Abstract
Effective and successful clinical trials are essential in developing new drugs and advancing new treatments. However, clinical trials are very expensive and easy to fail. The high cost and low success rate of clinical trials motivate research on inferring knowledge from existing clinical trials in innovative ways for designing future clinical trials. In this manuscript, we present our efforts on constructing the first publicly available Clinical Trials Knowledge Graph, denoted as \(\mathop {\mathtt {CTKG}}\limits\). \(\mathop {\mathtt {CTKG}}\limits\) includes nodes representing medical entities in clinical trials (e.g., studies, drugs and conditions), and edges representing the relations among these entities (e.g., drugs used in studies). Our embedding analysis demonstrates the potential utilities of \(\mathop {\mathtt {CTKG}}\limits\) in various applications such as drug repurposing and similarity search, among others.
Similar content being viewed by others
Introduction
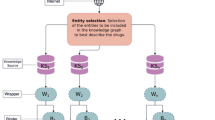
Clinical trials are studies aiming at determining the safety and efficacy of interventions, treatments or investigational drugs on human subjects1. Effective and successful clinical trials are essential in developing new drugs and advancing new treatments2. However, clinical trials are very expensive. As reported in Sertkaya et al.3, the average cost of a single phase in clinical trials ranges from 1.4 million up to 52.9 million US dollars. In addition, the success rate of the clinical trials is considerably low. As reported in Wong et al.4, for certain therapeutic groups like Oncology, the overall success rate of clinical trials could be as low as 3.4%. The high cost and low success rate of clinical trials motivate deliberate analysis of existing clinical trials, inferring knowledge from them, utilizing existing clinical trials in innovative ways, and accordingly carefully designing future clinical trials. The Access to Aggregate Content of ClinicalTrials.gov (AACT) database5 represents an effort in enhancing the accessibility and analysis of the clinical trial data. However, as a relational database, AACT is not formatted for the purpose of inferring new knowledge from existing clinical trials6. A Knowledge Graph (KG), instead, is a graph representation in which information entities are represented as nodes, and their relations are coded as edges connecting the corresponding nodes. In contrast to relational databases, KG has been proven7,8,9,10 to be an effective representation for knowledge inference purposes. Constructing a KG over clinical trial data is vital for advancing the analysis and research of clinical trials. In this manuscript, we present our work on constructing a such KG, referred to as Clinical Trials Knowledge Graph, denoted as \(\mathop {\mathtt {CTKG}}\limits\), and also release \(\mathop {\mathtt {CTKG}}\limits\) to the research community to facilitate advanced research using clinical trial data. \(\mathop {\mathtt {CTKG}}\limits\) includes nodes representing medical entities (e.g., studies, drugs and conditions), and edges representing relations among these entities (e.g., drugs used in studies). Different from the recently released knowledge base11 that focuses only on extracting medical entities from the eligibility criteria in clinical trials, \(\mathop {\mathtt {CTKG}}\limits\) includes more medical entities (e.g., adverse events and outcomes) and also the relations among these entities. The rich information in \(\mathop {\mathtt {CTKG}}\limits\) could enable more biomedical applications (e.g., adverse drug event prediction, outcome prediction) than the existing knowledge base in clinical trials. Figure 1 presents the schema of \(\mathop {\mathtt {CTKG}}\limits\). The detailed descriptions of nodes and edges in \(\mathop {\mathtt {CTKG}}\limits\) will be presented in “Nodes in CTKG” section. To the best of our knowledge, \(\mathop {\mathtt {CTKG}}\limits\) is the first publicly available clinical trials knowledge graph in the scientific research community. The results of the embedding analysis over \(\mathop {\mathtt {CTKG}}\limits\) demonstrate its potential utilities in various applications such as drug repurposing and similarity search, among others.
Results
\(\mathop {\mathtt {CTKG}}\limits\) schema
Figure 1 presents the schema of \(\mathop {\mathtt {CTKG}}\limits\). The schema presents the different information entities involved in clinical trials, represented as nodes, and the relations among them, represented as edges. There are 18 types of nodes and 21 types of edges in \(\mathop {\mathtt {CTKG}}\limits\). Each node and edge type has attributes describing the properties of the nodes and edges. The statistics of different nodes and edges are presented in Tables 1 and 2, respectively. Detailed descriptions of node and edge attributes are available in the online documentation of \(\mathop {\mathtt {CTKG}}\limits\)12. We developed a web portal (https://u.osu.edu/ning.104/data-set/ctkg/ctkg-webportal/) for users to visualize \(\mathop {\mathtt {CTKG}}\limits\) and access its nodes and edges interactively.
Nodes in \(\mathop {\mathtt {CTKG}}\limits\)
Each study node represents a clinical trial and is associated with the primary properties of that clinical trial as node attributes. The properties of each study node describe the purposes, phases and the protocols of the corresponding clinical trial. Each study node links to condition nodes, drug nodes, outcome nodes and multiple types of group nodes via one-to-many relationships. Each condition node describes a disease or syndrome that is extracted from the AACT and studied by some clinical trials.
Each drug-term node represents the drug used in clinical trials, and is identified by the extracted drug mention (“Drug mentions and normalization” section). The drug-term nodes connect with study nodes via StudiedDrug and UsedDrug relations. The StudiedDrug relation connects studies and drug terms that are studied in at least one study group of the corresponding clinical trial, and the UsedDrug relation connects studies and the auxiliary drug terms such as pain reducers. Please refer to “Drug mentions and normalization” section for more details.
Each outcome node represents an outcome measure used to evaluate the efficacy of interventions in the clinical trials, and has the name and the description of the outcome measure as attributes. For example, the study “NCT04322526” uses the outcome “changes in blood oxygenation level-dependent (BOLD)” to evaluate the efficacy of interventions. Each outcome node is connected to a study node, representing that this specific outcome is used within the study. Note that unlike the condition node linking to multiple study nodes, each outcome node links to a unique study node. This is due to the complexity and the diversity of outcome measures, which makes it difficult to be shared across multiple study nodes. Each outcome node also links to one cluster-outcome node and multiple standard-outcome nodes. The connection between the outcome node and the cluster-outcome node represents that the name of the outcome can be assigned to the cluster represented by the cluster-outcome node, while the connection between the outcome node and the standard-outcome node represents that the name or the description of the outcome contains the standard outcome measure. Please refer to “Outcome extraction and outcome clustering” section for more details.
Each group node represents a study arm or a comparison group, that is, a group of participants who receive a specific intervention. There are multiple types of group nodes as follows:
-
event-group node. The information described by each event-group node is the number of participants within the group affected by specific types of adverse events. Each event-group node is connected to multiple drug nodes representing the drugs used in the event group, and adverse-event nodes representing the specific adverse events that occurred in the event group. Each adverse-event node also links to an organ-system node representing the affected organ system.
-
baseline-group node. Each baseline-group node represents a group of participants with their demographic attributes (e.g., “Age” and “Ethnicity”) or study-specific attributes (e.g., “Baseline Modified Gingival Index”). Each baseline-group node is connected to one or multiple baseline-record nodes.
-
drop-group node. Each drop-group node represents a group of participants with their withdrawal information. Each drop-group node is connected to one or multiple period nodes. Each period node represents an interval of the study (e.g., “First Intervention” and “Part 1: Treatment Period 1”), and has attributes describing the number of participants at the beginning and the end of the period. Each period node can link to multiple drop-record nodes. Each drop-record node includes a withdrawal reason and documents the number of the participants in the group withdrawing with this reason in a period.
-
outcome-group node. An outcome-group node has the information on the efficacy of the studied interventions on the participants. The efficacy is evaluated by different outcome measures and analyzed by different statistical test methods with the measurements. For example, an outcome-group node could represent a group of 17 participants receiving Naltrexone as the intervention, and the efficacy of Naltrexone was evaluated using the results from this group of participants; these 17 participants and the efficacy evaluation are included in an outcome-group node. Other nodes related to the efficacy measures of interventions are as below:
-
method node. Each method node represents a statistical hypothesis testing method that is used to make inference or draw conclusion statistically from the data collected in clinical trials. For example, the variance analysis method “ANOVA” could be a method node that is used to test the superiority of an intervention compared with a control in a clinical trial. Each method node is connected to multiple outcome-analysis nodes, representing that the method is used to conduct the analyses. Please refer to “Statistical analysis method normalization” section for more details about the normalization of method names.
-
outcome-measurement node. Each outcome-measurement node represents the measurement of a specific outcome measure (i.e., outcome node) on the corresponding group of participants. For example, the measurement of the average changes in BOLD (i.e., outcome node) on the 17 participants in the outcome-group node is 1.23 with a standard deviation 1.07, and is represented as a outcome-measurement node. Each outcome-measurement node links to one outcome node and one outcome-group node.
-
outcome-analysis node. Each outcome-analysis node represents a statistical analysis on a specific outcome measure by comparing multiple outcome groups using a statistical testing method. For example, the analysis of the outcome “average changes in BOLD” via the statistical testing method “paired t-test” on two groups of participants receiving the intervention “Naltrexone” and placebo, is represented as an outcome-analysis node; the p-value of the analysis is 0.002, indicating that the alternative “Naltrexone will block contextual processing” can be accepted at the significance level of 0.005. Each outcome-analysis node links to one outcome node, one method node and multiple outcome-group nodes.
-
Note that the different types of group nodes for a study could represent the same participant group with different information. According to AACT, using a single group to uniquely represent a participant group in the study is impossible due to the complicated designs of clinical trials. Therefore, we followed AACT and used different types of group nodes to represent different types of information of the clinical trials.
Embedding analysis
We conducted an embedding analysis to evaluate the quality of \(\mathop {\mathtt {CTKG}}\limits\) and demonstrate its utilities in various important applications. We applied TransE 13, a state-of-the-art graph embedding method, to generate embeddings for nodes in \(\mathop {\mathtt {CTKG}}\limits\). These computable embeddings can benefit various downstream tasks. For example, we could establish similarities among nodes using their embeddings. The similarities enable fast retrieval of nodes corresponding to similar medical entities and could facilitate applications such as drug repurposing and similar study search, as will be discussed below. Note that TransE generates node embeddings based on the topology of the graph (i.e., node attributes are not considered). We used the TransE implementation in DGL-KE 14, which is a high-performance python library on top of the Deep Graph Library (DGL 15). DGL is a python library for deep learning on graphs and enables training models on large-scale graphs in a convenient way. Based on DGL, DGL-KE provides many popular KG embedding algorithms like TransE for users to learn KG embeddings that can be used for many different applications 9. Other KG embedding methods are also applicable for the following analyses. Detailed information on KG embedding methods is available in a survey 16.
\(\mathop {\mathtt {CTKG}}\limits\) for drug repurposing
In this analysis, we evaluated the utilities of \(\mathop {\mathtt {CTKG}}\limits\) for drug repurposing—a strategy to identify new therapeutic indications for existing drugs 17. Particularly, we assessed if the high similarities between the condition node embeddings and drug-term node embeddings indicate the high potential of the corresponding drugs in treating the conditions. For the evaluation, we calculated the cosine similarities between all the condition nodes and drug-term nodes, and identified the top-10 most similar pairs. Among these 10 pairs, we found that 5 of them have evidence demonstrated by the literature indicating potential repurposability, as presented in Table 3. For example, the condition node “Diabetes Mellitus, Type 2” has a similarity 0.597 with the drug-term node “Benzoates”; as demonstrated in the literature 18, Alogliptin Benzoates, an agent of Benzoates, is now available for the treatment of Type 2 Diabetes. Similarly, the condition node “Lung Neoplasms” has a similarity 0.574 with the drug-term node “Triterpenes”, and as demonstrated in the literature 19, Triterpenes have anti-cancer properties against Lung Neoplasms. Please note the average similarity between condition nodes and drug-term nodes is -0.032, and thus the above similarities are significantly high. In addition, the above drugs are not studied for their highly-similar conditions in any \(\mathop {\mathtt {CTKG}}\limits\) studies (i.e., no existing edges connecting the condition nodes and drug-term nodes). Thus, the above results demonstrate the utilities of \(\mathop {\mathtt {CTKG}}\limits\) for drug repurposing. Other similar condition and drug-term node pairs, for example, “Squamous Cell Carcinoma of Head and Neck” and “Naloxone” with similarity 0.565, and “Lung Neoplasms” and “Uric Acid” with similarity 0.564, may enable new hypothesis generation for innovative investigation and findings.
\(\mathop {\mathtt {CTKG}}\limits\) for similar medical entity retrieval
In this analysis, we evaluated whether \(\mathop {\mathtt {CTKG}}\limits\) enables high-quality node embeddings for similar medical entity retrieval tasks. Particularly, we focused on the retrieval of similar studies, and the retrieval of similar conditions, drugs, adverse events and outcomes. All these retrieval tasks are common and useful in designing new clinical trials 20.
Similar study retrieval
\(\mathop {\mathtt {CTKG}}\limits\) can support the search and retrieval of similar studies. To demonstrate this, we first identified the top-5 most similar pairs of study nodes using cosine similarity over their embeddings. In each identified pair, we randomly selected one study node, and identified its top-5 most similar study nodes. Table 4 presents the selected study nodes and their top-5 most similar study nodes. As presented in Table 4, the identified similar studies all investigated similar drugs or conditions. For example, study “NCT00795769” and its top-5 most similar studies investigated the prevention of the side effects caused by the stem cell transplant, or conditions that could be treated by the stem cell transplant; study “NCT01431274” and its top-5 most similar studies all investigated the therapies for the Chronic Obstructive Pulmonary Disease (COPD). These results show the utilities of \(\mathop {\mathtt {CTKG}}\limits\) for retrieving similar studies, which could facilitate new clinical trial design.
Similar medical concept retrieval
\(\mathop {\mathtt {CTKG}}\limits\) can also support the retrieval of other similar medical concepts. To demonstrate this, we identified the top-10 most similar pairs of condition nodes, drug-term nodes, adverse-event nodes, and standard-outcome nodes, as in Tables 5, 6, 7, and 8, respectively, using cosine similarities over their embeddings. As presented in Table 5, the identified similar condition nodes all share some commonalities. For example, condition node “Nephritis” and “Lupus Nephritis” have a similarity 0.997 (average condition similarity is 0.331), and Lupus Nephritis is a common sub-type of Nephritis. We also found a similar trend in Table 6, 7 and 8. For example, durg-term nodes “ABT-267” and “Macrocyclic Compound” have a similarity 0.997 (average drug-term similarity is 0.254), and both drugs could be used to treat Hepatitis C Virus (HCV) infection 21,22. In addition, the two drugs are studied together in multiple studies such as NCT01458535, NCT01464827 and NCT01563536. In Table 7 for adverse-event nodes, “Blood Luteinising Hormone” is very similar to “Uterus Myomatosus” (cosine similarity 0.995; average adverse-event similarity is 0.329). The high similarity could be due to the fact that Luteinising Hormone can affect the development and growth of Uterus Myomatosus by stimulating the production of estrogen 23. Note that Luteinising Hormone and Uterus Myomatosus are not present together in any of the \(\mathop {\mathtt {CTKG}}\limits\) studies; therefore, such similar pairs demonstrate the effectiveness of \(\mathop {\mathtt {CTKG}}\limits\) on retrieving similar/related adverse events. In Table 8 for standard-outcome nodes, “Aspartate Aminotransferase” is very similar to “Alanine Aminotransferase” in their embeddings (cosine similarity 0.986; average standard-outcome similarity is 0.315), and both measure the amount of two enzymes made by liver in the blood and can be tested to check the liver damage. These results demonstrate that \(\mathop {\mathtt {CTKG}}\limits\) can facilitate the search and retrieval of medical entities in the context of clinical trials that carry similar/related information.
\(\mathop {\mathtt {CTKG}}\limits\) for other applications
\(\mathop {\mathtt {CTKG}}\limits\) could also enable other potential applications such as adverse drug event prediction and outcome prediction, etc. Specifically, for the adverse drug event prediction, we could employ knowledge reasoning methods24 over \(\mathop {\mathtt {CTKG}}\limits\), and infer new adverse events of drugs using the existing or predicted paths from drug-term nodes to adverse-event nodes in \(\mathop {\mathtt {CTKG}}\limits\). For the outcome prediction, we could employ link prediction methods9,13 to infer new edges between study nodes and outcome nodes based on the existing ones in \(\mathop {\mathtt {CTKG}}\limits\). Overall, \(\mathop {\mathtt {CTKG}}\limits\) could facilitate new knowledge discovery and benefit the design of new clinical trials, and also improve the success rate of future clinical trials. We released the code for drug repurposing and similar node retrieval (“Data availability” section). For the link prediction applications, please refer to the examples in DGL 25 for a concrete implementation.
Discussion
In this manuscript, we presented and released a new knowledge graph \(\mathop {\mathtt {CTKG}}\limits\) for clinical trials. We also described our methods in generating \(\mathop {\mathtt {CTKG}}\limits\). We demonstrated the potential utilities of \(\mathop {\mathtt {CTKG}}\limits\) in drug repurposing and similarity search, among others, via embedding analysis over \(\mathop {\mathtt {CTKG}}\limits\). Currently, \(\mathop {\mathtt {CTKG}}\limits\) only includes studies that have both drug interventions and reported outcomes. However, incomplete studies (e.g., studies not started or without reported outcomes), and studies without drug interventions (e.g., studies for medical devices) could also contain valuable knowledge for the design of future clinical trials. Therefore, we will enrich \(\mathop {\mathtt {CTKG}}\limits\) with more studies in the future research. In addition, current \(\mathop {\mathtt {CTKG}}\limits\) does not contain all the important information for drug discovery and development. For example, \(\mathop {\mathtt {CTKG}}\limits\) does not have the interactions between drugs/molecules and proteins/diseases, nor the interactions among proteins. Missing such information may limit the potential of \(\mathop {\mathtt {CTKG}}\limits\) for a much wider range of applications (e.g., to predict if a new molecule for a disease can survive from clinical trials). In the future research, we will align \(\mathop {\mathtt {CTKG}}\limits\) with other knowledge bases10,26,27 and integrate more and diverse information into \(\mathop {\mathtt {CTKG}}\limits\) to enable more applications using \(\mathop {\mathtt {CTKG}}\limits\). Moreover, \(\mathop {\mathtt {CTKG}}\limits\) embeds rich textual information (e.g., title and description) and heterogeneous data types (e.g., numerical, categorical and textual data) as node attributes, which encourages a much borader spetrum of research (e.g., deep graph embedding, link prediction) and more complicated methods to be developed to leverage such information for better translational clinical trial design. We will also investigate attribute-sensitive KG embedding methods 28 to better leverage \(\mathop {\mathtt {CTKG}}\limits\).
Methods
\(\mathop {\mathtt {CTKG}}\limits\) represents each medical entity (e.g., a clinical trial, also referred to as a study; a drug; an adverse event) as a single node. To develop \(\mathop {\mathtt {CTKG}}\limits\), we extracted the medical entities from the Access to Aggregate Content of ClinicalTrials.gov (AACT) database 5. We then normalized multiple expressions of a same medical entity into a single one. Figure 2 presents the overview of development of \(\mathop {\mathtt {CTKG}}\limits\).
Clinical trials data
The clinical trials data in \(\mathop {\mathtt {CTKG}}\limits\) is collected from the AACT database. AACT is a publicly available relational database, which contains the information of every clinical trial registered in ClinicalTrials.gov, and is updated on a daily basis. In AACT, each clinical trial, also referred to as a study, is associated with a unique National Clinical Trial (NCT) ID, and all the information of a clinical trial is stored in 45 different tables. For example, information representing the medications, procedures and other actions provided or conducted in a clinical trial is stored in two tables: “interventions” and “browse interventions”; information representing the measurements used to evaluate the safety and efficacy of drugs or procedures studied in clinical trials is stored in the table “outcomes.” All the tables and their schemas are publicly available 29. Until July 2020, 344,500 clinical trials have been registered in ClinicalTrials.gov and included in AACT. We selected all the studies that have drug interventions, conditions and outcome analyses into \(\mathop {\mathtt {CTKG}}\limits\). Specifically, we excluded 232,274 studies that do not have drug interventions, and 103,047 studies that do not have outcome analyses. Among 9,179 remaining studies, we excluded 969 studies without the conditions, resulting 8210 studies in \(\mathop {\mathtt {CTKG}}\limits\). Note that we did not consider clinical trials that are not on drug inventions, such as physical therapies, behavioral therapies or medical devices. We will update \(\mathop {\mathtt {CTKG}}\limits\) with more studies in the future as new studies on drug interventions become available.
Note that \(\mathop {\mathtt {CTKG}}\limits\) does not include all the tables in AACT. For example, \(\mathop {\mathtt {CTKG}}\limits\) does not include tables such as “Sponsors”, “Overall officials” and “Result contacts” because they are not directly related to the design and results of clinical trials, and including them may not significantly benefit the knowledge graph in analyzing the relations among medical entities. Other AACT tables such as “Provided documents” and “Documents” contain the links to detailed study protocols, informed consent forms and statistical analysis plans, etc. These documentations have rich textual information that might be complementary to the structural relations represented by \(\mathop {\mathtt {CTKG}}\limits\). However, such information is highly specific to each individual clinical trial, and does not help establish new relations across clinical trials if no natural language processing is applied first, which by itself is highly non-trivial. Therefore, \(\mathop {\mathtt {CTKG}}\limits\) does not include such tables; instead, \(\mathop {\mathtt {CTKG}}\limits\) uses AACT’s original study IDs so that all such information can still be retrieved from AACT if needed. \(\mathop {\mathtt {CTKG}}\limits\) does not include other AACT tables such as “Calculated values”, “Design outcomes” and “Design group interventions” because information in such tables is already included in other tables that \(\mathop {\mathtt {CTKG}}\limits\) includes. Table 9 summarizes the AACT tables that are included and are not included in \(\mathop {\mathtt {CTKG}}\limits\).
Adverse event normalization
In AACT, we could find the adverse events (\(\mathop {\mathtt {AE}}\limits\)) , represented by AE terms, happened among the participants in the “reported events” table. Many \(\mathop {\mathtt {AE}}\limits\) terms listed in the table could be mapped to the Medical Dictionary for Regulatory Activities (\(\mathop {\mathtt {MedDRA}}\limits\) \(\circledR\)30). \(\mathop {\mathtt {MedDRA}}\limits\) \(\circledR\) is the international medical terminology developed under the auspices of the International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use (ICH). More specifically, we found 28,677 unique \(\mathop {\mathtt {AE}}\limits\) terms in which 13,995 terms could be directly mapped to the \(\mathop {\mathtt {MedDRA}}\limits\) dictionary. In \(\mathop {\mathtt {CTKG}}\limits\), such terms are also referred to as \(\mathop {\mathtt {MedDRA}}\limits\) terms. We normalized the remaining 14,682 \(\mathop {\mathtt {AE}}\limits\) terms that are not in the \(\mathop {\mathtt {MedDRA}}\limits\) \(\circledR\) to \(\mathop {\mathtt {MedDRA}}\limits\) terms as follows:
-
We removed parenthesized contents (e.g., “Altered pitch perception (pitch seemed lower)”). The contents in parentheses are typically explanations or afterthoughts so removing them would not significantly affect the major meanings.
-
We removed words or phrases that specify the auxiliary information (e.g., “left”, “right”, “Baseline Phase”) or the time frame (e.g., “for 12 hours”) of adverse events. We observed that these words or phrases are study-specific, and not in the \(\mathop {\mathtt {MedDRA}}\limits\) terms. For example, by removing the phrase “Baseline Phase”, the \(\mathop {\mathtt {AE}}\limits\) term “Throat tightness - Baseline Phase” can be normalized to the \(\mathop {\mathtt {MedDRA}}\limits\) term “Throat tightness”. The phrase “Baseline Phase” is given to specify the initial phase of assessment involving collection of initial data in the study, and thus unrelated to the adverse event itself.
-
We removed the stop words and lemmatized \(\mathop {\mathtt {AE}}\limits\) terms using the NLTK library31, and Stanza NLP Library32, respectively.
-
We mapped an \(\mathop {\mathtt {AE}}\limits\) term to its most similar \(\mathop {\mathtt {MedDRA}}\limits\) term if their edit distance is less than 4. For example, the adverse event term “Cholecyctitis” will be normalized to the \(\mathop {\mathtt {MedDRA}}\limits\) term “Cholecystitis”. This process can correct simple misspellings.
After each step above, if the normalized \(\mathop {\mathtt {AE}}\limits\) term is a \(\mathop {\mathtt {MedDRA}}\limits\) term, we will stop the normalization. With the above normalization, we successfully normalized 7,296 \(\mathop {\mathtt {AE}}\limits\) terms to \(\mathop {\mathtt {MedDRA}}\limits\) terms. In total, we got 15,976 unique \(\mathop {\mathtt {MedDRA}}\limits\) terms and had 7,393 \(\mathop {\mathtt {AE}}\limits\) terms that cannot be normalized.
In order to construct a one-to-one mapping between the adverse events and the \(\mathop {\mathtt {MedDRA}}\limits\) terms, we used the \(\mathop {\mathtt {MedDRA}}\limits\) dictionary to further group multiple \(\mathop {\mathtt {MedDRA}}\limits\) terms of the same adverse event into a unique \(\mathop {\mathtt {MedDRA}}\limits\) term. According to the definition of \(\mathop {\mathtt {MedDRA}}\limits\), each \(\mathop {\mathtt {MedDRA}}\limits\) term is assigned to one of the five hierarchical levels33. Specifically, the \(\mathop {\mathtt {MedDRA}}\limits\) terms with the lowest level (i.e., level 1), which are used to communicate the adverse events in practice, could correspond to the same adverse event. For example, “Eye itching” and “Ocular itching” are two \(\mathop {\mathtt {MedDRA}}\limits\) terms with level 1 and represent the same event. Such \(\mathop {\mathtt {MedDRA}}\limits\) terms corresponding to the same event have a common parent, which is a \(\mathop {\mathtt {MedDRA}}\limits\) term with level 2 (e.g., “itchy eyes” in the above example). Therefore, we normalized each \(\mathop {\mathtt {MedDRA}}\limits\) term with level 1 to its linked \(\mathop {\mathtt {MedDRA}}\limits\) term with level 2. In total, we converted 15,976 \(\mathop {\mathtt {MedDRA}}\limits\) terms into 11,153 more abstract \(\mathop {\mathtt {MedDRA}}\limits\) terms. Each term among these 11,153 \(\mathop {\mathtt {MedDRA}}\limits\) terms and 7,393 non-\(\mathop {\mathtt {MedDRA}}\limits\) terms represents an adverse event, which is further represented as an adverse-event node in \(\mathop {\mathtt {CTKG}}\limits\). Note that due to the licensing restriction of \(\mathop {\mathtt {MedDRA}}\limits\) \(\circledR\), we didn’t specify which adverse-event nodes represent \(\mathop {\mathtt {MedDRA}}\limits\) terms in \(\mathop {\mathtt {CTKG}}\limits\) and only kept the terms as the attribute of adverse-event nodes.
Drug mentions and normalization
In AACT, the drugs used in studies (i.e., clinical trials) could be found in the intervention table, in which the “name” field stores the information about medicines and administrations used in each intervention. For example, we could find that the drug Naltrexone is used in the study NCT04322526 via its intervention “Naltrexone 50 Mg Oral Tablet.”
In \(\mathop {\mathtt {CTKG}}\limits\), we used Medical Text Indexer (MTI)34 to automatically extract drug mentions. MTI is developed by the National Library of Medicine (NLM) to recognize medical entities (e.g., anatomy, drugs and conditions) from plain text. We used this tool to extract drug mentions following 2 steps:
-
We used MTI to automatically recognize all the medical entities from the interventions.
-
We found drug entities from the medical entities recognized by MTI. Specifically, for each recognized entity, MTI will output its MeSH code if available. MeSH is a hierarchically-organized vocabulary from NLM to index and categorize biomedical and health-related information35. Given the MeSH code, we first identified entities with MeSH codes starting with character “D”, which indicates drug entities (e.g., D02.241.223.701.430 for Ibuprofen). After that we removed the entities not representing specific drugs by excluding those with the MeSH code D26.310 (drug combination), D26 (pharmaceutical preparations), D23.101 (biomarkers) and D26.255 (dosage forms). We also noticed that a few recognized entities were not associated with MeSH codes. For these entities, we did a manual check and identified the ones representing specific drugs.
After the above 2 steps, there were still 1,775 unique interventions in which MTI did not find any drug mentions. For these interventions, we did a manual search and identified the drugs mentioned. Eventually, from the intervention table, we found 3,487 mentioned drugs in total. Among these drugs, 860 (24.7%) of them are found manually. Most of the manually found drugs are investigational drugs (e.g., pf-06669571), or drugs mentioned in abbreviations (e.g., tvr and umec).
Besides the drugs in interventions, there were also drugs mentioned in the titles or descriptions of the study groups (e.g., event group). For example, from the title “tramadol/diclofenac 25/25”, we could find the drugs Tramadol and Diclofenac. We also extracted drugs mentioned in the titles or descriptions of study groups to generate a complete list of drug mentions. Specifically, we first used the above 2 steps to automatically extract the mentioned drugs in titles and descriptions of study groups. For groups that we did not find any drugs automatically, we manually searched their titles and descriptions, and identified the mentioned drugs. In the end, we found 4585 drug mentions from the interventions and the study groups.
From the drug mentions, we observed that one drug could be represented by different names. For example, the drug “losartan potassium” could be represented by its brand name “cozaar” or its generic name “losartan.” Therefore, we normalized the drug mentions found in texts to normalized terms. Specifically, we first used MTI to map all the 4585 drugs to their MeSH terms. For example, MTI could automatically map the drugs “losartan potassium”, “cozaar” and “losartan” to the MeSH term “losartan.” For the drugs that MTI can find their MeSH terms, the MeSH terms were used as their normalized terms. For the other drugs, if they are in abbreviations (e.g., tvr), we first found their full names (e.g., Telaprevir), and used the MeSH terms of their full names for normalization; if they are not in abbreviations, we used their generic names for normalization. We noticed that investigational drugs may not have generic names. For these drugs, their identifiers mentioned in studies (e.g., pf-06669571) were used as their normalized terms. After the normalization, the 4585 drug mentions were normalized to 2548 normalized terms. Each of the normalized term is represented as a drug-term node in \(\mathop {\mathtt {CTKG}}\limits\).
Statistical analysis method normalization
We observed that one statistical analysis method could be represented by different names in the table. For example, the method “paired t-test” could be represented as “paired t test”, “paired t-tests” and “paited t-test” in the table. Therefore, we normalized the names of the methods using the 3 steps as follows:
-
We preprocessed the method names from the table by removing the space and punctuation in the text.
-
We calculated the edit distance among the preprocessed names, and normalized the preprocessed names with edit distance less than 4 to a same normalized term. We also did a manual check to correct possible mis-normalization. For the names that will be normalized to a same term, we used the names with the highest frequency as the normalized term.
-
We further refined the normalized terms by merging the terms with the same words. We noticed that after the second step, there were still normalized terms that represent the same method with the same words but of different orders. For example, the normalized terms “pairedttest” and “ttestpaired” represent the same method “paired t-test” with the same words but of different orders. We manually merged such terms to the one with the highest frequency.
After all the steps, we normalized the 1,299 unique method names mentioned in the table to 907 normalized terms. Each of the normalized terms is represented as a method node in \(\mathop {\mathtt {CTKG}}\limits\).
Outcome extraction and outcome clustering
In AACT, the outcome measures used to test the effectiveness of the interventions could be found in the “title” or the “description” fields of the outcome table. Most of the titles in the outcome table are long phrases and could involve multiple standard outcome measures (e.g., in the title “Change From Baseline in Platelet Count and White Blood Cell Count”, where “Platelet Count” and “White Blood Cell Count” represent standard outcome measures). These standard outcome measures are common assessment tools that are used to assess the effectiveness of an intervention. The complex relations between the outcome titles and the standard outcome measures make it difficult to directly represent the outcomes with the extracted standard outcome measures. Therefore, we incorporated the identified standard outcome measures as nodes into \(\mathop {\mathtt {CTKG}}\limits\) and built connections between the outcome nodes and the involved standard-outcome nodes. Through such connections, we can infer which standard outcome measures are used in each study to assess the efficacy of interventions. We observed that some popular phrases within the titles or the descriptions of outcome records represent standardized assessment tools used to measure the outcome of clinical trials, for example, “Visual Analogue Scale” is a tool widely used as a measure for pain. Incorporating such standard outcome measures into the \(\mathop {\mathtt {CTKG}}\limits\) could enable the comparison on the outcome measurements across different studies, and also could provide a reference regarding the choice of standardized assessment tools in the design of clinical trials. Therefore, we extracted the phrases that could represent standard outcome measures as below:
-
We found the abbreviations and identified the definitions of abbreviations from the titles or the descriptions of the outcomes using the Schwartz-Hearst algorithm36. We observed in the titles that many standard outcome measures are associated with their corresponding abbreviations. For example, we could identify the abbreviation “BI” and the corresponding definitions “Bleeding Index” from the outcome name “Gingival Health Measured by Bleeding Index (BI).”
-
We kept only the definitions containing the following words: scale, index, score, test, questionnaire, value, count, inventory, assessment, level, rate. We observed that most standard outcome measures would contain such words (e.g., “Visual Analogue Scale”, “Social Responsiveness Scale”).
-
We manually normalized different variants of the same standard outcome measures and removed the extracted phrases that are not outcome measures. We also manually added some popular standard outcome measures (e.g., “Overall Survival”, “blood pressure”, “triglyceride”) that do not contain the above words or do not have any abbreviations.
All the extracted phrases are represented as the standard-outcome nodes in the \(\mathop {\mathtt {CTKG}}\limits\). In the end, we got 492 standard-outcome nodes from 50,342 outcome records (i.e., 56.96% over all the 88,386 outcome records), and connected the standard-outcome nodes with the corresponding outcome nodes.
With the extracted standard outcome measures, there were still more than 40% of the outcome nodes not connected to any standard-outcome nodes. Therefore, to aggregate similar outcome nodes, we also grouped all the outcome titles (including those containing the standard outcome measures) into several clusters. Specifically, we represented each outcome title using its term frequency-inverse document frequency (TF-IDF) vectors. We then grouped the TF-IDF vectors of outcome titles using the CLUTO37, a clustering toolkit, into 200 clusters. Each cluster is presented as a cluster-outcome node and has attributes describing the cluster size, that is, the number of outcomes within the cluster, and the most representative words of these outcomes. Specifically, for each cluster, the representative words of outcomes include 5 descriptive words and 5 discriminating words derived by CLUTO that can best describe or discriminate each cluster. Each word is associated with a percentage computed by CLUTO (details in its manual) which indicates the importance of this word with respect to describing or discriminating the cluster. We converted the descriptive words and the discriminating words as two attributes of each cluster, by combining the words and their corresponding percentages. For example, after clustering, one cluster-outcome node has these descriptive features: “circumference 56.4%, waist 43.0%, head 0.1%, abdominal 0.1%, change 0.1%”, and most outcome nodes connected with it have the titles related to “circumference” and “waist”, such as “Change in Waist-to-hip Ratio” and “Mean Change From Baseline in Waist Circumference”.
Data availability
The \(\mathop {\mathtt {CTKG}}\limits\) dataset and the code for the embedding analyses are released in GitHub: https://github.com/ninglab/CTKG.
References
Elkin, M. E. & Zhu, X. Predictive modeling of clinical trial terminations using feature engineering and embedding learning. Sci. Rep. 11, 1–12 (2021).
Friedman, L. M., Furberg, C. D., DeMets, D. L., Reboussin, D. M. & Granger, C. B. Fundamentals of Clinical Trials (Springer, 2015).
Sertkaya, A., Wong, H.-H., Jessup, A. & Beleche, T. Key cost drivers of pharmaceutical clinical trials in the united states. Clin. Trials 13, 117–126 (2016).
Wong, C. H., Siah, K. W. & Lo, A. W. Estimation of clinical trial success rates and related parameters. Biostatistics 20, 273–286 (2019).
Tasneem, A. et al. The database for aggregate analysis of ClinicalTrials.gov (AACT) and subsequent regrouping by clinical specialty. PLoS ONE 7, e33677 e33677 (2012).
Macak, M., Stovcik, M. & Buhnova, B. The suitability of graph databases for big data analysis: A benchmark. In International Conference on Internet of Things, Big Data and Security (IoTBDS) (2020).
Pujara, J., Miao, H., Getoor, L. & Cohen, W. Knowledge graph identification. In International Semantic Web Conference (ISWC) 542–557 (Springer, 2013).
Ma, Y., Crook, P. A., Sarikaya, R. & Fosler-Lussier, E. Knowledge graph inference for spoken dialog systems. In 2015 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) 5346–5350 (IEEE, 2015).
Ji, S., Pan, S., Cambria, E., Marttinen, P. & Yu, P. S. A survey on knowledge graphs: Representation, acquisition, and applications. IEEE Trans. Neural Netw. Learn. Syst. 33, 494–514. https://doi.org/10.1109/TNNLS.2021.3070843 (2022).
Ioannidis, V. N. et al. DRKG—Drug Repurposing Knowledge Graph for Covid-19. https://github.com/gnn4dr/DRKG/ (2020). Accessed: 2022-01-29.
Liu, H., Chi, Y., Butler, A., Sun, Y. & Weng, C. A knowledge base of clinical trial eligibility criteria. J. Biomed. Inform. 117, 103771 (2021).
Peng, B. et al.https://github.com/ninglab/CTKG/blob/main/Schema.pdf (2021). Accessed: 2022-01-29.
Bordes, A., Usunier, N., Garcia-Durán, A., Weston, J. & Yakhnenko, O. Translating embeddings for modeling multi-relational data. In Proceedings of the 26th International Conference on Neural Information Processing Systems, Vol. 2, 2787–2795 (Curran Associates Inc., 2013).
Zheng, D. et al. DGL-KE: Training knowledge graph embeddings at scale. In Proceedings of the 43rd International ACM SIGIR Conference on Research and Development in Information Retrieval, SIGIR ’20, 739–748 (Association for Computing Machinery, 2020).
Wang, M. et al. Deep graph library: A graph-centric, highly-performant package for graph neural networks. arXiv preprint arXiv:1909.01315 (2019).
Wang, Q., Mao, Z., Wang, B. & Guo, L. Knowledge graph embedding: A survey of approaches and applications. IEEE Trans. Knowl. Data Eng. 29, 2724–2743 (2017).
Pushpakom, S. et al. Drug repurposing: Progress, challenges and recommendations. Nat. Rev. Drug Discov. 18, 41–58 (2019).
Rendell, M., Drincic, A. & Andukuri, R. Alogliptin benzoate for the treatment of type 2 diabetes. Expert Opin. Pharmacother. 13, 553–563 (2012).
Chudzik, M., Korzonek-Szlacheta, I. & Król, W. Triterpenes as potentially cytotoxic compounds. Molecules 20, 1610–1625 (2015).
Hao, T., Rusanov, A., Boland, M. R. & Weng, C. Clustering clinical trials with similar eligibility criteria features. J. Biomed. Inform. 52, 112–120 (2014).
Feld, J. J. et al. Treatment of HCV with ABT-450/r-ombitasvir and dasabuvir with ribavirin. N. Engl. J. Med. 370, 1594–1603 (2014).
Pillaiyar, T., Namasivayam, V. & Manickam, M. Macrocyclic hepatitis C virus NS3/4A protease inhibitors: An overview of medicinal chemistry. Curr. Med. Chem. 23, 3404–3447 (2016).
Maruo, T., Ohara, N., Wang, J. & Matsuo, H. Sex steroidal regulation of uterine leiomyoma growth and apoptosis. Hum. Reprod. Update 10, 207–220 (2004).
Chen, X., Jia, S. & Xiang, Y. A review: Knowledge reasoning over knowledge graph. Expert Syst. Appl. 141, 112948 (2020).
Zheng, D. et al. Scalable graph neural networks with deep graph library. In Proceedings of the 14th ACM International Conference on Web Search and Data Mining 1141–1142 (2021).
Santos, A. et al. Clinical knowledge graph integrates proteomics data into clinical decision-making. bioRxiv https://doi.org/10.1101/2020.05.09.084897 (2020).
Xu, J. et al. Building a PubMed knowledge graph. Sci. Data 7, 205 (2020).
Wu, Y. & Wang, Z. Knowledge graph embedding with numeric attributes of entities. In Proceedings of The Third Workshop on Representation Learning for NLP 132–136 (Association for Computational Linguistics, 2018).
Clinical Trials Transformation Initiative. Improving Public Access to Aggregate Content of ClinicalTrials.gov. https://aact.ctti-clinicaltrials.org/. Accessed: 2022-1-28.
Brown, E. G., Wood, L. & Wood, S. The medical dictionary for regulatory activities (MedDRA). Drug Saf. 20, 109–117 (1999).
Bird, S., Klein, E. & Loper, E. Natural Language Processing with Python (O’Reilly UK Ltd., 2009).
Zhang, Y., Zhang, Y., Qi, P., Manning, C. D. & Langlotz, C. P. Biomedical and clinical English model packages for the stanza python NLP library. J. Am. Med. Inform. Assoc. 28, 1892–1899 (2021).
Yokotsuka, M., Aoyama, M. & Kubota, K. The use of a medical dictionary for regulatory activities terminology (MedDRA) in prescription-event monitoring in japan (J-PEM). Int. J. Med. Inform. 57, 139–153 (2000).
Mork, J. G., Jimeno-Yepes, A. & Aronson, A. R. The NLM medical text indexer system for indexing biomedical literature. In Proceedings of the First Workshop on Bio-Medical Semantic Indexing and Question Answering, a Post-Conference Workshop of Conference and Labs of the Evaluation Forum, vol. 1094 of CEUR Workshop Proceedings (eds Ngomo, A. N. & Paliouras, G.) (CEUR-WS.org, 2013).
Chapman, D. Advanced search features of PubMed. J. Can. Acad. Child Adolesc. Psychiatry 18, 58 (2009).
Schwartz, A. S. & Hearst, M. A. A simple algorithm for identifying abbreviation definitions in biomedical text. In Pac. Symp. Biocomput. 451–462 (World Scientific, 2003).
Karypis, G. Cluto: A clustering toolkit. Tech. Rep. 02-017, Dept. of Computer Science, University of Minnesota (2002).
Bourbon, J. R. & Farrell, P. M. Fetal lung development in the diabetic pregnancy. Pediatr. Res. 19, 253–267 (1985).
Yadav, P. et al. Experimental and theoretical investigation of synthesized pregnenolone derivatives via palladium catalyzed cross coupling reactions, their anticancer activity against lung cancer cells. J. Mol. Struct. 1245, 131115 (2021).
Pasquini, M. et al. Differences in depressive thoughts between major depressive disorder, ifn-\(\alpha\)-induced depression, and depressive disorders among cancer patients. J. Psychosom. Res. 65, 153–156 (2008).
Anestopoulos, I. et al. Surface active agents and their health-promoting properties: Molecules of multifunctional significance. Pharmaceutics 12, 688 (2020).
Lancaster, D., Lennard, L., Rowland, K., Vora, A. & Lilleyman, J. Thioguanine versus mercaptopurine for therapy of childhood lymphoblastic leukaemia: A comparison of haematological toxicity and drug metabolite concentrations. Br. J. Haematol. 102, 439–443 (1998).
Clissold, S. P., Todd, P. A. & Campoli-Richards, D. M. Imipenem/cilastatin. Drugs 33, 183–241 (1987).
Sumiyoshi, T., Jayathilake, K. & Meltzer, H. Y. The effect of melperone, an atypical antipsychotic drug, on cognitive function in schizophrenia. Schizophr. Res. 59, 7–16 (2003).
Zhang, Y., Liu, J., Chen, X. Q. & Chen, C. Y. O. Ubiquinol is superior to ubiquinone to enhance Coenzyme Q10 status in older men. Food Funct. 9, 5653–5659 (2018).
Wilck, M. B., Xu, Z. J., Stek, J. E. & Lee, A. W. Safety and immunogenicity of a fully-liquid DTaP-IPV-Hib-HepB vaccine (Vaxelis) in premature infants. Hum. Vaccines Immunother. 17, 191–196 (2021).
Croxtall, J. D. & Keating, G. M. Pneumococcal polysaccharide protein D-conjugate vaccine (Synflorix; PHiD-CV). Pediatr. Drugs 11, 349–357 (2009).
Bellandi, F. et al. Long-term efficacy and safety of propafenone and sotalol for the maintenance of sinus rhythm after conversion of recurrent symptomatic atrial fibrillation. Am. J. Cardiol. 88, 640–645 (2001).
Keller, J. E. & Neale, E. A. The role of the synaptic protein snap-25 in the potency of botulinum neurotoxin type A. J. Biol. Chem. 276, 13476–13482 (2001).
Graves, N., Birrell, F. & Whitby, M. Effect of pressure ulcers on length of hospital stay. Infect. Control Hosp. Epidemiol. 26, 293–297 (2005).
Weijmer, M. C. et al. Randomized, clinical trial comparison of trisodium citrate 30% and heparin as catheter-locking solution in hemodialysis patients. J. Am. Soc. Nephrol. 16, 2769–2777 (2005).
Chiusolo, P. et al. Luteinizing hormone-releasing hormone analogue: Leuprorelin acetate for the prevention of menstrual bleeding in premenopausal women undergoing stem cell transplantation. Bone Marrow Transplant. 21, 821–823 (1998).
Pabon, J. E. et al. Human skin contains luteinizing hormone/chorionic gonadotropin receptors. J. Clin. Endocrinol. Metab. 81, 2738–2741 (1996).
Tenke, P. et al. European and Asian guidelines on management and prevention of catheter-associated urinary tract infections. Int. J. Antimicrob. Agents 31, 68–78 (2008).
Meldgaard, T. & Brock, C. Diabetes and the gastrointestinal tract. Medicine 47, 454–459 (2019).
Vollenhoven, B. J., Lawrence, A. S. & Healy, D. L. Uterine fibroids: A clinical review. BJOG Int. J. Obstet. Gynaecol. 97, 285–298 (1990).
Williams, A. L. & Hoofnagle, J. H. Ratio of serum aspartate to alanine aminotransferase in chronic hepatitis relationship to cirrhosis. Gastroenterology 95, 734–739 (1988).
Smolen, J. S. et al. Validity and reliability of the twenty-eight-joint count for the assessment of rheumatoid arthritis activity. Arthritis Rheum. 38, 38–43 (1995).
National Heart, Lung, and Blood Institute. Types of Blood Tests. U.S. Department of Health and Human Services. https://www.nhlbi.nih.gov/health-topics/blood-tests#Types. Accessed: 2021-10-24.
Therasse, P. et al. New guidelines to evaluate the response to treatment in solid tumors. J. Natl Cancer Inst. 92, 205–216 (2000).
Baum, N., Dichoso, C. C. & Carlton, C. Blood urea nitrogen and serum creatinine. Urology 5, 583–588 (1975).
Radin, M. S. Pitfalls in hemoglobin a1c measurement: When results may be misleading. J. Gen. Intern. Med. 29, 388–394 (2013).
Inoue, E., Yamanaka, H., Hara, M., Tomatsu, T. & Kamatani, N. Comparison of disease activity score (DAS)28- erythrocyte sedimentation rate and DAS28- C-reactive protein threshold values. Ann. Rheum. Dis. 66, 407–409 (2007).
Acknowledgements
\(\mathop {\mathtt {MedDRA}}\limits\) \(\circledR\) trademark is registered by ICH. We thank Michael Lin for his efforts in constructing the \(\mathop {\mathtt {CTKG}}\limits\) web portal.
Funding
This project was made possible, in part, by support from the National Science Foundation under Grant Number IIS-1855501. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the funding agencies.
Author information
Authors and Affiliations
Contributions
X.N. and G.K. conceived the research. All the authors designed the research; Z.C. and B.P. conducted the research, including data curation, formal analysis, methodology design and implementation, result analysis and visualization; X.N. supervised and mentored Z.C. and B.P.; Z.C. and B.P. drafted the original manuscript; X.N. edited and revised the manuscript; V.I.L., M.L. and G.K. provided comments; all authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Z., Peng, B., Ioannidis, V.N. et al. A knowledge graph of clinical trials (\(\mathop {\mathtt {CTKG}}\limits\)). Sci Rep 12, 4724 (2022). https://doi.org/10.1038/s41598-022-08454-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-08454-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.