Abstract
The widespread use of electronic cigarettes (e-cig) is a serious public health concern; however, mechanisms by which e-cig impair the function of airway epithelial cells—the direct target of e-cig smoke—are not fully understood. Here we report transcriptomic changes, including decreased expression of many ribosomal genes, in airway epithelial cells in response to e-cig exposure. Using RNA-seq we identify over 200 differentially expressed genes in air–liquid interface cultured primary normal human bronchial epithelial (NHBE) exposed to e-cig smoke solution from commercial e-cig cartridges. In particular, exposure to e-cig smoke solution inhibits biological pathways involving ribosomes and protein biogenesis in NHBE cells. Consistent with this effect, expression of corresponding ribosomal proteins and subsequent protein biogenesis are reduced in the cells exposed to e-cig. Gas chromatography/mass spectrometry (GC/MS) analysis identified the presence of five flavoring chemicals designated as ‘high priority’ in regard to respiratory health, and methylglyoxal in e-cig smoke solution. Together, our findings reveal the potential detrimental effect of e-cig smoke on ribosomes and the associated protein biogenesis in airway epithelium. Our study calls for further investigation into how these changes in the airway epithelium contribute to the current epidemic of lung injuries in e-cig users.
Similar content being viewed by others
Introduction
The widespread use of electronic cigarettes (e-cig) is a significant public health concern. In 2017, 6.9 million (1 in 36) of adults in the US were current e-cig users1. In addition, more than 3.6 million youth, including 20.8% high school students and 4.9% middle school students, in the US currently used e-cig in 20182. Recently, multiple studies reported a cluster of cases with acute, server respiratory distress in e-cig users3,4,5. The Centers for Disease Control and Prevention reported 2807 hospitalized lung injury cases from all 50 states, the District of Columbia, and two U.S. territories and 68 deaths in 29 states and the District of Columbia from people have a history of e-cig use or vaping as of February 18, 20206. Although recent investigations have shown that vitamin E acetate, an additive in some tetrahydrocannabinol (THC)-containing e-cigarette or vaping products, is strongly linked to this outbreak, contribution of other chemicals in either THC or non-THC products still needs further investigations.
E-cig fluid contains numerous chemicals known to be toxic to the respiratory system, such as carbonyl compounds, aldehydes, fine particulate matter, metals, propylene glycol, glycerol, formaldehyde, volatile organic compounds (VOCs), and flavoring chemicals7,8,9,10,11,12,13,14,15,16,17, and bacterial endotoxins and fungal glucans18. The recent cases of lung injuries are characterized by non-infectious pneumonia-like symptoms with widely varying severity and types of inflammation3,4,5,19. This heterogeneity likely stems from the combinational effects of multiple chemicals and toxins in e-cig fluid19. Airway epithelium is the first line of defense in the lung and the direct target of chemicals in e-cig products. Although recent studies reported adverse impact of e-cig on airway epithelium20,21,22,23,24,25,26, the underlying mechanisms are not fully understood.
To fill these knowledge gaps, this study was aimed to identify transcriptomic changes and impacted biological pathways in human airway epithelium exposed to e-cig smoke solution from commercial e-cig products. We performed global transcriptomic profiling using RNA-seq in primary normal human bronchial epithelial (NHBE) cells, cultured at an air–liquid interface (ALI) to mimic the in vivo airway characteristics27, exposed to e-cig smoke solution obtaining by puffing commercial e-cig cartridges. We then conducted DAVID pathway analysis followed by qRT-PCR and in vitro mechanistic study to further validate RNA-seq results. We also analyzed flavoring chemicals and dicarbonyls in the e-cig smoke solution using Gas chromatography/mass spectrometry (GC/MS).
Results
Chemical analysis of e-cig smoke solution
To generate e-cig smoke solution that closely mimics the real-world exposure to e-cig users, commercial e-cig cartridges were puffed using a smoking machine. The TE-2B smoking machine puffed twenty e-cig cartridges and the e-cig downstream smoke was collected through an impinger filled with ultrapure water (Fig. 1a). We then used GC/MS analysis to identify and quantify chemicals present in the e-cig smoke solution. Table 1 lists the compounds quantified in rank order from highest to lowest concentrations. This includes five flavoring chemicals designated as ‘high priority’ by a flavoring industry report on respiratory hazards associated with flavoring chemicals (diacetyl, benazaldehyde, acetaldehyde, propionaldehyde, and acetoin)17. In addition to these flavoring chemicals, we also identified the presence of methylglyoxal, an alpha dicarbonyl that is structurally related to diacetyl which can form after propylene glycol is heated in e-cig28. Three vanilla flavorings including ethylvanillin, 4-methoxybenzaldehyde and vanillin represented 99.7% of the compounds quantified as the vanilla e-cig flavor was used to generate the solution.
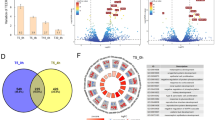
Identification of differential gene expression in NHBEs exposed to e-cigarette smoke solution by RNA-seq. (A) Preparation of e-cig smoke solution. (B) Schematic workflow of the study. (C) Cytotoxicity of e-cig smoke solution in NHBE cells. (D) Volcano plot of RNA-seq results with top 10 genes annotated in NHBE cells exposed to e-cig smoke solution for 24 h. Block dots represent gene with Padj < 0.05. Grey dots represent genes that do not meet the significance threshold. (E) qPCR validation of top 10 genes identified by RNA-seq with e-cig smoke solution. *, P < 0.05 compared to control. N = 3 subjects.
Transcriptomic profiling of human bronchial epithelial cells exposed to e-cig smoke solution
To examine the effect of e-cig on human airway epithelium, we utilized air–liquid interface (ALI) cultures of primary NHBE cells. NHBE cells cultured under ALI after 14 days differentiate into a mixture of ciliated cells, goblet cells as well as some remaining basal cells, thus closely mimicking human airway epithelium in vivo27. Immunofluorescence staining of NHBE cells confirms the presence of ciliated and goblet cells at ALI day 14 indicating the well-differentiated state of the cells (Supplementary Fig. S1). We first exposed NHBE cells to control (DI water dissolved in culture medium, 2% v/v) or e-cig smoke solution (2% v/v, containing 2 ppm diacetyl) for 24 h (Fig. 1b). The rationale for choosing 2 ppm diacetyl was that the maximum diacetyl concentration/e-cig was about 2 ppm based on the calculation using the data from the previous study17. LDH (lactate dehydrogenase) assay confirmed that e-cig smoke solution did not induce significant cytotoxicity at 6 or 24 h (Fig. 1c). After 24-h incubation, total RNAs from the cells were extracted and used for RNA-Seq-based transcriptional profiling. RNA-seq libraries were constructed, each with a unique barcode that allows multiplexing. We obtained an average of ~ 18 million reads per sample and tested for differential expression of GRCh37 Ensemble-annotated genes. Following multiple testing corrections, we identified a total of 201 genes that were differentially regulated with e-cig treatment as shown in the volcano plot in Fig. 1d (Black dots, padj ≤ 0.05). A list of all statistically significant results is included in Excel File Table S1.
qPCR validation of differentially regulated genes by e-cig smoke solution
From the list of differentially regulated genes (Excel File Table S1), we ranked the differentially regulated genes by fold change. We then selected a total of 10 genes (five most upregulated and five most downregulated genes) with each treatment for qRT-PCR validation. (Fig. 1d; Table 2). We treated ALI cultures of primary NHBE cells derived from three different donors with vehicle control or e-cig smoke solution. Out of the top 10 differentially expressed genes (Table 2), qRT-PCR verified 5 genes that were significantly changed compared to control. E-cig treatment increased expression of CYP1B1, CYP1A1 while suppressing the expression of RPL30, RPL37A, and CLIP (Fig. 1e, p < 0.05). Because our previous study reported the transcriptional profiling of NHBE cells exposed to flavoring chemicals in e-cig products29, we compared the lists of significantly regulated genes exposed to diacetyl, 2,3-pentanedione, or e-cig smoke solution. We identified 24 common genes that were significantly changed with all three treatments (padj < 0.05, Supplementary Table S3). In addition, based on our previous findings showing the downregulation of cilia-involved genes by flavoring chemicals29, we tested the effect of e-cig smoke solution on selected cilia-related genes in NHBE cells. Consistent with previous findings, e-cig treatment downregulated expression of cilia genes in NHBE cells as shown in Supplementary Fig. S2.
Functional annotation enrichment using DAVID
We then used the DAVID pathway analysis tool to identify ontological categories that are enriched among the differentially expressed genes induced by exposures to e-cig smoke solution. As shown in Table 3, pathways related to “SRP-dependent cotranslational protein targeting to membrane”, “Translation initiation”. “Ribosomal protein”, “Inflammatory response”, and “Cytokine”. Interestingly, the first cluster with the highest enrichment score includes pathways related to mRNA translation, ribosomes, and protein biogenesis, suggesting e-cig may perturb ribosomal functions and protein biogenesis. The full list of enriched terms is available in Excel File Table S2.
Expression of ribosomal protein genes
Because the pathways involved in translation and ribosomal proteins have the lowest p-values and highest enrichment score (Table 3), we focused our follow up studies on the genes involved in ribosomal proteins. Our RNA-seq data showed that expression of 14 genes related to ribosome (RPL30, RPL37A, RPS14, RPS19, RPL7A, RPS12, RPS11, RPL32, RPL10A, RPL 19, RPL35, RPS5, RPS21, and RPS 27) was significantly down-regulated in NHBE cells treated with with e-cig treatment except that expression of RPS29 was up-regulated (Table 4). Using qRT-PCR, we validated the down-regulation of selected genes in NHBE cells from three different donors upon e-cig treatment. Consistent with RNA-seq data, mRNA expression of RPS14, RPS19, RPL30, and RPL37A was significantly down-regulated with e-cig smoke solution (Fig. 2a, p < 0.05), suggesting that e-cig may impair ribosomal functions and protein biogenesis.
Effect of e-cig smoke solution on expression of ribosomal proteins in NHBE cells (A) Expression of ribosomal protein-encoding genes measured by qPCR. (B) Expression of ribosomal protein-encoding genes in NHBE cells at varying concentrations of e-cig solution measured by qRT-PCR. (C) Protein quantification of ribosomal proteins measured by Western blot. *p < 0.05 significant compared to control, N = 3 experiments.
To investigate the concentration-dependent effect of e-cig on ribosomal protein genes, we exposed NHBE cells to 0, 1, 2, and 5 ppm of e-cig smoke solution, then performed qPCR for RPL30, RPL37A, RPS14, and RPS19. Down-regulation of RPL37A was observed with e-cig smoke solution at levels as low as 1 ppm (diacetyl) (Fig. 2b, p < 0.05). Treatment with the higher concentration of e-cig solution (5 ppm diacetyl) also significantly suppressed expression of RPS14, RPS19, RPL30, and RPL37A (Fig. 2b, p < 0.05). To further validate these findings, protein expression of selected targets were measured by Western blot assay. The three top hits RPL30, RPL37A, and RPS14 were selected from Table 4. RPS 14 expression showed a significant, but modest decrease at 2 ppm at 24 h. RPL 30 expression decreased at 2 and 5 ppm at 24 h, but was not significant (Fig. 2c). We suggest that the abundance of ribosomal proteins in the cells, the modest transcriptional changes, and relative insensitivity of western blot assay may have led to the difficulty in detecting protein changes. The full length blots are shown in Supplementary Fig. S4. RPL37A was not included due to non-specific binding of its antibody.
Effect of e-cig smoke solution on ribosomal RNA transcription and protein synthesis
To further investigate the effect of e-cig on ribosomal biogenesis, we measured transcriptional levels of ribosomal RNAs including pre-rRNA transcripts and mature rRNAs. Pre-rRNA processing begins on the 47S pre-rRNA transcript, which gives rise to 45S pre-rRNA and ultimately to mature 18S, 5.8S, and 25/28S rRNAs in functional ribosomal subunits through a complex series of processing, modification, and folding steps30,31,32. Within 47S pre-rRNA transcript, the 18S, 5.8S, and 28S rRNAs are flanked by the 5′ and 3′ external transcribed spacers (ETS) and two internal transcribed spacers (ITS1 and ITS2)30,31,32. E-cig treatment at 2 ppm for 24 h decreased the expression of 47S pre-rRNA as indicated by the 5′-ETS level and increased ITS2 in NHBE cells (Supplementary Fig. S3). However, levels of ITS1 and mature rRNAs, including 28S, 18S, and 5.8S were not altered by e-cig treatment (Supplementary Fig. S3).
To investigate the effect of e-cig on protein biogenesis, we exposed primary NHBE cells to e-cig smoke solution for 24 or 48 h and performed protein synthesis assay. As shown in Fig. 3, protein synthesis at 24 h was significantly decreased by 88% and 23% at 2 and 5 ppm, respectively, compared to control (p < 0.05). At 48 h, protein synthesis was significantly decreased by 79%, 74%, and 18% at 1, 2, and 5 ppm, respectively (p < 0.05). These data indicate that exposure to e-cig may disrupt protein biogenesis in lung epithelium.
Discussion
The recent outbreak of e-cig or vaping associated lung injuries calls for immediate studies on their impact on human lung. Although human airway epithelium is the first line of defense in the lung and is the direct target of external chemicals, little is known about how chemicals in e-cig smoke may impair the lung epithelium and its function. The present study shows that e-cig smoke solution induces significant transcriptomic changes, including those related to ribosomal proteins and translation, affects transcription of rRNAs, and decreases new protein synthesis in primary human airway epithelial cells. The findings in this study implicate potential mechanisms by which e-cig smoking impact lung epithelium, leading to increased risks for lung diseases.
The present study performed chemical analysis on dicarnonyls and flavors in e-cig smoke solution collected from commercial e-cig products. The analysis identified “high priority” flavoring chemicals (diacetyl, benazaldehyde, acetaldehyde, propionaldehyde, and acetoin) that are associated with respiratory injuries17. Among them, exposure to diacetyl after heating and inhaling has been attributed to increased risk of developing bronchiolitis obliterans33. Our previous study found that diacetyl induce significant transcriptomic changes, including those related to ciliogenesis, and decrease the number of ciliated cells in NHBE cells29, implicating potential impairment of the cilia function in human airway epithelium. In addition to flavoring chemicals, we also identified the presence of methylglyoxal, an alpha dicarbonyl that is structurally related to diacetyl which can form after propylene glycol is heated in e-cigs28. Recent toxicological evidence found that methylglyoxal causes necrosis of respiratory epithelium and is more cytotoxic than diacetyl34. Mixtures of chemicals found in e-cig smoke solution may act synergistically targeting lung epithelium. Whether such cumulative influence extends to compromised cell function and lung injuries needs further investigations.
We identified that exposure to e-cig smoke solution led to significant transcriptomic changes related to protein synthesis in primary human airway epithelial cells. Specifically, expression of genes encoding ribosomal proteins, such as RPS14, RPS19, RPL30, and RPL37A, is down-regulated in NHBE cells exposed to e-cig smoke solution. Consistent with this, protein biogenesis was suppressed in NHBE cells exposed to e-cig smoke solution. In addition, we found that transcription of ribosomal RNAs was differentially regulated with e-cig smoke solution although the impact of these changes on overall protein translation needs further studies. Because ribosome biogenesis is a complex process requiring tight coordination of ribosomal RNA and ribosomal protein production35, our data implicates potential imbalance of rRNA and ribosome protein synthesis by e-cig exposure, thereby leading to disruption of ribosome biogenesis. Ribosome biogenesis and protein translation are finely coordinated with and essential for cell growth, proliferation, differentiation, and development. Impairment of any of these cellular processes can perturb cell growth and development36. It is reported that p53 play critical roles in sensing and regulating ribosomal stress and biogenesis, which in turn regulate cell cycle progression and cell growth to prevent carcinogenesis37. Furthermore, the balance between rRNA and ribosomal protein synthesis has been shown to control the function of p53 in mammalian cells38. A recent oral transcriptome analysis in by RNA-Seq revealed “cancer” as the top disease associated with dysregulated genes in e-cig users39. Further study on the role of p53 activation in the regulation of e-cig-mediated ribosomal stress and cell cycle regulation in relevant with carcinogenesis would be of great interest.
Regulation of translation modulates immune functions by directly affecting antigen presentation, cytokine production, as well as the survival of dendritic cells40. Cells need to rapidly activate the inflammatory protein synthesis in response to infection and shut it down when no longer needed41. Therefore, downregulation of ribosomal proteins and protein translation may potentially compromise immune functions against pathogens. On the other hand, it is reported that suppression of ribosomal function and protein synthesis activates innate immune signaling through activation of the NLRP3 inflammasome in murine bone marrow-derived macrophages42. In addition, the present study showed that e-cig smoke solution activates pathways related to inflammatory response and cytokines in NHBE cells (Table 3). Similarly, it is reported that e-cigs with flavorings induce inflammatory and pro-senescence responses in oral epithelial cells and periodontal fibroblasts43. Further study on the interaction between ribosomal proteins and immune responses and its consequences on the functions of human airway epithelium will be warranted.
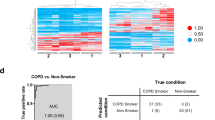
Among the differentially regulated genes with e-cig smoke solution treatment, qRT-PCR validated up-regulation of CYP1A1 and CYP1B1, which encode Cytochromes P450 (CYPs), key metabolizing enzymes for drugs and xenobiotics44. This is consistent with the previous reports showing induction of CYPs by treatment of e-cig in vitro or in vivo. For example, treatment with nicotine and e-cig fluid led to significant induction of mRNA expression for CYP2A6, CYP2E1, CYP1A1 and CYP2S1 following in an in vitro model of human brain blood barrier, the hCMEC/D3 cell line45. In addition, exposure to e-cig vapor increased expression of CYP1A1/2, CYP2B1/2, and CYP3A in the lungs of rats, implicating potential risks for lung cancer by bioactivation of procarcinogenes in e-cig vapor46. Indeed, recent studies showed that mice exposed to e-cig smoke resulted in adenocarcinomas, bladder urothelial hyperplasia, and DNA damage in mice lungs and bladders implicating e-cig smoke as a potential carcinogen in mice47,48. Because the present study showed the upregulation of CYP genes in human primary lung epithelial cells, further study will be warranted to investigate potential carcinogenic effects on human lungs. Interestingly, differential expression of ribosomal proteins has been also implicated in human cancer including lung cancer49,50. It is implicated that dysregulation of ribosomal proteins and CYPs by exposure to e-cig may have combinational/synergistic effect on cell proliferation and carcinoma in the lung.
Our previous study showed that exposure to flavoring chemicals found in e-cig such as diacetyl and 2,3-pentanedione downregulated genes related to ciliogenesis in primary NHBE cells29. GC/MS analysis in the present study identified diacetyl in e-cig smoke solution (Table 1). We further showed that treatment with e-cig smoke solution led to suppression of cilia-related genes in NHBE cells consistent with the effect of flavoring chemicals on these genes29, although this effect of e-cig smoke solution may not be solely mediated by flavoring chemicals. In addition to cilia genes, we found 24 common genes that are significantly regulated in all three groups (diacetyl, 2,3-pentanedione, and e-cig smoke solution) (Supplementary Table S3). However, different concentrations and mixture effect of various chemicals in e-cig smoke solution may contribute to the divergent responses in the expression of same genes. Full spectrum analysis of chemical composition in e-cig smoke solution will be necessary to fully understand the effect of e-cig smoke solution on lung epithelium and identify key players mediating the effect.
This is the first whole transcriptomic profiling in NHBE cells exposed to e-cig smoke solution from commercial e-cig products, however, there are a few limitations. First, mechanisms by which e-cig smoke solution impact protein synthesis is not known. Although the present study implicates that downregulation of ribosomal proteins-encoding genes may lead to retarded protein translation, it is not clear whether this phenomenon is through dysregulation of ribosomal functions or through extra-ribosomal functions that are involved in cell proliferation, differentiation, apoptosis, DNA repair, and other cellular processes50. We detected a modest change in protein expression of RPS 14 and RPL 30 and transcription of rRNAs, this may not fully explain the dramatic decrease of protein biosynthesis in Fig. 3A. Second, we also do not know which specific chemicals in e-cig smoke solution contribute to this outcome. Although we performed GC/MS for dicarbonyls and flavors, there is a lack of information on other chemical components such as nicotine, aldehydes, fine particulate matter, metals, propylene glycol, glycerol, formaldehyde7,8,9,10,11,12,13,14,15,16,17. Besides, since the e-cig smoke solution used in this study was collected in 2016, it may not accurately reflect the current formulations of e-cig products. Therefore, comprehensive chemical analysis on the latest e-cig formulation will be needed. Third, NHBE cells were exposed to aqueous solution of e-cig smoke that may not reflect real human exposure to e-cig. Further study using vapor exposure system will be warranted to confirm the results from this study.
Our findings reveal that e-cig smoke solution affect biological pathways related to ribosomal proteins and protein synthesis in NHBE cells. We further showed that exposure to e-cig smoke solution down-regulated expression of ribosomal proteins-encoding genes and decreased new protein synthesis from the cells. Because of the associations of increasing popularity of e-cig use among people and associations of e-cig usage and severe respiratory diseases, further mechanistic studies are warranted to evaluate the effects of e-cig on airway epithelium.
Methods
Preparation and analysis of e-cig smoke solution
The e-cig solution was prepared by drawing the e-cig downstream smoke through a 25 mm impinger filled with ultrapure water (Fig. 1A). The cigarettes were puffed using a TE-2B smoking machine (Teague Enterprises, Woodland, Ca). Flow rate through each cigarette was set at the minimum to ignite the e-cig. (1 to 1.5 lpm). Puffs were two seconds in duration spaced one minute apart. The e-cig cartridges using rechargeable batteries were used until no visible smoke could be seen entering the system after each puff. Twenty e-cig cartridges (Blu vivid vanilla, 2.8%) were used to make the solution. The solution was generated in a darken room over a five-date period at a rate of 4 e-cigarette cartridges per day. Ultrapure water was added to the solution each day to return the volume to 10 ml to account for solution lost to evaporation. At the initial analysis of the e-cig smoke solution detected by gas chromatography with flame ionization detector (GC-FID), the concentration of diacetyl was 89 ppm (ALS Laboratories, Cincinnati, OH). Based on this concentration, the solution was diluted into cell culture medium to treat NHBE cells in vitro. To analyze additional flavoring chemicals and dicarbonyls in the e-cig smoke solution, we performed headspace(HS)—solid-phase microextraction (SPME)—GC/MS for flavorings and gas chromatography/mass Spectrometry (GC/MS) for Di-carboynls (Labstat International, Kitchener, Ontario). The detailed methods on GC/MS are provided in Supplementary Materials.
Cell culture and exposure
Primary normal Human Bronchial Epithelial (NHBE) cells were isolated from left over tissues after lung transplantation following the approved protocol51 and were received from Marsico Lung Institute/Cystic Fibrosis Center at the University of North Carolina, Chapel Hill (Chapel Hill, NC). Then NHBE cells were cultured as previously described29,52,53. Cells at passage 2 were transferred to microporous polyester inserts (0.4 um pore size, Transwell-Clear; Corning Costar, Corning, NY) and fed with a 1:1 mixture of BEBM and Dulbecco’s Modification of Eagle’s Media (Mediatech, Herndon, VA). Media was applied apically and basally until the cells were confluent and then basally after an air–liquid interface (ALI) was established. Cells were cultured at ALI for 14 days to promote relatively stable expression of goblet and ciliated cells before exposure to e-cig smoke solution. Mature, well-differentiated monolayers of cells were then exposed to control (ultrapure water dissolved in culture medium, water/medium: 2% v/v) or e-cig smoke solution (containing 2 ppm diacetyl, water/medium: 2% v/v) on the apical side for 6 or 24 h (n = 3 subjects, each treatment was performed in duplicate). After incubation, LDH release in the culture medium was measured for cytotoxicity assay. Total RNAs were isolated from the cells using miRNeasy kit (Qiagen) for RNA -Seq analysis.
RNA-seq library preparation and sequencing
Libraries for sequencing were prepared as previously described29. Polyadenylated mRNAs were selected from total RNA samples using oligo-dT-conjugated magnetic beads on an Apollo324 automated workstation (PrepX PolyA mRNA isolation kit, Takara Bio USA). Entire poly-adenylated RNA samples were immediately converted into stranded Illumina sequencing libraries using 200 bp fragmentation and sequential adapter addition on an Apollo324 automated workstation following manufacturer’s specifications (PrepX RNA-Seq for Illumina Library kit, Takara Bio USA). Libraries were enriched and indexed using 12 cycles of amplification (LongAmp Taq 2 × MasterMix, New England BioLabs Inc.) with PCR primers which include a 6 bp index sequence to allow for multiplexing (custom oligo order from Integrated DNA Technologies). Excess PCR reagents were removed using magnetic bead-based cleanup on an Apollo324 automated workstation (PCR Clean DX beads, Aline Biosciences). Resulting libraries were assessed using a 2200 TapeStation (Agilent Technologies) and quantified by QPCR (Kapa Biosystems). Libraries were pooled and sequenced on one lane of a HiSeq 2500 high output v3 flow cell using single end, 50 bp reads (Illumina).
RNA-seq data analysis
RNA-Seq data was analyzed as previously described29. Taffeta scripts (https://github.com/blancahimes/taffeta) were used to analyze the RNA-Seq data, which included trimming of adapters using trimmomatic (v.0.32)54 and using FastQC55 (v.0.11.2) to obtain overall QC metrics. Trimmed reads for each sample were aligned with STAR (v. 2.5.2b) to the reference homo sapiens build 38 UCSC file (hg38) genome obtained from the Illumina, Inc. iGenomes resource56. Additional QC parameters were obtained to assess whether reads were appropriately mapped. Bamtools (v.2.3.0)57 was used to count/summarize the number of mapped reads, including junction spanning reads. The Picard Tools (v.1.96; http://picard.sourceforge.net) RnaSeqMetrics function was used to compute the number of bases assigned to various classes of RNA, according to the hg38 refFlat file available as a UCSC Genome Table. For each sample, HTSeq (v.0.6.1) was used to quantify genes based on reads that mapped to the provided hg38 reference files58. The DESeq2 R package (v. 1.12.4) was used to measure significance of differentially expressed genes between the exposed (N = 4) and control (N = 4) samples and create plots of the results59. The reported adjusted p-values are false-discovery rate corrected to 5% according to the procedure in DESeq2 that accounts for the large number of comparisons made. An adjusted p-value < 0.05 was considered significant. The NIH Database for Annotation, Visualization and Integrated Discovery (DAVID) was used to perform gene functional annotation clustering using Homo Sapiens as background, and default options and annotation categories (Disease: OMIM_DISEASE; Functional Categories: COG_ONTOLOGY, SP_PIR_KEYWORDS, UP_SEQ_FEATURE; Gene_Ontology: GOTERM_BP_FAT, GOTERM_CC_FAT, GOTERM_MF_FAT; Pathway: BBID, BIOCARTA, KEGG_PATHWAY; Protein_Domains: INTERPRO, PIR_SUPERFAMILY, SMART)60.
qPT-PCR
RNA was reverse transcribed using iScript cDNA Synthesis kit (Biorad). For rRNA measurement, cDNA was synthesized using random hexamers and SuperScript III Reverse Transcriptase (Invitrogen). The resulting cDNA was amplified using 2 × SYBR mix (Qiagen) and 1 μM of each primer in a StepOne Plus Thermocycler (Applied Biosystems) in Quantitative Reverse Transcriptase Polymerase Chain Reaction (qRT-PCR). Melt curves were checked for single-length amplification products. Fold changes were calculated using the 2−ΔΔCt method. GAPDH is the housekeeping gene used for normalization for protein coding genes. B2M was an internal control for rRNAs whose expression is not dependent on Polymerase I activity61. All primer sequences used in this study are listed in Supplemental Table S2.
Western blot
Whole-cell lysates were prepared in Radioimmunoprecipitation Assay (RIPA) buffer supplemented with Protease and Phosphatase Inhibitor Cocktails (Roche, Indianapolis, Ind). Cleared lysates were heated at 70 °C for 10 min and run in SDS-PAGE gel under reducing conditions. Primary antibodies included anti RPS14 (Adcam), anti-RPL30 (Invitrogen), and anti-GAPDH (Santa Cruz Biotechnology).
Protein synthesis assay
To test whether exposure to e-cig smoke solution impacts cells’ ability to produce new proteins, we performed protein synthesis assay using a kit from Cayman Chemical (Ann Arbor, MI, USA). This kit uses O-Propargyl-puromycin (OPP) as a probe for labeling translating polypeptide chains. OPP-labeled proteins are detected by 5-Fluorescein (FAM)-Azide. Primary NHBE cells were plated in a black, clear-bottomed 96-well plate at a density of 10,000 cells/well. After 24 h, cells were exposed to control (ultrapure water) or e-cig smoke solution (containing 1, 2, and 5 ppm of diacetyl) for 24 or 48 h. Protein synthesis was quantified following the manufacture’s protocol. Cycloheximide (500 μg/ml) was included as a positive control.
Immunofluorescence staining
Immunofluorescence staining on ALI-cultured NHBE cells was performed as described previously with some modifications29,62. First, cells were fixed using 4% paraformaldehyde. Then, cells were blocked with PBS supplemented with 5% BSA and 0.2% Triton X-100 for 1 h at room temperature. Primary antibody incubation was performed overnight at 4 °C in PBS supplemented with 1% BSA and 0.2%Triton X-100 using anti-β-tubulin IV (Sigma) or anti-MUC5AC (Thermo) at 1:100 dilution. Secondary antibodies conjugated with Alexa-fluor 488 (Life Technologies) were used at 1:100 dilution. 4′-6-Diamidino-2-phenylindole, dihydrochloride was used to label the nuclear DNA and samples were mounted with Vectashield antifade mounting medium (Vector Labs, Burlingame, Calif). Confocal images were taken using Zeiss AxioObserver Z1 or Leica STP8000 and processed using ImageJ.
Statistical analysis
Statistical analysis for in vitro studies was performed with GraphPad Prizm version 6 (La Jolla, CA 92037, USA). Data were analyzed by either t-test or one-way analysis of variance (ANOVA). If significant effects were detected, the ANOVA was followed by Tukey post-hoc comparison of means. A p < 0.05 was considered statistically different. Data were expressed as means ± SEM.
References
Wang, T. W. et al. Tobacco product use among adults: United States, 2017. MMWR Morb. Mortal Wkly. Rep. 67, 1225–1232. https://doi.org/10.15585/mmwr.mm6744a2 (2018).
Cullen, K. A. et al. Notes from the field: Use of electronic cigarettes and any tobacco product among middle and high school students: United States, 2011–2018. MMWR Morb. Mortal. Wkly. Rep. 67, 1276–1277. https://doi.org/10.15585/mmwr.mm6745a5 (2018).
Henry, T. S., Kanne, J. P. & Kligerman, S. J. Imaging of vaping-associated lung disease. N. Engl. J. Med. https://doi.org/10.1056/NEJMc1911995 (2019).
Layden, J. E. et al. Pulmonary illness related to e-cigarette use in Illinois and Wisconsin: Preliminary report. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa1911614 (2019).
Maddock, S. D. et al. Pulmonary lipid-laden macrophages and vaping. N. Engl. J. Med. https://doi.org/10.1056/NEJMc1912038 (2019).
CDC. Outbreak of Lung Injury Associated with E-Cigarette Use, or Vaping (2019).
Bekki, K. et al. Carbonyl compounds generated from electronic cigarettes. Int. J. Environ. Res. Public Health 11, 11192–11200. https://doi.org/10.3390/ijerph111111192 (2014).
Callahan-Lyon, P. Electronic cigarettes: Human health effects. Tob. Control 23(Suppl 2), 36–40. https://doi.org/10.1136/tobaccocontrol-2013-051470 (2014).
Cheng, T. Chemical evaluation of electronic cigarettes. Tob. Control 23(Suppl 2), 11–17. https://doi.org/10.1136/tobaccocontrol-2013-051482 (2014).
Goniewicz, M. L. et al. Levels of selected carcinogens and toxicants in vapour from electronic cigarettes. Tob. Control 23, 133–139. https://doi.org/10.1136/tobaccocontrol-2012-050859 (2014).
Hutzler, C. et al. Chemical hazards present in liquids and vapors of electronic cigarettes. Arch. Toxicol. 88, 1295–1308. https://doi.org/10.1007/s00204-014-1294-7 (2014).
Jensen, R. P., Luo, W., Pankow, J. F., Strongin, R. M. & Peyton, D. H. Hidden formaldehyde in e-cigarette aerosols. N. Engl. J. Med. 372, 392–394. https://doi.org/10.1056/NEJMc1413069 (2015).
Orr, M. S. Electronic cigarettes in the USA: A summary of available toxicology data and suggestions for the future. Tob. Control 23(Suppl 2), 18–22. https://doi.org/10.1136/tobaccocontrol-2013-051474 (2014).
Pellegrino, R. M. et al. Electronic cigarettes: An evaluation of exposure to chemicals and fine particulate matter (PM). Ann Ig 24, 279–288 (2012).
Uchiyama, S., Ohta, K., Inaba, Y. & Kunugita, N. Determination of carbonyl compounds generated from the E-cigarette using coupled silica cartridges impregnated with hydroquinone and 2,4-dinitrophenylhydrazine, followed by high-performance liquid chromatography. Anal. Sci. 29, 1219–1222 (2013).
Williams, M., To, A., Bozhilov, K. & Talbot, P. Strategies to reduce tin and other metals in electronic cigarette aerosol. PLoS ONE 10, e0138933. https://doi.org/10.1371/journal.pone.0138933 (2015).
Allen, J. G. et al. Flavoring chemicals in e-cigarettes: Diacetyl, 2,3-pentanedione, and acetoin in a sample of 51 products, including fruit-, candy-, and cocktail-flavored e-cigarettes. Environ. Health Perspect. 124, 733–739. https://doi.org/10.1289/ehp.1510185 (2016).
Lee, M. S., Allen, J. G. & Christiani, D. C. Endotoxin and [formula: see text] contamination in electronic cigarette products sold in the United States. Environ. Health Perspect. 127, 47008. https://doi.org/10.1289/EHP3469 (2019).
Christiani, D. C. Vaping-induced lung injury. N. Engl. J. Med. https://doi.org/10.1056/NEJMe1912032 (2019).
Antherieu, S. et al. Comparison of cellular and transcriptomic effects between electronic cigarette vapor and cigarette smoke in human bronchial epithelial cells. Toxicol. In Vitro 45, 417–425. https://doi.org/10.1016/j.tiv.2016.12.015 (2017).
Czekala, L. et al. Toxicological comparison of cigarette smoke and e-cigarette aerosol using a 3D in vitro human respiratory model. Regul. Toxicol. Pharmacol. 103, 314–324. https://doi.org/10.1016/j.yrtph.2019.01.036 (2019).
Iskandar, A. R. et al. Application of a multi-layer systems toxicology framework for in vitro assessment of the biological effects of Classic Tobacco e-liquid and its corresponding aerosol using an e-cigarette device with MESH technology. Arch. Toxicol. 93, 3229–3247. https://doi.org/10.1007/s00204-019-02565-9 (2019).
Iskandar, A. R. et al. A framework for in vitro systems toxicology assessment of e-liquids. Toxicol. Mech. Methods 26, 389–413. https://doi.org/10.3109/15376516.2016.1170251 (2016).
Ghosh, A. et al. Chronic e-cigarette exposure alters the human bronchial epithelial proteome. Am. J. Respir. Crit. Care Med. 198, 67–76. https://doi.org/10.1164/rccm.201710-2033OC (2018).
Clapp, P. W. et al. Cinnamaldehyde in flavored e-cigarette liquids temporarily suppresses bronchial epithelial cell ciliary motility by dysregulation of mitochondrial function. Am. J. Physiol. Lung Cell Mol. Physiol. 316, L470–L486. https://doi.org/10.1152/ajplung.00304.2018 (2019).
Rowell, T. R. et al. Flavored e-cigarette liquids reduce proliferation and viability in the CALU3 airway epithelial cell line. Am. J. Physiol. Lung Cell Mol. Physiol. 313, L52–L66. https://doi.org/10.1152/ajplung.00392.2016 (2017).
Pezzulo, A. A. et al. The air-liquid interface and use of primary cell cultures are important to recapitulate the transcriptional profile of in vivo airway epithelia. Am. J. Physiol. Lung Cell Mol. Physiol. 300, L25-31. https://doi.org/10.1152/ajplung.00256.2010 (2011).
Klager, S. et al. Flavoring chemicals and aldehydes in e-cigarette emissions. Environ. Sci. Technol. 51, 10806–10813. https://doi.org/10.1021/acs.est.7b02205 (2017).
Park, H. R. et al. Transcriptomic response of primary human airway epithelial cells to flavoring chemicals in electronic cigarettes. Sci. Rep. 9, 1400. https://doi.org/10.1038/s41598-018-37913-9 (2019).
Aubert, M., O’Donohue, M. F., Lebaron, S. & Gleizes, P. E. Pre-Ribosomal RNA processing in human cells: From mechanisms to congenital diseases. Biomolecules https://doi.org/10.3390/biom8040123 (2018).
Mullineux, S. T. & Lafontaine, D. L. Mapping the cleavage sites on mammalian pre-rRNAs: Where do we stand?. Biochimie 94, 1521–1532. https://doi.org/10.1016/j.biochi.2012.02.001 (2012).
Peculis, B. A. Ribosome biogenesis: Ribosomal RNA synthesis as a package deal. Curr. Biol. 12, R623-624. https://doi.org/10.1016/s0960-9822(02)01135-1 (2002).
Kreiss, K. et al. Clinical bronchiolitis obliterans in workers at a microwave-popcorn plant. N. Engl. J. Med. 347, 330–338. https://doi.org/10.1056/NEJMoa020300 (2002).
Hubbs, A. F. et al. Flavorings-related lung disease: A brief review and new mechanistic data. Toxicol. Pathol. 47, 1012–1026. https://doi.org/10.1177/0192623319879906 (2019).
Albert, B. et al. A ribosome assembly stress response regulates transcription to maintain proteome homeostasis. Elife https://doi.org/10.7554/eLife.45002 (2019).
Zhou, X., Liao, W. J., Liao, J. M., Liao, P. & Lu, H. Ribosomal proteins: Functions beyond the ribosome. J. Mol. Cell Biol. 7, 92–104. https://doi.org/10.1093/jmcb/mjv014 (2015).
Golomb, L., Volarevic, S. & Oren, M. p53 and ribosome biogenesis stress: The essentials. FEBS Lett. 588, 2571–2579 (2014).
Donati, G. et al. The balance between rRNA and ribosomal protein synthesis up- and downregulates the tumour suppressor p53 in mammalian cells. Oncogene 30, 3274–3288. https://doi.org/10.1038/onc.2011.48 (2011).
Tommasi, S. C. et al. Deregulation of biologically significant genes and associated molecular pathways in the oral epithelium of electronic cigarette users. Int. J. Mol. Sci. 20, 738 (2019).
Pierre, P. Immunity and the regulation of protein synthesis: Surprising connections. Curr. Opin. Immunol. 21, 70–77. https://doi.org/10.1016/j.coi.2009.03.003 (2009).
Mazumder, B., Li, X. & Barik, S. Translation control: A multifaceted regulator of inflammatory response. J. Immunol. 184, 3311–3319. https://doi.org/10.4049/jimmunol.0903778 (2010).
Vyleta, M. L., Wong, J. & Magun, B. E. Suppression of ribosomal function triggers innate immune signaling through activation of the NLRP3 inflammasome. PLoS ONE 7, e36044. https://doi.org/10.1371/journal.pone.0036044 (2012).
Sundar, I. K. J., Romanos, G. E. & Rahman, I. E-cigarettes and flavorings induce inflammatory and prosenescence responses in oral epithelial cells and periodontal fibroblasts. Oncotarget 7, 77196–77204 (2016).
Estabrook, R. W. A passion for P450s (rememberances of the early history of research on cytochrome P450). Drug. Metab. Dispos. 31, 1461–1473. https://doi.org/10.1124/dmd.31.12.1461 (2003).
Zuikova, E. I. C., Bailey, A. & Marczylo, T. European Network for Smoking and Tobacco Prevention (Springer, 2020).
Canistro, D. et al. E-cigarettes induce toxicological effects that can raise the cancer risk. Sci. Rep. 7, 2028. https://doi.org/10.1038/s41598-017-02317-8 (2017).
Lee, H. W. et al. E-cigarette smoke damages DNA and reduces repair activity in mouse lung, heart, and bladder as well as in human lung and bladder cells. Proc. Natl. Acad. Sci. USA 115, E1560–E1569. https://doi.org/10.1073/pnas.1718185115 (2018).
Tang, M. S. et al. Electronic-cigarette smoke induces lung adenocarcinoma and bladder urothelial hyperplasia in mice. Proc. Natl. Acad. Sci. USA 116, 21727–21731. https://doi.org/10.1073/pnas.1911321116 (2019).
Yang, M. et al. Down-regulation of ribosomal protein L22 in non-small cell lung cancer. Med. Oncol. 30, 646. https://doi.org/10.1007/s12032-013-0646-0 (2013).
Wang, W. et al. Ribosomal proteins and human diseases: Pathogenesis, molecular mechanisms, and therapeutic implications. Med. Res. Rev. 35, 225–285. https://doi.org/10.1002/med.21327 (2015).
Fulcher, M. L. & Randell, S. H. Human nasal and tracheo-bronchial respiratory epithelial cell culture. Methods Mol. Biol. 945, 109–121. https://doi.org/10.1007/978-1-62703-125-7_8 (2013).
Park, J. A. & Tschumperlin, D. J. Chronic intermittent mechanical stress increases MUC5AC protein expression. Am. J. Respir. Cell Mol. Biol. 41, 459–466. https://doi.org/10.1165/rcmb.2008-0195OC (2009).
Randell, S. H. F. Human nasal and tracheo-bronchial respiratory epithelial cell culture. Epithelial Cell Cult. Protoc. 1, 109–121 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics https://doi.org/10.1093/bioinformatics/btu170 (2014).
Andrews, S. FastQC A Quality Control tool for High Throughput Sequence Data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Barnett, D. W., Garrison, E. K., Quinlan, A. R., Stromberg, M. P. & Marth, G. T. BamTools: A C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27, 1691–1692. https://doi.org/10.1093/bioinformatics/btr174 (2011).
Anders, S., Pyl, P. T. & Huber, W. HTSeq: A Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169. https://doi.org/10.1093/bioinformatics/btu638 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
da Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57. https://doi.org/10.1038/nprot.2008.211 (2009).
Zhang, D. et al. Stem cell and neurogenic gene-expression profiles link prostate basal cells to aggressive prostate cancer. Nat. Commun. 7, 10798. https://doi.org/10.1038/ncomms10798 (2016).
Cao, X. et al. Tight junction disruption by cadmium in an in vitro human airway tissue model. Respir. Res. 16, 30. https://doi.org/10.1186/s12931-015-0191-9 (2015).
Acknowledgements
This study was funded by the National Institutes of Health (NIH)/National Institute of Environmental Health Science (NIEHS) R01 Grant ES029097 and the Harvard NIEHS Center Grant (P30ES000002).
Author information
Authors and Affiliations
Contributions
Q.L. and J.A. conceived the project. H.-R.P. and Q.L. designed the experiments. H.-R.P., J.V., M.O., C.W., R.A.P., and G.W. performed the experiments. M.S., B.E.H., and H.-R.P. analyzed the data. H.-R.P. visualized the data. H.-R.P., J.V., J.A., Q.L. wrote the paper. All authors reviewed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Park, HR., Vallarino, J., O’Sullivan, M. et al. Electronic cigarette smoke reduces ribosomal protein gene expression to impair protein synthesis in primary human airway epithelial cells. Sci Rep 11, 17517 (2021). https://doi.org/10.1038/s41598-021-97013-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-97013-z
This article is cited by
-
Single-cell transcriptomics reveals e-cigarette vapor-induced airway epithelial remodeling and injury
Respiratory Research (2024)
-
Integrated mRNA- and miRNA-sequencing analyses unveil the underlying mechanism of tobacco pollutant-induced developmental toxicity in zebrafish embryos
Journal of Translational Medicine (2024)
-
Electronic cigarette liquids impair metabolic cooperation and alter proteomic profiles in V79 cells
Respiratory Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.