Abstract
Marine sponges (phylum Porifera) are leading organisms for the discovery of bioactive compounds from nature. Their often rich and species-specific microbiota is hypothesised to be producing many of these compounds. Yet, environmental influences on the sponge-associated microbiota and bioactive compound production remain elusive. Here, we investigated the changes of microbiota and metabolomes in sponges along a depth range of 1232 m. Using 16S rRNA gene amplicon sequencing and untargeted metabolomics, we assessed prokaryotic and chemical diversities in three deep-sea sponge species: Geodia barretti, Stryphnus fortis, and Weberella bursa. Both prokaryotic communities and metabolome varied significantly with depth, which we hypothesized to be the effect of different water masses. Up to 35.5% of microbial ASVs (amplicon sequence variants) showed significant changes with depth while phylum-level composition of host microbiome remained unchanged. The metabolome varied with depth, with relative quantities of known bioactive compounds increasing or decreasing strongly. Other metabolites varying with depth were compatible solutes regulating osmolarity of the cells. Correlations between prokaryotic community and the bioactive compounds in G. barretti suggested members of Acidobacteria, Proteobacteria, Chloroflexi, or an unclassified prokaryote as potential producers.
Similar content being viewed by others
Introduction
Sponges (phylum Porifera) are sessile filter-feeding animals found worldwide in shallow and deep, marine and freshwater habitats. Sponges host an exceptionally rich and diverse associated microbiota, together referred to as a ‘holobionts’1, and are frequently grouped into high and low microbial abundance sponges (HMA and LMA sponges). Besides common marine prokaryotes, a considerable part of the prokaryotic communities within sponges is affiliated with so-called ‘sponge-enriched clusters’ indicating their adaptation to the sponge hosts2. These sponge-associated prokaryotic communities are acquired both through horizontal and vertical transmission of prokaryotic symbionts1,3,4 though recent work has challenged both the specificity of sponge-associated prokaryotes5 as well as the fidelity of their vertical inheritance6. Indeed, despite a high number of microbiome studies and their links to various temporal, spatial or environmental aspects, factors governing assembly and variation of the prokaryotic communities are still a matter of debate. As work on sponge prokaryotic community composition is usually based on shallow-water specimens7, this lack of understanding is particularly apparent in the deep-sea, where sampling is sparse8,9,10. In sponges from surface waters to 200 m depth, shifts in prokaryotic communities have been attributed to a combination of factors such as nutrient availability via light availability11,12,13. One study of a diverse set of deeper sponges (472–4160 m) found no evidence for the influence of depth on the sponge-associated microbiota9. Meanwhile, another study reported sponge-associated clusters of ASVs to vary in the proximity of a seamount (575 –2184 m)10. Depth is a proxy for correlated variables that may directly influence the sponge holobionts, that is, water mass with associated physiochemical and nutrient properties, hydrostatic pressure, light, and food supply.
The high diversity of sponge-associated prokaryotes, each producing its own specific set of compounds, has been used to partially explain the exceptional diversity of chemical compounds in sponges14,15 and positions them as leading sources for novel marine bioactive compounds16,17. However, the extent of the contribution of microbes to the holobiont chemodiversity is still poorly understood and a topic of high interest. To address this connection, studies have targeted specific bacteria or bacterial groups as producers of specific compounds18,19,20,21. Profiling of (mostly unknown) specialized metabolites with untargeted metabolomics is a less common approach in sponges and so far, it has essentially been used in demosponge chemotaxonomy22,23,24,25,26,27. Only two works connected the microbial community and the metabolome28,29, neither of which investigated a deep-sea gradient.
In this study, we addressed (a) whether shifts in prokaryotic community composition extend beyond the euphotic zone, (b) whether changes in depth and/or prokaryotic community composition are reflected in the metabolome and (c) whether we can use these potential variations to link microbes and metabolome and suggest putative producers of bioactive compounds. We address these questions through 16S rRNA gene sequencing and ultra-high performance liquid chromatography high-resolution mass spectrometry (UPLC-HRMS) in three North Atlantic deep-sea demosponge species, Geodia barretti Bowerbank, 185830, Stryphnus fortis (Vosmaer, 1885)31 and Weberella bursa Vosmaer, 188531. G. barretti is an HMA sponge32 and S. fortis was hypothesised to be an HMA sponge, too, whereas W. bursa was hypothesised to be an LMA sponge (N. Boury-Esnault pers. comm.). We collected 20, 15 and 17 specimens respectively, along a depth gradient ranging from 244 to 1476 m.
Methods
Sampling
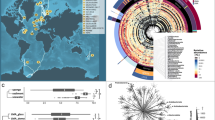
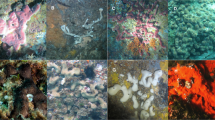
Sponge specimens were sampled by the crews of the R/V Pâmiut of the Greenland Institute of Natural Resources during cruises conducted by Fisheries and Oceans Canada in the Davis Strait using Alfredo/Cosmos benthic trawls taken between a depth of 244 m and 1476 m (Fig. 1). All samples were taken during the same season (September–October, Autumn) from 2011 to 2015. Samples were frozen on board (− 20 °C) and shipped to the Bedford Institute of Oceanography (BIO, Dartmouth, Nova Scotia, Canada) for identification. Freeze-dried subsamples from the choanosome (i.e. inside of the sponge, to reduce the risks of surface contamination by epibionts) were sent to Uppsala University and stored at 8 °C until processing for global metabolite profiling. Sample identification was reassessed and all specimens of S. fortis were checked for the presence of Hexadella dedritifera33 overgrowth. Frozen subsamples were sent to Wageningen University & Research and stored at − 20 °C until processing for prokaryotic community analyses.
Map of the slope in the Davis Strait in the North Atlantic, where the samples were collected. The insert shows the geographic location of the sampling area (black box) between Canada and Greenland. Samples were collected at depths ranging from 244 to 1476 m. Figure made with R v3.5.1 (https://www.r-project.org) with packages marmap and ggplot2.
Detailed sample information can be found in Table S1 (deposited at PANGAEA https://doi.pangaea.de/10.1594/PANGAEA.909246).
Study site, geography and oceanography
The maximal geographic distance across the sampling site was 786 km, covering a depth range of 1232 m. To identify the potential effect of oceanographic conditions (e.g. seawater properties, currents and vertical velocities, water mass origin, age and mixing history) on sponge prokaryotic community compositions and metabolomes, all oceanographic data available for the study area were assembled. These oceanographic observations came from the World Ocean Database (https://www.nodc.noaa.gov/OC5/WOD/pr_wod.html) and Bedford Institute of Oceanography archives, and from Argo float profiles34. All vertical profiles of temperature and salinity obtained from discrete (e.g., water samples and reversing thermometers) and initially continuous and later subsampled measurements were checked for errors and vertically interpolated for every 5 m.
16S rRNA gene amplicon sequencing and analyses
For each sponge individual a match head of tissue was cut out of the frozen tissue samples with a sterile scalpel and washed three times in artificial seawater to remove any attached debris35. DNA extraction and two-step PCR reaction (16S rRNA gene: 515F 5′- GTG YCA GCM GCC GCG GTA A-3′, 806R 5′- GGA CTA CNV GGG TWT CTA AT-3′) were performed following procedures previously described36. The barcoded 16S rRNA gene amplicons were sequenced at GATC Biotech AG (Constance, Germany; now part of Eurofins Genomics Germany GmbH) by Illumina Miseq sequencing. Raw data was analysed using NG-Tax (Galaxy version 1.0), for detail see Supplementary methods37. Forward and reverse paired-end reads were trimmed to 70 nucleotides and concatenated, resulting in sequences of 140 bp used for subsequent sequence data processing. Taxonomy was assigned with a customized version of the SILVA 128 SSU database38, and ASVs (amplicon sequence variants) classified as chloroplasts were removed. Sequencing data was deposited at the NCBI Sequence Read Archive (SRA) with accession number SRP142603 under file numbers SRX3993926–SRX3993970.
Metabolite extraction, UPLC-HRMS acquisition and analyses
Freeze-dried sponge tissue was ground and 50 mg of the powder was extracted with 4.5 mL 70% methanol/Milli-Q (MQ) water for one hour at room temperature on a shaker at low speed. The vials were centrifuged at 2000 × g for 10 min at 4 °C and 2 mL of the supernatant was dispensed into two glass vials and dried on a heating block (43 °C) under a stream of nitrogen. Dry extracts were kept at − 80 °C until further analyses.
All samples were analysed with UPLC-HRMS, in hydrophilic interaction liquid chromatography (HILIC) and reversed-phase liquid chromatography (RP), in positive and negative ionization mode39.
Dried extracts were dissolved in 50 µL MQ water and 175 µL acetonitrile (MeCN) for HILIC, or 140 µL MQ water and 10 µL MeCN for RP analyses. Upon addition of the organic solvent for HILIC samples they separated into two immiscible layers. The vials were centrifuged for 3 min at 2000 × g to achieve complete separation and only the top layer (~ 150 µL) was transferred to an MS-vial for analysis. A 5 µL aliquot from each individual MS-vial for HILIC and RP chromatography, respectively, was combined to produce a quality control (QC) sample. The extracts were analysed on an Acquity I-Class UPLC coupled to a G2S Synapt Q-TOF with an electrospray ionization (ESI) ion source (all Waters Corp., Milford, Massachusetts, USA). Chromatographic separation in HILIC mode was performed on an Acquity UPLC BEH Amide column (1.7 μm, 2.1 mm i.d. × 50 mm, Waters Corp.) and in RP mode on an Acquity UPLC BEH C18 column (1.7 μm, 2.1 mm i.d. × 50 mm, Waters). The flow rate was 0.4 mL/min and the injection volume was 5 µL in all experiments. Data were acquired in MSE mode, and lock mass correction was applied in both positive and negative mode. Prior to each analysis, the instrument was calibrated in the m/z range 50 to 1500 and QC injections were made to condition the column and to ensure stable retention times and signal intensities. The study samples were analysed in randomized order with QC injections interspaced every sixth injection.
Raw data from the four UPLC-HRMS experiments were processed separately. Files were converted to CDF files with Databridge/MassLynx (Waters Corp., Milford, Massachusetts, USA), sorted into folders by species and processed with the R packages XCMS40 and CAMERA41. The output is a table with signal intensity per sample and ‘feature’. Each feature can be annotated as metabolite, adduct, isotope, or lack an annotation. The four resulting feature tables were filtered in R in three different ways (“cleaned”, “pc_group”, “ion”), as follows. Features eluting in the void (< 40 s) and features with a coefficient of variation > 30% in the QC samples were removed (data sets called “cleaned”). Based on these “cleaned” feature tables, adducts and overrepresented features were removed by retaining only the feature with the greatest cumulative signal per CAMERA pc_group (“pc_group”). In addition, based on the “cleaned” feature tables, only features explicitly annotated as [M + H]+ in positive ESI mode and [M-H]− in negative ESI mode were retained (“ion”). Metabolomic data is deposited at MetaboLights (www.ebi.ac.uk/metabolights/MTBLS1388) and extended methods are found in the supplementary information.
Computational and statistical analyses
Data sets (ASV table, features tables, meta data) produced as described above were analysed in R v3.5.1. For the prokaryotes, the effect of different data transformations was evaluated by principal component analysis (PCA) and non-metric multidimensional scaling (NMDS) using Bray–Curtis distance (Fig. S1). PERMANOVA, function ‘adonis’ in R package vegan42, was used to test for significant differences in prokaryotic community composition between sponge species. Alpha diversity of the samples was calculated with the R package phyloseq. For each sponge species, ASVs were classified as common if their average relative abundance was > 0.25%. ASVs classified as common were cross-checked for being sponge enriched following a procedure described by Dat et al.43. For subsequent ecological analyses, we assessed the rank correlation between dissimilarity indices and gradient separation based on different data transformations and applied the method yielding highest similarity with the Bray–Curtis index. To investigate the effect of depth on the sponge-associated prokaryotes, we fitted all available environmental variables (latitude, longitude, depth, salinity, temperature, sampling year) onto unconstrained ordinations (NMDS with Bray–Curtis distance) of the three species’ prokaryotic communities. In addition, we assessed multicollinearity among the environmental variables and then included only those environmental variables with a variance inflation factor < 10 to build a full constrained model as part for an automated stepwise model building approach evaluated with an ANOVA. The relative abundances of ASVs were correlated (Pearson) to sample depth to infer whether ASVs were ‘increasing’ or ‘decreasing’ with increasing sample depth (p ≤ 0.05), and noted as ‘increasing trend’ or ‘decreasing trend’ with increasing sample depth for non-significant correlations (p > 0.05). Pairwise sequence similarity was calculated with seqinr and a t-test was performed comparing the relative abundance above and below 1000 m for each ASV. The resulting p-values were corrected for multiple testing (FDR). The phylogeny of the ASVs was inferred with RAxML analyses implemented in CIPRES Science Gateway V. 3.344 and all annotations were mapped on the phylogram in iToL45.
To investigate the effect of depth on the holobiont metabolomes, we used orthogonal projection of latent structures (OPLS) models, a supervised ordination method separating predicted variation (depth) from all orthogonal variation (R package ROPLS46). OPLS models predicting variation with depth (n = 36) were built for all combinations of experimental setups (n = 4), species (n = 3), and feature filtering (n = 3). Seven-fold cross-validation is performed by the function resulting in a pQ2 value. Models were rejected due to overfitting if they yielded a pQ2 > 0.05. Model fit was evaluated to assess whether depth was affecting the variation of the metabolomes’ features. In an OPLS, all variables (metabolome features) are assigned a “variable influence on projection” (VIP) score: a variable with a VIP score > 1 is considered important to the model. The VIP score is not a statistic test and needs to be supported by such for validity. Known metabolite features from G. barretti and S. fortis, and the features with the highest VIP scores in the OPLS model of the HILIC positive “cleaned” data set were manually annotated in the metabolome and their signal intensities across the samples were extracted. As the patterns and models based on unidentified metabolomic features underlying the OPLS model should be interpreted with caution, we applied a dual feature identification approach: (1) manually extracting signal intensities of known bioactive compounds and (2) de novo identification of features designated as VIPs by a general OPLS model including all sponge samples combined. Standards were ordered to confirm the identity of VIPs in a repeated UPLC-HRMS/MS experiment, and annotations were confirmed/rejected based on retention time and matching MS/MS spectra. The correlation of the signal intensity and sample depth was tested.
To test overall similarity between the prokaryote and metabolome data sets, we used a Mantel test on the distance matrices (Bray–Curtis for the prokaryotes and Euclidean dissimilarity for the metabolome) and Procrustes rotations.
The microbial interaction network for the G. barretti-associated prokaryotes was generated following suggestions by Weiss et al.47. ASVs present in less than three samples were removed and the sparsity and Neff of the remaining ASV table was calculated. The ASV table containing 289 remaining ASVs was formatted according to the requirements of each network inference tool. Prokaryotic interaction networks were generated using MENA (using Pearson and Spearman correlation coefficients)48 (http://ieg4.rccc.ou.edu/mena), SparCC49, fastLSA50 and MIC (R package minerva)51. The resulting networks were filtered to only retain positive interactions (edges). The consensus network contained edges with p ≤ 0.05 in four or five of the algorithms.
All ASVs in G. barretti were correlated (Spearman) with the signal of the diketopiperazines and tryptophan derivatives. In these two compound groups, the ASVs producing the ten strongest positive correlations were compared.
The online supplement including analysis code can be found at https://ksteffen.github.io/Paamiut/.
Results
The oceanographic setting revealed five water masses in the Davis Strait
The Davis Strait where the samples were collected was found to be characterized by five water masses from shallow to deep: Shelf Water (ShW), Slope Water (SW), the Irminger Current (IC), Labrador Sea Water (LSW), and Icelandic Slope Water (ISW) (Fig. 2). The water masses can be detected by bends in the lines representing the continuous measurements of salinity and temperature with depth. Based on the sample depth, the samples in this study originated mainly from two water masses: those shallower than 1000 m were attributed to the IC with its core at about 600 m, a salinity of 34.93 psu, a temperature of 4.5 °C, and those from localities deeper than 1000 m were attributed to the LSW with its core at 1200 m, a salinity of 34.89 psu, a temperature of 3.7 °C. The limit of 1000 m depth was chosen to approximate the midpoint in depth between the two water masses. The LSW was more recently ventilated through winter convection in the Labrador Sea and therefore was richer in oxygen than the IC above it, which in turn was mainly advected from the North Atlantic outside the Labrador Sea. The strong currents intensify mixing between these two water masses and thus horizontal and vertical transfer of matter52. Even though the distinction between the water masses persists, this exchange may help transfer critical substances and enrich the water mass interfaces. The two shallowest specimens of Weberella bursa (224 and 331 m respectively) are influenced by the SW water mass in the Hudson Strait.
Temperature-salinity diagram showing the superimposed water masses at different depths in the Davis Strait. Shelf Water (ShW) and Slope Water (SW) are shallower than the samples in this study and display a range of salinities. We detect the core of the Irminger current (IC) at approx. 600 m depth, the core of the Labrador Sea Water (LSW) at approx. 1200 m depth, and the Icelandic Slope Water at approx. 1600 m and below. The abbreviations for the water masses were placed to approximate the cores of the water masses. Samples were attributed to the water masses based on their sample depth (see Table S1).
Prokaryotic communities changed with depth
We recovered a total of 420 amplicon sequence variants (ASVs) belonging to 17 prokaryotic phyla associated to the 14 G. barretti specimens, 461 ASVs belonging to 20 prokaryotic phyla associated to the 15 S. fortis specimens, and 135 ASVs belonging to 13 prokaryotic phyla associated to 16 W. bursa specimens (Fig. 3). For individual specimens, the richness ranged from 148 to 229 (mean 192) ASVs for G. barretti, from 129 to 204 (mean 176) ASVs for S. fortis, and 16–58 (mean 29) ASVs for W. bursa. Although S. fortis is frequently overgrown with the sponge Hexadella dedritifera, we detected no signs of a contamination of the S. fortis microbiota as its composition is very different from that of H. dedritifera microbiota (Fig. S2 in53). The prokaryotic alpha diversity in G. barretti and S. fortis was higher than in W. bursa for all metrics assessed (Fig. S2). This is evidence of S. fortis being an HMA sponge and W. bursa being an LMA sponge54. There were 98 (78.6% of the reads from G. barretti), 89 (77.6% of the reads from S. fortis), and 20 (94.8% of the reads from W. bursa) ASVs above the threshold of 0.25% average relative abundance. Among these abundant ASVs, 72 (73.5%), 64 (71.9%) and 11 (55%) were from sponge-enriched clusters (in G. barretti, S. fortis, and W. bursa respectively). As the prokaryotic communities were different for each sponge species (PERMANOVA of all 45 samples and all 691 ASVs by sponge species, p = 0.001), we performed further analyses per sponge species hereafter. The results of constrained and unconstrained analyses of environmental parameters and the prokaryotic communities, identified depth as significant factor for the microbiota composition, sometimes in combination with other environmental parameters (Table 1).
Prokaryotic community composition based on relative abundance of 16S rRNA gene sequences, aggregated at the phylum level. Specimens were grouped by sponge species and ordered by increasing depth from left to right, Geodia barretti (Gb, 407–1462 m), Stryphnus fortis (Sf, 483–1476 m) and Weberella bursa (Wb, 244–1271 m).
In G. barretti, 149 ASVs (35.5% of the ASVs representing an average relative abundance per sample of 45%) correlated with depth (86 increased, 63 decreased in relative abundance with increasing depth). In S. fortis, 99 ASVs (21.5% of the ASVs, average relative abundance per sample 21.2%) correlated with depth (62 increased, 37 decreased) and in W. bursa, 23 ASVs (17.0% of the ASVs, average relative abundance per sample 13.5%) correlated with depth (12 increased, 11 decreased) (Fig. 4). There was no apparent taxonomic trend or clustering among ASVs changing with depth. Hierarchical clustering of the prokaryotes revealed clusters of samples below 1000 m in G. barretti and S. fortis but not in W. bursa (Fig. S3). The sponge microbiota in the different water masses were significantly different in G. barretti and S. fortis, but not in W. bursa (PERMANOVA p = 0.004, p = 0.001, and p = 0.135 respectively, samples grouped above/below 1000 m, approximating the midpoint between the IC and LSW).
ASVs in G. barretti, S. fortis and W. bursa for which average relative abundance correlated with depth. The left panel shows ASVs increasing with depth, the right panel shows ASVs decreasing with depth in the three sponge respectively. ASV relative abundance is scaled from 0 to 1 for visualisation purposes. ASV labels coloured in red are sponge-enriched ASVs.
Sister group ASVs were defined as pairs of ASVs with sequence similarity ≥ 97% and opposing depth response. We found 19 pairs of sister group ASVs in G. barretti, two pairs in S. fortis and no instance of sister ASVs in W. bursa. These sister group ASVs included members of Acidobacteria (Subgroups 6, 9, 15, and 16), Proteobacteria (JTB23), Chloroflexi (SAR202 clade, TK10, and Anaerolineae) and Gemmatimonadetes (BD2-11 terrestrial group) (Figs. S4, S5, Table S2).
Metabolomes changed with depth
Metabolomes contained 3507, 2808, 4673, and 3166 features for HILIC positive, HILIC negative, RP positive and RP negative, respectively (“cleaned”). Further filtering to retain only entries identified as ions recovered 2212, 1351, 2736 and 1678 features respectively. Filtering to retain only the feature with the strongest signal per ‘pc_group’ (i.e., computationally approximating individual metabolites) recovered 105, 123, 171, and 105 features respectively. Overall, the metabolomes of the three holobionts were significantly different from each other (PERMANOVA, p = 0.001 in the HILIC positive “cleaned” data set) (Fig. S6). Therefore, we treated the metabolomes for the three species separately hereafter. Two major peaks containing barettin and 8,9-dihydrobarettin clearly decreased in deep specimens of G. barretti (Fig. 5). In contrast, the profiles for S. fortis and W. bursa did not show obvious variations in major peaks.
Overlain are the chromatograms acquired on a HILIC column (positive ESI) of (a) G. barretti, (b) S. fortis, and (c) W. bursa. Black lines indicate chromatograms from shallow specimens (< 1000 m), red lines indicate chromatograms from deep specimens (> 1000 m). For G. barretti, the green area highlights the peak of barettin (E- and Z-form), the blue area highlights the peak of 8,9-dihydrobarettin.
To evaluate whether depth affects the metabolomes as a whole, OPLS depth model fit (Table S3) was evaluated. Of the 36 depth models generated, 27 performed well (R2X > 0.9 and Q2 > 0.5), 11 of G. barretti, 9 of S. fortis, and 7 of W. bursa. Despite the many variables, cross validation indicated only six cases of overfitting (pQ2 > 0.05). The high number of good models is interpreted as support for the hypothesis that metabolome changed with depth.
Since the HILIC positive data set contained the most steadily acquired data (Fig. S7), we extracted compound signals from it when possible. We identified VIPs (features with importance to the depth model) (Fig. 6), and confirmed their identity with reference compounds for arsenobetaine, carnitine, phosphocholine, acetylcholine, choline sulphate, creatine. For VIPs uranidine and 2-methylbutyroylcarnitin no reference compounds were available. Significant variation with depth was annotated in the corresponding plot, for the complete results see (Table S4). Acetylcholine is the only VIP not correlated with depth in any of the three species.
In G. barretti, we identified 12 previously reported bioactive compounds diketopiperazines Trp-Arg (barettin55, 8,9-dihydrobarettin56, 8,9-dihydro-8-hydroxybarrettin57, geobarrettin A and B58), diketopiperazine Pro-Arg57, bromotryptophane derivatives (6-bromo-8-hydroxyconicamin25, 6-bromoconicamin25, geobarrettin C58, L-6-bromohypaphorine58), and the peptides barrettides A and B59. We could not find bromobenzisoxalone barettin60 (Fig. 7), possibly due to its very low concentration in the animal61 or production under specific circumstances only. In S. fortis, ianthelline and stryphnusin, two previously reported compounds were annotated (Fig. 7). Significant variation with depth was stated in the corresponding plot, for the complete results see (Table S4). There are no previously reported compounds in W. bursa.
Signal intensities of previously known bioactive compounds plotted against sample depth. Correlation tests and t-tests were performed to assess all compounds variation with depth. Significant results (pFDR < 0.05) were annotated in the plot, correlation (ρ and pFDR) on top, and t-test (pFDR) below. Compounds associated with G. barretti: top panel (1–6) are diketopiperazines, middle panel (7–10) are indole derivatives, bottom left are peptides. Compounds associated with S. fortis: bottom right panel (11, 12).
Inter-omics analyses and identification of microbiota-metabolite links
Directed analysis of the G. barretti ASVs and specialised metabolites hypothesised to be of prokaryotic origin (diketopiperazines, tryptophan derivatives), revealed that four bacteria yielded strong positive correlations with three of the six diketopiperazines: ASVs 144, 264, 284, all members of Acidobacteria and ASV 171, a member of Actinobacteria. Six prokaryotes yielded strong positive correlations with three of the four tryptophan derivatives: ASV 144, 283, 284 all members of Acidobacteria, 320 a member of Proteobacteria, 350 a member of Chloroflexi, and 307, an unclassified prokaryote. ASVs 144, 171, 283 and 320 were categorised as “common ASVs” and all ASVs with strong positive correlations are found both in G. barretti and S. fortis.
To visualise patterns among the G. barretti ASVs, we constructed a consensus network of positive microbial interactions, containing 117 nodes (ASVs) and 143 edges (Fig. S8A). ASVs mentioned previously as correlating with specialised metabolites were highlighted with bold read labels. ASVs in the network correlating with depth were mainly found in distinct clusters (Fig. S8B). Results for testing overall congruency or correlation between microbiota and metabolome matrices were inconclusive and can be found in the supplement (Tables S5, S6).
Discussion
Sponge-associated prokaryotic communities are influenced by the water mass
Sponge-associated prokaryotic communities varied across meso- and bathypelagic depths (Fig. 4), a pattern which was masked when ASVs were aggregated at phylum level (Fig. 3). This finding contradicted the generally perceived stability of the sponge microbiome62,63,64,65,66 which changes mainly under lethal stress or disease67. The extent of ASV change was more pronounced in HMA than LMA sponges, but we were able to detect it in both types. For G. barretti, we also found a significant effect of longitude and salinity on the associated prokaryotic community (Table 1), but the variance inflation factors indicate that caution must be taken in interpreting these environmental variables due to their collinearity with depth (Table S7). For the S. fortis prokaryotic community composition, longitude may play an additional role (if only in the form of distance-decay). For W. bursa, the impact of environmental variables on prokaryotic community composition were inconclusive because they varied with the method used.
We conceptualized depth as a proxy for other environmental (biotic/abiotic) factors, as depth per se is unlikely to affect the sponge or its prokaryotic community. Oceanic depth correlates with light availability, temperature, salinity, and hydrostatic pressure, yet none of these parameters are likely to affect the samples in this study. Light intensity was ruled out as influencing factor in this study as even the shallowest sponge samples originated from localities beyond the euphotic zone and cyanobacteria were mostly absent68. Further, neither temperature62,69 nor salinity64 on their own necessarily affect sponge prokaryotic communities in shallow water except under fatal temperature stress conditions70. Hydrostatic pressure affects enzyme activity and thereby metabolic rates in fish in abyssal and hadal depths71. While these depths are beyond our sample range, the effect of hydrostatic pressure on the holobionts in this study could not be ruled out. Two previous studies investigating the overall effect of depth found little to no evidence for an effect on sponge prokaryotic communities9,65. This was likely due to the sampling within the same water masses and low number of samples in general.
Additional conceivable parameters for explaining the shifts in prokaryotic communities would be nutrients, minerals, dissolved oxygen and carbon dioxide, but the number of in situ measurements for these were insufficient to support quantitative conclusions. However, many of these parameters are dependent on the oceanographic setting, i.e. water masses. In context of geographical position and depth, temperature and salinity are known to sufficiently resolve the water mass structure, ventilation history and mixing conditions34,72,73. In particular, it has been demonstrated that the LSW formed by winter convection in the Labrador Sea is much better ventilated (higher dissolved oxygen and carbon dioxide) and is more recent in formation (younger in age) than the Atlantic Water that is advected to the region with the IC. We thus attributed the change of prokaryotic communities to variations in the vertical water mass structure and characteristics as detected by our oceanographic analyses. Since the dynamics and oceanographic structure of the water column changed in transition from the sea interior onto the shelf, water depth could be used as a variable effectively representing the cross-slope changes in seawater and ecosystem characteristics at the bottom level and throughout the water column.
In the HMA sponges G. barretti and S. fortis, where the effect of depth on prokaryotic community composition was most pronounced, we found samples below 1000 m forming distinct clusters (Fig. S3). This grouping was supported by the approximate position of the oceanic front between the two water masses: the IC and the LSW. This indicated the shallower sponge specimens (< 1000 m depth) were mainly influenced by the warmer and saltier IC (core at approx. 500 m depth). Meanwhile the deeper specimens (> 1000 m depth) were mainly influenced by the colder and fresher LSW (core at approx. 1200 m depth).
This grouping was further supported by the observation that differences in prokaryotic communities in deep and shallow sponges were sometimes caused by substitutions of taxonomically close ASVs (“sister group ASVs”) potentially representing microbial ecotypes of two different water masses. We reported such a pattern mainly among the G. barretti prokaryotes. However, in terms of the number of such ecotype pairs recovered, we found their detection to be strongly influenced by the stringency of the significance threshold (p < 0.05 versus pFDR < 0.05) and detection strategy (correlation test or t-test for depth response) highlighting the difficulty of assessing the phenomenon. Generally, less stringent thresholds revealed more pairs of sister group ASVs and additional phyla (Table S3, Figs. S4, S5).
Previous findings support the hypothesis of the effect of water masses on the microbiota, as water masses are known to structure microbial distributions in the sea74,75. They act as dispersal barrier for microorganisms in the deep sea, and each water mass harbours specific prokaryotic communities with different ecotypes76,77,78,79. Even communities of unicellular eukaryotes80,81, fish or benthic invertebrates seem structured by water masses82,83,84,85,86. Recent work linking clusters of sponge microbiota to a seamount could likewise be interpreted as sign of the effect of water masses and currents around the structure10.
In addition to this general effect of water masses on marine life, we acknowledge that population structure in the sponge hosts themselves could be a factor contributing to the shifts observed in the prokaryotic community composition of the three sponges. It is conceivable that water masses stratify dispersal of sponge larvae thereby leading to isolated populations86,87. However, as a part of the sponge prokaryotic community would still be derived from the different water masses, sponge-host population structure is not mutually exclusive with the hypothesis proposed herein.
Taken together, the present study has shown for the first time that water masses have a major impact in the structuring of sponge-associated prokaryotic communities, and thereby could explain the prokaryotic community variation observed in the three deep-sea sponge species.
Given the resilience of the prokaryotic taxa associated with sponges, even upon disturbance63,64, the sponge microbiome appears set once fully acquired. It is further recognized that the microbiota composition is dictated mainly by the sponge host identity2,9,12.
This poses the question as to how and when the adult sponge microbiota is established. Despite ample evidence for vertical inheritance of microbes1,3,4, its consistency in terms of which taxa are passed on has been challenged recently6 giving more weight to environmental acquisition. The pattern described herein, i.e. i) the compositional stability at phylum level despite ii) variations of underlying ASVs could be interpreted as evidence for a selective process for the required taxa, where representation of the taxa at ASV strain level depends on the availability in the environment. The presence of corresponding ASV ecotypes in samples from different water masses suggests a functional and taxonomic redundancy or complementarity88,89. Therefore, we hypothesize these taxa to participate in equivalent functions in the holobiont.
The chemical profiles of sponge holobionts vary with depth
Thus far, factors affecting specialized metabolite production in sponges have often been inconclusive or contradictory26,27,90,91, e.g. regarding seasonality28,92 or reproduction26,27,93. In the present study, evidence for variation of the chemical profiles with depth could be found at all levels of the analyses, with the most pronounced effects in the HMA sponges. The multivariate analyses attributed a proportion of the variation of all three holobiont metabolomes to depth and these models in turn aided in the identification of specific features varying with depth. Signals of previously known metabolites (Fig. 7) as well as additionally reported compounds (Fig. 6) extracted from the chemical profiles further confirmed variations with depth.
Among the VIP compounds creatine, carnitine and choline sulphate, are known osmolytes94,95 also called osmoprotectants or compatible solutes, regulating the osmolarity of the cell94. Arsenobetaine96 and acetylcholine95 and phosphocholine (and choline) are suggested to be involved or precursor to betaine synthesis, another osmolyte. The compound 2-methylbutyroylcarnitine97 was found to vary in intertidal mussels (Fig. S9, Table S4 for extended results) and thus could also be an osmolyte. The variation of osmolyte levels seems logical as salinity of the surrounding water varies with depth (Fig. 2). Although quantitative comparisons cannot be made based on our data, it appears that the major osmolytes differed in each holobiont, as did their respective variation with depth.
Ianthelline (a bromotyrosine derivative) and stryphnusin (a bromophenetylamine derivate) were originally isolated from verongiid sponges and are hence likely to be produced by the encrusting verongiid Hexadella dedritifera commonly overgrowing S. fortis98. As we found that most of the samples of S. fortis had traces of overgrowth, we hypothesize that H. dedritifera produces more of the compounds with depth (Fig. 7). Although ianthelline production may vary98, the clear increase with depth is an unexpected finding. In the shallow water verongiid Aplysina aerophoba, bromotyrosine derivative variations are correlated with water temperature and season92, whereas our results suggested other factors associated with depth. The signals of bromotryptophane derivatives and barrettides A and B, as well as C99 in G. barretti remained unaffected by depth.
Bioactivity perspective
In general, diketopiperazines (DKPs) are implicated in bacterial communication as quorum sensing molecules100. G. barretti-associated DKPs (Fig. 7 compounds 1–6) changed strongly in signal intensity across the different depth which would seem illogical under the assumption that quorum sensing was their only function. Indeed, several other bioactivities have been assayed for the barettins (all DKPs except for Pro-Arg). Barettin and 8,9-dihydrobarettin are antifouling56, selective 5-HT serotonin receptor ligands60 and acetylcholine esterase inhibitors25; and geobarrettin B has anti-inflammatory properties58. Interestingly, similar activities have been reported for the tryptophan derivatives (Fig. 7 compounds 7–10). 6-bromoconicamin inhibits acetylcholine esterase25, geobarrettin C has anti-inflammatory properties58 and L-6-bromohypaphorine is an agonist of α7 nicotinic acetylcholine receptor101. It is difficult to disentangle complex interactions e.g. when hypothesising about producers of these compounds among the rich and diverse sponge microbiota but from ranking the correlation analysis in these two compound groups, we found members of Acidobacteria accounted for half the ASV top hits. There are no cultured members from sponges and little is known about their lifestyle, but they are found globally as a dominant group in HMA sponges102. Although only few classes of Acidobacteria have been investigated for their functional potential it is suggested that their genomes frequently encode a large number of specialized metabolites103,104.
However, questions remain as all the ASV top hits were shared by G. barretti and S. fortis, and yet the barettins are characteristic to the former only. Among the ASV top hits, the abundant ASVs 144 and 171 present at high relative abundance are more likely to produce a compound as abundant as barettin (compared to low abundance ASVs 264 and 284 present at low relative abundance).
Conclusions
This study showed that depth impacted prokaryotic communities and metabolites associated with sponge holobionts. Water masses, as approximated by depth, are known to structure marine life and were shown here to stratify holobiont microbiota alike. For both HMA sponges, but especially G. barretti, substitutions of distinct pairs of highly related ASVs present in deeper or shallower specimens, respectively, suggested that a part of the functional potential of the holobionts is preserved despite ASV replacement. Although sponge-associated prokaryotes and metabolites covary with depth, it is challenging to link specific metabolites to specific prokaryotes (or the sponge itself) without additional orthogonal data or study design. Correlating ASV abundance with metabolite signals showed an overrepresentation of members of Acidobacteria among the results. From a bioprospecting and chemical ecology point of view, our finding that sponge samples from different water masses harboured unique ASVs and displayed increase and decrease of bioactive compounds further suggests that the deep-sea and its inhabitants are an untapped source for novel compounds, even for species that have already been screened in shallower waters.
Data availability
Sequencing data was deposited at the NCBI Sequence Read Archive (SRA) with accession number SRP142603 under file numbers SRX3993926–SRX3993970 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA453542). UPLC-HRMS data was deposited at MetaboLights (www.ebi.ac.uk/metabolights/MTBLS1388). Extended methods and data sets were documented in the supplementary information for this article. Analysis code can be found at https://ksteffen.github.io/Paamiut/.
References
Taylor, M. W., Radax, R., Steger, D. & Wagner, M. Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev. 71, 295–347 (2007).
Thomas, T. et al. Diversity, structure and convergent evolution of the global sponge microbiome. Nat. Commun. 7, 11870 (2016).
Webster, N. S. et al. Deep sequencing reveals exceptional diversity and modes of transmission for bacterial sponge symbionts. Environ. Microbiol. 12, 2070–2082 (2010).
Sipkema, D. et al. Similar sponge-associated bacteria can be acquired via both vertical and horizontal transmission: Microbial transmission in Petrosia ficiformis. Environ. Microbiol. 17, 3807–3821 (2015).
Cleary, D. F. R. et al. The sponge microbiome within the greater coral reef microbial metacommunity. Nat. Commun. 10, 1644 (2019).
Björk, J. R., Díez-Vives, C., Astudillo-García, C., Archie, E. A. & Montoya, J. M. Vertical transmission of sponge microbiota is inconsistent and unfaithful. Nat. Ecol. Evol. 3, 1172–1183 (2019).
Webster, N. S. & Taylor, M. W. Marine sponges and their microbial symbionts: Love and other relationships. Environ. Microbiol. 14, 335–346 (2012).
Kennedy, J. et al. Evidence of a putative deep sea specific microbiome in marine sponges. PLoS ONE 9, e91092 (2014).
Steinert, G. et al. Compositional and quantitative insights into bacterial and archaeal communities of south pacific deep-sea sponges (Demospongiae and Hexactinellida). Front. Microbiol. 11, 716 (2020).
Busch, K. et al. On giant shoulders: How a seamount affects the microbial community composition of seawater and sponges. Biogeosciences 17, 3471–3486 (2020).
Olson, J. B. & Gao, X. Characterizing the bacterial associates of three Caribbean sponges along a gradient from shallow to mesophotic depths. FEMS Microbiol. Ecol. 85, 74–84 (2013).
Steinert, G. et al. In four shallow and mesophotic tropical reef sponges from Guam the microbial community largely depends on host identity. PeerJ 4, e1936 (2016).
Morrow, K. M., Fiore, C. L. & Lesser, M. P. Environmental drivers of microbial community shifts in the giant barrel sponge, Xestospongia muta, over a shallow to mesophotic depth gradient. Environ. Microbiol. 18, 2025–2038 (2016).
Ebada, S. S. & Proksch, P. The chemistry of marine sponges. In Handbook of Marine Natural Products (eds Fattorusso, E. et al.) 191–293 (Springer, 2012). https://doi.org/10.1007/978-90-481-3834-0_4.
Kornprobst, J.-M. Porifera (Sponges). Encyclopedia of Marine Natural Products (Wiley, 2014).
Leal, M. C., Puga, J., Serôdio, J., Gomes, N. C. M. & Calado, R. Trends in the discovery of new marine natural products from invertebrates over the last two decades—Where and what are we bioprospecting?. PLoS ONE 7, e30580 (2012).
Blunt, J. W., Copp, B. R., Keyzers, R. A., Munro, M. H. G. & Prinsep, M. R. Marine natural products. Nat. Prod. Rep. 34, 235–294 (2017).
Unson, M. D., Holland, N. D. & Faulkner, D. J. A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Mar. Biol. 119, 1–11 (1994).
Bewley, C. A., Holland, N. D. & Faulkner, D. J. Two classes of metabolites from Theonella swinhoei are localized in distinct populations of bacterial symbionts. Experientia 52, 716–722 (1996).
Wilson, M. C. et al. An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506, 58–62 (2014).
Tianero, M. D., Balaich, J. N. & Donia, M. S. Localized production of defence chemicals by intracellular symbionts of Haliclona sponges. Nat. Microbiol. 4, 1149–1159 (2019).
Ivanišević, J., Thomas, O. P., Lejeusne, C., Chevaldonné, P. & Pérez, T. Metabolic fingerprinting as an indicator of biodiversity: Towards understanding inter-specific relationships among Homoscleromorpha sponges. Metabolomics 7, 289–304 (2011).
Pérez, T. et al. Oscarella balibaloi, a new sponge species (Homoscleromorpha: Plakinidae) from the Western Mediterranean Sea: Cytological description, reproductive cycle and ecology: O. balibaloi: Description, reproductive cycle and ecology. Mar. Ecol. (Berl.) 32, 174–187 (2011).
Reveillaud, J. et al. Relevance of an integrative approach for taxonomic revision in sponge taxa: Case study of the shallow-water Atlanto-Mediterranean Hexadella species (Porifera: Ianthellidae: Verongida). Invertebr. Syst. 26, 230–248 (2012).
Olsen, E. K. et al. Marine AChE inhibitors isolated from Geodia barretti: Natural compounds and their synthetic analogs. Org. Biomol. Chem. 14, 1629–1640 (2016).
Reverter, M., Perez, T., Ereskovsky, A. V. & Banaigs, B. Secondary metabolome variability and inducible chemical defenses in the Mediterranean Sponge Aplysina cavernicola. J. Chem. Ecol. 42, 60–70 (2016).
Reverter, M., Tribalat, M.-A., Pérez, T. & Thomas, O. P. Metabolome variability for two Mediterranean sponge species of the genus Haliclona: Specificity, time, and space. Metabolomics 14, 114 (2018).
Villegas-Plazas, M. et al. Variations in microbial diversity and metabolite profiles of the tropical marine sponge Xestospongia muta with season and depth. Microb. Ecol. 78, 243–256 (2019).
Mohanty, I. et al. Multi-omic profiling of Melophlus sponges reveals diverse metabolomic and microbiome architectures that are non-overlapping with ecological neighbors. Mar. Drugs 18, 124 (2020).
Bowerbank, J. S. On the anatomy and physiology of the Spongiadae. Part I. On the spicula. Philos. Trans. R. Soc. Lond. 148, 279–332 (1858).
Vosmaer, G. C. J. The sponges of the ‘Willem Barents’ expedition 1880 and 1881. Bijdragen tot de Dierkunde 12, 1–47 (1885).
Radax, R. et al. Metatranscriptomics of the marine sponge Geodia barretti: Tackling phylogeny and function of its microbial community. Environ. Microbiol. 14, 1308–1324 (2012).
Topsent, E. Spongiaires provenant des campagnes scientifiques de la ‘Princesse Alice’ dans les Mers du Nord (1898–1899—1906–1907). Résultats des campagnes scientifiques accomplies par le Prince Albert I. Monaco 45, 1–67 (1913).
Yashayaev, I. & Loder, J. W. Further intensification of deep convection in the Labrador Sea in 2016. Geophys. Res. Lett. 44, 1429–1438 (2017).
Gutleben, J. et al. Diversity of tryptophan halogenases in sponges of the genus Aplysina. FEMS Microbiol. Ecol. 95, fiz108 (2019).
Indraningrat, A. et al. Cultivation of sponge-associated bacteria from Agelas sventres and Xestospongia muta collected from different depths. Mar. Drugs 17, 578 (2019).
Ramiro-Garcia, J. et al. NG-Tax, a highly accurate and validated pipeline for analysis of 16S rRNA amplicons from complex biomes. F1000 Res. 5, 1791 (2018).
Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucl. Acids Res. 42, D643–D648 (2014).
Erngren, I., Smit, E., Pettersson, C., Cárdenas, P. & Hedeland, M. The effects of sampling and storage conditions on the metabolite profile of the marine sponge Geodia barretti. Front. Chem. 9:662659 (2021)
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Kuhl, C., Tautenhahn, R., Böttcher, C., Larson, T. R. & Neumann, S. CAMERA: An integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal. Chem. 84, 283–289 (2012).
Oksanen, J. et al. vegan: Community Ecology Package (2017).
Dat, T. T. H., Steinert, G., Thi Kim Cuc, N., Smidt, H. & Sipkema, D. Archaeal and bacterial diversity and community composition from 18 phylogenetically divergent sponge species in Vietnam. PeerJ 6, e4970 (2018).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES science gateway for inference of large phylogenetic trees. In 2010 Gateway Computing Environments Workshop (GCE) 1–8 (IEEE, 2010). https://doi.org/10.1109/GCE.2010.5676129.
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucl. Acids Res. 47, W256–W259 (2019).
Thévenot, E. A., Roux, A., Xu, Y., Ezan, E. & Junot, C. Analysis of the human adult urinary metabolome variations with age, body mass index, and gender by implementing a comprehensive workflow for univariate and OPLS statistical analyses. J. Proteome Res. 14, 3322–3335 (2015).
Weiss, S. et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J. 10, 1669–1681 (2016).
Deng, Y. et al. Molecular ecological network analyses. BMC Bioinform. 13, 113 (2012).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Durno, W. E., Hanson, N. W., Konwar, K. M. & Hallam, S. J. Expanding the boundaries of local similarity analysis. BMC Genom. 14, S3 (2013).
Reshef, D. N. et al. Detecting novel associations in large data sets. Science 334, 1518–1524 (2011).
Hall, M. M., Torres, D. J. & Yashayaev, I. Absolute velocity along the AR7W section in the Labrador Sea. Deep Sea Res. Part 1 Oceanogr. Res. Pap. 72, 72–87 (2013).
Reveillaud, J. et al. Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J. 8, 1198–1209 (2014).
Moitinho-Silva, L. et al. Predicting the HMA-LMA status in marine sponges by machine learning. Front. Microbiol. 8, 752 (2017).
Lidgren, G., Bohlin, L. & Bergman, J. Studies of Swedish marine organisms VII. A novel biologically active indole alkaloid from the sponge Geodia barretti. Tetrahedron Lett. 27, 3283–3284 (1986).
Sjögren, M. et al. Antifouling activity of brominated cyclopeptides from the marine sponge Geodia barretti. J. Nat. Prod. 67, 368–372 (2004).
Sölter, S. Identifizierung und Synthese von Naturstoffen aus Borealen Schwämmen (Universität Hamburg, 2004).
Di, X. et al. 6-Bromoindole derivatives from the Icelandic marine sponge Geodia barretti: Isolation and anti-inflammatory activity. Mar. Drugs 16, 437 (2018).
Carstens, B. B. et al. Isolation, characterization, and synthesis of the barrettides: Disulfide-containing peptides from the marine sponge Geodia barretti. J. Nat. Prod. 78, 1886–1893 (2015).
Hedner, E. et al. Brominated cyclodipeptides from the marine sponge Geodia barretti as selective 5-HT ligands. J. Nat. Prod. 69, 1421–1424 (2006).
Hedner, E. et al. Antifouling activity of a dibrominated cyclopeptide from the marine sponge Geodia barretti. J. Nat. Prod. 71, 330–333 (2008).
Erwin, P. M., Pita, L., López-Legentil, S. & Turon, X. Stability of sponge-associated bacteria over large seasonal shifts in temperature and irradiance. Appl. Environ. Microbiol. 78, 7358–7368 (2012).
Cárdenas, C. A., Bell, J. J., Davy, S. K., Hoggard, M. & Taylor, M. W. Influence of environmental variation on symbiotic bacterial communities of two temperate sponges. FEMS Microbiol. Ecol. 88, 516–527 (2014).
Glasl, B., Smith, C. E., Bourne, D. G. & Webster, N. S. Exploring the diversity-stability paradigm using sponge microbial communities. Sci. Rep. 8, 8425 (2018).
Schöttner, S. et al. Relationships between host phylogeny, host type and bacterial community diversity in cold-water coral reef sponges. PLoS ONE 8, e55505 (2013).
Lurgi, M., Thomas, T., Wemheuer, B., Webster, N. S. & Montoya, J. M. Modularity and predicted functions of the global sponge-microbiome network. Nat. Commun. 10, 992 (2019).
Luter, H. M. et al. Microbiome analysis of a disease affecting the deep-sea sponge Geodia barretti. FEMS Microbiol. Ecol. 93, fix074 (2017).
Thistle, D. Ecosystems of the Deep Oceans (Elsevier, 2003).
Pita, L., Erwin, P. M., Turon, X. & López-Legentil, S. Till death do us part: Stable sponge-bacteria associations under thermal and food shortage stresses. PLoS ONE 8, e80307 (2013).
Webster, N. S., Cobb, R. E. & Negri, A. P. Temperature thresholds for bacterial symbiosis with a sponge. ISME J. 2, 830–842 (2008).
Gerringer, M. E., Drazen, J. C. & Yancey, P. H. Metabolic enzyme activities of abyssal and hadal fishes: Pressure effects and a re-evaluation of depth-related changes. Deep Sea Res. Part 1 Oceanogr. Res. Pap. 125, 135–146 (2017).
Yashayaev, I. Hydrographic changes in the Labrador Sea, 1960–2005. Prog. Oceanogr. 73, 242–276 (2007).
Rhein, M., Steinfeldt, R., Kieke, D., Stendardo, I. & Yashayaev, I. Ventilation variability of Labrador Sea Water and its impact on oxygen and anthropogenic carbon: A review. Philos. Trans. A Math. Phys. Eng. Sci. 375, 20160321 (2017).
Galand, P. E., Potvin, M., Casamayor, E. O. & Lovejoy, C. Hydrography shapes bacterial biogeography of the deep Arctic Ocean. ISME J. 4, 564–576 (2010).
Frank, A. H., Garcia, J. A. L., Herndl, G. J. & Reinthaler, T. Connectivity between surface and deep waters determines prokaryotic diversity in the North Atlantic Deep Water: North Atlantic dark ocean prokaryotic biogeography. Environ. Microbiol. 18, 2052–2063 (2016).
Agogué, H., Lamy, D., Neal, P. R., Sogin, M. L. & Herndl, G. J. Water mass-specificity of bacterial communities in the North Atlantic revealed by massively parallel sequencing. Mol. Ecol. 20, 258–274 (2011).
Djurhuus, A., Boersch-Supan, P. H., Mikalsen, S.-O. & Rogers, A. D. Microbe biogeography tracks water masses in a dynamic oceanic frontal system. R. Soc. Open Sci. 4, 170033 (2017).
Müller, O. et al. Spatiotemporal dynamics of ammonia-oxidizing Thaumarchaeota in distinct Arctic water masses. Front. Microbiol. 9, 1–13 (2018).
Kraemer, S., Ramachandran, A., Colatriano, D., Lovejoy, C. & Walsh, D. A. Diversity and biogeography of SAR11 bacteria from the Arctic Ocean. ISME J. https://doi.org/10.1038/s41396-019-0499-4 (2019).
Monier, A. et al. Upper Arctic Ocean water masses harbor distinct communities of heterotrophic flagellates. Biogeosciences 10, 4273–4286 (2013).
Monier, A. et al. Oceanographic structure drives the assembly processes of microbial eukaryotic communities. ISME J. 9, 990–1002 (2015).
Corrège, T. The relationship between water masses and benthic ostracod assemblages in the western Coral Sea, Southwest Pacific. Palaeogeogr. Palaeoclimatol. Palaeoecol. 105, 245–266 (1993).
Muhling, B. A., Beckley, L. E., Koslow, J. A. & Pearce, A. F. Larval fish assemblages and water mass structure off the oligotrophic south-western Australian coast: SW Australian larval fish assemblages. Fish. Oceanogr. 17, 16–31 (2007).
Eerkes-Medrano, D. et al. A community assessment of the demersal fish and benthic invertebrates of the Rosemary Bank Seamount Marine Protected Area (NE Atlantic). Deep Sea Res. Part 1 Oceanogr. Res. Pap. https://doi.org/10.1016/j.dsr.2019.103180 (2019).
Puerta, P. et al. Influence of water masses on the biodiversity and biogeography of deep-sea benthic ecosystems in the North Atlantic. Front. Mar. Sci. 7, 239 (2020).
Roberts, E. et al. Water masses constrain the distribution of deep-sea sponges in the North Atlantic Ocean and Nordic Seas. Mar. Ecol. Prog. Ser. 659, 75–96 (2021).
Kenchington, E. et al. Connectivity modelling of areas closed to protect vulnerable marine ecosystems in the northwest Atlantic. Deep Sea Res. Part 1 Oceanogr. Res. Pap. 143, 85–103 (2019).
Louca, S. et al. Function and functional redundancy in microbial systems. Nat. Ecol. Evol. 2, 936–943 (2018).
McCauley, M., Chiarello, M., Atkinson, C. L. & Jackson, C. R. Gut microbiomes of freshwater mussels (Unionidae) are taxonomically and phylogenetically variable across years but remain functionally stable. Microorganisms 9, 411 (2021).
Page, M., West, L., Northcote, P., Battershill, C. & Kelly, M. Spatial and temporal variability of cytotoxic metabolites in populations of the New Zealand Sponge Mycale hentscheli. J. Chem. Ecol. 31, 1161–1174 (2005).
Ternon, E., Perino, E., Manconi, R., Pronzato, R. & Thomas, O. P. How environmental factors affect the production of guanidine alkaloids by the Mediterranean sponge Crambe crambe. Mar. Drugs 15, 181 (2017).
Sacristán-Soriano, O., Banaigs, B. & Becerro, M. A. Temporal trends in the secondary metabolite production of the sponge Aplysina aerophoba. Mar. Drugs 10, 677–693 (2012).
Ivanisevic, J. et al. Biochemical trade-offs: Evidence for ecologically linked secondary metabolism of the sponge Oscarella balibaloi. PLoS ONE 6, e28059 (2011).
Burg, M. B. & Ferraris, J. D. Intracellular organic osmolytes: Function and regulation. J. Biol. Chem. 283, 7309–7313 (2008).
Nau-Wagner, G., Boch, J., Le Good, J. A. & Bremer, E. High-affinity transport of choline-O-sulfate and its use as a compatible solute in Bacillus subtilis. Appl. Environ. Microbiol. 65, 560–568 (1999).
Popowich, A., Zhang, Q. & Le, X. C. Arsenobetaine: The ongoing mystery. Natl. Sci. Rev. 3, 451–458 (2016).
Connor, K. M. & Gracey, A. Y. High-resolution analysis of metabolic cycles in the intertidal mussel Mytilus californianus. Am. J. Physiol. Regul. Integr. Comp. Physiol. 302, R103–R111 (2012).
Cárdenas, P. Who produces Ianthelline? The Arctic sponge Stryphnus fortis or its sponge Epibiont Hexadella dedritifera: A probable case of sponge–sponge contamination. J. Chem. Ecol. 42, 339–347 (2016).
Steffen, K. et al. Barrettides: A peptide family specifically produced by the deep-sea sponge Geodia barretti. J. Nat. Prod. 84, 3138–3146 (2021).
Abbamondi, G. R., De Rosa, S., Iodice, C. & Tommonaro, G. Cyclic dipeptides produced by marine sponge-associated bacteria as quorum sensing signals. Nat. Prod. Commun. 9, 229–232 (2014).
Kasheverov, I. et al. 6-Bromohypaphorine from Marine Nudibranch Mollusk Hermissenda crassicornis is an agonist of human α7 nicotinic acetylcholine receptor. Mar. Drugs 13, 1255–1266 (2015).
Moitinho-Silva, L. et al. The sponge microbiome project. Gigascience 6, 1–7 (2017).
Kielak, A. M., Barreto, C. C., Kowalchuk, G. A., van Veen, J. A. & Kuramae, E. E. The ecology of acidobacteria: Moving beyond genes and genomes. Front. Microbiol. 7, 744 (2016).
Crits-Christoph, A., Diamond, S., Butterfield, C. N., Thomas, B. C. & Banfield, J. F. Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis. Nature 558, 440–444 (2018).
Acknowledgements
We gratefully acknowledge Wojciech Walkusz and Tim Siferd, Fisheries & Oceans Canada (DFO), Central and Arctic Division, Winnipeg, Manitoba, for collecting the specimens, and M. Best, G. Thompson, L. Anstey and E. Baker for identification of the specimens on board the R/V Pâmiut. We thank Xiaxia Di, University of Iceland, Reykjavik, Iceland for reference extracts and Luke Odell, Dept. of Medicinal Chemistry, Uppsala University, Uppsala, Sweden for the reference compounds. Thanks also to F. Javier Murillo, DFO and Sarahi Garcia, Stockholm University for insightful discussions on data analyses.
Funding
Open access funding provided by Uppsala University. This study was financially supported by the SponGES project from the European Union’s Horizon 2020 research and innovation programme under Grant Agreement No. 679849. This document reflects only the authors’ view and the Executive Agency for Small and Medium-sized Enterprises (EASME) is not responsible for any use that may be made of the information it contains. Canadian contributions were funded by DFO’s International Governance Strategy “Marine Biological Diversity Beyond Areas of National Jurisdiction (BBNJ): 3-Tiers of Diversity (Genes-Species-Communities)” awarded to Ellen Kenchington. Anak Agung Gede Indraningrat received a PhD fellowship from the Indonesia Endowment Fund for Education (LPDP), Grant Number 20140812021557.
Author information
Authors and Affiliations
Contributions
D.S. and P.C. conceived and designed the study. C.P., L.B., H.S. helped with planning and discussion of the project. A.A.G.I. conducted the microbiome lab and data preparation. I.E., J.H., K.S., P.C. conducted the metabolome lab and data preparation. I.E. identified features and confirmed them with standards. I.Y. performed oceanographic analyses. K.S. analysed the data. K.S., P.C., A.A.G.I., D.S., E.K., I.Y., I.E. drafted the manuscript. All authors commented and accepted the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Steffen, K., Indraningrat, A.A.G., Erngren, I. et al. Oceanographic setting influences the prokaryotic community and metabolome in deep-sea sponges. Sci Rep 12, 3356 (2022). https://doi.org/10.1038/s41598-022-07292-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-07292-3
This article is cited by
-
Prokaryotic communities of the French Polynesian sponge Dactylospongia metachromia display a site-specific and stable diversity during an aquaculture trial
Antonie van Leeuwenhoek (2024)
-
Distribution and diversity of ‘Tectomicrobia’, a deep-branching uncultivated bacterial lineage harboring rich producers of bioactive metabolites
ISME Communications (2023)
-
Candidatus Nemesobacterales is a sponge-specific clade of the candidate phylum Desulfobacterota adapted to a symbiotic lifestyle
The ISME Journal (2023)
-
A Study of Sponge Symbionts from Different Light Habitats
Microbial Ecology (2023)
-
Sponge holobionts shift their prokaryotic communities and antimicrobial activity from shallow to lower mesophotic depths
Antonie van Leeuwenhoek (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.