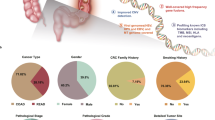
Abstract
Colorectal cancer (CRC) is a complex disease that can be caused by a spectrum of genetic variants ranging from low to high penetrance changes, that interact with the environment to determine which individuals will develop the disease. In this study, we sequenced 20 early-onset CRC patients to discover novel genetic variants that could be linked to the prompt disease development. Eight genes, CHAD, CHD1L, ERCC6, IGTB7, PTPN13, SPATA20, TDG and TGS1, were selected and re-sequenced in a further 304 early onset CRC patients to search for rare, high-impact variants. Although we found a recurring truncating variant in the TDG gene shared by two independent patients, the results obtained did not help consolidate any of the candidates as promising CRC predisposing genes. However, we found that potential risk alleles in our extended list of candidate variants have a tendency to appear at higher numbers in younger cases. This supports the idea that CRC onset may be oligogenic in nature and may show molecular heterogeneity. Further, larger and robust studies are thus needed to unravel the genetics behind early-onset CRC development, coupled with novel functional analyses and omic approaches that may offer complementary insight.
Similar content being viewed by others
Introduction
Heritability in colorectal cancer (CRC) is estimated to be between 8 and 40%1,2. However, only 5–10% is explained by rare, high-penetrance germline mutations in Mendelian susceptibility genes, such as those in Lynch syndrome—caused by pathogenic variants in mismatch repair (MMR) genes, adenomatous polyposis syndromes—caused by pathogenic variants in APC, MUTYH, NTHL1 or in the exonuclease domain of POLE and POLD13,4.
Except for the identification of RPS20 as a causal gene for hereditary MMR-proficient nonpolyposis CRC, the studies undertaken in the past decade to identify new nonpolyposis CRC predisposing genes, have been mostly unsuccessful5,6. This suggests that the missing CRC heritability is presumably complex and polygenic in nature, and is caused by common, low-penetrance risk variants (such as those identified by genome-wide association studies), or by moderately penetrant rarer variants playing an important role in modulating neoplastic transformation7,8.
In this study, we performed whole-exome sequencing (WES) in 20 unrelated patients diagnosed with non-polyposis CRC at young age. Our purpose was to identify novel rare pathogenic variants that could explain such a premature occurrence of the disease.
Results
Patient description and sequencing
Whole-exome sequencing was performed on 20 early-onset CRC patients (< = 50 years). Median age at diagnosis was 45 years. All presented with MMR-proficient tumours, assessed as the conserved expression of the four MMR proteins MLH1, MSH2, MSH6 and PMS2. The clinical features of the patients are described in Supplementary Table 1.
Germline DNA obtained from peripheral blood was sequenced to achieve an average depth of 62.82X, where 77% of the targeted regions were covered by ≥ 10 reads. This provides a good assessment of inherited variant presence. After alignment, calling and annotation, a median of 76,122 variants were identified per individual (see Materials and Methods for a detailed description). No pathogenic variants were found in any of the known hereditary cancer genes that could account for the observed phenotype.
Variant prioritization
Raw data were analyzed with a prioritization pipeline to reduce the number of candidate variants. Briefly, from the initial set of 337,011 unique variants, only those common to both the LifeScope™ (Thermo Fisher Scientific, MA, USA) and GATK calling algorithms were selected9, provided that they were found in the same patient and with the same genotype (n = 218,803). Variants annotated as synonymous or unknown were removed (n = 187,144), followed by restriction to exonic and splicing changes (n = 34,283). Common variants (MAF > 1% and > 0.1% for homozygous and heterozygous calls, respectively) were filtered out (n = 6,716 selected rare variants). Protein truncating—frameshift, nonsense -, variants at canonical splice sites, and missense changes predicted to have high-functional impact by at least 3 in silico predictors (Dann, Polyphen 2, CADD) and/or present at highly conserved positions (GERP + + ≥ 2), were selected (n = 358)10,11,12,13. Lastly, to exclude population frequency mismatches, we eliminated any variants found in a Spanish cohort of 267 non-cancer controls from the MPG cohort of the CIBERER Spanish Variant Server14. A total of 262 variants in 254 genes were finally selected after the implementation of this prioritization algorithm (Supplementary Table 2).
Data analysis
Variant and gene-based analysis
Considering this extended list of 254 candidate genes (Supplementary Table 2), there was a median number of 15 variants and 17.5 alleles per patient. We find a tendency for younger patients to have a higher number of rare, high-impact variants. This correlation is both true for the number of variants (p = 0.023) and the number of alleles (p = 0.033) (Fig. 1). Eight genes were found to have 2 candidate variants each after prioritization, and we found recurring variants, present in at least 3 individuals, in 11 genes The low presence of variants in recurring genes precluded us from doing gene-burden tests from the candidate gene list.
Relationship between number of candidate variants/alleles and age. Scatter plots depicting the significant trend between number of (A) risk variants (p = 0.021) and (B) risk alleles (p = 0.033) and age in the discovery cohort. Figure was created using Stata v11 (StataCorp—www.stata.com).
Pathway-based analysis
We performed pathway-based enrichment analyses based on KEGG pathways (https://www.genome.jp/kegg/) based on all 262 variants, to test whether there was an overrepresentation of potentially damaging rare variants in our cohort based on cancer-related pathways15. We focused on the following terms: Wnt signalling, TGF-β signalling, DNA repair, Pathways in cancer, Apoptosis, Cell adhesion molecules, Colorectal cancer, DNA replication, Hippo signalling pathway, MAPK signalling pathway, MicroRNAs in cancer, PI3K-Akt and PPAR signalling. However, none of the results were statistically significant after performing Fisher´s exact tests (Supplementary Table 3). We also performed hypothesis-free Gene Ontology enrichment analyses with PANTHER using Reactome pathway descriptions, also with no statistically significant results (Supplementary Table 4)16. Interestingly however, some of the top hit pathways correspond to known carcinogenetic routes, such as TET1 demethylation, ion channel transport or glycogenolysis (as per the Warburg effect)17.
Candidate gene selection
We selected 8 genes for replication in an independent familial/early-onset CRC cohort: CHAD, CHD1L, ERCC6, ITGB7, PTPN13, SPATA20, TDG and TGS1 (Table 1)18,19,20,21. These were selected based on their prior description in early-onset sequencing studies, involvement on cancer-related KEGG pathways, presence of the mRNA/protein in healthy colonic mucosa (Protein Atlas, www.proteinatlas.org), or variants found in patients under 40 years (Supplementary Table 2).
Candidate gene resequencing
Replication cohort
The replication cohort consisted of 304 non-related, MMR-proficient, familial and/or early-onset non-polyposis CRC patients recruited at the Hereditary Cancer Program of the Catalan Institute of Oncology, IDIBELL (Catalonia, Spain) (Supplementary Table 5)22.
Mutational screening of CHAD, CHD1L, ERCC6, ITGB7, PTPN13, SPATA20, TDG and TGS1 was performed using a combination of PCR amplification in pooled DNAs and targeted massively parallel sequencing (See Materials and Methods for detailed description).
Candidate-gene variants
Prioritization filters were applied to the results of the targeted sequencing of the 8 genes, in an identical manner to those in the discovery phase. Thirteen variants in 6 of the candidate genes were found to fulfil our criteria. All variants were found in heterozygosity. None of the patients in the replication phase had more than one rare, high-impact variant in any of these genes (Table 2).
Interestingly, the same variant in TDG (p.(Q23X)) was found in a single patient form the validation cohort as well as in the discovery cohort. We hence performed immunohistochemistry (IHC) assays on tumour biopsy samples from one of the patients to assess potential second allele inactivation. However, these revealed no differences in protein expression between the tumour tissue and the adjacent normal (Supplementary Figure S1).
Next, we compared the results found in our discovery and replication round with those found in the MPG Spanish reference cohort (control population)14. Twenty-three rare, high impact variants in 7 of the candidate genes were found in this cohort (Supplementary Table 5). Furthermore, we found no evidence for enrichment in rare variants in these 7 genes when comparing the cumulative allele count in the CRC patients with those in the MPG Spanish population reference dataset (Supplementary Table 6).
In parallel, we also looked into the prevalence of high-impact, rare variants in the eight candidate genes in the data published by Chubb and colleagues19. This included data on 1,006 early-onset familial CRC cases and 1609 healthy controls. Of the eight candidate genes, only TDG showed enrichment for nonsense, frameshift, missense predicted to be damaging and splice donor/acceptor-site variants in early-onset CRC patients compared to controls (n 8/1006 cases vs. 4/1609 controls) (Supplementary Table 7).
TCGA germline variants in the candidate genes
TCGA germline samples from colorectal cancer patients (from the COAD and READ cohorts; n = 219 Caucasians) were used to search for further evidence of variants in the 8 replicated genes being relevant to early-onset CRC23. Variants were assessed in the same way as described for the prioritization algorithm in “Variant prioritization” section. We found 21 heterozygous, rare, high-impact germline variants in 5 of the 8 replicated genes (Table 3). Seven of these were present in patients that had been diagnosed at 50 or earlier. We also observed a shift of variants in the complete list of genes in younger TCGA patients, although in this case, it was not statistically significant (p = 0.817) (Supplementary Fig. S2).
Tumour variation profiles were also inspected in the search for a second somatic hit in the TCGA patients carrying these mutations (https://portal.gdc.cancer.gov/). We found an additional somatic missense variant (p.(P677L)) in a patient carrying the germline ERCC6 p.(R683Q) change. Because ERCC6 is implicated in DNA nucleotide-excision repair (NER) repair, we inspected the mutation signature profile of the tumour with the help of the MuSiCa software24. It showed that signature 3 (associated with homologous recombination) was actually the most prevalent, whereas patterns related to NER (albeit not with CRC), such as signatures 4, 7, 11, 22 and 24, contributed only marginally, which did not support our hypothesis that the ERCC6 mutations were the drivers behind the early development of the tumour (Supplementary Table 8) (https://cancer.sanger.ac.uk/cosmic/signatures_v2.tt).
Interestingly, for the remaining TCGA patients carrying germline changes in these candidate genes, over half of these mutation profiles showed a predominant effect of signature 3. This signature has been related to deficient repair processes involving BRCA1 mutations but was not expected directly for a tumour arising from the variants in our candidate genes (Supplementary Table 8).
Discussion
In this study, we performed whole-exome sequencing to discover novel rare CRC susceptibility variants that could be responsible for the early onset of disease observed in these patients. For this purpose, we selected candidate high-impact rare variants obtained from the sequencing of 20 unrelated early-onset CRC patients. Then, we selected 8 genes as potential candidates, to be replicated in a validation cohort.
Remarkably, TDG turned out to be the most interesting gene in our analyses. It is a BER gene that has been proposed as a tumour suppressor as well as a Wnt pathway regulator and epigenetic modifier in CRC25,26,27,28,29. In our study, a TDG truncating variant p.(Q23X) appeared in two unrelated individuals. Unfortunately, the IHC results did not show any differences between normal and tumour protein levels to account for a second somatic event. Hence, the results obtained for these 8 genes from our analyses did not provide support enough to claim any of them as a strong candidate for CRC susceptibility19.
The causes for this lack of conclusive results may be several. Firstly, the fact that all patients present with an early onset of the disease does not necessarily mean that the underlying genetic cause is homogeneous4. This is supported by the fact that we hardly found recurring variants or genes in our complete list of rare high-impact variants. If so, then larger discovery cohorts would be needed to assess this heterogeneity reliably.
Secondly, CRC development is likely due to arise from the interaction of multiple genetic variants plus environmental factors (i.e. it is oligogenic in nature)30,31. Interestingly, we found a trend that younger patients have a higher proportion of rare, high-impact variants and alleles. Moreover, we do observe that some of the top hits in our pathway analyses correspond to known CRC carcinogenetic pathways, including TET1 methylation32,33, ion channel transport34 and colorectal carcinogenesis35,36. These would support the idea of multiple, moderately penetrant variants being responsible for the early-onset phenotype.
Lastly, the vast amount of data produced by sequencing makes it necessary to utilize prioritization algorithms that are somewhat arbitrary. The approach chosen for prioritization depends on prior knowledge of the patient selection criteria (as described above), the damaging effect of variants encountered in the genes, the inheritance model and the function of the genes affected, which is often (if not always) unavailable37. In this complicated scenario, it is then easy to envisage why not only this current study, but also other previous works inspecting the genetic contribution to early onset CRC development have been underwhelming. In this sense, other strategies may be pursued in order to explore the data more comprehensively. For instance, integration of genetic variation with other data sources such as transcriptomic gene expression of methylation levels may be useful in prioritizing candidates in a more meaningful way38.
In any case, it is guaranteed that larger studies are needed. These ought to be appropriately designed and powered to detect the expected genetic heterogeneity. In the era of Open Data Science, we must hence walk towards making coordinated efforts in order to obtain robust results, particularly for cases that are rare within the CRC spectrum, and that are presumably molecularly heterogeneous. Hopefully too, the near future may also facilitate data interpretation via complementation with functional data, such as CRISPR assays or in vitro organoid models, which would be certainly helpful to increase the throughput in the functional screening of candidate genes, and may prove invaluable in validating novel CRC susceptibility loci.
Materials and methods
Study patients: discovery cohort
The study received the approval of the Ethics Committee (CEIC Comité Ético de Investigación Clínica de Galicia (2011-123). The discovery dataset consisted of 20 non-related and unexplained MMR-proficient CRC cases from the EPICOLON consortium39. All patients received informed consent and protocols were at all times in accordance with the tenets of the declaration of Helsinki. The selected patients were all diagnosed with CRC before/at the age of 50, with tumours showing microsatellite stability and/or conserved expression of the MMR proteins (MLH1, MSH2, MSH6, and PMS2). Patients either showed apparently sporadic, autosomal recessive or incomplete-penetrance dominant inheritance patterns. None of these patients had detectable germline mutations (point mutations or small indels) in the MMR, MUTYH or BMPR1A genes, as assessed by bidirectional Sanger sequencing and Multiplex Ligation-dependent Probe Amplification (MLPA) initially, and verified by exome sequencing in this study.
Whole-exome sequencing (WES)
WES was performed on genomic DNA extracted from peripheral blood cells of all patients. DNA extraction was undertaken using the CHEMAGEN robot (Chemagen Biopolymer-Technologie AG, Baesweiler, Germany) and the sequencing was performed using the SureSelect Human All Exon kit V5 for library preparation (Agilent Technologies, Santa Clara, CA, USA) and ran on a 5500xl SOLiD™ system (Thermo Fisher Scientific, Massachusetts, USA). The sequencing reads were aligned to the hg19 reference genome using the software provided by the sequencer. Variant calling was performed using both the Lifescope and GATK 3.0 suites. For LifeScope, variant QC was performed employing the diBayes parameter of low astringency level (dibayes.het.min.start.pos = 2 and dibayes.hom.min.nonref.start.pos = 2). Additionally, we removed variants with a depth < 30X, variants with a calculated strand bias p value > 0.05, and all alternative allele variants with < 4 reads to minimise artifacts and select for high quality variants. For GATK, variant QC comprised filtering by QUAL > = 30, a minimum depth per variant of 30 and removal of variants with strand bias p < 0.05. An additional filter was applied to exclude variants with < 20% and/or < 4 variant reads, to mimic LifeScope stringency filters. Variants were annotated using ANNOVAR (version 2019Apr09).
Variant prioritization
A prioritization pipeline was applied to reduce the number of variants of interest. Firstly, only variants common to both LifeScope and GATK calling were selected, given that they were found in the same patient with the same genotype. Later, rare variants (MAFgnomAD2.1.1_NFE ≤ 0.1% and 1% were chosen for heterozygous and homozygous and calls, respectively). High-functional impact changes: loss-of-function (LoF) variants resulting in truncated proteins (nonsense, frameshift) and variants at canonical splice sites (+ -1/2) were selected, together with high-impact missense variants predicted by at least three in silico tools: PolyPhen-2 (selected if probably/possibly Damaging—D/P), CADD_phred ≥ 15, Dann ≥ 0.995), and/or located at conserved sites (GERP + + ≥ 2). Next, variants present in a representative population cohort of 267 control Spanish were eliminated to discard population ancestry bias. These belong to exome sequencing data from “non-cancer” healthy controls from the CIBERER Spanish Variant Server [ref same as above]. Sequencing artefacts were removed by curating using an in-house database of around 1,300 exomes produced with the same technologies. Variants in the eight candidate genes were validated by bidirectional Sanger sequencing.
Variant-based analysis
Frequency deviations between the detected in our cohort and the expected (as per gnomAD v2.1.1 counts in non-Finnish Europeans—NFE) were calculated for variants with at least 3 observed alleles using chi-squared tests of Fisher’s exact test if allele counts < 5.
Gene and pathway-based analyses
We selected the following cancer-related KEGG pathways to test whether there was an enrichment of potentially damaging rare variants in our cohort: hsa04310 (WNT signalling), hsa04350 (TGF-β signalling), hsa03430 (MisMatch Repair, MMR), hsa03410 (Base-Excision Repair, BER), hsa03420 (Nucleotide Excision Repair, NER), hsa03440 (Homologous Recombination, HR), map03450 (Non-Homologous End Joining, NHEJ, NHJ), hsa03460 (Fanconi Anaemia), hsa05200 (Pathways in cancer), hsa04210 (Apoptosis), hsa04514 (Cell adhesion molecules), hsa05210 (Colorectal cancer), hsa03030 (DNA replication), hsa04390 (Hippo signalling pathway), hsa04010 (MAPK signalling pathway), hsa05206 (MicroRNAs in cancer), hsa04151 (PI3K-Akt)], hsa03320 (PPAR signalling). For this, we performed a Fisher’s exact test using R40. A nominal p value of 0.05 was considered significant.
Gene prioritization and candidate-gene selecion for replication
We prioritized genes that had been described in previous works on early-onset CRC patients and/or those belonging to cancer KEGG pathways. Amongst those, we selected 8 genes for replication based on their involvement on cancer-related KEGG pathways (as described for the pathway analysis), presence of the mRNA/protein in healthy colonic mucosa (Protein Atlas, www.proteinatlas.org), genes with variants in patients under 40 years. All candidate variants in the 8 selected genes were validated using Sanger sequencing.
Candidate-gene resequencing
Replication cohort
For the replication, a total of 304 non-related unexplained MMR-proficient early-onset non-polyposis CRC patients were included. All cases were affected with CRC. The mean age at cancer diagnosis was 41.73 (range: 16–50). A detailed description of the series has been described in Belhadj et al.22. All patients were of European origin, and were assessed at the Hereditary Cancer Program of the Catalan Institute of Oncology (Spain) between 1999 and 2017. The study received the approval of the Ethics Committee of the Institut d’Investigació Biomèdica de Bellvitge (IDIBELL) (PR247/15). As for the discovery cohort, all patients received informed consent and recruitment complied with the tenets of the declaration of Helsinki.
Candidate gene sequencing
Targeted resequencing of the 8 novel candidate CRC susceptibility genes was performed on genomic DNA extracted from peripheral blood cells extracted using the FlexiGene DNA kit (Qiagen, Valencia, CA). Mutational screening was performed using a combination of PCR amplification in pooled DNAs and targeted massively parallel sequencing41. Primers used for amplification were described in the original publication. DNA pools were obtained adding equimolecular quantities of each sample (# samples/pool: 48-96). The resulting pools were used as templates for PCR amplification of each region of interest using the Phusion High-Fidelity DNA Polymerase (New England Biolabs, Ipswich, MA, USA). PCR products were checked by electrophoresis, purified (QIAquick PCR purification Kit, Qiagen, Valencia, CA, USA) and quantified (NanoDropTM, Thermo Fisher Scientific, Waltham, MA, USA). Equimolecular amounts of each purified amplicon were pooled, ligated and fragmented using a Covaris S2 (Covaris, Inc. MS, USA). DNA libraries and next generation sequencing (NGS) at high coverage were performed on a HiSeq-2000 (Illumina, San Diego, CA, USA) at Centro Nacional de Análisis Genómico (CNAG, Barcelona, Spain). FASTQ files were mapped to the reference genome GRCh37/hg19 using the Burrows-Wheeler Aligner (BWA-MEM). BAM files were generated using SAMtools. Only the bases with base quality ≥ 30 were used for the analysis. Base-level metrics of all positions were extracted using Bam-Readcount. The generated data was used to calculate the estimated number of mutated alleles per pool (ENMA), which depends on the number of samples included in each DNA pool. All genomic positions were annotated with ANNOVAR and common variants present in the Genome Aggregation Database (gnomAD v.2.1.1) with a minor allele frequency (MAF) ≥ 1% were filtered out. The median number of reads per base obtained for all coding regions and + /− 5 bp flanking regions of the 8 genes analysed was 9421(5–31,562 reads/base). The selection of the high-impact functional rare variants was performed using a prioritization pipeline identical as described above for the discovery cohort.
Variant validation
Variant-specific KASP genotyping assays (LGC Genomics, Hoddesdon, UK) and/or direct automated (Sanger) sequencing were used to validate all the variants in the candidate genes. Sequencing was performed at STAB VIDA (Caparica, Portugal) and Macrogen (Amsterdam, the Netherlands) and data was analysed with SeqMan Pro (Lasergene 13, DNASTAR, Madison, WI). The primers used for amplification and sequencing were the same as the ones used to amplify the pooled DNAs.
TCGA germline variants on candidate genes
TCGA germline variation was inspected for high-impact germline variant, as described in “Variant prioritization” section. For this, aligned bam files were obtained with permission access from the GDC data portal (https://portal.gdc.cancer.gov/). Germline variant calling was accomplished using VarScan2. This software is one of the callers used to generate somatic variation files, allows for the implementation of a germline calling pipeline as well. The results were filtered following the variant criteria described in the previous sections for colorectal COAD and READ samples. Additionally, results were restricted to samples of reported white ethnicity to account for population mismatches on allele frequencies.
TCGA second hit validation and somatic mutation profiles
VarScan2 annotated somatic vcf files were retrieved from the GDC data portal for the patients identified as carrying potential germline variants in the candidate genes, and analysed following the same variant selection criteria. Somatic mutation profiles were obtained using the MuSiCa software for mutational signatures using COSMIC v2 profiling.
TDG immunohistochemistry
Immunohistochemical assays were carried out using 4-μm-thick paraffin sections in an automatic immunostainer (Autostainer Link 48; Agilent, Santa Clara, CA, USA) equipped with a 2-step immunohistochemical staining system (EnVision FLEX/HPR; Dako, Glostrup, Denmark) that uses a peroxidase-labelled polymer conjugated to the secondary antibody. Before the immunostainer, the samples were treated for antigenic retrieval according to the manufacturer’s protocol in the pre-treatment module (PT link, Dako; Agilent). Samples were incubated with a primary antibody against TDG (thymine-DNA glycosylase) (polyclonal, pH6, dilution: 1:100, 40 min, Sigma Life Science, St Louis, MO, USA). Nonimmune serum samples were substituted for the primary antibodies as negative controls. Normal colonic mucosa was used as a positive control.
Conclusions
Sequencing of patients with early-onset CRC may be an important tool to discover novel susceptibility variants associated with an early-onset of the disease. In our study, we have discovered evidence that this early disease development may be the result of multiple and variable rare moderately-penetrant variants, i.e. CRC susceptibility is oligogenic and heterogeneous. Hence, further larger and appropriately powered studies are necessary in order to unveil the genetics behind it. Alternative complementary approaches such as using interrelated omic sources and high throughput functional studies may in the near future offer refinement on strategies based only on gene prioritization algorithms.
References
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer–analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 343, 78–85. https://doi.org/10.1056/NEJM200007133430201 (2000).
Graff, R. E. et al. Familial risk and heritability of colorectal cancer in the Nordic twin study of cancer. Clin. Gastroenterol. Hepatol. 15, 1256–1264. https://doi.org/10.1016/j.cgh.2016.12.041 (2017).
Valle, L., Vilar, E., Tavtigian, S. V. & Stoffel, E. M. Genetic predisposition to colorectal cancer: syndromes, genes, classification of genetic variants and implications for precision medicine. J. Pathol. 247, 574–588. https://doi.org/10.1002/path.5229 (2019).
Hahn, M. M. et al. The genetic heterogeneity of colorectal cancer predisposition—guidelines for gene discovery. Cell. Oncol. (Dordr.) 39, 491–510. https://doi.org/10.1007/s13402-016-0284-6 (2016).
Terradas, M., Capella, G. & Valle, L. Dominantly inherited hereditary nonpolyposis colorectal cancer not caused by MMR genes. J. Clin. Med. 2020, 9. https://doi.org/10.3390/jcm9061954 (2020).
Nieminen, T. T. et al. Germline mutation of RPS20, encoding a ribosomal protein, causes predisposition to hereditary nonpolyposis colorectal carcinoma without DNA mismatch repair deficiency. Gastroenterology 147, 595–598. https://doi.org/10.1053/j.gastro.2014.06.009 (2014).
de la Chapelle, A. Genetic predisposition to colorectal cancer. Nat. Rev. Cancer 4, 769–780. https://doi.org/10.1038/nrc1453 (2004).
Archambault, A. N. et al. Cumulative burden of colorectal cancer-associated genetic variants is more strongly associated with early-onset vs late-onset cancer. Gastroenterology 158, 1274–1286. https://doi.org/10.1053/j.gastro.2019.12.012 (2020).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. https://doi.org/10.1101/gr.107524.110 (2010).
Quang, D., Chen, Y. & Xie, X. DANN: a deep learning approach for annotating the pathogenicity of genetic variants. Bioinformatics 31, 761–763. https://doi.org/10.1093/bioinformatics/btu703 (2015).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249. https://doi.org/10.1038/nmeth0410-248 (2010).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894. https://doi.org/10.1093/nar/gky1016 (2019).
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913. https://doi.org/10.1101/gr.3577405 (2005).
Dopazo, J. et al. 267 Spanish exomes reveal population-specific differences in disease-related genetic variation. Mol. Biol. Evol. 33, 1205–1218. https://doi.org/10.1093/molbev/msw005 (2016).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377-386. https://doi.org/10.1093/nar/gks1118 (2013).
DeBerardinis, R. J. & Chandel, N. S. We need to talk about the Warburg effect. Nat. Metab. 2, 127–129. https://doi.org/10.1038/s42255-020-0172-2 (2020).
Esteban-Jurado, C. et al. Whole-exome sequencing identifies rare pathogenic variants in new predisposition genes for familial colorectal cancer. Genet. Med. 17, 131–142. https://doi.org/10.1038/gim.2014.89 (2015).
Chubb, D. et al. Rare disruptive mutations and their contribution to the heritable risk of colorectal cancer. Nat. Commun. 7, 11883. https://doi.org/10.1038/ncomms11883 (2016).
Arora, S. et al. Genetic variants that predispose to DNA double-strand breaks in lymphocytes from a subset of patients with familial colorectal carcinomas. Gastroenterology 149, 1872–2188. https://doi.org/10.1053/j.gastro.2015.08.052 (2015).
de Voer, R. M. et al. Identification of novel candidate genes for early-onset colorectal cancer susceptibility. PLoS Genet.. 12, e1005880. https://doi.org/10.1371/journal.pgen.1005880 (2016).
Belhadj, S. et al. Candidate genes for hereditary colorectal cancer: mutational screening and systematic review. Hum. Mutat. 41, 1563–1576. https://doi.org/10.1002/humu.24057 (2020).
Cancer Genome Atlas, N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337, https://doi.org/10.1038/nature11252 (2012).
Diaz-Gay, M. et al. Mutational Signatures in Cancer (MuSiCa): a web application to implement mutational signatures analysis in cancer samples. BMC Bioinform. 19, 224. https://doi.org/10.1186/s12859-018-2234-y (2018).
Sjolund, A. B., Senejani, A. G. & Sweasy, J. B. MBD4 and TDG: multifaceted DNA glycosylases with ever expanding biological roles. Mutat. Res. 743–744, 12–25. https://doi.org/10.1016/j.mrfmmm.2012.11.001 (2013).
Xu, X. et al. Thymine DNA glycosylase is a positive regulator of Wnt signaling in colorectal cancer. J Biol Chem 289, 8881–8890. https://doi.org/10.1074/jbc.M113.538835 (2014).
Cortazar, D. et al. Embryonic lethal phenotype reveals a function of TDG in maintaining epigenetic stability. Nature 470, 419–423. https://doi.org/10.1038/nature09672 (2011).
Vasovcak, P. et al. Unique mutational profile associated with a loss of TDG expression in the rectal cancer of a patient with a constitutional PMS2 deficiency. DNA Repair (Amst.) 11, 616–623. https://doi.org/10.1016/j.dnarep.2012.04.004 (2012).
Broderick, P. et al. Evaluation of NTHL1, NEIL1, NEIL2, MPG, TDG, UNG and SMUG1 genes in familial colorectal cancer predisposition. BMC Cancer 6, 243. https://doi.org/10.1186/1471-2407-6-243 (2006).
Ciavarella, M. et al. Somatic APC mosaicism and oligogenic inheritance in genetically unsolved colorectal adenomatous polyposis patients. Eur. J. Hum. Genet. 26, 387–395. https://doi.org/10.1038/s41431-017-0086-y (2018).
Morak, M., Massdorf, T., Sykora, H., Kerscher, M. & Holinski-Feder, E. First evidence for digenic inheritance in hereditary colorectal cancer by mutations in the base excision repair genes. Eur. J. Cancer 47, 1046–1055. https://doi.org/10.1016/j.ejca.2010.11.016 (2011).
Neri, F. et al. TET1 is a tumour suppressor that inhibits colon cancer growth by derepressing inhibitors of the WNT pathway. Oncogene 34, 4168–4176. https://doi.org/10.1038/onc.2014.356 (2015).
Guo, H., Zhu, H., Zhang, J., Wan, B. & Shen, Z. TET1 suppresses colon cancer proliferation by impairing beta-catenin signal pathway. J. Cell Biochem. 120, 12559–12565. https://doi.org/10.1002/jcb.28522 (2019).
Stock, C. How dysregulated ion channels and transporters take a hand in esophageal, liver, and colorectal cancer. Rev. Physiol. Biochem. Pharmacol. https://doi.org/10.1007/112_2020_41 (2020).
Abuli, A. et al. Case-control study for colorectal cancer genetic susceptibility in EPICOLON: previously identified variants and mucins. BMC Cancer 11, 339. https://doi.org/10.1186/1471-2407-11-339 (2011).
Guda, K. et al. Inactivating germ-line and somatic mutations in polypeptide N-acetylgalactosaminyltransferase 12 in human colon cancers. Proc Natl Acad Sci U S A 106, 12921–12925. https://doi.org/10.1073/pnas.0901454106 (2009).
MacArthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature 508, 469–476. https://doi.org/10.1038/nature13127 (2014).
Deelen, P. et al. Improving the diagnostic yield of exome- sequencing by predicting gene-phenotype associations using large-scale gene expression analysis. Nat. Commun. 10, 2837. https://doi.org/10.1038/s41467-019-10649-4 (2019).
Fernandez-Rozadilla, C. et al. Single nucleotide polymorphisms in the Wnt and BMP pathways and colorectal cancer risk in a Spanish cohort. PLoS ONE https://doi.org/10.1371/journal.pone.0012673 (2010).
R Core Team. R: A Language and Environment for Statistical Computing. https://www.R-project.org (2017).
Puente, X. S. et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475(7354):101–105. https://doi.org/10.1038/nature10113 (2011).
Acknowledgements
We are thankful to all patients participating in this study and all members of the EPICOLON Consortium. The results published here are in whole or partly based upon data generated by The Cancer Genome Atlas (TCGA) under dbGAP version phs000178.v11.p8. TCGA is managed by the NCI and NHGRI. Information about TCGA can be found at http://cancergenome.nih.gov.
Funding
This research was supported by Grants from Instituto de Salud Carlos III /FEDER: PI11/00681 (CRP), PI16/01057 (AC), PI17/00509 (CRP), PI17/00878 (SCB), PI19/00179 (CFR), PI19/01316 (JMC-T), PI20/01113 (SCB), PI20/00226 (CRP); CIBERONC-CB16/12/00234 (LV); and Fundación Olga Torres (LV and CFR); CERCA Program and Agència de Gestió d’Ajuts Universitaris i de Recerca (Generalitat de Catalunya, GRPRE 2017SGR21, GRC 2017SGR653). AL-N is supported by a Predoctoral Fellowship (GAIN, Xunta de Galicia, 2018 call) and IQ is supported by the FPI program of the Spanish Ministry of Science and Innovation—FEDER (SAF2016-80888-R). This article is based upon work from COST Action TransColonCan CA17118, supported by European Cooperation in Science and Technology (COST—www.cost.eu; www.transcoloncan.eu).
Author information
Authors and Affiliations
Contributions
Conceptualization, C.R.P.; Data curation, C.F.R., M.A.B. and A.L.N.; Formal analysis, C.F.R., I.Q., J.M.C.T. and L.V.; Funding acquisition, C.R.P.; Methodology, C.F.R., L.V. and C.R.P.; Project administration, C.R.P.; Resources, EPICOLON Consortium and C.R.P.; Supervision, C.R.P.; Validation, C.F.R. and L.V.; Writing—original draft, C.F.R., L.V. and C.R.P.; Writing—review and editing, J.A., J.M.C.T., E.R., D.G., X.L., L.B., X.B., R.J., F.B., A.C., S.C.-B., Á.C. and L.V.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernández-Rozadilla, C., Álvarez-Barona, M., Quintana, I. et al. Exome sequencing of early-onset patients supports genetic heterogeneity in colorectal cancer. Sci Rep 11, 11135 (2021). https://doi.org/10.1038/s41598-021-90590-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90590-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.