Abstract
Glioblastoma (GBM) is a lethal brain cancer known for its potent immunosuppressive effects. Loss of Methylthioadenosine Phosphorylase (MTAP) expression, via gene deletion or epigenetic silencing, is one of the most common alterations in GBM. Here we show that MTAP loss in GBM cells is correlated with differential expression of immune regulatory genes. In silico analysis of gene expression profiles in GBM samples revealed that low MTAP expression is correlated with an increased proportion of M2 macrophages. Using in vitro macrophage models, we found that methylthioadenosine (MTA), the metabolite that accumulates as a result of MTAP loss in GBM cells, promotes the immunosuppressive alternative activation (M2) of macrophages. We show that this effect of MTA on macrophages is independent of IL4/IL3 signaling, is mediated by the adenosine A2B receptor, and can be pharmacologically reversed. This study suggests that MTAP loss in GBM cells may contribute to the immunosuppressive tumor microenvironment, and that MTAP status should be considered for characterizing GBM immune states and devising immunotherapy-based approaches for treating MTAP-null GBM.
Similar content being viewed by others
Introduction
Immunotherapy has become a mainstay of cancer treatment and has reshaped the way we understand and approach certain cancer types1,2. Despite recent progress, however, the promise of immunotherapy-based approaches for treating brain tumors, in particular high-grade glioblastoma (GBM), remains to be fully realized3,4,5. GBM is the most common and lethal brain tumor, with a dismal median survival of 12–15 months from the time of diagnosis6. It is also characterized by its profoundly immune-suppressive nature7,8. It has been well-established that GBM cells actively employ multiple strategies to escape immune surveillance and to create an immunosuppressive microenvironment9,10. As such, to fully harness the power of immunotherapy for GBM requires better understanding and more effective strategies for addressing the tumors’ immuno-suppressive countermeasures.
Recent genomic studies have provided insight into the molecular mechanisms of GBM pathogenesis, revealing many commonly mutated genes in tumor cells11,12. Further studies, in both glioma and other types of cancer, have drawn association between genetic alterations and tumor evasion of immune surveillance, providing rationale for tailoring treatments to the cancer cells’ genetic composition13,14,15. As an example, recent discoveries have linked IDH1 mutations in glioma to immune evasion through interference of immune activation pathways, providing opportunities to develop highly specific immunological treatments16,17,18,19.
One of the most common genetic alterations in GBM, occurring in approximately 45% of all cases, is the homozygous deletion or epigenetic silencing of methylthioadenosine phosphorylase (MTAP)11,20. We have recently demonstrated that MTAP deletion is associated with increased tumorigenesis and with shortened disease-free survival in GBM patients21. MTAP is a metabolic enzyme that functions in the salvage pathway of adenine and methionine, and loss of MTAP results in the accumulation of its direct substrate, methylthioadenosine (MTA), which can be released into the tumor environment22,23,24,25. Aberrantly accumulated MTA is known to be functionally active within cells as an inhibitor of methyltransferases23,24, including interfering with intracellular protein methylation in T cells26, as well as acting on adenosine receptors on the cell surface of melanoma cell lines27. Studies of pathogen-induced host inflammatory responses have linked MTA to downregulation of TNFα production by macrophages through engaging adenosine receptors28, and have revealed a role of MTA in controlling the host inflammatory response29, such that MTA has been used as an immunosuppressive drug for treating colitis, liver inflammation, brain inflammation and autoimmunity in animal models30,31,32.
In this study, we investigated the link between MTAP loss in GBM cells and the GBM microenvironment. We reveal that in GBM tissues, low expression of MTAP (due to deletion or epigenetic silencing) is associated with immune cell populations indicative of an immunosuppressive state. We provide evidence that MTAP loss-induced MTA accumulation stimulates M2 alternative macrophage activation. We illustrate that this effect of MTA on macrophages is independent of interleukin (IL)-4/IL-13 signaling, is mediated by the adenosine A2B receptor, requires STAT3, and is distinct from the actions of adenosine. Finally, we show that the MTA-induced alternative macrophage activation can be pharmacologically reversed. These results provide a basis for pursuing adenosine receptor signaling as a component of tumor-mediated immunosuppression and potential target in MTAP-null GBM.
Results
Loss of MTAP expression is associated with an immunosuppressive molecular profile in GBM
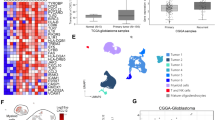
To explore the association of MTAP expression with the immune cell profile in GBM we employed CIBERSORT, a method for deconvoluting gene expression data and estimating immune cell fractions in human cancers33,34,35. This method has been successfully used for analyzing a large number of samples across multiple cancer types34, is readily compatible with the microarray-based platform of the largest available GBM expression dataset, and gives unique consideration to data normalization and noise control, which is essential when analyzing heterogeneous and diffusive tumors, such as GBM36. We applied this analysis to the GBM dataset from TCGA (385 GBM patient samples)11 to examine the link between MTAP loss and the GBM immune microenvironment. As MTAP alteration can occur through homozygous gene deletion or epigenetic silencing, we compared the two groups of patients with the highest and lowest MTAP expression (upper and lower quartiles, n = 96 each) (Supplementary Fig. S1). This revealed reduced fractions of activated CD4 T cells and increased resting CD4 T cells in tumors with low MTAP expression (Fig. 1A, Supplementary Fig. S1). Additionally, two types of innate immune cells were identified as differentially represented between these two groups of tumors. There was a lower fraction of γδT cells (Fig. 1B), and a significantly higher fraction of M2 macrophages in the low-MTAP expressing tumors (Fig. 1C).
MTAP loss is correlated with an immunosuppressive gene signature in GBM samples. (A) Comparison of upper and lower MTAP expression quartiles (n = 96 each category) using CIBERSORT analysis of GBM gene expression microarray data shows lower activated CD4 T cells in tumors with low MTAP expression, t test P value < 0.03. (B) GBM samples with the lowest quartile MTAP expression have fewer gamma delta T cells, t test P value = 0.0002. (C) GBM samples with low MTAP expression have higher proportions of M2 macrophages, t test P value = 0.0064. (D) Heatmap showing expression of 239 immune signature genes that are differentially expressed between the lower and upper MTAP expression quartiles in GBM patients. The gene list for the heatmap was derived from 782 immune-specific genes identified by Charoentong et al. Only 641 of those 782 genes were measured in the TCGA GBM microarray data set. Of the 641 genes in the GBM dataset, 239 genes were expressed at significantly different levels (Benjamini–Hocbberg corrected t test P value < 0.0185) in samples with low MTAP expression compared to samples with high MTAP expression. All differentially expressed genes were included in the heatmap. In order to classify the upregulated and downregulated genes identified, each clustered gene set was analyzed using data from the immunologic genome project (https://www.immgen.org), which uses a compendium of microarray data (366 microarrays/37 studies) from specific immune cell types to identify which immune cells express the genes of interest. Our analysis revealed two unique expression patterns. Genes expressed at higher levels in the low-MTAP samples are more representative of innate immune cells such as macrophages and monocytes, while genes downregulated in low-MTAP cells are more representative of activated CD4 T cells and innate lymphocytes.
To corroborate the findings from CIBERSORT analysis, we utilized a second recently developed platform for identifying individual immune cell types, which is based on a dataset of 366 microarrays compiled from multiple independent studies37. This data set was previously used to generate a list of 782 genes that are specific to immune cell subpopulations (i.e. not expressed in tumor cells or normal tissue)37. We analyzed this list of genes in the TCGA GBM dataset and found that among the 641 of these genes for which expression data was available, 239 of them were differentially expressed between the MTAP-low and MTAP-high GBM populations (upper quartile vs lower quartile, n = 96 each group, Benjamini–Hochberg corrected p-value < 0.0185). Hierarchical clustering revealed two groups of genes that could be evaluated to compare immune cell types upregulated and downregulated in MTAP deficient GBM (104 genes upregulated in MTAP-low tumors, 135 genes downregulated in MTAP-low tumors) (Fig. 1D; Supplementary Table 1). Using a database compiled by the Immunologic Genome Project37, we found that the genes upregulated in low-MTAP tumors were predominantly associated with macrophages and monocytes, while those genes downregulated in low-MTAP tumors represent activated CD4 T cells and innate lymphocytes (Fig. 1D; Supplementary Figs. S2–S4). Collectively, the findings from various analyses of the gene expression profiles in GBM samples with low versus high MTAP expression suggest a reduction in immune-reactive T cells38 and an increase in immunosuppressive M2 macrophages39 in samples with low MTAP expression, opening the possibility that MTAP deficiency is linked to the immunosuppressive GBM microenvironment. Of note, CIBERSORT and Immunologic Genomic project analyses revealed differences in expression of several immune cell types including plasma cells, memory B cells, NK cells, and dendritic cells. While all these differences in the tumor immune profile may be linked or regulated by a similar mechanism and warrant additional investigation, further work presented in this study focuses on macrophage regulation as a model for MTAP deficiency regulating the immune microenvironment.
MTA stimulates alternative activation of macrophages
Macrophages are the most abundant immune cell type in GBM, representing as many as half of all cells in the tumor mass40, where they are known to play an immunosuppressive/pro-tumor role39,41. Macrophages are traditionally classified as M1 (inflammatory) or M2 (anti-inflammatory). M2 macrophages are classically stimulated by cytokines IL-4, IL-13, IL-10, and M-CSF and are associated with reduction of inflammatory signals, resolution of the immune response, wound healing, and tumor promotion42,43. Higher numbers of M2 tumor-associated macrophages have been linked to poor cancer prognosis43,44. Upregulated numbers of M2 macrophages in MTAP deficient GBM samples from our data analysis suggests a more immunosuppressive environment with reduction of inflammatory signals and decreased antigen presentation, potentially contributing to the decreased numbers of activated CD4 lymphocytes observed in these samples as well.
Macrophages can be regulated by adenosine signaling45, which potentiates the effect of cytokines in promoting alternative macrophage activation through adenosine A2 receptors46. This led us to hypothesize that the MTAP substrate, MTA, an adenosine analogue that is secreted by MTAP deleted cells, impacts macrophage polarization through adenosine receptor signaling. In fact, studies focusing on pathogenic mechanism and treatment of pathogen-induced host inflammatory responses have found that MTA can suppress lipopolysaccharide (LPS)-induced expression of inflammatory response genes, such as TNF, via adenosine A2 receptors28,47.
To investigate the impact of MTA on macrophage activation, we utilized the well-established murine RAW 264.7 macrophage cell line model46,48. As a control, RAW 264.7 cells were treated with M2 macrophage-inducing Th2-type cytokines, IL-4 and IL-13, which drove the cells toward alternative (M2) activation as expected, demonstrated by the upregulated expression of Arginase 1 (Arg1), a canonical marker for M2 macrophages49 (Fig. 2A). When MTA was added to the IL-4/IL-13 treatment, induction of Arg1 expression was dramatically augmented (Fig. 2B). Remarkably, when MTA was used as a single agent to treat the cells, upregulated expression of Arg1 was also observed (Fig. 2C). This upregulation of Arg1 transcription was not limited to RAW 264.7 cells as it was also observed in the murine BV-2 microglial cell line, the human THP-1 monocytic cell line, and in primary human monocytes (Fig. 2D–F). This rise in transcription was followed by a rise in Arginase-1 protein levels (Fig. 2G). Notably, despite similarly upregulating Arg1 expression, the overall effect of MTA on macrophages was distinct from that of IL-4/IL-13, as treatment of cells with MTA attenuated the IL-4/IL-13-induced expression Pparg and Mrc1, marker genes associated with the M2a macrophage subtype (Supplementary Fig. S5A,B)50,51. Instead, MTA activated the expression of Vegfa, a classic feature of the M2d subtype and tumor associated macrophages (TAMs) (Fig. 2H)51. This phenotype was validated in human THP-1 cells and primary human monocytes differentiated into macrophage lineage (Fig. 2I,J). Prompted by this finding, we revisited the GBM gene expression data (TCGA) and confirmed that in GBM patients VEGFA expression was significantly higher in samples with low MTAP expression (Fig. 2K). Collectively, these results suggest that MTA promotes alternative macrophage activation resembling the M2d subtype and that this process is likely occurring in MTAP deficient GBMs.
MTA promotes upregulation of M2 macrophage marker genes. (A) RAW 264.7 cells were treated with IL-4 and IL-13 individually or in combination (5 ng/mL for each cytokine) for 12 h before total RNA was prepared for Arg1 expression analysis (all gene expression was measured by RT-qPCR, n ≥ 5). (B) RAW 264.7 cells were treated with the indicated cytokines as in (A), together with different doses of MTA, after which relative Arg1 expression was determined as above, n ≥ 5. (C) RAW 264.7, (D) BV-2, and (E) THP-1 cells were treated with different doses of MTA for 12 h and Arg1 expression was measured, n ≥ 5. (F) Primary human monocytes were differentiated to macrophages with M-CSF and treated with MTA or IL-4/IL-13 for 24 h and Arg1 expression levels were measured. Though the impact of cytokines on these cells was less robust than on RAW 264.7 cells, it is notable that MTA treatment still resulted in measurable differentiation while IL-4/IL-13 did not. (G) Immunoblots confirmed the MTA-stimulated expression of ARG1; “d” denotes “days”. (H) RAW 264.7, (I) THP-1, and (J) primary human monocyte cells were treated with MTA for 12 h, and total RNA was isolated for measuring VEGFA expression, n ≥ 4. In Fig. (J): “100” denotes “100 uM”, and “200” denotes “200 uM”. (K) TCGA human GBM microarray data were used for analyzing VEGFA expression in tumors with low or high MTAP expression, n = 96 patients per group. All experiments were repeated independently. All statistical comparisons were performed using ANOVA for group comparisons or unpaired student’s t-test for individual comparisons; error bars indicate mean ± SD; *P < 0.05, **P < 0.005, ***P < 5 × 10–4, n.s. = not significant.
To validate that the phenotype induced by exogenously administered MTA is relevant to the amount of MTA produced by tumor cells, we generated an isogenic MTAP-null derivative of the murine GBM cell line CT-2A. As expected, CRISPR-mediated homozygous deletion of Mtap in CT-2A cells led to the accumulation of MTA in the culture media (Supplementary Fig. S5C). We then tested the response of macrophages to spent media from the Mtap-null cell line as a way of simulating the tumor microenvironment. Exposure of RAW 264.7 cells to spent media from Mtap-null CT-2A cells indeed induced Arg1 expression, while exposure to the spent media from parental CT-2A (Mtap wildtype) cells had a minimal effect on Arg1 expression and the addition of exogenous MTA to the conditioned media was able to stimulate an alternative activation response (Supplementary Fig. S5D). Together, these results support the hypothesis that MTA in the extracellular compartment of MTAP-null GBM cells can work in concert with known M2 macrophage-stimulating cytokines in altering tumor associated macrophages, and/or directly promoting the alternative activation of macrophages as a single agent.
Regulation of macrophage activation by MTA requires the adenosine A2B receptor and STAT3
Previous work has shown that the action of adenosine in modulating the innate immune response to cytokine signaling is achieved through adenosine A2 receptors51. We tested whether the effects of MTA are also mediated through these receptors using specific antagonists of A2A and A2B receptors. We found that an A2A receptor antagonist, Istradefylline, had only minimal effect on MTA-induced Arg1 expression (Fig. 3A). In contrast, an antagonist of the A2B receptor, PSB 0788, potently attenuated the expression of Arg1 induced by MTA (Fig. 3B,C). Furthermore, PSB 0788 also blocked the induction of Arg1 expression by the spent media of MTAP-null GBM cells (Supplementary Fig. S5D), and attenuated other MTA-stimulated M2 marker genes, including Vegfa, Timp1, and IL10, while countering the inhibitory effect of MTA on Mrc1 expression (Fig. 3D–F; Supplementary Fig. S5B)46,48. The A2A receptor inhibitor, Istradefylline, again had negligible effects on any of these target genes’ transcription in response to MTA (Supplementary Fig. S6A–C). In order to rule out potential off-target effects of the pharmacologic inhibitor and confirm involvement of the A2B receptor in the phenomena we observed, we used CRISPR/CAS9 to genetically knock out the A2B receptor gene, Adora2b (Supplementary Fig. S7). We saw decreased basal expression levels of Arg1 and Timp1 and a greatly dampened response to MTA (Supplementary Fig. S6D,E). Collectively, these results suggest that MTA acts through the adenosine A2B receptor and that this receptor is involved in regulating macrophage activation states.
The impact of MTA on M2 macrophage marker genes is blocked by inhibition of the A2B adenosine receptor and STAT3. (A,B) RAW 264.7 cells were treated with MTA for 12 h and simultaneously with (A) A2A receptor antagonist Istradefylline or (B) A2B receptor antagonist PSB 0788 and expression of Arg1 was determined by RT-qPCR, error bars indicate mean ± SEM, n ≥ 5. (C) The BV-2 microglial cell line was treated with MTA with or without A2B receptor antagonist PSB 0788 for 12 h and Arg1 expression was measured by RT-qPCR, error bars indicate mean ± SEM, n ≥ 5. (D–F) RAW 264.7 cells were treated with MTA, with or without A2B receptor antagonist PSB 0788 for 12 h and expression of (D) Vegfa, (E) IL10 and (F) Timp1 were determined by RT-qPCR, error bars indicate mean ± SEM, n ≥ 5. (G–I) RAW 264.7 cells were treated with MTA, with or without STAT3 inhibitor Stattic for 12 h, and expression of (G) Arg1, (H) Vegfa, and (I) IL10 were determined by RT-qPCR, error bars indicate mean ± SEM, n ≥ 5. All statistical comparisons were done using ANOVA for group comparisons or unpaired student’s t-test for individual comparisons; *P < 0.05, **P < 0.005, ***P < 5 × 10–4, n.s. = not significant.
We next asked whether a sustained contribution of the A2B receptor signaling pathway is required for maintaining the alternative macrophage activation state. To address this question, we treated the cells with the A2B receptor antagonist at a delayed time point following treatment with MTA. We found that treatment of the already activated macrophages with the adenosine A2B receptor antagonist was still able to abolish Arg1 expression (Supplementary Fig. S5E), suggesting the effect of MTA can be pharmacologically reversed, an important point for determining any translational therapeutic potential.
To illuminate the downstream mediators of MTA-stimulated A2B receptor signaling, we tested the role of transcriptional regulators CREB and STAT3, as these are reported to function downstream of adenosine receptors and are known to regulate Arg1, Vegfa, and IL10, among other genes52,53,54,55,56. We found that the inhibition of STAT3 completely abrogated the MTA-induced expression of all three genes. In contrast, inhibition of CREB had only marginal or no effect on MTA-induced gene expression (Fig. 3G–I; Supplementary Fig. S8A–C). We then utilized a dominant-negative construct of STAT3 to genetically reduce STAT3 activity and again observed decreased response to MTA (Supplementary Fig. S8D)57. Interestingly, we were not able to reliably measure any change in STAT3 phosphorylation (p-Tyr705) levels in response to MTA at the time points measured (Supplementary Fig. S8E), suggesting such a phosphorylation change was not necessary for MTA-stimulated A2B receptor signaling in potentiating alternative macrophage polarization. This finding is consistent with known roles of STAT3 in regulating gene expression and chromatin organization, and as an important component of cancer and immune biology as previously demonstrated58,59,60,61.
MTA and adenosine activate distinct yet overlapping signaling pathways
As both adenosine and MTA signal through adenosine receptors, we sought to determine if there were any quantitative or qualitative differences in their effect on macrophage activation. We treated RAW 264.7 macrophages with equimolar concentrations of adenosine or MTA and tested the expression of macrophage activation markers in response to each metabolite. To our surprise, we found that MTA much more potently stimulated expression of Arg1, Vegfa, and Timp1 than did adenosine (Fig. 4A–C). In addition, MTA was unique in its ability to suppress Pparg expression (Fig. 4D). Furthermore, while MTA and adenosine similarly upregulated IL10 expression (Fig. 4E), the impact of adenosine on IL10 (and Timp1) was impervious to PSB0788, suggesting adenosine was acting through a different mechanism to induce expression of these genes.
Distinct MTA and adenosine signaling. (A–C) RAW 264.7 cells were treated with the MTA or adenosine with or without A2B receptor antagonist PSB 0788 for 12 h and (A) Arg1, (B) Vegfa, and (C) Timp1 expression were measured by RT-qPCR, error bars indicate mean ± SD, n ≥ 4. (D) RAW 264.7 cells were treated with MTA or Adenosine for 12 h and Pparg expression was measured by RT-qPCR, n ≥ 5. (E,F) Raw 264.7 cells were treated with MTA or adenosine for 12 h with or without (E) A2B receptor antagonist PSB 0788 or (F) C/EBP inhibitor betulinic acid (B. Acid) and IL10 expression was measured by RT-qPCR, n ≥ 5. (G) IL10 expression was measured by RT qPCR following adenosine treatment with or without STAT3 inhibition (Stattic), CREB inhibition (666-15), or A2AR antagonist Istradefylline, n ≥ 5. (H,I) RAW 264.7 cells were treated with MTA or adenosine for 12 h with or without the indicated inhibitors and IL6 expression was measured by RT-qPCR, n ≥ 5. (J) RAW 264.7 cells were treated with the indicated components, denoted by color codes, and cytokines in spent media were measured using Luminex technology after 2 (IL-6) or 3 (IL-10, VEGFA) days. The diagram shows the concentrations of each cytokine in the media. All statistical comparisons were performed using ANOVA for group comparisons or unpaired student’s t-test for individual comparisons; Error bars indicate mean ± SD; *P < 0.05, **P < 0.005, ***P < 5 × 10–4, n.s. = not significant.
We then tested the role of downstream mediators in producing the response to MTA and adenosine signaling. We found the effect of MTA and adenosine on IL10 expression were both effectively blocked by betulinic acid, an inhibitor of transcription factor C/EBP (CEBPA/CEBPB) (Fig. 4F), consistent with reports of C/EBP being a critical regulator of IL10 expression62 and indicating a convergence of the adenosine and MTA signaling pathways through this transcription factor. Furthermore, similar to what we had observed with MTA (Fig. 3G), induction of IL10 expression by adenosine was also blocked by stattic, indicating adenosine-stimulated IL10 expression requires STAT3 (Fig. 4G). However, whereas MTA-induced expression of IL10 was independent of CREB (Supplementary Fig. S8B), addition of CREB inhibitor 666-15 effectively blocked IL10 expression in response to adenosine (Fig. 4G), indicating that in the downstream signaling pathways there are distinct mechanisms responsible for MTA and adenosine-mediated responses in macrophages.
To further illuminate the distinct/overlapping roles of MTA and adenosine, we investigated the expression of IL6, another important regulator of macrophage activation63,64 that has been reported to be upregulated in response to adenosine signaling65,66,67,68,69. We found that adenosine much more potently upregulated IL6 than did MTA, and that while inhibition of STAT3 decreased IL6 expression in the context of MTA treatment, STAT3 inhibition had no significant effect on IL6 expression following adenosine treatment (Fig. 4H). Furthermore, adenosine-mediated upregulation of IL6 expression was further potentiated by A2A receptor antagonist Istradefylline, as well as by inhibition of CREB or C/EBP, while the effect of MTA was only potentiated by the C/EBP inhibition, not CREB or A2A receptor blockade, suggesting there are different negative feedback signaling pathways for MTA and adenosine (Fig. 4I). Thus, while MTA and adenosine both upregulate IL6 expression (adenosine to a greater extent), the dramatic differences in IL6 regulation following MTA or adenosine stimulation further demonstrate the distinctive receptors and signaling pathways activated by these two metabolites.
To confirm the cytokine production measured by gene expression analysis, we utilized multiplex cytokine quantification to measure IL-6, IL-10, and VEGFA protein levels in spent media from RAW 264.7 cells treated with MTA or adenosine. This analysis of cell protein production confirmed the gene expression findings detailed above, with increased VEGFA and IL-10 levels in response to MTA exposure and clear IL-6 and IL-10 production following adenosine treatment (Fig. 4J; Supplementary Fig. S9).
Collectively these results suggest that MTA and adenosine each engage a distinct balance of adenosine receptor signaling, resulting in divergent pathway activation, gene expression profiles, and cytokine production. The unique effect of MTA on macrophages, which is not shared by adenosine and requires the adenosine A2B receptor, STAT3, and C/EBP, supports an immunosuppressive, M2-polarizing effect of aberrant MTA accumulation on macrophages in MTAP-deficient GBM.
Discussion
Understanding and targeting the immuno-suppressive mechanisms of GBM is a critical step toward improving treatment for this lethal cancer type9. In this study, we provide multiple lines of evidence to link MTAP loss, one of the most common genetic/epigenetic events in GBM, to changes in the tumor immune microenvironment. We demonstrate using patient data that MTAP deleted GBMs contain an immunosuppressive gene expression signature, with increased numbers of M2 macrophages and decreased lymphocyte activation. Our findings resonate with a recent study showing that loss of the CDKN2A/MTAP locus confers an immunosuppressive microenvironment in human cancers70, raising the possibility that MTAP loss is an underlying factor. Utilizing in vitro macrophage models, we verified that the process of M2 polarization is influenced by MTA signaling through the adenosine A2B receptor, resulting in upregulation of M2 marker genes Arg1, IL10, and Vegfa. This represents a unique mechanism of alternative macrophage activation and a potentially significant contribution to the immunosuppressive tumor microenvironment in GBM, which is known for an abundance of immunosuppressive tumor-associated macrophages71. It has previously been demonstrated that MTA can directly suppress the proliferation and function of T lymphocytes26,72. Our findings complement these studies by showing that MTA can also influence innate immune cells.
The finding that the macrophage response to MTA is distinct from the response to adenosine suggests that MTAP-deficient GBMs with aberrant MTA accumulation likely have an impact on the immune environment in a manner that is qualitatively and/or quantitatively different from tumors which simply accumulate adenosine as a mechanism of immune evasion73,74. The role of adenosine receptor signaling in regulating inflammation, as well as its tumor-promoting functions, are beginning to be more widely recognized75. Adenosine receptors and the adenosine-generating enzymes, CD39 and CD73, are being investigated as therapeutic targets, either through stimulating or countering the adenosine receptor signaling pathways to control pathogenic inflammation or treat advanced cancers76,77,78. In GBM, adenosine (and upregulation of CD39 and CD73) has been shown to contribute to tumor-mediated immunosuppression79. Additionally, A2B receptor signaling has been shown to promote tumor metastasis and stimulate tumor angiogenesis80,81. Our finding that the effect of MTA on macrophage functionality is mediated by the adenosine A2B receptor and is amenable to pharmacological intervention adds a new line of evidence to support the rationale of targeting A2B receptor signaling in MTAP-deficient GBM. Development of MTAP-deficient, immune-competent GBM models will be necessary for further investigating the effectiveness of this strategy in vivo. We speculate that given the abundance of tumor-associated macrophages and the characteristic immunosuppressive microenvironment of GBM, altering MTA-mediated purinergic signaling has the potential to significantly impact tumor growth and augment immunotherapeutic interventions in MTAP-deleted GBM.
We note that experimental findings from the in vitro cell line models bear major limitations and may not fully explain the observed link between immunosuppression and MTAP status. In addition, those experimental results represent only a limited view of the complex time scale of the signaling mechanisms and responses involved. We suggest that further research utilizing human GBM specimens and in vivo, immunocompetent orthotopic GBM models will be necessary to assess the functional interplay between MTAP loss and the innate immune cells in GBM, and how they collectively influence the adaptive immune characteristics within the tumor.
Materials and methods
Cell lines and cell culture
RAW 264.7 and THP-1 macrophage cell lines were obtained from the Duke Cell culture Facility. BV-2 cells were a generous gift from Dr. Tso-Pang Yao. RAW 264.7 and BV-2 cells were maintained in DMEM with 4.5 g/L l-glucose (Sigma Cat #D6429) supplemented with 10% heat-inactivated FBS and anti-anti (antibiotic/antimycotic). THP-1 cells were cultured in RPMI 1640 with 10% heat-inactivated FBS and anti-anti. CT-2A cells were a generous donation from Dr. Darrell Bigner. The cells were maintained in DMEM/F12 (Gibco Cat #11330-032) supplemented with B-27 (Gibco Cat #17504-044), EGF (Stemcell), and FGF (Stemcell) and grown in suspension. Primary tissue cultures were derived with consent from patient tumor samples obtained by the Duke Brain Tumor Center. These patient-derived cultures were maintained in human neural stem cell (NSC) media (STEMCELL, cat# 05751), supplemented with EGF, FGF, and Heparin and plated onto laminin coated plates. All experiments were performed within the first 20 passages.
Plasmid construction and generation of derivative cell populations
The CRISPR system was used for knockout of Mtap in CT-2A. Two double-stranded oligonucleotides that encode sgRNA targeting exon 1 (CCTCGGGCTCCGCCTGCACGGCG) and exon 3 (CCATCCGATGCCTTAATTTTGGG) of Mtap were cloned into the px552-pEASY plasmid. CT-2A cells were transfected simultaneously with sgRNA plasmid and the px458 plasmid containing cas9 and GFP. For transient plasmid transfection, plasmids (2 plasmids at 1:1 ratio for achieving the desired gene deletion/mutations) and Transfex (ATCC, cat# ACS-4005) were mixed and used for cell transfection according to manufacturer’s instructions. Three to four days after the transfection, green fluorescent protein–positive (GFP+) cells were sorted via fluorescence-activated cell sorting (BD FACSVantage SE cell sorter, Duke Cancer Institute) to obtain the GFP+ population. Sorted cells were plated at single-cell densities and allowed to expand for 21 days, at which point DNA was prepped from each colony to screen for a deletion in Mtap (exon 1–exon 3) using PCR amplification across the deleted region. RAW264.7 cell lines with A2bR gene knockout (ko) was obtained by CRISPR using sgRNA#1 (GACGCAAGACGCGCTGTACG) or sgRNA#2 (GGCCACGTCTGCCGTCGCCA), delivered by lentivirus construct LentiVRISPR-E (Addgene #78852). Transduced cells were selected with puromycin and resistant populations were genotyped by Sanger sequencing. For generating RAW264.7- STAT3-DN derivative line, RAW264.7 cells were transduced with lentivirus (Addgene, plasmid # 24984, EF.STAT3DN.Ubc.GFP, expressing dominant negative STAT3 and GFP), and GFP + cells were sorted for further experiments. sgRNA and primer sequences are shown in Supplementary Table 1.
In vitro macrophage polarization
Raw 264.7 cells were plated in a 12 well dish and stimulated with methylthioadenosine (MTA) (Cayman Cat #15593) or adenosine (Sigma Cat #A9251) for 12 h, at which point they were collected for analysis. Cytokines IL4 (Peprotech Cat #200-04) and IL-13 (Peprotech Cat #200-13) were added within one hour of MTA/adenosine administration at a dose of 5 ng/mL. THP-1 stimulation with MTA was identical to RAW 264.7 cells except that the cells were first differentiated using PMA (Cayman Cat #10008014) for 24 h, then cultured in fresh media for 24 h prior to stimulation. BV2 cells were cultured in the same conditions as RAW 264.7 cells. Primary human monocytes were obtained frozen from STEMCELL Technologies (Cat # 70034). Cells were cultured in macrophage differentiation medium (STEMCELL Technologies, Cat # 10961) supplemented with M-CSF (STEMCELL, Cat # 78057.1) for 48 h, after which MTA was added and cells were harvested after 24 h. To test the impact of A2B and A2A receptors on MTA-mediated macrophage polarization, A2AR inhibitor Istradefylline (Selleckchem Cat #S2790) or A2BR inhibitor PSB 0788 (Tocris Cat #3199) were added simultaneously with MTA or adenosine, except in the indicated experiments where delayed administration of the A2BR inhibitor was tested. To further explore downstream signaling pathways STAT3 inhibitor Stattic (Tocris Cat # 2798), CREB inhibitor 666-15 (Tocris, Cat #5661) and C/EBP inhibitor betulinic acid (Tocris Cat #3906) were added at the time of MTA/adenosine administration. To test the effect of physiological accumulations of MTA from tumor cells, spent media from CT-2A parental or MTAP knockout cells was collected and added to the macrophage media in a 2:1 ratio and cells were plated for 12 h prior to measuring response.
Preparation of RNA and RT- qPCR
Total RNA was extracted using quick-RNA mini prep kit (Zymo Research, cat# 11-328) following the manufacturer’s protocols. Concentration of RNA was determined by Nanodrop Lite Spectrophotometer (Thermo Scientific). For gene expression analysis, reverse transcription was performed to convert total RNA into complementary DNA (cDNA) using the RNA to cDNA EcoDry Premix (Clontech, cat #639547). Subsequently, real-time qPCR was performed following the aforementioned qPCR procedure. Each reaction included a cDNA template equivalent of 10 ng of total RNA. The glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and beta-Actin genes were used as internal expression controls for RT-qPCR with reliable results. When beta Actin was used as the control amplicon the following program was followed: 95 °C, 3 min; 41 cycles of 95 °C 10 s and 68 °C 20 s, then a standard dissociation curve from 65 to 95 °C of 5 s/5° increment. When Gapdh was utilized as the internal control the following program was used: 95 °C, 3 min; 40 cycles of 95 °C 10 s, 60 °C 20 s, and 72 °C 1 s, then a standard dissociation curve from 65 to 95 °C of 5 s/5° increment. Relative quantification of expression was performed by comparing the Ct of target amplicons to the Ct of internal controls (GAPDH, beta-Actin) using the 2−ΔΔCt method.
Oligos and primers
All oligos and primers used for the study were synthesized by Eton Bio and are listed in Supplementary Table 1. Quantitative PCR was performed using KAPA SYBR Fast 2 × Universal master mix (KK4602) according to the manufacturer’s protocols on a BIO-RAD CFX96 Real-Time System.
Western Blot
Arginase1 antibody was obtained from Thermofisher (Cat # 711765). Cells were treated with MTA for 24 h and cells were lysed with RIPA buffer and ran on a Bio-Rad ready-made gel and transferred to nitrocellulose membrane. Total STAT3 and p-Tyr705 STAT3 were obtained from Cell Signaling Technologies (Cat # 9139, 9145).
Milliplex/luminex assays
To quantify cytokine levels in cell culture media, we utilized the multiplex cytokine quantification technology available from Milliplex. Custom kits were ordered that measure IL-6, IL-10, and VEGFA. Samples were processed according to manufacturer’s instructions and read on a Luminex machine in the Immunology Unit of the Duke Regional Biocontainment Laboratory (RBL).
Analysis of The Cancer Genome Atlas (TCGA) data
All TCGA data was downloaded from https://tcga-data.nci.nih.gov/docs/publications/tcga/ and through cbioportal.org82,83. The most recently published 2013 GBM data set was used for all analyses. For each analysis, the maximum number of complete cases available (confirmed IDH1/2 wildtype) were used unless otherwise stated, as IDH mutations are known to independently influence epigenetics and cellular differentiation. Analysis was carried out using Graphpad Prism, Microsoft excel, Genesis, and Statgraphics software. To perform this analysis, samples were ordered by MTAP expression level or equally divided into quartiles based on MTAP expression and statistical comparisons were made between the “low” and “high” expression groups to identify genes that may be impacted by MTAP status. Benjamini–Hochberg correction was performed to account for multiple testing error. We utilized MTAP expression levels to categorize patients rather than gene copy number because MTAP is known to be silenced epigenetically in a variety of cancer types84,85,86,87, and our analysis of DNA methylation and gene expression data in patients suggests it can also be epigenetically silenced in GBM21.
Statistical analysis
Statistical tests (student’s t test, ANOVA) were performed using Graphpad Prism. All experiments were repeated to ensure reproducibility of results. Unless otherwise indicated, pooled data from multiple experiments was used for each figure. A P value cutoff of 0.05 was used to determine significance in all cases except where corrections were applied for larger data sets (i.e., Benjamini-Hochberg, Bonferonni).
References
Beatty, G. L. & Gladney, W. L. Immune escape mechanisms as a guide for cancer immunotherapy. Clin. Cancer Res. 21, 687–692. https://doi.org/10.1158/1078-0432.CCR-14-1860 (2015).
Motz, G. T. & Coukos, G. Deciphering and reversing tumor immune suppression. Immunity 39, 61–73. https://doi.org/10.1016/j.immuni.2013.07.005 (2013).
Lin, Y. & Okada, H. Cellular immunotherapy for malignant gliomas. Expert Opin. Biol. Ther. 16, 1265–1275. https://doi.org/10.1080/14712598.2016.1214266 (2016).
Reardon, D. A. et al. An update on vaccine therapy and other immunotherapeutic approaches for glioblastoma. Expert Rev. Vaccines 12, 597–615. https://doi.org/10.1586/erv.13.41 (2013).
McGranahan, T., Li, G. & Nagpal, S. History and current state of immunotherapy in glioma and brain metastasis. Ther. Adv. Med. Oncol. 9, 347–368. https://doi.org/10.1177/1758834017693750 (2017).
Stupp, R. et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 10, 459–466. https://doi.org/10.1016/S1470-2045(09)70025-7 (2009).
Chongsathidkiet, P. et al. Sequestration of T cells in bone marrow in the setting of glioblastoma and other intracranial tumors. Nat. Med. 24, 1459–1468. https://doi.org/10.1038/s41591-018-0135-2 (2018).
Woroniecka, K. et al. T-cell exhaustion signatures vary with tumor type and are severe in glioblastoma. Clin. Cancer Res. 24, 4175–4186. https://doi.org/10.1158/1078-0432.CCR-17-1846 (2018).
Nduom, E. K., Weller, M. & Heimberger, A. B. Immunosuppressive mechanisms in glioblastoma. Neuro Oncol. 17(Suppl 7), vii9–vii14. https://doi.org/10.1093/neuonc/nov151 (2015).
Woroniecka, K. I., Rhodin, K. E., Chongsathidkiet, P., Keith, K. A. & Fecci, P. E. T-cell dysfunction in glioblastoma: Applying a new framework. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-18-0047 (2018).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477. https://doi.org/10.1016/j.cell.2013.09.034 (2013).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812. https://doi.org/10.1126/science.1164382 (2008).
Hodges, T. R., Ferguson, S. D. & Heimberger, A. B. Immunotherapy in glioblastoma: Emerging options in precision medicine. CNS Oncol. 5, 175–186. https://doi.org/10.2217/cns-2016-0009 (2016).
Chen, C. F. et al. ATR mutations promote the growth of melanoma tumors by modulating the immune microenvironment. Cell Rep. 18, 2331–2342. https://doi.org/10.1016/j.celrep.2017.02.040 (2017).
Coelho, M. A. et al. Oncogenic RAS signaling promotes tumor immunoresistance by stabilizing PD-L1 mRNA. Immunity 47, 1083-1099 e1086. https://doi.org/10.1016/j.immuni.2017.11.016 (2017).
Schumacher, T. et al. A vaccine targeting mutant IDH1 induces antitumour immunity. Nature 512, 324–327. https://doi.org/10.1038/nature13387 (2014).
Kohanbash, G. et al. Isocitrate dehydrogenase mutations suppress STAT1 and CD8+ T cell accumulation in gliomas. J. Clin. Investig. 127, 1425–1437. https://doi.org/10.1172/JCI90644 (2017).
Amankulor, N. M. et al. Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes Dev. 31, 774–786. https://doi.org/10.1101/gad.294991.116 (2017).
Zhang, X. et al. IDH mutant gliomas escape natural killer cell immune surveillance by downregulation of NKG2D ligand expression. Neuro Oncol. 18, 1402–1412. https://doi.org/10.1093/neuonc/now061 (2016).
Nobori, T. et al. Absence of methylthioadenosine phosphorylase in human gliomas. Can. Res. 51, 3193–3197 (1991).
Hansen, L. J. et al. MTAP loss promotes stemness in glioblastoma and confers unique susceptibility to purine starvation. Cancer Res. 79, 3383–3394. https://doi.org/10.1158/0008-5472.CAN-18-1010 (2019).
Kamatani, N. & Carson, D. A. Abnormal regulation of methylthioadenosine and polyamine metabolism in methylthioadenosine phosphorylase-deficient human leukemic cell lines. Cancer Res. 40, 4178–4182 (1980).
Kryukov, G. V. et al. MTAP deletion confers enhanced dependency on the PRMT5 arginine methyltransferase in cancer cells. Science 351, 1214–1218. https://doi.org/10.1126/science.aad5214 (2016).
Mavrakis, K. J. et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science 351, 1208–1213. https://doi.org/10.1126/science.aad5944 (2016).
Barekatain, Y. et al. Homozygous MTAP deletion in primary human glioblastoma is not associated with elevation of methylthioadenosine. Nat. Commun. 12, 4228. https://doi.org/10.1038/s41467-021-24240-3 (2021).
Henrich, F. C. et al. Suppressive effects of tumor cell-derived 5’-deoxy-5’-methylthioadenosine on human T cells. Oncoimmunology 5, e1184802. https://doi.org/10.1080/2162402X.2016.1184802 (2016).
Limm, K., Wallner, S., Milenkovic, V. M., Wetzel, C. H. & Bosserhoff, A. K. The metabolite 5’-methylthioadenosine signals through the adenosine receptor A2B in melanoma. Eur. J. Cancer 50, 2714–2724. https://doi.org/10.1016/j.ejca.2014.07.005 (2014).
Keyel, P. A. et al. Methylthioadenosine reprograms macrophage activation through adenosine receptor stimulation. PLoS One 9, e104210. https://doi.org/10.1371/journal.pone.0104210 (2014).
Wang, L. et al. Human genetic and metabolite variation reveals that methylthioadenosine is a prognostic biomarker and an inflammatory regulator in sepsis. Sci. Adv. 3, e1602096. https://doi.org/10.1126/sciadv.1602096 (2017).
Lathia, J. D., Mack, S. C., Mulkearns-Hubert, E. E., Valentim, C. L. & Rich, J. N. Cancer stem cells in glioblastoma. Genes Dev. 29, 1203–1217. https://doi.org/10.1101/gad.261982.115 (2015).
Benight, N. M., Stoll, B., Marini, J. C. & Burrin, D. G. Preventative oral methylthioadenosine is anti-inflammatory and reduces DSS-induced colitis in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 303, G71-82. https://doi.org/10.1152/ajpgi.00549.2011 (2012).
Moreno, B. et al. Methylthioadenosine reverses brain autoimmune disease. Ann. Neurol. 60, 323–334. https://doi.org/10.1002/ana.20895 (2006).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457. https://doi.org/10.1038/nmeth.3337 (2015).
Gentles, A. J. et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med. 21, 938–945. https://doi.org/10.1038/nm.3909 (2015).
Chen, B., Khodadoust, M. S., Liu, C. L., Newman, A. M. & Alizadeh, A. A. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol. Biol. 1711, 243–259. https://doi.org/10.1007/978-1-4939-7493-1_12 (2018).
Newman, A. M., Gentles, A. J., Liu, C. L., Diehn, M. & Alizadeh, A. A. Data normalization considerations for digital tumor dissection. Genome Biol. 18, 128. https://doi.org/10.1186/s13059-017-1257-4 (2017).
Charoentong, P. et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 18, 248–262. https://doi.org/10.1016/j.celrep.2016.12.019 (2017).
Lamb, L. S. Jr. Gammadelta T cells as immune effectors against high-grade gliomas. Immunol. Res. 45, 85–95. https://doi.org/10.1007/s12026-009-8114-9 (2009).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: From mechanisms to therapy. Immunity 41, 49–61. https://doi.org/10.1016/j.immuni.2014.06.010 (2014).
Charles, N. A., Holland, E. C., Gilbertson, R., Glass, R. & Kettenmann, H. The brain tumor microenvironment. Glia 60, 502–514 (2012).
Wu, S. Y. & Watabe, K. The roles of microglia/macrophages in tumor progression of brain cancer and metastatic disease. Front. Biosci. 22, 1805–1829 (2017).
Martinez, F. O. & Gordon, S. The M1 and M2 paradigm of macrophage activation: time for reassessment. F1000Prime Rep. 6, 13. https://doi.org/10.12703/P6-13 (2014).
Murray, P. J. & Wynn, T. A. Protective and pathogenic functions of macrophage subsets. Nat. Rev. Immunol. 11, 723–737. https://doi.org/10.1038/nri3073 (2011).
Sorensen, M. D., Dahlrot, R. H., Boldt, H. B., Hansen, S. & Kristensen, B. W. Tumour-associated microglia/macrophages predict poor prognosis in high-grade gliomas and correlate with an aggressive tumour subtype. Neuropathol. Appl. Neurobiol. 44, 185–206. https://doi.org/10.1111/nan.12428 (2018).
Hasko, G. & Pacher, P. Regulation of macrophage function by adenosine. Arterioscler. Thromb. Vasc. Biol. 32, 865–869. https://doi.org/10.1161/ATVBAHA.111.226852 (2012).
Csoka, B. et al. Adenosine promotes alternative macrophage activation via A2A and A2B receptors. FASEB J. 26, 376–386. https://doi.org/10.1096/fj.11-190934 (2012).
Ara, A. I., Xia, M., Ramani, K., Mato, J. M. & Lu, S. C. S-adenosylmethionine inhibits lipopolysaccharide-induced gene expression via modulation of histone methylation. Hepatology 47, 1655–1666. https://doi.org/10.1002/hep.22231 (2008).
Koscso, B. et al. Adenosine augments IL-10-induced STAT3 signaling in M2c macrophages. J. Leukoc. Biol. 94, 1309–1315. https://doi.org/10.1189/jlb.0113043 (2013).
Gordon, S. Alternative activation of macrophages. Nat. Rev. Immunol. 3, 23–35. https://doi.org/10.1038/nri978 (2003).
Chawla, A. Control of macrophage activation and function by PPARs. Circ. Res. 106, 1559–1569. https://doi.org/10.1161/CIRCRESAHA.110.216523 (2010).
Ferrante, C. J. et al. The adenosine-dependent angiogenic switch of macrophages to an M2-like phenotype is independent of interleukin-4 receptor alpha (IL-4Ralpha) signaling. Inflammation 36, 921–931. https://doi.org/10.1007/s10753-013-9621-3 (2013).
Rebe, C., Vegran, F., Berger, H. & Ghiringhelli, F. STAT3 activation: A key factor in tumor immunoescape. Jak-Stat 2, e23010. https://doi.org/10.4161/jkst.23010 (2013).
Ruffell, D. et al. A CREB-C/EBPbeta cascade induces M2 macrophage-specific gene expression and promotes muscle injury repair. Proc. Natl. Acad. Sci. U.S.A. 106, 17475–17480. https://doi.org/10.1073/pnas.0908641106 (2009).
Vasquez-Dunddel, D. et al. STAT3 regulates arginase-I in myeloid-derived suppressor cells from cancer patients. J. Clin. Investig. 123, 1580–1589. https://doi.org/10.1172/JCI60083 (2013).
Nakamura, R. et al. IL10-driven STAT3 signalling in senescent macrophages promotes pathological eye angiogenesis. Nat. Commun. 6, 7847. https://doi.org/10.1038/ncomms8847 (2015).
Hasita, H. et al. Significance of alternatively activated macrophages in patients with intrahepatic cholangiocarcinoma. Cancer Sci. 101, 1913–1919. https://doi.org/10.1111/j.1349-7006.2010.01614.x (2010).
Hillion, J. et al. The high-mobility group A1a/signal transducer and activator of transcription-3 axis: An achilles heel for hematopoietic malignancies?. Can. Res. 68, 10121–10127. https://doi.org/10.1158/0008-5472.CAN-08-2121 (2008).
Yang, J. et al. Unphosphorylated STAT3 accumulates in response to IL-6 and activates transcription by binding to NFkappaB. Genes Dev. 21, 1396–1408. https://doi.org/10.1101/gad.1553707 (2007).
Timofeeva, O. A. et al. Mechanisms of unphosphorylated STAT3 transcription factor binding to DNA. J. Biol. Chem. 287, 14192–14200. https://doi.org/10.1074/jbc.M111.323899 (2012).
Yang, J. et al. Novel roles of unphosphorylated STAT3 in oncogenesis and transcriptional regulation. Can. Res. 65, 939–947 (2005).
Yuan, J., Zhang, F. & Niu, R. Multiple regulation pathways and pivotal biological functions of STAT3 in cancer. Sci. Rep. 5, 17663. https://doi.org/10.1038/srep17663 (2015).
Brenner, S. et al. cAMP-induced Interleukin-10 promoter activation depends on CCAAT/enhancer-binding protein expression and monocytic differentiation. J. Biol. Chem. 278, 5597–5604. https://doi.org/10.1074/jbc.M207448200 (2003).
Wang, Q. et al. Vascular niche IL-6 induces alternative macrophage activation in glioblastoma through HIF-2alpha. Nat. Commun. 9, 559. https://doi.org/10.1038/s41467-018-03050-0 (2018).
Mauer, J. et al. Signaling by IL-6 promotes alternative activation of macrophages to limit endotoxemia and obesity-associated resistance to insulin. Nat. Immunol. 15, 423–430. https://doi.org/10.1038/ni.2865 (2014).
Zhang, L., Franchini, M., Wehrli Eser, M. & Dip, R. Enhanced IL-6 transcriptional response to adenosine receptor ligands in horses with lower airway inflammation. Equine Vet. J. 44, 81–87. https://doi.org/10.1111/j.2042-3306.2010.00350.x (2012).
Schwaninger, M., Neher, M., Viegas, E., Schneider, A. & Spranger, M. Stimulation of interleukin-6 secretion and gene transcription in primary astrocytes by adenosine. J. Neurochem. 69, 1145–1150 (1997).
Schwaninger, M. et al. Adenosine-induced expression of interleukin-6 in astrocytes through protein kinase A and NF-IL-6. Glia 31, 51–58 (2000).
Fiebich, B. L. et al. IL-6 expression induced by adenosine A2b receptor stimulation in U373 MG cells depends on p38 mitogen activated kinase and protein kinase C. Neurochem. Int. 46, 501–512. https://doi.org/10.1016/j.neuint.2004.11.009 (2005).
Rees, D. A. et al. Adenosine-induced IL-6 expression in pituitary folliculostellate cells is mediated via A2b adenosine receptors coupled to PKC and p38 MAPK. Br. J. Pharmacol. 140, 764–772. https://doi.org/10.1038/sj.bjp.0705488 (2003).
Han, G. et al. 9p21 loss confers a cold tumor immune microenvironment and primary resistance to immune checkpoint therapy. Nat. Commun. 12, 5606. https://doi.org/10.1038/s41467-021-25894-9 (2021).
Chen, Z. et al. Cellular and molecular identity of tumor-associated macrophages in glioblastoma. Can. Res. 77, 2266–2278. https://doi.org/10.1158/0008-5472.CAN-16-2310 (2017).
Vandenbark, A. A., Ferro, A. J. & Barney, C. L. Inhibition of lymphocyte transformation by a naturally occurring metabolite: 5’-methylthioadenosine. Cell. Immunol. 49, 26–33 (1980).
Morello, S., Pinto, A., Blandizzi, C. & Antonioli, L. Myeloid cells in the tumor microenvironment: Role of adenosine. Oncoimmunology 5, e1108515. https://doi.org/10.1080/2162402X.2015.1108515 (2016).
Ohta, A. A metabolic immune checkpoint: Adenosine in tumor microenvironment. Front. Immunol. 7, 109. https://doi.org/10.3389/fimmu.2016.00109 (2016).
Hasko, G. & Cronstein, B. Regulation of inflammation by adenosine. Front. Immunol. 4, 85. https://doi.org/10.3389/fimmu.2013.00085 (2013).
Hay, C. M. et al. Targeting CD73 in the tumor microenvironment with MEDI9447. Oncoimmunology 5, e1208875. https://doi.org/10.1080/2162402X.2016.1208875 (2016).
Silverman, M. H. et al. Clinical evidence for utilization of the A3 adenosine receptor as a target to treat rheumatoid arthritis: data from a phase II clinical trial. J. Rheumatol. 35, 41–48 (2008).
Chen, J. F., Eltzschig, H. K. & Fredholm, B. B. Adenosine receptors as drug targets—What are the challenges?. Nat. Rev. Drug Discovery 12, 265–286. https://doi.org/10.1038/nrd3955 (2013).
Xu, S. et al. Synergy between the ectoenzymes CD39 and CD73 contributes to adenosinergic immunosuppression in human malignant gliomas. Neuro Oncol. 15, 1160–1172. https://doi.org/10.1093/neuonc/not067 (2013).
Sorrentino, C., Miele, L., Porta, A., Pinto, A. & Morello, S. Myeloid-derived suppressor cells contribute to A2B adenosine receptor-induced VEGF production and angiogenesis in a mouse melanoma model. Oncotarget 6, 27478–27489. https://doi.org/10.18632/oncotarget.4393 (2015).
Desmet, C. J. et al. Identification of a pharmacologically tractable Fra-1/ADORA2B axis promoting breast cancer metastasis. Proc. Natl. Acad. Sci. U.S.A. 110, 5139–5144. https://doi.org/10.1073/pnas.1222085110 (2013).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404. https://doi.org/10.1158/2159-8290.CD-12-0095 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1. https://doi.org/10.1126/scisignal.2004088 (2013).
Behrmann, I. et al. Characterization of methylthioadenosin phosphorylase (MTAP) expression in malignant melanoma. Am. J. Pathol. 163, 683–690. https://doi.org/10.1016/S0002-9440(10)63695-4 (2003).
Hellerbrand, C. et al. Promoter-hypermethylation is causing functional relevant downregulation of methylthioadenosine phosphorylase (MTAP) expression in hepatocellular carcinoma. Carcinogenesis 27, 64–72. https://doi.org/10.1093/carcin/bgi201 (2006).
Ishii, M. et al. Methylthioadenosine phosphorylase gene is silenced by promoter hypermethylation in human lymphoma cell line DHL-9: Another mechanism of enzyme deficiency. Int. J. Oncol. 26, 985–991 (2005).
Leal, M. et al. Promoter hypermethylation of CDH1, FHIT, MTAP and PLAGL1 in gastric adenocarcinoma in individuals from Northern Brazil. World J. Gastroenterol. 13, 2568–2574 (2007).
Acknowledgements
We thank Ivan Spasojevic and Ping Fan of the Duke Cancer Institute’s Pharmacokinetic/Pharmacodynamic core laboratory for the LC-MS/MS metabolite analysis. Biomarker profiling was performed under the management of Dr. Andrew N. Macintyre and direction of Dr. Gregory D. Sempowski in the Immunology Unit of the Duke Regional Biocontainment Laboratory (RBL), which received partial support for construction from the National Institutes of Health, National Institute of Allergy and Infectious Diseases (UC6-AI058607). This work was supported by a National Cancer Institute National Research Service Award (F30CA206336; L.J.H), a National Comprehensive Cancer Network Young Investigator Award (Y.H.), and the National Institute of Neurological Disorders And Stroke of the National Institutes of Health, Award Number R01NS101074 (Y.H). This work was also supported by funding from the National Institutes of Health Duke SPORE in Brain Cancer (P50 CA190991), as well as the Circle of Service Foundation (Y.H) and the Southeastern Brain Tumor Foundation (Y.H).
Author information
Authors and Affiliations
Contributions
L.J.H. designed and performed the experiments, analyzed the data, and wrote the manuscript. R.Y., K.R., W.W. performed the experiments and analyzed the data. L.C. and Q.Y. performed statistical analyses of the data. C.J.P. and Y.H. designed and oversaw the implementation of the experiments and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hansen, L.J., Yang, R., Roso, K. et al. MTAP loss correlates with an immunosuppressive profile in GBM and its substrate MTA stimulates alternative macrophage polarization. Sci Rep 12, 4183 (2022). https://doi.org/10.1038/s41598-022-07697-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-07697-0
This article is cited by
-
MTAP loss: a possible therapeutic approach for glioblastoma
Journal of Translational Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.