Abstract
HCV screening depends mainly on a one-assay anti-HCV testing strategy that is subject to an increased false-positive rate in low-prevalence populations. In this study, a two-assay anti-HCV testing strategy was applied to screen HCV infection in two groups, labelled group one (76,442 people) and group two (18,415 people), using Elecsys electrochemiluminescence (ECL) and an Architect chemiluminescent microparticle immunoassay (CMIA), respectively. Each anti-HCV-reactive serum was retested with the other assay. A recombinant immunoblot assay (RIBA) and HCV RNA testing were performed to confirm anti-HCV positivity or active HCV infection. In group one, 516 specimens were reactive in the ECL screening, of which CMIA retesting showed that 363 (70.3%) were anti-HCV reactive (327 positive, 30 indeterminate, 6 negative by RIBA; 191 HCV RNA positive), but 153 (29.7%) were not anti-HCV reactive (4 positive, 29 indeterminate, 120 negative by RIBA; none HCV RNA positive). The two-assay strategy significantly improved the positive predictive value (PPV, 64.1% & 90.1%, P < 0.05). In group two, 87 serum specimens were reactive according to CMIA screening. ECL showed that 56 (70.3%) were anti-HCV reactive (47 positive, 8 indeterminate, 1 negative by RIBA; 29 HCV RNA positive) and 31 (29.7%) were anti-HCV non-reactive (25 negative, 5 indeterminate, 1 positive by RIBA; none HCV RNA positive). Again, the PPV was significantly increased (55.2% & 83.9%, P < 0.05). Compared with a one-assay testing strategy, the two-assay testing strategy may significantly reduce false positives in anti-HCV testing and identify inactive HCV infection in low-seroprevalence populations.
Similar content being viewed by others
Introduction
Hepatitis C virus (HCV) infection is one of the leading causes of liver cirrhosis, hepatocellular carcinoma, and mortality worldwide1,2,3. Approximately 71 million people around the world have chronic HCV infection, with an annual increase of 1.75 million new infections4. However, only 20% and < 1% of patients were diagnosed and treated in high-income settings and in low- and middle-income countries (LMICs), respectively5. Currently, the prevalence of hepatitis C is estimated at 2.8% globally, varying widely in different geographical regions and populations6. Among the nations of the world, Egypt is estimated to have the highest HCV prevalence, at 11.9%7, whereas Iran has the lowest prevalence, at 0.30%8. Most previous studies have focused on screening strategies for populations at high risk for HCV, such as people who inject drugs, men who have sex with men9,10,11, but few studies have paid attention to screening strategies for the general populations.
The World Health Organization (WHO) has developed evidence-based guidelines that focus on who to test and how to test for chronic hepatitis C infection12. The current diagnostic algorithm prioritizes the detection of HCV antibodies as evidence of past or current HCV infection through rapid diagnostic tests (RDTs) or laboratory-based serum samples such as enzyme immunoassays (EIAs), electrochemiluminescence immunoassays (ECLs), and chemiluminescence immunoassays (CLIAs). Anti-HCV-reactive serum requires further supplemental testing for HCV RNA or HCV core antigen to confirm whether there is an HCV infection with viraemia.
Several previous studies have attempted to establish the optimal S/CO ratio in various laboratory-based serological assays to minimize the need for supplemental tests and avoid the possibility of reporting false-positive results as much as possible13,14. However, the conclusions were not always consistent14,15. Only large-scale screening of the general population could substantially accelerate the rate of HCV elimination. A recent study demonstrated that the correlation coefficients of anti-HCV low S/CO ratios were poor between the Architect, Elecsys, and Vitros assays, leading to the inference that combining two serological assays may be useful to eliminate false-positive results16. Therefore, our study aims to assess the diagnostic performance of two-assay serological testing strategies in low-prevalence populations.
Results
One-assay serological testing strategy using ECL screening
Among the 76,442 people in group one, 518 showed anti-HCV reactivity in the ECL assay, of which 2 cases failed to show reactivity in a subsequent repeated test with the same assay (the S/CO values were 1.01 and 1.05, respectively, in the first Elecsys Screening testing, and the RIBA and HCV RNA results were also negative). Therefore, a total of 516 cases were included in this study. Among these sera, RIBA showed that 331 were anti-HCV positive, 59 were indeterminate, and 126 were negative. NAT confirmed that 191 (37.0%) cases were HCV RNA positive (Fig. 1). According to the manufacturer's instructions, samples with S/CO ≥ 1.0 were considered positive for anti-HCV, with a PPV of 64.1% (331/516).
Two-assay serological testing strategy using ECL screening and CMIA retesting
The 516 specimens that were anti-HCV reactive on ECL were retested using the second serological assay (CMIA), which showed that 363 (70.3%) were anti-HCV reactive, while the other 153 (29.7%) specimens were non-reactive. Among 363 specimens that were anti-HCV reactive on both serological tests, RIBA showed that 327 were anti-HCV positive, 30 were indeterminate, and 6 were negative, for a PPV of 90.1% (327/363), significantly higher than the PPV of the one-assay serological test (Fig. 1, χ2 = 94.81, P = 0.001). Moreover, 191 cases were HCV RNA positive in the above 363 specimens, which was significantly different from the results of the one-assay serological test (Fig. 1, χ2 = 21.11, P = 0.001).
Among the 153 sera with inconsistent results, 4 were anti-HCV positive, 29 were indeterminate, and 120 were negative by RIBA (Fig. 1), showing a significant difference compared with the one-assay serological strategy (Fig. 1, χ2 = 187.93, P < 0.001). Moreover, these 153 specimens were all HCV RNA negative.
The optimal algorithm for anti-HCV screening with ECL and CMIA
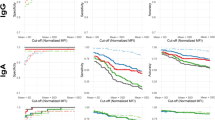
Figure 2a shows that the correlation coefficient of the Elecsys and Architect S/CO ratios was low (R2 = 0.12). The optimal S/CO ratio of CMIA for anti-HCV positivity was 5.6 according to ROC curve analysis (Fig. 2b). The anti-HCV-reactive specimens were further categorized into three groups according to the results of the two serological assays: ECL ≥ 1.0 and CMIA < 1.0, ECL ≥ 1.0 and 1.0 ≤ CMIA < 5.6, ECL ≥ 1.0 and CMIA ≥ 5.6 (Table 1). Their PPVs were 2.6%, 44.4% and 98.1%, respectively, and the difference was statistically significant (χ2 = 415.51, P = 0.001). Their HCV RNA positive rates were 0.0%, 5.6%, and 60.8%, respectively, and the difference was significant (χ2 = 188.08, P = 0.001).
The correlation coefficients of the Elecsys and Architect S/CO ratios was poor (R2 = 0.12). (b) Receiver characteristic curve of the RIBA test at different cutoff levels using CMIA for the anti-HCV test. The diagnostic sensitivity and specificity (95% CIs) were 91.5% (88.1–94.1%) and 96.8.7% (93.2–98.5%), respectively, for an S/CO ratio of 5.6. The area under the curve (95% CI) was 0.971 (0.953–0.990).
Two-assay serological testing strategy using CMIA screening and ECL retesting
It was observed that 87 serum specimens in group two (18,415 people) had S/CO ≥ 1.0 using CMIA screening. Of these, RIBA results indicated that 48 were anti-HCV positive (PPV, 55.2%), 13 were indeterminate, and 26 were negative. NAT showed that 29 cases were HCV RNA positive. However, the second serological test (ECL) showed that 56 specimens had S/CO ≥ 1.0 (47 positive, 8 indeterminate, 1 negative by RIBA; 29 HCV RNA positive), and 31 specimens had S/CO < 1.0 (1 positive, 5 indeterminate, 25 negative by RIBA; none HCV RNA positive). The two-assay serological testing strategy improved the PPV from 55.2% to 83.9% (Fig. 3, χ2 = 18.50, P < 0.01).
Discussion
Among 76,442 people in group one who were screened with a one-assay testing strategy, 516 (0.68%) were anti-HCV reactive, of which 331 were confirmed to be anti-HCV positive by RIBA, for an HCV prevalence of 0.43% (331/76,442). The prevalence in group two was 0.26% (48/18, 415). Both were lower than the initial screening results. Several similar previous studies in populations with seroprevalence rates of 1.0%, 4.6%, and 20.8% revealed that the corresponding PPVs of the Elecsys anti-HCV II assay were 88.04%, 98.9%, and 99.5%, respectively17,18,19. It is well known that prevalence affects PPV. Some previous studies have shown that the sensitivity of the Elecsys anti-HCV II assay to anti-HCV is relatively high (ranging from 99.3% to 100%)17,20. This means that almost all HCV-infected people will be found by serological screening. Nevertheless, even with high specificity (99%), the application of a one-assay test for anti-HCV may lead to a considerable number of false-positive results as well as low PPV, particularly in low-seroprevalence settings or populations21.
The causes of false reactivity are diverse, including the interference of many autoantibodies, nonspecific immune responses, antibody cross-reactions to other pathogens, and procedural errors22,23. Most manufacturers recommend retesting anti-HCV-reactive sera in duplicate. However, our data suggested that all reactive cases in the first screening test remained anti-HCV reactive in subsequent duplicate testing, except for two borderline results. The consistency rate of the two repeat tests of the Elecsys anti-HCV II assay was 99.6% (516/518). This result suggested that repeated testing using the same assay has no significant ability to discriminate false-positive results.
However, the consistency rate of Roche ECL and Abbott CMIA was only 70.3% (363/516, Fig. 1). This was consistent with Ha J’s report16. Surprisingly, using two-assay serological testing, 23.3% (120/516) RIBA-negative cases and 47.1% (153/325) HCV RNA-negative cases were further excluded (Fig. 1). Therefore, our data showed that two-assay serological testing, as an alternative to re-examination in duplicate with the same method recommended by the manufacturer, can significantly reduce false-positive results. Furthermore, compared with a one-assay approach, the two-assay strategy eliminates one test and reduces the cost by 50%, in addition to possibly reducing the number of HCV RNA tests that must be conducted.
Several recent studies have suggested that a high anti-HCV S/CO ratio is accurate at predicting the presence of viraemic HCV infection24,25. However, in this study, a high S/CO ratio merely provides insight into the real anti-HCV status and is not able to accurately predict the presence of viraemia, consistent with the report by Ha J et al.16. Notably, this would lead to some HCV-infected people being missed if a high anti-HCV S/CO ratio were set as a reference value. However, only 4 specimens were anti-HCV positive by RIBA, and no HCV RNA was positive in the 153 sera that produced discordant results on the two-assay serological tests (ECL S/CO ≥ 1.0 and CMIA < 1.0, Fig. 1).
Moreover, we analysed 87 anti-HCV-reactive sera in CMIA screening at Shanghai First Maternity and Infant Health Hospital. We observed a PPV of 55.2% by RIBA. However, the PPV rose to 83.9% with two-assay serological tests (Fig. 2). Again, this suggests that the two-assay testing strategy could improve the PPV of anti-HCV screening, which would increase the accuracy of epidemiological investigations of HCV infection.
Although the positive predictive value of RIBA is close to 100%, avoiding a high false-positive rate, it is more complicated and expensive than other assays and prone to a higher rate of indeterminate results26. NAT technology is also costly, requiring skilled staff and specialized laboratory equipment. Unfortunately, only 60% of anti-HCV-reactive individuals return for HCV RNA testing27. A survey of current HCV testing practices in 23 LMICs showed that the majority of these programs provide anti-HCV assays. However, HCV RNA testing is available in only 5% to 30% of countries. HCV core antigen testing is currently not reported in any of these LMICs28. The main reasons are lack of public awareness, a low level of knowledge among health professionals, limitations of diagnostic sites, deficiency of funding for HCV testing, and lack of follow-up after the diagnosis of viraemic HCV infection28. HCV RNA testing was mainly performed in highly resourced settings, but the majority of HCV-infected individuals were in LMICs. Therefore, a two-assay serological testing strategy may be an excellent alternative strategy to reduce the false-positive rate of anti-HCV testing and rule out inactive HCV infection among anti-HCV-reactive individuals in low-seroprevalence populations. These goals are also increasingly important for accurately assessing the global epidemiology and burden of HCV infection.
In 2016, the WHO set ambitious goals to eliminate hepatitis as a major public health threat. Most countries committed to achieving the bold targets of 90% diagnosis and 80% of diagnosed patients eligible for treatment by 203029. Recently, a new era of highly effective, well-tolerated oral direct-acting antiviral therapy for chronic HCV infection has emerged30,31. However, the overwhelming majority of people infected with HCV remain unaware of their infection, and the COVID-19 pandemic has further impacted the elimination of HCV32, which has further increased the potentially massive burden of undiagnosed infection worldwide. Therefore, there is an urgent need to establish an effective algorithm for broad-scale diagnosis of potentially HCV-infected individuals while simultaneously maintaining affordable testing costs for individuals and appropriate performance standards for laboratories, which will be critical for curbing the epidemic of HCV worldwide. In order to achieve the 2030 elimination target, the concept of micro-elimination has been proposed; in this strategy, the wider population would be divided into smaller subgroups through targeted treatment and prevention33. Similarly, different screening strategies should be targeted and implemented for certain prevalence populations. With the implementation of the hepatitis C elimination plan in 2030, the overall prevalence of HCV infection in the general population will gradually decline32. At that time, the two-assay serological test strategy may play an important role in the substantial reduction of false-positive anti-HCV results among low-prevalence populations.
There were several limitations to this research. Since serological tests cannot detect anti-HCV if it is absent or very low in the early stage of HCV infection, there is inevitably a window in which occasional false-negative results occur. Especially in groups with a high HCV prevalence, such as people who inject drugs and men who have sex with men, the two-assay strategy might cause a high number of false negatives. Under these circumstances, NAT is recommended to detect HCV RNA. In addition, the two-assay strategy is not suitable for immunocompromised individuals because they may not be able to produce enough antibodies even if exposed to a high serum HCV load.
In conclusion, one-assay anti-HCV serological tests play an important role in screening for HCV infection, but they also carry an increased risk of false-positive results in low-seroprevalence populations. Supplementary HCV RNA experiments could effectively detect viraemic HCV infection in anti-HCV-reactive individuals, but this method may not be feasible to implement effectively in low-income settings. Our data suggest that two-assay serological testing rather than repeated assays with the same kit could significantly reduce false-positive results and identify inactive HCV infection in HCV screening of low-seroprevalence populations.
Methods
Subjects
Two groups of people were included this study. The first group consisted of 76,442 people who planned to receive blood transfusions, surgery34 or pregnancy tests35 at Minhang Hospital, Fudan University, from September 2016 to December 2018. Among them, 28,943 (37.9%) were males with a mean age of 52.4 ± 17.0 years, and 47,499 (62.1%) were females with a mean age of 42.4 ± 14.5 years. Serum anti-HCV screening of the first group was completed using electrochemiluminescence (ECL) immunoassay systems within four hours after collection. The second group included 2,489 males (37.2 ± 7.5 years old) and 15,926 females (33.2 ± 7.4 years old) who visited Shanghai First Maternity and Infant Health Hospital for blood transfusions, surgery or pregnancy tests from May 2018 to December 2018. Serum anti-HCV screening for the second group was completed using a chemiluminescent microparticle immunoassay (CMIA) within 4 h of collection. Each anti-HCV-reactive serum sample was divided into 3 aliquots (each 500 μl). The first was stored at 4 ℃ and retested within 24 h with the other method. The other two were stored at − 80 ℃ for NAT and RIBA, which were performed as supplemental tests to confirm anti-HCV positivity or active HCV infection within 7 days and 30 days, respectively. All experimental procedures were conducted according to The Declaration of Helsinki; written informed consent was obtained from participants, and the study was approved by the institutional ethics committee of Minhang Hospital, Fudan University.
ECL immunoassay for anti-HCV
An ECL immunoassay was applied for anti-HCV testing using the Elecsys anti-HCV II assay on the Cobas 601 analyser (Roche Diagnostics, Mannheim, Germany). The kit was a third-generation test using peptides and recombinant antigens representing the core, NS3, and NS4 to capture the corresponding antibodies. The results were expressed as signal-to-cutoff (S/CO) ratios: S/CO < 1.0 indicated anti-HCV nonreactivity, and S/CO ≥ 1.0 indicated anti-HCV reactivity. All S/CO ≥ 1.0 sera were retested in duplicate according to the manufacturer's instructions. If either of the two results remained S/CO ≥ 1.0, then the subject was considered anti-HCV reactive. The anti-HCV reactive sera were retested using an Architect anti-HCV reagent kit.
CMIA for anti-HCV
A CMIA was performed to test the serum for anti-HCV using an Architect anti-HCV reagent kit on an Architect i2000 analyser (Abbott Diagnostics, IL, USA). The CMIA kit detected HCV antibodies against the fusion protein HCr43 (HCV core antigen and NS3-c33) and the recombinant protein c100-3 (NS4). If S/CO was ≥ 1.0, the serum was retested in duplicate according to the manuals. If either was S/CO ≥ 1.0, the subject was considered anti-HCV reactive and retested using the Elecsys anti-HCV II assay.
Recombinant immunoblot assay for anti-HCV
Specimens with reactive anti-HCV results were further tested with RIBAs to confirm anti-HCV positivity using a recombinant immunoblot kit for antibodies against the hepatitis C virus (Beijing Wantai Biopharm, Beijing, China). The nitrocellulose strips contained seven bands for the core, NS3, NS4-1, NS4-2, and NS5 antigens as well as control A and control B. The result was defined as negative, ± , 1 + , or 2 + by comparing the colour of the antigen band with control A. Anti-HCV positivity was defined as the presence of at least two antigens with greater than or equal to 1 + (≥ 1 +) reactivity. An indeterminate result was defined by only one band scoring ≥ 1 + . Anti-HCV negativity was defined by the absence of antigens scoring ≥ 1 + . The tests were performed according to the manufacturer’s instructions by technicians with more than 10 years of experience.
Active HCV infection
Nucleic acid extraction from serum samples was performed with a QIAamp Viral RNA Mini Kit (Qiagen Inc.) according to the manufacturer’s instructions. HCV RNA amplification and quantitative determination were performed using the HCV RNA Quantitation Kit (Shanghai Kehua Bio-Engineering, Shanghai, China), which has a lower limit of detection of 500 IU/mL and is linear from 103 to 107 IU/ml based on TaqMan real-time PCR on the ABI 7500 real-time PCR system (Applied Biosystems, CA, USA). The following PCR program was run: one cycle of UNG enzyme reaction at 50 °C for 2 min and pre-denaturation at 94 °C for 2 min, followed by 40 cycles of denaturation at 94 °C for 10 s and annealing/extension at 60 °C for 30 s.
Statistical analysis
Statistical analysis was performed with Stata 13.0 software (StataCorp LP, College Station, Texas, USA). The frequency data were analysed using the chi-square (χ2) test. Receiver operating characteristic (ROC) curve analysis was performed to evaluate the predictive accuracy of anti-HCV S/CO compared with the RIBA results. Positive predictive value (PPV) is presented as percentages. A P-value < 0.05 was considered significantly different.
Abbreviations
- HCV:
-
Hepatitis C virus
- anti-HCV:
-
Antibody against HCV
- ECL:
-
Electrochemiluminescence
- CMIA:
-
Chemiluminescent particle immunoassay
- RIBA:
-
Recombinant immunoblot assay
- PPV:
-
Positive predictive value
- S/CO:
-
Signal-to-cutoff
- LMICs:
-
Low- and middle-income countries
- EIA:
-
Enzyme immunoassay
- NAT:
-
Nucleic acid amplification testing
- CI:
-
Confidence interval
References
Beste, L. A. et al. Trends in Burden of cirrhosis and hepatocellular carcinoma by underlying liver disease in US veterans, 2001–2013. Gastroenterology 149, 1471–1482 (2015).
Kulik, L. & El-Serag, H. B. Epidemiology and management of hepatocellular carcinoma. Gastroenterology 156, 477–491 (2019).
Global, Regional, and National Age-Sex Specific All-Cause and Cause-Specific Mortality for 240 Causes of Death, 1990–2013:a Systematic Analysis for the Global Burden of Disease Study 2013. The Lancet. 385, 117–171 (2015).
World Health Organization. https://www.who.int/news/item/17-11-2020-who-commissioned-global-systematic-review-finds-high-hcv-prevalence-and-incidence-among-men-who-have-sex-with-men. Accessed 17 November 2020.
World Health Organization. Guidelines on Hepatitis B and C Testing. https://apps.who.int/iris/bitstream/handle/10665/254621/9789241549981-eng.pdf?sequence=1. Accessed February 2017.https://apps.who.int/iris/bitstream/handle/10665/254621/9789241549981-eng.pdf?sequence=1. Accessed February 2017.
Roudot-Thoraval, F. Epidemiology of hepatitis C virus infection. Clin. Res. Hepatol. Gas. 45, 101596 (2021).
Kouyoumjian, S. P., Chemaitelly, H. & Abu-Raddad, L. J. Characterizing hepatitis C virus epidemiology in Egypt: systematic reviews, meta-analyses, and meta-regressions. Sci. Rep UK 8, 150 (2018).
Mahmud, S., Akbarzadeh, V. & Abu-Raddad, L. J. The epidemiology of hepatitis C virus in Iran: systematic review and meta-analyses. Sci Rep 8, 73 (2018).
Martin, N. K., Skaathun, B., Vickerman, P. & Stuart, D. Modeling combination HCV prevention among HIV-infected men who have sex with men and people who inject drugs. AIDS Rev. 19(2), 97–104 (2017).
Rossetti, B. et al. Evolution of the prevalence of hepatitis C virus infection and hepatitis C virus genotype distribution in human immunodeficiency virus-infected patients in Italy between 1997 and 2015. Clin. Microbiol. Infec. 24, 422–427 (2018).
He, T. et al. Prevention of hepatitis C by screening and treatment in U.S. Prisons. Ann. Intern. Med. 164, 84 (2016).
Easterbrook, P. J. Who to test and how to test for chronic hepatitis C infection-2016 WHO testing guidance for low- and middle-income countries. J. Hepatol. 65, S46–S66 (2016).
Lai, K. K. et al. Improved reflexive testing algorithm for hepatitis C infection using signal-to-Cutoff ratios of a hepatitis C virus antibody assay. Clin. Chem. 57, 1050–1056 (2011).
Oethinger, M., Mayo, D. R., Falcone, J., Barua, P. K. & Griffith, B. P. Efficiency of the ortho VITROS assay for detection of hepatitis C virus-specific antibodies increased by elimination of supplemental testing of samples with very low sample-to-cutoff ratios. J. Clin. Microbiol. 43, 2477–2480 (2005).
Choi, M. S. et al. The role of the signal-to-cutoff ratio in automated anti-HCV chemiluminescent immunoassays by referring to the nucleic acid amplification test and the recombinant immunoblot assay. Ann. Lab. Med. 38, 466–472 (2018).
Ha, J., Park, Y. & Kim, H. Signal-to-cutoff ratios of current anti-HCV assays and a suggestion of new algorithm of supplementary testing. Clin. Chim. Acta. 498, 11–15 (2019).
Kim, B., Ahn, H. J., Choi, M. H. & Park, Y. Retrospective analysis on the diagnostic performances and signal to-cut-off ratios of the Elecsys anti-HCV II assay. J. Clin. Lab. Anal. 32, e22165 (2018).
Yue, Z., Xia, C. & Wang, H. Performance evaluation of the mindray anti-HCV assay for the detection of hepatitis C virus infection. J. Clin. Lab. Anal. 32, e22600 (2018).
Park, Y., Seok, Y., Choi, J. & Kim, H. Performance evaluation of the vitros anti-hepatitis C virus antibody assay for use in clinical laboratories. Clin. Biochem. 45, 175–177 (2012).
Yoo, S. J. et al. Evaluation of the Elecsys® anti-HCV II assay for routine hepatitis C virus screening of different Asian Pacific populations and detection of early infection. J. Clin. Virol. 64, 20–27 (2015).
Parry, J. V., Easterbrook, P. & Sands, A. R. One or two serological assay testing strategy for diagnosis of HBV and HCV infection? The use of predictive modelling. BMC Infect. Dis. 17, 705–770 (2017).
Heinrichs, A., Antoine, M., Steensels, D., Montesinos, I. & Delforge, M. HCV false positive immunoassays in patients with LVAD: a potential trap. J. Clin. Virol. 78, 44–46 (2016).
Sims, D. B., Kataria, R., Rangasamy, S. & Jorde, U. P. Seroreversion of positive anti-hepatitis C virus antibodies in left ventricular assist device recipients: now you see them now you don’t. Artif. Organs. 43, 791–795 (2019).
Seo, Y. S. et al. Significance of anti-HCV signal-to-cutoff ratio in predicting hepatitis C viremia. Korean J. Intern. Med. 24, 302 (2009).
Kao, H., Chen, K., Lin, C., Chang, J. & Lee, C. Utilization of signal-to-cutoff ratio of hepatitis C virus antibody assay in predicting HCV viremia among hemodialysis patients. Nephron 130, 127–133 (2015).
Dufour, D. R. et al. Testing for HCV infection: an update of guidance for clinicians and laboratorians. MMWR Morb. Mortal. Wkly Rep. 62, 362–365 (2013).
Tolan, N. V. et al. New therapies for treating hepatitis C virus: impact on laboratory testing recommendations and clinical management. Clin. Chem. 63, 1799–1805 (2017).
Reipold, E. I. et al. Values, preferences and current hepatitis B and C testing practices in low- and middle-income countries: results of a survey of end users and implementers. BMC Infect. Dis. 17, 154 (2017).
Razavi, H. et al. Hepatitis C virus prevalence and level of intervention required to achieve the WHO targets for elimination in the European Union by 2030: a modelling study. Lancet Gastroenterol. Hepatol. 2, 325–336 (2017).
Ioannou, G. N. & Feld, J. J. What are the benefits of a sustained virologic response to direct-acting antiviral therapy for hepatitis C virus infection?. Gastroenterology 156, 446–460 (2019).
Asselah, T. et al. Efficacy and safety of glecaprevir/pibrentasvir in patients with chronic hepatitis C virus genotype 5 or 6 infection (ENDURANCE-5,6): an open-label, multicentre, phase 3B trial. Lancet Gastroenterol. Hepatol. 4, 45–51 (2019).
Blach, S. et al. Global prevalence and genotype distribution of hepatitis C virus infection in 2015: a modelling study. Lancet Gastroenterol. Hepatol. 2, 161–176 (2017).
Lazarus, J. V., Wiktor, S., Colombo, M., Thursz, M. & Easl, I. L. F. Micro-elimination-a path to global elimination of hepatitis C. J. Hepatol. 67, 665–666 (2017).
Yurong, L. et al. Prevalence and characteristics of hepatitis C virus infection in Shenyang City, Northeast China, and prediction of HCV RNA positivity according to serum anti-HCV level: retrospective review of hospital data. BMC Infect. Dis 18, 145 (2018).
Schillie, S., Wester, C., Osborne, M., Wesolowski, L. & Ryerson, A. B. CDC recommendations for hepatitis C screening among adults-United States, 2020 morbidity and mortality weekly report (MMWR). Recomm. Rep. 69(2), 1–17 (2020).
Funding
This work was supported by the Ministry of Science and Technology of the People’s Republic of China (Grant numbers 2015AA021107-035).
Author information
Authors and Affiliations
Contributions
Y.H. study concept and design; acquisition data; analysis and interpretation of data; statistical analysis; drafting of the manuscript. H.P. acquisition data; analysis and interpretation of data; study supervision. Q.G. acquisition data; analysis and interpretation of data. P.L. and X.X. acquisition data. Z.Z. study concept and design; analysis and interpretation of data; statistical analysis; drafting of the manuscript; obtained funding; administrative and material support; study supervision; drafting of the manuscript. All authors participated in revising the manuscript for important intellectual content. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, Y., Pan, H., Gao, Q. et al. The role of a two-assay serological testing strategy for anti-HCV screening in low-prevalence populations. Sci Rep 11, 8689 (2021). https://doi.org/10.1038/s41598-021-88138-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-88138-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.