Abstract
Gut microbial dysbiosis has been shown to be an instrumental factor in severe acute malnutrition (SAM) and particularly, the absence of Methanobrevibacter smithii, a key player in energy harvest. Nevertheless, it remains unknown whether this absence reflects an immaturity or a loss of the microbiota. In order to assess that, we performed a case–control study in Mali using a propensity score weighting approach. The presence of M. smithii was tested using quantitative PCR on faeces collected from SAM children at inclusion and at discharge when possible or at day 15 for controls. M. smithii was highly significantly associated with the absence of SAM, detected in 40.9% controls but only in 4.2% cases (p < 0.0001). The predictive positive value for detection of M. smithii gradually increased with age in controls while decreasing in cases. Among children providing two samples with a negative first sample, no SAM children became positive, while this proportion was 2/4 in controls (p = 0.0015). This data suggests that gut dysbiosis in SAM is not an immaturity but rather features a loss of M. smithii. The addition of M. smithii as a probiotic may thus represent an important addition to therapeutic approaches to restore gut symbiosis.
Similar content being viewed by others
Introduction
Methanogenic archaea play a critical role in host-microbiota mutualism1 by removing fermentative dihydrogen (H2), which is a central metabolite in overall organic matter degradation2. This H2 removal allows for a more complete oxidation of substrates, thereby improving energy harvest and the production of key molecules for the host, such as butyrate3 and ATP2,4. For instance, accumulation of H2 inhibits bacterial NADH dehydrogenases, thus reducing the yield of ATP5,6. The very low H2 utilization threshold of M. smithii (10 Pa6) compared to that of acetogens makes it the most efficient gut microbe for depleting H2 from the gut environment7. Moreover, M. smithii can alter the specificity and efficiency of digestion of some glycans in animal models8,9. These observations suggest a critical clinical relevance of this archaeon for human health.
The critical role of M. smithii in energy harvest and glycan digestion regulation has been shown in vitro and in vivo8,9. Paradoxically, these earlier publications have neglected the clinical relevance of M. smithii in human health, weight regulation and severe acute malnutrition10,11,12,13,14. Archaeal methanogens are poorly detected by metagenomics methods. Indeed, specific approaches had to be devised because metagenomics based on the amplification of hypervariable regions of the 16S ribosomal RNA gene was not efficient15,16. In a healthy adult European population, we showed that 100% of individuals harbour M. smithii in their stool samples, while the second known human gut methanogen, Methanosphaera stadtmanae, was rarely found17. The ubiquity of M. smithii in healthy adults reinforces the idea that this methanogen plays a crucial role in gut microbiota physiology.
Because M. smithii has been associated with weight gain and adiposity in animals, it has been speculated that M. smithii may promote obesity and that its eradication could treat obesity8,9. However, we found that M. smithii was associated with normal weight and that obesity was associated with M. smithii depletion18,19. This has been confirmed by other teams20,21. The apparent contradiction between the experimental8,9 and clinical18,19 results disappears when considering that the presence of M. smithii could be associated with normal weight and adiposity and that the relationship between M. smithii and body mass index is an inverted U curve. M. smithii has also been shown to improve the production of acetate9 as well as butyrate, a key molecule for human health3, through microbe-microbe interactions22. All these unique properties make M. smithii one of the best candidates as a marker of healthy digestion and nutritional status.
Severe acute malnutrition is a major public health problem affecting nearly 20 million children under five and causing up to 1 million deaths annually in low- or middle-income countries in Africa and South Asia22. It is a severe disease primarily related to inadequate diet. Micronutrients as well as protein-energy deficiencies have been linked to severe acute malnutrition23,24,25. However, cases refractory to a therapeutic diet26 and the fact that antibiotics improve mortality rates26,27 suggest other instrumental factors, such as gut microbiota dysbiosis. A delay in maturation of the digestive microbiota has been reported11,13, suggesting a quantitative developmental abnormality or immaturity. However, this immaturity could be a consequence and not the main feature of gut dysbiosis. A preliminary study conducted using specific quantitative PCR has shown that no malnourished children were positive for M. smithii compared to 75% of healthy children28. This observation suggests an absence or a loss of M. smithii which could contradict the “immaturity hypothesis”. This prompted us to launch a clinical investigation of the key role of M. smithii in severe acute malnutrition.
Accordingly, we performed a large case–control study to clarify the association between M. smithii and severe acute malnutrition. We particularly investigated whether the findings were consistent with immaturity and whether M. smithii remains undetectable after therapeutic renutrition. This hypothesis would suggest that oral intake as a probiotic and/or in organic dairy29 of this cultivable archaeon, previously isolated from human milk and colostrum21, may be a useful addition in the treatment of severe acute malnutrition.
Results
Participant characteristics
A total of 180 malnourished children were screened. Of these, 16 could not be included because afflicted with moderate acute malnutrition. A total of 164 of them were eligible for study, but only 143 severely malnourished children were ultimately included, as 21 were excluded because of an unavailable or insufficient number of samples provided (Fig. 1a). Of 209 healthy children screened, 61 were excluded outright because they did not meet the adequate anthropometric criteria. Of the 148 eligible controls with adequate anthropometric data, 13 were excluded for the presence of clinical symptoms or diseases such as gastroenteritis, malaria, rhinitis/rhino bronchitis and chickenpox. We excluded 25 others for non-available stool samples. We finally included 110 control children (Fig. 1b).
Study flow-chart. (a) Case selection, malnourished children are selected based on the WHO severity criterion and before any treatment; the presence of any symptoms or infection was not an exclusion criterion, (b) Control selection, healthy children are selected based on WHO standard without symptoms and antibiotics into the last 15 days.
Our study population comprised 110 healthy controls against 143 severely malnourished infants. Among malnourished children, 129 (90.2%) suffered from marasmus, and the remaining 14 children, less than 10%, had oedema, of whom nine (6.3%) had kwashiorkor and five (3.5%) had marasmic-kwashiorkor (Table 1, Table S1). Malnourished cases and controls were not different regarding age overall (Table S1). Children with marasmus predominantly ranged from 6 to 12 months, while those with kwashiorkor and marasmic-kwashiorkor were older.
Controls were asymptomatic by definition and without any known affliction. Fever was more frequent in cases (11.9% (17/143) vs 0.9% (1/110) in controls, p < 0.001, Table S1). Diarrhoea was detected in 6.3% (7/111) of malnourished children, all marasmus cases. Emesis was recorded in only 1% (1/111) of malnourished children. Gastroenteritis, which symptoms include diarrhoea, emesis or both, was diagnosed in 29.7% (33/111) of malnourished children (Table S1).
Methanobrevibacter smithii is lost in severe acute malnutrition
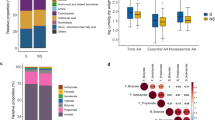
Key species such as M. smithii, Escherichia coli, Faecalibacterium prausnitzii and Staphylococcus aureus were quantified in severely malnourished children as well as healthy children. The concentration of M. smithii was significantly lower in malnourished children as shown in Fig. 2 (0.18 log10 DNA copies/ml, SD 0.89 vs 1.99 log10 DNA copies/ml, SD 2.56 in healthy children, Mann–Whitney test, p < 0.001). Conversely, the ubiquitous commensal E. coli was found in 99.6% (252/253) of the samples at similar mean concentrations in malnourished children and in healthy children (7.82 log10 DNA copies/ml, SD 1.39 vs 7.99 log10 DNA copies/ml, SD 1.42, Mann–Whitney test, p = 0.19) similarly as S. aureus (0.83 log10 DNA copies/ml, SD 1.75 in children with severe acute malnutrition vs 1.30 log10 DNA copies/ml, SD 2.34 in healthy children, Mann–Whitney test, p = 0.12). F. prausnitzii was also found in lower quantities in malnourished children (2.56 log10 DNA copies/ml, SD 2.35 vs 4.20 log10 DNA copies/ml, SD 2.93 in healthy children, Mann–Whitney test, p < 0.001).
The prevalence of M. smithii was dramatically lower in children with severe acute malnutrition compared to controls, 6/143 (4.2%) vs 45/110 (40.9%), respectively, p < 0.001. Strikingly, all positive malnourished children were marasmic (6/129), whereas none (0/14) of the malnourished children with oedema (isolated kwashiorkor or marasmic-kwashiorkor) were positive (Table 2). In order to assess the strength of our results and exclude a possible role of diarrhoea in the reduced prevalence of M. smithii in severely malnourished children, children with diarrhoea, emesis or gastroenteritis were excluded. The prevalence in M. smithii was still dramatically lower in children with severe acute malnutrition compared to controls, 4/89 (4.9%) vs 45/126 (35.7%), p < 0.001 (Table 3).
Detection of M. smithii increased with age only in controls, while prevalence decreased with age among malnourished children (Figs. 3 and 4). The prevalence rose to 90% in controls but dropped to 0% in severely malnourished children at 24 months (Fig. 3). The predictive positive value of detection of the M. smithii curve gradually increased to reach its maximum (all three children aged more than 55 months were positive) before 60 months of age in the healthy controls, unlike that of the malnourished children, which decreased abruptly and was nil after 15 months of age (Fig. 4). Controlling for age and gender, we found that the detection of M. smithii was associated with the absence of severe acute malnutrition (OR = 0.06, 95% confidence interval [0.02–0.15], p = 1·6 *10–9). We therefore performed a linear regression (Fig. 5) that showed that the concentration of M. smithii DNA increased with age only in controls (slope was positive (0.088) and was significantly different from zero in controls (p < 0.001) but negative (-0.0085) in severe acute malnourished children and not different from zero (p = 0.33)), and the difference between the two slopes was significant (p < 0 0.001).
Positive predictive value of Methanobrevibacter smithii in healthy and malnourished children according to age. Probability of detection of M. smithii relating to age is shown in healthy controls (CTL) and severely malnourished children (SAM); spike bars represent all the individuals from whom M. smithii has been detected in each group in the age range from 0 to 59 months. Green spike bars represent healthy controls children in top with the associated predictive value below. Red spike bars represent cases with the associated predictive value below.
DNA concentration of Methanobrevibacter smithii in healthy and severely malnourished children according to age. Linear regression model representing DNA concentrations (log10 DNA copies on the Y axis) according to the age of the subject (in months on the X axis). Regression lines are represented separately for each group and its 95% interval confidence range (coefficient of determination r2 of 0.29 and 0.006 for controls samples and SAM samples respectively) with the green points and line representing healthy children and the red points and line representing severely malnourished children.
Therapeutic diet does not reverse the loss of Methanobrevibacter smithii in severe acute malnutrition
Among severely malnourished children, therapeutic diet and renutrition were not associated with restoration of M. smithii but rather with a loss in the few initially positive children. Indeed, among the 89 severely malnourished children for whom a second sample was available, only one in three children who was initially positive remained positive, while 0/86 children who were initially negative became positive (Table 4). It was more difficult to obtain a second stool sample for the controls. Among 8 healthy children with 2 samples, 2/4 negative samples became positive, while 3/4 positive samples remained positive. The proportion of negative children becoming positive was significantly different according to nutritional status (severe acute malnutrition 0/86 versus healthy children 2/4, two-sided mid-P test, p = 0.001). The proportion of children who became positive or remained positive was significantly lower among severely malnourished children after renutrition (1/89 (1.1%) vs. 5/8 (62.5%), p < 0.001), which suggests that the therapeutic regimen is not able to restore or maintain M. smithii in the gut of these children.
Discussion
There is a paradox that the systematic analysis of the presence of M. smithii reported here has not been carried out before since this archaeon is a candidate of choice to explain good or bad digestion. Here, we confirmed that M. smithii, the main human gut archaeon and a critical human commensal found in virtually all human adults8,9,17, is lost (rather than decreased) in severe acute malnutrition. More than 80% of healthy children were positive when older than 20 months, while only 6 of 143 severely malnourished children were positive, all younger than 15 months.
These findings are robust thanks to the large sample size (n = 253) obtained in a different geographical area (Mali) than our previous work, which was located in Niger and Senegal28, and to the strict and rigorous selection of malnourished and healthy children, according to clinical data and WHO child growth standards 200630. Selection bias was controlled using a propensity score weighting approach based on the different age groups. All children were sampled during the same period of inclusion to avoid time bias. Stool samples of cases were collected before treatment with antibiotics and renutrition.
Using a DNA extraction protocol optimized for methanogen detection with quantitative real-time polymerase chain reaction, we showed that M. smithii was detectable before 24 months but not after 24 months, suggesting a loss of this microbe in children with severe acute malnutrition31. Rather than an immaturity of the microbiota, this suggests that the absence of M. smithii is a marker of dysbiosis, whether due to obesity or severe acute malnutrition12,13, and furthermore supports the association of this archaeon with weight gain in children32,33. Control samples were asymptomatic and without any known disease by definition as children selected as controls were excluded in case of gastroenteritis or a moderate or chronic form of malnutrition. In cases, the low prevalence of digestive symptoms such as diarrhoea, emesis as well as that of gastroenteritis could not explain this loss of M. smithii as confirmed by the analysis excluding the children suffering from severe acute malnutrition exhibiting the aforementioned symptoms.
We have long believed that M. smithii plays an essential role in nutrition. Indeed, Gordon's work in the 2000s showed that the digestion of complex carbohydrates requires the combined action of multiple bacterial strains equipped with a large abundance and diversity of glycosidases8,9. Digestion is accompanied by the production of hydrogen, which ultimately inhibits the metabolic activity of these bacteria. This demonstration prompted us to develop a specific method for the extraction and detection of M. smithii in faeces17. Indeed, 16S rRNA amplifications may miss methanogenic archaea, which have thick walls containing lysozyme-resistant pseudopeptidoglycan and thus require specific DNA extraction procedures17. The result of this earlier work was that in a normal French population, 100% of people were methanogen carriers17, in contrast with an earlier report that only 30% of the population were carriers34.
Since then, we have developed tools for better detection of M. smithii. Using these tools, we have noted the absence of M. smithii in malnourished patients in Africa28, an observation that had escaped other teams working on malnutrition, paradoxically including those of Gordon, whose work inspired our research on M. smithii.
In practice, we confirmed the significant absence of M. smithii in the faeces of malnourished children. We have previously shown that the source of M. smithii in the digestive tract of newborns originated from colostrum and breast milk21. Here, we observed a loss of the methanogenic archaea M. smithii in the gut microbiota of children with severe acute malnutrition. Severe acute malnutrition is an acute disease with an acute risk of death. Whether this loss is reversible after discharge remains unknown. It is noteworthy that M. smithii colonization is associated with organic dairy consumption29. Accordingly, organic dairy consumption may help these children recolonize their gut with M. smithii after the acute disease. This could be tested in further longitudinal studies with longer observation period post-discharge. Moreover, the acute loss of M. smithii may be associated with a sudden collapse of digestion, fermentation and butyrate production which is associated with death in these children35. Supplementation with missing microbes36, including M. smithii by probiotics/organic dairy may be required to prevent death in such a situation.
We believe that it is justified to consider the reintroduction of M. smithii in malnourished subjects in the form of a probiotic additive. Breastfeeding or the addition of milk-borne probiotics could be adequate since colonization has been associated with organic dairy consumption29. Future milk-based microbiome-directed therapies should investigate the potential benefit of the addition of M. smithii to seed the children’s gut and preserve this critical commensal as a key to restoring host-microbial mutualism. In addition, future studies should investigate the possible mechanisms leading to the loss of M. smithii and determine whether this loss is the cause of the consequence of other characteristics of the severe acute microbiota associated dysbiosis (loss of aero-intolerant bacterial species among others).
Methods
Participants/study design
This case–control study was reported according to the indications of the STROBE statement37 (STROBE checklist provided in Table S2) from May 2015 to January 2017 in the peri-urban area of Kalaban Coro located southeast of Bamako in Mali. The cases were severely malnourished children under five years of age and were recruited from the unit of recovery and nutritional education (URENI) of the Kalaban Coro reference health centre. The controls were children of the same age group with no form of acute or chronic illness that can modify the gut microbiota, such as fever, diarrhoea, and/or antibiotic intake, within fifteen days before inclusion.
To test the reversibility of the absence of M. smithii in cases and controls, we attempted to obtain a second sample. In the group of malnourished children, this second sample was collected after the acute phase, i.e., after recovery from acute clinical complications (diarrhoea, dehydration, fever, gastroenteritis, respiratory infections), return of appetite with an intake of at least 75% of the daily RUTF ration required for the child and a weight gain of 15 to 20% compared to the entry weight29. For control children, a second sample was collected 15 days after the first.
Malnourished infants who did not give stool samples before renutrition were excluded as well as those with acute or chronic illness that may explain their nutritional status. Cases of refusal of consent were also excluded. Case and control children were classified by age range of 0–6 months, 7–12 months, 13–24 months and > 24 months.
Ethical considerations
The study was started after the approval of the ethics committee of the Faculty of Medicine and Odonto-Stomatology of Bamako, Mali, under the number: N ° 2014/46/CE/FMPOS on May 22, 2014. Informed and signed consent was obtained for all children from their parents or legal guardian in accordance with the Helsinki declaration. Additionally, all experiments were performed in accordance with relevant guidelines and regulations.
Data sources/measurement/definitions
Anthropometric parameters, including weight, height, mid-upper arm circumference (MUAC) and age, were measured for all participants to determine the nutritional status of the children. We also calculated the weight-for-age, weight-for-height and height-for-age z-scores using the WHO Anthro software (https://www.who.int/childgrowth/software/en/) according to the date of inclusion, gender, date of birth, height measurement recumbent or not, and the presence or absence of oedema. Based on the WHO 2009 severity criterion on acute malnutrition (30), including weight-for-height Z-score (WHZ), weight-for-age Z-score (WAZ), height-for-age Z-score (HAZ), mid-upper arm circumference (MUAC) and the presence of oedema, cases were defined by WHZ <–3 standard deviations (SD), by the presence of nutritional oedema and/or by the MUAC < 115 mm for children over 6 months. Clinical data including temperature (to detect fever), respiratory symptoms and digestive symptoms among which diarrhoea, emesis and gastroenteritis were collected. Moreover, the presence of HIV infection as well as malaria was recorded.
Control children were enlisted in health centres during health monitoring or in the Kalaban Coro health district with anthropometric parameters meeting WHO standards including WAZ > -2 SD, HAZ > -2 SD, WHZ > -2 SD, and MUAC ˃ 125 mm in children older than 6 months and without known disease and who were asymptomatic and without oedema. All the children were screened during the same period and in the same geographical area to address potential sources of bias. A multivariate analysis including age and gender as confounding factors was also performed. In addition, we calculated the size of our sample by referring to the proportion of children positive for M. smithii in controls and in children with severe acute malnutrition in our previous study from Niger and Senegal (15/20 in controls vs 0/20 in cases)28. According to Fleiss with continuity correction38, we needed 8 malnourished compared to 8 controls to be powerful enough (80%) to confirm or infirm the results published in our previous work using a 95% two-sided confidence level28. Here, we included 143 cases and 110 controls, i.e. sample sizes well beyond what was required for this study.
Management of severe acute malnutrition
Management of severe acute malnutrition in Malian URENI (Unités de Récupération et d’Education Nutritionnelle Intensive) consists of three phases, which include, on one hand, nutritional cure with therapeutic milk and ready-to-use therapeutic food (RUTF) and, on the other hand, medical treatment with antibiotics, antiparasitic drugs including antimalarial, and vitamin A supplementation as described by the PECIMA, a programme on the integrated management of severe acute malnutrition (Fig. S1). To avoid therapeutic gut microbiota alteration, faecal samples were collected at admission before administration of any anti-infectious drugs.
Variables and parameters collected/techniques
Gut methanogenic archaea quantification
We performed M. smithii detection by targeting the 16S rRNA gene using real-time quantitative polymerase chain reaction using the optimized protocol of Dridi et al.17.
Real-time quantitative polymerase chain reaction
DNA was extracted manually from 30 mg of feces using the E.Z.N.A. Tissue DNA Kit (Omega Bio-tek, Norcross, GA, USA) according to the manufacturer’s instructions. The total DNA extracted was pure and diluted to the tenth and one-hundredth for real-time quantitative PCR, mainly targeting the 16S rRNA gene of M. smithii and F. prausnitzii, the sohB gene of E. coli and the nucA gene of S. aureus (Table S3). All PCRs were performed with a positive control series (plasmids) and negative controls (mix). Specific primers and probe systems were used for amplification (Table S2). Real-time PCR was performed in a total volume of 20 μL, including 10 μL of master mix (Roche Diagnostics GmbH, Mannheim, Germany), 3 μL of distilled water, 0.5 μL of primer Fwd, 0.5 μL of primer Rev, 0.5 μL of probe, 0.5 of uracil-DNA glycosylase (UDG) and 5 μL of DNA. The amplification reactions were performed using the Roche protocol, which consisted of in two minutes at 50 °C, five minutes at 95 °C followed by 40 cycles of five seconds at 95 °C and 30 s at 60 °C and analysed using the CFX96 real-time PCR detection system (Bio-Rad Life Science, Marnes-La-Coquette, France). The real-time PCR results were considered negative in the absence of an amplification curve.
Statistical methods
The data collected on a questionnaire (supplementary material) were entered in Microsoft Excel and analysed using SPSS software version 20.0 (IBM, Paris, France), SAS 9.4 statistical software (SAS Institute, Cary, NC) and GraphPad Prism 8.0 (GraphPad software, La Jolla, USA). Descriptive statistical analyses were performed for all parameters. The normality test of Shapiro–Wilk was first used for the distribution of quantitative data to apply parametric tests (t tests) or nonparametric tests (Mann–Whitney-Wilcoxon test). The chi-square test was used to test the differences in proportion between groups. All tests were two-tailed. The threshold of significance was set at a value of p ≤ 0.05.
Despite our attempt to recruit controls within the same age group and sex as the cases, controls were still not matched within age categories (p < 0.001, Table S1). In order to control for this confounding factor, we used a propensity score weighting approach on our entire study population. The propensity score was calculated using logistic regression on the age groups. The predicted probabilities from the propensity-score model were used to calculate the stabilized inverse-probability-weighting weights39. Associations between groups (cases/controls) and the different variables were then estimated using weighted regressions (normal or logistic depending on the outcome). The positive predictive value (PPV) of M. smithii according to age for healthy and malnourished children was calculated. PPV is an estimate of the specificity and sensitivity of a variable, calculated using the following formula: number of true positives/(number of true positives + number of false positives)40. We performed linear regression models on the DNA concentration to analyse the dynamics of M. smithii relating to age and to compare the speed of expected increase. For this purpose, the slope difference with the horizontal was determined to evaluate to what degree the DNA concentration of M. smithii was different from zero in each group and between groups. We performed these calculations on the slope of linear regression following the method described by Zar41 and used them in GraphPad Prism 8.0.
References
Bang, C. & Schmitz, R. A. Archaea associated with human surfaces: Not to be underestimated. FEMS Microbiol. Rev. 39, 631–648 (2015).
Chassard, C. & Bernalier-Donadille, A. H2 and acetate transfers during xylan fermentation between a butyrate-producing xylanolytic species and hydrogenotrophic microorganisms from the human gut. FEMS Microbiol. Lett. 254, 116–122 (2006).
Hamer, H. M. et al. Review article: The role of butyrate on colonic function. Aliment. Pharmacol. Ther. 27, 104–119 (2008).
Bui, T. P. N. et al. Mutual metabolic interactions in co-cultures of the intestinal Anaerostipes rhamnosivorans with an acetogen, methanogen, or pectin-degrader affecting butyrate production. Front Microbiol 10, 2449 (2019).
Samuel, B. S. et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. PNAS 105, 16767–16772 (2008).
Stams, A. J. Metabolic interactions between anaerobic bacteria in methanogenic environments. Antonie Van Leeuwenhoek 66, 271–294 (1994).
Hansen, E. E. et al. Pan-genome of the dominant human gut-associated archaeon, Methanobrevibacter smithii, studied in twins. Proc. Natl. Acad. Sci. U.S.A. 108(Suppl 1), 4599–4606 (2011).
Samuel, B. S. et al. Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc. Natl. Acad. Sci. U.S.A. 104, 10643–10648 (2007).
Samuel, B. S. & Gordon, J. I. A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc. Natl. Acad. Sci. U.S.A. 103, 10011–10016 (2006).
Blanton, L. V. et al. Gut bacteria that prevent growth impairments transmitted by microbiota from malnourished children. Science 351, 6275 (2016).
Gehrig, J. L. et al. Effects of microbiota-directed foods in gnotobiotic animals and undernourished children. Science 365, 6449 (2019).
Raman, A. S. et al. A sparse covarying unit that describes healthy and impaired human gut microbiota development. Science 365, eaau4735 (2019).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 6, 341 (2013).
Pausan, M. R. et al. Exploring the archaeome: Detection of archaeal signatures in the human body. Front. Microbiol. 10, 2796 (2019).
Koskinen, K. et al. First insights into the diverse human archaeome: Specific detection of archaea in the gastrointestinal tract, lung, and nose and on skin. mBio 8, e00824 (2017).
Dridi, B., Henry, M., El Khéchine, A., Raoult, D. & Drancourt, M. High prevalence of Methanobrevibacter smithii and Methanosphaera stadtmanae detected in the human gut using an improved DNA detection protocol. PLoS ONE 4, e7063 (2009).
Million, M. et al. Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int. J. Obes. (Lond) 37, 1460–1466 (2013).
Million, M. et al. Obesity-associated gut microbiota is enriched in Lactobacillus reuteri and depleted in Bifidobacterium animalis and Methanobrevibacter smithii. Int. J. Obes. (Lond) 36, 817–825 (2012).
Ignacio, A. et al. Correlation between body mass index and faecal microbiota from children. Clin. Microbiol. Infect. 22(258), e1-8 (2016).
Togo, A. H. et al. Culture of Methanogenic Archaea from human colostrum and milk. Sci. Rep. 9, 18653 (2019).
Black, R. E. et al. Maternal and child undernutrition and overweight in low-income and middle-income countries. Lancet 382, 427–451 (2013).
Golden, M. H. Specific deficiencies versus growth failure: Type I and type II nutrients. SCN News 10–14 (1995).
Hibberd, M. C. et al. The effects of micronutrient deficiencies on bacterial species from the human gut microbiota. Sci. Transl. Med. 9, eaa14069 (2017).
Rivera, J. A., Hotz, C., González-Cossío, T., Neufeld, L. & García-Guerra, A. The effect of micronutrient deficiencies on child growth: A review of results from community-based supplementation trials. J. Nutr. 133, 4010S-4020S (2003).
Trehan, I. et al. Antibiotics as part of the management of severe acute malnutrition. N. Engl. J. Med. 368, 425–435 (2013).
Schoonees, A., Lombard, M. J., Musekiwa, A., Nel, E. & Volmink, J. Ready-to-use therapeutic food (RUTF) for home-based nutritional rehabilitation of severe acute malnutrition in children from six months to five years of age. Cochrane Database Syst. Rev. 5, CD009000 (2019).
Million, M. et al. Increased gut redox and depletion of anaerobic and methanogenic prokaryotes in severe acute malnutrition. Sci. Rep. 6, 26051 (2016).
van de Pol, J. A. A. et al. Gut colonization by Methanogenic archaea is associated with organic dairy consumption in children. Front. Microbiol. 8, 355 (2017).
WHO. Child Growth Standards and the Identification of Severe Acute Malnutrition in Infants and Children: A Joint Statement by the World Health Organization and the United Nations Children’s Fund (World Health Organization, Geneva, 2009).
Million, M. & Raoult, D. Gut dysbiosis in severe acute malnutrition is not an immaturity: The irreversible quantitative-qualitative paradigm shift. Hum. Microbiome J. 15, 100067 (2020).
Mbakwa, C. A. et al. Gut colonization with methanobrevibacter smithii is associated with childhood weight development. Obesity (Silver Spring) 23, 2508–2516 (2015).
Chaudhary, P. P., Conway, P. L. & Schlundt, J. Methanogens in humans: potentially beneficial or harmful for health. Appl. Microbiol. Biotechnol. 102, 3095–3104 (2018).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Attia, S. et al. Mortality in children with complicated severe acute malnutrition is related to intestinal and systemic inflammation: An observational cohort study. Am. J. Clin. Nutr. 104, 1441–1449 (2016).
Tidjani Alou, M. et al. Gut bacteria missing in severe acute malnutrition, can we identify potential probiotics by culturomics?. Front. Microbiol. 8, 899 (2017).
von Elm, E. et al. Strengthening the reporting of observational studies in epidemiology (STROBE) statement: guidelines for reporting observational studies. BMJ 335, 806–808 (2007).
Fleiss, J. L., Levin, B. & Paik, M. C. Statistical Methods for Rates and Proportions (Wiley, New York, 2003).
Robins, J. M. Marginal structural models. J. Am. Stat. Assoc. Section on Bayesian Statistical Science, 1–10 (1997). https://cdn1.sph.harvard.edu/wp-content/uploads/sites/343/2013/03/msm-web.pdf
Blakely, T. & Salmond, C. Probabilistic record linkage and a method to calculate the positive predictive value. Int. J. Epidemiol. 31, 1246–1252 (2002).
Zar, J. H. Biostatistical Analysis (Prentice Hall, Upper Saddle River, 1999).
Acknowledgements
We thank all the IHU team for excellent growing work and Annick ABEILLE for technical help. We wish to honour here the memory of Professor Ogobara DOUMBO, who supervised this study and passed away during the execution of this work; he was a collaborator with unequalled human and professional qualities. We also thank Patricia CARRIERE for helping us to perform the group matching. This work has received financial support from the French Government through the Agence Nationale pour la Recherche (ANR), including the “Programme d’Investissement d’Avenir” under the reference Méditerranée Infection 10-IAHU-03. This work was supported by Région Provence Alpes Côte d’Azur and European funding FEDER PRIMMI (Fonds Européen de Développement Régional—Plateformes de Recherche et d'Innovation Mutualisées Méditerranée Infection).
Author information
Authors and Affiliations
Contributions
A.C., S.K., A.K. and A.T. performed the experiments; A.C. and S.K. collected samples and information from the included individuals; A.C., M.T.A., B.H. and M.M. wrote the manuscript; A.C., S.K., M.T.A., S.C., B.H. and M.M. analysed the data; M.A.T., O.K.D., D.R. and M.M. supervised the study; and O.K.D. and D.R. conceived the study. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Camara, A., Konate, S., Tidjani Alou, M. et al. Clinical evidence of the role of Methanobrevibacter smithii in severe acute malnutrition. Sci Rep 11, 5426 (2021). https://doi.org/10.1038/s41598-021-84641-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-84641-8
This article is cited by
-
Enhanced Biogas Production from Human and Agro-Waste: Waste to Wealth Initiative
Waste and Biomass Valorization (2024)
-
Novel complete methanogenic pathways in longitudinal genomic study of monogastric age-associated archaea
Animal Microbiome (2023)
-
The intersection of undernutrition, microbiome, and child development in the first years of life
Nature Communications (2023)
-
Species-targeted sorting and cultivation of commensal bacteria from the gut microbiome using flow cytometry under anaerobic conditions
Microbiome (2022)
-
Bacteria and Methanogens in the Human Microbiome: a Review of Syntrophic Interactions
Microbial Ecology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.