Abstract
Retinitis Pigmentosa (RP) is a blinding disease that arises from loss of rods and subsequently cones. The P23H rhodopsin knock-in (P23H-KI) mouse develops retinal degeneration that mirrors RP phenotype in patients carrying the orthologous variant. Previously, we found that the P23H rhodopsin protein was degraded in P23H-KI retinas, and the Unfolded Protein Response (UPR) promoted P23H rhodopsin degradation in heterologous cells in vitro. Here, we investigated the role of a UPR regulator gene, activating transcription factor 6 (Atf6), in rhodopsin protein homeostasis in heterozygous P23H rhodopsin (Rho+/P23H) mice. Significantly increased rhodopsin protein levels were found in Atf6−/−Rho+/P23H retinas compared to Atf6+/−Rho+/P23H retinas at early ages (~ P12), while rhodopsin mRNA levels were not different. The IRE1 pathway of the UPR was hyper-activated in young Atf6−/−Rho+/P23H retinas, and photoreceptor layer thickness was unchanged at this early age in Rho+/P23H mice lacking Atf6. By contrast, older Atf6−/−Rho+/P23H mice developed significantly increased retinal degeneration in comparison to Atf6+/−Rho+/P23H mice in all retinal layers, accompanied by reduced rhodopsin protein levels. Our findings demonstrate that Atf6 is required for efficient clearance of rhodopsin protein in rod photoreceptors expressing P23H rhodopsin, and that loss of Atf6 ultimately accelerates retinal degeneration in P23H-KI mice.
Similar content being viewed by others
Introduction
Retinitis pigmentosa (RP) is a group of retinal degenerative diseases that leads to irreversible blindness with a worldwide prevalence of 1:40001,2. RP causes progressive retinal degeneration that first results in the loss of rod photoreceptors which then results in the loss of cone photoreceptors, and gives rise to initial night blindness and loss of peripheral vision. Hundreds of gene mutations have been found to cause heritable forms of RP (RetNet, Retinal Information Network, at https://sph.uth.edu/retnet/disease.htm). Mutations in the rhodopsin gene are a common cause of autosomal dominant RP (adRP). More than 150 distinct rhodopsin mutations have been identified in adRP (RetNet[Retinal Information Network- www.sph.uth.tmc.edu/Retnet]2,3,4. Most mutations introduce missense changes throughout the rhodopsin protein coding region; the most common mutation in the USA is the substitution of proline to histidine at codon 23 (P23H) in the extracellular N-terminal domain, accounting for 15–20% of all adRP cases in the USA5,6,7.
The endoplasmic reticulum (ER) is vital for membrane protein packaging, secretion, and folding, and, consequently, protein misfolding in the ER can disrupt ER function and lead to “ER stress”. In response to ER stress, cells activate the Unfolded Protein Response (UPR), a cellular homeostatic mechanism that reduces ER stress by promoting the degradation of misfolded proteins and slowing new protein synthesis8. The UPR is controlled by 3 ER resident transmembrane proteins, inositol-requiring enzyme 1 (IRE1), activating transcription factor 6 (ATF6), and protein kinase RNA-like endoplasmic reticulum kinase (PERK). In response to ER stress, IRE1 activates its kinase and RNase functions, initiating the nonconventional splicing of X-box binding protein 1 (XBP-1) mRNA9,10,11. Spliced XBP-1 (XBP-1s) mRNA encodes a transcription activator that induces expression of ER chaperones and ER-associated degradation (ERAD) components that remove and degrade misfolded proteins via the ubiquitin–proteasome system12,13. The PERK arm of the UPR regulates protein synthesis. Upon ER stress, PERK oligomerizes via its cytoplasmic domain, which leads to phosphorylation of eukaryotic initiation factor 2α (eIF2α) and, as a result, attenuation of protein translation8,14,15,16. Under prolonged ER stress, PERK promotes cell death by activating stress-responsive transcription factors such as activating transcription factor 4 (ATF4) and CCAAT/enhancer-binding protein homologous protein (CHOP)17,18. ATF6 undergoes proteolytic processing in response to ER stress, releasing its cytosolic bZIP-containing transcription factor domain (ATF6f), which then travels to the nucleus to upregulate genes involved in ER protein folding and ERAD (that significantly overlap with the transcriptional targets of XBP-1s)19,20,21. Thus, activation of the UPR ultimately promotes cell viability and homeostasis by reducing misfolded protein levels.
Here, we used the P23H-KI mouse model to evaluate the UPR’s function in rod photoreceptors in the eye and in the pathogenesis/progression of retinal degeneration arising from misfolded rhodopsin proteins22,23. Within two weeks of age, photoreceptor and retinal degeneration begin in the Rho+/P23H mice22,23. At post-natal day 30 (P30) and P60, degeneration continues with outer nuclear layer (ONL) thinning, loss of rhodopsin protein levels, and poorer ERG responses by a-wave and b-wave analysis22,23,24. In addition, Rho+/P23H mice show progressive shortening of the rod outer segment and rod inner segment at P30 and P6022,23,24. In healthy retinas, rhodopsin is translated at the ER membrane and, when correctly folded, rhodopsin is exported to the outer segment of rod photoreceptor cells to initiate phototransduction in response to light. In contrast, biochemical and cellular studies reveal that P23H rhodopsin is misfolded, retained in the ER, and aggregates; as a result, P23H rhodopsin may be a source of ER stress and cause activation of the UPR6,25. The role of P23H rhodopsin as an ER stress inducer in rod photoreceptors is supported by the activation of the IRE1 pathway in the Rho+/P23H mice26, which was found using an IRE1-reporter ER stress activated indicator (ERAI) transgenic mouse crossed with P23H-KI mice. The Rho+/P23H mice also showed significant increase of XBP-1s mRNA and XBP-1s targets, such as DNA damage inducible transcript 3 (Ddit3), SEC24 homolog d (Sec24d), and homocysteine inducible ER protein with ubiquitin like domain 1 (Herpud1), in Rho+/P23H mice at P30, P60, P90, and P12026. These findings demonstrate increased IRE1 activity in rods expressing the P23H misfolded rhodopsin. We therefore proposed that misfolded rhodopsin generated by the P23H mutation activates the UPR, and that in turn may alleviate ER stress caused by P23H rhodopsin through protein clearance mechanisms such as ERAD.
Previously, we demonstrated that chemical-genetic activation of either IRE1 (to produce XBP-1s) or ATF6f expression promoted P23H rhodopsin protein degradation when expressed in heterologous HEK293 cells, while sparing wild-type rhodopsin protein22,27. By contrast, PERK signaling negatively impacted both misfolded and wild-type rhodopsin protein levels27. These findings suggested that IRE1, XBP-1s, and ATF6 are important for removing P23H rhodopsin from photoreceptors and, thereby, influence rod photoreceptor survival. Ire1−/− and Xbp-1s−/− mice develop widespread developmental defects and undergo early embryonic death28,29. In contrast, Atf6−/− mice are viable and have normal retinal morphology and function at birth and up to 3-month-old, but aged Atf6−/− mice develop rod and cone dysfunction and retinal degeneration (18-month-old)30. Here, we test the role of Atf6 in regulating rhodopsin protein levels and influencing retinal degeneration in P23H-KI mice by crossing Atf6−/− with P23H-KI animals23.
Results
Loss of Atf6 leads to impaired clearance of rhodopsin protein and hyperactivation of the IRE1-XBP-1s signaling pathway in early age Rho +/P23H mice
Previously, we found that P23H rhodopsin protein was rapidly degraded in photoreceptors of P23H-KI mice at early age (P15)22. We also found that chemical-genetic activation of Atf6 signaling pathways promoted P23H rhodopsin protein degradation in heterologous HEK293 cells, while sparing wild-type rhodopsin protein22,27. To investigate if Atf6 is important for P23H rhodopsin protein degradation in photoreceptors, we examined retinas of Rho+/P23H mice bred with Atf6−/− mice.
First, we examined steady-state levels of rhodopsin in retinal protein lysates collected from Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H at P12, an age before morphological defects in photoreceptors or retinal degeneration emerges in Rho+/P23H mice22. We saw significantly increased rhodopsin protein levels in Atf6−/− Rho+/P23H retinas compared to Atf6+/− Rho+/P23H retinas (191% increase, P = 0.04, Fig. 1a,b). This finding demonstrates that Atf6 is necessary for rhodopsin protein homeostasis in Rho+/P23H mice retinas. Rhodopsin mRNA levels were not significantly different between Atf6−/− Rho+/P23H and Atf6+/− Rho+/P23H retinas (P = 0.5, Fig. 1b) indicating that the differences seen in rhodopsin protein levels were not due to transcriptional differences. These findings support a role for Atf6 in maintaining rhodopsin protein homeostasis in the retina via post-transcriptional mechanisms such as regulating its protein degradation.
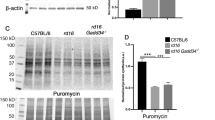
Elevated rhodopsin protein levels and increased IRE1 signaling activity in Rho + /P23H mice in the absence of Atf6 in P12. (a) Retinas were collected at P12 and processed. Total rhodopsin, IRE1a, and BiP/Grp78 from Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas were detected by immunoblotting and quantified. HSP90 served as loading control. (b) Total rhodopsin protein levels and mRNA levels were quantified in the retinas of Atf6+/−Rho+/P23H (n = 3) and Atf6−/−Rho+/P23H (n = 3) mice. (c) IRE1a and BiP/Grp78 protein levels were quantified in the retinas of Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice normalized to HSP90. Full-length blots are presented in Supplementary Fig. S2. (d) Xbp-1s and Syvn1 mRNA levels in Atf6−/−Rho+/P23H retinas (n = 3) were measured by real-time quantitative PCR and normalized to mRNA levels in Atf6+/−Rho+/P23H retinas (n = 3). (e) Chop mRNA levels in both genotypes (n = 3) were measured by real-time quantitative PCR. Data are shown as mean ± SEM, three sets of independent experiments were performed. Data were analyzed by Student’s t-test and showed significance at *P < 0.05.
Previously, we found that the IRE1-XBP-1 pathway was activated in Rho+/P23H mice retinas and promoted P23H rhodopsin protein degradation in vitro22,26. To determine how loss of Atf6 affected IRE1-XBP-1 in Rho+/P23H mice, we examined biochemical and molecular markers of IRE1-XBP-1 activity in retinal lysates of P12 Atf6+/− Rho+/P23H and Atf6−/−Rho+/P23H mice. Interestingly, Atf6−/−Rho+/P23H retinas showed significantly increased IRE1a and binding immunoglobulin protein/78-kDa glucose-regulated protein (BiP/Grp78) chaperone protein expression compared to Atf6+/−Rho+/P23H retinas (P = 0.04, Fig. 1c). To further investigate if IRE1 signaling pathway was increased in these retinas, we measured the mRNA levels of Xbp-1s (i.e. downstream target of IRE1 pathway) and ERAD-associated E3 ubiquitin-protein ligase HRD1/Synoviolin 1 (Hrd1/Syvn1) mRNA levels, an ER-associated protein degradation gene transcriptionally regulated by XBP-1s12. We found significant increase in the mRNA levels of Xbp-1s (P = 0.03) and an Xbp-1s target gene, Syvn1, (P = 0.04) in Atf6−/−Rho+/P23H retinas compared to Atf6+/−Rho+/P23H retinas (Fig. 1d). These findings provide evidence that IRE1 is hyper-activated in Rho+/P23H retina in the absence of Atf6 at this age.
By contrast to the changes observed in IRE1 pathway markers, Chop mRNA levels showed no significant differences between Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice (Fig. 1e) consistent with prior studies showing no induction of CHOP in mice expressing P23H rhodopsin31. In summary, we find that loss of Atf6 in Rho+/P23H mice leads to rhodopsin protein build-up in the retina at an early age, concomitant with increased activation of the IRE1 pathway.
No gross changes in retinal histology in the absence of Atf6 in young Rho +/P23H retinas
Overexpression of rhodopsin causes photoreceptor cell death and induces retinal degeneration in transgenic animals expressing wild-type rhodopsin or P23H rhodopsin32,33,34. With that in mind, we asked if the increased steady-state rhodopsin protein levels in Rho+/P23H mice lacking Atf6 corresponded with photoreceptor cell loss (Fig. 1a). We performed histologic studies to see if photoreceptors or retinal lamination was impacted. A previously published paper has shown that Rho+/P23H retinas have scattered pyknotic nuclei by P1522, with Rho+/P23H mice exhibiting a slight reduction in the ONL thickness. Furthermore, the outer segments and inner segments of rod photoreceptors were shorter compared to the wild-type mice. In both Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice, we observed scattered and disorganized nuclei in the ONL ofP15 mice. In addition, the thickness of the ONL, outer plexiform layer (OPL), inner nuclear (INL), and inner plexiform layer (IPL) appeared similar (Fig. 2a,b) in retinas (P > 0.05, two-way ANOVA analysis). Therefore, the ~ 2 × increase in rhodopsin protein levels found in the absence of Atf6 did not lead to detectable changes in the thickness of photoreceptor ONL or overall retinal anatomy at this age. These findings demonstrate that Rho+/P23H mice can tolerate increased amounts of rhodopsin protein when Atf6 is lost, at least at this young age.
Quantitative spider plot analysis of P15 Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (a) Light micrographs taken from vertical cryostat sections processed for H&E staining in P15 Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (b) The retinal layers of H&E-stained retinal sections through the optic nerve (0) were measured at 8 locations around the retina, four each in the dorsal and ventral hemispheres. Retinal layer thickness between the two genotypes was not significantly different (n = 3–5, Data represents mean ± SEM, Two-way ANOVA, P > 0.5). ONL outer nuclear layer; OPL outer plexiform layer; INL inner nuclear layer; IPL inner plexiform layer. Scale bar = 50 μm.
Rhodopsin protein levels stabilize in intermediate age of Rho +/P23H retina in the absence of Atf6
Next, we examined the retina in older Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice. At P30, the thickness of retinal layers between Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H showed no significant difference (P > 0.05, two-way ANOVA analysis, Fig. 3a,b). In both Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice, we observed scattered and disorganized nuclei in the ONL. To investigate if absence of Atf6 altered rhodopsin protein levels in P23H retina at this age, we performed immunoblot analyses on the retinas of Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H P30 mice. In contrast to the increase in rhodopsin protein levels observed in younger mice (P12), retinal protein lysates of Atf6−/−Rho+/P23H showed no significant difference in rhodopsin, BiP/Grp78, and IRE1a protein expression compared to Atf6+/−Rho+/P23H at this age (Fig. 3c,d). We also found no significant increase in the mRNA levels of Xbp-1s in Atf6−/−Rho+/P23H retinas compared to Atf6+/−Rho+/P23H retinas (P > 0.05, data not shown). These results show that loss of Atf6 does not alter rhodopsin protein levels or retinal anatomy in Rho+/P23H mice by this age. We propose that the equalization of rhodopsin protein levels in Atf6−/− Rho+/P23H mice to levels seen in Atf6+/−Rho+/P23H mice may be a result of the hyperactivation of IRE1-XBP-1s signaling observed in younger mice.
Rhodopsin protein levels in Rho + /P23H in the absence of Atf6 at P30. (a) Light micrographs taken from vertical cryostat sections processed for H&E staining in P30 Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (b) The retinal layers of H&E-stained retinal sections through the optic nerve (0) were measured at 8 locations around the retina, four each in the dorsal and ventral hemispheres. Quantification of retinal layer thickness between two genotypes showed no significant difference (n = 3–5, Data represents mean ± SEM, Two-way ANOVA, p > 0.5). ONL outer nuclear layer; OPL outer plexiform layer; INL inner nuclear layer; IPL inner plexiform layer. Scale bar = 50 μm. (c) Total rhodopsin, BiP/Grp78, and IRE1a from Atf6+/−Rho+/P23H (n = 5) and Atf6−/−Rho+/P23H (n = 4) retinas were detected by immunoblotting and quantified. HSP90 served as the loading control. Full-length blots are presented in Supplementary Fig. S3. (d) Rhodopsin, BiP/Grp78, and IRE1a protein levels were quantified and normalized to HSP90 in Atf6−/−Rho+/P23H retinas compared to Atf6+/−Rho+/P23H retinas. Data are shown as mean ± SEM, four to five sets of independent experiments were performed. Data were analyzed by Student’s t-test and showed significance at *P < 0.05.
Increased retinal degeneration in the absence of Atf6 in Rho +/P23H retina in older mice
Last, we examined Rho+/P23H mice lacking Atf6 at an older timepoint—P60. In contrast to P15 and P30 mice, morphological analysis of P60 retinas revealed significantly thinner ONL, OPL, INL, and IPL in Atf6−/−Rho+/P23H when compared to Atf6+/−Rho+/P23H retinas (Fig. 4a). Furthermore, the P60 Atf6−/−Rho+/P23H showed shortening of outer segments and inner segments of photoreceptors in the ONL as previously described22,23. In the ONL, the ventral retinal ONL appeared to be selectively degenerated while the dorsal ONL was preserved in Atf6−/−Rho+/P23H retinas (Fig. 4b; *P < 0.05; **P < 0.005, ***P < 0.0005, ****P < 0.0001). Consistent with the loss of photoreceptors, we observed a reduction in total rhodopsin protein levels in Atf6−/−Rho+/P23H retinas compared to Atf6+/−Rho+/P23H retinas (P = 0.03, Fig. 4c,d). Furthermore, no differences were observed in BiP/Grp78 or IRE1a levels at this age (Fig. 4c,d). These findings demonstrate that, by P60, loss of Atf6 leads to increased retinal degeneration in Rho+/P23H mice. We speculate that in the absence of Atf6, the duration and intensity of ER stress overwhelms the capacity of IRE1 to support ER homeostasis, leading to increased cell death at later ages of Atf6−/−Rho+/P23H mice35,36.
Loss of rhodopsin protein and attenuation of retinal lamina in Rho + /P23H in the absence of Atf6 at P60. (a) Light micrographs taken from vertical cryostat sections processed for H&E staining in P60 Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (b) The retinal layers of H&E-stained retinal sections through the optic nerve (0) were measured at 8 locations around the retina, four each in the dorsal and ventral hemispheres. The ventral retinal layers are significantly reduced compared to dorsal part of the retinas in Atf6−/−Rho+/P23H compared to Atf6+/−Rho+/P23H retinas at P60 (n = 3–5, Data represents mean ± SEM, Two-way ANOVA, *P < 0.05, **P < 0.005, ***P < 0.0005, ****P < 0.0001). ONL outer nuclear layer; OPL outer plexiform layer; INL inner nuclear layer; IPL inner plexiform layer. Scale bar = 50 μm. (c) Total rhodopsin, BiP/Grp78, and IRE1a from Atf6+/−Rho+/P23H (n = 4) and Atf6−/−Rho+/P23H (n = 4) retinas were detected by immunoblotting and quantified. HSP90 served as the loading control. Full-length blots are presented in Supplementary Fig. S4. (d) Rhodopsin, BiP/Grp78, and IRE1a protein levels were quantified in the retinas of Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H and normalized to HSP90. Reduction of rhodopsin protein levels was observed in Atf6−/−Rho+/P23H compared to Atf6+/−Rho+/P23H retinas while there were no changes in BiP/Grp78 protein levels in both genotypes. Data are shown as mean ± SEM, four sets of independent experiments were performed. Data were analyzed by Student’s t-test and showed significance at *P < 0.05.
Assessment of scotopic and photopic function with ERGs
Last, we evaluated rod and cone function in these P60 Rho+/P23H mice lacking Atf6. For the visual response of rods, we measured scotopic (Fig. 5) responses at P60 in Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H of both oculus sinister (OS) and oculus dexter (OD) by full-field ERG30. First, the scotopic ERG was recorded, and the amplitudes of the b-wave were analyzed (Fig. 5). In addition, an example of waveforms of the scotopic ERG responses from P60 Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) (Fig. 5a,b) retinas were generated. For the amplitudes of the resulting b-wave responses at all light intensities, we did not detect differences in scotopic rod response between P60 Atf6+/−Rho+/P23H and P60 Atf6−/−Rho+/P23H mice. Based on these data, the reduction of retinal layers in ventral retina observed in P60 in absence of Atf6 in Rho+/P23H retina were likely not detected with full-field ERG due to preservation of the dorsal ONL layer.
Scotopic ERG recordings from Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (a) Representative waveforms generated by scotopic intensity series (− 1.5 to 2 log cd s/m2 stimuli) for Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in OS retinas. (b) Representative waveforms generated by scotopic intensity series (− 1.5 to 2 log cd s/m2 stimuli) for Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in OD retinas. (c,d) In P60, the b-wave amplitudes of Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in across multiple intensities, ranging from − 1.5 to 2 log cd s/m2 showed no significant difference (n = 6, Data represents mean ± SEM, Two-way ANOVA, p > 0.5).
The photopic ERGs also demonstrated no noticeable changes between P60 Atf6+/−Rho+/P23H and P60 Atf6−/−Rho+/P23H retinas (Fig. 6). An example of waveforms of the photopic ERG responses from Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) (Fig. 6a,b) retinas are shown. For the amplitudes of the resulting b-wave responses at all light intensities, we were unable to detect differences in photopic cone response between Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H P60 mice. In addition, the pattern of waveforms between the two groups were consistent across all light intensities measured.
Photopic ERG recordings from Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H retinas. (a) Representative waveforms generated by photopic intensity series (− 0.31 to 2.81 log cd s/m2 stimuli) for Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in OS retinas. (b) Representative waveforms generated by photopic intensity series (− 0.31 to 2.81 log cd s/m2 stimuli) for Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in OD retinas. (c,d) In P60, the b-wave amplitudes of Atf6+/−Rho+/P23H (blue) and Atf6−/−Rho+/P23H (red) in across multiple intensities, ranging from − 0.31 to 2.81 log cd s/m2 stimuli showed no significant difference (n = 6, Data represents mean ± SEM, Two-way ANOVA, p > 0.5).
Discussion
Many disease variants in the human RHODOPSIN gene found in RP patients introduce missense mutations in the rhodopsin polypeptide that cause rhodopsin protein misfolding, retention in the ER, and inability to bind to 11-cis-retinal6,25,37,38. These molecular defects instigate rod photoreceptor decline by incompletely understood mechanisms and, ultimately, lead to the clinical manifestations of RP. Currently, there is no cure for RP caused by these misfolded rhodopsin proteins. We previously found that chemical-genetic activation of the ATF6 signaling pathway significantly reduced protein levels of several misfolded RP rhodopsin variants, such as T17M, Y178C, C185R, D190G, and K296E rhodopsin, while sparing wild-type rhodopsin when expressed in heterologous HEK293 cells39. Furthermore, activation of ATF6 reduced misfolded P23H mutant rhodopsin protein levels (monomer, dimer, and multimers) in HEK293 cell in vitro39. Here, we investigated how ATF6 signaling affected P23H mutant rhodopsin protein in photoreceptors in vivo. We examined the steady-state levels of total rhodopsin protein in retinal samples collected from Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice. The rhodopsin protein species in these heterozygous Rho+/P23H mice consist of wild-type and P23H rhodopsin. We found significantly more (nearly 2x) total rhodopsin protein in Atf6−/− Rho+/P23H compared to Atf6+/− Rho+/P23H while rhodopsin mRNA levels did not significantly change between these strains of mice at 12. These findings provide support that Atf6 is important for rhodopsin protein quality control in rod photoreceptors, because in the Atf6−/−Rho+/P23H mice, steady state rhodopsin protein levels increased almost 2x. ATF6 signaling likely ensures the efficient degradation of mutant P23H rhodopsin protein through transcriptional induction of factors involved in ER protein folding and ERAD20,21,40,41. Therefore, this model demonstrates that loss of Atf6 leads to accumulation of P23H rhodopsin protein that contributes to the ~ 2 × increase in steady-state rhodopsin protein levels at early ages. Our findings may provide mechanistic insight into prior studies demonstrating a protective role for ATF6 activity in RP models. For example, in vivo intravitreal AAV injection of one of ATF6’s downstream targets, the BiP/Grp78 chaperone, into P23H rhodopsin transgenic rats improved ERG responses42. This protective response could arise from increased elimination of the P23H rhodopsin protein through increased BiP/Grp78 chaperone-mediated increase in ERAD. Taken together, these findings underscore the importance of Atf6 plays in rhodopsin protein homeostasis in rods.
Variants in the human ATF6 gene cause achromatopsia and cone-rod dystrophy carrying bi-allelic disease alleles30,43,44,45,46,47. Patients with these ATF6 mutations showed malformation of the fovea, dysfunction of photoreceptors, and severe vision loss from infancy30,45. Furthermore, it is reported that abnormal retinal vasculature development may lead to malformation of the fovea48,49. However, none of these findings are apparent in young Atf6−/− mice or in young Atf6−/−Rho+/P23H mice30 (Supplemental Fig. S1). This difference may reflect a selective function for ATF6 in human cone and/or foveal development. For example, retinal organoids produced from the patients homozygous for ATF6 disease alleles showed significant defects in cone photoreceptor development accompanied by reduction in cone gene expression which included all cone phototransduction genes (CNGB3, CNGA3, PDE6C, PDE6H, and GNAT2) and red and green cone opsin genes50. Although cones do not appear to be selectively compromised in Atf6−/− mice or Atf6−/−Rho+/P23H mice, the absence of Atf6 does accelerate degeneration throughout the retina in Rho+/P23H mice. This is consistent with Atf6 expression in all retinal cell types, where it likely functions to ensure cell viability in the face of ER stress throughout life47.
In our study, we found that the IRE1 signaling pathway was hyper-activated in Atf6−/−Rho+/P23H mice when rhodopsin steady-state levels were increased at young ages. We propose that this hyper-activity in IRE1 signaling reflects a compensatory response to loss of Atf6. Specifically, the loss of Atf6 leads to reduced degradation of mutant rhodopsin protein in Rho+/P23H. In turn, this accumulation of misfolded rhodopsin hyper-activates the IRE1 signaling pathway to degrade the increased rhodopsin accumulating in P12 Atf6−/− Rho+/P23H. Consistent with a role for IRE1 signaling in rhodopsin degradation, we have previously demonstrated that the IRE1 signaling pathway of the UPR is selectively activated in photoreceptors of Rho+/P23HERAI+/− compared to Rho+/+ERAI+/− mice in a study that used the ERAI mouse GFP reporter line to indicate IRE1-XBP-1 activation22. We found that induction of ERAD by IRE1 signaling leads to ubiquitination of P23H rhodopsin in photoreceptors in Rho+/P23H mice22. This demonstrated that P23H rhodopsin is rapidly degraded by induction of ERAD in photoreceptors to eliminate misfolded rhodopsin from the ER in vivo. Furthermore, in a 2021 ARVO poster, Massoudi et al. selectively deleted the gene encoding IRE1a in rod photoreceptor in Rho+/P23H mice and demonstrated that ablation of Ire1a in rod photoreceptors damaged retinal function and increased retinal degeneration in Rho+/P23H mice51. Based on our previous and current study and the recent report by Massoudi et al. (2021), both ATF6 and IRE1a protect against ER stress in photoreceptors in Rho+/P23H mice. Our current study showed that the levels of Xbp-1s mRNA, BiP/Grp78 protein, and other transcriptional targets were significantly increased in the retinas of Atf6−/− Rho+/P23H mice compared to Atf6+/−Rho+/P23H mice at early age. Many of XBP-1’s target genes encode components of the ERAD pathway, and these genes have been found to be upregulated in the retinas of Rho+/P23H mice12,13,22. These findings suggest that degradation of P23H rhodopsin via downstream transcriptional activity of the IRE1-XBP-1s pathway and, consequently, ERAD, both work to alleviate ER stress caused by the accumulation of misfolded rhodopsin. Our model is further supported by the findings that E3 ubiquitin ligases, SORDD1/2, was able to facilitate degradation of Rh1P37H (the Drosophila equivalent of P23H rhodopsin) at larval and earlier stages of growth to allow for development of healthy adult eyes. Furthermore, SORDD1/2 and HRD1/SYVN1 were also able to prevent retinal degeneration in Drosophila with the G69D (glycine to aspartic acid at amino acid residue 69) rhodopsin mutation52. The lack of Atf6 in Rho+/P23H mice may initially increase the ability of E3 ubiquitin ligases downstream of IRE1-XBP-1s-ERAD to target misfolded rhodopsin in early stages of life. In contrast, Chop mRNA levels (an ER stress gene induced by PERK pathway)22,53 were not affected between Atf6+/− Rho+/P23H and Atf6−/−Rho+/P23H mice, which is consistent with previous studies showing that Chop was not induced during retinal degeneration in P23H rhodopsin mice and that the loss of CHOP had no impact on retinal degeneration based on histology or ERG31,54. Activation of PERK signaling also did not lead to greater reduction in rhodopsin protein levels in WT or P23H mice27.
There are several lines of evidence suggesting alterations of other degradation systems in Atf6−/−Rho+/P23H mice. We have previously reported in cell culture models that IRE1 relies on functioning proteasomes and lysosomes to degrade the mutated, misfolded rhodopsin27. Yao et al. (2018) also reported that P23H mice experience increase in autophagy secondary to ER stress, which leads to proteasome insufficiency and increase retinal degeneration. In contrast, genetic or pharmacologic inhibition of autophagy reduced retinal degeneration and improved proteasome levels55. Modulating the ratio between autophagy and proteasome activity (A:P) also helped to improve photoreceptor survival56. The authors demonstrated that normalizing the A:P ratio, either by improving folding of P23H rhodopsin or increasing proteasome activity to keep autophagy pathways down, increased photoreceptor survival and preserved retinal function. Taken together, we suggest that autophagy activity is increased as a result of the loss of Atf6.
We found increased retinal degeneration and diminished rhodopsin protein levels in P60 Atf6−/−Rho+/P23H retinas compared to P60 Atf6+/−Rho+/P23H. Furthermore, we found that the thickness of retinal layers including ONL, OPL, INL, and IPL were also significantly lower in the ventral part of the Atf6−/−Rho+/P23H retina compared to Atf6+/−Rho+/P23H. The reduction of ONL in the ventral part of the retina is consistent with previous histological data but the thickness of other retinal layers was not measured previously22,23,24. In Rho+/P23H mice, approximately half of the rod photoreceptor cells had disappeared between P14-P40 when compared to Rho+/+ retina, which showed no reduction of rod photoreceptors between P40 and P63 as described in previous studies22,23,24. Our data demonstrate that by P60, loss of Atf6 accelerates retinal degeneration in Rho+/P23H mice. Why does loss of Atf6 increase retinal degeneration in Rho+/P23H mice at P60, while not affecting younger animals? Our previous study showed that early wave of photoreceptor cell death and peak induction of the IRE1 reporter occur during the first postnatal month in Rho+/P23H mice22. The activation of IRE1 in Rho+/P23H is maintained throughout life to regulate proteostatic balance to remove P23H rhodopsin22. We propose that hyperactivation of IRE1 (as seen in the younger animals) restored rhodopsin protein homeostasis in the absence of Atf6 beginning at P12 (i.e., early stage), so that P30 Rho+/P23H retinas looked indistinguishable. Why can’t IRE1 hyperactivation keep rhodopsin and retina healthy at P60? We propose that the capacity of IRE1 to support ER homeostasis may ultimately be overwhelmed in the absence of Atf6, leading to increased rod photoreceptor cell death, and reduction of rhodopsin protein levels at later ages of Atf6−/−Rho+/P23H mice35,36. The ongoing photoreceptor cell death in RP retina likely causes widespread ER stress from oxidative damage, mitochondrial dysfunction, and other metabolic degenerative mechanisms57,58. Other sources of ER stress arising in the degenerating retina include damaged lipids, proteins, carbohydrates, enzymes, and DNA in photoreceptor cells, which ultimately results in further photoreceptor cell death through lipid peroxidation59. Thus, P23H rhodopsin-induced cell damage in addition to P23H rhodopsin protein itself could elicit too much ER stress, overwhelming the proteostatic balance maintained by the IRE1 in in the Atf6−/−Rho+/P23H mice.
Here, we observed no detectable difference in the function of rods and cones between P60 Atf6+/−Rho+/P23H and P60 Atf6−/−Rho+/P23H mice. Although, we observed a reduction of ONL and other retinal layers in Atf6−/−Rho+/P23H mice compared to Atf6+/−Rho+/P23H mice, no significant difference was noted in amplitude of either the scotopic or photopic b-wave in the strains of mice. Why did the reduction of retinal layers in Atf6−/−Rho+/P23H mice compared to Atf6+/−Rho+/P23H mice show no functional changes? We propose that the full-field flash ERG is relatively insensitive to detect smaller defects60,61 because it represents the global retinal function via summed electrical response of the whole retina excited by a flash of light62. Thus, the reduction of retinal layers in ventral retina observed in P60 in the absence of Atf6 in Rho+/P23H retina was likely not detected with full-field ERG due to preservation of the dorsal ONL layer.
In recent years, numerous small molecules have been identified that activate or inhibit ATF6 or IRE127,63,64,65,66,67,68. Agonists of IRE1 signaling include Type 1 IRE1 kinase inhibitors, which allosterically activate the RNAse function of IRE1, and IRE1 activators, which activate both RNAse and kinase function; however, these small molecule candidates (e.g. 474, IXA4, and IXA6), albeit showing no activation of IRE1-dependent cell death pathways, have yet to be fully tested for rhodopsin proteostatic properties67,69. By contrast, ATF6 agonists (e.g., AA147 and AA263) are effective in vivo and may have significant implications for amyloid related diseases and retinal development through ATF6 activation53,69,70. Furthermore, research from other groups similarly propose that BiP/Grp78, a prominent target of ATF6 upon ER stress, alleviates P23H RP symptoms42. We propose that ATF6 and IRE1-XBP-1 small molecule agonists are promising agents for further RP clinical studies if their rhodopsin proteostatic properties can be shown in vivo.
Methods
Animals
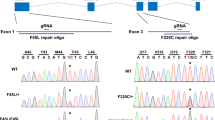
Transgenic Atf6+/+ and Atf6−/−30,41 and Rho P23H-KI mice23,24 on a pure C57BL/6J background were used to generate Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice for the experiments. First, breeding pairs of Atf6+/+Rho+/P23H and Atf6−/−Rho+/+ mice of C57BL/6J background were crossed to generate Atf6+/−Rho+/P23H. Breeding pairs of Atf6−/−Rho+/P23H with Atf6−/−Rho+/+ mice of C57BL/6J background were crossed to generate Atf6−/−Rho+/P23H.
There are no reported differences in the phenotypes between Atf6+/+ and Atf6+/− mice30,41,42,44. Injection of Atf6+/+, Atf6+/−, and Atf6−/− mice with tunicamycin led to kidney and liver toxicity only in Atf6−/− animals but not in Atf6+/+ or Atf6+/− mice; in addition, our previous study has shown normal morphology and normal rhodopsin expression when comparing Atf6+/+ to Atf6+/− mice30. All experiments used female or male Atf6−/−Rho+/P23H mice in comparison to control littermates Atf6+/−Rho+/P23H, at the postnatal (P) days 12, 15, 30, and 60 (number (n) = 3 ~ 6 respectively for each stage). For retinal vasculature assessment in Atf6−/− mice, female and male P30 Atf6+/+ and P30 Atf6−/− mice (n = 3 animals per group) on a C57BL/6 J background were used as described in previous studies30,41. For all experiments, animals were kept in cyclic 12-h light/dark conditions with free access to food and water. All mouse care and experimental procedures in this study were approved and conducted in strict accordance with relevant guidelines and regulations by the Institutional Animal Care and Use Committee at the Stanford University and in compliance with the Association for Research in Vision and Ophthalmology Statement for the Use of Animals in Ophthalmic and Vision Research and the ARRIVE (Animal Research: Reporting of in Vivo Experiments) guidelines.
Tissue preparation
The animals were euthanized by carbon dioxide euthanasia at P12, P15, P30, and P60. The eyes were enucleated for collection of retinal tissue. For secondary method, we performed cervical dislocation. The lens and the anterior segment were removed, and the eyecups were further dissected to collect whole retinal lysate for biochemistry or molecular biology, or eyecups were fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (PB), for 60 min at 4 °C. After fixation, the eyecups were processed for hematoxylin and eosin (H&E) staining71 and cryostat sectioning. For cryostat sectioning, eyecups were transferred from 10% for 1 h to 20% for 1 h to 30% sucrose overnight at 4 °C, then eyecups were embedded in Optimal Cutting Temperature (OCT) medium (Tissue-Tek, Elkhart, IN), frozen in liquid nitrogen and subsequently vertically sectioned on a Leica cryostat (Leica Biosystems Inc, Buffalo Grove, IL) at a thickness of 20 μm. For wholemount retinal preparation, the retinas were isolated from the eyecups and dissected as wholemounts.
H & E staining
The detail protocols for H & E staining in retinal layer was performed as previously published71. Three to five left eyecups from three to five animals (n = 3–5) were sectioned along the vertical meridian on a cryostat at a thickness of 20 μm. Sections were then collected on gelatin-coated slides for H&E staining. Slides were dipped in Harris hematoxylin for 1 min then they were washed in tap water and dehydrated in alcohol. Slides were then dipped in Eosin-Phloxyine for 30 s, then dehydrated in a series of 95% ethanol and 100% ethanol followed by 5 min in xylene, and mounted in Vectashield mounting medium (Vector Labs, Burlingame, CA).
Immunoblotting analysis
The detail protocols for immunoblotting analysis were performed as previously published27,45. Three to five right retinas from three to five animals (n = 3–5) were lysed in lysis buffer (0.5 g/mL n-Dodecyl β-d-maltoside (Calbiochem EMD Bioscience, San Diego, CA) in PBS), protease inhibitor (Sigma-Aldrich, St. Louis, MO) and phosphatase inhibitor (Thermo Scientific, Rockford, IL). Protein concentrations of the total retinal lysates were determined by BCA protein assay (Pierce, Rockford, IL). Equal amounts of protein were applied onto 4–15% Mini-PROTEAN TGX precast gels (Bio-Rad, Hercules, CA) and analyzed by immunoblot. Antibodies B630N anti-rhodopsin 1:100039 (gift of W.C. Smith, Gainesville, FL); anti-BiP/Grp78 at 1:100027, anti-IRE1a at 1:100072, and anti-HSP90 at 1:100022 (GeneTex, Inc., Irvine, CA) were used. After overnight incubation with primary antibody in a 4 °C cold room, membranes were washed in TBS with 0.1% Tween-20, followed by incubation of a horseradish peroxidase-coupled secondary antibody (Cell Signaling, Danvers, MA). Immunoreactive bands were detected with the SuperSignal West chemiluminescent substrate (Pierce, Rockford, IL).
Quantitative PCR analysis (qPCR)
The detail protocols for qPCR analysis were performed as previously published44,45. Three to five right retinas from three to five animals (n = 3–5) were lysed and total RNA was collected with a RNeasy mini kit (Qiagen, Germany) and mRNA was reverse transcribed with the iScript cDNA Synthesis Kit (Bio Rad, Hercules, CA). Primers that were used included22,31: mouse Rhodopsin mRNA, 5′-TTCACCACCACCCTCTACACATCAC-3′ and 5′-CGGAAGTTGCTCATCGGCTTG-3′; mouse Xbp-1s mRNA, 5′- GAGTCCGCAGCAGGTG-3′ and 5′-GTGTCAGAGTCCATGGGA-3′; mouse Syvn1, 5’- ACACACTACTGGATGCTGCC-3’ and 5’- GCTTCAGGAATTGGTGGGGA-3’; mouse Chop: 5′- ACGGAAACAGAGTGGTCAGTGC-3′ and 5′-CAGGAGGTGATGCCCACTGTTC-3′, and mouse Rpl19: 5′-ATGCCAACTCCCGTCAGCAG- 3′ and 5′- TCATCCTTCTCATCCAGGTCACC-3′. Rpl19 mRNA levels were used as internal normalization standards for qPCR analysis as they were not altered by ER stress. qPCR conditions were 95 °C for 5 min; 95 °C for 10 s; 60 °C for 10 s; 72 °C for 10 s, with 50 cycles of amplification.
Electroretinography (ERG) and quantification
Mice were dark adapted for 24 h prior to recordings. ERGs (Diagnosys LLC, Lowell, MA) were recorded from both eyes of Atf6−/−Rho+/P23H (n = 6) mice and compared to ERGs from eyes of Atf6+/−Rho+/P23H control littermates (n = 6) at P60 as described previously30. Mice were anaesthetized using a combination of ketamine (20 mg/kg; KETASET, Fort Dodge, IA, USA) and xylazine (5 mg/kg, X-Ject SA; Butler, Dublin, OH, USA) using similar procedures as our published protocols30. Under a dim red light, the pupils were dilated with Atropine sulfate ophthalmic solution 1% (Akorn Inc, Lake Forest, IL, USA). The recording electrodes attached to two gold wire rings were placed on the cornea of both eyes. The eye lubricant hypromellose ophthalmic gel, USP 2.5% (HUB pharmaceuticals, LLC, Rancho Cucamonga, CA, USA) was applied to keep the hydration and conductivity between the cornea and recording electrodes. The ground and reference electrodes were placed at the tail and tongue, respectively. The eyes were then given scotopic ERG responses (a series of white light flashes varying from -1.5 to 2 log cd s/m2). After 10 min of light adaptation, photopic ERG responses of -0.31 to 2.81 log cd s/m2 were recorded. The amplitudes for the resulting b-wave responses at the series of light flash intensity were plotted.
Retinal vasculature staining in Atf6 −/− mice
For wholemount immunohistochemical staining, the same procedures described in our previous studies were used30,71. Three right retinas from three animals (n = 3) were used for wholemount staining. Wholemounts were treated with 1% Triton X-100 in 0.1 M PBS (40 min) before NDS (1 h), and antibody against isolectin B4-Alexa 488 (IB4, molecular probe, 1:200)73 was diluted in 0.5% Triton X-100 in 0.1 M PBS (48 h at 4 °C). After incubation, wholemounts were washed for 30 min with 0.1 M PB and cover slipped with Vectashield mounting medium.
Equipment and settings
IB-4 staining wholemount (excitation 488, emission 552, 63 × 1.40 oil objective) images were acquired using a Leica SP8 DLS confocal microscope. Images were processed with the Leica application suite-X software (3.0.11.20652, Leica Mcirosystems—Dimension X × Y—local size 1024 × 1024 pixels, 8 Bit for superficial layer, intermediate layer, and deep layer; Dimension X × Y-local size 4753 × 4753 pixels, 8 Bit for entire wholemount images). For retinal vasculature assessment in Atf6−/− mice, confocal micrographs of the wholemounts (n = 3, animals per group) were taken at the nerve fiber layer (superficial layer), at the IPL (intermediate layer), and at the OPL (deep layer) of the dorsal regions (1 mm away from optic disc) of the retina. At these regions, serial optical section (Dimension z, 2 µm intervals) was made using a confocal microscope. H & E staining sections (20 × objective) images under brightfield were acquired using a NanoZoomer 2.0-HT slide scanner NDP scan 2.5 and viewed using NDP view 2 (Hamamatsu Photonics). The Hamamatsu NanoZoomer uses 3-chip time-delay integration (TDI) sensor signal. Images were processed with the Image Lab Touch Software version 3.0 (Bio Rad). For retinal layer thickness measurement, thickness of the retina was measured at 0.5 mm intervals beginning from the optic nerve. For each retinal section, three measurements of the ONL, OPL, INL, and IPL thickness were taken for each section (fields covering 350 µm × 350 µm), spaced approximately 100 µm apart, which were then averaged74. Layer thickness measurements were collected from three to five retinas from separate Atf6+/−Rho+/P23H and Atf6−/−Rho+/P23H mice. The results were plotted as a spider plot with distance from the optic nerve as the x-axis and thickness of retinal layer as the y-axis. Intensity of Immunoreactive bands in all blots were measured with National Institute of Health (NIH) Image J software version 1.50i. For presentation, all Photoshop (Adobe photoshop CC 2020) adjustments (brightness and contrast only) were carried out equally in each figure.
Statistical analysis
All the statistics were expressed as mean ± standard error of the mean (SEM). Student’s t-test was used for comparison. Two-way ANOVA and Fisher's least significant difference procedure (LSD test) were used to examine the differences among the group of means. All the statistical tests were performed using GraphPad Prism Version 8.3.1. The difference between the means of separate experimental groups was considered statistically significant at P < 0.05.
References
Berson, E. L. Retinitis pigmentosa. The Friedenwald lecture. Invest. Ophthalmol. Vis. Sci. 34, 1659–1676 (1993).
Hartong, D. T., Berson, E. L. & Dryja, T. P. Retinitis pigmentosa. Lancet 368, 1795–1809. https://doi.org/10.1016/S0140-6736(06)69740-7 (2006).
Berger, W., Kloeckener-Gruissem, B. & Neidhardt, J. The molecular basis of human retinal and vitreoretinal diseases. Prog. Retina Eye Res. 29, 335–375. https://doi.org/10.1016/j.preteyeres.2010.03.004 (2010).
Verbakel, S. K. et al. Non-syndromic retinitis pigmentosa. Prog. Retina Eye Res. 66, 157–186. https://doi.org/10.1016/j.preteyeres.2018.03.005 (2018).
Dryja, T. P. et al. Mutations within the rhodopsin gene in patients with autosomal dominant retinitis pigmentosa. N. Engl. J. Med. 323, 1302–1307. https://doi.org/10.1056/NEJM199011083231903 (1990).
Sung, C. H. et al. Rhodopsin mutations in autosomal dominant retinitis pigmentosa. Proc. Natl. Acad. Sci. U S A 88, 6481–6485. https://doi.org/10.1073/pnas.88.15.6481 (1991).
Wang, Q. et al. Update on the molecular genetics of retinitis pigmentosa. Ophthalmic Genet. 22, 133–154. https://doi.org/10.1076/opge.22.3.133.2224 (2001).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086. https://doi.org/10.1126/science.1209038 (2011).
Calfon, M. et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 415, 92–96. https://doi.org/10.1038/415092a (2002).
Cox, J. S., Shamu, C. E. & Walter, P. Transcriptional induction of genes encoding endoplasmic reticulum resident proteins requires a transmembrane protein kinase. Cell 73, 1197–1206. https://doi.org/10.1016/0092-8674(93)90648-a (1993).
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891. https://doi.org/10.1016/s0092-8674(01)00611-0 (2001).
Lee, A. H., Iwakoshi, N. N. & Glimcher, L. H. XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol. Cell. Biol. 23, 7448–7459. https://doi.org/10.1128/mcb.23.21.7448-7459.2003 (2003).
Shoulders, M. D. et al. Stress-independent activation of XBP1s and/or ATF6 reveals three functionally diverse ER proteostasis environments. Cell. Rep. 3, 1279–1292. https://doi.org/10.1016/j.celrep.2013.03.024 (2013).
Harding, H. P., Zhang, Y. & Ron, D. Protein translation and folding are coupled by an endoplasmic-reticulum-resident kinase. Nature 397, 271–274. https://doi.org/10.1038/16729 (1999).
Ron, D. & Harding, H. P. Protein-folding homeostasis in the endoplasmic reticulum and nutritional regulation. Cold Spring Harb. Perspect. Biol. 4, a013177. https://doi.org/10.1101/cshperspect.a013177 (2012).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell. Biol. 8, 519–529. https://doi.org/10.1038/nrm2199 (2007).
Han, J. et al. ER stress signalling through eIF2alpha and CHOP, but not IRE1alpha, attenuates adipogenesis in mice. Diabetologia 56, 911–924. https://doi.org/10.1007/s00125-012-2809-5 (2013).
Tabas, I. & Ron, D. Integrating the mechanisms of apoptosis induced by endoplasmic reticulum stress. Nat. Cell. Biol. 13, 184–190. https://doi.org/10.1038/ncb0311-184 (2011).
Chen, C. Y. et al. Signal peptide peptidase functions in ERAD to cleave the unfolded protein response regulator XBP1u. EMBO J. 33, 2492–2506. https://doi.org/10.15252/embj.201488208 (2014).
Haze, K., Yoshida, H., Yanagi, H., Yura, T. & Mori, K. Mammalian transcription factor ATF6 is synthesized as a transmembrane protein and activated by proteolysis in response to endoplasmic reticulum stress. Mol. Biol. Cell 10, 3787–3799. https://doi.org/10.1091/mbc.10.11.3787 (1999).
Ye, J. et al. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 6, 1355–1364. https://doi.org/10.1016/s1097-2765(00)00133-7 (2000).
Chiang, W. C. et al. Robust endoplasmic reticulum-associated degradation of rhodopsin precedes retinal degeneration. Mol. Neurobiol. 52, 679–695. https://doi.org/10.1007/s12035-014-8881-8 (2015).
Sakami, S. et al. Probing mechanisms of photoreceptor degeneration in a new mouse model of the common form of autosomal dominant retinitis pigmentosa due to P23H opsin mutations. J. Biol. Chem. 286, 10551–10567. https://doi.org/10.1074/jbc.M110.209759 (2011).
Sakami, S., Kolesnikov, A. V., Kefalov, V. J. & Palczewski, K. P23H opsin knock-in mice reveal a novel step in retinal rod disc morphogenesis. Hum. Mol. Genet. 23, 1723–1741. https://doi.org/10.1093/hmg/ddt561 (2014).
Saliba, R. S., Munro, P. M., Luthert, P. J. & Cheetham, M. E. The cellular fate of mutant rhodopsin: quality control, degradation and aggresome formation. J. Cell Sci. 115, 2907–2918 (2002).
Alavi, M. V. et al. In vivo visualization of endoplasmic reticulum stress in the retina using the ERAI reporter mouse. Invest. Ophthalmol. Vis. Sci. 56, 6961–6970. https://doi.org/10.1167/iovs.15-16969 (2015).
Chiang, W. C., Hiramatsu, N., Messah, C., Kroeger, H. & Lin, J. H. Selective activation of ATF6 and PERK endoplasmic reticulum stress signaling pathways prevent mutant rhodopsin accumulation. Invest. Ophthalmol. Vis. Sci. 53, 7159–7166. https://doi.org/10.1167/iovs.12-10222 (2012).
Reimold, A. M. et al. An essential role in liver development for transcription factor XBP-1. Genes Dev. 14, 152–157 (2000).
Zhang, K. et al. The unfolded protein response sensor IRE1alpha is required at 2 distinct steps in B cell lymphopoiesis. J. Clin. Invest. 115, 268–281. https://doi.org/10.1172/JCI21848 (2005).
Kohl, S. et al. Mutations in the unfolded protein response regulator ATF6 cause the cone dysfunction disorder achromatopsia. Nat. Genet. 47, 757–765. https://doi.org/10.1038/ng.3319 (2015).
Chiang, W. C. et al. Ablation of chop transiently enhances photoreceptor survival but does not prevent retinal degeneration in transgenic mice expressing human P23H rhodopsin. Adv. Exp. Med. Biol. 854, 185–191. https://doi.org/10.1007/978-3-319-17121-0_25 (2016).
LaVail, M. M. et al. Phenotypic characterization of P23H and S334ter rhodopsin transgenic rat models of inherited retinal degeneration. Exp. Eye Res. 167, 56–90. https://doi.org/10.1016/j.exer.2017.10.023 (2018).
Olsson, J. E. et al. Transgenic mice with a rhodopsin mutation (Pro23His): A mouse model of autosomal dominant retinitis pigmentosa. Neuron 9, 815–830. https://doi.org/10.1016/0896-6273(92)90236-7 (1992).
Tan, E. et al. The relationship between opsin overexpression and photoreceptor degeneration. Invest. Ophthalmol. Vis. Sci. 42, 589–600 (2001).
Paschen, W. & Frandsen, A. Endoplasmic reticulum dysfunction: A common denominator for cell injury in acute and degenerative diseases of the brain?. J. Neurochem. 79, 719–725. https://doi.org/10.1046/j.1471-4159.2001.00623.x (2001).
Rao, R. V. & Bredesen, D. E. Misfolded proteins, endoplasmic reticulum stress and neurodegeneration. Curr. Opin. Cell. Biol. 16, 653–662. https://doi.org/10.1016/j.ceb.2004.09.012 (2004).
Illing, M. E., Rajan, R. S., Bence, N. F. & Kopito, R. R. A rhodopsin mutant linked to autosomal dominant retinitis pigmentosa is prone to aggregate and interacts with the ubiquitin proteasome system. J. Biol. Chem. 277, 34150–34160. https://doi.org/10.1074/jbc.M204955200 (2002).
Kaushal, S. & Khorana, H. G. Structure and function in rhodopsin. 7. Point mutations associated with autosomal dominant retinitis pigmentosa. Biochemistry 33, 6121–6128. https://doi.org/10.1021/bi00186a011 (1994).
Chiang, W. C., Messah, C. & Lin, J. H. IRE1 directs proteasomal and lysosomal degradation of misfolded rhodopsin. Mol. Biol. Cell 23, 758–770. https://doi.org/10.1091/mbc.E11-08-0663 (2012).
Adachi, Y. et al. ATF6 is a transcription factor specializing in the regulation of quality control proteins in the endoplasmic reticulum. Cell Struct. Funct. 33, 75–89. https://doi.org/10.1247/csf.07044 (2008).
Wu, J. et al. ATF6alpha optimizes long-term endoplasmic reticulum function to protect cells from chronic stress. Dev. Cell 13, 351–364. https://doi.org/10.1016/j.devcel.2007.07.005 (2007).
Gorbatyuk, M. S. et al. Restoration of visual function in P23H rhodopsin transgenic rats by gene delivery of BiP/Grp78. Proc. Natl. Acad. Sci. USA 107, 5961–5966. https://doi.org/10.1073/pnas.0911991107 (2010).
Ansar, M. et al. Mutation of ATF6 causes autosomal recessive achromatopsia. Hum. Genet. 134, 941–950. https://doi.org/10.1007/s00439-015-1571-4 (2015).
Chiang, W. C. et al. Achromatopsia mutations target sequential steps of ATF6 activation. Proc. Natl. Acad. Sci. USA 114, 400–405. https://doi.org/10.1073/pnas.1606387114 (2017).
Lee, E. J. et al. Multiexon deletion alleles of ATF6 linked to achromatopsia. JCI Insight https://doi.org/10.1172/jci.insight.136041 (2020).
Skorczyk-Werner, A. et al. Autosomal recessive cone-rod dystrophy can be caused by mutations in the ATF6 gene. Eur. J. Hum. Genet. 25, 1210–1216. https://doi.org/10.1038/ejhg.2017.131 (2017).
Xu, M. et al. ATF6 is mutated in early onset photoreceptor degeneration with macular involvement. Invest. Ophthalmol. Vis. Sci. 56, 3889–3895. https://doi.org/10.1167/iovs.15-16778 (2015).
Kroeger, H. et al. The unfolded protein response regulator ATF6 promotes mesodermal differentiation. Sci. Signal. 11, eaan5785. https://doi.org/10.1126/scisignal.aan5785 (2018).
Liu, L. et al. Targeting the IRE1alpha/XBP1 and ATF6 arms of the unfolded protein response enhances VEGF blockade to prevent retinal and choroidal neovascularization. Am. J. Pathol. 182, 1412–1424. https://doi.org/10.1016/j.ajpath.2012.12.020 (2013).
Kroeger, H., Grandjean, J. M. D. & Lin, J. H. ATF6 is essential for human cone development. PNAS (revision).
Massoudi, D. et al. The UPR transducer IRE1α is required for photoreceptor health and protection against retinal degeneration. IOVS 62, 3073 (2021).
Xu, J., Zhao, H. & Wang, T. Suppression of retinal degeneration by two novel ERAD ubiquitin E3 ligases SORDD1/2 in Drosophila. PLoS Genet. 16, e1009172. https://doi.org/10.1371/journal.pgen.1009172 (2020).
Kroeger, H., Chiang, W. C. & Lin, J. H. Endoplasmic reticulum-associated degradation (ERAD) of misfolded glycoproteins and mutant P23H rhodopsin in photoreceptor cells. Adv. Exp. Med. Biol. 723, 559–565. https://doi.org/10.1007/978-1-4614-0631-0_71 (2012).
Adekeye, A., Haeri, M., Solessio, E. & Knox, B. E. Ablation of the proapoptotic genes CHOP or Ask1 does not prevent or delay loss of visual function in a P23H transgenic mouse model of retinitis pigmentosa. PLoS ONE 9, e83871. https://doi.org/10.1371/journal.pone.0083871 (2014).
Yao, J. et al. Inhibiting autophagy reduces retinal degeneration caused by protein misfolding. Autophagy 14, 1226–1238. https://doi.org/10.1080/15548627.2018.1463121 (2018).
Qiu, Y., Yao, J., Jia, L., Thompson, D. A. & Zacks, D. N. Shifting the balance of autophagy and proteasome activation reduces proteotoxic cell death: A novel therapeutic approach for restoring photoreceptor homeostasis. Cell Death Dis. 10, 547. https://doi.org/10.1038/s41419-019-1780-1 (2019).
Campian, J. L., Qian, M., Gao, X. & Eaton, J. W. Oxygen tolerance and coupling of mitochondrial electron transport. J. Biol. Chem. 279, 46580–46587. https://doi.org/10.1074/jbc.M406685200 (2004).
Portera-Cailliau, C., Sung, C. H., Nathans, J. & Adler, R. Apoptotic photoreceptor cell death in mouse models of retinitis pigmentosa. Proc. Natl. Acad. Sci. USA 91, 974–978. https://doi.org/10.1073/pnas.91.3.974 (1994).
Komeima, K., Rogers, B. S. & Campochiaro, P. A. Antioxidants slow photoreceptor cell death in mouse models of retinitis pigmentosa. J. Cell Physiol. 213, 809–815. https://doi.org/10.1002/jcp.21152 (2007).
Azarmina, M. Full-field versus multifocal electroretinography. J. Ophthalmic Vis. Res. 8, 191–192 (2013).
Dettoraki, M. & Moschos, M. M. The role of multifocal electroretinography in the assessment of drug-induced retinopathy: A review of the literature. Ophthalmic Res. 56, 169–177. https://doi.org/10.1159/000446321 (2016).
Holder, G. E. Electrophysiological assessment of optic nerve disease. Eye (Lond) 18, 1133–1143. https://doi.org/10.1038/sj.eye.6701573 (2004).
Blackwood, E. A. et al. Pharmacologic ATF6 activation confers global protection in widespread disease models by reprograming cellular proteostasis. Nat. Commun. 10, 187. https://doi.org/10.1038/s41467-018-08129-2 (2019).
Gallagher, C. M. et al. Ceapins are a new class of unfolded protein response inhibitors, selectively targeting the ATF6alpha branch. Elife https://doi.org/10.7554/eLife.11878 (2016).
Gallagher, C. M. & Walter, P. Ceapins inhibit ATF6alpha signaling by selectively preventing transport of ATF6alpha to the Golgi apparatus during ER stress. Elife https://doi.org/10.7554/eLife.11880 (2016).
Martindale, J. J. et al. Endoplasmic reticulum stress gene induction and protection from ischemia/reperfusion injury in the hearts of transgenic mice with a tamoxifen-regulated form of ATF6. Circ. Res. 98, 1186–1193. https://doi.org/10.1161/01.RES.0000220643.65941.8d (2006).
Plate, L. et al. Small molecule proteostasis regulators that reprogram the ER to reduce extracellular protein aggregation. Elife https://doi.org/10.7554/eLife.15550 (2016).
Torres, S. E. et al. Ceapins block the unfolded protein response sensor ATF6alpha by inducing a neomorphic inter-organelle tether. Elife https://doi.org/10.7554/eLife.46595 (2019).
Grandjean, J. M. D. & Wiseman, R. L. Small molecule strategies to harness the unfolded protein response: where do we go from here?. J. Biol. Chem. 295, 15692–15711. https://doi.org/10.1074/jbc.REV120.010218 (2020).
Paxman, R. et al. Pharmacologic ATF6 activating compounds are metabolically activated to selectively modify endoplasmic reticulum proteins. Elife https://doi.org/10.7554/eLife.37168 (2018).
Ji, Y., Zhu, C. L., Grzywacz, N. M. & Lee, E. J. Rearrangement of the cone mosaic in the retina of the rat model of retinitis pigmentosa. J. Comp. Neurol. 520, 874–888. https://doi.org/10.1002/cne.22800 (2012).
Roos, A. et al. Cellular signature of SIL1 depletion: Disease pathogenesis due to alterations in protein composition beyond the ER machinery. Mol. Neurobiol. 53, 5527–5541. https://doi.org/10.1007/s12035-015-9456-z (2016).
Jones, I., Hagglund, A. C. & Carlsson, L. Reduced mTORC1-signaling in retinal ganglion cells leads to vascular retinopathy. Dev. Dyn. https://doi.org/10.1002/dvdy.389 (2021).
Pak, J. S., Lee, E. J. & Craft, C. M. The retinal phenotype of Grk1-/- is compromised by a Crb1 rd8 mutation. Mol. Vis. 21, 1281–1294 (2015).
Acknowledgements
Research results reported in this publication was supported by NIH awards R01EY027735; CIRM DISC2-10973 award; and VA Merit I01BX002284. Portions of this work were supported by NIH grants R01 DK113171 and R01 AG062190 (RJK).
Author information
Authors and Affiliations
Contributions
E.L.: Conceptualization, Data curation, Formal analysis, Investigation, Project administration, Validation, Visualization: All figures; Writing—original draft, Writing—review & editing. P.C.: Data curation, Formal analysis, Investigation, Validation, Visualization: All figures; Writing—original draft, Writing—review & editing. L.C.: Data curation, Formal analysis, Investigation, Validation, Visualization: Figs. 2, 3, 4; Writing—review & editing. K.K.: Investigation, Validation; Figs. 2, 3, 4; Writing—review & editing. R.J.K.: Atf6 knockout mice, Writing—review & editing. J.L.: Conceptualization, Funding acquisition, Data curation, Investigation, Project administration, Resources, Supervision, Validation, Visualization: All figures; Writing—review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, EJ., Chan, P., Chea, L. et al. ATF6 is required for efficient rhodopsin clearance and retinal homeostasis in the P23H rho retinitis pigmentosa mouse model. Sci Rep 11, 16356 (2021). https://doi.org/10.1038/s41598-021-95895-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-95895-7
This article is cited by
-
TDP43 pathology in chronic traumatic encephalopathy retinas
Acta Neuropathologica Communications (2023)
-
Imbalanced unfolded protein response signaling contributes to 1-deoxysphingolipid retinal toxicity
Nature Communications (2023)
-
The generation of detergent-insoluble clipped fragments from an ERAD substrate in mammalian cells
Scientific Reports (2023)
-
Cellular stress signaling and the unfolded protein response in retinal degeneration: mechanisms and therapeutic implications
Molecular Neurodegeneration (2022)
-
Network biology analysis of P23H rhodopsin interactome identifies protein and mRNA quality control mechanisms
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.