Abstract
Alien impact on native diversity could be a function of both the relatedness of alien species to native community and resources availability. Here, we investigated whether alien plants expand or decrease the functional and phylogenetic space of native plant communities, and how this is affected by alien relatedness to natives and by resources availability. We used a trait-environment dataset of 33 alien and 130 native plants in 83 pairs of invaded and non-invaded plots, covering a gradient of soil resources (organic matter-nitrogen) in Saint-Katherine-Protectorate, Egypt. First, we compared the changes in native composition and calculated alien relatedness to natives within each pair of plots. Second, we tested the effects of resources availability and relatedness on the magnitude of alien impact (defined as a change in native diversity). We found that native composition was phylogenetically less but functionally more diverse in invaded plots compared to non-invaded ones. Moreover, in resources-rich plots, dissimilar aliens to natives significantly increased native diversity, while in resource-limited ones, similar aliens to natives declined native diversity. These results suggest that the assessment of alien impacts in arid-regions is significantly linked to resources-availability and relatedness to natives. Hence, future studies should test the generality of our findings in different environments.
Similar content being viewed by others
Introduction
Invasions by alien plant species are increasingly impacting native biodiversity, communities and ecosystem functioning1,2,3. Invasive alien plants can cause a local loss of native species richness and change the dominance relationships in a community, and consequently alter ecosystem processes1,4,5,6,7. The awareness of these potential consequences of biological invasions on native biodiversity is increasing. However, still little is known about how biological invasions affect taxonomic, phylogenetic and functional aspects of biodiversity. Furthermore, how the impacts depend on the functional and phylogenetic relatedness (similarity or dissimilarity of alien species to native species), and how the impacts vary along gradients of abiotic environmental conditions deserves more research.
Alien species are likely to change the richness, functional and phylogenetic diversity of the native resident community. The direction of this change could depend on the ecological similarity or dissimilarity between aliens and natives, and on resources availability8,9,10,11. In this sense, Darwin's naturalization conundrum (DNC; Darwin 1859), which includes both pre-adaptation and naturalization as important factors for alien success, has encountered a considerable amount of attention12,13,14,15. According to the pre-adaptation hypothesis, if an alien species is phylogenetically closely related to native resident species, it will likely share similar traits and occupy a similar niche as the closely-related species due to the strong effect of environmental filtering12,16. This filtering restricts the range of trait values of the alien species to be more similar to those of natives17,18,19,20,21,22,23. For example, in resource-limited environments, a pattern of aliens co-occurring with functionally similar natives is expected. This increasing similarity might cause the alien species to replace natives and to occupy a portion of the functional and possibly of the phylogenetic space originally occupied by the native species they replaced, leading to the contraction of the functional and phylogenetic diversity of the native resident communities24,25.
According to Darwin’s Naturalization hypothesis (DNH), if the alien species is distantly related to resident natives, it will occupy a distinct niche and its competitive effect is assumed to be low with resident natives. Under this scenario, alien species can fill vacant niches that are not used by natives leading to niche differentiation. For example, in resource-rich environments, exploitative aliens might be more likely to naturalize if they are phylogenetically and functionally dissimilar to the native species, because then they can exploit unfilled ecological niches26. Such alien species could expand the native phylogenetic and functional diversity, compared to non-invaded communities27, and provide a novel suit of traits and evolutionary origins to the invaded communities28,29,30.
Significantly, competitive interactions between aliens and natives within low and high resource environments can also cause the dominance of clades of native species bearing traits related to greater competitive abilities31,32. This dominance arises from the relative differences in the competitive abilities among alien and native species11,31,33,34,35,36. Such competition can select strong competitors among the environmentally adapted aliens that can coexist within local native communities, leading to a decline in native richness and abundance37,38,39,40,41,42. For example, natives are expected to be eliminated by aliens if natives are weak competitors and overlap too much in their soil preferences with aliens, and vice-versa28,31. This phenomenon is linked to the tendency of alien species to become dominant in a plant community outside its native range and to locally replace native species37,38,39.
In this study, we investigated (1) whether the presence of alien plants changes the richness and abundance of natives, (2) whether alien plants occupy a portion of the functional and phylogenetic space of native communities, (3) whether alien plants can expand the trait space and phylogenetic (evolutionary) space to the native community, (4) to what extent the impact of alien plants on native communities is affected by soil resources availability and (5) by alien relatedness to natives.
We expect an increase in the richness, functional and phylogenetic space of invaded-native communities compared to non-invaded ones in resources rich environments. This increase could be attributed to the presence of alien plants that tend to be functionally and phylogenetically dissimilar to natives (i.e. Darwin’s Naturalization Hypothesis). This dissimilarity enables alien plants to employ different strategies of resource use (i.e. niche differentiation and fitness differences), exploit resources unused by natives, and fill niches unoccupied by natives. As a consequence of dissimilarity, a competitive exclusion10,31,43,44 may arise to eliminate alien /native species that are too similar in their trait values45,46,47.
By contrast, in limited resources environments, we expect a decline in the native diversity measures (richness, functional and phylogenetic matrices) as well as the phylogenetic signal of native traits in invaded communities compared to non-invaded ones. This could be attributed to the negative impact of alien plants on resident natives if aliens and natives are functionally and phylogenetically similar (i.e. Darwin’s Pre-adaptation Hypothesis). In this case, aliens and natives have similar fitness and equivalent competitive effects or sharing similar niches due to the effect of environmental filtering that will cause alien plants to occupy a portion of the phylogenetic and functional space originally occupied by natives.
Methods
Vegetation plots, species inventories and soil data
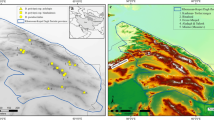
Field surveys were carried out during the spring and summer seasons (March to July) of 2018 in Saint Katherine Protectorate (SKP), South Sinai, Egypt (Supplementary Appendix A). The diversity in geomorphological and geological structures of SKP resulted in six types of microhabitats, namely Wadis (valleys), Terraces, Slopes, Gorges, Cliffs, Farsh (basins) and Caves (SKP Management Plan 200348,49). Wadis are the most common microhabitats in the current study area, and act as drainage systems collecting water from catchment areas for plant growth. The wadis in SKP are very narrow, have very steep slopes, short in length and occur at higher elevations, ranging from 1190 to 1900 m (a.s.l). Across the entire study area, we selected 83 plots of 10 m2 that were invaded by at least one alien plant species. The maximum number of alien species in a plot was three. Close to each invaded plot, at approximately 10 m distance, we set up a 10 m2 plot with similar vegetation but with no invader. This resulted in 83 pairs of invaded and non-invaded plots. We identified all plants to species (33 aliens and 130 natives), and we counted the numbers of alien and native individuals per plot (Supplementary Appendix A, Supplementary Table A1).
To describe the abiotic environment and resource availability, we obtained for each plot measurements of soil moisture, soil nitrogen, and organic matter content. Soil moisture measurements were taken directly in the field in the early morning, using soil hygrometer. Three soil samples were collected at randomly chosen positions within each plot and air-dried to constant mass. Then, soil water extracts at 1:5 were prepared for the determination of soil nitrogen and organic matter content. The percentage of organic matter was calculated as the difference between total C and CaCO3%50. The total concentration of available nitrogen (mg/L) was calculated using a CHN analyser (EA1108, Carlo Erba Instruments, USA) and standard methods51. These measures were highly correlated; we created a compound variable, named “soil resources”, as the coordinates of the first axis of a PCA for these three variables (the first axis accounted for 90% of the total variability).
Species trait data
To quantify the functional similarity of the 33 alien and 130 native species, we measured three traits non-destructively on all plants in the plots. These traits were plant height from the ground [cm], the number of leaves, and the number of reproductive organs (flowers and fruits). In addition to these traits, we also determined specific leaf area (SLA [cm2/g]) and aboveground biomass [kg]. For the alien plants, these two traits were measured destructively on individuals collected from the field plots. For each plant, we scanned the leaves and measured the total leaf area using the IMAGEJ software, version 1.4952. Then, we determined the leaf dry weight, and calculated the SLA as the leaf area divided by the leaf weight53. To obtain measurements for above-ground biomass of alien plants, all aboveground parts (leaves and stems) were dried in a drying oven (VWR International) at 50 °C for three days, and then weighed using a Mettler Toledo ML Series Precision Balance (ML Analytical balance).
Phylogenetic and functional trait analyses
To quantify phylogenetic diversity and relatedness, we constructed a phylogeny of the 166 species (33 aliens, 130 natives) using four commonly sequenced genes available in GenBank54: rbcL, matK, ITS1 and 5.8 s. Of the 133 native species, 120 species had at least one gene represented in GenBank. For the 13 native species without sequence data, we used instead the available sequences from congeneric relatives as a proxy (see phylogenetic guidelines by Jin and Cadotte 2015). We also included the genetic sequences of Amborella trichopoda Baill., a species that diverged early in angiosperm evolution, to serve as an outgroup species. Sequences were aligned for each of the four genes independently using FASconCAT v1.055 and combined into a single matrix. We then selected best-fit maximum likelihood (ML) models of nucleotide substitution for each gene sequence by jModeltest56. The ML phylogeny was generated using the PhyML algorithm with a BIONJ starting tree57,58 to estimate the phylogeny. Nodal support was estimated using approximate likelihood-ratio test (aLRT) scores, which have been shown to correlate with ML-bootstrap scores but require much less computational time58. We then used a semiparametric rate-smoothing method59 to transform the phylogeny to an ultrametric tree using the R package ape60.We iterated these functions across a suite of rate-smoothing parameters and found that the parameter value that maximized the likelihood was ʎ = 1. The final ultrametric phylogenetic tree, including 130 native species and 33 alien species, is provided in Supplementary Appendix A, Supplementary Fig. A.
We calculated Faith’s phylogenetic diversity (PD), which quantifies the total independent evolutionary history of a subset of taxa61,62 of all native species in each of the 166 plots. We also calculated the native phylogenetic relatedness (NMPD) within each non-invaded and invaded plot63, as well as the alien phylogenetic relatedness to natives (ANMPD) for the invaded plots64. These phylogenetic matrices were calculated using the functions PD and MPD in the R package picante (1.8 version65).
To test whether the phenotypic resemblance of native species is affected by the presence of alien species, we quantified the phylogenetic signal in five phenotypic traits (height, biomass, leaf production, floral production, SLA) of the native species in each plot using Blomberg's K and pagel’s lambda for continuous traits66, using the Phylosignal function in the R package picante. For each plot, we used for each of the native species the trait value averaged across the measured individuals. Significance of the phylogenetic signal in each plot was estimated through 999 randomizations with the trait distribution randomly shuffled across phylogenetic tips.
To quantify the functional diversity in each plot, we used our five phenotypic traits to calculate multi-trait functional richness (FRic) and functional dispersion (FDis). FRic is the amount of functional space filled by the community, an analogue of trait range in a multidimensional space61. FRic is calculated as the pairwise functional dissimilarity across species, using the Euclidean distance in multi-trait space after standardizing each trait to a mean of zero and a standard deviation of one. FDis is the multidimensional trait space or mean distance of each species, weighted by its relative abundance, to the centroid of all species in a community67,68. FDis for each plot was calculated using function dbFD which is available in the FD package67 in R version 3.2.5 (R Development Core Team 2014).
In addition to the functional diversity of the native communities, we also quantified the alien functional relatedness (ANMFD) to native species as well as the native functional relatedness (NMFD), using the same set of traits as used for the calculation of FRic and FDis. ANMFD was calculated as the mean weighted (by abundance) pairwise Euclidian distance of each alien species to the native community, NMFD was calculated as the mean weighted pairwise Euclidian distance among natives within each pair plot36,69, using the ‘dist’ function in R (package ‘stats’ version 3.7.0).
Comparing native diversity between invaded and non-invaded plots
To assess effects of the alien species on native plant diversity, we compared native species richness (SR), abundance, phylogenetic diversity indices (PD, NMPD), functional diversity indices (NMFD, FDis, FRic) of the invaded versus non-invaded plots using t-test for paired comparisons. To account for false discovery rates in the multiple comparisons, we applied Bonferroni correction for each test. We then used ANOVA (with Bonferroni correction) to test whether these native indices differed between invaded and non-invaded plots.
To visualize whether there was a difference in native species composition between invaded and non-invaded plots, we performed non-metric multidimensional scaling analysis (NMDS70) based on the Jaccard index of dissimilarity. To test whether the native species composition differed between the invaded and non-invaded plots, we used permutational analysis of variance (PERMANOVA, Anderson 2001). We performed these analyses using the metaMDS, and adonis functions of the Vegan package71.
To assess alien impact on species richness, abundance, FDis, FRic and PD of native communities, we calculated the changes in the different native diversity indices between the invaded and non-invaded plot of each pair using the following equation72,73 where x is diversity index of interest (richness, abundance, FDis, FRic, PD). So, for each index, we had 83 Alien Impactx values, where positive values indicate an increase in the diversity index due to alien invasion, and negative Alien Impactx values indicates a decline.
To test how Alien Impactx depends on the resource availability and the similarity of the alien species to the native community, we constructed series of Generalized Additive Models (GAM). Here, Alien Impactx for the different native diversity indices (SR, abundance, FDis, FRic, PD) were the response variables, and soil variables (nitrogen, organic carbon and moisture contents) and the dissimilarity between aliens and natives (ANMFD and ANMPD) were the explanatory variables. All models were compared using Akaike Information Criterion and Akaike weights to find an optimal model structure that best explains Alien Impactx74. We checked diagnostic plots (e.g. residuals versus fitted values and observed versus fitted values) to identify potential outliers, and whether the assumptions of homogeneity of variance and normality were not violated. In addition, we tested for normality of the residuals using the Shapiro–Wilk test75. To overcome the large spread of fitted values, phylogenetic measures were log transformed in order to improve the normality of the error distribution. All correlations between explanatory variables were < 0.6, indicating that multicollinearity should not be a problem. All analyses were done using R v.3.3.1 (RStudio Team 2018).
Results
In total, 33 alien and 130 native plant species were recorded within the sampled plots, covering a gradient of soil resources. The two major findings are the following: first, comparisons between invaded and non-invaded plots revealed that native functional diversity indices (FDis, FRic) were significantly higher in invaded plots than non-invaded ones. Interestingly, we found an evidence that supports Darwin’s pre-adaptation hypothesis; native phylogenetic indices (PD, NMPD), richness and abundance were marginally lower in invaded plots compared to non-invaded ones (Table 1, Fig. 1). Additionally, there was a significant decline in the phylogenetic signals of native traits in invaded plots compared to non-invaded ones (Table 3). Second, aliens were both functionally and phylogenetically dissimilar to natives in richer environments; they had high positive impact on native richness, functional and phylogenetic diversity indices (SR, FD, FR, PD), indicating that the presence of aliens increased native diversity in such environments. By contrast, aliens were both functionally and phylogenetically similar to natives in limited resources environments; they had a significant negative impact on native diversity measures, indicating that presence of aliens was associated with negative effects on native diversity measures in such environments (Table 2, Fig. 2).
Non-metric multidimensional scaling analysis (NMDS) showing the differences in the native species composition between invaded plots and non-invaded ones (stress = 0.09 for the two axes). Red crosses represent invaded plots, blue crosses non-invaded ones. The colored ellipses are centered on the centroid of both groups and their size is proportional to the standard error of the coordinates of the corresponding points.
Relative differences in native diversity indices between invaded/non-invaded plots
In total, there were significant differences in native diversity between invaded and non-invaded plots, indicating significant effects of alien species on native diversity indices (Table 1, Fig. 1). Paired comparisons revealed that native richness and abundance were slightly lower (− 10.65% and − 9.83% respectively) in invaded plots compared to non-invaded plots (t-test, z = 1.496, p < 0.6) (Table 1). Native phylogenetic diversity and phylogenetic relatedness were marginally lower in invaded plots than in non-invaded ones (t-test, z = − 1.55, p < 0.05; t-test = − 1.78, p < 0.07) and declined by 9.7% and 10%, respectively, compared to non-invaded plots. However, native functional dispersion and functional richness were significantly higher in invaded plots compared to non-invaded ones (t- test, z = − 0.672, p < 0.01; z = − 2.713, p < 0.01 respectively) (Table 1, Fig. 1).
Relative differences in native composition between invaded/non-invaded plots
The NMDS analysis and PERMANOVA test revealed statistically significant differences in native species composition between invaded and non-invaded plots. NMDS analysis (Fig. 2) showed that the non-invaded plots tended to have similar species composition and formed a subset of the invaded plots whose compositions were highly dispersed. PERMANOVA analyses confirmed this pattern by unravelling significant dispersion in the native species composition in invaded plots compared to non-invaded ones (R2 = 0.044, F-value = 5.18, p < 0.001, 999 permutations), which was influenced by the presence of alien species.
Relative differences in the phylogenetic signal of traits of native species between invaded/non-invaded plots
There was a decline in the phylogenetic signal of measured native traits (height-biomass-leaf production-floral production-SLA) in invaded plots compared to non-invaded ones (Table 3). For shoot biomass, height and floral production of native species, the Blomberg’s K values declined significantly in invaded plots (K = 0.67, p < 0.01), but such decline was non-significant for leaf production and SLA. This indicates that the phylogenetic signal of native species traits in invaded plots was weaker than expected by a Brownian motion model for trait evolution. In parallel, Pagel’s lambda values of such native traits were significantly lower in invaded plots compared to non-invaded ones (Table 3).
Magnitude of Alien Impact along soil resources gradients
GAM models revealed that aliens tended to have significant positive impacts on native diversity indices in plots with high contents of soil organic carbon, nitrogen and moisture, but negative impacts in resource-limited plots (Table 2, Fig. 3). For native richness, abundance, functional dispersion, and native phylogenetic diversity, Alien Impactx was positive in richer plots and negative in limited plots, with soil resources having highly significant effects (F = 23.08, p < 0.001; F = 9.30, p < 0.001; F = 3.75, p < 0.001; F = 24.21, p < 0.001, respectively). However, AI on native functional richness was non-significant and not affected by soil resources (F = 0.007, p < 0.93).
Alien impact (AI) on native richness, abundance, PD, FDis, FRic, in responding to soil resource availability (nitrogen, organic matter, soil moisture summarized in composite variable with a PCA) and the alien relatedness (functional and phylogenetic) to natives in SKP. Blue solid lines represent the average response expected by GAM model with 95% confidence intervals (red dashed lines). Green points represent the observed data points.
Magnitude of Alien Impact along biotic relatedness
Alien species had significant positive impacts on native diversity measures, which increased with increasing functional and phylogenetic relatedness between aliens and natives (Table 2, Fig. 3). Aliens tended to have positive impacts on native diversity if they were functionally and phylogenetically dissimilar to natives; such impacts tended to be negative if aliens were more closely related to natives. For native richness and abundance, and native functional dispersion and richness, Alien Impactx had positive effects with greater alien functional (ANMFD) and phylogenetic relatedness (ANMPD) to natives (F = 7.65, p < 0.0001; F = 31.13, p < 0.001; F = 57.577, p < 0.01; F = 35.649, p < 0.0001; F = 3.821, p < 0.001; F = 887.017, p < 0.001; F = 7.808, p < 0.001; F = 24.904, p < 0.001, respectively).
Discussion
Documenting how alien species modify native community structure from a functional and phylogenetic point of view is a critical task to understanding the mechanisms driving alien impacts on native species assembly6,7,29,76. Interestingly, we found that invaded native communities are phylogenetically less, but functionally more diverse than non-invaded communities within our study area, indicating that the phylogenetic diversity of resident native communities can diverge from trait-based assembly processes. For example, in highly competitive environments, communities of phylogenetically under-dispersed species can be functionally over-dispersed to adapt to the intense competitive interactions77 among different functional species groups. Although, on average, the presence of alien plants affects the diversity components of invaded native communities, we highlight here that the magnitude of this impact strongly depends on resource availability and alien relatedness to natives.
First, our results support the hypothesis of niche-filling26, in which native species in invaded communities can be functionally more dissimilar to alien species and to each other than in non-invaded communities, which gives the possibility to alien species to contribute to the creation of gaps in the niche space by excluding functionally similar natives. The establishment of functionally-different native species is then possible, which would increase the functional dissimilarity of native communities26,27,78. By contrast, the presence of native species exploiting similar niches (i.e. intact native communities with filled niches) as the potential invader would confer high native biotic resistance due to the lack of an ‘empty niche’79,80,81, which would decrease the likelihood of establishment success even further. For example, this could happen when the spatial niche space is filled by productive native species that efficiently utilize local resources82,83 and occupy the same niche as the invader. However, this process at the functional level may not be mirrored by the phylogenetic diversity of native species, which in our case decreased in presence of aliens. Our results are consistent with the meta-analysis of Loiola et al., who proposed that aliens can fill empty gaps and occupy the existing functional niche space of the displaced species, rather than add novel evolutionary origins of resident native species7,8,84.
Ecological studies speculated that if the phylogenetic and functional components of invaded communities are concordant, the measured traits should exhibit a strong phylogenetic signal85. By contrast, our finding indicated an inconsistency between native PD and FD, which is supported by an observed decline in the phylogenetic signal of native species traits in the invaded communities compared to non-invaded ones. This inconsistency can thus be attributed to the fact that phylogenies are not completely capturing ecologically relevant traits. The weak phylogenetic signal for these traits could be due to their lack of phylogenetic conservatism or to convergent evolution in our study area86. Hence, although it is frequently assumed that the phylogenetic diversity is a surrogate for functional diversity in invasion studies28,84,87; but see, for example, Lososova et al., our results indicate that the phylogenetic structure can be complementary to, but not a substitute for, the functional trait structure88.
Indeed, the loss of native phylogenetic divergence in the invaded plots supports the prediction that competition can eliminate native species with conserved trait-based ecological strategies from phylogenetically-closely related taxa31. In other words, native species that have high phylogenetic convergence in their traits or similar trait values may decrease their potential advantage in competitive abilities89. For example, convergence in height may decrease the opportunity for individuals of taller species to outcompete those of smaller species in light‐limited environments90.
The inconsistency between phylogenetic under-dispersion and trait over-dispersion in invaded communities may also arise from the present-day strong negative interactions among phylogenetically related similar species (alien/native). This could lead to mutual exclusion of phylogenetically similar species and ultimately to divergence in trait space77. For example, highly competitive alien plants have a strong negative effect on native diversity, especially through increasing competition for resources such as light91,92,93,94,95. These interspecific interactions are particularly important for natives with more divergent traits, which are expected to be weakly conserved within phylogenetic lineages77.
We also observed a decline in native richness of invaded communities. A possible explanation of this change could be attributed to the covariation between environmental changes and presence of alien species96,97,98. For example, milder winters changed the environmental space of deciduous forests to conditions that are now more suitable for evergreen broad-leaved species99. Consequently, resident native species can become increasingly poorly adapted to the local environment, which will then provide opportunities for better-adapted aliens, ultimately declining the native richness through competitive exclusion.
Second, we found that aliens that were functionally and phylogenetically dissimilar to natives in resources rich environments100 are likely to expand the functional and phylogenetic space of native community (see Refs.29,101,102. This expansion may be attributed to the differences in the competitive abilities of both species groups31,34,103. For example, aliens with superior competitive ability tend to eliminate resident natives that have similar trait values as well as similar resources preferences. Thereby, if both species groups compete for different resources, and aliens outcompete resident natives, this should promote only dissimilar natives to co-exist with aliens18,104.
By contrast, we found that aliens functionally and phylogenetically similar to natives in resource-limited environments100 are expected to occupy a portion of the native functional space, leading to a reduction in native functional diversity. Previous studies (e.g. Refs.105,106,107) have suggested that both species groups should be more similar under stressful conditions108,109. This similarity is due to the strong effect of the environmental filtering that select species sharing analogous responses to the limiting environmental resources, a pattern that is consistent with the pre-adaptation hypothesis (PAH)12,17 even though these classic theories do not include facilitation as a potential mechanism. Eventually, alien species are presumed to occupy a portion of the functional space originally occupied by native species, leading to an increased functional similarity between both species groups24,25 and a reduction in native richness and subsequently in their functional and phylogenetic space.
Conclusion
The presented paired-design empirical study highlighted that alien plants pose significant impacts at the native community structure1, and that such impacts are significantly linked to soil resources availability and to alien relatedness with resident natives. Significantly, we found evidence for the role of environmental filtering (pre-adaptation hypothesis) that regulate alien impacts on resident natives in harsh environments, while niche gap filling (Darwin’s Naturalization Hypothesis DNH) was the main mechanism determining such impacts in rich-resources environments. Our quantitative approach to value alien impacts could be further developed as the basis for underscoring alien species and recipient ecosystems for risk assessment of invasions110. We hope this contribution helps to invigorate this area of research by highlighting the association between invasion impacts and biotic and abiotic gradients at several levels of ecological complexity. As such, future studies should conduct similar ecological surveys in different ecosystems at fine and coarse scales to test the generality of our findings. For example, arid disturbed-invaded ecosystems generally undergo rapid human-driven disturbances that can strongly change the species composition similarity111, and thereby affect significantly the relatedness between alien and native groups. Further, It would be interesting to test the roles of different nutrient‐niches and biotic relatedness between aliens and natives on the magnitude of alien impacts, which might be an effective strategy for a global risk assessment of plant invasions.
References
Vilà, M. et al. Ecological impacts of invasive alien plants: A meta-analysis of their effects on species, communities and ecosystems: Ecological impacts of invasive alien plants. Ecol. Lett. 14, 702–708 (2011).
Pyšek, P. et al. A global assessment of invasive plant impacts on resident species, communities and ecosystems: The interaction of impact measures, invading species’ traits and environment. Glob. Change Biol. 18, 1725–1737 (2012).
Pyšek, P. et al. Scientists’ warning on invasive alien species. Biol. Rev. 95, 1511–1534 (2020).
Gordon, D. R. Effects of invasive, non-indigenous plant species on ecosystem processes: Lessons from Florida. Ecol. Appl. 8, 975–989 (1998).
Vieites-Blanco, C. & González-Prieto, S. J. Effects of Carpobrotus edulis invasion on soil gross N fluxes in rocky coastal habitats. Sci. Total Environ. 619–620, 966–976 (2018).
Loiola, P. P. et al. Invaders among locals: Alien species decrease phylogenetic and functional diversity while increasing dissimilarity among native community members. J. Ecol. 106, 2230–2241 (2018).
de la Riva, E. G., Godoy, O., Castro-Díez, P., Gutiérrez-Cánovas, C. & Vilà, M. Functional and phylogenetic consequences of plant invasion for coastal native communities. J. Veg. Sci. 30, 510–520 (2019).
Ordonez, A. Functional and phylogenetic similarity of alien plants to co-occurring natives. Ecology 95, 1191–1202 (2014).
Bezeng, S. B., Davies, J. T., Yessoufou, K., Maurin, O. & der Bank, M. V. Revisiting Darwin’s naturalization conundrum: Explaining invasion success of non-native trees and shrubs in southern Africa. J. Ecol. 103, 871–879 (2015).
Li, S. et al. The effects of phylogenetic relatedness on invasion success and impact: Deconstructing Darwin’s naturalisation conundrum. Ecol. Lett. 18, 1285–1292 (2015).
Carboni, M. et al. What it takes to invade grassland ecosystems: Traits, introduction history and filtering processes. Ecol. Lett. 19, 219–229 (2016).
Cadotte, M. W., Campbell, S. E., Li, S., Sodhi, D. S. & Mandrak, N. E. Preadaptation and naturalization of nonnative species: Darwin’s two fundamental insights into species invasion. Annu. Rev. Plant Biol. 69, 661–684 (2018).
Diez, J. M., Sullivan, J. J., Hulme, P. E., Edwards, G. & Duncan, R. P. Darwin’s naturalization conundrum: Dissecting taxonomic patterns of species invasions. Ecol. Lett. 11, 674–681 (2008).
Ma, C. et al. Different effects of invader–native phylogenetic relatedness on invasion success and impact: A meta-analysis of Darwin’s naturalization hypothesis. Proc. R. Society B: Biological Sciences 283, 20160663 (2016).
Bennett, J. A. Similarities between invaders and native species: Moving past Darwin’s naturalization conundrum. J. Veg. Sci. 30, 1027–1034 (2019).
Funk, J. L., Standish, R. J., Stock, W. D. & Valladares, F. Plant functional traits of dominant native and invasive species in mediterranean-climate ecosystems. Ecology 97, 75–83 (2016).
Daehler, C. C. Darwin’s naturalization hypothesis revisited. Am. Nat. 158, 324–330 (2001).
Duncan, R. P. & Williams, P. A. Ecology: Darwin’s naturalization hypothesis challenged. Nature 417, 608 (2002).
Ferreira, R. B., Beard, K. H., Peterson, S. L., Poessel, S. A. & Callahan, C. M. Establishment of introduced reptiles increases with the presence and richness of native congeners. Amphibia-Reptilia 33, 387–392 (2012).
Allen, C. R. et al. Predictors of regional establishment success and spread of introduced non-indigenous vertebrates. Glob. Ecol. Biogeogr. 22, 889–899 (2013).
Maitner, B. S., Rudgers, J. A., Dunham, A. E. & Whitney, K. D. Patterns of bird invasion are consistent with environmental filtering. Ecography 35, 614–623 (2012).
Park, D. S. & Potter, D. Why close relatives make bad neighbours: Phylogenetic conservatism in niche preferences and dispersal disproves Darwin’s naturalization hypothesis in the thistle tribe. Mol. Ecol. 24, 3181–3193 (2015).
Park, D. S. & Potter, D. A reciprocal test of Darwin’s naturalization hypothesis in two mediterranean-climate regions. Glob. Ecol. Biogeogr. 24, 1049–1058 (2015).
Kembel, S. W. & Hubbell, S. P. The phylogenetic structure of a neotropical forest tree community. Ecology 87, S86–S99 (2006).
Catford, J. A., Jansson, R. & Nilsson, C. Reducing redundancy in invasion ecology by integrating hypotheses into a single theoretical framework. Divers. Distrib. 15, 22–40 (2009).
Thuiller, W. et al. Resolving Darwin’s naturalization conundrum: a quest for evidence. Divers. Distrib. 16, 461–475 (2010).
Funk, J. L., Cleland, E. E., Suding, K. N. & Zavaleta, E. S. Restoration through reassembly: Plant traits and invasion resistance. Trends Ecol. Evol. 23, 695–703 (2008).
Lapiedra, O., Sol, D., Traveset, A. & Vilà, M. Random processes and phylogenetic loss caused by plant invasions. Glob. Ecol. Biogeogr. 24, 774–785 (2015).
Castro-Díez, P., Pauchard, A., Traveset, A. & Vilà, M. Linking the impacts of plant invasion on community functional structure and ecosystem properties. J. Veg. Sci. 27, 1233–1242 (2016).
Hulme, P. E. & Bernard-Verdier, M. Evaluating differences in the shape of native and alien plant trait distributions will bring new insights into invasions of plant communities. J. Veg. Sci. 29, 348–355 (2018).
Mayfield, M. M. & Levine, J. M. Opposing effects of competitive exclusion on the phylogenetic structure of communities: Phylogeny and coexistence. Ecol. Lett. 13, 1085–1093 (2010).
de Bello, F. et al. Functional species pool framework to test for biotic effects on community assembly. Ecology 93, 2263–2273 (2012).
Chesson, P. Mechanisms of maintenance of species diversity. Annu. Rev. Ecol. Syst. 31, 343–366 (2000).
Kunstler, G. et al. Competitive interactions between forest trees are driven by species’ trait hierarchy, not phylogenetic or functional similarity: Implications for forest community assembly. Ecol. Lett. 15, 831–840 (2012).
Gallien, L. et al. Contrasting the effects of environment, dispersal and biotic interactions to explain the distribution of invasive plants in alpine communities. Biol. Invasions 17, 1407–1423 (2015).
Gallien, L. & Carboni, M. The community ecology of invasive species: Where are we and what’s next?. Ecography 40, 335–352 (2017).
Parker, I. M. et al. Impact: Toward a framework for understanding the ecological effects of invaders. Biol. Invasions 1, 3–19 (1999).
Byers, J. E. et al. Directing research to reduce the impacts of nonindigenous species. Conserv. Biol. 16, 630–640 (2002).
Hejda, M., Pyšek, P. & Jarošík, V. Impact of invasive plants on the species richness, diversity and composition of invaded communities. J. Ecol. 97, 393–403 (2009).
Pyšek, P. & Pyšek, A. Invasion by Heracleum mantegazzianum in different habitats in the Czech Republic. J. Veg. Sci. 6, 711–718 (1995).
Hejda, M. & Pyšek, P. What is the impact of Impatiens glandulifera on species diversity of invaded riparian vegetation?. Biol. Conserv. 132, 143–152 (2006).
Chmura, D. et al. The influence of invasive Fallopia taxa on resident plant species in two river valleys (southern Poland). Acta Soc. Bot. Pol. 84, 23–33 (2015).
MacDougall, A. S., Gilbert, B. & Levine, J. M. Plant invasions and the niche. J. Ecol. 97, 609–615 (2009).
Li, S. et al. Contrasting effects of phylogenetic relatedness on plant invader success in experimental grassland communities. J. Appl. Ecol. 52, 89–99 (2015).
Macarthur, R. & Levins, R. The limiting similarity, convergence, and divergence of coexisting species. Am. Nat. 101, 377–385 (1967).
Abrams, P. The theory of limiting similarity. Annu. Rev. Ecol. Syst. 14, 359–376 (1983).
Davies, T. J. Evolutionary ecology: When relatives cannot live together. Evol. Ecol. 16, R645–R647 (2006).
Omar Kariem, A. Eco-geographical analysis on mountain plants—Kariem Omar—Livres spécialisés. Africa Vivre. https://www.laboutiqueafricavivre.com/livres-specialises/156599-eco-geographical-analysis-on-mountain-plants-9783847331537.html (2012).
Omar Karim A. Extinction—Towards Plant Conservation. (Lap Lambert Academic Publ, 2014).
Klute, A. Water retention: Laboratory methods. In: (ed. Klute, A.) Methods of Soil Analysis, Part 1, Physical and Mineralogical Methods, ASA and SSSA, Madison, 635–662. https://doi.org/10.2136/sssabookser5.1.2ed. (1986)
Allen, S. E., Grimshaw, H. M., Parkinson, J. A. & Quarmby, C. Chemical Analysis of Ecological Materials. (Blackwell Scientific Publications, 1974).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Pérez-Harguindeguy, N. et al. New handbook for stand-ardised measurement of plant functional traits worldwide. Aust. J. Bot. 61, 167–234 (2013).
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Wheeler, D. L. GenBank. Nucleic Acids Res. 33, D34-38 (2005).
Kück, P. & Meusemann, K. FASconCAT: Convenient handling of data matrices. Mol. Phylogenet. Evol. 56, 1115–1118 (2010).
Posada, D. jModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 25, 1253–1256 (2008).
Anisimova, M. & Gascuel, O. Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. Syst. Biol. 55, 539–552 (2006).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Sanderson, M. J. Estimating absolute rates of molecular evolution and divergence times: A penalized likelihood approach. Mol. Biol. Evol. 19, 101–109 (2002).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Villéger, S., Mason, N. W. H. & Mouillot, D. New multidimensional functional diversity indices for a multifaceted framework in functional ecology. Ecology 89, 2290–2301 (2008).
Veron, S., Davies, T. J., Cadotte, M. W., Clergeau, P. & Pavoine, S. Predicting loss of evolutionary history: Where are we?. Biol. Rev. 92, 271–291 (2017).
Swenson, N. Functional and Phylogenetic Ecology in R. (Springer, 2014). https://doi.org/10.1007/978-1-4614-9542-0.
Webb, C. O., Ackerly, D. D., McPeek, M. A. & Donoghue, M. J. Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 33, 475–505 (2002).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Blomberg, S. P., Garland, T. Jr. & Ives, A. R. Testing for phylogenetic signal in comparative data: Behavioral traits are more labile. Evolution 57, 717–745 (2003).
Laliberté, E. & Legendre, P. A distance-based framework for measuring functional diversity from multiple traits. Ecology 91, 299–305 (2010).
Spasojevic, M. J. & Suding, K. N. Inferring community assembly mechanisms from functional diversity patterns: The importance of multiple assembly processes. J. Ecol. 100, 652–661 (2012).
Carboni, M. et al. Functional traits modulate the response of alien plants along abiotic and biotic gradients. Glob. Ecol. Biogeogr. 27, 1173–1185 (2018).
Kruskal, J. B. Multidimensional scaling by optimizing goodness of fit to a nonmetric hypothesis. Psychometrika 29, 1–27 (1964).
Oksanen, J. et al. Vegan: community ecology package. R Package version 2.4-1. https://cran.r-project.org (2016).
Vila, M. et al. Local and regional assessments of the impacts of plant invaders on vegetation structure and soil properties of Mediterranean islands. J. Biogeogr. 33, 853–861 (2006).
Dong, L.-J., Yu, H.-W. & He, W.-M. What determines positive, neutral, and negative impacts of Solidago canadensis invasion on native plant species richness? Sci Rep. 5(1), 1–9 (2015).
Johnson, J. B. & Omland, K. S. Model selection in ecology and evolution. Trends Ecol. Evol. 19, 101–108 (2004).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples). Biometrika 52, 591–611 (1965).
Jucker, T., Carboni, M. & Acosta, A. T. R. Going beyond taxonomic diversity: deconstructing biodiversity patterns reveals the true cost of iceplant invasion. Divers. Distrib. 19, 1566–1577 (2013).
Prinzing, A. et al. Less lineages—More trait variation: Phylogenetically clustered plant communities are functionally more diverse. Ecol. Lett. 11, 809–819 (2008).
Blonder, B. Do hypervolumes have holes?. Am. Nat. 187, E93–E105 (2016).
Levine, J. M. & D’Antonio, C. M. Elton revisited: A review of evidence linking diversity and invasibility. Oikos 87, 15–26 (1999).
Fargione, J., Brown, C. S. & Tilman, D. Community assembly and invasion: An experimental test of neutral versus niche processes. PNAS 100, 8916–8920 (2003).
Zavaleta, E. S. & Hulvey, K. B. Realistic variation in species composition affects grassland production, resource use and invasion resistance. Plant Ecol 188, 39–51 (2007).
Case, T. J. Invasion resistance arises in strongly interacting species-rich model competition communities. Proc. Natl. Acad. Sci. 87, 9610–9614 (1990).
Kennedy, T. A. et al. Biodiversity as a barrier to ecological invasion. Nature 417, 636 (2002).
Gerhold, P. et al. Phylogenetically poor plant communities receive more alien species, which more easily coexist with natives. Am. Nat. 177, 668–680 (2011).
de Bello, F. et al. Decoupling phylogenetic and functional diversity to reveal hidden signals in community assembly. Methods Ecol. Evol. 8, 1200–1211 (2017).
Cadotte, M. W., Carboni, M., Si, X. & Tatsumi, S. Do traits and phylogeny support congruent community diversity patterns and assembly inferences?. J. Ecol. 107, 2065–2077 (2019).
Lososová, Z. et al. Alien plants invade more phylogenetically clustered community types and cause even stronger clustering. Glob. Ecol. Biogeogr. 24, 786–794 (2015).
Hulme, P. E. & Bernard-Verdier, M. Comparing traits of native and alien plants: Can we do better?. Funct. Ecol. 32, 117–125 (2018).
Luo, Y.-H. et al. Trait-based community assembly along an elevational gradient in subalpine forests: Quantifying the roles of environmental factors in inter- and intraspecific variability. PLoS One 11, e0155749 (2016).
Luo, Y.-H. et al. Forest community assembly is driven by different strata-dependent mechanisms along an elevational gradient. J. Biogeogr. 46, 2174–2187 (2019).
Jakobs, G., Weber, E. & Edwards, P. J. Introduced plants of the invasive Solidago gigantea (Asteraceae) are larger and grow denser than conspecifics in the native range. Divers. Distrib. 10, 11–19 (2004).
Chmura, D. & Sierka, E. The invasibility of deciduous forest communities after disturbance: A case study of Carex brizoides and Impatiens parviflora invasion. For. Ecol. Manag. 242, 487–495 (2007).
Szymura, M. & Szymura, T. H. The dynamics of growth and flowering of invasive Solidago species. Steciana 19, 143–152 (2015).
Divíšek, J. et al. Similarity of introduced plant species to native ones facilitates naturalization, but differences enhance invasion success. Nat. Commun. 9, 4631 (2018).
Czarniecka-Wiera, M., Kącki, Z., Chytrý, M. & Palpurina, S. Diversity loss in grasslands due to the increasing dominance of alien and native competitive herbs. Biodivers. Conserv. https://doi.org/10.1007/s10531-019-01794-9 (2019).
Tilman, D. Species richness of experimental productivity gradients: How important is colonization limitation?. Ecology 74, 2179–2191 (1993).
Burke, M. J. W. & Grime, J. P. An experimental study of plant community invasibility. Ecology 77, 776–790 (1996).
Naeem, S. et al. Plant diversity increases resistance to invasion in the absence of covarying extrinsic factors. Oikos 91, 97–108 (2000).
Berger, S., Söhlke, G., Walther, G.-R. & Pott, R. Bioclimatic limits and range shifts of cold-hardy evergreen broad-leaved species at their northern distributional limit in Europe. Phytocoenologia 37, 523–539 (2007).
El-Barougy, R. F. et al. Functional similarity and dissimilarity facilitate alien plant invasiveness along biotic and abiotic gradients in an arid protected area. Biol. Invasions 22, 1997–2016 (2020).
Ordonez, A., Wright, I. J. & Olff, H. Functional differences between native and alien species: A global-scale comparison: Functional differences of native and alien plants. Funct. Ecol. 24, 1353–1361 (2010).
Godoy, O. & Levine, J. M. Phenology effects on invasion success: Insights from coupling field experiments to coexistence theory. Ecology 95, 726–736 (2014).
Cavender-Bares, J., Ackerly, D. D., Baum, D. A. & Bazzaz, F. A. Phylogenetic overdispersion in Floridian Oak Communities. Am. Nat. 163, 823–843 (2004).
Richardson, D. M. & Pyšek, P. Plant invasions: Merging the concepts of species invasiveness and community invasibility. Prog. Phys. Geogr. 30, 409–431 (2006).
Pyšek, P., Prach, K. & Smilauer, P. Relating invasion success to plant traits: An analysis of the Czech alien flora. Plant Invasions Gen. Aspects Spec. Probl. 39–60 (1995).
Pyšek, P. et al. Alien plants in temperate weed communities: Prehistoric and recent invaders occupy different habitats. Ecology 86, 772–785 (2005).
Prinzing, A., Durka, W., Klotz, S. & Brandl, R. Which species become aliens?. Evol. Ecol. Res. 4, 385–405 (2002).
van Kleunen, M., Weber, E. & Fischer, M. A meta-analysis of trait differences between invasive and non-invasive plant species. Ecol. Lett. 13, 235–245 (2010).
Jauni, M. & Hyvönen, T. Interactions between alien plant species traits and habitat characteristics in agricultural landscapes in Finland. Biol. Invasions 14, 47–63 (2012).
Nentwig, W., Kühnel, E. & Bacher, S. A generic impact-scoring system applied to alien mammals in Europe. Conserv. Biol. 24, 302–311 (2010).
Liu, P. et al. Urbanization increases biotic homogenization of zooplankton communities in tropical reservoirs. Ecol. Indic. 110, 105899 (2020).
Acknowledgements
The study was financed and supported by the University of Fribourg, Department of Biology, Ecology and Evolution, Switzerland. Louis-Félix Bersier acknowledges the Swiss National Science Foundation grant 31003A_165800. We are grateful to Prof. Mark Van-Kleunen (Biology Dept., Universität Konstanz, Germany) for his comments on the earlier drafts of the manuscript.
Author information
Authors and Affiliations
Contributions
R.F.E. conceived the approach with substantial contributions from L.F.B. I.A.E. and A.A.K. collected and compiled the datasets. R.F.E. carried out the analyses and wrote the manuscript and prepared figures and tables. L.F.B. helped with the writing of the manuscript. All authors gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El-Barougy, R.F., Elgamal, I.A., Khedr, AH.A. et al. Contrasting alien effects on native diversity along biotic and abiotic gradients in an arid protected area. Sci Rep 11, 13557 (2021). https://doi.org/10.1038/s41598-021-92763-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-92763-2
This article is cited by
-
Composition and phylogenetic structure of Pampean grasslands under distinct land use and presence of alien species
Community Ecology (2023)
-
Plant invasion drives liana and tree community assemblages and liana-tree network structure in two moist semi-deciduous forests in Ghana
Biological Invasions (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.