Abstract
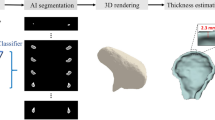
Temporal bone CT-scan is a prerequisite in most surgical procedures concerning the ear such as cochlear implants. The 3D vision of inner ear structures is crucial for diagnostic and surgical preplanning purposes. Since clinical CT-scans are acquired at relatively low resolutions, improved performance can be achieved by registering patient-specific CT images to a high-resolution inner ear model built from accurate 3D segmentations based on micro-CT of human temporal bone specimens. This paper presents a framework based on convolutional neural network for human inner ear segmentation from micro-CT images which can be used to build such a model from an extensive database. The proposed approach employs an auto-context based cascaded 2D U-net architecture with 3D connected component refinement to segment the cochlear scalae, semicircular canals, and the vestibule. The system was formulated on a data set composed of 17 micro-CT from public Hear-EU dataset. A Dice coefficient of 0.90 and Hausdorff distance of 0.74 mm were obtained. The system yielded precise and fast automatic inner-ear segmentations.
Similar content being viewed by others
Introduction
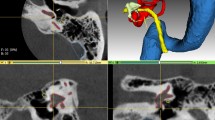
Temporal bone CT-scan is largely used for diagnostic and surgical preplanning in diseases involving the inner ear such as hearing loss and balance disorders1. In routine practice, this technique offers a series of 2D images which are browsed back and forth by the practitioner to mentally deduce 3D information and to this end 3D reconstructions have been applied to training and surgical planning2. On independent 2D image slices, multiple structures can be confounded with inner ear and the mental reconstruction of inner ear structures can be difficult (Fig. 1). Commercially available volume rendering techniques may provide useful information about large structures such as large vessels or lungs3,4 but are not accurate and robust enough for otological applications because the small size of the structures and large variations in background intensity values around the inner ear boundaries may generate artefacts5. Nevertheless, 3D anatomical information is crucial before an ear surgery to predict difficulties and to adapt instrumentation and approach6. For example, when an electrode array is inserted into the cochlea during a cochlear implantation, the information about the anatomy of the cochlea (e.g. malformations, lumen obstruction or narrowing) or its size influences the choice of the array7. Inner ear has a complex 3D anatomy and is surrounded by critical structures like facial nerve and blood vessels, with only few visible landmarks during surgery8,9,10. Moreover, its anatomy is subject to great inter individual variability justifying even more the use of preoperative CT-scan10,11,12.
Automated or aided tridimensional reconstruction of anatomical regions including the temporal bone and the inner ear can be conducted by commercially available softwares13,14,15,16. The main drawback of these generic softwares is the need for interactions and significant anatomical expertise to conduct the segmentation, which could make the work tedious and time-consuming in clinical practice. Softwares specific to temporal bone often use basic algorithms such as region growing, boundary detection and thresholding to segment the cochlea17,18,19,20. This often leads to incomplete segmentation since the identification of inner ear structures solely based on intensity generates multiple errors at each slice and generally yields an aberrant final 3D image. As an alternative, inner ear segmentations are often carried out manually during a tedious and time-consuming process21. Several projects have proposed solutions via semi-automatic algorithms such as 3D-level sets and interactive contour algorithms22,23,24. However, they still require user interaction and introduce human error into the system. Fully automatic algorithms based on active statistical shape modelling have been applied to cochlear segmentation25,26, but to attain an accurate statistical shape in different scenarios, a large amount of annotated data would be required27. Other proposed solutions such as atlas-based frameworks11,12 and iterative random-walks algorithm with shape prior integration produced encouraging results for cochlea segmentation but are computationally expensive28,29. Moreover, with shape priors and atlas-based methods, segmentation might fail if the analysed image diverges from the average shape model, and this is often the case in malformations. Furthermore, these methods can reach theirs limits with clinical CTs at low resolutions which do not include elaborative details on the cochlea.
Deep-learning strategies have gained immense popularity in segmentation tasks where they outperform conventional approaches30. In the medical field, large image sizes and limited availability of expert annotations are the main challenges of deep-learning algorithms which have been recently tackled31,32. To optimize performances, U-Net has been the most popular base-architecture algorithm in this field for several years32. This architecture allows processing a considerable amount of data with a relatively low computational cost and without loss of detail. It has been named after the “U” shape of its architecture: the system downsamples the data while extracting higher level features for a faster processing, and upsamples it again by integrating stored details to deliver a high-quality result33.
To deal with a limited set of data, patch-based learning is effective. This method consists of selecting and analysing patches of relevant features inside an image to classify it instead of using the entire image34,35. This method was developed to reduce the computational burden in selected scenarios, but inherently, it relies on local features instead of using the contextual information. In particular, processing large stacks of high-resolution CT-scan slices (> 1000) by this method is still computationally expensive.
Multi-view algorithms have been proposed to integrate contextual information into the segmentation framework. These algorithms segment structures independently from each orthogonal view and then combine the output using either majority voting, connected component analysis, sigmoid/softmax output scores or FCN output layer36,37,38. However, these architectures do not take the outputs of other views into account in the segmentation architectures. Auto-context is an alternative algorithm that combines low-level appearance features to high-level shape information extracted from all three orthogonal planes39. This architecture was conceived as an alternative to cascaded network schemes in which the output of one network is fed to a second network to incorporate spatial context40. By the combination of these two different approaches, auto-net was developed41. This algorithm fuses the posterior distribution of labels with image features, and it has shown encouraging results in segmenting brain images41. However, it is also performed on image patches and is an iterative algorithm.
We hypothesized that by using deep-learning architectures in an auto-context framework, and by avoiding patch-based classifiers and relatively extensive iterative schemes which are computationally expensive, we could obtain a rapid and precise segmentation of the inner ear from a highly-detailed data set (micro-CT scans).
The aim of this work was to develop and evaluate a fully automatic framework for segmentation of the inner ear structures from micro-CT images. We present an auto-context cascaded convolutional neural network using a three-stage orthogonal U-Net model with connected component extraction after each stage to robustly segment inner ear structures from the micro-CT images.
Material and methods
Dataset preprocessing
The Hear-EU cochlear public dataset was chosen for this project42. It consists of micro CT-scans from 17 human temporal bone specimens. The ground truth labels, manually delineated by an expert neuroradiologist on each slice, consisted of cochlear scalae, semicircular canals, and the vestibule. The images were acquired at 16.3 μm for 13 specimens and 19.5 μm voxel resolutions for the remaining 4. The original volume size of the CT-scans ranged from 618 × 892 × 600 to 1500 × 1500 × 1500. CT-scans were resampled to a fixed size of 256 × 256 × 256 voxels (using spline interpolation), corresponding to 4352 image slices in the dataset for computational and memory requirements. The ground truth labels were not aligned with the raw CT-scan images in 5 specimens and these data were manually aligned with the CT-scan data. The intensity values were zero-centered and normalized by the standard deviation of the training dataset in each experiment and for each cross-validation fold. No parallelization was employed for data processing.
Auto-cascaded net (AutoCasNet)
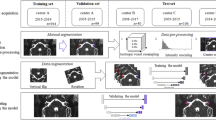
The proposed methodology (Fig. 2) employed a 2D segmentation algorithm to segment each image slice individually in a three-stage cascaded framework to incorporate 3D information. The system began by partitioning the input volumes into axial, coronal and sagittal slices. In the first stage, coronal slices of each CT were fed to a 2D segmentation architecture. The output was then assembled to form a segmented 3D volume. The largest connected component region was extracted from this volume and partitioned into sagittal slices. This output of the first stage was fed to the second stage as binary masks and additional image channels along with their corresponding image slices from the original CT volume. Again, a segmented 3D volume was obtained and fed to the third stage. Finally, in the third stage, axial slices from all previous stages and original CT volume were used as input. The output of this last stage was used as the final system output.
The architecture of the 2D segmentation network was based on encoder-decoder style networks (Fig. 3). The network took a 256 × 256 image slice with ‘X’ image channels as input and output the image mask, where ‘X = 1, 2 or 3’. Every step on the contracting stage consists of two consecutive blocks each having a 3 × 3 convolutional layer followed by batch normalization (BN) and rectified linear unit (ReLU) activation. This was followed by a 2 × 2 max pooling layer. The expanding stage was composed of a similar block preceded by deconvolutional layer and concatenation. Finally, we added a dropout regularization with a 20% dropout rate in the bottleneck layer to avoid overfitting. The output layer was a 1 × 1 convolutional layer with sigmoid activation function.
2D segmentation architectures. (a) U-Net, (b) residual U-Net, (c) SEU-Net with constant ratios X = {1,2,3} in stages {1,2,3} respectively. See Fig. 2 for details.
Training
The network was trained using individual slices of the input volumes. The weights were updated using Adam optimizer with a learning rate of 0.0001. The training was carried out with a batch size of 20 for maximum 200 epochs with early-stopping criteria in each stage, where the best model on training dataset (with the lowest training loss) was checkpointed and stored for testing. The training was carried out with a batch size of 20 for 200 epochs in each stage, where the best model was checkpointed and stored for testing. The binary cross entropy was used as the cost function. The first network was trained until convergence and its output was post-processed and fed into the second network as input for training and similarly for the last stage. The model was implemented using Keras and TensorFlow libraries on a standard Intel i7 computer with 32 GB RAM and a dedicated GPU (Nvidia Titan X processor, 12 GB RAM) and validated using a fourfold cross validation approach with all slices belonging to a patient kept in the same fold. The fourfold evaluation, with hold-out test sets, was performed to reduce the influence of biases in the quantitative analysis43. The dataset included in this study was obtained from two different scanners and acquisition protocols and was randomly placed into the different folds. During the evaluation, different combinations of the orthogonal axes were also tested to determine the optimum ordering of orientation. The code is available at https://github.com/raabid236/AutoCasNet.
As comparison, the runtime of the framework was also tested on a standard Intel Xeon E5-2609 computer with 16 GB RAM.
2D-segmentation architectures
Three segmentation architectures were compared. Each of them was used independently (on the best performing slice orientation) and as the basic segmentation algorithm in our proposed framework:
-
1.
U-Net: Our design was adapted from the original paper33 with batch normalization integration and a change in number of layers and filter sizes (Fig. 3). Dropout regularization was performed on the bottleneck layer. A post-processing step (largest region extraction) was also applied to refine the output. The basic block of the architecture is presented in detail in Fig. 3a.
-
2.
Residual U-Net (ResU-Net): ResU-Net followed the same architecture as in Fig. 3 but incorporated residual blocks44,45 instead of convolution blocks in down- and up-sampling layers. The residual blocks consisted of 3 × 3 and 1 × 1 convolutions summed together (Fig. 3b). Like U-Net, a post processing step was also integrated.
-
3.
Squeeze and Excitation U-Net (SEU-Net): SEU-Net also followed the same architecture as in Fig. 3 but incorporated squeeze and excitation blocks (with constant ratio in all layers)46 along with convolution blocks in down- and up-sampling layers (Fig. 3c). Like previous networks, a post processing step was also integrated.
Evaluation
The evaluation was based on the following volume and distance-based metrics: Dice coefficient (\(\frac{2|X\cap Y|}{\left|X\right|+|Y|}\)), Jaccard index (\(\frac{|X\cap Y|}{|XUY|}\)), segmentation accuracy, precision (\(\frac{|X\cap Y|}{|X|}\)), recall (\(\frac{|X\cap Y|}{|Y|}\)), specificity, area under receiver operating characteristic (ROC) curve and 3D Hausdorff distance (\(\mathrm{max}\{{sup}_{x\in X}{inf}_{y\epsilon Y}d\left(x,y\right),{sup}_{y\in Y}{inf}_{x\epsilon X}d(x,y)\}\)), where X and Y are the segmentation results and the groundtruth respectively47. A Wilcoxon signed-rank test was also performed on the segmentation outputs to analyze the impact of AutoCasNet frameworks. A p-value < 0.05 was considered as significant.
Results
All networks were trained from scratch until the training error reached a plateau state. The dropout regularization helped to prevent overfitting. Table 1 depicts the quantitative results of the segmentation. In general, AutoCasNet based networks outperformed their corresponding state of the art networks48 in most of the evaluation metrics for all the patients in the dataset with AutoCasU-Net yielding the best segmentation. The largest improvement was seen in case of SEU-Net. Moreover, during the experiments similar results were obtained when the order of the slice orientations was changed, ensuring repeatability. Figure 4 depicts the qualitative segmentation results on different image slices. The Wilcoxon signed-rank test revealed a significant improvement in most test cases. The 3D reconstruction of the segmentation output was anatomically realistic with smooth surfaces and very small irregularities (Fig. 5).
Different viewpoints of 3D reconstruction of the inner ear with the cochlea from the AutoCasU-Net segmentation output. ITK-SNAP65 (v 3.4.0, http://www.itksnap.org/) was used for the 3D visualisation of the segmentation output.
In the comparative evaluation, we noted that when contextual information (intermediate outputs from previous stages) was not taken into account, incomplete and unwanted regions that have similar 2D features were also segmented (Fig. 4c,e,g). The proposed framework yielded refined output masks which were coherent with the ground truth labels for all the proposed basic architectures. However, as we can observe from rows 5 and 6 in Fig. 4, the entire region was not segmented in some slices. From visual inspection of 400 random slices, it was found that 10.5% of the slices contained such fragmented segmentation labels. Due to the spiral shape of the structures with small gaps between regions, smoothing and morphological operators were not considered to deal with such fragmented regions.
Segmentation of one 3D volume took 11 s on the GPU and 240 s on the conventional desktop computer. This time included loading the resampled data and the model and predicting the segmentation output.
Discussion
In this study, we presented a fully automatic neural network for segmenting inner ear structures from micro-CT images. The system yielded a precise 3D reconstruction in a few seconds on a computer equipped with a GPU. A PC without a GPU produced the result in less than 5 min. The precision and the rapidity of this program are compatible with its use in routine practice.
Although the segmentation performance might increase with the integration of contextual information through 3D architectures or shape prior modelling as a post-processing step49,50 due to the limited size of the dataset and the large volume size, these options were excluded and a 2D architecture that processed each slice individually was adopted instead. Although patch-based 3D approaches can reduce the high-memory requirements, such approaches are still computationally expensive and they do not incorporate global context which may lead to incomplete segmentations. Another advantage of using slices instead of whole volumes was to extend the dataset size to 4352 individual and independent input cases. The disadvantage of a 2D architecture was that slices do not include all the geometric information of the inner ear and targets can be more easily confounded with similar regions (Fig. 1).
To overcome this weakness, a three-stage cascaded convolutional neural network design was adopted to incorporate 3D information. It employed a 2D U-Net architecture followed by 3D largest-region extraction at each stage in an auto-context framework. We compared the performance of the proposed framework with other deep-learning architectures, and despite the limited size of the dataset, the architecture outperformed other state-of-the-art segmentation algorithms according to several conventional metrics. ResU-Net, and SEU-Net are generally considered to be improved modifications of the original U-Net51,52, but even with the optimal network parameters (e.g. number of layers, filter size, learning rate), they could not compete with U-Net. This advantage was also observed for AutoCasNet frameworks with the same parameters. This difference may be explained by the complex nature of inner ear anatomy and the common choice of hyperparameters in all the frameworks. Indeed, systematic comparisons of U-Net performance in the field of virus identification has shown that a judicious choice of hyperparameters for each framework are pivotal especially if a limited number of trainable weights are used53.
The runtime was adequate for clinical use and could be further improved with a C + + implementation (instead of Python which is comparatively a higher-level language)54. The relative high standard deviation for some metrics could be due to the fact that temporal bones contain air cells that resemble inner ear cavities on many sections. These air cells have variable sizes and locations. This difficulty is inherent to the model. It could hamper the inner ear identification and the reproducibility of the procedure.
Although deep-learning-based methods were used in the AutoCasNet framework in this project, conventional 2D segmentation algorithms could replace them as the basic segmentation technique in this framework. However, conventional algorithms exhibiting good performance for cochlear segmentation have been observed to be computationally expensive11,12,28,29. In a future step, we propose to integrate learned prior shape models in the 2D segmentation algorithm through deep generative networks50,55. This would help to generate a proposal for the shape of the inner ear in a given image slice based on its CT image which can be used to refine the segmentation output.
In future, our framework can be used to build a detailed and robust inner ear model using statistical shape modelling from a high number of micro-CT segmentations42. Also, it will be adapted to abnormal inner ear anatomy to extend its clinical and research applications. Statistical shape modelling algorithms are often used to represent anatomical variations of target structures in a compact parametric model by first registering segmentations obtained from different patients and generating a mesh structure through deformation field generation56,57. This detailed statistical model can then be registered with the segmentation from a patient's clinical CT using automatically extracted anatomical points or intensity-based algorithms58,59. The resulting high-quality co-registered data set of the human bony labyrinth can be used to study microscopic inner ear morphology in detail, for developing efficient design of neuroprostheses and for surgical planning during minimally invasive treatment42,58,60.
Understanding the congenital or acquired (e.g. otosclerosis, fracture, fibrosis) abnormal inner ear anatomy via 3D reconstruction has major consequences on diagnosis, management, and surgical preplanning61. Although the focus of this study was to provide a 3D segmentation of inner ear structure for surgical or clinical applications, the reconstruction of complex living tissues has other applications such as developing finite-element models and 3D printing. For example, the segmentations can be used to modify an already existing finite-element model of the ear in order to estimate the behaviour of an ear with its specific anatomical characteristics, especially when considering the significant interindividual variations of the human inner ear morphology62. Our study is also a first step towards the automatic 3D-reconstruction of the inner ear in abnormal cases. To adapt our system to these complicated cases, specific training with CT-scans from inner ear malformations can be envisaged. Since data on malformation and other acquired abnormalities are relatively rare, artificial data augmentation and transfer learning strategies should be considered to increase the training of the network63,64.
Conclusion
The AutoCasU-Net framework yielded accurate ear segmentations on micro-CT images of human ear in a few seconds. This method has the potential to be applied to routine CT-scans for diagnostic and surgical preplanning purposes.
Data availability
The data i.e. the micro-CTs that were used in this study are publically available at: https://www.smir.ch/objects/204388.
References
Vlastarakos, P. V. et al. CT scan versus surgery: How reliable is the preoperative radiological assessment in patients with chronic otitis media?. Eur. Arch. Otorhinolaryngol. 269(1), 81–86 (2012).
Alenzi, S., Dhanasingh, A., Alanazi, H., Alsanosi, A. & Hagr, A. Diagnostic value of 3D segmentation in understanding the anatomy of human inner ear including malformation types. Ear Nose Throat J. 13, 145561320906621, https://doi.org/10.1177/0145561320906621 (2020).
Oliveira, D. A., Feitosa, R. Q. & Correia, M. M. Segmentation of liver, its vessels and lesions from CT images for surgical planning. Biomed. Eng. Online 10(1), 1–23. https://doi.org/10.1186/1475-925X-10-30 (2011).
Wei, Q., Hu, Y., Gelfand, G. & MacGregor, J. H. Segmentation of lung lobes in high-resolution isotropic CT images. IEEE Trans. Biomed. Eng. 56(5), 1383–1393 (2009).
Ferreira, A., Gentil, F. & Tavares, J. M. R. Segmentation algorithms for ear image data towards biomechanical studies. Comput. Methods Biomech. Biomed. Eng. 17(8), 888–904 (2014).
Saikawa, E. et al. Cochlear implantation in children with cochlear malformation. Adv. Otorhinolaryngol. 77, 7–11 (2016).
Vu, T. H. et al. CT-scan contouring technique allows for direct and reliable measurements of the cochlear duct length: Implication in cochlear implantation with straight electrode-arrays. Eur. Arch. Otorhinolaryngol. 276(8), 2135–2140 (2019).
Dhanasingh, A. Variations in the size and shape of human cochlear malformation types. Anat. Rec. 302(10), 1792–1799 (2019).
van der Jagt, A. M., Kalkman, R. K., Briaire, J. J., Verbist, B. M. & Frijns, J. H. Variations in cochlear duct shape revealed on clinical CT images with an automatic tracing method. Sci. Rep. 7(1), 17566 (2017).
Meng, J., Li, S., Zhang, F., Li, Q. & Qin, Z. Cochlear size and shape variability and implications in cochlear implantation surgery. Otol. Neurotol. 37(9), 1307–1313 (2016).
Al-Dhamari, I. et al. Automatic cochlear length and volume size estimation. In Proceedings of OR 2.0 Context-Aware Operating Theatres, Computer Assisted Robotic Endoscopy, Clinical Image-Based Procedures, and Skin Image Analysis, 54–61, https://doi.org/10.1007/978-3-030-01201-4_7 (2018).
Iyaniwura, J. E., Elfarnawany, M., Ladak, H. M. & Agrawal, S. K. An automated A-value measurement tool for accurate cochlear duct length estimation. Otolaryngol. Head Neck Surg. 47(1), 5 (2018).
Rutel, I. B., Stoner, J., Kota, P., Dormer, K. & Alleman, A. Orientation of the round window membrane: A normative study of inner ear anatomical orientation using 2D projections of 3D volumes. Anat. Rec. (Hoboken), Early View, https://doi.org/10.1002/ar.24327 (2019).
Elfarnawany, M. et al. Micro-CT versus synchrotron radiation phase contrast imaging of human cochlea. J. Microsc. 265(3), 349–357 (2017).
Schart-Morén, N., Agrawal, S. K., Ladak, H. M., Li, H. & Rask-Andersen, H. Effects of various trajectories on tissue preservation in Cochlear implant surgery: A micro-computed tomography and synchrotron radiation phase-contrast imaging study. Ear Hear. 40(2), 393–400 (2019).
Vezhnevets, V. & Konouchine, V. GrowCut: Interactive multi-label ND image segmentation by cellular automata. Proc. Graphicon 1(4), 150–156 (2005).
Franz, D. et al. Wizard-based segmentation for cochlear implant planning. in Proceedings of Bildverarbeitung für die Medizin (BVM), 258–263, https://doi.org/10.1007/978-3-642-54111-7_49 (2017).
Gerber, N. et al. Surgical planning tool for robotically assisted hearing aid implantation. Int. J. Comput. Assist. Radiol. Surg. 9(1), 11–20 (2014).
Folowosele, F. O., Camp, J. J., Brey, R. H., Lane, J. I. & Robb, R. A. 3D imaging and modeling of the middle and inner ear. Proc. SPIE Med. Imaging 5367, 508–516, https://doi.org/10.1117/12.535364 (2004).
Rodt, T. et al. 3D visualisation of the middle ear and adjacent structures using reconstructed multi-slice CT datasets, correlating 3D images and virtual endoscopy to the 2D cross-sectional images. Neuroradiology 44(9), 783–790 (2002).
Bonne, N. X., Dubrulle, F., Risoud, M. & Vincent, C. How to perform 3D reconstruction of skull base tumours. Eur. Ann. Otorhinolaryngol. Head Neck Dis. 134(2), 117–120 (2017).
Poznyakovskiy, A. A. et al. A segmentation method to obtain a complete geometry model of the hearing organ. Hear. Res. 282(1–2), 25–34 (2011).
Xianfen, D., Siping, C., Changhong, L. & Yuanmei, W. 3D semi-automatic segmentation of the cochlea and inner ear. in Proceedings of IEEE Annual Conference of Engineering in Medicine and Biology (EMBC), 6285–6288, https://doi.org/10.1109/IEMBS.2005.1615934 (2006).
Yoo, S. K., Wang, G., Rubinstein, J. T. & Vannier, M. W. Semiautomatic segmentation of the cochlea using real-time volume rendering and regional adaptive snake modeling. J. Digit. Imaging 14(4), 173–181 (2001).
Noble, J. H., Gifford, R. H., Labadie, R. F. & Dawant, B. M. Statistical shape model segmentation and frequency mapping of cochlear implant stimulation targets in CT. in Proceedings of International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 421–428, https://doi.org/10.1007/978-3-642-33418-4_52 (2012).
Noble, J. H., Labadie, R. F., Majdani, O. & Dawant, B. M. Automatic segmentation of intracochlear anatomy in conventional CT. IEEE Trans. Biomed. Eng. 58(9), 2625–2632 (2011).
Heimann, T. & Meinzer, H. P. Statistical shape models for 3D medical image segmentation: A review. Med. Image Anal. 13(4), 543–563 (2009).
Pujadas, E. R., Piella, G., Kjer, H. M. & Ballester, M. A. G. Random walks with statistical shape prior for cochlea and inner ear segmentation in micro-CT images. Mach. Vis. Appl. 29(3), 405–414 (2018).
Pujadas, E. R., Kjer, H. M., Vera, S., Ceresa, M. & Ballester, M. Á. G. Cochlea segmentation using iterated random walks with shape prior. in Proceedings of SPIE Medical Imaging, Vol. 9784, 97842U, https://doi.org/10.1117/12.2208675 (2016).
Sultana, F., Sufian, A. & Dutta, P. Evolution of image segmentation using deep convolutional neural network: A survey. Knowl. Based Syst. 201–202, 106062, https://doi.org/10.1016/j.knosys.2020.106062 (2020).
Chen, C. et al. Deep learning for cardiac image segmentation: A review. Front. Cardiovasc. Med. 7, 25 (2020).
Bernard, O. et al. Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: Is the problem solved?. IEEE Trans. Med. Imaging 37(11), 2514–2525 (2018).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. in Proceedings of International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 234–241, https://doi.org/10.1007/978-3-319-24574-4_28 (2015).
Anders, C. J., Montavon, G., Samek, W. & Müller, K. R. Understanding patch-based learning of video data by explaining predictions. in Lecture Notes in Computer Science, Vol. 11700, 297–309, https://doi.org/10.1007/978-3-030-28954-6_16 (2019).
Hou, L. et al. Patch-based convolutional neural network for whole slide tissue image classification. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2424–2433, https://doi.org/10.1109/CVPR.2016.266 (2016).
Wang, S. et al. A multi-view deep convolutional neural networks for lung nodule segmentation. in 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 1752–1755, https://doi.org/10.1109/EMBC.2017.8037182 (2017).
Ma, L., Stückler, J., Kerl, C. & Cremers, D. September. Multi-view deep learning for consistent semantic mapping with rgb-d cameras. in IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), 598–605, https://doi.org/10.1109/IROS.2017.8202213 (2017).
Mortazi, A., Karim, R., Rhode, K., Burt, J. & Bagci, U. CardiacNET: Segmentation of left atrium and proximal pulmonary veins from MRI using multi-view CNN. in International Conference on Medical Image Computing and Computer-Assisted Intervention(MICCAI), 377–385, https://doi.org/10.1007/978-3-319-66185-8_43 (2017).
Tu, Z. & Bai, X. Auto-context and its application to high-level vision tasks and 3d brain image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 32(10), 1744–1757 (2009).
Littmann, E. & Ritter, H. Cascade network architectures. in Proceedings of International Joint Conference on Neural Networks (IJCNN), Vol. 2, 398–404, https://doi.org/10.1109/IJCNN.1992.226955 (1992).
Salehi, S. S. M., Erdogmus, D. & Gholipour, A. Auto-context convolutional neural network (auto-net) for brain extraction in magnetic resonance imaging. IEEE Trans. Med. Imaging 36(11), 2319–2330 (2017).
Gerber, N. et al. A multiscale imaging and modelling dataset of the human inner ear. Sci. Data 4, 170132, https://doi.org/10.1038/sdata.2017.132 (2017).
James, G., Witten, D., Hastie, T. & Tibshirani, R. An Introduction to Statistical Learning Vol. 112, 18. (Springer, New York, 2013).
Alom, M. Z., Yakopcic, C., Hasan, M., Taha, T. M. & Asari, V. K. Recurrent residual U-Net for medical image segmentation. J. Med. Imaging 6(1), 014006 (2019).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. in Proceedings of IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 770–778, https://doi.org/10.1109/CVPR.2016.90 (2016).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 7132–7141, https://doi.org/10.1109/CVPR.2018.00745 (2018).
Taha, A. A. & Hanbury, A. Metrics for evaluating 3D medical image segmentation: Analysis, selection, and tool. BMC Med. Imaging 15(1), 29 (2015).
Hesamian, M. H., Jia, W., He, X. & Kennedy, P. Deep learning techniques for medical image segmentation: Achievements and challenges. J. Digit. Imaging 32(4), 582–596 (2019).
Zhou, T., Ruan, S., & Canu, S. A review: Deep learning for medical image segmentation using multi-modality fusion. Array 3–4, 100004, https://doi.org/10.1016/j.array.2019.100004 (2019).
Girum, K. B., Créhange, G., Hussain, R., Walker, P. M., & Lalande, A. Deep generative model-driven multimodal prostate segmentation in radiotherapy. in Artificial Intelligence in Radiation Therapy (AIRT): Lecture Notes in Computer Science Vol. 11850, 119–127, https://doi.org/10.1007/978-3-030-32486-5_15. (2019).
Liu, S. et al. Segmenting nailfold capillaries using an improved U-net network. Microvasc. Res. 130, 104011, https://doi.org/10.1016/j.mvr.2020.104011 (2020).
Roy, A. G., Navab, N. & Wachinger, C. Recalibrating fully convolutional networks with spatial and channel “squeeze and excitation” blocks. IEEE Trans. Med. Imaging 38(2), 540–549 (2019).
Matuszewski, D. J. & Sintorn, I. M. Reducing the U-Net size for practical scenarios: Virus recognition in electron microscopy images. Comput. Methods Programs Biomed. 178, 31–39 (2019).
Fourment, M. & Gillings, M. R. A comparison of common programming languages used in bioinformatics. BMC Bioinform. 9(1), 82 (2008).
Girum, K. B., Lalande, A., Hussain, R. & Créhange, G. A deep learning method for real-time intraoperative US image segmentation in prostate brachytherapy. Int. J. Comput. Assist. Radiol. Surg. 15(9), 1467–1476 (2020).
Frangi, A. F., Rueckert, D., Schnabel, J. A. & Niessen, W. J. Automatic construction of multiple-object three-dimensional statistical shape models: Application to cardiac modeling. IEEE Trans. Med. Imaging 21(9), 1151–1166 (2002).
Lüthi, M. et al. Statismo-A framework for PCA based statistical models. Insight J. 2012, 1–18 (2012).
Wimmer, W. et al. Human bony labyrinth dataset: Co-registered CT and micro-CT images, surface models and anatomical landmarks. Data Brief 27, 104782, https://doi.org/10.1016/j.dib.2019.104782 (2019).
Hussain, R., Lalande, A., Girum, K.B., Guigou, C. & Grayeli, A.B. Augmented reality for inner ear procedures: Visualization of the cochlear central axis in microscopic videos. Int. J. Comput. Assist. Radiol. Surg., Early Access, 1–9 (2020).
Sieber, D. et al. The OpenEar library of 3D models of the human temporal bone based on computed tomography and micro-slicing. Sci. Data. 6, 180297, https://doi.org/10.1038/sdata.2018.297 (2019).
Dhanasingh, A., Dietz, A., Jolly, C. & Roland, P. Human inner-ear malformation types captured in 3D. J. Int. Adv. Otol. 15(1), 77 (2019).
Avci, E., Nauwelaers, T., Lenarz, T., Hamacher, V. & Kral, A. Variations in microanatomy of the human cochlea. J. Comp. Neurol. 522(14), 3245–3261 (2014).
Bae, H. J. et al. A Perlin noise-based augmentation strategy for deep learning with small data samples of HRCT images. Sci. Rep. 8(1), 1–7. https://doi.org/10.1038/s41598-018-36047-2 (2018).
Kim, D. H. & MacKinnon, T. Artificial intelligence in fracture detection: Transfer learning from deep convolutional neural networks. Clin. Radiol. 73(5), 439–445 (2018).
Yushkevich, P. A. et al. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31(3), 1116–1128 (2006).
Acknowledgements
Authors acknowledge the financial support of Oticon Medical, France for this project. The authors are also grateful to the NVIDIA GPU grant program for donating the TITAN X processor.
Author information
Authors and Affiliations
Contributions
R.H.: initial idea, writing manuscript, software development, architecture design. A.L.: initial idea, writing manuscript, resource management, data analysis. K.B.G.: software development, architecture design. C.G.: data preprocessing, medical expertise. A.B.G.: initial idea, experiment design, resource management, medical expertise. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hussain, R., Lalande, A., Girum, K.B. et al. Automatic segmentation of inner ear on CT-scan using auto-context convolutional neural network. Sci Rep 11, 4406 (2021). https://doi.org/10.1038/s41598-021-83955-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83955-x
This article is cited by
-
Landmark-based registration of a cochlear model to a human cochlea using conventional CT scans
Scientific Reports (2024)
-
Towards fully automated inner ear analysis with deep-learning-based joint segmentation and landmark detection framework
Scientific Reports (2023)
-
Interobserver variability of cochlear duct measurements in pediatric cochlear implant candidates
European Archives of Oto-Rhino-Laryngology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.