Abstract
Microsatellites (simple sequence repeats, SSRs) are co-dominant nuclear markers that are widely used in population genetic studies. Population genetic parameters from different studies might be significantly influenced by differences in marker number. In our study, 265 sequences with polymorphic microsatellites were obtained from SLAF-seq data. Then, subpopulations containing different numbers (5, 6, 7,…, 15, 20, 25, 30, 35, 40) of markers were genotyped 10 times to investigate the impact of marker numbers on population genetic diversity results. Our results show that genotyping with less than 11 or 12 microsatellite markers lead to significant deviations in the population genetic diversity or genetic structure results. In order to provide markers for population genetic and conservation studies for Rhododendron, 26 SSR primers were designed and validated in three species.
Similar content being viewed by others
Introduction
Microsatellites (simple sequence repeats, SSRs) have been the most frequently used genetic marker in population genetics over the past 20 years1. Although single nucleotide polymorphic loci (SNPs) have been used in a variety of genetic studies2 recently, they still have not replaced SSRs completely as microsatellites are highly informative, codominant, exhibit high specificity, are transferable among related species and have relatively low costs. As such, microsatellites continue to be widely used, especially for wild species, particularly when sample sizes are large. The number of loci used for genotyping is one of the key issues concerning the use of microsatellites. Koskinen et al. show a substantial decrease in standard deviation estimates of the interpopulation genetic distances by increasing the number of loci from six to seventeen. They indicate that the stability of commonly used genetic distances and phylograms is determined by the number of microsatellites investigated3. In addition, the results obtained from 205 red deer indicate significant effects on population genetic parameter if the number of microsatellite loci smaller than six4. However, in experimental studies of Drosophila nigrosparsa, Arthofer et al. demonstrate that a large proportion of individuals can still be correctly assigned to population of origin when using eight loci, and the population structure is still retained when using only two highly polymorphic loci5.
Unfortunately, to investigate the impact of different numbers of loci on population genetic results, the subpopulations containing different loci were generated from the total set of 16 or 17 microsatellite loci in previous studies3,4. Compared with the total number of microsatellites in genome, 16 or 17 microsatellite loci are too low and may be inconsistent with actual experimental design. This is limited by previous de novo SSR development, which can be a tedious and costly process6. More recently, next generation sequencing (NGS) technology has facilitated the development of hundreds microsatellite loci based on sequence data with reduced cost and effort7,8,9,10, offering the possibility of resampling different numbers of microsatellites from genome.
In our previous study, we estimated the population genetic structure and demographic history of two closely related species of Rhododendron, R. dauricum, and R. mucronulatum, distributed in northeastern China, using 664,406 SNPs based on specific-locus amplified fragment sequencing (SLAF-seq), a recently developed, high-resolution strategy for the discovery of large-scale de novo genotyping of SNPs11,12.
Here, using SSR sequences from this dataset, we evaluated the impact of using different numbers of SSR loci, generated from the whole genome, on the stability of population genetic results. Furthermore, we used the SLAF-seq data to develop polymorphic microsatellite markers for R. dauricum and R. mucronulatum. Rhododendron species are widely distributed around the world ranging from tropical to polar climates, and used as valuable horticultural plants due to their beautiful vegetative forms and remarkably bright-colored flowers13. The microsatellite markers developed in this study will aid genetic diversity studies of Rhododendron.
Materials and methods
SLAF data and microsatellite mining
On the basis of our previous SLAF sequencing data of R. dauricum and R. mucronulatum (accession number: PRJNA589346), we removed some populations that contain only one individual and merged some populations were close to each other geographically. Our dataset for this part consisted of 38 R. dauricum and 25 R. mucronulatum samples, and the sample vouchers have been deposited at the Northeast Normal University Herbarium (NENU, Table S1). All samples were identified by an expert taxonomist, Dr. Mingzhou Sun and Prof. Hongxing Xiao, Northeast Normal University, China. All filtered SLAF reads were clustered by the BLAT software according to sequence similarity to create SLAF tag sequences14. Polymorphic SLAF tags showed sequence polymorphisms between different samples.
Microsatellite motifs from di-nucleotides to hexa-nucleotides were identified from the polymorphic SLAF tag sequences in MISA-web (http://misaweb.ipk-gatersleben.de/). The lowest threshold of repeats for dinucleotides was set to six, while all others were set to five. Two SSRs motifs with the maximum interruption less than 100 bp were considered as one compound microsatellite.
Additionally, insertions/deletions (INDELs) were called using the program SAMtools 15 and Genome Analysis Toolkit (GATK)16 with our previously used parameters11. Raw INDELs were filtered using our custom Perl scripts with the cutoff “mapping quality (MQ) > 30, read depth (DP) > 3.” Moreover, PLINK 217 was used to further filter with the minor allele frequency (MAF) of 0.04 and maximum missing rate of 0.1. Finally, microsatellite genotypes for each individual were determined based on the sequence length of core motifs.
Genetic diversity and structure based on different number of microsatellite loci
Inbreeding coefficients (FIS) and corresponding p-values, which indicate whether markers or populations deviate from Hardy–Weinberg equilibrium, were tested by 1000 random permutations using FSTAT version 2.9.3.218. When FIS values for a locus deviated significantly from zero (p < 0.01), loci were excluded from further analyses. In addition, number of alleles (NA), allelic richness (Ar) and genetic diversity (Hs)19 were calculated for each species using FSTAT software. The population genetic structure was analyzed using the Bayesian clustering program STRUCTURE version 2.3.320. The admixture model with correlated allele frequencies was chosen, as recommended for faint population structures. The number of clusters (K) assumed was set to [1, 10], and each value of K was run 10 times. Each run was performed with 20,000 MCMC iterations and an initial burn-in of 180,000. The final posterior probability of K, ln p(K), and Delta K (ΔK) was calculated using STRUCTURE HARVESTER21 to determine the most likely K value.
To assess the effect of the number of microsatellites on the stability of the genetic diversity and genetic structure, population genetic analysis was assessed by the following procedure: data files consisting of 5, 6, 7,…, 15, 20, 25, 30, 35, 40 microsatellite loci were created by resampling from the complete data set randomly using a python script and repeated 10 times for each subset of microsatellite loci. Genetic diversity parameters and genetic structure analysis were constructed for each replicated data set as described above. Statistical analysis was done used one sample t-test with the IBM-SPSS package version 24.
Development of highly polymorphic microsatellite markers derived from SLAF sequences
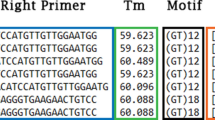
Polymorphism is one of the important criteria for judging the usability of microsatellite markers. To determine if SLAF data of populations can be used for highly polymorphic microsatellite marker development, we selected 66 loci with both highly polymorphic microsatellite motif (at least 4 alleles/locus) and at most one individual missing data for primer design, and finally only 40 pairs of primers were synthesized since their flanking regions were long enough. All primers were designed by the program Primer v3 (http://bioinfo.ut.ee/primer3-0.4.0/). The primer size ranged from 18 to 22 bp with the optimal size of 20 bp. The optimum GC content was 50%, the optimum melting temperature was 60 °C (ranged from 50 to 65 °C), and the maximum acceptable difference between the melting temperatures of the forward and reverse primers was 5 °C.
PCR validation and polymorphism examination
To test the use of polymorphic microsatellite markers we designed in Rhododendron species, total genomic DNA was extracted from 18 samples of population AES (R. dauricum), 10 samples of population JC (R. mucronulatum), moreover, and 12 individuals of R. aureum following a modified CTAB procedure22 and verified by electrophoresis on 1% agarose gel. PCR amplifications were performed in 20 μL reactions containing 50 ng genomic DNA, 1 × PCR buffer (plus Mg2+), 0.2 mM of dNTPs, and 0.5 μM of each primer, with each forward primer labeled with fluorescent dye (FAM, TAMRA, or HEX) (Invitrogen) and 1 unit (U) of Taq polymerase (Takara). Thermal cycling began with an initial denaturation step at 95 °C for 5 min, followed by 35 cycles of 30 s at 94 °C, 30 s at an optimal annealing temperature (Table 1), and 30 s at 72 °C, and a final elongation step at 72 °C for 8 min. The amplified fragments were resolved using an ABI 3730 DNA Analyzer (Applied Biosystems) using GeneScan 500 ROX as an internal size standard (Applied Biosystems, USA). Allele sizes were determined with the Peakscanner 2.0 software (Thermofisher Scientific, Germany).
Results
Composition and characteristics of microsatellites in Rhododendron SLAF data
By analyzing the SLAF data of 63 individuals, we obtained 555,834 sequences. After screening these sequences, a total of 57,951 microsatellites were identified from 41,121 sequences and 10,705 sequences (26.03%) contained more than one microsatellite motif. Di-nucleotides were the most abundant repeat motif (86.10%, 49,584), followed by tri-nucleotides (11.69%), tetra-nucleotides (1.38), penta-nucleotides (0.48%) and hexa-nucleotides (0.34%). In addition, the highest number of repeats per locus was 6 (13,187, 22.90%) (Table 2). Of the di-nucleotides, the most frequent motifs were AG/CT repeats (38,339, 77.32%) (Fig. 1). The dominant repeats in the tri-nucleotide were AAG/CTT (2050, 30.44%), followed by ACC/GGT (1133, 16.82%) while ACG/CGT had the lowest frequency (117, 1.74%) Among the tetra-nucleotide motifs, AAAT/ATTT (227, 28.66%) was the most dominant motif (Fig. 1).
Effects of the number of microsatellites on the stability of population genetics
Among the 41,121 sequences containing microsatellites, a total of 275 polymorphic microsatellites were selected after filtering by PLINK 2. The FIS values of ten loci deviated significantly from zero (p < 0.01), so they were excluded from subsequent analyses. Among the remaining 265 loci, the number of alleles per locus was 7, 6, 5, 4, 3 and 2 for 91, 48, 39, 18, 25 and 44 loci, respectively. For the remaining 265 loci, allelic richness (Ar) and the genetic diversity (Hs) measured 1.508 and 0.511 for R. dauricum and 1.445 and 0.444 for R. mucronulatum, respectively.
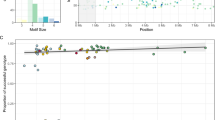
Though the total mean values from all measurement repetitions did not deviate significantly from the value of 265 loci for any of the population genetic parameters (Fig. S1), the standard deviations of Ar and Hs decreased dramatically with increasing number of loci (Fig. 2). With five loci, the standard deviations of the Ar and Hs were very high, being up to 25% of the Ar and Hs based on all 265 loci (Fig. 2). Moreover, the absolute deviations were statistically significant when there were less than 11 microsatellites (p < 0.01). The average Ar for R. dauricum with 5 to 10 markers were from 1.553 to 1.745 (Fig. S1a), deviated 7.15% to 22.28% and a maximum of 44.16% to 67.04% from the number based on 265 loci (data not show). And for R. mucronulatum, the average Ar with 5 to 10 markers were from 1.483 to 1.630 (Fig. S1b), deviated 6.61% to 18.17% from the number based on 265 loci (data not show).
STRUCTURE analysis based all SSR markers (265 loci) showed clear differentiation between species (Fig. 3), similar to that detected by SNPs in our previous analysis11. All individuals were divided into two clusters according to the highest ΔK (Fig. S2). The resulting STRUCTURE plots for K = 2 of different microsatellite loci with the highest ln p(k) are given in Fig. 4. The number of admixed individuals decreased as more loci were used, especially in R. mucronulatum. Compared with the full set of 265 loci, with fewer than twelve loci, at least one incorrect cluster was detected in all datasets. Remarkably, when data of only five or six loci were used, the error rate of populations clustering reached 50%.
PCR validation of SSR primers
Among the 265 polymorphic microsatellites, a total of 40 primer pairs were designed for evaluating PCR amplification efficiency and polymorphism in congeneric species. Of the primers tested, 14 primers were excluded from further analysis because these primers did not generate clear microsatellite peaks or failed to be amplified. The remaining 26 primers exhibited high amplification success and were screened in 40 samples from three species. Of the 26 polymorphic SSR loci, 23 were dinucleotides, two were trinucleotides and one was a tetranucleotide.
In total, 274 alleles were detected across all individuals, with the number of alleles per locus (NA) ranging from 4 to 19. The Ar values ranged from 3.023 to 9.339, with an average of 6.082, while the HS varied from 0.297 to 0.917, 0 to 0.929 and 0 to 0.968 in R. dauricum, R. mucronulatum and R. aureum, respectively (Table 1).
Discussion
Simple sequence repeats (SSRs) are co-dominant nuclear markers that are widely used in population genetic studies, which provide insights and guidelines for preserving the genetic diversity of populations23,24. Large number of papers published in the last few years have involved the use of microsatellites. However, as shown here and elsewhere, the low number of loci used may lead to erroneous conclusions when comparing populations3.
For example, in our study, significant deviations in Ar were found when using less than 11 loci, compared to the value from the full dataset of 265 loci. Moreover, the genetic diversity parameter Hs of R. dauricum was significantly different from the Hs based 265 loci (0.511) when compared to analyses of less than 10 microsatellites (Fig. 2).While for R. mucronulatum, the Hs value already deviated significantly from the Hs based 265 loci (0.444) when used 25 microsatellite loci (Fig. 2), it may be caused by few individuals of R. mucronulatum in this study. Experimental studies in red deer showed that significant deviations from the actual values for sample sizes of less than 30 per population4. Allelic richness (Ar), one of the most reported measures of genetic variation, is also referred to as allelic diversity or mean number of alleles per locus. And in FSTAT software, the Ar is corrected for sample size, thus compared with Ar, Hs is more sensitive to the sample size.
These effects could also be confirmed for the population genetic structure, which was investigated by STRUCTURE. Arthofer et al. demonstrate that the population structure was still retained, though about a quarter of a individuals cannot be correctly assigned, when using only two loci with the Arwere 12.94 and 13.54, , respectively5, which were much larger than the average Ar of microsatellite sites in the other study25,26,27. Thus, it is difficult to de novo develop microsatellite primers with such high polymorphism. Therefore, two microsatellite loci are far from enough in actual genetic structure research. Moreover, our previous study illustrated R. dauricum and R. mucronulatum clustered into distinct groups and showed majority populations collected from the Changbai Mountains (MES, SL, LTS, MH, HC, LJ, CB, WT) of R. dauricum with some admixture from R. mucronulatum11. However, in our study, the differentiation between species was not obvious with low microsatellite loci used. Remarkably, the incorrect assignment was still possible with fewer than 12 loci. With decreasing number of microsatellites, a reliable comparison between species cannot be achieved, even if Bayesian methods are used.
In addition, 26 polymorphic microsatellite loci were validated and characterized for individuals of R. dauricum, R. mucronulatum and R. aureum. The levels of diversity observed at these microsatellite loci, measured as allelic richness (Ar) and genetic diversity (Hs), were similar to those in previous studies11. Rhododendron is a familiar ornamental plant worldwide, ranging from tropical to polar climates13. This study provides a potentially highly polymorphic SSR markers library for the research of Rhododendron subgen. Rhodorastrum, which will facilitate the further study of the genetics of Rhododendron subgen. Rhodorastrum, even Rhododendron. Furthermore, we explored a simple route to develop polymorphic SSR markers from non-model species based on SLAF-seq, which is well suited for polymorphic SSR marker discovery in non-model organisms.
Conclusion
Previous studies showed considerable differences of genetic diversity and genetic structure with regard to the number of microsatellite loci. Our results indicated significant effects on population genetic parameters if the number of microsatellite loci was less than 12. With decreasing marker numbers, the accuracy of population genetic of and the genetic structure decreases. Fortunately, the SLAF-seq data of populations offers an effective approach to develop polymorphic microsatellite markers for non-model species. The 26 polymorphic microsatellite markers we developed for Rhododendron species will be important for investigating population genetic diversity and genetic structure, and these results in turn will provide crucial information for conservation and management of Rhododendron species.
References
Vieira, M. L. C. et al. Microsatellite markers: What they mean and why they are so useful. Genet. Mol. Biol. 39(3), 312–328 (2016).
Kumar, D. et al. Characterization of genetic diversity and population structure in wheat using array based SNP markers. Mol. Biol. Rep. 47(1), 293–306 (2020).
Koskinen, M. T. et al. The benefits of increasing the number of microsatellites utilized in genetic population studies: An empirical perspective. Hereditas 141(1), 61–67 (2004).
Reiner, G., Lang, M. & Willems, H. Impact of different panels of microsatellite loci, different numbers of loci, sample sizes, and gender ratios on population genetic results in red deer. Eur. J. Wildl. Res. 65(2), 25 (2019).
Arthofer, W. et al. Identifying the minimum number of microsatellite loci needed to assess population genetic structure: A case study in fly culturing. Fly 12(1), 13–22 (2018).
Hodel, R. G. et al. The report of my death was an exaggeration: A review for researchers using microsatellites in the 21st century. Appl. Plant Sci. 4(6), 1600025 (2016).
Davey, J. W. et al. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 12(7), 499–510 (2011).
Jia, X. et al. Single-molecule long-read sequencing of the full-length transcriptome of Rhododendron lapponicum L.. Sci. Rep. 10(1), 1–11 (2020).
Qin, H. et al. Using MiddRAD-seq data to develop polymorphic microsatellite markers for an endangered yew species. Plant Divers. 39(5), 294–299 (2017).
Jia, B. et al. Development of novel EST microsatellite markers for genetic diversity analysis and correlation analysis of velvet antler growth characteristics in Sika deer. Hereditas 157(1), 1–14 (2020).
Yang, B. et al. A genomewide scan for genetic structure and demographic history of two closely related species, Rhododendron dauricum and R. mucronulatum (Rhododendron, Ericaceae). Front. Plant Sci. 11, 1093 (2020).
Lyu, Y.-Z. et al. SLAF-seq uncovers the genetic diversity and adaptation of Chinese Elm (Ulmus parvifolia) in Eastern China. Forests 11(1), 80 (2020).
Xing, W. et al. De novo assembly of transcriptome from Rhododendron latoucheae Franch. using Illumina sequencing and development of new EST-SSR markers for genetic diversity analysis in Rhododendron. Tree Genet. Genomes 13(3), 53 (2017).
Kent, W. J. BLAT—The BLAST-like alignment tool. Genome Res. 12(4), 656–664 (2002).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25(16), 2078–2079 (2009).
Depristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43(5), 491 (2011).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81(3), 559–575 (2007).
Goudet J. FSTAT, a Program to Estimate And Test Gene diversitIes and Fixation Indices, Version 2.9, Vol. 3 (2001). Accessed 1 Oct 2020. http://www2.unil.ch/popgen/softwares/fstat.htm.
Nei, M. Molecular Evolutionary Genetics (Columbia University Press, 1987).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155(2), 945–959 (2000).
Earl, D. A. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4(2), 359–361 (2012).
Doyle, J. & Doyle, J. Genomic plant DNA preparation from fresh tissue-CTAB method. Phytochem. Bull. 19(11), 11–15 (1987).
Feng, X., Wang, Y. & Gong, X. Genetic diversity, genetic structure and demographic history of Cycas simplicipinna (Cycadaceae) assessed by DNA sequences and SSR markers. BMC Plant Biol. 14(1), 187 (2014).
Liu, Y. et al. Genetic structure and diversity of Glycyrrhiza populations based on transcriptome SSR markers. Plant Mol. Biol. Rep. 37(5–6), 401–412 (2019).
Dachapak, S. et al. Genetic diversity and structure of the zombi pea (Vigna vexillata (L.) A. Rich) gene pool based on SSR marker analysis. Genetica 145(2), 189–200 (2017).
Torokeldiev, N. et al. Genetic diversity and structure of natural Juglans regia L. populations in the southern Kyrgyz Republic revealed by nuclear SSR and EST-SSR markers. Tree Genet. Genomes 15(1), 1–12 (2019).
Zhou, Q. et al. Analysis of genetic diversity of ancient Ginkgo populations using SSR markers. Ind. Crops Prod. 145, 111942 (2020).
Acknowledgements
We would like to thank Mr. Mingzhou Sun for his help in sample collection. We appreciate Ling Zhao for their technical assistance in PCR validation of SSR primers.
Funding
This work was supported by the Natural Science Foundation of the Science and Technology Department of Jilin Province (Subject layout project: 20190201184JC).
Author information
Authors and Affiliations
Contributions
Conceptualization, Huaying Wang and H.X.; Formal analysis, Huaying Wang and Huan Wang; Funding acquisition, H.X.; Methodology, Huaying Wang, B.Y., Huan Wang and H.X.; Supervision, H.X.; Writing—original draft, Huaying Wang and B.-M.Y.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, H., Yang, B., Wang, H. et al. Impact of different numbers of microsatellite markers on population genetic results using SLAF-seq data for Rhododendron species. Sci Rep 11, 8597 (2021). https://doi.org/10.1038/s41598-021-87945-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87945-x
This article is cited by
-
Complete plastid genome structure of 13 Asian Justicia (Acanthaceae) species: comparative genomics and phylogenetic analyses
BMC Plant Biology (2023)
-
Development of twenty-one novel microsatellite loci for Gila topminnow, Poeciliopsis occidentalis occidentalis
Molecular Biology Reports (2023)
-
An integrative study of species distribution modelling and conservation genetics: Magnolia in Hispaniola
Biodiversity and Conservation (2023)
-
Performance comparison of gel and capillary electrophoresis-based microsatellite genotyping strategies in a population research and kinship testing framework
BMC Research Notes (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.