Abstract
Parkinson’s disease is a common neurodegenerative disease. The differential expression of alpha-synuclein within Lewy Bodies leads to this disease. Some missense mutations of alpha-synuclein may resultant in functional aberrations. In this study, our objective is to verify the functional adaptation due to early and late-onset mutation which can trigger or control the rate of alpha-synuclein aggregation. In this regard, we have proposed a computational model to study the difference and similarities among the Wild type alpha-synuclein and mutants i.e., A30P, A53T, G51D, E46K, and H50Q. Evolutionary sequence space analysis is also performed in this experiment. Subsequently, a comparative study has been performed between structural information and sequence space outcomes. The study shows the structural variability among the selected subtypes. This information assists inter pathway modeling due to mutational aberrations. Based on the structural variability, we have identified the protein–protein interaction partners for each protein that helps to increase the robustness of the inter-pathway connectivity. Finally, few pathways have been identified from 12 semantic networks based on their association with mitochondrial dysfunction and dopaminergic pathways.
Similar content being viewed by others
Introduction
One of the common neurodegenerative disorders (NDs) is Parkinson’s disease (PD) which has been reported to be associated with the protein alpha-synuclein(Asyn)1,2. Asyn\(_s\) are usually known for their robust expression within lewy bodies (LB). For example, the association between expression levels of the Asyn especially through the Lewy Bodies, and functional disorder of the dopaminergic neuron are reported3. Propensity of the Asyn towards aggregation might be responsible for functional disorder of its protein partners4. These genetic aberrations due to mutations can lead to functional modifications. For example, the EGFR1 signaling pathway is highly dependent on the activities of the asyn proteins. Any mutational modification can lead to functional changes in EGFR1 signaling pathways. In 1997, a point mutation at G209A has been reported. The substitution has resulted in point mutation A53T which is an early onset mutation linked with PD. However, this mutation is highly debatable as threonine can naturally be identified at rodent synuclein homologs, followed by two more mutations Viz., E46K, and A30P. The observation has justified the occurrence of functional similarity and pathway orchestrations associated with synuclein homologs and paralogs. Early-onset and late-onset mutations Viz., A53T, E46K, H50Q and G51D are reported in5. Usually, alpha-synuclein has three domains. Among them, A53T, E46K and H50Q are shown to enhance the aggregation propensity6,7 whereas G51D is associated with slower rate of aggregation. Similarly , non-beta component of Alzheimer’s disease amyloid plaque (NAC) domain i.e., residue range 61–95 plays significant role during aggregation (residue range 61–95). On other hand, \(\beta \)- and \(\gamma \)-synucleins8 helps to inhibit the aggregation process Asyn. The mutations on genomic encoding of SNCA gene have been identified as major element in Lewy bodies. So far 16 loci (PARK1 to PARK16) and 11 genes have been associated with pathogenic progression of PD.
Impacts of such sequential changes are significant. For example, the amyloid formation mechanism of alpha-synuclein is mostly associated with a monomeric primary level of nucleation9. Usually the aggregation rate is too slow to detect. However, the phenotypical modifications due to mentioned mutations would have been affecting the residues at NAC domain (which is the prime point of aggregation). Therefore, intra-molecular allosteric effects would lead mutant to behave like ancestral phenotypes. However, these activities are directly affecting the functional profiles of the targeted proteins. Modifications, enhanced due to phenotypical changes, are correlated with the ontology terms. These are shared as common semantics among the pathways separately for wild type (WT) protein their mutants and closest neighbors from each of the cases. During the pathogenic progression, phenotypical modifications can explain the internal orchestration of the pathways based on the mutated candidate and their interacting neighbors. Mostly, these connections are observed for the mutated candidates by following the ancestral protein neighbors. A point mutation at a certain residual position cannot change the whole structure. However, it can affect the co-evoluting residues. Therefore, the relation among pathways is modified which may lead to the initiation of neurodegenerative progression.
In this study, one frame has been proposed by considering the sequential-structural modifications, and pathway semantics networks for Asyn and their mutants. The experiment has been initiated focusing on five mutations from early and late onset PD cases have been A30P, A53T, G51D, E46K, and H50Q. Firstly, the sequential trait of the synuclein family is observed. Then evolutionary conserved co-evolving patches are studied by performing the Direct Coupling Analysis (DCA). Subsequently the information fetched from sequence space has been mapped to the structure. For five mutated cases, two distinct structures has been predicted applying ab-initio method. For the wild type protein, the ensemble model has been selected from the known structures. There after, the structure network has been formed based on normal mode scores of each residue. Comparing these information, the effective residues due to mutations have been fetched. Subsequently, BLAST helps to find the sequential similarity with ancestors from the same family. This may provide different lists of neighboring interaction partners for the wild type and mutant. Finally, weighted networks are formed based on pathway semantic relations. The pathways and their relations represent the nodes and the edges of the network. The weight of the edges are computed based on the semantic similarities.
Results
It is known that \(\alpha \)-Synuclein is the candidate protein of Parkinson’s disease. In order to understand the effect of this protein and its mutated samples towards the neurodegenerative diseases, a frame is proposed based on sequential and structural changes along with the pathway semantics.
Sequence analysis
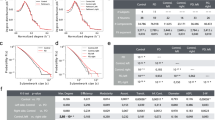
In sequence space, the family trait of the Synuclein family is analyzed in terms of order and disorder. In this regard, Shannon entropy is calculated for each protein belongs to this family, shown in Fig. 1. Moreover, the entropic score of consensus sequence is 2.58. This suggests that the family trait of Synuclein family is ordered, whereas many proteins belong to this family are disorder in nature. This gives an in-depth idea of evolution.
To support the evolutionary changes Direct Coupling Analysis (DCA) is performed. The co-varying residue patches are represented in Fig. 2a. DCA score indicates the change in one residue is responsible for the change in other coupled residues. The result depicts that the residue changing for point mutation is also responsible for the change of its co-varying patches which eventually contributes to slow as well as accelerating aggregation of \(\alpha \)-Synuclein. From the analysis, among five mutations E46K, H50Q and G51D mutated samples show conservation throughout the evolution. In parallel, the residue patches are clustered depending on their DCA score and residue in one cluster denotes same evolution rate. Interestingly, the mutated residues are present in one cluster shown in Fig. 2b, which indicates that the residue present in that cluster are varying accordingly.
Selection of structure and analysis
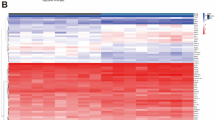
\(\alpha \)-Synuclein has multiple dynamic structure with frequent mutation. Due to this, an ensemble of all the structures is performed to understand the fluctuation rate, shown in Fig. 3. Moreover, a heatmap (Fig. 4) of all the structures also implies the minimum energy and fluctuation rate of structure ID 1XQ8. In this regard, we selected this PDB structure for the further analysis.
In order to have a comprehensive grasp of the changes in sequence space, the PDB structures of \(\alpha \)-Synuclein and the mutated samples are constructed from the IntFOLD protein prediction server10. Based on the PDB models, structure networks are established. The structure network provides a new insight towards the nature and essential factors of structure-function dynamicity and folding process of a protein.
The community detection of a protein is crucial to understand evolution. In this study, every residue pair, obtained from structure network (as shown in Fig. 5a–f) belongs to the same cluster or in a different cluster with a strong association between them as shown in Fig. 6 for 5 mutation along with WT-human. This result also supports the DCA score. Additionally, the betweeness centrality of the mutations are shown in Supplementary Figure F1.
Furthermore, the residue present in modules are compared with cluster residue of Fig. 2b in which the mutated residues are available. The common residues present in both the cluster in the stretch of the NAC domain are mutated accordingly for three models in Fig. 5. The stretch are further consider for performing BLAST. The highest similarity sequence is identified and applied for establishing a PPI network.
The sequence with highest similarity of each sequential models after BLAST are used to understand the interaction with other protein by PPI network. From the BLAST result, it is found that mutation G51D-slow, E46K-fast and H50Q, A53T, A30P have more similarity with Pan paniscus, Pan troglodytes, Sus scrofa, Gorilla gorilla gorilla and Erythrocebus patas respectively. With the cutoff, 0.5 top 11 including SNCA (\(\alpha \)-Synuclein) are selected. The details list of associated proteins with their scoring depending on the shortest distance from SNCA are listed in Table 1. From this table it is clear that most of the associating proteins are remain common among organisms which depicts that, these proteins remain connected through the evolution of \(\alpha \)-Synuclein protein. It is concluded that protein may remain to interact but their connection scores change from one species to other.
Pathway semantic similarity
The six proteins are further studied in details to understand the role of pathways during evolution. From Reactome11 and Enrichr12,13 associated pathways of the six proteins are curated. We consider those pathways having more than two proteins including \(\alpha \)-Synuclein. The biological process and cellular component are considered for each pathway and a semantic similarity graph is constructed shown in Fig. 9. The semantic network of mutation G51D-slow and A30P are shown in Supplementary Figure F2. Each colour of the nodes represent a particular pathway and the edges between them depict the semantic scores. Moreover, the bold edges of the pathway graphs represent the connection weight between two pathways depending on the common biological terms. Additionally, we have included a Table 2 where we have validated few outcomes from literature (rest are shown in Supplementary Table T1). The table helps to conclude that this model provides an anonymous pipeline. There is no such model which targets the desired biological questionnaire best of our knowledge.
In Fig. 7, the certain courses of Van der Waals interaction are shown of the domain 40–95 where mutations mostly occur for three sequential models for he structure network of (a) WT-human, (b) G51D-slow, (c) E46K-fast other three mutations are reported in Supplementary Table T2. Along with the non-covalent force the hydrophobic profiling is represented through a graph in Fig. 8.
An adjacency matrix of each type is obtained during pathway semantic calculation. This matrix is used as input for PR algorithm to rank the pathways according to their importance. From the resulted rank list we noticed that apart from Parkinson disease and Alzhimer’s disease dopaminergic synapse, Amphetamine addiction and Cocaine addiction secured a higher position in all five mutations. However in the WT-type parkin ubiquitin proteasomal system pathway ranked high which is absent in both the mutations. A table of each mutation of alpha-synuclein along with their PR score is reported in Supplementary Table T3.
Discussion
The dynamic nature of the \(\alpha \)-synuclein lies on the NAC domain. As this domain has a higher propensity of aggregation, it can be considered as the most unstable region. As discussed before, the \(\beta \)-synuclein and \(\gamma \)-synuclein proteins are paralogous to \(\alpha \)-synuclein. However, \(\alpha \)-synuclein has two more domains. SE scores on each member from the synuclein family indicate that overall trait of the family is highly ordered. In the case of \(\alpha \)-synuclein, a certain range of disorder has been observed. This trait indicates the unstructured region of the \(\alpha \)-synuclein. Furthermore, from coupling analysis, 291 potential coupling pairs have been found. Among them, very few have fallen under NAC domain range. However, the mutated samples, namely, A53T, H50Q, G51D and E46K, are not known as part of the aggregation domain. So, the preliminary level of aggregations is somehow influenced by such mutations. Residual association within the protein is majorly controlled by the covalent and non-covalent interactions. In folding purposes, non-covalent forces such as weak van der Waals and ionic interactions play a vital role. Also, the hydrophobic residual shields guard the hydrophilic residues against misfolding. Following this theory, the abruption at non-covalent forces is not allowing the proper folding activity. In Fig. 7, the certain courses of non-covalent interaction have been shown at the affected targeted region at three sequential models. With that, hydrophobic profiling of the models are assist to describe the unstructured nature. In Fig. 8, the individual hydrophobic profiling of the WT and five mutants have been shown. Strikingly, the profiling of WT and E46k-fast are more similar including A30P. H50Q and G51D are also similar. Therefore, the folding pattern of these mutations are supposed to be similar. On other hand, generic extensive effects of the co-evolution at a group of residues have been shown through network \(G_{DCA}\) (shown in Fig. 2a). To make a comprehensive understanding, the sequence space information is mapped to individual structural information corresponding to the structure networks WT, G51D-slow and E46K-fast (shown in Fig. 5). From the colour modules of the \(G_{NMA}\), it is observed that the distribution of the clusters is giving proper evidence of the aggregation. As each networks has the same tree cutter threshold, the distribution of the modules should be unbiased. Following this trait, it has been observed that G51D-slow has a higher propensity of aggregation than the other five. Mohite et al.60 shows the strong effect of G51D phenotypical changes. In the rest of mutation, structure network can not provide much relevant information. All these monomeric models are highly prone to aggregation that reflect in the normal mode based weighted networks. Most of the residue nodes are conserved within fewer communities. From Fig. 2a, the coupling propensity of the NAC domain has shown. Interestingly, the distribution of co-evolutionary patches of the NAC domain is large. Comparing with individual structure networks, the individual lists of the affected residues due to an individual set of mutations are considered to be found the ancestral sequential traits. Eventually, the sequence traits are indicating the type of structural aberration from the WT. Subsequently, structural modifications in each case can provide a different list of top interaction partners. In Table 1, the list of interaction partners for the individual types are given. Few of the pathways are usually performed in every eukaryotic cell. However, the activities of the pathways are revised based on their metabolome. In Fig. 9, networks based on semantic similarity have been shown. From the networks, all the non-pathogenic pathways associated mostly with the common top neighbors of \(\alpha \)-synuclein (with few protein sample specific neighbors) are semantically associated with each other. BP has been considered for the similarity calculation. In WT-human, the non-pathogenic pathways have almost homogeneous association where the connection between blue and yellow node is slightly strong. In E46K-fast, this association between blue and yellow node is extremely strong. This information is comparable with the module distribution of the \(G_{NMA}\) where the propensity of aggregation is highly strong in G51D-slow.
The activities of Parkin-Ubiquitin Proteasomal System pathway and Ectoderm Differentiation are highly influential in the activity of the Dopaminergic Neurons which is key point of PD initiations61,62. However, strong association between the selected pathways has shown in semantic networks. From the pathway list, it has been observed that few pathways viz., Mitophagy, Dopaminergic synapses, MAPK signaling pathway, Amphetamine addiction are available in almost all prime mutations, and WT type alpha-synuclein. As per previous evidence, the association between the dopaminergic synapses and mitophagy can be explained. In63, the association of mitochondrial dysfunction and PD has been discussed. Dopaminergic Neurons are one of the highest consumers of ATP which explains the contribution of mitochondrial dysfunction in neuronal death64. Two perspectives so far can explain the relation clearly- firstly, the presence of the unpaired electrons accelerates ROS. ROS elevation facilitates aging by activating antioxidant enzymes as well as transporters. Subsequently, this also helps to relocate dopamine from the intercellular medium to synaptic vesicles. This whole process is associated with cell death where ROS levels increase throughout tissues including the brain for affected patients. Likewise, mitophagy involves the maintenance of a highly interconnected network throughout neurons. Balancing the mitochondrial activity in healthy cases needs the fusion and fission process65. Any sort of disruption main leads to aggregation and loss of direct movement. The common key between this activity is the mutations of mtDNA66 which disrupts the complex I and stops the mitophagy process. Interestingly, early-onset A53T mutation also follows the same path which leads to PD67. As per our results, most of the mutations are associated with mitophagy and dopaminergic synapses. Therefore, we are expecting that the mutations may have followed a similar path through mitochondrial dysfunction. However, two early-onset mutations i.e., A53T and A30P are observed to be involved in the MAPK signaling pathway. This explains the alternative path through MAPK kinase68. The dysfunction of such kinase leads during oxidative stress can responsible for mitochondrial stress and lead to mitochondrial dysfunction. Similarly, upregulating E46k is directly associated with the autophagy mechanism which controls the mitochondrial fission69. As per our study, some of the nodes from the pathway semantic networks (shown in Fig. 9) can be explained through previous researches. The other nodes from each network are expected to be partially associated with the regulation of the influential nodes. In Table, results have been validated through existing literature where we focused on the associated interaction partners of the mutants. Hence, the semantic networks can be considered as a summarization of the mutant specific molecular mechanism.
The pathway semantic graph of (a) WT-Human, (b) A53T, (c) E46K-fast and (d) H50Q-late onset mutations of Alpha-synuclein. In the graph, the color nodes represent a particular pathway associated with the mutation type and the edges represents the weighted connection between the pathways. The bold edges indicate the higher association between those pathways.
Aging is one of the prime reasons for losing the cognitive senses. WT type Asyn is usually coming up with cellular precipitation which leads to dementia. However, the early onset mutation can trigger the precipitation within 30–40 years of the age range. Mutational subtypes i.e., A30P, A53T, G51D-slow and E46K-fast, are studied to unveil the functional and molecular orchestration during such mutations. The associated list of pathways individually for these samples has been considered. The pathways are connected through the sharing of interacting neighbors. Here, the list of protein interacting neighbors is modifying with mutations which brings us the distinct list of pathways as well as semantic connectivity. Twelve networks are based on two cognitive parameters i.e., Cellular component and, Biological process. From the networks, all the associated pathways are clearly sharing almost the same subcellular localization. In WT type, the pathways excluding Parkinson’s Disease and Alzheimer’s Disease are associated with two distinct types of the function where some pathways are associated with prevention from unusual activity within the neurons whereas the rest of the pathways are associated with the pathogenic progression of the diseases. For example, monoamine transport systems are neurotransmitters that prevent the extracellular vesicles of the neuronal cells from excess dopamine, serotonin, noradrenaline, etc. Dopaminergic pathway, found in the list, is one of the prime pathways from the neuronal cells which helps to control the cognitive senses. Monoamine transport systems are the key modulator of the dopaminergic pathways. It can also control the effects of nanomolar elements such as cocaine, amphetamine, etc. On the other hand, pathways like Parkin-Ubiquitin Proteasomal System regulates the misfolding of the proteins which are further responsible for the pathogenic progression of the diseases such as PD. The pathway semantic networks (shown in Fig. 9) show the connectivity within the pathways based on sharing list of biological processes and cellular components. In the BP based network, the Parkin-Ubiquitin Proteasomal System pathway is the highest-ranked pathway as per PR outcomes. This shows the influence of the pathway in the network which increases the possibility of the Asyn misfolding.
The pathways of the selected mutations Viz., G51D-slow and E46K-fast, are mostly associated with external nanomolar elements. More elaborately, dopaminergic pathways are strongly associated with the proteins which are differentially regulated due to cocaine addiction, alcoholism, amphetamine addiction70,71,72. Amphetamine, cocaine are known neurotransmitters. Although appropriate etiology associated with amphetamine is not known, amphetamines are initially used to treat PD73. However, some recent studies have shown that long use of amphetamine-type stimulators increases the risk of PD. The studies explain that amphetamine-type stimulators bind in the intrinsically disordered regions of the Asyn protein and facilitates the molecular mechanism of the aggregation. The studies also suggest that it promotes the post-translational modification of the Asyn directly or indirectly. Interestingly, the involvement of the Asyn mutants specially, H50Q and G51D have largely been observed in PD cases who are long users of such stimulators74,75. The decreasing number of dopaminergic neurons at Substantial nigra is also considered as one of the key reasons76. Also, the mutations can inhibit the activities of the neurotransmitters such as monoamine transport systems77. Due to that reason, the excess hormones in the cellular systems cannot be removed. As per the pathway semantic networks, E46K-fast shows the aforementioned strong connectivity. However, the amphetamine addiction pathway is disconnected in terms of BP for G51D-slow.
We have included three more mutations to explain the pd etiology clearly. From the pathway list, it has been observed that few pathways viz., Mitophagy, Dopaminergic synapses, MAPK signaling pathway, Amphetamine addiction are available in almost all prime mutations, and WT type alpha-synuclein. As per previous evidence, the association between the dopaminergic synapses and mitophagy can be explained. In63, the association of mitochondrial dysfunction and PD has been discussed. Dopaminergic Neurons are one of the highest consumers of ATP which explains the contribution of mitochondrial dysfunction in neuronal death64. Two perspectives so far can explain the relation clearly–firstly, the presence of the unpaired electrons accelerates ROS. ROS elevation facilitates aging by activating antioxidant enzymes as well as transporters. Subsequently, this also helps to relocate dopamine from the intercellular medium to synaptic vesicles. This whole process is associated with cell death where ROS levels increase throughout tissues including the brain for affected patients. Likewise, mitophagy involves the maintenance of a highly interconnected network throughout neurons. Balancing the mitochondrial activity in healthy cases needs the fusion and fission process65. Any sort of disruption main leads to aggregation and loss of direct movement. The common key between this activity is the mutations of mtDNA66 which disrupts the complex I and stops the mitophagy process. Interestingly, early-onset A53T mutation also follows the same path which leads to PD67. As per our results, most of the mutations are associated with mitophagy and dopaminergic synapses. Therefore, we are expecting that the mutations may have followed a similar path through mitochondrial dysfunction. However, two early-onset mutations i.e., A53T and A30P are observed to be involved in the MAPK signaling pathway. This explains the alternative path through MAPK kinase68. The dysfunction of such kinase leads during oxidative stress can responsible for mitochondrial stress and lead to mitochondrial dysfunction. Similarly, upregulating E46k is directly associated with the autophagy mechanism which controls the mitochondrial fission69.
Outcomes of the PR algorithm shows affinity of all the mutations with Amphetamine addictions, Cocaine addictions, dopaminergic synapses, etc. Also, H50Q and A30P have synaptic vesicles among the top ranking pathways. These observation supports the etiology associated with mitochondrial dysfunction. Interestingly, few pathways which are mostly associated with dopaminergic disorientation, are identified under A53T mutation.
Conclusion
From this study, the sequential variation of \(\alpha \)-synuclein is observed depending on six mutational conditions, considering the structural consequences. The objective of the study is also to observe changes in pathways due to structural aberration. Among the mutations, the pathway semantic networks of A53T and G51D show maximum involvement of pathways. Also, few pathways play vital roles in mutational perspectives viz., Amyloid fiber formation, Parkin-Ubiquitin Proteasomal System pathway, Ectoderm Differentiation, Mitophagy, MAPK signaling pathway, synaptic vesicles, etc. Finally, the proposed frame can provide a comprehensive outlook on the mutation mediated structural aberrations and their affects on functional pathways.
Method
This study aims to understand the structural changes along with the pathway semantics of \(\alpha \)-Synuclein concerning evolution. The detailed method is described in Fig. 10.
Sequence space analysis
Primarily, the Synuclein family is considered to unveil the family trait with the help of Shannon entropy (SE) calculation. It is evident that Shannon entropy is directly proportional to the rate of disorder. This implicates if the entropic score increases it signifies the higher disorderness of a protein. In78, 2.9 is considered as the threshold of Shannon entropy score, i.e. sequence with score less than 2.9 is treated as ordered sequence and vice-versa. This idea is utilized in this study and applied on each protein sequences of Synuclein family. Shannon entropy is calculated as follows:
where \(H_i\) was the probability of given amino acids and L was the number of letters in a sequence. The summation run over the 20 residues that normally were present in a protein sequence. The probability \(H_i\) represent the composition of the consensus sequence. So the entropy range lied between 0 and the \(log_2(20) = 4.32\).
Moreover, multiple sequence alignment is performed of the protein sequences belong to PF01387. It is noticed that multiple sequence alignment have a significant role in the field of structure and function analysis of biological sequences. Due to this, full alignment FASTA format sequences were collected, which comprises of tree orderings and all lower case letters including dashes to indicate the gaps. Additionally, a consensus sequence is generated from the aligned sequences. The consensus sequence is a set of amino acids with their occurrence frequency. These frequencies indicate the signature amino acids remain conserved throughout the evolution. In this regard, Shannon entropy is calculated for the consensus sequence also.
The aligned sequences are further considered for DCA. This is a statistical framework which possesses the idea of direct co-evolution coupling among the residue pairs. The drawback of the Mutual Information (MI) is, unable to extricate direct correlation from indirect ones. This problem is easily solved by Direct Information (DI) theory. Depending on the DI theory, DCA is calculated. Hence, the computation of the DCA score of an aligned sequence indicates how directly the selected coinciding residues are coupled with each other and contributed toward evolution. The DI score is defined as:
Here, \(F_{st}^{(dir)}\) represents reweighted frequency counts to introduce two residues for DI. where \(F(X,X')\) is considered as joint probability, and F(X) and \(F(X')\) are individual probability. \(F_{s}(X)\) and \(F_{t}(X')\) are for amino acid type A at \(s\)th position and similarly B at \(t\)th position.
The cascading effect of the co-evolution is not conserved in a couple of residues. It is highly possible that mutational changes at a residual point can affect a distant residue by means of a cascading effect. To understand such an effect, a weighted network \(G_{DCA}\) has been defined considering DI score between residue. The weighted undirected network \(G_{DCA}\) = (\(V_{res}\), \(E_{DI}\)) where \(V_{res}\) are set of nodes, consist of residue whereas \(E_{DI}\) are weighted edges, consists of DI score between the residues. Subsequently, the color modules have been formed applying Girvan-Newman algorithm79.
Root mean square fluctuation
The information related to sequence space or structure space individually is not enough to understand the changes. For the study, the WT type protein is taken from Protein Data Bank where the respective PDB id is 1XQ8. Due to the dynamic structure and frequent mutations, the structure possess lowest energy is considered. In this regard, Root Mean Square Fluctuation (RMSF) is performed to measure the particle deviation. In RMSF, a mean over time is considered for a residue r at the current position and some reference position. The definition of the RMSF is given in Equation 4.
Where S is time over which the mean has been taken for reference position of the particle i , \(R_i^{re}\). The RMSF has been observed based on the reference position of the particle i over time.0
Structure network analysis
To grasp this observation we have built the structure of those sequences. These complex models are analyzed with the help of a structure network80. The interaction between the elements of the networks is represented through nodes and edges. Generally, a secondary structure and folding arrangement mechanism are used to understand the structure of a protein. Another promising methodology for understanding the structure is through network. The equation is represented below.
\(E_c\) is the threshold of interaction strength, the default value is 4%. Here, \(sc_{ab}\) was the number of side chain atom pairs of residues a and b. \(sc_a\) and \(X_b\) were the normalization factor for residues types a and b81.
In this paper, depending on the normal mode analysis (NMA) a correlation matrix is obtained. The matrix is applied to establish a full residue weighted network \(G_{NMA}\) = (\(V_{res}\), \(E_{NM}\)) where \(V_{res}\) is a set of nodes representing residues whereas \(E_{NM}\) is set of weighted edges where weights are obtained from correlation matrix. Consequently, the network is split into a highly correlated coarse-grained community cluster network by using Girvan-Newman79 clustering method where the highly interacting residues were clumped together in the clusters with a threshold value 0.7.
Pathway semantic
Furthermore, mutation G51D and E46K i.e., slow and accelerating mutation on early onset are considered for further analysis. During mutation, co-varying residue cluster according to the DCA score are compared with the structure network module. The residues common in both the cases and also belong with the residue number 51 and 46 are mutated accordingly. As it is known that residue range 61–95 is the NAC domain of \(\alpha \)-Synuclein, the residue stretch from 40 to 95 is utilized for performing Basic Local Alignment Tool (BLAST). The protein sequences are matched with the sequence database and statistical significance is calculated for the matched areas. The highest similarity sequence is selected and Protein-Protein Interaction (PPI)82 is executed to detect the associated proteins. PPI is carried out for \(\alpha \)-Synuclein of three species with a threshold value of 0.5. Pathways responsible for at least two proteins including \(\alpha \)-Synuclein are considered. Here the semantic similarity is calculated over a set of a biological process responsible for the selected pathways. Wang method83 is applied to evaluate the similarity established on graph-like structure of gene ontology (GO). The aggregated contribution is done by the semantic value of GO term T to the terms in \(DAG_T\) which is semantic of GO term T is firstly defined in Wang method. GO terms closer to T in \(DAG_T\) implies more contribution toward it semantics. Hence, it is defined that the contribution of GO term p to the semantics of T as the \(S{\text{-}}value\) of GO term p related to GO term T. For whatever term of p in \(DAG_T\), the \(S{\text{-}}value\) associated GO term T, \(S_T(p)\) is calculated as:
Here \(C_e\) is defined as semantic contribution factor for the edge \(e \in E_T\) linking GO term p with its child term \(p'\). After calculating the \(S{\text{-}}value\) for the GO term in \(DAG_T\), the semantic value of GO term T, SV(T) is defined as:
For two given GO term such as T and Q, semantic similarity is calculated between them is as follows:
Here \(S_T(P)\) is the \(S{\text{-}}value\) of GO term t related to term T and \(S_Q(p)\) is the \(S{\text{-}}value\) of GO term p related to term Q.
Moreover, based on the semantic similarity of GO terms, Best-Match Average (BMC)84 strategy is performed to compute semantic similarity among sets of GO terms associated with the protein associated with a particular pathway and column, which defined as:
where gene G1 annotated by GO terms sets \(GO1=(go_{11},go_{12} \ldots go_{1i})\) and G2 annotated by \(GO2=(go_{21},go22 \ldots go_{2j})\).
Ranking the pathways
The pathway semantic similarity graphs reveal the semantic strength between two pathway. In order to understand the importance of the pathways PageRank (PR)85 Algorithm introduced by Google is applied on the resulted networks. This algorithm used probability distribution based on the weight among different nodes. In following equation 9, pathways are represented as nodes and the edges are weighted. The node rank has been defined as:
where the rank of node n is relied on the PR values for each connected node m \(\in \) A\(_{n}\), divided by E(m), edges from node n. Therefore, according to the the PR values the pathways are ranked based on their importance in the network.
References
Pujols, J. et al. Small molecule inhibits \(\alpha \)-synuclein aggregation, disrupts amyloid fibrils, and prevents degeneration of dopaminergic neurons. Proc. Natl. Acad. Sci. USA 115, 10481–10486 (2018).
Stefanis, L. \(\alpha \)-Synuclein in Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2, a009399 (2012).
Venda, L. L., Cragg, S. J., Buchman, V. L. & Martins, R. W. \(\alpha \)-Synuclein and dopamine at the crossroads of Parkinson’s disease. Trends Neurosci. 33, 559–568 (2010).
Laura, M., Giuseppe, N., Lesley, C., Michele, V. & Annalisa, P. Functional interactions as a survival strategy against abnormal aggregation. FASEB J. 25, 45–54 (2010).
Ozansoy, M. & Basa, A. N. The central theme of Parkinson’s disease: \(\alpha \)-Synuclein. Mol. Neurobiol. 47, 460–465 (2013).
Flagmeier, P. et al. Mutations associated with familial Parkinson’s disease alter the initiation and amplification steps of \(\alpha \)-synuclein aggregation. Proc. Natl. Acad. Sci. USA 113, 10328–10333 (2016).
Sakai, R. et al. E46K mutant \(\alpha \)-synuclein is more degradation resistant and exhibits greater toxic effects than wild-type \(\alpha \)-synuclein in Drosophila models of Parkinson’s disease. PLoS ONE 14, e0218261 (2019).
Liberles, J. S. Evolution of structurally disordered proteins promotes neostructuralization. Mol. Biol. Evol. 28, 59–62 (2011).
Oliveira, G. A. P. & Silva, J. L. Alpha-synuclein stepwise aggregation reveals features of an early onset mutation in Parkinson’s disease. Commun. Biol. 2, 1–13 (2019).
McGuffin, L. J. et al. Intfold: An integrated server for modelling protein structures and functions from amino acid sequences. Nucl. Acids Res. 43, W169–W173 (2015).
Croft, D. et al. Reactome: A database of reactions, pathways and biological processes. Nucl. Acids Res. 39, D691–D697 (2011).
Chen, E. Y. et al. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 1–14 (2013).
Kuleshov, M. V. et al. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucl. Acids Res. 44, W90–W91 (2016).
Cresto, N. & Gardier, C. The unlikely partnership between LRRK2 and \(alpha \)-synuclein in Parkinson’s disease. Eur. J. Neurosci. 49, 339–363 (2019).
Torres, C. A. M. et al. The role of alpha-synuclein and other Parkinson’s genes in neurodevelopmental and neurodegenerative disorders. Int. J. Mol. Sci. 21, 1–20 (2020).
Yedlapudi, D. et al. Targeting alpha synuclein and amyloid beta by a multifunctional, brain-penetrant dopamine D2/D3 agonist D-520: Potential therapeutic application in Parkinson’s disease with dementia. Sci. Rep. 9, 1–12 (2019).
Waxman, E. A. & Giasson, B. I. Molecular mechanisms of \(\alpha \)-synuclein neurodegeneration. Biochim. Biophys. Acta 1792, 616–624 (2009).
Xiong, Y., Dawson, T. M. & Dawson, V. L. Models of LRRK2 associated Parkinson’s disease. Adv. Neurobiol. 14, 163–191 (2017).
O’Hara, D. M., Pawar, G., Kalia, S. K. & Kalia, L. V. LRRK2 and \(\alpha \)-synuclein: Distinct or synergistic players in Parkinson’s disease?. Front. Neurosci 14, 577 (2020).
Buchman, V. L. & Ninkina, N. Modulation of alpha-synuclein expression in transgenic animals for modelling synucleinopathies-is the juice worth the squeeze?. Neurotox. Res. 14, 329–341 (2015).
Indrieri, A., Pizzarelli, R., Franco, B. & Leonibus, E. D. Dopamine, alpha-synuclein, and mitochondrial dysfunctions in parkinsonian eyes. Front. Neurosci 14 (2019).
Nuytemans, K. et al. Genetic etiology of Parkinson disease associated with mutations in the SNCA, PARK2, PINK1, PARK7, and LRRK2 genes: A mutation update. Hum. Mutat. 31, 763–780 (2010).
Eschbach, J. & Danzer, K. M. \(\alpha \)-Synuclein in Parkinson’s disease: Pathogenic function and translation into animal models. Neurodegener. Dis. 14, 1–17 (2014).
Kim, E. K. & Choi, E. J. Pathological roles of MAPK signaling pathways in human diseases. Biochim. Biophys. Acta Mol. Basis Dis. 1802, 396–405 (2010).
Tong, Y. & Shen, J. \(\alpha \)-Synuclein and LRRK2: Partners in crime. Neuron 64, 771–773 (2009).
Martin, I., Dawson, V. L. & Dawson, T. M. The impact of genetic research on our understanding of Parkinson’s disease. Prog. Brain Res. 183, 21–41 (2010).
Chen, J. et al. Phosphorylation of Parkin at serine 131 by p38 MAPK promotes mitochondrial dysfunction and neuronal death in mutant A53T \(\alpha \)-synuclein model of Parkinson’s disease. Cell Death Dis. 9, 1–15 (2018).
Si, X., Pu, J. & Zhang, B. Structure, distribution, and genetic profile of \(\alpha \)-synuclein and their potential clinical application in Parkinson’s disease. J. Mov. Disord. 10, 69–79 (2017).
Kuhlmann, N. & Milnerwood, A. J. A critical LRRK at the synapse? The neurobiological function and pathophysiological dysfunction of LRRK2. Front. Mol. Neurosci. 13, 1–19 (2020).
Cacabelos, R. Parkinson’s disease: From pathogenesis to pharmacogenomics. Int. J. Mol. Sci. 18, 1–28 (2017).
Cacabelos, R. The genetics of Parkinson disease. J. Geriatr. Psychiatry Neurol. 13, 228–242 (2010).
Lesage, S. & Brice, A. Parkinson’s disease: From monogenic forms to genetic susceptibility factors. Hum. Mol. Genet. 18, R48–R59 (2009).
Twohig, D. & Nielsen, H. M. \(\alpha \)-Synuclein in the pathophysiology of Alzheimer’s disease. Mol. Neurodegener. 14, 1–19 (2019).
Weber, O. C. & Uversky, V. N. Insights into the molecular mechanisms of Alzheimer’s and Parkinson’s diseases with molecular simulations: Understanding the roles of artificial and pathological missense mutations in intrinsically disordered proteins related to pathology. Int. J. Mol. Sci 19, 1–62 (2018).
Spillantini, M. G. & Goedert, M. Neurodegeneration and the ordered assembly of \(\alpha \)-synuclein. Cell Tissue Res. 373, 137–148 (2018).
Siddiqui, I. J., Pervaiz, N. & Abbasi, A. A. The Parkinson disease gene SNCA: Evolutionary and structural insights with pathological implication. Sci. Rep. 6, 1–11 (2016).
Meade, R. M., Fairlie, D. P. & Mason, J. M. Alpha-synuclein structure and Parkinson’s disease-lessons and emerging principles. Mol. Neurodegener. 14, 1–14 (2019).
Liu, G., Aliaga, L. & Cai, H. \(\alpha \)-synuclein, LRRK2 and their interplay in Parkinson’s disease. Future Neurol. 7, 145–153 (2014).
Vacchi, E., Kaelin-Lang, A. & Melli, G. Tau and alpha synuclein synergistic effect in neurodegenerative diseases: When the periphery is the core. Int. J. Mol. Sci. 21, 1–23 (2020).
Marcoa, I. I. et al. E46K \(\alpha \)-synuclein pathological mutation causes cell-autonomous toxicity without altering protein turnover or aggregation. PNAS 114, E8274–E8283 (2017).
Aryal, S., Skinner, T., Bridges, B. & Weber, J. T. The pathology of Parkinson’s disease and potential benefit of dietary polyphenols. Molecules 25, 1–27 (2020).
Ma, S. X. et al. Complement and coagulation cascades are potentially involved in dopaminergic neurodegeneration in \(\alpha \)-synuclein-based mouse models of Parkinson’s disease. bioRxiv 1–49 (2020).
Cannon, J. R. et al. Expression of human E46K-mutated \(\alpha \)-synuclein in BAC-transgenic rats replicates early-stage Parkinson’s disease features and enhances vulnerability to mitochondrial impairment. Exp. Neurol. 240, 44–56 (2013).
Butler, B. et al. Dopamine transporter activity is modulated by \(alpha \)-synuclein. J. Biol. Chem. 290, 29542–29554 (2015).
Hernandez, S. M., Tikhonova, E. B. & Karamyshev, A. L. Protein–protein interactions in alpha-synuclein biogenesis: New potential targets in Parkinson’s disease. Front. Aging Neurosci. 12, 1–8 (2020).
Selvaraja, S. & Piramanayagam, S. Impact of gene mutation in the development of Parkinson’s disease. Genes Dis. 6, 120–128 (2019).
Zhong, X. L. et al. A genome-wide association study of \(\alpha \)-synuclein levels in cerebrospinal fluid. Neurotoxicity Res. 35, 41–48 (2019).
Lesage, S. et al. Genetic and phenotypic basis of autosomal dominant Parkinson’s disease in a large multi-center cohort. Front. Neurol. 11, 1–11 (2020).
Vasili, E., Dominguez-Meijide, A. & Outeiro, T. F. Spreading of \(\alpha \)-synuclein and tau: A systematic comparison of the mechanisms involved. Front. Mol. Neurosci. 12, 1–23 (2019).
Obergasteiger, J., Frapporti, G., Pramstaller, P. P., Hicks, A. A. & Volta, M. A new hypothesis for Parkinson’s disease pathogenesis: GTPase-p38 MAPK signaling and autophagy as convergence points of etiology and genomics. Mol. Neurodegener. 13, 40 (2018).
Pei, Y. & Maitta, R. W. Alpha synuclein in hematopoiesis and immunity. Heliyon 5, e02590 (2018).
Mor, D. E., Daniels, M. J. & Ischiropoulos, H. The usual suspects, dopamine and alpha-synuclein, conspire to cause neurodegeneration. Mov. Disord. 34, 167–179 (2019).
Mash, D. C. et al. Cocaine abusers have an overexpression of alpha-synuclein in dopamine neurons. J. Neurosci. 23, 2564–2571 (2019).
Pathak, D. et al. Loss of \(\alpha \)-synuclein does not affect mitochondrial bioenergetics in rodent neurons . Eneuro 4 (2019).
Bernal-Conde, L. D. et al. Alpha-synuclein physiology and pathology: A perspective on cellular structures and organelles. Front. Neurosci. 13, 1399 (2020).
Poehler, A. M. & Xiang, W. Autophagy modulates SNCA/\(alpha \)-synuclein release, thereby generating a hostile microenvironment. Autophagy 13, 2171–2192 (2014).
Cartier, A. E. et al. Differential effects of UCHL1 modulation on alpha-synuclein in PD-like models of alpha-synucleinopathy. PLoS ONE 7, e34713 (2012).
Protter, D., Lang, C. & Cooper, A. A. \(\alpha \)Synuclein and mitochondrial dysfunction: A pathogenic partnership in Parkinson’s disease?. Parkinson’s Dis. 2012, (2012).
Gao, F. et al. Mitophagy in Parkinson’s disease: Pathogenic and therapeutic implications. Front. Neurol. 8, 527 (2017).
Mohite, G. M. et al. Parkinson’s disease associated \(\alpha \)-synuclein familial mutants promote dopaminergic neuronal death in Drosophila melanogaster. ACS Chem. Neurosci. 9, 2628–2638 (2018).
Lim, K. L. & Tan, J. M. Role of the ubiquitin proteasome system in Parkinson’s disease. BMC Biochem. 8, 1–10 (2007).
Jadalannagari, S. et al. Ectodermal differentiation of Wharton’s jelly mesenchymal stem cells for tissue engineering and regenerative medicine applications. Tissue Eng. Part B Rev. 21, 314–322 (2015).
Arduino, D. M., Esteves, A. R., Swerdlow, R. H. & Cardoso, S. M. A cybrid cell model for the assessment of the link between mitochondrial deficits and sporadic Parkinson’s disease. Methods Mol. Biol. 1265, 415–424 (2015).
Haddad, D. & Nakamura, K. Understanding the susceptibility of dopamine neurons to mitochondrial stressors in Parkinson’s disease. FEBS Lett. 589, 3702–3713 (2015).
Chen, H. & Chan, D. C. Mitochondrial dynamics-fusion, fission, movement, and mitophagy-in neurodegenerative diseases. Hum. Mol. Genet. 18, R169–R176 (2009).
Giannoccaro, M. P., Morgia, C. L., Rizzo, G. & Carelli, V. Mitochondrial DNA and primary mitochondrial dysfunction in Parkinson’s disease. Mov. Disord. 32, 346–363 (2017).
Devoto, V. M. P. et al.\(\alpha \)-Synuclein control of mitochondrial homeostasis in human-derived neurons is disrupted by mutations associated with Parkinson’s disease. Sci. Rep. 7, 1–13 (2017).
Bohush, A., Niewiadomska, G. & Filipek, A. Role of mitogen activated protein kinase signaling in Parkinson’s disease. Int. J. Mol. Sci. 19, 1–17 (2018).
Yan, J. Q., Yuan, Y. H., Chu, S. F., Li, G. H. & Chen, N. H. E46K mutant \(\alpha \)-synuclein is degraded by both proteasome and macroautophagy pathway. Molecules 23, 1–13 (2018).
Illes, A. et al. Open Access Dynamic interaction of genetic risk factors and cocaine abuse in the background of Parkinsonism—a case report. BMC Neurol. 19, 1–6 (2019).
Bettiol, S. S., Rose, T. C., Hughes, C. J. & Smith, L. A. Alcohol consumption and Parkinson’s disease risk: A review of recent findings. J. Parkinson’s Dis. 5, 425–442 (2015).
Christine, C. W., Garwood, E. R., Schrock, L. E., Austin, D. E. & McCulloch, C. E. Parkinsonism in patients with a history of amphetamine exposure. Mov. Disord. 25, 228–231 (2010).
Parkes, J. D. et al. Amphetamines in the treatment of Parkinson’s disease. J. Neurol. Neurosurg. Psychiatry 38, 232–237 (1975).
Biagioni, F. et al. Methamphetamine persistently increases alpha-synuclein and suppresses gene promoter methylation within striatal neurons. Brain Res. 1719, 157–175 (2019).
Hayakawa, H. & Nakatani, R. Structurally distinct \(alpha \)-synuclein fibrils induce robust parkinsonian pathology. Mov. Disord. 35, 256–267 (2020).
Rumpf, J. J., Albers, J., Fricke, C., Mueller, W. & Classen, J. Structural abnormality of substantia nigra induced by methamphetamine abuse. Mov. Disord. 32, 1784–1788 (1975).
Oaks, A. W. & Sidhu, A. Synuclein modulation of monoamine transporters. FEBS Lett. 585, 1001–1006 (2011).
Romero, P. et al. Sequence complexity of disordered protein. Proteins 42, 38–48 (2000).
Girvan, M. & Newman, M. E. J. Community structure in social and biological networks. Proc. Natl. Acad. Sci. USA 99, 7821–7826 (2002).
Greene, L. H. Protein structure networks. Brief. Funct. Genomics 11, 469–478 (2012).
Brinda, K. V. & Vishveshwara, S. A network representation of protein structures: Implications for protein stability. Biophys. J. 894159–4170, 4159–4170 (2005).
Makino, T. & Gojobori, T. Evolution of protein–protein interaction network. Genome Dyn. 3 (2007).
Wang, J. Z. et al. A new method to measure the semantic similarity of GO terms. Bioinformatics 23, 1274–1281 (2007).
Pesquita, C. et al. Metrics for GO based protein semantic similarity: a systematic evaluation. In BMC Bioinformatics, Vol. 9 (2008).
Page, L., Brin, S., Motwani, R. & Winograd, T. The pagerank citation ranking: Bringing order to the web. Tech. Rep., Stanford InfoLab (1999).
Acknowledgements
The work of SS and AD is supported by DST-INSPIRE fellowship. We would also like to thank the reviewers for their precious comments which enhance the quality of the article.
Author information
Authors and Affiliations
Contributions
S.S. and A.D. conceived and conducted the experiment. S.S., A.D., and U.M. analysed the results and drafted the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sen, S., Dey, A. & Maulik, U. Studying the effect of alpha-synuclein and Parkinson’s disease linked mutants on inter pathway connectivities. Sci Rep 11, 16365 (2021). https://doi.org/10.1038/s41598-021-95889-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-95889-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.