Abstract
Alignment-free methods for sequence comparison and phylogeny inference have attracted a great deal of attention in recent years. Several algorithms have been implemented in diverse software packages. Despite the great number of existing methods, most of them are based on word statistics. Although they propose different filtering and weighting strategies and explore different metrics, their performance may be limited by the phylogenetic signal preserved in these words. Herein, we present a different approach based on the species-specific amino acid neighborhood preferences. These differential preferences can be assessed in the context of vector spaces. In this way, a distance-based method to build phylogenies has been developed and implemented into an easy-to-use R package. Tests run on real-world datasets show that this method can reconstruct phylogenetic relationships with high accuracy, and often outperforms other alignment-free approaches. Furthermore, we present evidence that the new method can perform reliably on datasets formed by non-orthologous protein sequences, that is, the method not only does not require the identification of orthologous proteins, but also does not require their presence in the analyzed dataset. These results suggest that the neighborhood preference of amino acids conveys a phylogenetic signal that may be of great utility in phylogenomics.
Similar content being viewed by others
Introduction
It is a well-established fact that different genes from the same set of organisms often lead to different phylogenetic trees1. That happens even with mitochondrial-encoded genes2,3, despite the fact that such genes are inherited together without recombination, and the risk of confusing orthologous with paralogous sequences is non-existent. If, in addition, phenomena such as horizontal gene transfer, recombination, unrecognized paralogy, and highly variable rates of evolution are in place, the task of reconstructing accurate phylogenetic topologies can be seriously compromised. Not surprisingly, species phylogenies derived from comparison of single genes are seldom consistent with each other. To overcome this problem, two strategies are currently used when resolving phylogenies based on multiple alignments. In the so-called supermatrix approach, individual aligned genes or proteins are concatenated into a supermatrix, which is then subjected to phylogenetic analyses using either maximum likelihood or Bayesian inference4. In the alternative supertree method, gene or protein data sets are analyzed separately. Afterwards, the trees derived from these independent analyses are used to infer a single joined phylogeny5,6. Each of these alternatives has its own strengths and weaknesses, which has led to extensive discussions regarding the best strategy to conduct phylogenetic analyses of sequence data from multiple genes or proteins7,8,9,10. Nevertheless, both approaches have in common that they are time consuming, and they often require manual intervention. On the other hand, among the diverse sources of error in molecular phylogenies, incorrect sequence alignments rank high11,12. Therefore, those methods based on sequence alignments are prone to artefacts when used in phylogenomics13. Indeed, a number of previous studies have shown that the alignment method can have a considerable impact on tree topology14,15,16,17,18. Although attempts have been made to deal with multiple sequence alignment uncertainty during phylogeny reconstruction19, a satisfying and computationally tractable way to deal with alignment uncertainty is still lacking. Alignment artefacts have become even a bigger problem in the era of phylogenomics, where thousands of genes are automatically analyzed without accounting for alignment uncertainty14.
With the advent of modern genome sequencing techniques, it is now possible to consider phylogeny inference based on total genome sequences. However, given that most genomes contain millions of nucleotides, the standard approach based on positional homology (where each column from a multiple sequence alignment is considered as a homologous character) represents a daunting challenge that becomes impractical. Consequently, alternative approaches to compare whole genomes have been proposed. Thus, gene arrangement20, gene content21,22, protein domain-abundance23 and presence/absence of protein folds24, are all strategies tha have been explored to compare whole genomes. More recently, a wide number of alignment independent methods to compare sequences have been developed, and their utility in phylogenomics has been evaluated25. Thus, the so-called alignment-free approach include methods based on words-counting26,27, some of which implement diverse strategies to discriminate signal from noise28,29,30,31,32,33. Other published methods are based on matching statistics (i.e., they compute the length of common substrings with or without allowing mismatches)34,35, information theory36,37, splits driven by common subsequences38, or even based on micro-alignments39,40.
In this study, we describe a new and fast method for generating molecular phylogenies using multiple proteomes or protein-coding genomes. This method, which does not require sequence alignment or the identification of orthologous proteins, is based on a rationale previously unexplored in the context of phylogeny: the preference of each amino acid to be surrounded by other amino acids41,42,43. These species-specific preferences seem to posse a phylogenetic signal enough to reconstruct accurate tree topologies, even when the proteins analyzed from each species are functionally unrelated to the proteins selected from the other species.
Results
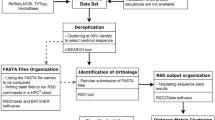
The new method, which is presented in detail in the Methods section, is briefly outlined in Fig. 1. As a proof of concept, the phylogenetic relationships of 11 species of bovids were addressed using their protein-coding mitogenomes and the new method described below, hereinafter referred to as Env-NJ. The topology of the reconstructed tree is shown in Fig. 2. This topology fully matches that of the tree inferred using traditional alignment-based methods (reference tree), which reflect the accepted phylogeny for this groups of bovids. For comparative purposes, the same mitogenomes were employed with the following alignment-free tree-building packages: Feature Frequency Profile (FFP)26, alfpy (a stand-alone Phyton application that implements different approaches as well as different metrics to assess vectors distances)27, CVTree32, ALFRED-G35, SANS-serif38 and Prot-SpaM39. In all cases, the resulting trees were compared to the reference tree. Table 1 shows the corresponding normalized Robinson-Foulds distances (nRF).
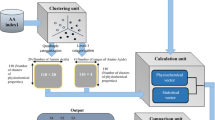
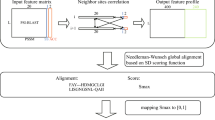
Encoding species as vectors. (A) For the sake of concreteness, we will focus on the sequence environment around methionine residues using a radius of 10 residues. Thus, a given proteome can be characterized by a matrix whose elements (mij) provide the absolute frequency of the amino acid i at the position j in the environment of methionine residues. For instance, in this example, the element m2,2 gives the number of arginines found 9 residues away (toward the N-terminal) from any methionine residue. (B) Now, considering not just methionine but the 20 proteinogenic amino acids, the protein-coding genome of a given species of interest can be characterized by a set of 20 square matrices of order 20 or, equivalently, by a vector, \(u\in {U}^{8000}\), of dimension 8000. It should be noted that, when coding species as vectors, the dimension of these vectors will depend on the radius chosen. Thus, in general, \(u\in {U}^{n}\), where \(n=800 radius\). (C) In this way, each vector is used to represent an organism (its protein-coding genome) within a set evolutionarily related species. (D) The direction, rather than the norm, of these vectors reflect the preference of the different amino acids at the different positions of the sequence environments. The work of encoding a set of genomes in a set of vectors is conveniently carried out by the function otu.space() from the R package accompanying the current paper.
We next challenged the Env-NJ method with a larger, more diverse, and more controversial dataset consisting in the mitogenomes of 34 mammalian species spanning 13 orders. The phylogenetic relationships between the organisms of this set were first analyzed by Reyes and coworkers using a maximum likelihood approach44. Figure 3A reproduces the topology of the tree obtained by these authors. Nevertheless, since alternative hypotheses for the phylogeny of this set of species have been proposed by different authors29,36, in order to adopt a reference tree we resorted to the community resource VerLife45 to draw the topology of the tree (Fig. 3B) that relates the species under study according to this source45. As shown in Fig. 3, the Env-NJ method yields a credible phylogeny where primates, carnivores, cetartiodactyls and perissodactyls are some of the well-established mammalian lineages that appear as uninterrupted groupings within the Env-NJ tree. In order to obtain a more quantitative comparison, we next built 14 trees using the same dataset and different alignment-free tree building approaches. Afterwards, the normalized Robinson-Foulds symmetric difference between these trees and the reference tree was computed (Table 2). As it can be observed in this table, the Env-NJ with the Jensen-Shannon metrics provided the best result, understood as the one that provided the lowest Robinson-Foulds distance to the reference tree.
Comparison of phylogenetic tree topologies. The same mitogenomes of 34 mammalian species spanning 13 orders were employed with different tree-building methods. (A) Reproduces the topology of the tree obtained by Reyes et al. 2000 using a maximum likelihood approach based on multiple sequence alignments of nucleotides. (For protein genes, only first and second positions, P12, of the codons were considered. In addition, the ND6 gene encoded by the L-strand was also excluded). (B) The topology of the tree provided by VerLife45 for the relevant species is drawn. (C) The topology of the tree obtained using Env-NJ with a radius of 46 and the Jensen-Shannon metric is shown. (D) The topology of the tree constructed using singular value decomposition (SVD) to analyzed 4-mer string frequencies derived from unaligned sequences is also shown.
At this point, we reason as follows. If the amino acid neighborhood preference in sequence environments is a species feature, then perhaps it may be possible to reconstruct phylogenetic relationships using non-orthologous proteins sets. To explore the potential of the Env-NJ method to provide such an achievement, we selected five species (three animals and two plants) whose phylogenetic relationships are well established. For each species a random set consisting of 180 proteins was selected, with the only restriction that no protein belonging to this set could be homologous to any of the proteins belonging to the remaining species. These random sets of non-orthologous protein sequences were used to generate the Env-NJ tree. To compare the performance of our method with that of previously proposed alignment-free approaches, the same dataset was subjected to 7 alternative methods, including Prot-SpaM39, W-metric27,33, FFP26, CVTree32, ALFRED-G35, normalized compression distances (NCD)27 and SANS-serfi38. As shown in Fig. 4, the new methodology yielded the correct tree topology even when non-orthologous proteins were employed, and under this specific conditions it seems to outperform other tree building methods.
Alignment-free trees built using a dataset of non-orthologous protein sequences. For each species a random set consisting of 180 proteins was selected, with the only restriction that no protein belonging to this set could be homologous to any of the proteins belonging to the remaining species. These random sets of non-orthologous protein sequences were used to generate the corresponding trees.
Finally, all proteins encoded by the full genome of 11 plant species (Table 3) were used as input data to assess the performance of Env-NJ and other alignment-free alternative methods. The results are summarized in Table 4. As it can be observed in these tables, the Env-NJ approach was able to analyze 425,115 proteins accounting for 169,094,374 amino acids in less than 1 min, providing a reliable phylogeny.
Discussion
Traditionally, the starting point to construct a molecular phylogeny has been identifying and gathering a set of evolutionary related (orthologous) sequences. However, before using these sequences to build a tree, it is important to ensure that each nucleotide or amino acid in each sequence is compared only with the corresponding homologous nucleotide or amino acid in the other sequences, what is referred to as positional homology. This preliminary task, that is one of the trickiest parts of the whole phylogenetic reconstruction process, is performed by aligning the sequences to one another. It should be noted that none of the frequently used alignment programs is capable of consistently producing perfect alignments, even when moderately divergent sequences are employed46. For that reason, it is always important to check the alignment quality before continuing with the phylogenetic reconstruction procedure. Obviously, this protocol is not scalable to phylogenomics. Since most genomes contain millions of sequence characters, these traditional methods based on positional homology comparisons, carried out over ambiguously resolved large-scale alignments, are unbusinesslike14. Thus, it seems that the problem of phylogenomic reconstruction based on site-evolution has no solution in the near future. To overcome this problem, different approaches have been explored.
One of these approaches to whole genomes phylogenetic analysis has focused on the ordering of the genes along the chromosomes, others have resorted to the gene content as its primary data. Since proteins (gene products) are modular and many of them are mosaics of diverse domains47, phylogenomic strategies based on domain-abundance or the presence/absence of protein folds may performe even better than those focused on genes23,24. The use of gene-order and gene/fold-content data in the context of phylogeny is the subject of important research efforts. However, there remain important challenges. Thus, mapping a full genome is a demanding task. Furthermore, the posterior analysis of the annotated genome is computationally expensive and time consuming because of the extreme mathematical complexity of gene orders. For instance, for a chromosome with n distinct single-copy genes, the number of possible states is \({2}^{n-1}\left(n-1\right)!\)48. This computational burden means that all reconstruction methods face a considerable challenge, even on small datasets consisting of only a few genomes. Furthermore, in these approaches the information contained into a genome is largely simplified, in the sense that point mutations are completely ignored, that is, these methods somehow make use of lossy data compression, so that relevant information contained in a genome is not used to infer its evolutionary history.
In this report, we describe a phylogenetic approach based on sequence environments, that may be valuable for the future development of new methods for generating phylogenies from whole genomes without resorting to lossy data compression. The protein-coding genomes of the set of organisms being analyzed are converted into a matrix, where each column vector represents a species. More concretely, these vectors represent the species-specific amino acid neighborhood preferences (Fig. 1). During our previous investigations, we had observed that different species exhibited a differential preference for amino acids in the vicinity of their methionine residues, even though the relative frequencies of the proteinogenetic amino acids were very similar in the analyzed species. This observation prompted us to explore the potential of sequence environments to accurately reconstruct phylogenies using genome/proteome datasets of unaligned sequence information.
As a first approach, we decided to carry out a pilot study using an optimal set of genomes, in the sense that the expected tree topology is widely accepted. To this end, we chose the protein-coding mitogenomes of 11 species of bovids. This dataset is small and simple, coding for only 143 proteins whose sequences are curated by the NCBI and are expected to be very accurate. Furthermore, the orthologous relationships among the proteins belonging to this set are obvious and undisputed. Moreover, mitochondrial sequences are often used to generate metazoan phylogenies, hence the tree generated by Env-NJ can be easily compared to those generated by other methods, either based on sequence alignments or alignment-independent. Given that the group of organisms was formed by species closely related, and the optimal conditions discussed above, not surprisingly, most methods consistently produced the same tree topology (Fig. 2 and Table 1).
Encouraged by this success, we next tested the Env-NJ method with a larger, more diverse and more controversial dataset consisting in the mitogenomes of 34 mammalian species spanning 13 orders. This dataset, first analyzed by Reyes and coworkers using alignment-based methods44, has been used later by different authors employing different tree building methods. Thus, the same genome set has been analyzed by Stuart and coworkers using the SVD-4-Gram method29 and also by Li and colleagues, using a method that also works on unaligned sequences, but in this case exploiting the Kolmogorov complexity concept to estimate distances between genomes36. In the R package EnvNJ accompanying the current paper (throughout the text we use the term Env-NJ to indicate the method, while EnvNJ refers to the software) we have implemented, in addition to the Env-NJ method, those utilities required to reproduce the trees reported by Reyes et al. (Fig. 3A) and Stuart and coworkers (Fig. 3D), which are shown herein for comparative purposes. The method based on the Kolmogorov complexity was not included in the comparison because, although it faithfully reproduced the tree obtained by Cao and coworkers for a smaller and less conflictive set of mammalian species3, it offered a rather poor phylogeny, showing polytomy, for the taxa we are addressing herein (see Fig. 2 from Li et al. 2001).
Figure 3 and Table 2 summarize the topologies comparison of the trees obtained with the different methods being compared. Overall, the Env-NJ tree seems to be a reasonably good approximation to the reference tree, at least as good as any of the trees obtained by alternative methods. This was also true when mitogenomes from other groups of vertebrates were analyzed (Fig. S1). To this respect, a group of 25 species of fish, which is widely used to benchmark alignment-independent phylogenetic methods25, was analyzed using different approaches. Again, the Env-NJ yielded excellent results, both in term of computation time as well as regarding tree-topology reliability.
In the current work, the accuracy of the different alignment-independent methods has been evaluated by a comparison of topology between the reconstructed tree using a given method and the corresponding reference tree. For this purpose, we computed the nRF distance between the trees, which is a straightforward to interpret metric (Fig. 5). Despite of being a metric widely used in the literature to quantify similarity between pairs of phylogenetic trees25,26,35,39,40,49,50, the nRF is known to present certain shortcomings such as rapid saturation and imprecise values51,52. Therefore, to rule out that these drawbacks could be biasing the results presented herein, we also computed a generalized RF metric (GRF) designed to avoid the limitations of the nRF53. Using two different datasets (Fig. S1 for the fish group, and Table 4 for the plant group) we found a good positive correlation between nRF and GRF (R-squared = 0.97, p-value = 2.4 10–9), and the conclusions obtained regarding the benchmark analyses are equally well supported regardless the metric employed.
Summary of the performance of different programs with different data sets. Four sets of sequences were analyzed with the programs EnvNJ, alfpy, FFP, CVTree, ALFRED-G, SANS-serif and Prot-SpaM, using different parameters combinations. The perform of each method, in terms of nRF distances to the reference tree is given in Table 1 for the bovid group, Table 2 for the group of mammals, Table 4 for the plant species and Figure S1 for the fish species. This figure summarizes and shows the results that each program yielded with the optimal selection of parameters.
We have extensively assessed the performance of the Env-NJ method on mitogenomes. In this context, the new method seems to be a valid alternative for phylogenomics since it has three valuable properties: (i) accuracy, (ii) speed and (iii) independence of positional homology. Indeed, Env-NJ does not rest on positional homology, and it does not require to identify orthologous proteins to proceed with the computation. However, one thing is that the method does not require identification of orthologous proteins, and quite another is that the method does not require the presence of orthologous proteins in the dataset. The latter is guaranteed when working with mitogenomes, where the presence of one-to-one orthologous proteins is guaranteed. On the other hand, when all proteins encoded by the full genome of the analyzed species are used as input, the success of the Env-NJ approach (Table 4) could be due to the presence of a high proportion of orthologous proteins in the input dataset. Therefore, we next wondered whether the Env-NJ method would be able to reconstruct a phylogeny analyzing non-orthologous proteins? That is, when each species contributes a set of proteins completely unrelated to the protein sets contributed by the other species under analysis.
To address this issue, we chose a small set of species formed by three animals (human, chimp and gorilla) and two plants (Arabidopsis thaliana and A. lyrate). For each species we randomly sampled 180 protein sequences from its proteome. The selection process was random with the only restriction that there were no pairs of orthologous proteins among the 900 sequences that made up the dataset (both the script to sample the sequences and the sequences themselves can be obtained at https://bitbucket.org/jcaledo/envnj/src/master/AncillaryCode/Oma_PlantAnimal.R, and https://bitbucket.org/jcaledo/envnj/src/master/Datasets/oseq.Rda, respectively). When this dataset was subjected to Env-NJ, the recovered tree was the expected one (Fig. 4), where human was closer to chimp than gorilla, and the two plants appeared as sister operational taxonomic units (OTUs). More interestingly, under these challenging conditions, the strategy based on sequence environments was the only one that provided an acceptable result. Having a tree building method that does not require orthologs identification is a good thing, and Env-NJ is indeed such a method. Having a method that does not even require the presence of orthologs in the dataset, is even better and Env-NJ may fulfil this feature. A drawback of Env-NJ, as well as most alignment-free approaches, is that they are distance methods. That is, there is no evolutionary model behind them, which precludes the use of maximum likelihood techniques to explore tree spaces. Undoubtedly, further research effort will be required before we can witness a significant breakthrough in the field of phylogenomics.
Conclusion
In the current report we describe a new tree-building method and its implementation into an R package (EnvNJ). This new method presents many advantages: (i) it does not resort to lossy data compression; (ii) it is computationally very fast, making it suitable for addressing whole genomes; (iii) because the method makes use of whole genomes/proteomes, there is no gene tree versus species tree problem; (iv) there is no need for multiple sequence alignment, which contributes to the speed of the method and avoids the impact of misalignments on the tree topology; (v) it does not require orthology identification, which further contributes to shortening computation times. Finally, the possibility that Env-NJ may perform well even with non-orthologous protein datasets, is a line of research that deserves further work in the future.
Material and methods
The species vector space
It has been shown that every amino acid has a characteristic sequence environment in proteins41,54. In previous works, we have analyzed the sequence environment (10 residues on each side) around methionine residues in human proteins42,43,55. Thus, and just based on methionine residues, the human proteome can be characterized by a matrix, M, whose elements (mij) provide the absolute frequency of the amino acid i at the position j in the environment of methionine residues (Fig. 1A). Similarly, for each proteinogenic amino acid, \(X\in \left\{A,R,N,D,C,Q,E,G,H,I,L,K,M,F,P,S,T,W,Y,V\right\}\), a matrix \(\left({x}_{ij}\right)\in {\mathcal{M}}_{20}({\mathbb{N}})\) can be computed. In this way, the protein-coding genome of a given species of interest can be characterized by a set of 20 square matrices of order 20 or, equivalently, by a vector, \(u\in {U}^{8000}\), of dimension 8000 (Fig. 1B). However, when coding species as vectors, the dimension of these vectors will depend on the radius chosen. Thus, in general, \(u\in {U}^{n}\), where \(n=20 \left(2 radius\right) 20=800 radius\). In this way, each vector is used to represent an organism (its protein-coding genome). The components (coordinates) of the vector reflect the preference of the different amino acids at the different positions of the sequence environments.
A suitable metric for the species vector space
Once species are encoded as high-dimensional vectors, we can make use of the extensive mathematical tools of numerical linear algebra. Since we are interested in assessing distances between species, we must endow this vector space with a suitable metric for our purpose. To this end, we must look for functions, d, able to provide a distance between vectors.
In general, any function, d, to be considered a distance must satisfy the following 4 properties. (i) Positive definiteness: \(d\left({u}_{i},{u}_{j}\right)\ge 0 \forall {u}_{i},{u}_{j}\in {U}^{n}\); (ii) coincidence axiom: \(d\left({u}_{i},{u}_{j}\right)=0\Leftrightarrow {u}_{i}={u}_{j}\); (iii) symmetry: \(d\left({u}_{i},{u}_{j}\right)=d\left({u}_{j},{u}_{i}\right) \forall {u}_{i},{u}_{j}\in {U}^{n}\); and (iv) triangle inequality: \(d\left({u}_{i},{u}_{j}\right)\le d\left({u}_{i},{u}_{k}\right)+d\left({u}_{k},{u}_{j}\right) \forall {u}_{i},{u}_{j},{u}_{k}\in {U}^{n}\). A wide variety of metrics can be used to measure relatedness between vectors. The function vect2tree() from the EnvNJ package accompanying this paper, implements 29 different metrics previously described56,57. However, as illustrated in Fig. 1C,D, not all of them will be equally suitable for our purpose of establishing evolutionary relationships between species. Furthermore, the link between a given metric and its performance is not always obvious58. Nevertheless, for the sequence datasets we have used in the current study, we have noticed that the so-called Jensen-Shannon and cosine-based dissimilarities perform better than other metrics. Although many of the offered methods compute proper distances, that is not always the case. For instance, the ‘cosine’ method we described next, does not satisfy the coincidence axiom, so it cannot be considered a true distance. This fact, far from being a drawback, can be an advantage (as it will be argued below) for our phylogenetic purposes. In the context of latent semantic analysis, a common measure of similarity between two vectors is the cosine of the angle between them58,59. Since protein sequence data can be regarded as a complex written language, Stuart and coworkers have proposed the use of the cosine between two vectors as a suitable measure of vector similarity when the vectors being considered contain information related to protein sequences28,29. For instance, if we have the protein-coding genome of the species i and j (Fig. 1C), their similarity can be assessed by the expression:
where \({u}_{i}^{T}{u}_{j}\) is the dot product of the vectors \({u}_{i}\) and \({u}_{j}\), and \(\Vert \Vert \) is the Euclidean vector norm. It should be noted that the function \(f\left({u}_{i},{u}_{j}\right)=cos{\theta }_{ij}\) is not a distance properly speaking. For instance, suppose that two species have identical genomes, in this case \({u}_{i}={u}_{j}\) and we would expect a null distance between them. However, \(f\left({u}_{i},{u}_{j}\right)=cos{\theta }_{ij}=cos0=1\). Nevertheless, pairwise cosine values can be converted into pairwise evolutionary distances using the following formula:
This formula converts a similarity measure into a distance measure28. It is important to note that this evolutionary distance is not a proper distance metric as it violates the coincidence axiom. For instance, \(d\left({u}_{i},{2u}_{j}\right)=0\) but \({u}_{i}\ne 2{u}_{i}\). However, this violation is very convenient for our goals. A concrete example will be useful to understand this assertion. Suppose that we have a population that splits into two species. Suppose, further, that one of this species undergoes a genome duplication event, but otherwise their proteomes are identical. In such a scenario, the computed species vectors would be \({u}_{j}=2{u}_{i}\). Although both vectors have different lengths, since their directions are identical, we obtain \(d\left({u}_{i},{u}_{j}\right)=0\), which conveniently reflects the fact that their proteomes are equal and therefore the neighborhood preferences of their sequence environments are the same in both species (Fig. 1D).
Environment-based trees
After encoding the genome of each species into a vector, as described above, these vectors are used to obtain a matrix of pairwise cosine values that are subsequently converted into a matrix of pairwise evolutionary distances using the formula given in the previous section. Alternatively, other metrics can be used to obtain a distance matrix. In the EnvNJ package accompanying this paper, we have implemented 29 different metrics among which the user can choose. However, in our experience the cosine-based dissimilarity and the Jensen-Shannon distance are among the best performing metrics for phylogenetic analyses using sequence environments. In any case, the obtained distance matrix can be used to produce a phylogenetic tree employing the neighbor joining algorithm60.
Implementation
The Env-NJ tree building method has been implemented in an R package, EnvNJ. The package, which works on all major operating systems (Windows, MacOS and Linux) can be installed either from CRAN, install.packages(“EnvNJ”), or from its bitbucket repository, typing consecutively the following three commands in an R terminal: install.packages(“devtools”), library(devtools), install_bitbucket(“jcaledo/envnj”, subdir = “REnvNJ”). Since the protein sequence datasets analyzed in the current work (see below) have also been included into the package, once it has been installed, the trees shown in Fig. 2 and 3C can be easily obtained with the commands envnj(bovids, r = 2) and envnj(reyes, r = 46), respectively. Further help can be obtained from the package documentation by introducing into the R terminal: ?envnj. A vignette about the use of the EnvNJ package can be found as Supplementary Material.
In addition to the Env-NJ method described in this paper, the package EnvNJ also implements the method based on the SVD-n-Gram approach, previously described by Stuart and coworkers28. The aim was to facilitate its use for comparative purposes (check the documentation, ?svdgram).
Mitogenome Datasets
The mtDNA-encoded protein sequences were obtained from the NCBI genome database (https://www.ncbi.nlm.nih.gov/genome/organelle). Two sets of mitogenomes have been analyzed in the current work. The first set is formed by 11 species of bovids including Bison bison, Bison bonasus, Bos grunniens, Bos indicus, Bos javanicus, Bos primigenius, Bos taurus, Bubalus bubalis, Bubalus depressicornis, Pseudoryx nghetinhensis, Syncerus caffer. An R dataframe containing these sequences can be loaded and examined by typing data(bovids) after having installed the R package EnvNJ.
A second set of mitogenomes analyzed in this study is the one formed by 34 mammalian species spanning 13 orders, first used by Reyes and coworkers44, which includes the following species: Artibeus jamaicensis (Ajam), Balaenoptera musculus (Bmus), Balaenoptera physalus (Bphy), Bos taurus (Btau), Canis lupus (Clup), Cavia porcellus (Cpor), Ceratotherium simum (Csim), Dasypus novemcinctus (Dnov), Didelphis virginiana (Dvir), Equus asinus (Easi), Equus caballus (Ecab), Felis catus (Fcat), Glis glis (Ggli), Gorilla gorilla (Ggor), Halichoerus grypus (Hgry), Hippopotamus amphibius (Hamp), Homo sapiens (Hsap), Hylobates lar (Hlar), Loxodonta africana (Lafr), Macropus robustus (Mrob), Mus musculus (Mmus), Ornithorhynchus anatinus (Oana), Orycteropus afer (Oafe), Oryctolagus cuniculus (Ocun), Ovis aries (Oari), Pan paniscus (Ppan), Pan troglodytes (Ptro), Papio hamadryas (Pham), Phoca vitulina (Pvit), Pongo pygmaeus (Ppyg), Rattus norvegicus (Rnor), Rhinoceros unicornis (Runi), Sciurus vulgaris (Svul), Sus scrofa (Sscr). Again, this dataset can be obtained in a suitable format (as dataframe) by typing in the R terminal: data(reyes).
Env-NJ trees using non-orthologous protein datasets
We chose three closely related primate species (human, chimp and gorilla) and two Arabidopsis species (A. thaliana and A. lyrate). Since we wanted to make sure that no orthology could be established between any pair of proteins from the dataset subjected to analysis, we proceeded as described next. First, we started by identifying a set of 907 one-to-one orthologous proteins present in the five species. To achieve that, we took advantage of the REST API for the OMA orthology database61,62. Both, the dataset (oseq.Rda) and the script (Oma_PlantAnimal.R) used to obtain it, can be downloaded from https://bitbucket.org/jcaledo/envnj/src/master/Datasets and https://bitbucket.org/jcaledo/envnj/src/master/AncillaryCode, respectively. In this way, the oseq.Rda object is a dataframe with five columns (one per species) and 907 rows (one per orthologous protein), and each entry contains the corresponding protein sequence. To form a dataset of non-orthologous proteins we proceeded as follows. For the first column (species) we randomly chose 180 rows (proteins). Afterward, the randomly selected rows were discarded from the dataframe before proceeding with the next column (species). Among the remaining rows, again we randomly selected 180, and the corresponding proteins from the second species were selected before removing the randomly selected rows from the dataframe. This operation was repeated until reaching the last species, at which point we had a collection of 900 non-orthologous proteins (180 per species). This randomly selected dataset formed by 900 non-orthologous proteins was then subjected to Env-NJ.
Plant proteomes
AFproject (http://afproject.org) is a publicly available web-based service for objective performance comparison of alignment-free sequence comparison tools on different datasets25. They provide a benchmark dataset formed by the full genome sequences for 14 plant species and the corresponding reference species tree. Since the Env-NJ approach uses protein sequences and to avoid pre-processing (identification of open reading frames and translation) we resorted to the UniProt Proteomes (https://www.uniprot.org/proteomes) to search for protein sequences belonging to this group of plant species. In this way, we managed to assemble a dataset formed by 425,115 proteins from 11 species accounting for around 170 million amino acids (Table 3). The three species for which we could not find enough data were pruned from the reference tree.
Robinson–Foulds distance
As a measure of the accuracy attributable to each phylogeny, the normalized Robinson-Foulds (nRF) distances between the reconstructed trees and the reference trees were computed. The Robinson-Fould algorithm to compute distances between trees topologies63, as implemented in the R package phangorn64, was used for this purpose. Briefly, Let T1 and T2 be two sets formed by all the splits at internal edges for tree 1 and tree 2, respectively (the two trees whose topologies we want to compare), then the cardinal of the symmetric difference of these two sets provides the Robinson-Foulds distance.
In other words, the Robison-Foulds is the number of splits appearing in one tree but not the other. The normalized Robinson-Foulds distance, nRF, is obtained by dividing RF by the maximal possible distance, that is
Normalization forces this metric to take values between 0 and 1, which makes its interpretation straightforward: 0 indicating identical tree topologies and 1 pointing to the most dissimilar topologies.
Data availability
The Env-NJ method is implemented in the R package EnvNJ. Release versions are available via CRAN and work on all major operating systems. The development version is maintained at https://bitbucket.org/jcaledo/envnj/src/master. The mtDNA-encoded protein sequences in the 11 species of bovids can be obtained from the EnvNJ package just typing, after loading the package, data(bovids). Similarly, the 442 protein sequences that make up the dataset referred to as Reyes, can be obtained typing data(reyes). Alternatively, all the data employed in the current work, together with their corresponding descriptions can be obtained from the Bitbucket repository at https://bitbucket.org/jcaledo/envnj/src/master/Datasets.
References
Hedges, S. B. Molecular evidence for the origin of birds. Proc. Natl. Acad. Sci. USA 91, 2621–2624 (1994).
Russo, C. A. M., Takezaki, N. & Nei, M. Efficiencies of different genes and different tree-building methods in recovering a known vertebrate phylogeny. Mol. Biol. Evol. 13, 525–536 (1996).
Cao, Y. et al. Conflict among individual mitochondrial proteins in resolving the phylogeny of eutherian orders. J. Mol. Evol. 47, 307–322 (1998).
de Queiroz, A. & Gatesy, J. The supermatrix approach to systematics. Trends Ecol. Evol. 22, 34–41 (2007).
Bininda-Emonds, O. R. P. The evolution of supertrees. Trends Ecol. Evol. 19, 315–322 (2004).
Liu, L., Yu, L., Kubatko, L., Pearl, D. K. & Edwards, S. V. Coalescent methods for estimating phylogenetic trees. Mol. Phylogenet. Evol. 53, 320–328 (2009).
Gatesy, J., Matthee, C., DeSalle, R. & Hayashi, C. Resolution of a supertree/supermatrix paradox. Syst. Biol. 51, 652–664 (2002).
Bininda-Emonds, O. R. P. et al. Supertrees are a necessary not-so-evil: A comment on gatesy. Syst. Biol. 52, 724–729 (2003).
Bininda-Emonds, O. R. P. Trees versus characters and the supertree/supermatrix ‘paradox’. Syst. Biol. 53, 356–359 (2004).
Janies, D. A., Studer, J., Handelman, S. K. & Linchangco, G. A comparison of supermatrix and supertree methods for multilocus phylogenetics using organismal datasets. Cladistics 29, 560–566 (2013).
Thorne, J. L. Models of protein sequence evolution and their applications. Curr. Opin. Genet. Dev. 10, 602–605 (2000).
Lake, J. A. & Moore, J. E. Phylogenetic analysis and comparative genomics. Trends Guid. Bioinf. Trends J. Suppl. 1, 22–23. https://doi.org/10.1136/jmg.38.11.807 (1998).
Springer, M. S. & Gatesy, J. On the importance of homology in the age of phylogenomics. Syst. Biodivers. 16, 210–228 (2018).
Wong, K. M., Suchard, M. A. & Huelsenbeck, J. P. Alignment uncertainty and genomic analysis. Science 319, 473–476 (2008).
Lake, J. A. The order of sequence alignment can bias the selection of tree topology. Mol. Biol. Evol. 8, 378–385 (1991).
Mugridge, N. B. et al. Effects of sequence alignment and structural domains of ribosomal DNA on phylogeny reconstruction for the protozoan family sarcocystidae. Mol. Biol. Evol. 17, 1842–1853 (2000).
Morrison, D. A. & Ellis, J. T. Effects of nucleotide sequence alignment on phylogeny estimation: A case study of 18S rDNAs of apicomplexa. Mol. Biol. Evol. 14, 428–441 (1997).
Ogden, T. H. & Rosenberg, M. S. Multiple sequence alignment and phylogenetic inference. Syst. Biol. 55, 314–332 (2006).
Wu, M., Chatterji, S. & Eisen, J. A. Accounting for alignment uncertainty in phylogenomics. PLoS ONE 7, 1–10 (2012).
Boore, J. L. & Brown, W. M. Big trees from little genomes: Mitochondrial gene order as a phylogenetic tool. Curr. Opin. Genet. Dev. 8, 668–674 (1998).
Fitz-Gibbon, S. T. & House, C. H. Whole genome-based phylogenetic analysis of free-living microorganisms. Nucl. Acids Res. 27, 4218–4222 (1999).
Snel, B., Bork, P. & Huynen, M. A. Genome phylogeny based on gene content. Nat. Genet. 21, 108–110 (1999).
Caetano-Anollés, G. & Caetano-Anollés, D. An evolutionarily structural universe of protein architecture. Genome Res. 13, 1563–1571 (2003).
Yang, S., Doolittle, R. F. & Bourne, P. E. Phylogeny determined by protein domain content. Proc. Natl. Acad. Sci. USA 102, 373–378 (2005).
Zielezinski, A. et al. Benchmarking of alignment-free sequence comparison methods. Genome Biol. 20, 144 (2019).
Sims, G. E., Jun, S. R., Wu, G. A. & Kim, S. H. Alignment-free genome comparison with feature frequency profiles (FFP) and optimal resolutions. Proc. Natl. Acad. Sci. USA 106, 2677–2682 (2009).
Zielezinski, A., Vinga, S., Almeida, J. & Karlowski, W. M. Alignment-free sequence comparison: Benefits, applications, and tools. Genome Biol. 18, 1–17 (2017).
Stuart, G. W., Moffett, K. & Leader, J. J. A comprehensive vertebrate phylogeny using vector representations of protein sequences from whole genomes. Mol. Biol. Evol. 19, 554–562 (2002).
Stuart, G. W., Moffett, K. & Baker, S. Integrated gene and species phylogenies from unaligned whole genome protein sequences. Bioinformatics 18, 100–108 (2002).
Qi, J., Luo, H. & Hao, B. CVTree: A phylogenetic tree reconstruction tool based on whole genomes. Nucl. Acids Res. 32, 45–47 (2004).
Xu, Z. & Hao, B. CVTree update: A newly designed phylogenetic study platform using composition vectors and whole genomes. Nucl. Acids Res. 37, 174–178 (2009).
Zuo, G. CVTree: A parallel alignment-free phylogeny and taxonomy tool based on composition vectors of genomes. Genom. Proteom. Bioinf. https://doi.org/10.1016/j.gpb.2021.03.006 (2021).
Vinga, S., Gouveia-Oliveira, R. & Almeida, J. S. Comparative evaluation of word composition distances for the recognition of SCOP relationships. Bioinformatics 20, 206–215 (2004).
Leimeister, C. A. & Morgenstern, B. Kmacs: The k-mismatch average common substring approach to alignment-free sequence comparison. Bioinformatics 30, 2000–2008 (2014).
Thankachan, S. V., Chockalingam, S. P., Liu, Y., Krishnan, A. & Aluru, S. A greedy alignment-free distance estimator for phylogenetic inference. BMC Bioinf. 18, 1–8 (2017).
Li, M. et al. An information-based sequence distance and its application to whole mitochondrial genome phylogeny. Bioinformatics 17, 149–154 (2001).
Wiskunde, C., Vitanyi, P. M. B., Wiskunde, C., Cilibrasi, R. L. & Vit, P. M. B. Fast Whole-genome phylogeny by compression: The COVID-19 case Fast Whole-Genome Phylogeny by Compression : the. 0–7 (2021).
Rempel, A. & Wittler, R. SANS serif: Alignment-free, whole-genome-based phylogenetic reconstruction. Bioinformatics 1, 1–3. https://doi.org/10.1093/bioinformatics/btab444 (2021).
Leimeister, C. A. et al. Prot-SpaM: Fast alignment-free phylogeny reconstruction based on whole-proteome sequences. Gigascience 8, 1–14 (2018).
Dencker, T. et al. ‘Multi-SpaM’: a maximum-likelihood approach to phylogeny reconstruction using multiple spaced-word matches and quartet trees. NAR Genom. Bioinforma. 2, 1–10 (2020).
Cserzo, M. & Simon, I. Regularities in The Primary Structure of Proteins. Int. J. Pept. Prot. Res. 34, 184–195 (1989).
Aledo, J. C., Cantón, F. R. & Veredas, F. J. Sulphur atoms from methionines interacting with aromatic residues are less prone to oxidation. Sci. Rep. 5, 16955 (2015).
Veredas, F. J., Cantón, F. R. & Aledo, J. C. Methionine residues around phosphorylation sites are preferentially oxidized in vivo under stress conditions. Sci. Rep. 7, 40403 (2017).
Reyes, A., Gissi, C., Pesole, G., Catzeflis, F. M. & Saccone, C. Where do rodents fit? Evidence from the complete mitochondrial genome of Sciurus vulgaris [2]. Mol. Biol. Evol. 17, 979–983 (2000).
Upham, N. S., Esselstyn, J. A. & Jetz, W. Inferring the mammal tree: Species-level sets of phylogenies for questions in ecology, evolution, and conservation. PLoS Biol. 17, 1 (2019).
De Bruyn, A., Martin, D. P. & Lefeuvre, P. Phylogenetic reconstruction methods: An overview. Methods Mol. Biol. 1115, 257–277 (2014).
Doolittle, R. F. The Multiplicity of Domains in Proteins. Mult. Dreams 64, 287–314 (1995).
Moret, B. M. E. & Warnow, T. Advances in phylogeny reconstruction from gene order and content data. Methods Enzymol. 395, 673–700 (2005).
Ferreira, A. P. S. et al. Active glutaminase C self-assembles into a supratetrameric oligomer that can be disrupted by an allosteric inhibitor. J. Biol. Chem. 288, 28009–28020 (2013).
Li, Y. et al. Feature frequency profile-based phylogenies are inaccurate. Proc. Natl. Acad. Sci. USA 117, 31580–31581 (2020).
Lin, Y., Rajan, V. & Moret, B. M. E. A metric for phylogenetic trees based on matching. Lect. Notes Comput. Sci. (including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics) 6674 LNBI, 197–208 (2011).
Kuhner, M. K. & Yamato, J. Practical performance of tree comparison metrics. Syst. Biol. 64, 205–214 (2015).
Smith, M. R. Information theoretic generalized Robinson-Foulds metrics for comparing phylogenetic trees. Bioinformatics 36, 5007–5013 (2020).
Tüdös, É., Fiser, A. & Simon, I. (1994) Different sequence environments of amino acid residues involved and not involved in long-range interactions in proteins. Int. J. Pept. Protein Res. 43, 205–208 (1994).
Aledo, J. C. & Aledo, P. Susceptibility of protein methionine oxidation in response to hydrogen peroxide treatment–ex vivo versus in vitro: A computational insight. Antioxidants 9, 1 (2020).
Luczak, B. B., James, B. T. & Girgis, H. Z. A survey and evaluations of histogram-based statistics in alignment-free sequence comparison. Brief. Bioinf. 20, 1222–1237 (2018).
Cha, S.-H. Comprehensive survey on distance/similarity measures between probability density functions. Int. J. Math. Model. Methods Appl. Sci. 1, 300–307 (2007).
Jones, W. P. & Furnas, G. W. Pictures of relevance: A geometric analysis of similarity measures. J. Am. Soc. Inf. Sci. 38, 420–442 (1987).
Berry, M. W., Drmač, Z. & Jessup, E. R. Matrices, vector spaces, and information retrieval. SIAM Rev. 41, 335–362 (1999).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Altenhoff, A. M. et al. OMA orthology in 2021: Website overhaul, conserved isoforms, ancestral gene order and more. Nucleic Acids Res. 49, D373–D379 (2021).
Kaleb, K., Vesztrocy, A. W., Altenhoff, A. & Dessimoz, C. Expanding the orthologous matrix (OMA) programmatic interfaces: REST API and the OmaDB packages for R and Python [version 2; peer review: 2 approved]. F1000Research 8, 1–21 (2019).
Robinson, D. F. & Foulds, L. R. Comparison of phylogenetic trees. Math. Biosci. 53, 131–147 (1981).
Schliep, K. P. phangorn: Phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Acknowledgements
The author is in debt to Pablo Aledo for his advice and help with all the aspects related to the linear algebra. The author also thanks Alicia Esteban and Elena Aledo for helpful discussion during the preparation of this work. This work was supported by European Regional Development Fund and the University of Málaga [UMA18-FEDERJA-149].
Author information
Authors and Affiliations
Contributions
J.C.A. designed and executed the present study. J.C.A. wrote the manuscript text and prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aledo, J.C. Phylogenies from unaligned proteomes using sequence environments of amino acid residues. Sci Rep 12, 7497 (2022). https://doi.org/10.1038/s41598-022-11370-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-11370-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.