Abstract
Alterations to cell polarization or to intercellular junctions are often associated with epithelial cancer progression, including breast cancers (BCa). We show here that the loss of the junctional scaffold protein MAGI1 is associated with bad prognosis in luminal BCa, and promotes tumorigenesis. E-cadherin and the actin binding scaffold AMOTL2 accumulate in MAGI1 deficient cells which are subjected to increased stiffness. These alterations are associated with low YAP activity, the terminal Hippo-pathway effector, but with an elevated ROCK and p38 Stress Activated Protein Kinase activities. Blocking ROCK prevented p38 activation, suggesting that MAGI1 limits p38 activity in part through releasing actin strength. Importantly, the increased tumorigenicity of MAGI1 deficient cells is rescued in the absence of AMOTL2 or after inhibition of p38, demonstrating that MAGI1 acts as a tumor-suppressor in luminal BCa by inhibiting an AMOTL2/p38 stress pathway.
Similar content being viewed by others
Introduction
Breast cancers (BCa) arise from mammary epithelial cells, and show tremendous diversity. Based on transcriptome, BCa are typically divided into five main subtypes: luminal A, luminal B, HER2-enriched, normal-like, and basal (reviewed in1,2), partly overlapping with previous classifications (such as ER/PR/HER2 receptors expression status). Basal tumors, overlapping strongly with triple-negative breast cancers, are the most aggressive and present the worst prognosis. However, even though luminal A and B subtypes (typically ER/PR positives) usually respond well to current treatments, when “luminal” tumors relapse, they become very difficult to treat with very poor outcome.
The apico-basal (A/B) polarity of mammary epithelial cells is established and maintained by the asymmetric segregation of evolutionarily conserved protein complexes3. A/B polarity governs the correct position of different intercellular junctions that ensure the integrity of the epithelial sheet. Adherens Junctions (AJs) constitute apical adhesive structures mediated by the homophilic trans-interactions of E-cadherin/catenin complexes4,5. At the very apical part of epithelial cells, Tight Junctions (TJs) ensure the impermeability of tissues in part through the action of Claudins/Zonula Occludens proteins complexes6. The transmembrane proteins of these different junctions (E-cadherin, Claudins…) are coupled and anchored to the actin cytoskeleton via adaptor scaffold proteins, thus coupling adhesion and cell contacts to actin dynamics. As such AJs represent mechano-sensory structures, where tensile or compressive forces, arising from internal or external sources, will be sensed to promote short term responses such as changes in adhesion and actomyosin dynamics, but also long term responses to control cell density, in particular through the mechano-sensitive Hippo pathway7,8. In epithelial cells, the compressive forces exerted by the cortical actin cytoskeleton or by cell packing, sensed at AJs, antagonize the proliferative and anti-apoptotic effects of YAP/TAZ in part through α-catenin conformational change, or through the release of Merlin/NF29,10,11,12,13,14. Other signaling pathways such as β-catenin, ERK, or JNK/SAPK stress pathways can also be influenced by polarity and forces sensed at junctions11,15,16,17,18. BCa are derived from transformed mammary epithelia where frequent alterations in TJs and/or AJs composition and structure, and the associated abnormal mechano-sensory responses have been linked to cell dysfunction such as uncontrolled cell proliferation and metastasis19,20.
At apical junctions, cortical scaffold proteins such as MAGI (MAGUKs with Inverted domain structure), assemble large and dynamic molecular complexes. In humans, among the three MAGI proteins (MAGI1-3), MAGI1 is the most widely expressed and contains several protein–protein interaction domains including six PDZ and two WW domains21 mediating interactions with TJ proteins22. In Drosophila, we have demonstrated that Magi, the sole fly MAGI homolog, is required for E-cadherin belt integrity and AJ dynamics ultimately restricting cell numbers in the developing eye23,24. The regulation of Cadherin complexes by MAGIs, is evolutionarily conserved from C. elegans25 to vertebrates, and in particular vertebrate endothelial cells where MAGI1 links VE-cadherin complexes with the underlying cortical actin cytoskeleton26. A tumor suppressive role of MAGI1 (reviewed by Feng et al.27) is suggested by the correlation between low MAGI1 expression and poor prognosis in various cancers, such as hepatocellular carcinoma28. The anti-tumoral activity of MAGI1 is further supported by its ability to bind the tumor-suppressor PTEN29,30,31, the fact that it is often targeted by viral oncoproteins32,33, and by the observation that in colorectal cancer cells, MAGI1 was upregulated in response to cyclooxygenase-2 inhibitor and prevented metastasis34. Thus it appeared that the apical junction-localized MAGI1 scaffold protein participates in multiple complexes to fine-tune adhesion and signaling, and may therefore be considered as a tumor suppressor.
The angiomotin (AMOT) family of membrane-associated scaffold proteins is composed of three members: AMOT, AMOTL1, and AMOTL2, playing important roles in the regulation of intercellular junctions35. In endothelial cells and in zebrafish developing embryos, AMOTL2 has been shown to link cadherins and the actin cytoskeleton. In particular, AMOTL2 is required for the maintenance of tension at the level of cadherin junctions to properly shape strong blood vessels26,36. Moreover, AMOTs are also known to control the Hippo tumor-suppressor signaling pathway. Canonical Hippo pathway, through sequential activation of the Hippo/MST and Wts/LATS kinases, promotes the phosphorylation and cytoplasmic retention of the transcriptional co-activators Yki/YAP/TAZ (reviewed in37). Specific PPxY motifs found in AMOTs interact with the WW domains of YAP and TAZ38 and AMOTs act thus as negative regulators of YAP/TAZ mediated transcription by trapping YAP38,39 and TAZ40 in the cytoplasm. Recent studies identified interactions between MAGI1 and AMOTs, in particular in the regulation of intercellular junctions41,42,43.
We report here that low levels of MAGI1 are associated with bad prognosis in luminal ER + BCa. Impairing MAGI1 expression in luminal BCa cells promoted their growth in 2D, and their ability to grow in low attachment conditions in-vitro and in subcutaneously grafted nude mice. We further document that MAGI1 deficient cells accumulate specific junctional proteins, including E-cadherin and AMOTL2. Mechanistically, we show that all MAGI1 deficient phenotypes are suppressed by down-regulating AMOTL2 suggesting that AMOTL2 and its effects on junctions is the primary cause of the increased tumorigenicity after MAGI1 loss. Consistent with these observations, we could further show that MAGI1 deficient luminal BCa cells experience higher stiffness as evidenced by increased Young’s modulus, and ROCK activity as evidenced by increased ROCK-specific Ser19 phosphorylation of the regulatory Myosin Light Chain 2. They activate the p38 stress signaling pathway while YAP activity is antagonized. Finally, we provide evidence that releasing actin cytoskeletal strength, or impairing p38 activity can revert the effects of MAGI1 loss, supporting a model by which, in response to MAGI1 loss, elevated AMOTL2/E-cadherin and junction dysfunction, together with actin cytoskeletal tension activate the p38 stress pathway fueling tumorigenicity.
Results
The loss of MAGI1 enhances tumorigenicity of luminal breast cancer (BCa) cells
In order to study the role of the apical scaffold MAGI1 during breast carcinogenesis, we first analyzed the expression of MAGI1 by western blot in both luminal and basal BCa cell lines: MAGI1 expression was restricted to luminal ER+ lineages (T47D and MCF7 luminal A, and to a lesser extent in ZR75 luminal B), while no expression could be detected in basal ER− lineages (immortalized MCF10A and triple negative MDA-MB-468 and BT549; Supplementary Fig. S1A). Moreover, public database mining (http://www.kmplot)44 and recent studies45, indicated that low MAGI1 expression levels were associated with worse prognosis in relapse-free survival for BCa patients, but only in ER+ (mainly luminal) molecular BCa subtypes (Kaplan Meier curve; Supplementary Fig. S1B).
We thus decided to investigate the functional role of MAGI1 and the consequence of MAGI1 knockdown in luminal BCa, and generated MCF7 and T47D cell lines in which MAGI1 was targeted by constitutive shRNA. Two independent shRNA constructs targeting different parts of the MAGI1 transcript were used, shMAGI1(3-1) and shMAGI1(1-1), which led respectively to more than 90% and around 60% MAGI1 knockdown at the protein level as shown by western blot and immunofluorescence analyses (Supplementary Fig. S1D&E). The knockdown was specific for MAGI1 and did not induce compensatory up-regulation of MAGI2 and MAGI3, the two remaining MAGI family members (Supplementary Fig. S1D&F). shMAGI1(3-1) was the most potent resulting in almost knock-out like down-regulation, and was therefore used in all subsequent studies and referred to as shMAGI1 thereafter. It should be noted that in the subsequent analyses of MAGI1 function similar effects were obtained using the independent shMAGI1(1-1) ruling out non-specific phenotypes of the shMAGI1(3-1) (Supplementary Fig. S2A-F).
First, we studied the effect of MAGI1 knockdown on cell numbers and proliferation. MAGI1 depletion increased 2D cell growth of MCF7 cells (20–25% increase at day 4 and day 7) as assayed by MTT cell growth assay (Fig. 1A). At day 7, it was associated with an increase in the proportion of cells in the S phase of the cell cycle from 41% in control MCF7shLuc to 54% in MCF7shMAGI1 cells as assessed by BrdU incorporation (Fig. 1B) with no discernable change in apoptotic cells (see Legend of Fig. 1B). More importantly, in anchorage-independent soft agar cell growth conditions, MCF7shMAGI1 cell lines formed circa twice as many colonies compared to shLuc controls (Fig. 1C). Similarly, MCF7shMAGI1 cells showed enhanced clonal mammosphere capacities which were correlated with an expanded area occupied by the spheres (60% increase; Fig. 1D). This increase in mammosphere formation is associated with a slight increase in the proportion of cells presenting stemness markers (to circa 4% of total cells) as evidenced by increase in CD44High/CD24Low (1.7x) and ALDH activity (2.6x). Then, we studied the effect of MAGI1 knock-down on cell migration and could not detect any effect on collective cell migration (scratch wound assay), and on invasion (boyden chamber assay; Supplementary Fig. S3A-C). Importantly, similar results were obtained in T47D, a second luminal A BCa cell line, in which MAGI1 knockdown (shMAGI1) led to elevated 2D and anchorage-independent cell growth (Supplementary Fig. S4A,B&G). Taken together, these results show that the loss of MAGI1 in luminal BCa cells promotes cancer cell proliferation and anchorage-independent growth, but not migration/invasion.
MAGI1 impairment induces tumorigenic phenotypes in epithelial cells. (A) MTT assay (OD 560 nm) representing 2D cell growth of MCF7shMAGI1 as compared to MCF7shLuc cells. Bars represent mean ± Standard Deviation (SD; n = 10 wells as replicates) of a representative experiment (out of 3). Unpaired two-tailed Student's t-test; **p < 0.01; ***p < 0.001. (B) Cell cycle phases of MCF7shLuc and MCF7shMAGI1 cells assessed after BrdU incorporation and analyzed by flow cytometry, showing an increased proportion of cells in S phase at the expense of G0/G1 after MAGI1 invalidation. The pie chart also showed no major difference in the percentage of apoptotic cells in SubG1 phase (around 2% in shLuc and 1% in shMAGI1 cells). (C) Left: quantification of colony numbers of MCF7shMAGI1 cells grown in anchorage independent conditions (soft agar assay) and represented as fold increase compared to MCF7shLuc cells. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; ***p < 0.001. Right: western blot analysis of whole protein extracts issued from the cells used in the soft agar assays and showing the relative amounts of MAGI1 in the different cell lines; Tubulin was used as a loading control. Uncropped blots can be found in the Supplementary Information. (D) Top: quantification of mammospheres represented as total area issued from MCF7shMAGI1 cells and shown as fold increase compared to MCF7shLuc cells, starting from 200 cells. Data are presented as the means ± SD (n = 4). Unpaired two-tailed Student's t-test; ***p < 0.001. Bottom: images of mammosphere formation assay showing a representative experiment.
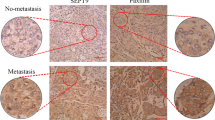
Significantly, when MCF7shMAGI1 cells were injected subcutaneously in nude mice, they grew as a tumor mass much more rapidly and extensively than control MCF7shLuc cells (Fig. 2A). This dramatic increase in tumor growth was not due to obvious changes in the tumor microenvironment and comparable density of immune cells, fibroblasts, and vascular cells infiltrates were observed in shMAGI1 tumors compared to controls as reflected by CD45, α-SMA, and CD31 staining respectively (Supplementary Fig. S5A-J). Importantly, higher Ki67 reflecting increased number of mitotic cells was observed in the MCF7shMAGI1 tumors (Fig. 2B–D) and among the proliferative tumor cells, the Ki67 staining was more intense (Fig. 2E), showing that indeed MAGI1-deficient cells proliferate more, and suggesting that the increased tumor growth upon MAGI1 depletion is primarily due to increased tumor cells burden. Even though BCa was the prime focus of this study, the effects of MAGI1 knockdown appeared not restricted to mammary cells, and similar observations on anchorage-free growth in soft agar, and orthotopic tumor growth in nude mice were also observed using HCT116 colon cancer cells (Supplementary Fig. S4E-G) extending earlier reports34. Altogether, our results show that the loss of MAGI1 in luminal A BCa cells enhances their tumorigenic behaviors.
The loss of MAGI1 in primary tumors induces tumor cell proliferation. (A) Primary tumor growth of MCF7shLuc and MCF7shMAGI1 cells injected subcutaneously in nude mice. Primary tumor growth was assessed by measuring the tumor volume over time until the tumors were too big and the mice had to be euthanized. Bars correspond to the mean ± Standard Error to the Mean (SEM; n = 8 mice per group). Unpaired two-tailed Student’s t-test; *p < 0.05. (B,C) Representative immunohistochemical staining of Ki67 in primary tumors obtained after subcutaneous engraftment of either MCF7shLuc (B) or MCF7shMAG1 cells (C). Brown staining indicates positive immunoreactivity. Scale bar = 250 µM. In the magnified area, the red * highlights an increased number of mitotic cells in the MCF7shMAGI1-derived primary tumor. (D,E) Quantification of the immunohistochemical analysis in (B,C), showing the percentage of primary tumor cells stained positively for Ki67 (D) and the normalized mean intensity of Ki67 positive cells (E) in MCF7shMAGI1 and MCF7shLuc primary tumors. Bars correspond to the mean ± SD (n = 4 for shLuc and n = 5 for shMAGI1). Unpaired two-tailed Student’s t-test; **p < 0.01 and ***p < 0.001.
The loss of MAGI1 promotes the accumulation of epithelial junctions components
In order to further investigate the role of MAGI1 in breast tissue, we sought to determine its localization in mammary epithelial cells. Immunohistochemistry on BCa patient tissue micro-array revealed that in normal breast cells, MAGI1 was expressed only in the luminal epithelial cells and not in the underlying basal myo-epithelial layer, consistent with MAGI1 expression exclusively in luminal-type BCa cell lines (Supplementary Fig. S1A). At the sub-cellular level, MAGI1 was localized at the apical pole of luminal breast cells (Supplementary Fig. S1C, arrows). Using MCF7 cancer cells, immortalized hMEC (human mammary epithelial cells), and polarized canine MDCK cells, we further confirmed by immunofluorescence that MAGI1 localized near the plasma membrane, overlapping with junction components such as E-cadherin (AJ), ZO1, and Claudin3 (TJ), suggesting that MAGI1 is a TJ and/or AJ resident protein (Fig. 3A, arrows and data not shown).
The loss of MAGI1 affects E-cadherin levels and localization in MCF7 cells. (A) Representative immunofluorescence experiments performed in MCF7 cells monitoring MAGI1 localization (red) compared to E-cadherin, ZO1, and Claudin3 (green). DAPI (blue) was used to stain DNA and the nuclei. White arrows indicate staining overlap. Scale bar = 10 µm. (B) Western blot analysis of whole protein extracts (n = 4) monitoring the expression the junctional components Claudin-1, ZO-1, β-catenin, PARD3, E-cadherin and AMOTL2 in MCF7shMAGI1 cell lines compared to MCF7shLuc. Tubulin was used as a loading control. Note the increase in E-cadherin levels. Western blot experiments have been repeated at least four times. Protein expression levels were quantified as compared to Tubulin. Uncropped blots can be found in the Supplementary Information. (C) Representative immunofluorescence experiments (n = 3) were performed on MCF7shMAGI1 compared to MCF7shLuc cells monitoring the localization of β-catenin, PARD3, Claudin-3, ZO-1 and E-cadherin and AMOTL2 (green). DAPI (blue) was used to stain DNA and the nuclei. Scale bar = 10 µm. Note the slight sub-cortical accumulation of E-cadherin shown in the high magnification images. (D) Representative E-cadherin staining (green) of MCF7shLuc and MCF7shMAGI1 shown through the Z axis. DAPI (blue) was used to stain DNA and the nuclei.
The localization of MAGI1 at apical junctions prompted us to explore whether MAGI1 could control their biology. Performing western-blot analyses on whole protein extracts, the loss of MAGI1 did not affect the total protein abundance of the typical TJs component ZO-1 nor of the AJs component PARD3 (Fig. 3B). However, in MCF7shMAGI1 we observed increases in the junctional proteins β-catenin (× 1.25), E-cadherin (× 1.6) and AMOTL2 (× 1.8) (Fig. 3B). Consistently, performing immunofluorescence on fixed MCF7shMAGI1 cells, the membrane levels and localization of ZO-1 and PARD3 were unchanged, while E-cadherin staining was increased, and less tightly restricted to the plasma membrane, and extending more basally than controls (Fig. 3C,D). The altered distribution of E-cadherin in shMAGI1 cells is reminiscent of the role of Magi in Drosophila where we showed that it controls E-cadherin belt at AJs in epithelial cells of the developing eye24.
In epithelial cells, the increase in E-cadherin material is often associated with increased junctional strength and, through the linkage to the underlying actin, to increases in tension and overall compressive forces (reviewed by7). While we did not observe obvious changes in the intensity or morphology of F-actin stained with phalloidin upon MAGI1 knockdown (Supplementary Fig. S6A), MCF7shMAGI1 cells exhibited behaviors compatible with increased compression. First, when cultured as spheroids, MCF7shMAGI1 cells grew as round masses of cells, rounder than MCF7shLuc controls with circa 30% smaller perimeter (Fig. 4A,B) and better circularity (0.95 vs 0.62 respectively; Fig. 4C), a sign that MCF7shMAGI1 cells aggregates were more compact, even though composed of the same number of cells as MCF7shLuc controls. It is noteworthy that MCF7shMAGI1 spheroids grew more (more cells) than controls reminiscent of the effect of MAGI1 knock-down in 2D and anchorage-independent cell growth assays (Fig. 4D). This compaction was also observed in T47D shMAGI1 cells (Supplementary Fig. S4C&D). Second, using Atomic Force Microscopy (AFM), we showed that MCF7shMAGI1 cells were stiffer than controls (higher Young’s modulus factor; Fig. 4E), strongly suggesting that MCF7shMAGI1 cells may have a stronger cytoskeleton and might experience higher internal pressure. Since AFM was performed on small MCF7 cells clusters, our data indicate that the increased compressive forces were generated from within the cells, likely through increased actin contractility. Indeed, elevated ROCK activity was detected in MAGI1 deficient cells as shown by the increase in ROCK-specific serine 19 phosphorylation of the regulatory myosin light chain MLC2 (Fig. 4F,G), suggesting that the actin cytoskeleton was under higher tension. Consistently, more actin-associated phosphorylated MLC2 (Ser19) positive foci were detected in MCF7shMAGI1 cells compared to MCF7shLuc cells (Supplementary Fig. S6A&B). Furthermore, treating MCF7shMAGI1 cells with ROCK inhibitors completely abolished the increased cell plasma membrane strength and the Young’s modulus factors measured by AFM after inhibitors treatment reached similarly low levels in shMAGI1 and shLuc controls (Supplementary Fig. S6C).
The loss of MAGI1 affects MCF7 cell compaction, ROCK activity, and compressive forces. (A) Representative phase contrast and fluorescence images of MCF7shLuc and MCF7shMAGI1 cells grown in 3D spheroid cultures and stained with E-cadherin (red) and DAPI (blue). (B) Calculated perimeters for 3D spheroid cultures of MCF7shMAGI1 normalized by the perimeter of MCF7shLuc cells. Bars represent mean ± SD (n = 10 spheroids) of five independent experiments. Unpaired two-tailed Student’s t-test; *p < 0.05. (C) Calculated circularity for 3D spheroid cultures of MCF7shLuc and MCF7shMAGI1 (calculations were done with the ImageJ software where a value of 1 is considered as a perfect circle). Bars represent mean ± SD (n = 10 spheroids) of five independent experiments. Unpaired two-tailed Student’s t-test; *p < 0.05. (D) MTT assay (OD 560 nm) representing 3D spheroid cell growth of MCF7shMAGI1 compared to MCF7shLuc cells 5 days after cell seeding. Bars represent mean ± SD (n = 10 spheroids) of five independent experiments. Unpaired two-tailed Student’s t-test; *p < 0.05. (E) Elastic Young’s modulus (EYM) of cells: Hertz contact mechanics model for spherical indenters was used. In MCF7shMAGI1 cells, the apical surface EYM is significantly elevated when compared to control MCF7shLuc cells. Data are represented as mean + /− SD: MCF7shLuc EYM = 258.8 + /− 101, MCF7shMAGI1 EYM = 302.4 + /− 91. Unpaired two-tailed Student’s t-test; *p < 0.05 (n = 132 and 94 for MCF7shLuc and MCF7shMAGI1 respectively). (F) Western blot analysis on whole protein extracts (n = 3) of ROCK-specific ser19 phosphorylation of Myosin Light Chain 2, total MLC2 and MAGI1 in MCF7shMAGI1 cell lines compared to MCF7shLuc. GAPDH was used as a loading control. Uncropped blots can be found in the Supplementary Information. (G) Quantification of the representative Western blot showing protein expression represented in panel E. Phosphorylated proteins were quantified as compared to their total protein counterparts to evaluate their activation.
MAGI1 loss does not promote YAP nor β-catenin signaling
In epithelial cells, E-cadherin-based AJs regulate the activity of mechanosensitive pathways, and in particular the Hippo and Wnt/β-catenin pathways, two major pathways controlling cell proliferation and frequently mis-regulated during carcinogenesis8,46. We thus explored whether these two pathways could be affected following MAGI1 depletion.
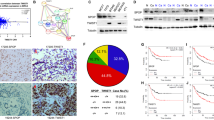
Monitoring Hippo pathway and YAP activity in stable MCF7shMAGI1 showed that in the absence of MAGI1, total YAP and phosphorylated YAP (on both serine residues S127 and S397) accumulated in the cells while TAZ levels remained unchanged (Western blot analyses; Fig. 5A). By immunofluorescence, we then showed that unlike what was observed for control cells grown at similar confluence, YAP did not accumulate in the nuclei of MCF7shMAGI1 cells (Fig. 5B). These findings were further confirmed in luminal BCa patients TMA, where we observed a strong correlation in ER+ tumors between MAGI1 apical membrane localization and YAP nuclear localization: only 27% of the patients with no MAGI1 expression or mis-localized MAGI1 (no membrane staining) showed YAP nuclear localization while 92% of patients with membranous MAGI1 showed nuclear YAP (Fig. 5C). Accordingly, MAGI1 knockdown in MCF7 cells resulted in impaired YAP transcriptional activity with decreased expression of canonical YAP target genes such as CTGF, CYR61, BIRC2, AXL, and AREG (Fig. 5D). The expression of AMOTL2, a YAP target gene encoding a negative YAP regulator, was increased after MAGI1 knockdown, suggesting that despite its regulation by YAP, AMOTL2 transcription is also affected by other inputs in response to MAGI1 knockdown. Surprisingly, even though YAP phosphorylations on S127 and S397 are mediated by LATS kinases (Reviewed in47,48), the loss of MAGI1 did not promote a higher activity of the Hippo pathway and the levels of phosphorylated activated LATS1 kinase and of phosphorylated MOB1 were unaffected (Fig. 5A). This suggests that the elevated phosphorylated YAP were not the consequence of increased upstream Hippo pathway activity (MST kinases), but might reflect higher p-YAP stability, either because it was protected from destruction, or because it was not dephosphorylated as efficiently, or a combination of the two. These results are in agreement with earlier studies showing that E-cadherin junctional tension in epithelial cells promote YAP/TAZ nuclear exclusion either through α-catenin conformational change12,13,14, or through Merlin/NF2-mediated cytoplasmic retention10.
The loss of MAGI1 tumorigenic phenotypes are not associated with active YAP. (A) Western blot analysis on whole protein extracts (n = 3) of phosphorylated YAP (S397 and S127), total YAP and TAZ, phosphorylated LATS1, total LATS1, phosphorylated MOB1, non-phosphorylated (active) β-catenin, total β-catenin and MAGI1 in MCF7shMAGI1 cell lines compared to MCF7shLuc. Tubulin was used as a loading control. Uncropped blots can be found in the Supplementary Information. (B) Representative immunofluorescence images (n = 3) of endogenous MAGI1 and YAP protein expression in MCF7shLuc and MCF7shMAGI1 cells cultured to similar confluence (60.4% and 63.2% respectively). Scale bar = 10 µm. (C) Quantification of MAGI1 and YAP staining in a tissue micro-array (TMA) from luminal breast cancer patients in a cohort of 28 patients, showing the strong correlation between MAGI1 membranous and YAP nuclear localizations; Fisher exact test, ***p < 0.001. Histogram shows the percentages of patients having a nuclear or a cytoplasmic YAP staining according to the absence or the presence of MAGI1 membranous staining. (D) Real-time qRT-PCR showing the expression of canonical Hippo pathway target genes (CTGF, CYR61, BIRC2, AXL and AREG) as well as AMOTL2 in MCF7shMAGI1 compared to the control MCF7shLuc cells and normalized with GAPDH expression. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; *p < 0.05; **p < 0.01; ***p < 0.001. (E) Real-time qRT-PCR showing the expression of two distinct Wnt pathway target genes (AXIN2 and CCND1) in MCF7shMAGI1 compared to the control MCF7shLuc cells and normalized with GAPDH expression. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test revealed no statistical differences. (F) WNT reporter activity assay showing the TOP/FOP ratio in MCF7shMAGI1 as compared to MCF7shLuc control cells. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test revealed no statistical differences.
Similarly, even though higher overall protein levels of β-catenin could be observed in MCF7shMAGI1 cells compared to controls, this was not reflected by an increase in the active unphosphorylated β-catenin pool (Fig. 5A). Accordingly, β-catenin transcriptional activity remained unchanged as evidenced by following by qPCR the expression of the AXIN2 and CCND1 canonical target genes or by TOP/FOP luciferase reporter assay (Fig. 5E,F). These results are in agreement with studies showing that increased compression transduced by apical junctions in epithelial cells prevent β-catenin nuclear accumulation11,16. Together, these results thus show that the loss of MAGI1 in luminal BCa cells does not promote YAP nor β-catenin signaling, and suggest that these pathways do not mediate the increased tumorigenicity of MAGI1 deficient cells.
p38 stress activated protein kinase mediates tumorigenicity of MAGI1 deficient cells
We thus asked which mechanisms mediate the increase in 2D cell proliferation and in anchorage-independent cell growth observed upon MAGI1 loss (see Fig. 1), focusing on other known oncogenic pathways. The ERK/MAP Kinase and JNK stress kinase pathways could be influenced by junctions and the forces they sense15,17,18. In response to MAGI1 loss, we did not detect any alterations of the main oncogenic pathways: ERK1/2, JNK and Akt pathways (Fig. 6A,B) in contrast with recently published results on MAGI1 function45. Interestingly, the p38 stress kinase was strongly activated in MCF7shMAGI1 compared to MCF7shLuc: the amount of phosphorylated p38, normalized to total p38 protein level, was increased 2 fold (Fig. 6A,B) leading to increased p38 activity as evidenced by the elevated expression of the p38 transcriptional target genes49 ATF3, SOX9, and GATA6 (SOX2 levels remained unchanged) (Fig. 6C). Importantly, similar results regarding the activity of the different signaling pathways upon MAGI1 knock-down, and in particular the increased activity of p38, were obtained in other cell lines (T47D luminal BCa and colon HCT116, Supplemental Fig. S4G), showing that the effects observed are not restricted to MCF7 cells and represent a more general cellular response to MAGI1 loss. The p38 pathway is activated in response to a wide variety of cellular stresses and has been implicated either as a tumor-suppressor or as an oncogene in various cancers, including BCa50,51,52,53,54. Using specific inhibitors and siRNA mediated silencing, we then tested whether the p38 pathway could mediate, at least in part, the increased tumorigenicity of MCF7 luminal cells. In 2D cell growth and in soft agar anchorage-independent cell growth, MCF7shMAGI1 treated with LY2228820 p38α/β inhibitor (a.k.a. Ralimetinib) grew less than untreated cells (2 fold suppression at 8 days of growth, and 2.5 fold decrease respectively; Fig. 6D,E). Similar effects were obtained using another independent p38 inhibitor, ARRY-614 (p38 and Tie2 inhibitor), even though we observed higher toxicity (data not shown) and importantly using p38α knockdown by siRNA (Fig. 6F,G), demonstrating that the effects observed were the consequence of p38 inhibition and not any unspecific action of the drugs. It should be noted that p38 inhibition also affected MCF7shLuc control cells (2D growth and soft agar), but the effect were far less pronounced than for MCF7shMAGI1 cells, strongly supporting that the increased p38 activity is mediating, at least in part, the increased tumorigenicity upon MAGI1 depletion. Taken together, our results demonstrate the essential role of the p38 signaling pathway for MAGI1-deficient MCF7 luminal BCa cells tumorigenicity, consistent with previous report identifying the critical role of p38α in mammary tumorigenesis using mouse BCa models50.
The loss of MAGI1 tumorigenic phenotypes are associated with p38 stress signaling pathway activation. (A) Western blot analysis showing protein expression and/or activation of Akt and MAPK proliferation signaling pathways as well as p38 and JNK stress signaling pathways in MCF7shMAGI1 compared to MCF7shLuc cells. Tubulin was used as a loading control. Western blot experiments have been repeated at least three times. Uncropped blots can be found in the Supplementary Information. (B) Quantification of the representative Western blot showing protein expression represented in panel A. Phosphorylated proteins were quantified as compared to their total protein counterparts to evaluate their activation. (C) Real-time qRT-PCR showing the expression of some p38 pathway target genes (ATF3, SOX2, SOX9 and GATA6) in MCF7shMAGI1 compared to the control MCF7shLuc cells and normalized with GAPDH expression. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; *p < 0.05; **p < 0.01; ***p < 0.001. (D) 2D cell growth assay of MCF7shMAGI1 treated, or not, with the p38 MAPK inhibitor LY2228820 (LY in the figure) compared to MCF7shLuc. Bars represent mean ± SD (n = 6 wells as replicates) of a representative experiment (n = 3). Unpaired two-tailed Student’s t-test; *p < 0.05. Note the increased percentage of drop for MCF7shMAGI1 cells at day 8. (E) Quantification of colony numbers of MCF7shMAGI1 cells non treated or treated with LY2228820 grown in anchorage independent conditions (soft agar assay) and represented as fold increase compared to MCF7shLuc cells non treated or treated with the p38 inhibitor (LY in the figure). Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; ***p < 0.001. (F) 2D cell growth assay (MTT) representing 2D cell growth of MCF7shMAGI1 and MCF7shLuc after transfection of p38asiRNA or CtlsiRNA. Bars represent mean ± SD (n = 10 wells as replicates) of a representative experiment (out of 3). Unpaired two-tailed Student's t-test; *p < 0.05. (G) Quantification of colony numbers of MCF7shMAGI1 and MCF7shLuc after transfection of p38a siRNA or Ctl siRNA, grown in anchorage independent conditions (soft agar assay) and represented as fold increase compared to MCF7shLuc cells transfected with Ctl siRNA. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; ***p < 0.001. (H) Western blot analysis showing protein levels of phosphorylated p38 and total p38 in MCF7shMAGI1 cells treated or non-treated (NT) with Blebbistatin (Blebb) or Y-27632. Both inhibitors were used at 10 µM during 2 h and Tubulin that was used as a loading control. Uncropped blots can be found in the Supplementary Information. (I) Quantification of the representative Western blot showing protein expression and represented in panel E. Phosphorylated p38 was quantified as compared to total p38 to evaluate its activation.
Strikingly, treating MCF7shMAGI1 cells with the Rho Kinase (ROCK) inhibitors Y-27632 or Blebbistatin that prevent Myosin-mediated F-actin tension, abolished p38 activation (Fig. 6H,I) and largely decreased elastic cell Young’s modulus (Supplementary Fig. S6C), showing that increased ROCK activity and actin strength represent critical events to activate p38 signaling. However, due to their cell toxicity, these ROCK inhibitors treatments could only be maintained for short periods, preventing us to assay formally, whether releasing actin tension in MAGI1 deficient cells could suppress their tumorigenicity as would be expected from their effects on p38.
MAGI1 interacts with the angiomotin family members AMOT and AMOTL2 at the junctions
In order to better understand how the loss of MAGI1 and the associated alterations in ROCK activity and in E-cadherin-based junctions could lead to p38 stress-kinase activation, we decided to focus our studies on the junctional MAGI1 molecular complexes. Several studies, including proteomic approaches, have described a physical association between MAGI1 and AMOT or AMOTL2, two members of the angiomotin family of apical scaffolds, acting both as junctions/actin linkers as well as negative regulators of YAP transcriptional activity35,41,42,43,55. We thus, investigated whether AMOTs could mediate the effects of MAGI1 to restrict tumorigenesis in breast luminal cells.
First, using co-immunoprecipitation experiments, we confirmed that overexpressed Flag-tagged MAGI1 or endogenous MAGI1 interacted with endogenous AMOT and AMOTL2 in MCF7 luminal BCa cells (Fig. 7A,B). Building on the availability of numerous constructs generated by the group of KL. Guan38, we have then used AMOT as a paradigm to study the interaction between MAGI1 and AMOTs through co-immunoprecipitation experiments between Flag-tagged MAGI1 and HA-tagged AMOT constructs. Using point mutations affecting key structural Proline residues in the first and second WW motifs of MAGI1, we showed that the second WW motif was required for the interaction with AMOT, since co-immunoprecipitation was abolished by a point mutation on Proline residue 390 (mutated to Alanine MAGI1 P390A; Fig. 7C,D). The first WW domain appeared dispensable as no change in binding was observed when mutating Proline 331 (MAGI1 P331A). Similar results were obtained performing co-immunoprecipitations on endogenous AMOTL2 (Fig. 7E), showing that as for AMOT, the second WW domain of MAGI1 was required for AMOTL2 binding, confirming earlier findings43. The N-terminus part of AMOT contains two PPXY motifs, which represent canonical WW domains interactors. Using HA-tagged AMOT point mutants, we then determined that the interaction between MAGI1 and AMOT occurred through the second PPXY motif (Fig. 7F,G) since co-immunoprecipitation was lost in AMOT Y287A (Tyrosine 287 mutated to Alanine; the mutation AMOT Y242A did not show any effect). Together these results confirm using precise point mutations that MAGI1 binds to AMOT and AMOTL2 through its second WW domain, and at the level of the conserved second PPXY motifs of the N-terminal domains of AMOT family scaffolds.
MAGI1 interacts with AMOT and AMOTL2. (A) Western blot of Flag-MAGI1 immunoprecipitates: Flag-MAGI1 transfections were performed in MCF7 cells and endogenous AMOT (upper panels) and AMOTL2 (lower panels) were revealed; Flag blotting was used as the immunoprecipitation control. Note that we could hardly detect Flag-MAGI1 by Western blot in the inputs, even though MAGI1 was well immunoprecipitated, due to sensitivity issues associated with single Flag tag in Flag-MAGI1 used in this typical experiment. Uncropped blots can be found in the Supplementary Information. (B) Western blot analysis of endogenous MAGI1 immunoprecipitates revealing the presence of endogenous AMOT (upper panels) and AMOTL2 (lower panels). MAGI1 blotting was used as the immunoprecipitation control (*shows non-specific bands with the anti-MAGI1 antibody). Uncropped blots can be found in the Supplementary Information. (C) Schematic of full length MAGI1 and of the point MAGI1 WW domain mutants used (P331A and P390A). (D) Western blot analysis of Flag-MAGI1 immunoprecipitates on protein extracts from MCF7 cells transfected with HA-AMOT and different Flag tagged MAGI1 constructs, showing that the interaction with AMOT occurred through the second WW domain of MAGI1 (interaction lost in P390A; red). Uncropped blots can be found in the Supplementary Information. (E) Western blot analysis of Flag-MAGI1 immunoprecipitates on protein extracts from MCF7 cells transfected with Flag-MAGI1 or Flag-MAGI1 P390A mutant, revealing the interaction of endogenous AMOTL2 with Flag-MAGI1 but not Flag-MAGI1 P390A (red). Uncropped blots can be found in the Supplementary Information. (F) Schematic of full-length AMOT and of the point AMOT PPXY mutants used (Y242A and Y287A). (G) Western-blot analysis of Flag-MAGI1 immunoprecipitates on protein extracts from MCF7 cells transfected with Flag-MAGI1 and different HA-AMOT constructs, showing that the interaction occurs through the second PPXY domain of AMOT (interaction lost in Y287A; red). Similar levels of interaction were obtained for HA-AMOT/Flag-MAGI1 and HA-AMOT Y242A/Flag-MAGI1 complexes (quantified using the ImageJ software). Uncropped blots can be found in the Supplementary Information. (H) Representative immunofluorescence images of endogenous staining for MAGI1 (red; top panels) co-localizing at the plasma membrane (white arrows) with AMOT (green; left) or AMOTL2 (green; right) in MCF7 cells. Note the non-specific nuclear staining for AMOT. Scale bar = 10 µm. (I) Western blot analysis after subcellular fractionation showing the relative amount of AMOTL2 protein in the cytoplasmic and membrane fractions in MCF7shMAGI1 compared to MCF7shLuc cells. Tubulin was used as a cytoplasmic control for the fractionation and for normalization. Uncropped blots can be found in the Supplementary Information.
Then, using immunofluorescence on MCF7 fixed cells, MAGI1, AMOT, and AMOTL2 localized in overlapping membrane domains corresponding to cellular junctions (Fig. 7H; the AMOT nuclear staining is non-specific). Even though, MAGI1 colocalized with AMOTL2 and AMOT (Fig. 7H, arrows), it was not required for their proper membrane localizations as they were unaffected in shMAGI1, suggesting that even though they physically interact once in proximity, they must reach their membrane localization independently (Fig. 7I). Nevertheless, even though MAGI1 was not required for AMOTL2 localization, it did however control AMOTL2 levels, and AMOTL2 accumulated 1.8 fold in MCF7shMAGI1 compared to controls as measured by western blots on whole protein extracts (Fig. 3B). These results suggest that MAGI1 regulates the stability and/or degradation of AMOTL2, but using classic drugging approaches, we were unable to identify the mechanisms involved. It is noteworthy here that AMOTL2 mRNA levels were slightly increased in MAGI1-deficient cells (Fig. 5D), suggesting that the accumulation of AMOTL2 protein could also be a consequence of increased transcription.
Together our results confirm the physical interaction between MAGI1 and AMOTs and show that MAGI1 restricts the total AMOTL2 protein levels, through a mechanism that remains to be established.
AMOTL2 mediates the increased tumorigenicity of MAGI1-deficient luminal BCa cells
We observed that in luminal BCa cells, the loss of MAGI1 triggered both (i) an accumulation of AJs material including its binding partner AMOTL2, and (ii) an activation of the p38 pathway driving an increase in tumorigenicity. We thus studied next whether these two different aspects were linked or independent consequences of MAGI1 loss. Generating double knockdown MCF7 cells for MAGI1 and AMOTL2, we could show that the increased 2D growth and anchorage-independent growth after MAGI1 loss were suppressed by the knockdown of AMOTL2 (Fig. 8A,B). Consistently, AMOTL2 knockdown abolished the accumulation of E-cadherin and the activation of the p38 stress kinase (phospho-p38 levels; Fig. 8C,D). These results identify AMOTL2 accumulation as a critical mediator in the p38 activation and tumorigenesis induced by impaired MAGI1.
AMOTL2 mediates the effects of MAGI1 on junctions and on p38 signaling. (A) MTT assay (OD 560 nm) representing 2D cell growth of MCF7shMAGI, MCF7shAMOTL2 and MCF7shMAGI1/shAMOTL2 cells as compared to MCF7shLuc. Bars represent mean ± SD (n = 10 wells as replicates) of a representative experiment (n = 3). Unpaired two-tailed Student's t-test; **p < 0.01; ***p < 0.001. (B) Quantification of colony numbers of MCF7 cells (shMAGI1, shAMOTL2 and shMAGI1/shAMOTL2) grown in anchorage independent conditions (soft agar assay) and represented as fold increase compared to MCF7shLuc cells. Data are presented as the means ± SD (n = 3). Unpaired two-tailed Student's t-test; *p < 0.05. Western blot analyses confirmed the 90% knockdown for MAGI1 in MCF7shMAGI1 and MCF7shMAGI1/shAMOTL2 and QPCR data confirmed the 50 to 60% knockdown for AMOTL2 in MCF7shAMOTL2 and MCF7shMAGI1/shAMOTL2 respectively (determined by RT-qPCR and/or Western Blot ; Data not shown). (C) Western blot analysis showing protein expression and/or phosphorylation (activation) of p38 and JNK stress signaling pathways as well as junctions’ components in MCF7shMAGI1 as compared to MCF7shLuc cells when AMOTL2siRNA was transfected. Tubulin was used as a loading control. Western blot experiments have been repeated at least three times. Uncropped blots can be found in the Supplementary Information. (D) Quantification of the representative Western blot showing protein expression and represented in panel C. Phosphorylated proteins were quantified as compared to their total protein counterparts to evaluate their activation and junctions’ proteins were quantified as compared to tubulin that was used as a loading control. Upper: Phosphorylated p38/Total p38 protein quantification and Lower: protein/Tubulin quantification. (E) Model for the role of MAGI1 during Luminal BCa. MAGI1 prevents the accumulation of junctional AMOTL2 and E-cadherin as well as ROCK activity thus releasing cellular stiffness. Increased AMOTL2 and ROCK then activate p38 stress signaling responsible for the increased tumorigenicity of MAGI1-deficient cells. Anti and Pro tumorigenic events are highlighted in green and red respectively.
Together, these results show that both AMOTL2 and ROCK activity are required for p38 activation in MAGI1 deficient luminal BCa cells. They support a model in which, the AMOTL2 accumulation in MAGI1 deficient cells governs an increase of E-cadherin-based junctions, which together with ROCK activity and cortical actin tension, activate the tumorigenic activity of the p38 signaling pathway (Fig. 8E).
Discussion
In this study, we explored the function of the junctional component MAGI1 in breast carcinogenesis and showed that the loss of MAGI1 in luminal BCa cell lines promoted tumorigenesis. At the cellular level, we showed that the loss of MAGI1 induced an accumulation of AJ components and increased cell compression behaviors. Importantly, we showed that the increased tumorigenicity of MAGI1 deficient luminal BCa cells was mediated by p38 stress signaling. Molecularly, we confirmed that MAGI1 interacted with the AMOT family of apical scaffolds and actin linkers, and showed that decreasing AMOTL2 levels, completely suppressed the effect of MAGI1 loss on the activation of p38 and on tumorigenicity. Finally, we provided evidence that inhibiting ROCK activity could alleviate the p38 activation, supporting a model in which MAGI1 suppresses luminal BCa by preventing both ROCK and AMOTL2-mediated Junction dysfunction, and subsequent p38 stress signaling.
The loss of MAGI1 led to an increase in tumorigenicity, and an accumulation of AJ material, in particular E-cadherin, reminiscent to the role we described previously for the unique MAGI homologue in Drosophila24. This correlation could appear counterintuitive as E-cadherin knockdown promotes invasion and migration in cultured cells56, and is critical during EMT and for aggressiveness (recently reviewed in57). However, in several cancers including some BCa, E-cadherin expression is maintained with a proposed tumor supporting role58,59 and recent mouse models of luminal and basal invasive ductal carcinomas have demonstrated that mammary cancer cells proliferation, resistance to apoptosis, and distant metastatic seeding potential, require E-cadherin60. These observations reconcile the high E-cadherin levels and increased tumorigenicity observed upon MAGI1 loss, and suggest that at least in specific phases of tumorigenesis (e.g. mass proliferation, colony formation), E-cadherin could play a positive role during BCa.
One striking feature of MAGI1 depleted cells is the protein accumulation of AMOTL2. Even though the exact mechanisms remain to be described, the stability of the related AMOT is controlled by the activity of the RNF146/Tankyrase pathway61,62, and by phosphorylation by the Hippo pathway core kinase LATS1 (reviewed in37). Importantly AMOTL2, a binding partner of MAGI1, is required for the increased tumorigenicity of MAGI1-depleted luminal BCa cells, showing that AMOTL2 is a critical mediator in this context. AMOTs are apical scaffolds playing important roles in cadherin-based junction regulations and its linkage to the actin cytoskeleton26,35,36. Previous reports have shown that AMOTL1 is overexpressed in invasive ductal carcinomas, leading to invasive behaviors. However, this effect of AMOTL1 appears specific to ER-negative subtypes63. AMOTL1 is not expressed in breast luminal lineages, in which AMOTL2 is the most abundant (twice more abundant than AMOT in MCF7 cells; our unpublished observation). We propose thus that in ER-positive luminal breast lineages, AMOTL2 overexpression could play similar roles as marker and mediator of tumorigenesis. Interestingly, a short isoform of AMOTL2 (AMOTL2 p60 in contrast to the long AMOTL2 p100) was reported overexpressed in in situ and invasive ductal carcinoma cells64. The AMOTL2 p60 isoform lacks the PPxY domain critical for its interaction with MAGI1, and it is possible this isoform is thus insensitive to MAGI1-dependent destabilization, even though this remains to be studied. Hypoxia and Fos signaling govern the specific expression of AMOTL2 p6064, and the accumulation of AMOTL2 in the context of MAGI1 mutation could thus represent an alternative mechanism for AMOTL2 expression, extending the relevance of AMOTL2 up-regulation during carcinogenesis.
AMOTL2 couples adhesion complexes to the actin cytoskeleton to allow F-actin tension and thus morphogenesis in endothelial cells (coupling with VE-cadherin26) and in epithelial cell during embryogenesis (E-cadherin35). Importantly, the specific cellular morphology defects in AMOTL2 knockdown could be mimicked by inhibiting ROCK kinase and Myosin phosphorylation showing that AMOTL2 and actin tension are linked35. Relaxing actin tension using two ROCK inhibitors (Y-27632 and Blebbistatin), or preventing AMOTL2 accumulation (siRNA), both robustly suppressed p38 activation, a key feature mediating tumorigenesis after MAGI1 loss. So how could AMOTL2 and ROCK interact to mediate the effects of MAGI1 loss? They could either (i) be two independent consequences of MAGI1 loss or (ii) they could be linked causally. Due to the toxicity of long term exposures to ROCK inhibitors, we were unable to study whether AMOTL2 stability could be a consequence of ROCK activity. Preliminary results would indicate that AMOTL2 does not control ROCK activity (data not shown), suggesting that ROCK might act upstream, or in parallel to AMOTL2, but more experiments are needed to better determine how MAGI1 controls ROCK activity, and how ROCK and AMOTL2 regulate p38 activity.
p38 signaling is activated by many upstream signals including a wide variety of environmental stresses such as UV, genotoxic agents, heat shock, or hyperosmotic conditions18. During osmotic shock, the cellular cortex is subjected to external pressure reminiscent to compressive forces, and we propose therefore that E-cadherin and AMOTL2 enrichment, together with increased ROCK activity and actin tension result in similar compressive forces that could represent a new stress signal activating p38. Indeed, amongst the genes regulated transcriptionally by p38, SOX9, a gene mostly responsive in osmotic shock49, was the most robustly activated upon MAGI1 knock-down. Amongst the four different p38 kinases (p38α, β, γ, δ), P38α is the most ubiquitously expressed. p38 signaling has been implicated either as a tumor suppressor or as an oncogene depending on cancer type and stage18,52,53,54. Indeed, the tumor suppressive and dormancy promoting role of p38 has been extensively documented, showing that the role of p38 in cancer is complex and multiple65,66,67. These differing and opposite roles are not uncommon for different cancer associated genes (e.g. Notch…) and may depend on context. In BCa, targeting p38δ resulted in a reduced rate of cell proliferation and enhanced cell-matrix adhesion68. More importantly, BCa mouse models, demonstrated the critical role of p38α (Mapk14) during the initiation, and proliferation of mammary tumors. The tumor promoting role of p38α involves a protective role against deleterious DNA damage in the mammary epithelial cancer cells50. We observed in MAGI1-deficient cells an increased proportion of cells in S-phase, indicating that these cells might have a slower S-phase. Slow S-phase is classically observed when cells have to overcome a high level of DNA damage. These results highlight the need for further studies to better understand the link between MAGI1 loss, p38 activation, DNA damage, and breast tumorigenesis.
Besides elevated p38 signaling, MAGI1-deficient cells exhibited low YAP nuclear activity, consistent with earlier reports showing that in epithelial cells with high E-cadherin and/or under compressive forces α-Catenin and NF2/Merlin act to exclude YAP from the nucleus10. Here we demonstrated the critical role of AMOTL2 downstream of MAGI1 to mediate E-cadherin accumulation. Since AMOTs have been shown to trap YAP in the cytoplasm38, further studies are thus required to better understand the interactions between MAGI1/AMOTL2 and α-Cat/NF2 in the control of YAP localization and activity. This low YAP signaling could appear surprising since the oncogenic role of nuclear YAP is well established69. However, the oncogenic role of YAP in luminal BCa remains debated. While elevated YAP/TAZ activity gene signatures have been reported to correlate with more aggressive BCa70,71,72, aggressive BCa are enriched in basal/triple negative sub-types. Furthermore, several studies report conflicting or no correlation between YAP staining levels and clinical outcomes in BCa patients (reviewed in69) suggesting that YAP/TAZ levels and nucleo/cytoplasmic distributions in luminal BCa could be re-examined.
Methods
Plasmids, mutant constructs and shRNA cloning
All expression plasmids were generated with the Gateway system (Invitrogen). The Entry vector pDONR223-MAGI1 was a gift from William Hahn (Addgene plasmid #23523)73 and it was used to introduce the different point mutations in MAGI1 by mutagenesis PCR using PfuTurbo (Agilent). All the Gateway destination vectors are listed in the supplementary methods. pEZY-EGFP-MAGI1 was constructed by Gateway recombination between the pDONR223-MAGI1 and the pEZY-EGFP destination vector that was a gift from Zu-Zhu Zhang (Addgene plasmid #18671)74. pcDNA3 HA-AMOT, pcDNA3 HA-AMOT Y242A, pcDNA3 HA-AMOT Y287A and pcDNA3 HA-AMOT Y242/287A were gifts from Kunliang Guan (Addgene plasmids #32821, #32823, #32824 and #32822 respectively)38.
shRNA directed against human MAGI1 and AMOTL2 were constructed and cloned in the pSIREN-RetroQ vector (TaKaRa) according to the manufacturer's conditions between BamHI and EcoRI cloning sites as previously described for other genes75. Targeted sequences are listed in the supplementary methods section.
Cell culture and cell transfection
MCF7, T47D and HCT116 human cell lines originated from the TumoroteK bank (SIRIC Montpellier Cancer) and have been authenticated by STR (Short-tandem repeat) profiling and certified mycoplasm-free. Details for culture are added in the supplementary methods.
Plasmid transfections were performed in MCF7 cells using Lipofectamine2000 reagent (Invitrogen) according to the manufacturer's directions. Cell extracts were assayed for protein expression 24–48 h post-transfection. Concerning AMOTL2siRNA and p38α siRNA transfection (Dharmacon siGENOME #M-01323200-0005 and #L-003512-00-0005 respectively), a final concentration of 100 nM was used using Lipofectamine2000 reagent as well.
Western blotting
Proteins issued from transfected MCF7, MCF7shRNA, T47DshRNA or HCT116shRNA cell lines were extracted, analyzed by SDS-PAGE. For dilutions and antibodies references, please refer to the supplementary methods.
RNA extraction, reverse transcription and real time RT-qPCR
RNAs were extracted from cells using RNeasy plus kit (Qiagen) and cDNAs were prepared starting from 1 µg of RNAs using the Superscript III reverse transcriptase (Invitrogen) and following manufacturer's directions. Real time quantitative PCR was then performed on cDNAs using SYBR Green I Mastermix (Roche) according to the manufacturer's conditions on a Light Cycler 480 device (Roche). All primers used are listed in the supplementary methods.
Patients and TMA construction
Breast cancer samples were retrospectively selected from the Institut régional du Cancer de Montpellier (ICM) pathology database using the following inclusion criteria: chemotherapy-naïve at the time of surgery and estrogen (ER), progesterone (PR) receptor and HER2 status available. TMA construction are detailed in the supplementary methods section. Tumor samples were collected following French laws and declared to the French Ministry of Higher Education and Research (Biobanque BB-0033-00059; declaration number DC-2008-695). Informed consent was obtained from all participants (adult breast cancer patients). All patients were informed about the use of their tissue samples for biological research. All experimental protocols were approved by the ethics and licensing committee CORT (Comité de Recherche Translationnelle de l’Institut regional du Cancer de Montpellier (ICM)).
Immunohistochemistry
Four µm thin sections of the TMA were mounted on Flex microscope slides (Dako) and allowed to dry overnight at room temperature before immunohistochemistry (IHC) processing. For further details, please refer to the supplementary methods.
Immunofluorescence
Cells seeded on glass coverslips were fixed 10 min in paraformaldehyde (4%), before being permeabilized in PBS/0.1% TritonX-100 for 10 min. After blocking in PBS/0.5% BSA, cells were incubated with primary antibodies overnight at 4C. Antibodies used are listed in the supplementary methods. Secondary Alexa Fluor Antibodies (1/200; Invitrogen) were used as described previously24 for 1 h at room temperature before mounting the coverslips with Vectashield (Vector Laboratories #H-1200) and imaging on Zeiss Apotome or Leica Thunder microscopes.
Atomic Force Microscopy (AFM) measurements
Cells were plated on glass 35 mm fluorodish at intermediate (1/3) dilution density, grown overnight, and placed in 2% serum medium before AFM experiments. Cells grew as clusters of cells. AFM Force Spectroscopy (AFM-FS) experiments76 on living cell clusters were performed on a JPK Nanowizard 4 microscope equipped with a CellHesion stage (JPK). The AFM is mounted on an inverted Axiovert 200 M microscope system (Carl Zeiss) equipped with a 100x (1.5NA, plan-Apochromat) objective lens (Carl Zeiss). We employed a heating module (JPK) placed at the sample level to maintain cells at the physiological temperature of 37 °C during measurements. We used CP-CONT-SIO-D cantilevers with 10 µm colloidal beads as tip (NanoAndMore). Cantilever stiffness and optical lever sensitivities were both calibrated in liquid environment using the Contact-Free-Method provided by JPK AFM, and based on a mix of a thermal and Sader77 calibration methods. For calibration details, please refer to the supplementary methods. At least 5 force maps were acquired in each experiment on each cell category and 3 separated experiments were performed. Analyses were carried out using the JPK AFM data processing software. The elastic Young’s modulus (E; Pa) was evaluated by fitting each force versus tip-cell distance curve with the Hertz contact model for indenting an infinite isotropic elastic half space with a solid sphere as described in78.
Mammosphere culture
MCF7shLuc and MCF7shMAGI1 cell suspensions were obtained by trypsinization and then filtered using 30 µm cell strainer (Miltenyi). Filtered cells were immediately seeded in 96 well plates coated with 1% Poly (2-hydroxyethyl methacrylate; Sigma Aldrich) at a density of 200 or 50 cells per well. Cells were grown in MEBM basal medium supplemented with B27 without vitamin A (50X), 20 ng/mL EGF, 20 ng/mL bFGF and 4 μg/mL heparin. Cells were incubated at 37 °C with 5% CO2 during 5 days. For mammosphere acquisition and analysis, please refer to the supplementary methods.
Immunoprecipitation and co-immunoprecipitation
Protein extracts were prepared in lysis buffer (NaCl 150 mM, Tris pH 7.4 10 mM, EDTA 1 mM, Triton X-100 1%, NP-40 0.5%, cOmplete, EDTA-free protease inhibitors (Roche #11873580001) for 30 min on ice before centrifugation. Immunoprecipitations were performed overnight at 4 °C on a rocking wheel using either mouse anti-MAGI1 antibody for endogenous immunoprecipitations or anti-HA antibody and EZview Red anti-FLAG M2 affinity gel (Sigma-Aldrich #F1804) for co-immunoprecipitations. Protein G sepharose was then added to the MAGI1- or HA- immunoprecipitates for 1 h at 4 °C before extensive washes. Concerning the FLAG immunoprecipitation, washes were performed followed by protein elution by competition with 3XFLAG peptide (150 ng/µL final concentration) during 1 h at 4 °C. The different immunoprecipitates were then submitted to Western blotting for detection of protein complexes.
Subcellular fractionation: isolation of cytoplasmic, nuclear and membrane protein fractions
Subcellular fractionation of cultured human cell lines was performed as previously described on-line (http://www.bio-protocol.org/e754).
Cell growth assay
2D cell growth assay was analyzed using MTT (Thiazolyl Blue Tetrazolium Bromide (Sigma Aldrich #M2128)). Briefly, cells seeded in 96 well plates were stopped from day 0 to day 7 by incubation with MTT solution (0.5 mg/mL final concentration) during 4 h at 37 °C. Before reading OD at 570 nm, cells were incubated with DMSO to solubilize the formazan crystals formed in the presence of MTT.
3D spheroid formation and measures
10,000 cells were seeded in Ultra-Low Attachment 96 well plates (Costar) cultured in their regular medium supplemented with 15% FCS. After 3 days, all cells formed 3D spheroids that can be further analyzed by perimeter and circularity measurements. Pictures of spheroids were taken 3 days after seeding cells to assay the ability of cell lines to form spheroids and both the perimeters and the circularity of spheroids were calculated. Circularity was calculated (4px(area/(perimeter)2) where 1 is considered as a perfect circle.
p38 MAPK inhibitor treatment
The different MCF7shRNA cell lines were plated at 3 × 106 cells per 60 mm plates. The day after, 10 µM of Ralimetinib (LY2228020; Selleckchem #S1494) was added and 24 h later cells were collected and counted to be plated for the different experiments (Ralimetinib treatment is maintained throughout the experiments). For Soft agar assay, 10,000 cells were plated in 12-well plates before proceeding to the assay detailed below. The same experiments were conducted with another p38 inhibitor using 10 µM of Pexmetinib (ARRY-614; Selleckchem #S7799). SRB proliferation assay was performed as detailed in the supplementary methods.
ROCK inhibitors treatment
The treatments of MCF7shRNA cell lines with either Blebbistatin (Selleckchem #7099) or Y-27632 (Selleckchem #S1049) were both performed at a final concentration of 10 µM and cells were treated during 2 h before being lysed for Western blot analyses.
Soft agar assay
To assess anchorage independent growth, soft agar assays were performed. A first layer of 0.8% Noble agar (Sigma Aldrich #A5431) was prepared in 12-well plates and left at 4C during 1 h for the agar to solidify gently. The plate was kept at 37C before the cell-containing agar layer was prepared. 5,000 or 10,000 cells were imbedded in the second layer containing 0.4% Noble agar on top of the first layer. When the second layer was solidified, fresh medium was added on top of the cells. MCF7 cell lines were cultured during 14 to 21 days before Crystal violet coloration (0.01% final concentration; Sigma Aldrich #C0775) or MTT assay (1 mg/mL final concentration).
Primary tumor growth assay
MCF7shRNA and HCT116shRNA cell lines were injected subcutaneously in six-week-old female athymic Nude-Foxn1 mice (Envigo). Both flanks of each mouse were injected with cells mixed with Matrigel (106 cells injected for HCT116; 10 × 106 cells injected for MCF7). Mice injected with MCF7 cells were subjected to neck dropping with 50 µl of β-estradiol (1.5 mg/mL diluted in ethanol) to stimulate MCF7 cell growth. Animal procedures were performed according to protocols approved by the ethics and licensing committee of the Direction Générale de la Recherche et de l’Innovation (Ministère de l’Enseignement Supérieur de la Recherche et de l’Innovation), and this study was carried out in compliance with the ARRIVE guidelines.
References
Holliday, D. L. & Speirs, V. Choosing the right cell line for breast cancer research. Breast Cancer Res. BCR 13, 215 (2011).
Guedj, M. et al. A refined molecular taxonomy of breast cancer. Oncogene 31, 1196–1206 (2012).
Knust, E. & Bossinger, O. Composition and formation of intercellular junctions in epithelial cells. Science 298, 1955–1959 (2002).
Coopman, P. & Djiane, A. Adherens Junction and E-Cadherin complex regulation by epithelial polarity. Cell. Mol. Life Sci. CMLS 73, 3535–3553 (2016).
Jeanes, A., Gottardi, C. J. & Yap, A. S. Cadherins and cancer: how does cadherin dysfunction promote tumor progression?. Oncogene 27, 6920–6929 (2008).
Balda, M. S. & Matter, K. Tight junctions as regulators of tissue remodelling. Curr. Opin. Cell Biol. 42, 94–101 (2016).
Charras, G. & Yap, A. S. Tensile forces and mechanotransduction at cell–cell junctions. Curr. Biol. CB 28, R445–R457 (2018).
Pinheiro, D. & Bellaïche, Y. Mechanical force-driven adherens junction remodeling and epithelial dynamics. Dev. Cell 47, 3–19 (2018).
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Cell adhesion. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015).
Furukawa, K. T., Yamashita, K., Sakurai, N. & Ohno, S. The epithelial circumferential actin belt regulates YAP/TAZ through nucleocytoplasmic shuttling of merlin. Cell Rep. 20, 1435–1447 (2017).
Hirata, H., Samsonov, M. & Sokabe, M. Actomyosin contractility provokes contact inhibition in E-cadherin-ligated keratinocytes. Sci. Rep. 7, 46326 (2017).
Kim, N.-G., Koh, E., Chen, X. & Gumbiner, B. M. E-cadherin mediates contact inhibition of proliferation through Hippo signaling-pathway components. Proc. Natl. Acad. Sci. USA 108, 11930–11935 (2011).
Schlegelmilch, K. et al. Yap1 acts downstream of α-catenin to control epidermal proliferation. Cell 144, 782–795 (2011).
Silvis, M. R. et al. α-catenin is a tumor suppressor that controls cell accumulation by regulating the localization and activity of the transcriptional coactivator Yap1. Sci. Signal. 4, ra33 (2011).
Aoki, K. et al. Propagating wave of ERK activation orients collective cell migration. Dev. Cell 43, 305-317.e5 (2017).
Hall, E. T., Hoesing, E., Sinkovics, E. & Verheyen, E. M. Actomyosin contractility modulates Wnt signaling through adherens junction stability. Mol. Biol. Cell 30, 411–426 (2019).
Pereira, A. M., Tudor, C., Kanger, J. S., Subramaniam, V. & Martin-Blanco, E. Integrin-dependent activation of the JNK signaling pathway by mechanical stress. PLoS ONE 6, e26182 (2011).
Wagner, E. F. & Nebreda, A. R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat. Rev. Cancer 9, 537–549 (2009).
Beavon, I. R. The E-cadherin-catenin complex in tumour metastasis: structure, function and regulation. Eur. J. Cancer Oxf. Engl. 1990(36), 1607–1620 (2000).
Bosch-Fortea, M. & Martín-Belmonte, F. Mechanosensitive adhesion complexes in epithelial architecture and cancer onset. Curr. Opin. Cell Biol. 50, 42–49 (2018).
Laura, R. P., Ross, S., Koeppen, H. & Lasky, L. A. MAGI-1: a widely expressed, alternatively spliced tight junction protein. Exp. Cell Res. 275, 155–170 (2002).
Hirabayashi, S. et al. JAM4, a junctional cell adhesion molecule interacting with a tight junction protein, MAGI-1. Mol. Cell. Biol. 23, 4267–4282 (2003).
Padash Barmchi, M., Samarasekera, G., Gilbert, M., Auld, V. J. & Zhang, B. Magi is associated with the Par complex and functions antagonistically with Bazooka to regulate the apical polarity complex. PLoS ONE 11, e0153259 (2016).
Zaessinger, S., Zhou, Y., Bray, S. J., Tapon, N. & Djiane, A. Drosophila MAGI interacts with RASSF8 to regulate E-Cadherin-based adherens junctions in the developing eye. Dev. Camb. Engl. 142, 1102–1112 (2015).
Lynch, A. M. et al. A genome-wide functional screen shows MAGI-1 is an L1CAM-dependent stabilizer of apical junctions in C. elegans. Curr. Biol. CB 22, 1891–1899 (2012).
Hultin, S. et al. AmotL2 links VE-cadherin to contractile actin fibres necessary for aortic lumen expansion. Nat. Commun. 5, 3743 (2014).
Feng, X., Jia, S., Martin, T. A. & Jiang, W. G. Regulation and involvement in cancer and pathological conditions of MAGI1, a tight junction protein. Anticancer Res. 34, 3251–3256 (2014).
Zhang, G., Liu, T. & Wang, Z. Downregulation of MAGI1 associates with poor prognosis of hepatocellular carcinoma. J. Investig. Surg. Off. J. Acad. Surg. Res. 25, 93–99 (2012).
Kozakai, T. et al. MAGI-1 expression is decreased in several types of human T-cell leukemia cell lines, including adult T-cell leukemia. Int. J. Hematol. 107, 337–344 (2018).
Zhang, G. & Wang, Z. MAGI1 inhibits cancer cell migration and invasion of hepatocellular carcinoma via regulating PTEN. Zhong Nan Da Xue Xue Bao Yi Xue Ban 36, 381–385 (2011).
Zmajkovicova, K. et al. MEK1 is required for PTEN membrane recruitment, AKT regulation, and the maintenance of peripheral tolerance. Mol. Cell 50, 43–55 (2013).
Kranjec, C., Massimi, P. & Banks, L. Restoration of MAGI-1 expression in human papillomavirus-positive tumor cells induces cell growth arrest and apoptosis. J. Virol. 88, 7155–7169 (2014).
Makokha, G. N. et al. Human T-cell leukemia virus type 1 Tax protein interacts with and mislocalizes the PDZ domain protein MAGI-1. Cancer Sci. 104, 313–320 (2013).
Zaric, J. et al. Identification of MAGI1 as a tumor-suppressor protein induced by cyclooxygenase-2 inhibitors in colorectal cancer cells. Oncogene 31, 48–59 (2012).
Hildebrand, S. et al. The E-cadherin/AmotL2 complex organizes actin filaments required for epithelial hexagonal packing and blastocyst hatching. Sci. Rep. 7, 9540 (2017).
Hultin, S. et al. AmotL2 integrates polarity and junctional cues to modulate cell shape. Sci. Rep. 7, 7548 (2017).
Maugeri-Saccà, M. & De Maria, R. The Hippo pathway in normal development and cancer. Pharmacol. Ther. 186, 60–72 (2018).
Zhao, B. et al. Angiomotin is a novel Hippo pathway component that inhibits YAP oncoprotein. Genes Dev. 25, 51–63 (2011).
Yu, F.-X. & Guan, K.-L. The Hippo pathway: regulators and regulations. Genes Dev. 27, 355–371 (2013).
Chan, S. W. et al. Hippo pathway-independent restriction of TAZ and YAP by angiomotin. J. Biol. Chem. 286, 7018–7026 (2011).
Wang, W. et al. Defining the protein-protein interaction network of the human hippo pathway. Mol. Cell. Proteomics MCP 13, 119–131 (2014).
Bratt, A. et al. Angiomotin regulates endothelial cell–cell junctions and cell motility. J. Biol. Chem. 280, 34859–34869 (2005).
Patrie, K. M. Identification and characterization of a novel tight junction-associated family of proteins that interacts with a WW domain of MAGI-1. Biochim. Biophys. Acta 1745, 131–144 (2005).
Györffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. Treat. 123, 725–731 (2010).
Alday-Parejo, B. et al. MAGI1, a new potential tumor suppressor gene in estrogen receptor positive breast cancer. Cancers 12, 223 (2020).
Nusse, R. & Clevers, H. Wnt/β-Catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Furth, N. & Aylon, Y. The LATS1 and LATS2 tumor suppressors: beyond the Hippo pathway. Cell Death Differ. 24, 1488–1501 (2017).
Misra, J. R. & Irvine, K. D. The Hippo signaling network and its biological functions. Annu. Rev. Genet. 52, 65–87 (2018).
Whitmarsh, A. J. A central role for p38 MAPK in the early transcriptional response to stress. BMC Biol. 8, 47 (2010).
Cánovas, B. et al. Targeting p38α increases DNA damage, chromosome instability, and the anti-tumoral response to taxanes in breast cancer cells. Cancer Cell 33, 1094-1110.e8 (2018).
Gupta, J. & Nebreda, A. R. Roles of p38α mitogen-activated protein kinase in mouse models of inflammatory diseases and cancer. FEBS J. 282, 1841–1857 (2015).
Gupta, J. et al. Dual function of p38α MAPK in colon cancer: suppression of colitis-associated tumor initiation but requirement for cancer cell survival. Cancer Cell 25, 484–500 (2014).
Hardwick, J. C., van den Brink, G. R., Offerhaus, G. J., van Deventer, S. J. & Peppelenbosch, M. P. NF-kappaB, p38 MAPK and JNK are highly expressed and active in the stroma of human colonic adenomatous polyps. Oncogene 20, 819–827 (2001).
Maik-Rachline, G., Zehorai, E., Hanoch, T., Blenis, J. & Seger, R. The nuclear translocation of the kinases p38 and JNK promotes inflammation-induced cancer. Sci. Signal. 11, eaao3428 (2018).
Couzens, A. L. et al. Protein interaction network of the mammalian Hippo pathway reveals mechanisms of kinase-phosphatase interactions. Sci. Signal. 6, rs15 (2013).
Frixen, U. H. et al. E-cadherin-mediated cell-cell adhesion prevents invasiveness of human carcinoma cells. J. Cell Biol. 113, 173–185 (1991).
Yang, J. et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-020-0237-9 (2020).
Kleer, C. G., van Golen, K. L., Braun, T. & Merajver, S. D. Persistent E-cadherin expression in inflammatory breast cancer. Mod. Pathol. Off. J. U. S. Can. Acad. Pathol. Inc 14, 458–464 (2001).
Rodriguez, F. J., Lewis-Tuffin, L. J. & Anastasiadis, P. Z. E-cadherin’s dark side: possible role in tumor progression. Biochim. Biophys. Acta 1826, 23–31 (2012).
Padmanaban, V. et al. E-cadherin is required for metastasis in multiple models of breast cancer. Nature 573, 439–444 (2019).
Campbell, C. I. et al. The RNF146 and tankyrase pathway maintains the junctional Crumbs complex through regulation of angiomotin. J. Cell Sci. 129, 3396–3411 (2016).
Wang, W. et al. Tankyrase inhibitors target YAP by stabilizing angiomotin family proteins. Cell Rep. 13, 524–532 (2015).
Couderc, C. et al. AMOTL1 promotes breast cancer progression and is antagonized by merlin. Neoplasia N. Y. N 18, 10–24 (2016).
Mojallal, M. et al. AmotL2 disrupts apical-basal cell polarity and promotes tumour invasion. Nat. Commun. 5, 4557 (2014).
Jahanban-Esfahlan, R. et al. Tumor cell dormancy: threat or opportunity in the fight against cancer. Cancers 11, 1207 (2019).
Ranganathan, A. C., Adam, A. P. & Aguirre-Ghiso, J. A. Opposing roles of mitogenic and stress signaling pathways in the induction of cancer dormancy. Cell Cycle Georget. Tex 5, 1799–1807 (2006).
Dolado, I. et al. p38alpha MAP kinase as a sensor of reactive oxygen species in tumorigenesis. Cancer Cell 11, 191–205 (2007).
Wada, M. et al. P38 delta MAPK promotes breast cancer progression and lung metastasis by enhancing cell proliferation and cell detachment. Oncogene 36, 6649–6657 (2017).
Zanconato, F., Cordenonsi, M. & Piccolo, S. YAP/TAZ at the roots of cancer. Cancer Cell 29, 783–803 (2016).
Bartucci, M. et al. TAZ is required for metastatic activity and chemoresistance of breast cancer stem cells. Oncogene 34, 681–690 (2015).
Cordenonsi, M. et al. The Hippo transducer TAZ confers cancer stem cell-related traits on breast cancer cells. Cell 147, 759–772 (2011).
Di Agostino, S. et al. YAP enhances the pro-proliferative transcriptional activity of mutant p53 proteins. EMBO Rep. 17, 188–201 (2016).
Johannessen, C. M. et al. COT drives resistance to RAF inhibition through MAP kinase pathway reactivation. Nature 468, 968–972 (2010).
Guo, F., Chiang, M.-Y., Wang, Y. & Zhang, Y.-Z. An in vitro recombination method to convert restriction- and ligation-independent expression vectors. Biotechnol. J. 3, 370–377 (2008).
Marzi, L. et al. FOXO3a and the MAPK p38 are activated by cetuximab to induce cell death and inhibit cell proliferation and their expression predicts cetuximab efficacy in colorectal cancer. Br. J. Cancer 115, 1223–1233 (2016).
Saavedra, V. O., Fernandes, T. F. D., Milhiet, P.-E. & Costa, L. Compression, rupture, and puncture of model membranes at the molecular scale. Langmuir ACS J. Surf. Colloids 36, 5709–5716 (2020).
Fernandes, T. F. D., Saavedra, O., Margeat, E., Milhiet, P.-E. & Costa, L. Synchronous, crosstalk-free correlative AFM and confocal microscopies/spectroscopies. Sci. Rep. 10, 7098 (2020).
Cartagena-Rivera, A. X., Van Itallie, C. M., Anderson, J. M. & Chadwick, R. S. Apical surface supracellular mechanical properties in polarized epithelium using noninvasive acoustic force spectroscopy. Nat. Commun. 8, 1030 (2017).
Acknowledgements
We acknowledge the contribution of G. Froment, D. Nègre and C. Costa from lentivectors' production facility of SFR Biosciences (UMS3444/CNRS, US8/Inserm, ENS Lyon, UCBL, France). We thank the ABIC (IRCM), MRI (IRCM), MGC (IGMM) platforms and M. Larroque from the URT (IRCM, Montpellier, France). We thank V. Chambon and T. Soirat for technical help with the mammosphere formation assay and PE Milhiet (CBS, Montpellier, France) for helpful discussion and AFM set up. All plasmids obtained from Addgene are detailed in the Material and Methods section and all references have been included. We also thank Drs C. Gongora, L. Holmgren, N. Tapon, and B. Zhao for discussion, tools and advice. The CBS is a member of the France-BioImaging (FBI), national infrastructures supported by the French National Research Agency ANR-10-INBS-04-01. This work was supported by grants from Fondation ARC and Ligue Régionale Contre le Cancer (34) to LHM and AD. DK is supported by Ligue Nationale Contre le Cancer. VS and EB were supported by Fondation de France. MM was supported by Fondation ARC.
Author information
Authors and Affiliations
Contributions
D.K., E.B.M., M.M., V.S., Y.B.S., L.C., E.F., F.B.M., B.O., C.B., and L.H.M. performed experiments. L.H.M. and A.D. designed the experiments. D.K., E.B.M., M.M., J.C., B.O., L.H.M., and A.D. analyzed the data. D.K., P.L., C.G., J.C., C.T., A.M., L.H.M., and A.D. interpreted the data. L.H.M. and A.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kantar, D., Mur, E.B., Mancini, M. et al. MAGI1 inhibits the AMOTL2/p38 stress pathway and prevents luminal breast tumorigenesis. Sci Rep 11, 5752 (2021). https://doi.org/10.1038/s41598-021-85056-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-85056-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.