Abstract
Severe asthma is the main reason for death and disability caused by asthma. However, effective biomarkers for severe asthma have not been identified. Here, we aimed to identify potential biomarkers in severe asthma. We identified 202 differentially expressed genes (DEGs) between severe asthma and mild-moderate asthma after integrating the results from GSE69683 and GSE27011 datasets. The enrichment analysis indicated that 202 DEGs were associated with metabolism- and immune-related processes. 10 hub genes were identified by Cytoscape and five of these genes’ AUC (area under the curve) values were greater than 0.6 in GSE69683. The AUC value reached to 0.701 when combined SEC61A1 and ALDH18A1 expression. The expression of the five hub genes was verified in an external dataset. The network analysis revealed that transcription factor (TF) WT1, ZEB1, RERE, FOSL1, and miR-20a may be involved in the development of asthma. In addition, we found cyclosporine and acetaminophen could interact with these hub genes and may be negatively associated with most of the five hub genes according to previous reports. Overall, key genes were identified between mild-moderate and severe asthmatics, which contributed to the understanding of the development of asthma.
Similar content being viewed by others
Introduction
Severe asthma is characterized by uncontrolled symptoms, airflow obstruction, and frequent exacerbations, which is the major reason for death and disability caused by asthma1. Mild-moderate to severe asthma is based on the level of treatment required to control symptoms or prevent asthma attacks2. Severe asthma affects only 3%–10% of the asthma population, but their health-care is estimated to contribute more than 60% of the cost with asthma3. It also imposes a considerable burden owing to poor symptoms, increased exacerbations, and medication side effects, which have profound effects on emotional and mental health, relationships, and careers4.
Substantial progress in understanding and treating type-2 asthma has been made in the past decades, including evidence-based biomarkers and the availability of novel targeted therapies5. However, more than 50% of patients with non-type-2 severe asthma remain poorly understood and treated6. The result of high-dose inhaled corticosteroids treatment in severe asthma patients is disappointing with uncontrolled symptoms and frequent exacerbations7. A greater mechanistic understanding of different biomarkers and phenotypes of severe asthma is an important step to personalized treatment, as well as being critical for discovering new efficacious therapies targeting non-type-2 pathways8.
The transcriptomic profiling in severe asthma has provided clues and insights on promising biomarkers and therapeutic targets9. A specific signature produced by transcriptome data of epithelial brushings could predict asthma patients’ response to inhaled corticosteroids (ICS) and associated with Th2-high inflammation10,11. In severe asthma, transcriptomic analyses demonstrated that novel biomarkers or phenotypes associated with clinical features12, adult-onset severe asthma13, Th2-high inflammation9,14, and corticosteroid insensitivity15. However, the underlying pathologic mechanism of several asthma is still unclear. In this study, we hypothesized key potential genes drive asthma progression and therefore, are associated with severe asthma. This study aimed to identify differentially expressed genes (DEGs) and potential mechanistic regulation networks between mild-moderate asthma and severe asthma.
Methods
Transcriptome data acquisition
We obtained microarray data for mild-moderate and severe asthma from Gene Expression Omnibus (GEO) database. The inclusive criteria are as follow: (1) Homo sapiens; (2) the number of sample size is more than 30; (3) complete information about the samples; (4) samples are blood source except for the validation dataset. At last, GSE69683 and GSE27011 were chosen for identifying potential biomarkers in the severe asthma; GSE76262 was considered as the validation dataset.
Identification of DEGs
All data was normalized by log2(x + 1) transformed. The “limma” package16 from R software was used to identify DEGs between mild-moderate and severe asthma groups. Only genes’ false discovery rate (FDR) < 0.05 were considered as DEGs.
Enrichment analysis
Gene set enrichment analysis (GSEA) analysis17 was performed using the GSEA software with the gene list of Gene Ontology (GO) gene sets (c5.all.v7.4.symbols.gmt). The "clusterProfiler" R package18 was used to perform GO and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses of these DEGs.
Construction of the protein–protein interaction (PPI) network
Online tool STRING (https://string-db.org/) was used to construct PPI network based on the 202 DEGs. Cytoscape software18 (v3.7.2) was used to visualize the interaction results from the STRING. The cytoHubba plugin19 was used to obtain hub genes according to the degree method.
Receiver operating characteristics (ROC) analysis
For ROC analysis of individual gene, outcome (mild-moderate or severe asthma) and individual gene expression were collected. Using "pROC" R package20, we calculated the AUC value. For ROC analysis of combined gene, outcome and each included gene expression were collected. Firstly, logistic regression was used to calculate the diagnostic score: β1 × gene1 expression + β2 × gene2 expression + β3 × gene3 expression… + βn × genen expression, where β was the correlation coefficient produced by the logistic regression analysis. Then, according to the outcome and diagnostic score, we calculated the AUC value using "pROC" R package. For the recommended model: diagnostic score = − 2.0892*SEC61A1 + (− 1.297)*ALDH18A1.
Construction of transcription factor (TF)-gene and gene-miRNA networks
TF-gene and gene-miRNA networks were constructed using the online tool of NetworkAnalyst (https://www.networkanalyst.ca/). Based on the ENCODE ChIP-seq data (https://www.encodeproject.org/), we constructed the TF-gene network. According to miRTarBase v8.0 (https://mirtarbase.cuhk.edu.cn/php/download.php), we constructed the gene-miRNA network. We extracted expression of TFs and hub genes from GSE69683. Spearman correlation analysis were used to explore the association between hub genes and TFs. "ggplot2" R package (https://ggplot2.tidyverse.org/) was used to visualize the results.
Prediction of interacted chemicals
According to the data from Comparative Toxicogenomics Database (CTD) (http://ctdbase.org/), we constructed the protein-chemical network using the NetworkAnalyst tool.
Statistics analysis
Statistical analyses of all data were performed by R software (version 4.1.1, https://www.r-project.org/). Student’s t-test was used to compare the differences between the two groups. P < 0.05 was considered statistically significant.
Results
Identification of DEGs
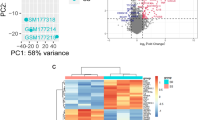
4327 DEGs were identified between severe and mild-moderate asthma in GSE69683, including 2575 downregulated genes and 1752 upregulated genes (Fig. 1A). 650 DEGs were identified between severe and mild-moderate asthma in GSE27011, including 548 downregulated genes and 102 upregulated genes (Fig. 1B). Taking the intersection of the results from GSE69683 and GSE27011, we obtained 202 genes for further explorations (Fig. 1C). Among 202 genes, 23 genes were upregulated and 179 genes were downregulated in GSE69683; 19 genes were upregulated and 183 genes were downregulated in GSE27011 (Fig. 1D). In addition, 8 of 202 genes was found upregulated in one study and downregulated in the other, including FFAR3, ANKRD11, SLC39A14, ZC3H7B, TMEM107, RPUSD3, NDUFA4, and CTBP1-AS2. We consider that two independent datasets presenting contrary expression differences in 8 of 202 genes is acceptable.
Identification of DEGs. The volcano plot for GSE69683 (A) and GSE27011 (B). Red dots represent upregulated genes; blue dots represent downregulated genes; grey dots represent genes with no significant expression differences. (C) Venn diagram for GSE69683 (the blue set) and GSE27011 (the red set). (D) The number of upregulated and downregulated genes of shared 202 genes in GSE69683 and GSE27011, respectively.
Enrichment analysis
GSEA analysis for all genes ranked by log2Fold Change showed that the nucleic acid metabolism-related and immune-related process were downregulated in severe asthma in both GSE69683 and GSE27011 datasets (Fig. 2A,B). The enrichment analysis of shared 202 genes indicated that arginine and proline metabolism, protein export, N-Glycan biosynthesis, cysteine and methionine metabolism and nucleic acid complex were associated with the progression of asthma (Fig. 2C,D).
Enrichment analysis. GSEA analysis for GSE69683 (A) and GSE27011 (B). The results include upper, middle, and lower parts. The upper part: the peak in the right represents core molecules were mainly concentrated in the mild-moderate asthma group; the middle part: each vertical line represents a molecule in the gene set; the lower part: visualization of values given by the uploaded normalized data. GO and KEGG analysis for DEGs displayed in the form of histogram (C) and the network diagram (D).
PPI network analysis
Next, we constructed PPI network of the 202 genes to understand the interactions between these functional proteins (Fig. 3A). Then, the cytoHubba plugin was used to identify hub genes in this network and we obtained 10 hub genes (Fig. 3B). The detailed information of 10 hub genes was shown in Table 1.
PPI network analysis. (A) PPI analysis for all DEGs. Network nodes represent proteins. Edges represent protein–protein associations. Light blue and purple edges mean known interactions; Dark blue, green, and red edges mean predicted interactions. (B) The network of 10 hub genes determined by the cytoHubba. The darker the color, the higher the degree value.
ROC curve of 10 hub genes in mild-moderate and severe asthma samples
In GSE69683, roc curve was used to evaluate 10 hub genes’ distinguishing ability between mild- moderate and severe asthma. The AUC value (> 0.6) of hub genes are as follows: SEC61A1 (0.681, Fig. 4A), ALDH18A1 (0.661, Fig. 4B), TP53 (0.636, Fig. 4C), PDCD11 (0.620, Fig. 4D), MCM3 (0.619, Fig. 4E). The AUC value of other five genes was between 0.5 and 0.6 (Fig. 4F). The five hub genes with AUC value > 0.6 were chosen for a further investigation. Since each single gene with AUC value < 0.7, we tried to combine these genes to increase the AUC value. The results showed that AUC values were 0.702 (combined 10 hub genes, Fig. 4G), 0.696 (combined AUC top 5 genes, Fig. 4H), and 0.701 (combined AUC top 2 genes, Fig. 4I), respectively. Considering the reliability and cost, we recommended combined SEC61A1 and ALDH18A1 genes as a diagnostic model.
Evaluation of the hub genes. ROC (Receiver operating characteristic) curves for SEC61A1 (A), ALDH18A1 (B), TP53 (C), PDCD11 (D), MCM3 (E), the other five genes (F), combined 10 hub genes (G), combined AUC top 5 genes (H), and combined AUC top 2 genes (I). AUC area under the curve, CI Confidence interval.
Verification of the hub gene expression in the external dataset
To confirm whether these hub genes were meaningful in the progression of asthma, we identified their expression in the external dataset (GSE76262, data produced from sputum samples). We consider that if results from different tissues are consistent, this may be more convincing. The results showed that SEC61A1 (Fig. 5A), ALDH18A1 (Fig. 5B), TP53 (Fig. 5C), PDCD11 (Fig. 5D) and MCM3 (Fig. 5E) were lowly expressed in severe asthma, which was consistent with the results in GSE69683 and GSE27011.
TF-gene interactions and gene-miRNA networks
To understand the potential regulation of hub genes, we performed a comprehensive network analysis, including TF and miRNA. The TF-gene interaction network consists of 24 nodes and 45 edges (Fig. 6A). The gene-miRNA network includes 23 nodes and 38 edges (Fig. 6B). Based on the hub genes and predicted TFs, enrichment analysis showed that these genes were associated with cell cycle, transcriptional misregulation and histone deacetylase complex (Fig. 6C). To further understand the relation of activation or repression of TFs with the hub genes, we conducted a correlation analysis in GSE69683. Interestingly, asthma-related TFs, including WT1, ZEB1, and RERE, were negatively associated with most of hub genes (Fig. 6D).
TF-gene interactions and gene-miRNA networks. (A) The TF-gene interaction network. Red spots represent hub genes; blue spots represent TFs. (B) The gene-miRNA network. Red spots represent hub genes; blue rectangles represent miRNAs. (C) GO and KEGG analysis for the TF-gene interaction network displayed in the form of histogram. (D) Heatmap of expression correlation in severe asthmatics from GSE69683. Five hub genes and TFs in the network were analyzed using Spearman method.
Protein–chemical interactions
We consider the hub genes could be potential therapeutic targets for severe asthma patients, so we constructed a network between five hub genes and chemicals. The results showed that cyclosporine can interact with all five genes and acetaminophen can react to four genes. The other four chemicals only interact with two genes (Fig. 7).
Discussion
Identification of novel biomarkers can contribute to the patient classification, therapy response, and clinical outcome prediction21. In asthma, most identified biomarkers are limited to the Th2 phenotype, and no effective biomarkers have been verified for severe asthma22. Blood samples serve as an alternative to the challenging and costly approach of obtaining airways samples in severe asthma23. In this study, we identified five potential biomarkers in severe asthma based on blood samples. TF-gene interactions and gene-miRNA networks associated with the five biomarkers were constructed. In addition, we found two chemicals that have an ability to interact with the identified biomarkers.
Nucleic acid metabolism-related pathways are associated with multiple biological processes, including DNA repair24, chronic inflammation25, collagen deposition26, and so on. Our data showed that dysregulated nucleic acid metabolism-related pathways were involved in the asthma progression. Some cytokines were considered to play important roles in asthma progression. Specifically, Eosinophil-associated fibrogenic factors (such as TGF-β) can lead to airway remodeling characterized by smooth muscle thickening, goblet cell metaplasia, and extracellular matrix protein deposition27. In addition, increased IL-1β and IL-17 also were considered with asthma disease severity28. Meanwhile, TGF-β, IL-1β, and IL-17 were considered as immunosuppressive factors29. Our GSEA results indicated immune-related processes were downregulated in severe asthma. We considered that chronic inflammation in lung may induce an immunosuppressive microenvironment. A typical example is that chronic inflammation promotes cancer progression through immune evasion29. However, immune status is a complex and dynamic process, whether severe asthmatics showing an immunosuppressive microenvironment compared with mild-moderate asthmatics is needed to be further explored.
To date, the five genes in this study have not been reported in severe asthma. Minichromosome maintenance 3 protein (MCM3) protein is a member of 6 highly conserved minichromosome maintenance proteins family (MCM2-MCM7) that bind to replication origins and permit them to maintain a single round of DNA replication. Even a minor reduction of the MCM expression destabilizes the genome and predisposes to increased incidence of diseases30. Previous studies focused on the role of MCM3 in tumor development and growth31 and there is no study about the relationship between MCM3 and asthma. We found MCM3 was lowly expressed in severe asthma, which indicated patients with severe asthma may suffered with the disorder of DNA replication and cell cycle. The transcription factor encoded by TP53 gene is a crucial hurdle to carcinogenesis and inactivation of TP53 is the most common mutation in human cancers32. Moreover, TP53 methylation is a suitable blood biomarker to predict late-onset asthma (LOA)33. PDCD11 (Programmed Cell Death 11) is a NF-kappa-B-binding protein that colocalizes with U3 RNA in the nucleolus and is required for rRNA maturation and generation of 18S rRNA34. Rare studies reported the function of PDCD11 in diseases and this gene is needed to be explored in the future. Aldehyde dehydrogenase 18 Family member A1 (ALDH18A1) is a NAD(P)H- and ATP-dependent enzyme that can convert glutamate to pyrroline-5-carboxylate (P5C), an important intermediate step in proline metabolism. ALDH18A1 is associated with poor prognosis in patients with hepatocellular carcinoma35 and breast cancer36. ALDH18A1 and glutaminase protein are highly expressed in proliferative estrogen receptor positive (ER+) tumor cells compared to ER− cells36. Increasing evidence has convincingly demonstrated that high expression of ALDH18A1 is a poor prognosis factor in different tumor types and increased proline biosynthesis is needed in cancer cells37. SEC61 translocon subunit alpha 1 (SEC61A1) is the main subunit of the SEC61 complex that plays a role in the main polypeptide conduction pathway of the endoplasmic reticulum membrane. Missense mutation of SEC61A1 can induce immune-related diseases, such as plasma cell deficiency38. We found PDCD11 was negatively associated with asthma-related TFs, including WT1, ZEB1, and RERE; ALDH18A1 was negatively associated with WT1 and RERE; SEC61A1 was negatively related with WT1 and ZEB1. Hence, PDCD11, ALDH18A1, and SEC61A1 may play a role in the development of asthma through transcriptional regulation-related mechanisms.
To elucidate the potential mechanism of the five genes in severe asthma, we conducted a TF-gene and gene-miRNA interaction analysis. The identified TFs and miRNAs may be involved in the pathological process of severe asthma. Wilms tumor 1 (WT1) is a suppressor of MMP-9, regulated by a NO-mediated signaling pathway in human lung epithelial cells. MMP-9 expression correlates with the severity of asthma and chronic obstructive pulmonary disease. Zinc finger E-box binding homeobox 1 (ZEB1), an epithelial-mesenchymal transition (EMT) key regulator, has been reported involved in the airway smooth muscle cell proliferation39 and nickel exposure-induced impairment of human lung epithelial cells40. Through transcriptome-wide association studies, transcription factor RERE was identified as a candidate causal gene of asthma41. The whole-genome expression analysis of peripheral blood mononuclear cells indicated that FOSL1 was positively associated with aspirin-intolerant asthma42. In addition, miR-20a epigenetically regulated the IL-10 production by targeting HDAC4 and attenuated allergic inflammation in the human mast cells. Hence, the identified hub genes, TFs and miRNAs may contribute to the progression of asthma.
Through the protein-chemical interaction analysis, we identified cyclosporine and acetaminophen may have effects on severe asthma. Cyclosporin, as an immunosuppressive agent, has benefits in the treatment of inflammatory disorders. Patients with chronic severe asthma are dependent on the oral corticosteroids and alternative therapies are needed to reduce the use of corticosteroids. Cyclosporin has been considered as a potential useful agent in the treatment of chronic severe asthma43,44. Moreover, for lung transplant recipients, improvements in lung function were observed in the group receiving inhaled cyclosporine45. Acetaminophen (paracetamol) is widely used in infants and toddlers as the drug of choice for fever for its recognized safety for most children. However, the findings of a case–control study raised a concern about the safety of acetaminophen that indicated frequent use of acetaminophen was associated with the occurrence and development asthma. This concern led to decades of debate about whether frequent acetaminophen use leads to asthma in children who would otherwise be asthma-free and worsens asthma in patients who already have asthma46. Until now, for pediatricians the question whether “to give or not to give” acetaminophen might be a bitter pill to swallow because ambiguous conclusions about the question. According to the Comparative Toxicogenomics Database and literature review, we found Cyclosporine results in increased expression of TP53 mRNA47,48, increased expression of SEC61A1 mRNA49,50,51, increased expression of PDCD11 mRNA50, decreased expression of ALDH18A1 mRNA50,52, and decreased expression of MCM3 mRNA48,50,53. Moreover, Acetaminophen results in increased expression of SEC61A1 mRNA54,55, increased activity of TP53 protein56, increased expression of TP53 mRNA57,58, increased expression of MCM3 mRNA55, and decreased expression of ALDH18A1 mRNA59,60. Although Cyclosporine and Acetaminophen were reported inhibition the expression of most of five hub genes, the proof were produced from non-asthma population. A correlation with the clinical and complex etiopathogenetic mechanisms of asthma is necessary. In the era of precision medicine, it is important to use specific drugs for increasingly specific asthmatic phenotypes.
Conclusion
We identified 202 DEGs between mild-moderate and severe asthmatics from GSE69683 and GSE27011. The results of GSEA analysis indicated that nucleic acid metabolism-related and immune-related process were downregulated in severe asthma. 10 hub genes were identified using cytoHubba plugin and five of these genes’ AUC values were greater than 0.6. In addition, a diagnostic model combined SEC61A1 and ALDH18A1 genes could effectively distinguish mild-moderate from severe asthmatics (AUC value: 0.701). Finally, cyclosporine and acetaminophen were able to interact with most of the five hub genes. Collectively, we identified key genes and potential pathways between mild-moderate and severe asthmatics and provided clues for the mechanisms of the development of asthma at the transcriptome level.
Data availability
The data can be acquired in GEO database (https://www.ncbi.nlm.nih.gov/geo/).
References
Kim, L. H. Y., Saleh, C., Whalen-Browne, A., O’Byrne, P. M. & Chu, D. K. Triple vs dual inhaler therapy and asthma outcomes in moderate to severe asthma: A systematic review and meta-analysis. JAMA 325(24), 2466–2479 (2021).
Michalovich, D. et al. Obesity and disease severity magnify disturbed microbiome-immune interactions in asthma patients. Nat. Commun. 10(1), 5711 (2019).
Israel, E. & Reddel, H. K. Severe and difficult-to-treat asthma in adults. N. Engl. J. Med. 377(10), 965–976 (2017).
Hyland, M. E., Whalley, B., Jones, R. C. & Masoli, M. A qualitative study of the impact of severe asthma and its treatment showing that treatment burden is neglected in existing asthma assessment scales. Qual. Life Res. 24(3), 631–639 (2015).
Papi, A., Brightling, C., Pedersen, S. E. & Reddel, H. K. Asthma. Lancet 391(10122), 783–800 (2018).
Wenzel, S. E. Severe adult asthmas: Integrating clinical features, biology, and therapeutics to improve outcomes. Am. J. Respir. Crit. Care Med. 203(7), 809–821 (2021).
Sánchez-Ovando, S., Simpson, J. L., Barker, D., Baines, K. J. & Wark, P. A. B. Transcriptomics of biopsies identifies novel genes and pathways linked to neutrophilic inflammation in severe asthma. Clin. Exp. Allergy 51, 1279–1294 (2021).
Robinson, D. et al. Revisiting Type 2-high and Type 2-low airway inflammation in asthma: Current knowledge and therapeutic implications. Clin. Exp. Allergy 47(2), 161–175 (2017).
Rossios, C. et al. Sputum transcriptomics reveal upregulation of IL-1 receptor family members in patients with severe asthma. J. Allergy Clin. Immunol. 141(2), 560–570 (2018).
Woodruff, P. G. et al. Genome-wide profiling identifies epithelial cell genes associated with asthma and with treatment response to corticosteroids. Proc. Natl. Acad. Sci. USA. 104(40), 15858–15863 (2007).
Woodruff, P. G. et al. T-helper type 2-driven inflammation defines major subphenotypes of asthma. Am. J. Respir. Crit. Care Med. 180(5), 388–395 (2009).
Hekking, P. P. et al. Transcriptomic gene signatures associated with persistent airflow limitation in patients with severe asthma. Eur. Respir. J. 50(3), 1609923 (2017).
Hekking, P. P. et al. Pathway discovery using transcriptomic profiles in adult-onset severe asthma. J. Allergy Clin. Immunol. 141(4), 1280–1290 (2018).
Kuo, C. S. et al. T-helper cell type 2 (Th2) and non-Th2 molecular phenotypes of asthma using sputum transcriptomics in U-BIOPRED. Eur. Respir. J. 49(2), 1602135 (2017).
Kuo, C. S. et al. A transcriptome-driven analysis of epithelial brushings and bronchial biopsies to define asthma phenotypes in U-BIOPRED. Am. J. Respir. Crit. Care Med. 195(4), 443–455 (2017).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47 (2015).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102(43), 15545–15550 (2005).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287 (2012).
Chin, C. H. et al. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8(Suppl 4), S11 (2014).
Robin, X. et al. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 12, 77 (2011).
Chiappori, A. et al. Biomarkers and severe asthma: A critical appraisal. Clin. Mol. Allergy 13, 20 (2015).
Hachim, M. Y. et al. Derangement of cell cycle markers in peripheral blood mononuclear cells of asthmatic patients as a reliable biomarker for asthma control. Sci. Rep. 11(1), 11873 (2021).
Shin, S. W. et al. Asthma-predictive genetic markers in gene expression profiling of peripheral blood mononuclear cells. Allergy Asthma Immunol. Res. 3(4), 265–272 (2011).
Glousker, G., Briod, A. S., Quadroni, M. & Lingner, J. Human shelterin protein POT1 prevents severe telomere instability induced by homology-directed DNA repair. EMBO J. 39(23), e104500 (2020).
Brégnard, C., Benkirane, M. & Laguette, N. DNA damage repair machinery and HIV escape from innate immune sensing. Front. Microbiol. 5, 176 (2014).
Senavirathna, L. K. et al. Long noncoding RNA FENDRR inhibits lung fibroblast proliferation via a reduction of β-catenin. Int. J. Mol. Sci. 22(16), 8536 (2021).
Kanda, A. et al. Multiple biological aspects of eosinophils in host defense, eosinophil-associated diseases, immunoregulation, and homeostasis: Is their role beneficial, detrimental, regulator, or bystander?. Biol. Pharm. Bull. 43(1), 20–30 (2020).
Hammad, H. & Lambrecht, B. N. The basic immunology of asthma. Cell 184(9), 2521–2522 (2021).
Garner, H. & de Visser, K. E. Immune crosstalk in cancer progression and metastatic spread: A complex conversation. Nat. Rev. Immunol. 20(8), 483–497 (2020).
Sedlackova, H. et al. Equilibrium between nascent and parental MCM proteins protects replicating genomes. Nature 587(7833), 297–302 (2020).
Løkkegaard, S. et al. MCM3 upregulation confers endocrine resistance in breast cancer and is a predictive marker of diminished tamoxifen benefit. NPJ Breast Cancer 7(1), 2 (2021).
Boutelle, A. M. & Attardi, L. D. p53 and tumor suppression: it takes a network. Trends Cell Biol. 31(4), 298–310 (2021).
Yuan, L. et al. The DNA methylation of FOXO3 and TP53 as a blood biomarker of late-onset asthma. J. Transl. Med. 18(1), 467 (2020).
Sweet, T., Khalili, K., Sawaya, B. E. & Amini, S. Identification of a novel protein from glial cells based on its ability to interact with NF-kappaB subunits. J. Cell. Biochem. 90(5), 884–891 (2003).
Ding, Z. et al. Metabolic pathway analyses identify proline biosynthesis pathway as a promoter of liver tumorigenesis. J. Hepatol. 72(4), 725–735 (2020).
Craze, M. L. et al. MYC regulation of glutamine-proline regulatory axis is key in luminal B breast cancer. Br. J. Cancer 118(2), 258–265 (2018).
Geng, P., Qin, W. & Xu, G. Proline metabolism in cancer. Amino Acids 53, 1769–1777 (2021).
Bolar, N. A. et al. Heterozygous loss-of-function SEC61A1 mutations cause autosomal-dominant tubulo-interstitial and glomerulocystic kidney disease with anemia. Am. J. Hum. Genet. 99(1), 174–187 (2016).
Gao, Y., Wang, B., Luo, H., Zhang, Q. & Xu, M. miR-217 represses TGF-β1-induced airway smooth muscle cell proliferation and migration through targeting ZEB1. Biomed. Pharmacother. 108, 27–35 (2018).
Jose, C. C. et al. Nickel exposure induces persistent mesenchymal phenotype in human lung epithelial cells through epigenetic activation of ZEB1. Mol. Carcinog. 57(6), 794–806 (2018).
Valette, K. et al. Prioritization of candidate causal genes for asthma in susceptibility loci derived from UK Biobank. Commun. Biol. 4(1), 700 (2021).
Wieczfinska, J. et al. The whole-genome expression analysis of peripheral blood mononuclear cells from aspirin sensitive asthmatics versus aspirin tolerant patients and healthy donors after in vitro aspirin challenge. Respir. Res. 16, 147 (2015).
Coren, M. E., Rosenthal, M. & Bush, A. The use of cyclosporin in corticosteroid dependent asthma. Arch. Dis. Child 77(6), 522–523 (1997).
Bush, A. & Saglani, S. Management of severe asthma in children. Lancet 376(9743), 814–825 (2010).
Groves, S. et al. Inhaled cyclosporine and pulmonary function in lung transplant recipients. J. Aerosol. Med. Pulm. Drug. Deliv. 23(1), 31–39 (2010).
Litonjua, A. A. Acetaminophen and asthma: A small sigh of relief?. N. Engl. J. Med. 375(7), 684–685 (2016).
Sereno, J. et al. Conversion to sirolimus ameliorates cyclosporine-induced nephropathy in the rat: Focus on serum, urine, gene, and protein renal expression biomarkers. BioMed Res. Int. 2014, 576929 (2014).
Jennen, D. G. et al. Comparison of HepG2 and HepaRG by whole-genome gene expression analysis for the purpose of chemical hazard identification. Toxicol. Sci. 115(1), 66–79 (2010).
Wolters, J. E. et al. Integrative “-omics” analysis in primary human hepatocytes unravels persistent mechanisms of cyclosporine A-induced cholestasis. Chem. Res. Toxicol. 29(12), 2164–2174 (2016).
Van den Hof, W. F. et al. Integrating multiple omics to unravel mechanisms of Cyclosporin A induced hepatotoxicity in vitro. Toxicol. In Vitro 29(3), 489–501 (2015).
Schaap, M. M. et al. A novel toxicogenomics-based approach to categorize (non-)genotoxic carcinogens. Arch. Toxicol. 89(12), 2413–2427 (2015).
Limonciel, A. et al. Transcriptomics hit the target: Monitoring of ligand-activated and stress response pathways for chemical testing. Toxicol. In Vitro 30(1), 7–18 (2015).
Magkoufopoulou, C., Claessen, S. M., Jennen, D. G., Kleinjans, J. C. & van Delft, J. H. Comparison of phenotypic and transcriptomic effects of false-positive genotoxins, true genotoxins and non-genotoxins using HepG2 cells. Mutagenesis 26(5), 593–604 (2011).
Yu, D. et al. Multiple microRNAs function as self-protective modules in acetaminophen-induced hepatotoxicity in humans. Arch. Toxicol. 92(2), 845–858 (2018).
Prot, J. M. et al. Predictive toxicology using systemic biology and liver microfluidic “on chip” approaches: Application to acetaminophen injury. Toxicol. Appl. Pharmacol. 259(3), 270–280 (2012).
Vatakuti, S., Schoonen, W. G., Elferink, M. L., Groothuis, G. M. & Olinga, P. Acute toxicity of CCl4 but not of paracetamol induces a transcriptomic signature of fibrosis in precision-cut liver slices. Toxicol. In Vitro 29(5), 1012–1020 (2015).
Verstraelen, S. et al. Phenotypic and biomarker evaluation of zebrafish larvae as an alternative model to predict mammalian hepatotoxicity. J. Appl. Toxicol. 36(9), 1194–1206 (2016).
Hrach, J., Mueller, S. O. & Hewitt, P. Development of an in vitro liver toxicity prediction model based on longer term primary rat hepatocyte culture. Toxicol. Lett. 206(2), 189–196 (2011).
Jiang, J. et al. Increased mitochondrial ROS formation by acetaminophen in human hepatic cells is associated with gene expression changes suggesting disruption of the mitochondrial electron transport chain. Toxicol. Lett. 234(2), 139–150 (2015).
Gore, M. A., Morshedi, M. M. & Reidhaar-Olson, J. F. Gene expression changes associated with cytotoxicity identified using cDNA arrays. Funct. Integr. Genomics 1(2), 114–126 (2000).
Funding
This work was supported by the science and technology development fund project of Nanjing Medical University (NMUB2019224; NMUB2019221; NMUB2020115; NMUB2020112).
Author information
Authors and Affiliations
Contributions
X.W. and R.L. designed this work. R.L. and Q.X. integrated and analyzed the data. R.L., X.W., F.L., and Y.J. wrote this manuscript. M.T. and M.Z. edited and revised the manuscript. All authors approved this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, X., Li, R., Xu, Q. et al. Identification of key genes and pathways between mild-moderate and severe asthmatics via bioinformatics analysis. Sci Rep 12, 2549 (2022). https://doi.org/10.1038/s41598-022-06675-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06675-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.