B., S.F., C.X., J.C., R.Z., and X.Z. edited the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, M., Wang, Z., Jiang, S. et al. Predictive performance of different NTCP techniques for radiation-induced esophagitis in NSCLC patients receiving proton radiotherapy. Sci Rep 12, 9178 (2022). https://doi.org/10.1038/s41598-022-12898-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-12898-8
This article is cited by
-
SAPT: a synchrotron-based proton therapy facility in Shanghai
Nuclear Science and Techniques (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.