Abstract
Here we report a jute endophyte Staphylococcus hominis strain MBL_AB63 isolated from jute seeds which showed promising antimicrobial activity against Staphylococcus aureus SG511 when screening for antimicrobial substances. The whole genome sequence of this strain, annotated using BAGEL4 and antiSMASH 5.0 to predict the gene clusters for antimicrobial substances identified a novel antimicrobial peptide cluster that belongs to the class I lantibiotic group. The predicted lantibiotic (homicorcin) was found to be 82% similar to a reported peptide epicidin 280 having a difference of seven amino acids at several positions of the core peptide. Two distinct peaks obtained at close retention times from a RP-HPLC purified fraction have comparable antimicrobial activities and LC–MS revealed the molecular mass of these peaks to be 3046.5 and 3043.2 Da. The presence of an oxidoreductase (homO) similar to that of epicidin 280- associated eciO or epilancin 15X- associated elxO in the homicorcin gene cluster is predicted to be responsible for the reduction of the first dehydrated residue dehydroalanine (Dha) to 2-hydroxypropionate that causes an increase of 3 Da mass of homicorcin 1. Trypsin digestion of the core peptide and its variant followed by ESI–MS analysis suggests the presence of three ring structures, one in the N-terminal and other two interlocking rings at the C-terminal region that remain undigested. Homicorcin exerts bactericidal activity against susceptible cells by disrupting the integrity of the cytoplasmic membrane through pore formation as observed under FE-SEM.
Similar content being viewed by others
Introduction
Lantibiotics are ribosomally synthesized antimicrobial peptides possessing unusual amino acids normally not found in nature like lanthionine (Lan)- or methyllanthionine (MeLan)1. These peptides are mostly produced by a variety of Gram-positive bacteria and usually show antagonizing activity against Gram-positive bacteria, especially closely related species2,3. A few Gram-negative bacteria were also found to produce lantibiotics reported in recent years4,5. Producer strains synthesize lantibiotics as inactive prepeptides that consist of an N-terminal leader sequence and a C-terminal prepeptide part. This prepeptide sequentially undergoes several posttranslational modifications to become the mature lantibiotic. During the posttranslational modification unusual amino acids like Lan and MeLan are formed through intermolecular cyclization of the thiol groups of cysteine residues with Dha and Dhb, which are the dehydrated products of specific serine and threonine residues, respectively2. These thio-ether ring structures are assumed to provide the rigidity and impart resistance to proteolytic enzymes, temperature, pH and other parameters. Biosynthesis genes of lantibiotics are organized in the same gene clusters, which include the genes coding for the precursor peptides and other essential proteins involved in post-translational modification, processing of leader peptide, transportation, immunity, and regulation6,7. Recently, lantibiotics were classified in four classes based on the modification enzymes responsible for dehydration and cyclization1. In the class I lantibiotics, a dedicated dehydratase LanB, and a cyclase LanC perform dehydration and cyclization1. However, in class II lantibiotics, a biofunctional enzyme LanM introduces Lan or MeLan rings. For class III and class IV lantibiotics, tridomain proteins LanKC and LanL catalyze the formation of lanthipeptides, respectively1. Finally, cytoplasmic membrane protein LanT (an ATP-binding cassette [ABC] transporter) exports the modified precursor peptide outside the cell and an extracellular protease LanP cleaves the leader peptide to release the active lantibiotic. However, in some cases, a single protein LanT is responsible to export the precursor peptide and cleavage of leader peptide simultaneously8. Producer strains protect themselves from their own lantibiotic by expressing immunity proteins, including ABC transporter proteins LanFE(G) and/or lipoprotein LanI9.
Due to the increasing number of antibiotic resistance cases10,11, the world is in an urgent need for novel compounds and innovative methods to minimize the spread and development of drug resistant infection. Currently, among the Staphylococcus aureus strains isolated in the hospitals, 60–70% are found to be multidrug resistant3,12. Therefore, new antimicrobial drugs not affected by existing resistance mechanisms are needed to prevent the potential epidemic outbreaks of infectious diseases. The prototypic lantibiotic nisin has been utilized in the food industry as food preservative for over 40 years in more than 80 countries without the development of stable resistance, possibly as a consequence of its multiple modes of action13.
Potentially rich sources of lantibiotics are the endophytes, a treasure trove of therapeutically important bioactive compounds14,15. They are microorganisms, mostly fungi and bacteria that reside in plant tissue typically causing no apparent disease symptoms but on the contrary maintain a symbiotic relationship with host plants16,17. Endophytes basically have gained enormous attention for their capacity to control plant pathogenic insects and promote plant establishment and growth under adverse conditions18,19,20,21. However, they are also reported to produce a plethora of bioactive secondary metabolites that have anti-arthritic, antimicrobial, anti-cancer, anti-diabetic, anti-insect, and immunosuppressant activities22,23,24,25. In particular, researchers have shown interest in antimicrobial peptides (AMPs) from different types of endophytes due to their chemical diversity and broad spectrum of activity26,27.
As lantibiotics have diverse applications, much attention has been paid in the last few decades on the identification of new peptides. In recent years, with the availability of abundant genomic sequence data in public databases, many new lantibiotics from different sources have been identified. In this study, we present a novel class I lantibiotic homicorcin and its variant homicorcin 1 isolated from a plant (jute) endophyte Staphylococcus hominis strain MBL_AB63. A large number of lantibiotics with diverse structure have been identified from different Staphylococcal strains. These include Pep528 and epicidin 28029 from Staphylococcus epidermidis strains 5 and BN 280 respectively, epilancin K730 and the closely related epilancin 15X31 from S. epidermidis strains K7 and 15X154, epidermin32,33 from S. epidermidis Tü3298, gallidermin34 from Staphylococcus gallinarum F16/P57 Tü3298, hominicin35 from Staphylococcus hominis MBBL 2–9, nukacin ISK-136 and the very similar warnericin RB437 from S. warneri ISK-1 and RB4 respectively, two component lantibiotic staphylococcin C5538 from S. aureus C55 and a few more. Most of the lantibiotics isolated from staphylococcal origin are found to be active against closely related species.The biosynthetic gene cluster of homicorcin possesses an additional oxido-reductase enzyme that further modifies the homicorcin N-terminal first residue of dehydroalanine to 2-hydroxypropionate forming homicorcin 1, a similar modification has been predicted for epicidin 28029 and epilancin 15X31,39. Homicorcin and its variant are equally produced in the culture supernatant without any induction. Further characterization of these two peptides reveals that they show almost similar spectrum of bactericidal activity against closely related species.
Results
In silico identification of the homicorcin gene cluster in Staphylococcus hominis strain MBL_AB63
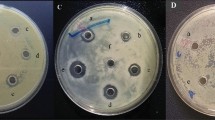
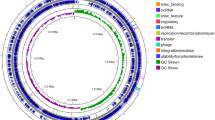
Jute endophyte Staphylococcus hominis strain MBL_AB63 isolated in our laboratory showed significant antibacterial activity against Staphylococcus aureus SG511 in a preliminary screening (Fig. 1A). Following whole genome sequencing of MBL_AB63 (DDBJ/ENA/GenBank accession number JAELVP000000000), the in silico tools BAGEL4 and anti-SMASH 5.0 were used to predict the biosynthetic gene cluster of the bioactive compound responsible for the antimicrobial activity. Both the tools identified a class I lantibiotic gene cluster of homicorcin that was found to be 82% similar to a reported lantibiotic; epicidin 280 with a difference of seven amino acids in the mature peptide (Fig. 1B,C).
Prediction of the homicorcin gene cluster and comparison with other class I lantibiotics. (A) Inhibitory activity of Staphylococcus hominis strain MBL_AB63 (activity overlay assay against Staphylococcus aureus SG511). (B) Biosynthetic gene cluster of different class I lantibiotics. Homicorcin gene cluster was predicted using BAGEL 4.0 (homO: 3-oxoacyl-[acyl-carrier-protein] reductase gene; homI: immunity gene; homA: homicorcin gene; homP: protease gene for leader peptide cleavage; homBC: genes for post translational modification enzyme). (C) Comparisons between the peptide sequences of homicorcin and other class I lantibiotics (residues not similar to Epicidin 280 are shown in red).
The open reading frame (ORF) for the biosynthetic machineries for homicorcin was found to be organized in a single gene cluster where the structural gene (homA) is followed by the protease (homP), modification genes (homBC) and immunity gene homI in the same orientation (Fig. 1B). Homology of the homicorcin gene cluster products were compared using BLASTp and most of the gene products have high sequence similarity with epicidin 280 gene cluster products (Table 1). The homA gene appears to be the structural gene encoding a 56-amino acid precursor peptide that is cleaved between glutamine (Q) and serine (S) following post-translational modifications by homBC, to form a 30-residue mature homicorcin pre-peptide (Fig. 1C). A 3-oxoacyl-[acyl-carrier-protein] reductase gene (homO) is also present in the cluster that might be responsible for the reduction of the first modified serine residue (Dha) to 2-hydroxypropionate (Hpo) similar to the epicidin 280 gene cluster. The core peptide of homicorcin possesses three Ser, four Thr and three Cys residues. Among them all Ser residues and three Thr residues are predicted to be dehydrated by HomB. One N-terminal Thr, one C-terminal Thr and one C-terminal Ser dehydrated residues are predicted to be cyclized with the C-terminal Cys residues by HomC. Homicorcin leader peptide and pre-peptide were found to be structurally similar to nisin A, pep5, epilancin 15X and epicidin 280 (Fig. 1C).
Purification and MS/MS confirms the degree of dehydration of homicorcin
An activity-guided purification was performed in four steps using ammonium sulfate precipitation extraction, size exclusion column chromatography, Sephadex cation exchange together with the C18 reversed phase-high performance liquid chromatography (RP-HPLC). Antimicrobial activity correlated with the peaks denoted as peak 1 and peak 2 eluted at 40% and 42% acetonitrile concentration respectively in the RP-HPLC chromatogram (Fig. 2A).
Purification of homicorcin through RP-HPLC and mass determination of active peaks by LC–MS. (A) RP-HPLC of active fractions (pooled from ion exchange); Two bioactive (peak-1 and peak-2) fractions were found eluting at 40% and 42% ACN gradient respectively . (B) Strong single peak found in each mass/charge (m/z) state. (C) Mass differences observed between homicorcin and its variant homicorcin 1.
Completely purified antimicrobial fractions (peak-1, peak-2) produced by S. hominis strain MBL _AB63 was traced to the components with a strong signal at [M + 3H]33+ = 1015.5 Da and 1016.2 Da respectively by liquid chromatography-mass spectrometry (LC–MS). LC–MS revealed that the corresponding active fractions have a mass of 3043.2 and 3046.5 Da respectively (Fig. 2B,C).
Correlating the predicted 3150 Da mass of homicorcin calculated from the genome sequence, gave insight into the degree of modification since the difference between these masses corresponded to the dehydration of six water molecules during maturation (18 Da reduction per dehydration). The active RP-HPLC fractions of peak 1 and peak 2 are named as homicorcin 1 and homicorcin respectively. Mass spectrometry revealed that homicorcin 1 is 3 Da larger than homicorcin in molecular mass (Fig. 2C).
Trypsin digestion of purified peptides deduces the possible variations in post translational modification and thio-ether ring position
The homicorcin leader peptide cleavage site and thio-ether ring patterns were predicted using RiPPMiner-Peptide webserver tool and found the cleavage site to be similar to class I lantibiotic cleavage site (Fig. 1C, Fig. S1) and the mature peptide contains three thio-ether rings as shown in Fig. 3A. Ring A is predicted to be between Thr9 and Cys13, ring B is between Thr21 and Cys24 and ring C is between Ser23 and Cys29 (Fig. 3A, Fig. S1).
Proposed structure of Homicorcin. (A) Posttranslationally modified residues are indicated as follows: Dha,- dehydroalanine; Dhb—dehydrobutyrine; Abu—aminobutyric acid; Abu-S-Ala -Methyllanthionine; Dha-S-Ala – Lanthionine. The first residue Dha to be modified to 2-hydroxypropionate (Hpo) in homicorcin 1 is shown in a yellow circle. (B) Trypsin digestion sites of homicorcin and identified fragments by LC–MS (Trypsin digestion sites are indicated by (|) and digested fragments of the peptide are marked as A–F). The digested fragments obtained by trypsin digestion are (blue colored box) A- 653.4 Da, B + C- 814.4 Da, D- 357.2 Da, B + C + D- 1153.4 Da, E + F- 1256.6 Da. A’ is the modified N-terminal 2- hydroxypropionate (Hpo) fragment with a molecular mass of 656.4 Da (B).
In vitro fragmentation and sequencing of homicorcin and homicorcin 1 using Edman degradation poses a significant challenge as they contain several ring structures. The homicorcin possesses five trypsin digestion sites and hence de novo structure prediction using trypsin digestion was used to analyse the modification of amino acids and thio-ether ring positions (Fig. 1, Table 2). The digestion sites located between thio-ether bridges are usually not influenced by the peptidase, hence only five peptide fragments (fragment A, B + C, D, B + C + D, E + F) were found from the trypsin digestion product after RP-HPLC and LC–MS spectroscopy (Table 2, Fig. 3B, Fig. S2). The intensity of E + F fragment that contain predicted rings B and C was found to be very low and could not be reproduced. It should be noted that LC–MS abundances are peptide specific depending on the ionisation response. A potentially poor ionisation response of the fragments with ring structures could lead to underestimation of their abundance. Other tryptic peptide fragments could not be obtained through RP-HPLC possibly due to poor binding to the column or low abundance in the fragment mixture (Table 2). The fragment patterns predicted the position of three ring structures for these active peptides where ring one was found in fragment B + C and interlocking rings two and three were positioned in E + F peptide fragment.
Besides the ring position, mass differences between the active peptides can also be explained from the trypsin digested fragments. The first dehydroalanine (Dha) in homicorcin is predicted to be oxidoreduced to 2-hydroxipropionate (Hpo) that causes a 3 Da mass increases in homicorcin 1. The homO gene product, an oxidoreductase similar to epicidin 280- associated eciO or epilancin 15X-associated elxO is responsible for the production of a mixture of peptides that have dehydroalanine (Dha) and 2-hydroxypropionyl (Hpo) groups at their N termini with a mass difference of 3 Da.
Homicorcin shows inhibitory effects against various Gram-positive strains
Antimicrobial susceptibility testing of homicorcin was done against indicator strains consisting of Staphylococcus simulans 22, Micrococcus luteus ATCC1856, Staphylococcus aureus SG511, Micrococcus luteus DSM1790, Lactococcus lactis NCTC497, Staphylococcus carnosus TM300, methicillin resistant Staphylococcus aureus (MRSA), methicillin sensitive Staphylococcus aureus (MSSA1), Bacillus subtilis 168 and Escherichia coli DH5α (Table 3). Homicorcin was found to exhibit antimicrobial activity only against the Gram-positive bacteria, and not against Gram-negative ones.
The minimum inhibitory concentration (MIC) of homicorcin was compared with nisin A against Staphylococcus simulans 22, Staphylococcus aureus SG511, Micrococcus luteus ATCC1856, methicillin resistant Staphylococcus aureus (MRSA), methicillin sensitive Staphylococcus aureus (MSSA1) and Escherichia coli DH5α (Table 4). Homicorcin was found to be less potent than nisin A against the tested strains.
Mode of antimicrobial mechanism of homicorcin and its variant
To understand the antimicrobial mechanism of homicorcin, inhibitory effect was assessed against Staphylococcus simulans 22 in liquid culture for an extended period of 148 h. CFU/ml was calculated at regular time intervals for both the control and homicorcin treated sample. CFU count decreased significantly when homicorcin was added to the culture medium compared to the control one (Fig. 4). This result indicates that homicorcin has a bactericidal effect on target organisms.
To confirm the bactericidal effect of homicorcin and its variant against Micrococcus luteus ATCC1856, a field emission-scanning electron microcopy (FE-SEM) was performed. The indicator strain Micrococcus luteus ATCC1856 was treated with homicorcin, homicorcin 1 and nisin A under normal growth conditions. In all treated samples morphological changes were observed in the indicator strain with an induced membrane deformation that ultimately led to bacterial cell death. Peptide treated Micrococcus luteus ATCC1856 generated numerous spike-like membrane protrusions on the surface following drastically deformed cell membranes that resulted in the loss of intact cell shape and size (Fig. 5).
Electron microscopy of surface morphological changes of bacteria triggered by homicorcin and its variant homicorcin 1. Electron micrographs showing membrane morphology of Micrococcus luteus ATCC 1856 under the following conditions: (A) without treatment, (B) nisin A-treated and (C) homicorcin and (D) homicorcin 1 treated. {Normal cell (green arrow); Pore on cell membrane (red arrow); Lysed cell with deformed cell membrane ( orange arrow)}.
Discussion
Endophytic bacteria are a source of a plethora of known and unknown novel biologically active metabolites15. The context of this study was set up with an aim to characterize a novel antimicrobial peptide (homicorcin) isolated from jute endophyte Staphylococcus hominis strain MBL_AB63. The preliminary assays for bioactive metabolites confirmed the production of a potential ribosomally synthesised antimicrobial peptide, commonly known as lantibiotic. Purification and characterization of this peptide was carried out using different quantitative and analytical methods including size exclusion, ion exchange and RP-HPLC chromatography and mass spectrometry. Scanning electron microscopy also helped us gain insights about the possible mode of action of homicorcin purified from Staphylococcus hominis strain MBL_AB63.
Lantibiotics are ribosomally synthesized antimicrobial peptides commonly produced by Gram-positive bacteria including genera such as Bacillus, Enterococcus, Micrococcus, Streptococcus, Staphylococcus, Actinomycetes, as well as some endophytic fungi40. However, lantibiotics produced by endophytic bacteria have not been reported so far. According to BACTIBASE (a database dedicated to bacteriocins), till now about 64 different lantibiotics have been reported; but only a few lantibiotics have been commercially applied or are under development for medical use in spite of their promising properties41,42,43. Research on lantibiotics has so far focused on bio-engineering to improve activity, to investigate the structure–activity relationship and to understand the modification process. Therefore, more focused research on clinical applications of lantibiotics is necessary to be carried out.
Lantibiotic biosynthesis requires the coordinated expression of a set of genes44,45. Whole genome sequence analysis for secondary metabolites of S. hominis strain MBL_AB63 identified a complete class I lantibiotic gene cluster. The gene cluster contains five genes (homABCOP) involved in biosynthesis of homicorcin, and one gene potentially involved in immunity (homI) (Table 1; Fig. 1B). The cluster organization resembles that of the lantibiotics, epicidin 28029, epilancin 15X39, and pep546 produced by different strains of staphylococci, suggesting that these clusters have evolved from a common ancestor. Our predicted peptide HomA has high amino acid sequence similarity (82%) to the epicidin 280 precursor peptide (EpiA) with seven amino acid differences (Table 1)29. HomA contains 26 amino acids in its N-terminal region as leader peptide and 30 amino acids in its C-terminal pre-peptide region. The leader region also contains the conserved motif F-(N/D)-L-(N/D/E) and an Ala at position -2 that are characteristic of class I lantibiotics47 (Fig. 1C). Downstream of HomA, there is a putative serine protease HomP which has high amino acid sequence similarity to EciP, the protease involved in the biosynthesis of epicidin 280 (Table 1)29. HomP likely removes the leader peptide before the mature peptide is transported outside the cell, similar to NisP or EpiP that are extracellularly located and remove the leader region once the peptide has been secreted. The cluster also contains HomB, downstream of HomP, which most likely catalyzes the dehydration of Ser or Thr residues of the HomA C-terminal pre-peptide region. Three Ser and three Thr residues are predicted to be dehydrated by HomB. The ORF for homC encodes a protein with high amino acid sequence similarity to EciC, the cyclase responsible for Lan and MeLan ring formation in epicidin 280 (Table 1). Comparing the sequence similarity of homicorcin and closely related peptides it can be predicted that homicorcin contains one Lan and 2 MeLan ring structures in its mature form. Based on the lantibiotic maturation pathway, for homicorcin we can predict that the precursor peptide HomA is modified by the dehydratase HomB and the cyclase HomC to produce the cross-linked peptide. Then, the leader peptide is removed by the protease HomP, producing an N-terminal dehydroalanine (Dha) present in equilibrium with a reduced form of Dha to 2-hydroxypropionate (Hpo) by the enzyme HomO29,39. The HomO of homicorcin similar to the oxidoreductase EciO (Table 1) is hypothesized to be involved in the reduction of N-terminal free Dha to 2-hydroxypropionate (Hpo) in the biosynthesis of homicorcin 129. The role of the N-terminal Hpo in homicorcin 1 is currently unknown. However, N-terminal modifications are common in lantibiotics and include (methyl)lanthionines, disulfides, pyruvate and lactate groups, 2-oxobutyrate groups, and acyl groups. These modifications might protect the terminal residues from exoproteases48,49. The homicorcin gene cluster also possesses a putative immunity gene, homI to protect the producer strain from its own antibiotic. HomI shows 81% amino acid similarity with EciI, the protein responsible for providing immunity to Staphylococcus epidermidis against epicidin 280 (Table 1)29. The Pep5 producing strain also gets protected by the PepI immunity protein which has 72% sequence similarity with EciI29. Therefore, the producer self-protection mechanism is likely to be mediated by HomI and could be based on the same molecular mechanism as the producer self-protection mediated by EciI and PepI.
Purified homicorcin demonstrated stability in a broad range of pH values (pH 4.0 – 10.6) and retained activity even after treatment at 100 °C (data not shown). Although they showed stable activity under biological pH but the activity began to decrease slightly around pH 10.6. Its stability in a broad range of pH values corroborates with previous studies which show that lanthionine rings and dehydrated residues included in a lantibiotic’s structure are highly stable at low pH50. Homicorcin was found to exhibit antimicrobial activity against closely related Gram-positive bacteria. Among the indicator strains tested homicorcin showed potent activity against Staphylococcus spp. and Micrococcus spp. including MRSA (methicillin-resistant Staphylococcus aureus) and MSSA1 (methicillin-susceptible Staphylococcus aureus) strains. Lantibiotics isolated from Staphylococcal spp. are mostly active against different strains of the same group3. Among 16 test strains of S. aureus, Pep5 and epidermin were found to inhibit 14 and 13 strains respectively including Brazilian MRSA clone A/22C51. Gallidermin have bactericidal activity against both MRSA and MSSA strains52. Hominicin displayed potent activity against MRSA ATCC 11,435, and vancomycin-intermediate S. aureus CCARM350135. Nukacin ISK-1 is active against a wide range of Gram positive bacteria including Bacillus and Lactobacillus strains53.
The position of the thio-ether rings in the mature homicorcin peptide was predicted using RiPPMiner-Peptide webserver tool which identified three rings, one before the hinge and other two are interlocking at C-terminal region, quite similar ring patterns to those predicted for epicidin 280 as well29. Trypsin digestion of the core peptide followed by ESI–MS analysis of the digested fragments has been proven to be a useful tool for analysing modifications and ring topology of different lantibiotics in vitro 54,55,56,57,58. Trypsin is the most common protease for generating peptides for MS analysis. Information regarding ring topology is based on the suppression of fragmentation within the rings. Both homicorcin and homicorcin 1 contain five trypsin cleavage sites, two of them are directly adjacent to potential ring structures (Table 1, Fig. 3B). It has been reported that in this case the sites are protected against trypsin cleavage59. Digested fragments of homicorcin and its variant indicate that trypsin cleavage sites are efficiently cleaved except for the sites within the ring structures (Fig. 3B, Table 1). The first tryptic peptide at the N-terminus of homicorcin and its variant was found to have a 3 Da mass difference that strongly correlates with the modification of initial Dha to 2-hydroxypropionate (Hpo) of homicorcin 1.
To understand the mode of action of homicorcin, a growth inhibition assay was performed against Staphylococcus simulans 22. Staphylococcus cell viability counts clearly indicated that peptide treated indicator strain showed significant decrease of viable cells as measured by CFU count over time. Both nisin and pep5 which belong to class I lantibiotic group also show similar bactericidal effect against the target organisms13,60. Cellular morphology of homicorcin treated cells assessed under field emission-scanning electron microscope (FE-SEM) clearly indicated damage on the cell surfaces of treated indicator strain and also shown rapid leakage of K+ ion in the surroundings (data not shown). Homicorcin is a cationic peptide having five positively charged amino acids. Therefore, it can be predicted that like other small cationic peptides, homicorcin can target the cytoplasmic membrane of sensitive cells61,62,63, where they act to dissipate the proton motive force (PMF) through the formation of discrete pores in the cytoplasmic membrane, and thus deprive cells of an essential energy source64. This effect most likely leads to an efflux of small molecules (potassium and amino acids) and results in the arrest of all cellular biosynthesis. Conformational studies of different lantibiotics have shown that, although the peptides are flexible in an aqueous environment65,66, in the presence of a lipophilic environment they adopt an amphipathic conformation with a central hinge region67, which enables the peptides to insert into the bacterial membrane, introduce temporary membrane perturbations and assemble into a pore. Lantibiotics also target the cell wall component lipid II as an alternative mode of action and inhibit cell wall biosynthesis68,69. Although little is known of lantibiotic resistance in comparison to resistance to commercial antibiotics, a few reports have shown that modification of cell wall composition of teichoic acids due to dltA gene alteration70, change of expression of penicillin binding proteins (PBPs)71, composition of cell membrane72 and presence of two-component sensing systems in target organisms like BceRS in B. subtilis73, BraRS of S. aureus74, AnrAB transporter of L. monocytogenes75, VraSR of S. aureus76 may greatly influence the development of lantibiotic resistance.
In conclusion, the newly identified class I lantibiotic homicorcin produced by Staphylococcus hominis strain MBL_AB63 has potent antimicrobial activity against Staphylococci and related species. Moreover, biosynthetic mechanism of this peptide is interesting compared to other reported lantibiotics. Heterologous expression of the homicorcin gene cluster, the current focus of our lab, will provide an in-depth understanding of the role of individual genes involved in the biosynthesis together with a comprehension of the structure function relationship of this peptide. The N-terminal modification of homicorcin might provide the peptide with higher stability and activity.
Material and methods
Homicorcin producer strains and in silico prediction of the lantibiotic gene cluster
The homicorcin producer strain Staphylococcus hominis strain MBL_AB63 was isolated as a jute endophyte in the Molecular Biology Laboratory (MBL), Dept. of Biochemistry and Molecular Biology, University of Dhaka. Whole genome sequencing of MBL_AB63 has been carried out using Illumina MiSeq platform and the genome sequence submitted to NCBI (DDBJ/ENA/GenBank accession number JAELVP000000000) database. For in silico prediction of lantibiotic biosynthesis, BAGEL4 and anti-SMASH 5.0 automated tools were used.
MBL_AB63 culture and homicorcin crude preparation
A single colony of Staphylococcus hominis strain MBL_AB63 was cultured overnight in 5 ml TSB (Tryptic Soy Broth) (OXOID, England) liquid broth at 37 °C. 4.0 ml of overnight culture was added in 1.0 L freshly prepared TSB broth to obtain an OD600 at 0.01 for large scale peptide production and incubated in a shaking incubator for 24 h at 37 °C. As the desired antimicrobial peptide is part of the extracellular proteins, cell free supernatant was collected after centrifugation at 7000 rpm for 10 min at 4 °C. The antimicrobial peptide was extracted and precipitated using 60% ammonium sulfate at 4 °C. The mixture was then centrifuged at 8000 rpm for 30 min at 4 °C and the pellets were dissolved in sterile nano-pure water. Antimicrobial activity was assayed using Staphylococcus simulans 22 as an indicator strain.
Purification of homicorcin and its variant using column chromatography
An activity guided purification of homicorcin and its variant were performed simultaneously using a discrete column chromatography method. Ammonium sulfate precipitated crude protein was applied to a Sephadex G-50 fine (Pharmacia, Uppsala, Sweden) size exclusion column previously equilibrated with 10 mM sodium phosphate buffer containing 50 mM NaCl of pH 6.0 and eluted with the same buffer at a flow rate of 1.0 ml/min. Gel filtrated active pooled fractions were diluted three times with 10 mM sodium phosphate buffer (pH 6.0) and passed through a weak cation exchanger CM Sephadex C-25 (Pharmacia, Uppsala, Sweden) column at a 1.0 ml/min flow rate. The column was washed with 10 mM sodium phosphate buffer (pH 6.0), and active peptides bound to the resin were eluted with 200 ml 10 mM sodium phosphate buffer with 1 M NaCl solution in a linear gradient manner. Reverse phase-high performance liquid chromatography (RP-HPLC) was used as the final purification step. Active pooled samples obtained from ion exchange were injected into RP-HPLC to get completely purified peptide using a multistep gradient using acetonitrile and nano-pure water with 0.05% TFA as mobile phase. Before RP-HPLC separation, all the organic solvents were filtered through polytetrafluoroethylene (PTFE) organic filter (Ultipor® N®66 Nylon 6, 6membrane, 0.45 μm) and deaerated. A reverse phase C18 column (Luna 5u C18 100A, 250 × 10.0 mm particle size 5 μm, pore size 110 A) was used and the peaks were recorded by UV (DAD – Diode Array Detector) detection at 220 nm.
Prediction of homicorcin leader peptide cleavage site and thio-ether cross links
“RiPPMiner-Peptide” tool was used to identify the homicorcin leader peptide cleavage site, cross links and similarity with different RiPPs (ribosomally synthesized and post-translationally modified peptides). RiPPMiner is a machine learning based webserver for deciphering chemical structures of RiPPs. It derives its predictive power using a manually curated database of more than 500 + experimentally characterized RiPPs belonging to 13 subclasses. These classes include lantipeptide, bottromycin, cyanobactin, glycocin, lasso peptide, linearazol, microcin, sactipeptide, thiopeptide, auto inducing peptide etc.
Mass spectrometry analysis (LC–MS/MS)
The LC–MS measurement was performed on a platform of Agilent 6420 LC/TQ, equipped with 1290 Infinity II LC System utilizing an Agilent rapid resolution HD ZORBAX Eclipse Plus C18 column (2.1 X 50 mm, 1.8-μm particle size) and sample purity was monitored by the 1290 Infinity II Diode Array Detector FS at 220 nm and 254 nm. MS measurement was carried out using the standard ESI (electrospray ionization) source that was equipped with turbo ion spray source operating at 350 °C with the fragmentation voltage set to 135 V and collision energy at 35 V.
Mass-spectra were acquired in centroid mode ranging from 100–2000 m/z in positive ionization mode with an auto MS2 fragmentation. Optimization of the MS parameters was conducted using standard solutions of each analyte prepared in acetonitrile: water (1:1) with 0.1% HCOOH and infused at a flow rate of 0.2 ml/min.
De novo peptide sequencing using trypsin digestion
Trypsin is a serine peptidase that predominantly cleaves proteins at the carboxyl side of the amino acids lysine (K) and arginine (R) except when either is bound to a C-terminal proline residue. Trypsin (Gibco ™ Trypsin dissolved in 0.2 N HCl) was added to the peptide solution as a final peptide: protein ratio of 1:20 (w/w) where it was desirable that protein concentration is at least 0.1 mg/ml. Then the sample was incubated overnight at 37 °C. Incubated samples were then applied in RP-HPLC and LC–MS to identify the digested fragments.
Homicorcin susceptibility test
On tryptic soy agar (TSA) plates, indicator strains with appropriate concentrations (106 cells/ml) were overlaid using TSB soft agar (3% TSB and 0.8% agar). Indicator strains used for susceptibility test were Staphylococcus simulans 22, Micrococcus luteus ATCC1856, Staphylococcus aureus SG511, Micrococcus luteus DSM1790, Lactococcus lactis NCTC497, Staphylococcus carnosus TM300, Escherichia coli DH5α. Wells were made over the soft agar and 30 μl of purified peptide was applied. After overnight incubation, wells containing the peptides with antimicrobial activities inhibited the growth of the test strains by forming clear zones around the wells.
Determination of minimum inhibitory concentration (MIC)
The MIC of homicorcin and nisin A were determined twice for each strain by spotting on a lawn antimicrobial assay. The method which has been described for the disk diffusion assays in the National Committee for Clinical Laboratory Standards protocol was used for the preparation of the plate77. The inoculum was prepared by suspension of colonies in sterile solution of TSB, grown overnight and the optical density was adjusted to 0.5 McFarland standard (1 × 108 cells/ml). Homicorcin and nisin A were diluted to obtain concentrations in the range 100.0–0.19 µM, and the MIC was determined after 16 h of incubation at 37 °C.
Determination of bactericidal/bacteriostatic activity
To determine the bacteriostatic or bactericidal nature of homicorcin, Staphylococcus simulans 22 was used as an indicator strain. 8XMIC of homicorcin was applied at the mid-log phase of indicator growth. The colony-forming unit (cfu/ml) was determined for 148 h in both peptide treated and untreated samples. CFU was counted by the spread plate method after plating the cells on TSA plate in tenfold serial dilutions.
Electron microscopy of homicorcin treated bacterial cells
For insights into the surface morphology of Micrococcus luteus ATCC1856 upon treatment with homicorcin and its variant, the bacterial samples were visualized under field emission-scanning electron microscope (FE-SEM) (model Jeol-JSM 7610F). For sample preparation, bacteria were collected in early stationary phase, incubated for growth upon treatment with 5 × MIC of homicorcin and its variant. 1 mL of each bacterial sample was centrifuged at 5000 rpm for 5 min at 4 °C and the pellet obtained was washed twice with 1X PBS buffer (pH-7.4). After washing, pellets were fixed by incubating cells in 0.25% glutaraldehyde in Na-phosphate buffer for 30 min. The fixed pellet was washed with 10 mM Na-phosphate buffer. Sequential dehydration was done using 30%, 50%, 70%, 90%, and absolute ethanol. These dehydrated samples were coated by platinum and visualized under FE-SEM.
References
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30(1), 108–160. https://doi.org/10.1039/c2np20085f.PMID:23165928 (2013).
Chatterjee, C., Paul, M., Xie, L. & Van Der Donk, W. A. Biosynthesis and mode of action of lantibiotics. Chem. Rev. 105(2), 633–684. https://doi.org/10.1021/cr030105v (2005).
de Freire Bastos, M. D. C., Miceli de Farias, F., Carlin Fagundes, P. & Varella Coelho, M. L. Staphylococcins: An update on antimicrobial peptides produced by staphylococci and their diverse potential applications. Appl. Microbiol. Biotechnol. 104(24), 10339–10368. https://doi.org/10.1007/s00253-020-10946-9 (2020).
Mohr, K. I. et al. Pinensins: The first antifungal lantibiotics. Angew Chem. Int. Ed. Engl. 54(38), 11254–11258. https://doi.org/10.1002/anie.201500927 (2015).
Caetano, T., van der Donk, W. & Mendo, S. Bacteroidetes can be a rich source of novel lanthipeptides: The case study of Pedobacter lusitanus. Microbiol. Res. 235, 126441. https://doi.org/10.1016/j.micres.2020.126441 (2020).
Bierbaum, G. & Sahl, H. G. Lantibiotics: mode of action, biosynthesis and bioengineering. Curr. Pharm. Biotechnol. 10(1), 2–18. https://doi.org/10.2174/138920109787048616 (2009).
Götz, F., Perconti, S., Popella, P., Werner, R. & Schlag, M. Epidermin and gallidermin: Staphylococcal lantibiotics. Int. J. Med. Microbiol. 304(1), 63–71. https://doi.org/10.1016/j.ijmm.2013.08.012 (2014).
Marsh, A. J., O’Sullivan, O., Ross, R. P., Cotter, P. D. & Hill, C. In silico analysis highlights the frequency and diversity of type 1 lantibiotic gene clusters in genome sequenced bacteria. BMC Genom. 11(1), 1–21. https://doi.org/10.1186/1471-2164-11-679 (2010).
Nagao, J. I. et al. Lantibiotics: Insight and foresight for new paradigm. J. Biosci. Bioeng. 102(3), 139–149. https://doi.org/10.1263/jbb.102.139 (2006).
Smith, R. & Coast, J. The true cost of antimicrobial resistance. BMJ 346, f1493. https://doi.org/10.1136/bmj.f1493 (2013).
Ogden, J. Tackling antimicrobial resistance: The progress so far. Prescr. 30(11), 27–31. https://doi.org/10.1002/psb.1803 (2019).
Taubes, G. The bacteria fight back. Science 321, 356–361. https://doi.org/10.1126/science.321.5887.356 (2008).
Breukink, E. & de Kruijff, B. Lipid II as a target for antibiotics. Nat. Rev. Drug. Discov. 5(4), 321–332. https://doi.org/10.1038/nrd2004 (2006).
Liu, L. et al. Isoprenylated chromone derivatives from the plant endophytic fungus Pestalotiopsis fici. J. Nat. Prod. 72(8), 1482–1486. https://doi.org/10.1021/np900308s (2009).
Guo, B., Wang, Y., Sun, X. & Tang, K. Bioactive natural products from endophytes: a review. Appl. Biochem. Microbiol. 44(2), 136–142. https://doi.org/10.1134/S0003683808020026 (2008).
Bodenhausen, N., Bortfeld-Miller, M., Ackermann, M. & Vorholt, J. A. A synthetic community approach reveals plant genotypes affecting the phyllosphere microbiota. PLoS Genet. 10(4), e1004283. https://doi.org/10.1371/journal.pgen.1004283 (2014).
Doolotkeldieva, T., Bobusheva, S. & Suleymankisi, A. Biological Control of Erwinia carotovora spp carotovora by Streptomyces Species. Adv. Microbiol. 6(02), 104. https://doi.org/10.4236/aim.2016.62011 (2016).
Anand, R., Grayston, S. & Chanway, C. N2-fixation and seedling growth promotion of lodgepole pine by endophytic Paenibacillus polymyxa. Microb. Ecol. 66(2), 369–374. https://doi.org/10.1007/s00248-013-0196-1 (2013).
Haidar, B. et al. Population diversity of bacterial endophytes from jute (Corchorus olitorius) and evaluation of their potential role as bioinoculants. Microbiol. Res. 208, 43–53. https://doi.org/10.1016/j.micres.2018.01.008 (2018).
Hallmann, J., Quadt-Hallmann, A., Rodrıguez-Kabana, R. & Kloepper, J. W. Interactions between Meloidogyne incognita and endophytic bacteria in cotton and cucumber. Soil Biol. Biochem. 30(7), 925–937. https://doi.org/10.1016/S0038-0717(97)00183-1 (1998).
Rajendran, G., Sing, F., Desai, A. J. & Archana, G. Enhanced growth and nodulation of pigeon pea by co-inoculation of Bacillus strains with Rhizobium spp. Bioresour. Technol. 99(11), 4544–4550. https://doi.org/10.1016/j.biortech.2007.06.057 (2008).
Joseph, B. & Priya, R. M. Bioactive Compounds from Endophytes and their Potential in. Am. J. Biochem. Mol. Biol. 1(3), 291–309. https://doi.org/10.3923/ajbmb.2011.291.309 (2011).
Pimentel, M. R., Molina, G., Dionísio, A. P., Maróstica Junior, M. R. & Pastore, G. M. The use of endophytes to obtain bioactive compounds and their application in biotransformation process. Biotechnol. Res. Int. https://doi.org/10.4061/2011/576286 (2011).
Shukla, S. T. et al. Hepatoprotective and antioxidant activities of crude fractions of endophytic fungi of Ocimum sanctum Linn in rats. Orient. Pharm. Exp. Med. 12(2), 81–91. https://doi.org/10.1007/s13596-012-0061-7 (2012).
Zhao, J., Shan, T., Mou, Y. & Zhou, L. Plant-derived bioactive compounds produced by endophytic fungi. Mini Rev. Med. Chem. 11(2), 159–168. https://doi.org/10.2174/138955711794519492 (2011).
Dickman, R., Mitchell, S. A., Figueiredo, A. M., Hansen, D. F. & Tabor, A. B. Molecular recognition of lipid II by lantibiotics: synthesis and conformational studies of analogues of nisin and mutacin rings A and B. J. Org. Chem. 84(18), 11493–11512. https://doi.org/10.1021/acs.joc.9b01253 (2019).
Jalgaonwala, R. E., Mohite, B. V. & Mahajan, R. T. A review: natural products from plant associated endophytic fungi. J. Microbiol. Biotechnol. Res. 1(2), 21–32 (2011).
Fontana, M. B., de Bastos, M. & C., & Brandelli, A. ,. Bacteriocins Pep5 and epidermin inhibit Staphylococcus epidermidis adhesion to catheters. Curr. Microbiol. 52, 350–353 (2006).
Heidrich, C. et al. Isolation, characterization, and heterologous expression of the novel lantibiotic epicidin 280 and analysis of its biosynthetic gene cluster. Appl. Environ. Microbiol. 64(9), 3140–3146. https://doi.org/10.1128/AEM.64.9.3140-3146.1998 (1998).
van de Kamp, M. et al. Elucidation of the primary structure of the lantibiotic epilancin K7 from Staphylococcus epidermidis K7 Cloning and characterisation of the epilancin-K7-encoding gene and NMR analysis of mature epilancin K7. Eur. J. Biochem. 230(2), 587–600. https://doi.org/10.1111/j.1432-1033.1995.tb20600.x (1995).
Ekkelenkamp, M. B. et al. Isolation and structural characterization of epilancin 15X, a novel lantibiotic from a clinical strain of Staphylococcus epidermidis. FEBS Lett. 579, 1917–1922 (2005).
Schnell, N. et al. Prepeptide sequence of epidermin, a ribosomally synthesized antibiotic with four sulphide-rings. Nature 333(6170), 276–278. https://doi.org/10.1038/333276a0 (1988).
Sahl, H. G. Staphylococcin 1580 is identical to the lantibiotic epidermin: implications for the nature of bacteriocins from gram-positive bacteria. Appl. Environ. Microbiol. 60, 752–755 (1994).
Kellner, R. et al. Gallidermin: a new lanthionine-containing polypeptide antibiotic. Eur. J. Biochem. 177, 53–59 (1988).
Kim, P. I. et al. Characterization and structure identification of an antimicrobial peptide, hominicin, produced by Staphylococcus hominis MBBL 2–9. Biochem. Biophys. Res. Commun. 399, 133–138 (2010).
Sashihara, T. et al. A novel lantibiotic, nukacin ISK-1, of Staphylococcus warneri ISK-1: cloning of the structural gene and identification of the structure. Biosci. Biotechnol. Biochem. 64(11), 2420–2428. https://doi.org/10.1271/bbb.64.2420 (2000).
Minamikawa, M., Kawai, Y., Inoue, N. & Yamazaki, K. Purification and characterization of warnericin RB4, anti-Alicyclobacillus bacteriocin, produced by Staphylococcus warneri RB4. Curr. Microbiol. 51, 22–26 (2005).
Navaratna, M. A., Sahl, H.-G. & Tagg, J. R. Two-component anti-Staphylococcus aureus lantibiotic activity produced by Staphylococcus aureus C55. Appl. Environ. Microbiol. 64, 4803–4808 (1998).
Velásquez, J. E., Zhang, X. & van der Donk, W. A. Biosynthesis of the antimicrobial peptide epilancin 15X and its N-terminal lactate. Chem. Biol. 18(7), 857–867. https://doi.org/10.1016/j.chembiol.2011.05.007 (2011).
Cooper, L. E., Li, B. & van der Donk, W. A. Biosynthesis and Mode of Action of Lantibiotics. Compr. Nat. Prod. II(5), 217–256. https://doi.org/10.1016/B978-008045382-8.00116-7 (2010).
Repka, L. M., Chekan, J. R., Nair, S. K. & van der Donk, W. A. Mechanistic understanding of lanthipeptide biosynthetic enzymes. Chem. Rev. 117(8), 5457–5520. https://doi.org/10.1021/acs.chemrev.6b00591 (2017).
Dischinger, J., Wiedemann, I., Bierbaum, G., & Sahl, H. G. Lantibiotics. In Handbook of Biologically Active Peptides (ed. Abba Kastin, J.), Second edition, Academic Press. 119–128. https://doi.org/10.1016/B978-0-12-385095-9.00019-1. (2013).
van Heel, A. J., Montalban-Lopez, M. & Kuipers, O. P. Evaluating the feasibility of lantibiotics as an alternative therapy against bacterial infections in humans. Expert. Opin. Drug Metab. Toxicol. 7(6), 675–680. https://doi.org/10.1517/17425255.2011.573478 (2011).
Will, S. E. et al. The limits to growth–energetic burden of the endogenous antibiotic tropodithietic acid in Phaeobacter inhibens DSM 17395. PLoS ONE 12(5), e0177295. https://doi.org/10.1371/journal.pone.0177295 (2017).
Montalbán-López, M., van Heel, A. J. & Kuipers, O. P. Employing the promiscuity of lantibiotic biosynthetic machineries to produce novel antimicrobials. FEMS Microbiol. Rev. 41(1), 5–18. https://doi.org/10.1093/femsre/fuw034 (2016).
Meyer, C. et al. Nucleotide sequence of the lantibiotic Pep5 biosynthetic gene cluster and functional analysis of PepP and PepC. Evidence for a role of PepC in thioether formation. Eur. J. Biochem. 232(2), 478–489. https://doi.org/10.1111/j.1432-1033.1995.tb20834.x (1995).
van der Meer, J. R. et al. Influence of amino acid substitutions in the nisin leader peptide on biosynthesis and secretion of nisin by Lactococcus lactis. J. Biol. Chem. 269(5), 3555–3562 (1994).
Cooper, L. E., McClerren, A. L., Chary, A. & van der Donk, W. A. Structure-activity relationship studies of the two-component lantibiotic haloduracin. Chem. Biol. 15(10), 1035–1045. https://doi.org/10.1016/j.chembiol.2008.07.020 (2008).
McClerren , A. L., Cooper, L. E., Quan, C., Thomas, P. M., Kelleher, N. L., & van der Donk, W. A. Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic. Pro. Nat. Acad. Sci.103 (46), 17243–17248; DOI: https://doi.org/10.1073/pnas.0606088103. (2006)
McAuliffe, O., Ross, R. P. & Hill, C. Lantibiotics: structure, biosynthesis and mode of action. FEMS Microbiol. Rev. 25(3), 285–308. https://doi.org/10.1111/j.1574-6976.2001.tb00579.x (2001).
Nascimento, J. S. et al. Bacteriocins as alternative agents for control of multiresistant staphylococcal strains. Lett. Appl. Microbiol. 42, 215–221 (2006).
Bengtsson, T., Lönn, J., Khalaf, H. & Palm, E. The lantibiotic gallidermin acts bactericidal against Staphylococcus epidermidis and Staphylococcus aureus and antagonizes the bacteria-induced proinflammatory responses in dermal fibroblasts. Microbiologyopen. 7, e00606 (2018).
Asaduzzaman, S. M. et al. Nukacin ISK-1, a bacteriostatic lantibiotic. Antimicrob. Agents Chemother. 53(8), 3595–3598. https://doi.org/10.1128/AAC.01623-08 (2009).
Caetano, T., Krawczyk, J. M., Mösker, E., Süssmuth, R. D. & Mendo, S. Heterologous expression, biosynthesis, and mutagenesis of type II lantibiotics from Bacillus licheniformis in Escherichia coli. Chem. Biol. 18(1), 90–100. https://doi.org/10.1016/j.chembiol.2010.11.010 (2011).
Garg, N., Tang, W., Goto, Y., Nair, S. K. & van der Donk, W. A. Lantibiotics from Geobacillus thermodenitrificans. Proc. Natl. Acad. Sci. U.S.A. 109(14), 5241–5246. https://doi.org/10.1073/pnas.1116815109 (2012).
Tang, W., Jiménez-Osés, G., Houk, K. N. & van der Donk, W. A. Substrate control in stereoselective lanthionine biosynthesis. Nat. Chem. 7, 57–64 (2015).
Shi, Y., Yang, X., Garg, N. & van der Donk, W. A. Production of lantipeptides in Escherichia coli. J. Am. Chem. Soc. 133(8), 2338–2341. https://doi.org/10.1021/ja109044r (2011).
Li, B. et al. Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proc. Natl. Acad. Sci. 107(23), 10430–10435. https://doi.org/10.1073/pnas.0913677107 (2010).
Lubelski, J., Overkamp, W., Kluskens, L. D., Moll, G. N. & Kuipers, O. P. Influence of shifting positions of Ser, Thr, and Cys residues in prenisin on the efficiency of modification reactions and on the antimicrobial activities of the modified prepeptides. Appl. Environ. Microbiol. 74(15), 4680–4685. https://doi.org/10.1128/AEM.00112-08 (2008).
Wiedemann, I. et al. Specific binding of nisin to the peptidoglycan precursor lipid II combines pore formation and inhibition of cell wall biosynthesis for potent antibiotic activity. J. Biol. Chem. 276(3), 1772–1779. https://doi.org/10.1074/jbc.M006770200 (2001).
Van Belkum, M. J. et al. The bacteriocin lactococcin A specifically increases permeability of lactococcal cytoplasmic membranes in a voltage-independent, protein-mediated manner. J. Bacteriol. Res. 173(24), 7934–7941. https://doi.org/10.1128/jb.173.24.7934-7941.1991 (1991).
Kordel, M. & Sahl, H. G. Susceptibility of bacterial, eukaryotic and artificial membranes to the disruptive action of the cationic peptides Pep 5 and nisin. FEMS Microbiol. Lett. 34(2), 139–144. https://doi.org/10.1111/j.1574-6968.1986.tb01393.x (1986).
Abee, T., Klaenhammer, T. R. & Letellier, L. Kinetic studies of the action of lactacin F, a bacteriocin produced by Lactobacillus johnsonii that forms poration complexes in the cytoplasmic membrane. Appl. Environ. Microbiol. 60(3), 1006–1013. https://doi.org/10.1128/AEM.60.3.1006-1013.1994 (1994).
Montville, T. J. & Bruno, M. E. C. Evidence that dissipation of proton motive force is a common mechanism of action for bacteriocins and other antimicrobial proteins. Int. J. Food Microbiol. 24(1–2), 53–74. https://doi.org/10.1016/0168-1605(94)90106-6 (1994).
van de Ven, F. J., van den Hooven, H. W., Konings, R. N. & Hilbers, C. W. NMR studies of lantibiotics: the structure of nisin in aqueous solution. Eur. J. Biochem. 202(3), 1181–1188. https://doi.org/10.1111/j.1432-1033.1991.tb16488.x (1991).
Lian, L. Y. et al. Solution structures of nisin A and its two major degradation products determined by NMR. Biochem. J. 283(2), 413–420. https://doi.org/10.1042/bj2830413 (1992).
van den Hooven, H. W. et al. NMR and circular dichroism studies of the lantibiotic nisin in non-aqueous environments. FEBS Lett. 319(1–2), 189–194. https://doi.org/10.1016/0014-5793(93)80065-3 (1993).
Islam, M. R. et al. Ring A of nukacin ISK-1: a lipid II-binding motif for type-A (II) lantibiotic. J. Am. Chem. Soc. 134(8), 3687–3690. https://doi.org/10.1021/ja300007h (2012).
Breukink, E. et al. Lipid II is an intrinsic component of the pore induced by nisin in bacterial membranes. J. Biol. Chem. 278(22), 19898–19903. https://doi.org/10.1074/jbc.M301463200 (2003).
Kovács, M. et al. A functional dlt operon, encoding proteins required for incorporation of d-alanine in teichoic acids in gram-positive bacteria, confers resistance to cationic antimicrobial peptides in Streptococcus pneumoniae. J. Bacteriol. 188(16), 5797–5805. https://doi.org/10.1128/JB.00336-06 (2006).
Gravesen, A., Sørensen, K., Aarestrup, F. M. & Knøchel, S. Spontaneous nisin-resistant Listeria monocytogenes mutants with increased expression of a putative penicillin-binding protein and their sensitivity to various antibiotics. Microb. Drug Resist. 7(2), 127–135. https://doi.org/10.1089/10766290152045002 (2001).
Mazzotta, A. S. & Montville, T. J. Nisin induces changes in membrane fatty acid composition of Listeria monocytogenes nisin-resistant strains at 10 degrees C and 30 degrees C. J. Appl. Microbiol. 82(1), 32–38. https://doi.org/10.1111/j.1365-2672.1997.tb03294.x (1997).
Dintner, S. et al. Coevolution of ABC transporters and two-component regulatory systems as resistance modules against antimicrobial peptides in Firmicutes Bacteria. J. Bacteriol. 193(15), 3851–3862. https://doi.org/10.1128/JB.05175-11 (2011).
Hiron, A., Falord, M., Valle, J., Débarbouillé, M. & Msadek, T. Bacitracin and nisin resistance in Staphylococcus aureus: a novel pathway involving the BraS/BraR two-component system (SA2417/SA2418) and both the BraD/BraE and VraD/VraE ABC transporters. Mol. Microbiol. 81(3), 602–622. https://doi.org/10.1111/j.1365-2958.2011.07735.x (2011).
Collins, B., Curtis, N., Cotter, P. D., Hill, C. & Ross, R. P. The ABC transporter AnrAB contributes to the innate resistance of Listeria monocytogenes to nisin, bacitracin, and various beta-lactam antibiotics. Antimicrob. Agents. Chemother. 54(10), 4416–4423. https://doi.org/10.1128/AAC.00503-10 (2010).
Kawada-Matsuo, M. et al. Three distinct two-component systems are involved in resistance to the class I bacteriocins, Nukacin ISK-1 and nisin A Staphylococcus aureus. PLoS ONE 8(7), e69455. https://doi.org/10.1371/journal.pone.0069455 (2013).
Kiehlbauch, J. A. et al. Use of the national committee for clinical laboratory standards guidelines for disk diffusion susceptibility testing in New York state laboratories. J. Clin. Microbiol. 38(9), 3341–3348. https://doi.org/10.1128/JCM.38.9.3341-3348.2000 (2000).
Acknowledgements
This study was partially supported by Ministry of Education, GoB GARE project no. 37.20.0000.004.033.020.2016. We deeply acknowledge Dipa Islam, Biomedical and Toxicological Research Institute (BTRI) and Institute of National Analytical Research and Service (INARS), BCSIR, Dhaka, Bangladesh. We are grateful to Prof. Dr. Gabriele Bierbaum, University of Bonn, Germany, Prof. Dr. Takeshi Zendo, Kyushu University, Japan and Prof. Dr. Kazuhisa Sekimizu, Teikyo University, Japan for their helpful advice and support.
Author information
Authors and Affiliations
Contributions
S.A., M.F., B.H., M.R.I. and H.K. conceived the study. S.A., M.A.U., M.R.I and H.K. designed the experiments. S.A., M.A.U., B.H., M.F., A.A., S.I.M.J. and M.R.I. performed the experiments and analysed the data. S.A., M.F., M.R.I. and H.K. wrote the manuscript and all authors critically revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aftab Uddin, M., Akter, S., Ferdous, M. et al. A plant endophyte Staphylococcus hominis strain MBL_AB63 produces a novel lantibiotic, homicorcin and a position one variant. Sci Rep 11, 11211 (2021). https://doi.org/10.1038/s41598-021-90613-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90613-9
This article is cited by
-
Rhizobacteria associated with Parkinsonia aculeata L. under semi desertic drought and saline conditions
Biologia (2024)
-
Impact of the endophytic and rhizospheric bacteria on crop development: prospects for advancing climate-smart agriculture
Journal of Crop Science and Biotechnology (2023)
-
Isolation and identification of grapevine endophytic bacteria with antagonistic potential against Fomitiporia mediterranea, a pathogen involved in grapevine trunk disease
Journal of Plant Diseases and Protection (2023)
-
Antimicrobial Peptides Against Microbial Biofilms: Efficacy, Challenges, and Future Prospect
International Journal of Peptide Research and Therapeutics (2023)
-
Strategies to access biosynthetic novelty in bacterial genomes for drug discovery
Nature Reviews Drug Discovery (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.