Abstract
Salvia miltiorrhiza is one of the most widely used traditional medicines. Natural antisense transcripts (NATs) are a class of long noncoding RNAs that can regulate gene expression. Here, we identified 812 NATs, including 168 cis-NATs and 644 trans-NATs from twelve root, flower, and leaf samples of S. miltiorrhiza using RNA-seq. The expression profiles for 41 of 50 NATs and their sense transcripts (STs) obtained from RNA-Seq were validated using qRT-PCR. The expression profiles of 17 NATs positively correlated with their STs. GO and KEGG pathway analyses mapped the STs for cis-NATs to pathways for biosynthesis of secondary metabolites. We characterized four NATs in detail, including NAT0001, NAT0002, NAT0004, and NAT00023. Their STs are kaurene synthase-like 1 and the homologs of UDP-glucose flavonoid 3-O-glucosyltransferase 6, UDP-glycosyltransferase 90A1, and beta-glucosidase 40, respectively. The first gene is involved in the biosynthesis of bioactive tanshinones, the next two are involved in anthocyanin biosynthesis, whereas the last is involved in phenylpropanoid biosynthesis. Besides, we found seven STs that are potential targets of miRNAs. And we found two miRNAs including miR156a and miR7208, might originate from NATs, NAT0112 and NAT0086. The results suggest that S. miltiorrhiza NATs might interact with STs, produce miRNAs, and be regulated by miRNAs. They potentially play significant regulatory roles in the biosynthesis of bioactive compounds.
Similar content being viewed by others
Introduction
Natural antisense transcripts (NATs) are a class of long noncoding RNAs, which transcribe in the opposite direction of protein-coding genes1. Relatively, the transcripts of protein-coding genes were called sense transcripts (STs). Base on the position of NATs and their STs, NATs can be divided into two categories, namely, cis-NATs and trans-NATs2,3. Cis-NATs originated from the same genomic loci with STs. In contrast, trans-NATs originated from different genomic loci with STs. NATs were initially detected in bacteria4, then they were found in prokaryotes5 and eukaryotes6. With the recent development of high-throughput DNA sequencing technologies and bioinformatics analysis methods, genome-wide NATs have been reported in various plants, such as Arabidopsis thaliana7, Oryza sativa8, Zea mays9, Saccharum officinarum10, and Petunia hybrida11.
It is found that NATs play an essential role in regulating gene expression through different mechanisms12, such as mechanisms related to transcriptional interference13, RNA–DNA interactions14, RNA–RNA interactions in the nucleus15, and RNA–RNA interactions in the cytoplasm16. In Arabidopsis thaliana, a cis-NAT producing from the opposite strand of MADS AFFECTING FLOWERING4 (MAF4) gene can positively regulate the expression of its ST MAF4, thereby affecting flowering time. It activates MAF4 by recruiting WDR5a to MAF4 to enhance histone 3 lysine 4 trimethylation (H3K4me3)17. In Oryza sativa, a cis-NAT was produced from an R2R3-MYB gene locus and suppressed the R2R3-MYB by mediating chromatin modifications18. In mutant plants with downregulation of the NAT by RNA interference (RNAi), the expression level of the R2R3-MYB transcript significantly increases, resulting in twisted leaf blades. Besides, a NAT originates from the CYCLING DOF FACTOR 5 (CDF5) gene locus in Arabidopsis thaliana. In mutant plants with a T-DNA insertion in non-overlapping regions of CDF5, the expression level of CDF5 transcript was strongly reduced compared with wild plants, while its NAT increased 2–4 folds19.
Salvia miltiorrhiza Bunge is one of the most important traditional medicines from the Lamiaceae family. Tanshinones is one of the major active compounds in S. miltiorrhiza and are used to treat cardiovascular diseases through anti-atherosclerosis20, anti-oxidation21, etc. Besides, this plant also produces many flavonoids22, such as rutin, isoquercitrin, and astragalin. Various genes in the biosynthetic pathway of tanshinones have been identified in S. miltiorrhiza. These include the SmCPS gene encoding copalyl diphosphate synthase, SmKSL gene encoding ent-kaurene synthase23, SmCYP76AH1 gene encoding cytochrome P450 enzymes24, etc. The whole-genome25,26, transcriptome27,28,29, and microRNAs30,31,32 of S. miltiorrhiza have been reported, respectively. However, a systematic analysis of NATs is not yet available for S. miltiorrhiza. Here, we conducted a genome-wide identification of NATs in S. miltiorrhiza. The results suggest that NATs might interact with different types of molecules such as the STs and miRNAs, and consequently play an important role in a wide range of biological processes such as the biosynthesis of active compounds and glycosylation.
Results
Global identification of NATs in Salvia miltiorrhiza
To identify the NATs in S. miltiorrhiza, we carried out strand-specific RNA-seq (ssRNA-seq) analysis of 12 RNA samples extracted from three tissues, including roots (r), flowers (f), and leaves (l). We obtained 25.3, 53.8, and 26.7 million paired-end reads from pooled flower, leaf, and root samples. And we obtained 44.5 (p01_f), 43.4 (p02_f), 42.9 (p03_f), 42.5(p01_l), 44.2 (p02_l), 41.6 (p03_l), 44.2 (p01_r), 41.9 (p02_r) and 40.9 (p03_r) million paired-end reads (Table S1).
We identified the NATs using the bioinformatic pipeline shown in Fig. 1. In step 1, 83.5%, 79.2%, and 84.3% reads from flowers, leaves, and roots, respectively, were mapped to the S. miltiorrhiza reference genome sequence25. We assembled a total of 63,858 transcripts. Among them, 30,478 were previously annotated protein-coding transcripts in the reference genome. Using an FPKM cutoff value of 1, we found a total of 44,596 transcripts, including 22,099 previously annotated protein-coding transcripts expressed in at least one of the three tissues. In step 2, we identified 21,790 candidate NATs. It includes 2,875 candidate cis-NATs and 18,915 candidate trans-NATs. Then, we identified 6,323 candidate trans-NATs based on the annealing potential of complementary regions. In step 3, we obtained 2,444 candidate cis-NATs and 5,085 candidate trans-NATs with a sequence length of more than 200 bp. These candidate NATs were subject to a series of filtering steps based on noncoding RNA identification. After removing transcripts with the maximum length of predicted open-reading frame ≥ 100 aa, transcripts matching the protein database, and transcripts with protein-coding potential (CPC score > 0), we finally annotated 812 NATs, which include 168 cis-NATs and 644 trans-NATs (Table S2, Sfile 1). The number of cis-NATs and trans-NATs correspond to 0.55% (168/30,478) and 2.11% (644/30,478) of predicted protein-coding genes, respectively.
The bioinformatics pipeline used to identify NATs in S. miltiorrhiza. The figure has three columns. The main steps of the pipeline are shown in the left column. The detailed analytical processes are shown in the middle column. The results produced by a particular process are shown in the right column.
Characterization of NATs in S. miltiorrhiza
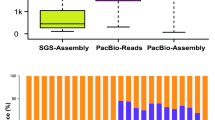
We analyzed the features of the identified NATs. It includes average size, exon number, and the length of overlapping regions. The length of cis-NATs ranged from 200 to 2641 bp, with an average size of 647.16 bp. The length of trans-NATs ranged from 200 to 4341 bp with an average size of 545.62 bp. The majority length distribution ranged from 200 to 300 bp in both cis-NATs and trans-NATs (Fig. 2A). There were also some NATs with lengths longer than 1000 bp. In general, the length distribution of cis-NATs and trans-NATs was similar. However, the number of exons in cis-NATs (the average number was 2.07) was more than that in trans-NATs (the average number was 1.10).
Characterization of S. miltiorrhiza NATs. (A) Length distribution of NATs. (B) Comparison of the proportion of the length of overlapping regions to that of STs and NATs with the length of the overlapping region. (C) Schematic representation of different types of cis-NATs based on the location relative to the STs. STs and NATs were depicted as gray and black bars, respectively. Arrows indicated the directions of transcription.
The mapping relationships between NATs and STs were found to be multiplex33. Some NATs are mapped to multiple STs. On the other hand, some STs were mapped to multiple NATs. This phenomenon was also observed in the SAT pairs (ST and NAT pairs) of S. miltiorrhiza. We classified the mapping relationships of SAT pairs into four types, including “1 versus 1”, “1 versus n”, “n versus 1”, and “n versus n” (Table S2). “1 versus 1” represents one NAT mapped to one ST, “1 versus n” represents one NAT mapped to multiple STs, “n versus 1” represents multiple NATs mapped to one ST, and “n versus n” represents multiple NATs mapped to multiple STs. We found a total of 1045 SAT pairs corresponding to 168 cis-NATs and 644 trans-NATs. For cis-NATs, 134 and 34 formed 1 versus 1 and n versus 1 relationship, respectively. However, we found no 1 versus n and n versus n relationships for cis-NATs. For trans-NATs, 411, 159, 155, and 152 formed 1 versus 1, 1 versus n, n versus 1, and n versus n relationships. The multiplex relationships are most likely due to duplicate genes in the genome. Then, we examined the proportion of the length of overlapping regions between STs and NATs (Fig. 2B). In general, the average proportion of the overlapping region to NATs (73.08%) is larger than that of STs (53.37%).
We classified the cis-NATs into four types based on the location relative to the STs. It includes convergent (with 3′-ends overlapping), divergent (with 5′-ends overlapping), STs containing NATs (S > N), and NATs containing STs (N > S) (Fig. 2C). Among the 168 cis-NATs, 40 were divergent, 60 were convergent, 60 were STs containing NATs, and 8 were NATs having STs. The primary type was convergent and STs containing NATs.
Conservation of NATs identified in S. miltiorrhiza
To determine the conservation of NATs identified in S. miltiorrhiza, we searched the PlncRNADB (http://bis.zju.edu.cn/PlncRNADB/index.php) and GreeNC34 databases for homologs. A total of 4 NATs, NAT0030, NAT0047, NAT0131, and NAT0346, were found to be conserved among the species of Medicago truncatula and Mimulus guttatus (Table 1).
Functional enrichment analyses of the STs having cis-NATs
To explore NATs’ putative function, we analyzed Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway35 analyses on all the STs having cis-NATs in three tissues analyzed. Almost all STs were mapped to GO terms. The STs were mapped to the subgroups of the biological processes, the cellular components subgroup, and the molecular functions subgroup counted for 68.1%, 94.1%, and 67.6%, respectively. Among them, five GO terms were significantly enriched. The terms include defense response by callose deposition, regulation of transport, cellular response to the extracellular stimulus, vesicle organization, and ADP binding (Table S3). Moreover, 14 STs were mapped to 19 KEGG pathways, including anthocyanin biosynthesis, beta-alanine metabolism, biosynthesis of antibiotics, biosynthesis of secondary metabolites, carbon metabolism, cutin, suberine and wax biosynthesis, cyanoamino acid metabolism, lipoic acid metabolism, metabolic pathways, phenylpropanoid biosynthesis, plant-pathogen interaction, propanoate metabolism, protein processing in the endoplasmic reticulum, purine metabolism, pyrimidine metabolism, riboflavin metabolism, RNA polymerase I, starch and sucrose metabolism and valine, leucine, and isoleucine degradation (Table S4).
Validation of ssRNA-seq experiments using quantitative real-time PCR (qPCR)
To validate the expression levels of cis-NATs and STs obtained from ssRNA-Seq in S. miltiorrhiza, we performed strand-specific reverse transcription-quantitative real-time PCR (ssRT-qPCR) on 25 SAT pairs with three replicates for each tissue. We selected 25 SAT pairs for validation if they met one of the following two conditions: (1) the expression profiles of NATs and parental genes were positively correlated with r values ≥ 0.9; and (2) the parental genes of these NATs were mapped to the GO terms or the KEGG pathways of interest. The SAT pairs satisfied condition 1 included SAT0005, SAT0010, SAT0011, SAT0013, SAT0015, SAT0018, SAT0021, SAT0024, SAT0025. The SAT pairs satisfied condition 2 included SAT0001, SAT0002, SAT0003, SAT0004, SAT0005, SAT0006, SAT0007, SAT0008, SAT0009, SAT0010, SAT0011, SAT0012, SAT0013, SAT0014, SAT0015, SAT0016, SAT0017, SAT0018, SAT0019, SAT0020, SAT0021, SAT0022, SAT0023. Detailed information for these 25 SAT pairs were shown in Table S5. The sequences of NATs and STs were showed in Sfile 2. For each transcript, we calculated the average expressed level of three replicates. Then we calculated the Pearson correlation coefficients (r) of the expressed level between ssRNA-Seq and ssRT-qPCR. Forty-one of 50 transcripts showed a positive correlation, twenty-three of which had an r valve more than 0.9 (Fig. 3, Figure S1). These results indicate that the data from RNA-Seq and qPCR are highly consistent. Moreover, the qPCR data also confirmed the differential expression of cis-NATs in the three tissues analyzed.
Validation of the expression level of 25 NATs in flower (F), root (R), and leaf (L) tissues of S. miltiorrhiza. The X-axis shows the expression levels in the three tissues using the two methods: ssRNA-seq and ssRT-qPCR. Y-axis shows the relative expression levels normalized to the mean expression levels across the three tissues. Error bars represent the standard errors. r: the correlation coefficient between the expression levels determined by the two experiments.
Differential expression of cis-NATs in flowers, leaves, and roots
The expression of cis-NATs in flowers, leaves, and roots was analyzed using nine RNA-Seq data from the three tissues with three biological replications from each tissue. We found six cis-NATs differentially expressed among three tissues in S. miltiorrhiza, namely, NAT0005, NAT0039, NAT0050, NAT0076, NAT0080, NAT0131 (Table S6). Among them, NAT0076 in leaves was expressed higher than that in flowers. NAT0039 and NAT0131 in roots were expressed lower than those in flowers. NAT0080 in roots were expressed higher than those in flowers. NAT0039, NAT0076, and NAT0131 in roots were expressed lower than those in leaves. NAT0005, NAT0050, and NAT0080 in roots were expressed higher than those in leaves.
Co-expression of NATs and cognate STs
It is shown that NATs could regulate the expression of corresponding STs12. To preliminarily explore the function of NATs in S. miltiorrhiza, we examined the correlation of expression levels between cis-NATs and the cognate STs using nine RNA-Seq data from the three tissues with three biological replications from each tissue (Table S7). A total of 19 of 168 (11.31%) cis-SATs pairs had a significantly positive correlation coefficient with r valve ≥ 0.9. And the trans-SATs pairs had a relatively low proportion. Only 37 of 644 (5.75%) pairs had a significantly positive correlation coefficient.
We also examined the correlation of expression levels between NATs and the cognate STs using the qPCR data; 22 of 25 pairs had a positive correlation coefficient, of which 17 pairs had r valve more than 0.9 (Fig. 4). The remaining three pairs had a negative correlation coefficient. The correlation coefficient values are − 0.06, − 0.61, and − 0.10, respectively (Fig. 4B, M, U). The results suggest a high correlation between 68% NATs and their cognate STs.
The correlation of expression levels of 25 NAT and ST pairs in flower (F), root (R), and leaf (L) tissues of S. miltiorrhiza. The X-axis shows the three tissues. Y-axis shows relative expression levels of NATs and STs analyzed using the ssRT-qPCR method. Error bars represent the standard error among three replicates. The values of r represent the correlation coefficient of expression levels between NATs and STs.
Differential expression and co-expression of NATs associated with tanshinone biosynthesis
Tanshinones are a class of active compounds mainly produced in the roots of S. miltiorrhiza. They are widely used for the treatment of cardiovascular diseases20. We found that the SmKSL1 gene (Sfile 3) involved in tanshinone biosynthesis had a cis-NAT. SmKSL1 converses diterpenoid precursor, (E,E,E)-geranylgeranyl diphosphate (GGPP) to the skeleton of miltiradiene (Fig. 5A)23. This SAT pair was named SAT0001, which contains ST0001 and NAT0001. There were four overlapped regions with a length of 181 bp, 38 bp, 165 bp, and 49 bp, respectively (Fig. 5A). Both NAT0001 and ST0001 had the highest expression level in roots and the lowest in flowers (Fig. 4A). The correlation coefficient of this SAT pair was 0.99, indicating a co-expression.
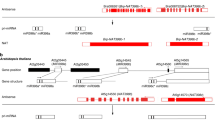
Gene structures of four SAT pairs. Panel (A) had two parts: the biosynthetic pathway of tanshinones in S. miltiorrhiza (above) and the gene structure of SAT0001 pairs (below). The enzymes were shaded in gray. The related ST and chemical compounds were highlighted with a white rectangle. Solid arrows indicated the reactions that have been experimentally validated, and dashed arrows indicated the hypothetical reactions. KSL1 that was encoded by ST0001 was marked by a red rectangle. In the area describing the gene structure, gray and black horizontal bars represent NAT and ST’s exons, respectively. The lines between bars represent introns. Arrows indicate the directions of transcription. The length of the overlapping region is marked above the structure. Similarly, panels (B, C, D) displayed the structures of SAT0002, SAT0004, and SAT0023 pairs in the same way as for SAT0001, respectively.
Differential expression and co-expression of NATs associated with glycosylation and de-glycosylation
Glycosylation is a key step in modifying the complexity and diversity of secondary metabolites in plants36. The uridine diphosphate-glucosyltransferase (UDP-glucosyltransferase, UGT) is one of the glycosyltransferases that use UDP-glucuronic acid as a glycosyl donor to catalyze glycosidic bond formation37. We found two transcripts (ST0002 and ST0004), which encode candidate UDP-glucosyltransferases, had cis-NATs (Sfiles 4, 5). Besides, ST0002 was mapped to the UDP-glucose flavonoid 3-O-glucosyltransferase 6 gene of Fragaria x ananassa that can catalyze quercetin to form the 3-O-glucosides38. SAT0002 belongs to the type ST containing NAT (Fig. 5B). Both NAT0002 and ST0002 had one exon and one overlapped region with a length of 330 bp. NAT0002 had the highest expression level in roots and the lowest in flowers. However, ST0002 had the highest expression level in leaves and the lowest in roots (Fig. 4B). The correlation coefficient of this SAT pair was − 0.06. The type of SAT0004 was convergent (with 3′-ends overlapping) (Fig. 5C). Both NAT0004 and ST0004 had two exons and overlapped in the second exon with 101 bp length. Both NAT0004 and ST0004 had the highest expression levels in roots and the lowest in flowers (Fig. 4D). The correlation coefficient of this SAT pair was 0.99.
β-glucosidase catalyzes the hydrolysis of terminal, non-reducing β-D-glucosyl residues to release β-D-glucose and the corresponding free aglycone39. ST0023 was mapped to the β-glucosidase 40 gene of A. thaliana (Sfile 6). ST0023 had a cis-NAT, termed NAT0023. The type of SAT0023 was convergent (with 3`-ends overlapping) (Fig. 5D). There were two overlapped regions with a length of 37 bp and 98 bp, respectively. NAT0023 had the highest expression level in roots and the lowest in leaves, whereas ST0023 had the highest expression level in roots and the lowest in flowers. The correlation coefficient of this SAT pair was 0.30 (Fig. 4W).
Identification of miRNA targeting STs for cleavage in S. miltiorrhiza
It was reported that NATs regulate the expression of their STs by increasing the stability of ST mRNA through RNA duplex formation. Precisely, NATs can mask specific sites leading to mRNA degradation, such as miRNA40. We found a high proportion of positive correlation between NATs and STs from ssRNA-seq and qPCR data. To explore putative mechanism of NATs in acting as a regulator in S. miltiorrhiza, we identified the STs having miRNA-binding sites in the region overlapped with NATs using psRNATarget41. A total of seven STs potentially to be cleaved were identified, namely, ST0003, ST0005, ST0009, ST0010, ST0018, ST0019, and ST0025 (Table 2). Besides, all the seven SAT pairs had a positive correlation with a correlation coefficient of more than 0.85 (Fig. 4C, E, I, J, R, S, Y).
Identification of pri-miRNAs originating from NATs
MiRNAs can regulate gene expression by direct cleavaging complementary transcripts to regulate the metabolite biosynthesis in plants42,43. We searched the NATs against miRNA sequences found in S. miltiorrhiza44,45 to identify the pri-miRNAs originating from NATs. Two miRNAs, namely, miR156a and miR7208, were aligned to the NAT0112 and NAT0086, respectively (Fig. 6A, B). We then analyzed the secondary structure of the region aligned to miRNAs in NATs, showing the secondary structure meets the criteria of plant miRNA annotation46. The miR156a originated from the overlapped region of the SAT. In a previous study, miR156 could target SQUAMOSA promoter binding protein likes (SPLs) transcript in S. miltiorrhiza47. And the miR7208 did not originate from the SAT's overlapped region, and its function was unknown.
Gene structures of two SAT pairs and the predicted hairpin structures of pre-miRNA. (A) and (B) panels showed information about SAT0112 and SAT0086, respectively. Each panel had two parts, gene structures (above) and predicted hairpin structure (below). In gene structure, the gray and black bars represent exons of NAT and ST, respectively. The lines between bars represent introns. Arrows indicate the transcription direction. The length of the overlapping region was marked above the structure. In the part of the predicted hairpin structure, the miRNA was marked by a red line. The gene structures were drawn using GSDS v2.062 and the hairpin structures of pre-miRNA were drawn using RNAstructure v6.0.163.
Discussion
NATs have been identified in many plant species and are important regulators of various biological processes12. Understanding the regulatory mechanism of NATs is important for the quality improvement of S. miltiorrhiza. A total of 812 NATs, including 168 cis-NATs and 644 trans-NATs, were identified in S. miltiorrhiza in this study through genome-wide analysis.
The general features of NATs from S. miltiorrhiza are similar to those found in other plants. However, the number of identified NATs was lower. The following reasons may cause it. Firstly, we defined NATs as transcripts complementary to protein-coding genes, so we only retain the SATs whose STs are protein-coding genes. Secondly, some NATs are polyadenylated, and some are not. The ratio of these two types of NATs varies in different species17,48. In this study, we identified only the polyadenylated NATs. Thirdly, NATs usually have lower expression levels. For instance, NATs were more than ten folds lower than their STs on average in some organisms49. We might not identify the NATs with very low expression levels. As a result, it is possible that performing polyA (-) RNA-seq will reveal additional NATs in S. miltiorrhiza.
To predict the function of NATs identified in S. miltiorrhiza, we performed GO and KEGG pathway enrichment analyses on the STs with cis-NATs. These STs were mapped to a broad range of terms from biological processes, cellular components, and molecular functions. Some STs were mapped to multiple pathways. For example, ST0117 was mapped to four pathways: beta-alanine metabolism, carbon metabolism, propanoate metabolism, valine, leucine, and isoleucine degradation. ST0087 was mapped to three pathways: purine metabolism, pyrimidine metabolism, and RNA polymerase I.
In contrast, multiple STs were mapped to the same pathway. For example, six STs, namely, ST0137, ST0023, ST0135, ST0076, ST0141, and ST0002, were all mapped to the biosynthesis of secondary metabolites pathway. Two STs, namely, ST0023, ST0135, were mapped to the phenylpropanoid biosynthesis pathway. These results suggest that NATs may play a comprehensive role in the biological processes in S. miltiorrhiza.
Several NATs are found to associate with genes involved in important pathways. SmKSL1 is a key enzyme involved in the biosynthesis of tanshinones. It catalyzes diterpenoid precursor (E,E,E)-geranylgeranyl diphosphate (GGPP) to miltiradiene23. The cis-NAT (NAT0001) was identified to pair with SmKSL1. SmKSL1 and NAT0001 showed a high level of co-expression with a correlation coefficient of 0.99. Both SmKSL1 and NAT0001 were highly expressed in roots. The result was consistent with the accumulation of tanshinones in the roots of S. miltiorrhiza. As a result, NAT0001 might represent a novel mechanism in the regulation of tanshinone biosynthesis by NATs.
Besides, NAT0004 is a cis-NAT paired with ST0004, which encode a candidate UDP-glucosyltransferase catalyzing the transfer of sugars to various receptors to form the glycosidic bond37. Secondary metabolites, such as flavonoids, are important receptors of this reaction36. This enzymatic reaction leads to the glycosylation of metabolites and is a key step in the biosynthesis of various bioactive compounds50. Like SmKSL1 and NAT0001, NAT0004 and ST0004 also showed a high co-expression pattern with a correlation coefficient of 0.99. As a result, NAT0004 might represent a novel mechanism in regulating flavonoids and glycosylation by NATs in S. miltiorrhiza.
NATs can regulate the expression of their STs at one or multiple steps of the gene expression process, such as pre-transcription, transcription, and post-transcription51. The regulation of gene expression by NATs can be either positive or negative51. We found 17 of the 25 SAT pairs had a significantly positive correlation. These NATs can positively regulate the expression of STs by the following two mechanisms. In the pre-transcription step, positive expression correlation may result from transcriptional activation. For example, NATs can enhance H3K4me3 deposition to their STs’ promoter, subsequently activate transcription initiation of STs17. In the post-transcription step, NATs can positively regulate the expression of their STs by limiting mRNA degradation. For example, NATs can mask specific sites that lead to mRNA degradation through binding with their STs, increasing the ST mRNA stability16.
In contrast, we found three pairs of ST and NAT had negative correlation coefficients. These NATs can negatively regulate the expression of their STs by the following mechanisms. In the pre-transcription step, NATs can regulate transcription initiation by affecting histone modifications52, DNA methylation53, and mediating chromatin modifications54. In the transcription step, NATs can negatively regulate their STs’ expression by co-transcription, resulting in transcriptional interference. For example, a DNA region is transcribed in both directions simultaneously. The two RNA polymerases on the opposite strand collide, causing transcription to pause55 simultaneously-transcription step, negative expression correlation may result from translational repression. For example, NAT can bind to the 5′ end of the NAT mRNA resulting in the binding site of the 30S ribosomal subunit, thereby inhibiting NAT translation56. The next step is to validate the function of NAT0001 and NAT0004 and explore the potential molecular mechanisms.
Materials and methods
Sample collection
Two-year-old S. miltiorrhiza (line 99-3) plants were grown under natural growth conditions in the garden of the Institute of Medicinal Plant Development, Beijing, China (Geospatial coordinates: N 40° 2′ 0" E 116° 16′ 5′).Flower (f), leaf (l) and root (r), samples were collected from three individual plants (named as p01, p02 and p03) and stored immediately in liquid nitrogen. We kept these samples in the − 80° refrigerator until use.
RNA extraction, library construction, and ssRNA sequencing
We divided these samples into two groups. In one group, the replicate samples were mixed by tissue type, named as p00_f, p00_l and p00_r respectively. In the other group, we extract RNAs from each of the samples (p01_f, p02_f, p03_f, p01_l, p02_l, p03_l, p01_r, p02_r. p03_r). We extracted RNAs from in total 12 RNA samples using the RNA extraction kit (TianGen, China). NanoDrop 2000 spectrophotometer (Thermo Scientific, USA) and Agilent 2100 Bioanalyzer (Agilent Technologies, USA) were used to detect the quantity and quality of total RNA, respectively. Then mRNA was isolated from total RNA using OligoTex mRNA midi kit (Qiagen) and cleaved into small fragments with 300 bp to 400 bp in length. The library construction was carried out using the ScriptSeq mRNA-Seq Library Preparation Kit (Illumina-compatible) according to the manufacturer’s recommendation. Sequencing was carried out using the Hiseq 2500 platform (Illumina, America). The raw sequence data have been submitted to GenBank under the Bioproject number: PRJNA683774.
Prediction of NATs
To globally identify NATs in S. miltiorrhiza, we developed a stringent selection pipeline that contains three steps (Fig. 1).
In step 1, RNA-seq data of each tissue were mapped to the reference S. miltiorrhiza genome25 using TopHat in the Tuxedo packages (v2.1.1)57. Mapped reads were assembled into transcripts and then annotated based on the reference genome using Cufflinks in the Tuxedo packages (v2.2.1)57. We merged the reassembled transcripts from three tissues into a single unified transcriptome.
In step 2, we searched the reassembled transcripts against protein-coding transcripts based on the annotation of reference genome using the BLASTN (v2.7.1) with an E-value cutoff of 1e-5. The BLASTN results were further analyzed using the Perl script in NATpipe58. We consider the transcripts meeting one of the following two conditions as candidate NATs: (1) the complementary region is longer than 50% of either transcript of the NAT pair; (2) the consecutive complementary region of the NAT pair should be 100 bp or longer. We selected the transcripts that complement transcripts annotated as genes and located in the same loci with the gene as cis-NATs. By contrast, we considered the transcripts located in different loci with the gene as trans-NATs. Then, we predicted RNA hybridization for trans-NATs using the RNAplex (v2.4.14). The results from BLASTN and RNAplex analyses were evaluated using the Perl script in NATpipe58. The transcripts meet the following two conditions were considered trans-NATs: (1) annealed region coincident with the complementary region and is longer than 80% of the complementary region, (2) any bubble within the annealed region shorter than 10% of the total region.
In step 3, we removed the transcripts with a length shorter than 200 bp and those with the potential to encode proteins. We considered the transcripts meet one of the following three conditions to have the protein-encoding potential: (1) transcripts with the maximum length of coding protein more than 100 aa, (2) transcripts matching the Swissprot or Pfam database, and (3) transcripts with Coding Potential Calculator (CPC) score more than 059. The transcripts with the length of amino acid greater than 100 that is the length of the open reading frame (ORF) greater than 300, were identified using the getorf program in EMBOSS package (v 6.5.7.0) with the following parameter: “-minsize 300 -find 1 -reverse N”. We searched these ORFs against the Swissprot database using the BLASTN (v2.7.1) with an e-value cutoff of 1e-5. Next, we searched the ORFs against the Pfam database using the PfamScan.pl (v1.6) with an e-value cutoff of 1e-5.
Conservation of NATs identified in S. miltiorrhiza
To determine the conservation of NATs identified in S. miltiorrhiza, we searched the PlncRNADB (http://bis.zju.edu.cn/PlncRNADB/index.php) and GreeNC34 databases for homologs using BLASTN with the following parameters: a word size 7, an e value cutoff of 10–5, and 50% alignment coverage of both sequences.
Functional enrichment analysis of STs
The sequences of STs having cis-NATs were extracted using the gffread module in Tuxedo packages (v2.1.1)57 and then annotated on the Metascape web server60. We further analyzed the results related to the GO and KEGG in detail35. We consider the terms with a q value < 0.05, a minimum count of 3, and an enrichment factor ≥ 1.5 as significantly enriched. The q value is the multi-test adjusted p value calculated using the Banjamini–Hochberg method. The enrichment factor is the ratio between the observed counts and the counts expected by chance.
Differential expression analysis of NATs
We analyzed the expression of NATs using the Cuffdiff in the Tuxedo packages (v2.2.1)57. The NATs were considered differentially expressed under the two conditions: \(\left| {{\text{log2}}\left( {\text{fold change}} \right)} \right| \, \ge {\text{ 1 and q }} \le \, 0.0{5}.\)
Strand-specific quantitative real-time PCR analysis
To validate the expression of NATs and their cognate STs in each tissue, we collected samples from root, stem, and leaf and extracted RNA using the RNA extraction kit (TianGen, China). We used the strand-specific quantitative real-time PCR (ssRT-qPCR) method61 to validate the expression level. The gene SmActin was used as internal control, as described previously29. First, the sequences of NATs or STs were extracted using the gffread module in Tuxedo packages (v2.1.1)57. Then, the primers were designed on the Integrated DNA Technologies (IDT) webserver. Nucleotide tags were added to the primers for reverse transcription to distinguish NATs and STs. The tag sequence for NATs was “GCTAGCTTCAGCTAGGCATC,” and that for STs was “CCAGATCGTTCGAGTCGT.” The primer sequences were listed in Table S8, Sfile 1. RNA was reversely transcribed to cDNA using strand-specific primers and the TransScript kit (TianGen, China). We carried qPCR with three technical replicates for each cDNA using the qPCR kit (Takara, Japan). We used approximately 1 µl cDNA, 1ul 10 µM each of the forward and reverse primer, 0.4 µl passive reference dye (50×) polymerase, 10 µl 2 × TransStart Top Green qPCR SuperMix and 6.6 µl ddH2O for qPCR with the following conditions: 94 °C for 30 s; 35 cycles of 94 °C for 5 s, and 60 °C for 34 s.
We used the average values from three technical replicates to calculate the Pearson correlation coefficients of expression level between ssRNA-Seq and ssRT-qPCR. The false discovery rate (q value) was calculated using R’s fdrtool package (v3.60).
Correlation of the expression profiles of NATs and their cognate STs
For ssRNA-Seq data, we calculated the Pearson correlation coefficients of expression level between NATs and their cognate STs based on each transcript’s FPKM values. For ssRT-qPCR data, we calculated the Pearson correlation coefficients based on the values of Ct[X] − Ct[ACT], in which ‘ACT’ stands for SmActin gene and ‘X’ stands for transcripts tested in this study. The false discovery rate (q value) was calculated using R’s fdrtool package (v3.60).
References
Katayama, S., Tomaru, Y., Kasukawa, T., Waki, K. & Wahlestedt, C. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566. https://doi.org/10.1126/science.1112009 (2005).
Lavorgna, G. et al. In search of antisense. Trends Biochem. Sci. 29, 88–94. https://doi.org/10.1016/j.tibs.2003.12.002 (2004).
Chen, J., Miao, S., Kent, W. J., Huang, X. & Rowley, J. D. Over 20% of human transcripts might form sense-antisense pairs. Nucl. Acids Res. 32, 4812–4820. https://doi.org/10.1093/nar/gkh818 (2004).
Barrell, B. G., Air, G. M. & Hutchison, C. A. 3rd. Overlapping genes in bacteriophage phiX174. Nature 264, 34–41. https://doi.org/10.1038/264034a0 (1976).
Tomizawa, J., Itoh, T., Selzer, G. & Som, T. Inhibition of ColE1 RNA primer formation by a plasmid-specified small RNA. Proc. Natl. Acad. Sci. USA. 78, 1421–1425. https://doi.org/10.1073/pnas.78.3.1421 (1981).
Williams, T. & Fried, M. A mouse locus at which transcription from both DNA strands produces mRNAs complementary at their 3’ ends. Nature 322, 275–279. https://doi.org/10.1038/322275a0 (1986).
Wang, X. J., Gaasterland, T. & Chua, N. H. Genome-wide prediction and identification of cis-natural antisense transcripts in Arabidopsis thaliana. Genome Biol. 6, R30. https://doi.org/10.1186/gb-2005-6-4-r30 (2005).
Lu, T. et al. Strand-specific RNA-seq reveals widespread occurrence of novel cis-natural antisense transcripts in rice. BMC Genom. 13, 721. https://doi.org/10.1186/1471-2164-13-721 (2012).
Xu, J. et al. Natural antisense transcripts are significantly involved in regulation of drought stress in maize. Nucl. Acids Res. 45, 5126–5141. https://doi.org/10.1093/nar/gkx085 (2017).
Lembke, C. G., Nishiyama, M. Y. Jr., Sato, P. M., de Andrade, R. F. & Souza, G. M. Identification of sense and antisense transcripts regulated by drought in sugarcane. Plant Mol. Biol. 79, 461–477. https://doi.org/10.1007/s11103-012-9922-1 (2012).
Zubko, E. & Meyer, P. A natural antisense transcript of the Petunia hybrida Sho gene suggests a role for an antisense mechanism in cytokinin regulation. Plant J. Cell Mol. Biol. 52, 1131–1139. https://doi.org/10.1111/j.1365-313X.2007.03309.x (2007).
Faghihi, M. A. & Wahlestedt, C. Regulatory roles of natural antisense transcripts. Nat. Rev. Mol. Cell Biol. 10, 637–643. https://doi.org/10.1038/nrm2738 (2009).
Prescott, E. M. & Proudfoot, N. J. Transcriptional collision between convergent genes in budding yeast. Proc. Natl. Acad. Sci. USA. 99, 8796–8801. https://doi.org/10.1073/pnas.132270899 (2002).
Bernstein, E. & Allis, C. D. RNA meets chromatin. Genes Dev. 19, 1635–1655. https://doi.org/10.1101/gad.1324305 (2005).
Hastings, M. L., Milcarek, C., Martincic, K., Peterson, M. L. & Munroe, S. H. Expression of the thyroid hormone receptor gene, erbAalpha, in B lymphocytes: alternative mRNA processing is independent of differentiation but correlates with antisense RNA levels. Nucl. Acids Res. 25, 4296–4300. https://doi.org/10.1093/nar/25.21.4296 (1997).
Faghihi, M. A. et al. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat. Med. 14, 723–730. https://doi.org/10.1038/nm1784 (2008).
Zhao, X. et al. Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA. Nat. Commun. 9, 5056. https://doi.org/10.1038/s41467-018-07500-7 (2018).
Liu, X. et al. A novel antisense long noncoding RNA, TWISTED LEAF, maintains leaf blade flattening by regulating its associated sense R2R3-MYB gene in rice. New Phytol. 218, 774–788. https://doi.org/10.1111/nph.15023 (2018).
Henriques, R. et al. The antiphasic regulatory module comprising CDF5 and its antisense RNA FLORE links the circadian clock to photoperiodic flowering. New Phytol. 216, 854–867. https://doi.org/10.1111/nph.14703 (2017).
Chang, C. C. et al. The anti-atherosclerotic effect of tanshinone IIA is associated with the inhibition of TNF-α-induced VCAM-1, ICAM-1 and CX3CL1 expression. Phytomed. Int. J. Phytother. Phytopharmacol. 21, 207–216. https://doi.org/10.1016/j.phymed.2013.09.012 (2014).
Tao, S. et al. Tanshinone I activates the Nrf2-dependent antioxidant response and protects against As(III)-induced lung inflammation in vitro and in vivo. Antioxid. Redox Signal. 19, 1647–1661. https://doi.org/10.1089/ars.2012.5117 (2013).
Zeng, H. et al. Comparative analysis of the major chemical constituents in Salvia miltiorrhiza roots, stems, leaves and flowers during different growth periods by UPLC-TQ-MS/MS and HPLC-ELSD methods. Molecules (Basel, Switzerland) https://doi.org/10.3390/molecules22050771 (2017).
Gao, W. et al. A functional genomics approach to tanshinone biosynthesis provides stereochemical insights. Org. Lett. 11, 5170–5173. https://doi.org/10.1021/ol902051v (2009).
Guo, J. et al. CYP76AH1 catalyzes turnover of miltiradiene in tanshinones biosynthesis and enables heterologous production of ferruginol in yeasts. Proc. Natl. Acad. Sci. USA. 110, 12108–12113. https://doi.org/10.1073/pnas.1218061110 (2013).
Xu, H. et al. Analysis of the genome sequence of the medicinal plant Salvia miltiorrhiza. Mol. Plant 9, 949–952. https://doi.org/10.1016/j.molp.2016.03.010 (2016).
Zhang, G. et al. Hybrid de novo genome assembly of the Chinese herbal plant danshen (Salvia miltiorrhiza Bunge). Gigascience 4, 62. https://doi.org/10.1186/s13742-015-0104-3 (2015).
Xu, Z. et al. Global identification of the full-length transcripts and alternative splicing related to phenolic acid biosynthetic genes in Salvia miltiorrhiza. Front. Plant Sci. 7, 100. https://doi.org/10.3389/fpls.2016.00100 (2016).
Xu, Z. et al. Full-length transcriptome sequences and splice variants obtained by a combination of sequencing platforms applied to different root tissues of Salvia miltiorrhiza and tanshinone biosynthesis. Plant J. Cell Mol. Biol. 82, 951–961. https://doi.org/10.1111/tpj.12865 (2015).
Chen, H., Wu, B., Nelson, D. R., Wu, K. & Liu, C. Computational identification and systematic classification of novel cytochrome P450 genes in Salvia miltiorrhiza. PLoS ONE 9, e115149. https://doi.org/10.1371/journal.pone.0115149 (2014).
Li, C., Li, D., Li, J., Shao, F. & Lu, S. Characterization of the polyphenol oxidase gene family reveals a novel microRNA involved in posttranscriptional regulation of PPOs in Salvia miltiorrhiza. Sci. Rep. 7, 44622. https://doi.org/10.1038/srep44622 (2017).
Zhou, H., Li, C., Qiu, X. & Lu, S. Systematic analysis of alkaline/neutral invertase genes reveals the involvement of Smi-miR399 in regulation of SmNINV3 and SmNINV4 in Salvia miltiorrhiza. Plants (Basel, Switzerland) https://doi.org/10.3390/plants8110490 (2019).
Shao, F., Qiu, D. & Lu, S. Comparative analysis of the Dicer-like gene family reveals loss of miR162 target site in SmDCL1 from Salvia miltiorrhiza. Sci. Rep. 5, 9891. https://doi.org/10.1038/srep09891 (2015).
Shao, J. et al. Genome-wide identification and characterization of natural antisense transcripts by strand-specific RNA sequencing in ganoderma lucidum. Sci. Rep. 7, 5711. https://doi.org/10.1038/s41598-017-04303-6 (2017).
Paytuví Gallart, A., Hermoso Pulido, A., Anzar Martínez de Lagrán, I., Sanseverino, W. & Aiese Cigliano, R. GREENC: a Wiki-based database of plant lncRNAs. Nucl. Acids Res. 44, 1161–1166. https://doi.org/10.1093/nar/gkv1215 (2015).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucl. Acids Res. 44, D457-462. https://doi.org/10.1093/nar/gkv1070 (2016).
Tiwari, P., Sangwan, R. S. & Sangwan, N. S. Plant secondary metabolism linked glycosyltransferases: an update on expanding knowledge and scopes. Biotechnol. Adv. 34, 714–739. https://doi.org/10.1016/j.biotechadv.2016.03.006 (2016).
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555. https://doi.org/10.1146/annurev.biochem.76.061005.092322 (2008).
Griesser, M. et al. Multi-substrate flavonol O-glucosyltransferases from strawberry (Fragaria×ananassa) achene and receptacle. J. Exp. Bot. 59, 2611–2625. https://doi.org/10.1093/jxb/ern117 (2008).
Escamilla-Treviño, L. L. et al. Arabidopsis thaliana beta-Glucosidases BGLU45 and BGLU46 hydrolyse monolignol glucosides. Phytochemistry 67, 1651–1660. https://doi.org/10.1016/j.phytochem.2006.05.022 (2006).
Yuan, J.-H. et al. The MBNL3 splicing factor promotes hepatocellular carcinoma by increasing PXN expression through the alternative splicing of lncRNA-PXN-AS1. Nat. Cell Biol. 19, 820–832. https://doi.org/10.1038/ncb3538 (2017).
Dai, X., Zhuang, Z. & Zhao, P. X. Computational analysis of miRNA targets in plants: current status and challenges. Brief. Bioinform. 12, 115–121. https://doi.org/10.1093/bib/bbq065 (2011).
Jones-Rhoades, M. W., Bartel, D. P. & Bartel, B. MicroRNAS and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218 (2006).
Tirumalai, V., Swetha, C., Nair, A., Pandit, A. & Shivaprasad, P. V. miR828 and miR858 regulate VvMYB114 to promote anthocyanin and flavonol accumulation in grapes. J. Exp. Bot. 70, 4775–4792. https://doi.org/10.1093/jxb/erz264 (2019).
Xu, X. et al. Deep sequencing identifies tissue-specific microRNAs and their target genes involving in the biosynthesis of tanshinones in Salvia miltiorrhiza. PLoS ONE 9, e111679. https://doi.org/10.1371/journal.pone.0111679 (2014).
Zhang, H. et al. Identification and characterization of Salvia miltiorrhizain miRNAs in response to replanting disease. PLoS ONE 11, e0159905. https://doi.org/10.1371/journal.pone.0159905 (2016).
Meyers, B. C. et al. Criteria for annotation of plant MicroRNAs. Plant Cell 20, 3186–3190. https://doi.org/10.1105/tpc.108.064311 (2008).
Zhang, L. et al. Genome-wide analysis and molecular dissection of the SPL gene family in Salvia miltiorrhiza. J. Integr. Plant Biol. 56, 38–50. https://doi.org/10.1111/jipb.12111 (2014).
Dutrow, N. et al. Dynamic transcriptome of Schizosaccharomyces pombe shown by RNA-DNA hybrid mapping. Nat. Genet. 40, 977–986. https://doi.org/10.1038/ng.196 (2008).
Ozsolak, F. et al. Comprehensive polyadenylation site maps in yeast and human reveal pervasive alternative polyadenylation. Cell 143, 1018–1029. https://doi.org/10.1016/j.cell.2010.11.020 (2010).
Saito, K. et al. The flavonoid biosynthetic pathway in Arabidopsis: structural and genetic diversity. Plant Physiol. Biochem. 72, 21–34. https://doi.org/10.1016/j.plaphy.2013.02.001 (2013).
Pelechano, V. & Steinmetz, L. M. Gene regulation by antisense transcription. Nat. Rev. Genet. 14, 880–893. https://doi.org/10.1038/nrg3594 (2013).
Yap, K. L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674. https://doi.org/10.1016/j.molcel.2010.03.021 (2010).
Latos, P. A. et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 338, 1469–1472. https://doi.org/10.1126/science.1228110 (2012).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076. https://doi.org/10.1038/nature08975 (2010).
Palmer, A. C., Ahlgren-Berg, A., Egan, J. B., Dodd, I. B. & Shearwin, K. E. Potent transcriptional interference by pausing of RNA polymerases over a downstream promoter. Mol. Cell 34, 545–555. https://doi.org/10.1016/j.molcel.2009.04.018 (2009).
Kawano, M., Aravind, L. & Storz, G. An antisense RNA controls synthesis of an SOS-induced toxin evolved from an antitoxin. Mol. Microbiol. 64, 738–754. https://doi.org/10.1111/j.1365-2958.2007.05688.x (2007).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515. https://doi.org/10.1038/nbt.1621 (2010).
Yu, D., Meng, Y., Zuo, Z., Xue, J. & Wang, H. NATpipe: an integrative pipeline for systematical discovery of natural antisense transcripts (NATs) and phase-distributed nat-siRNAs from de novo assembled transcriptomes. Sci. Rep. 6, 21666. https://doi.org/10.1038/srep21666 (2016).
Kang, Y.-J. et al. CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucl. Acids Res. 45, W12–W16. https://doi.org/10.1093/nar/gkx428 (2017).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523. https://doi.org/10.1038/s41467-019-09234-6 (2019).
Kawakami, E. et al. Strand-specific real-time RT-PCR for distinguishing influenza vRNA, cRNA, and mRNA. J. Virol. Methods 173, 1–6. https://doi.org/10.1016/j.jviromet.2010.12.014 (2011).
Hu, B. et al. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31, 1296–1297. https://doi.org/10.1093/bioinformatics/btu817(2015) (2015).
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinform. 11, 129. https://doi.org/10.1186/1471-2105-11-129 (2010).
Acknowledgements
This work was supported by the Chinese Academy of Medical Sciences Innovation Funds for Medical Sciences (CIFMS) [2016-I2M-3-016, 2017-I2M-1-013], the National Science Foundation Funds [81872966], and the National Science &Technology Fundamental Resources Investigation Program of China [2018FY100705], Qinghai Tibet Plateau phytochemistry Key Laboratory (2020-ZJ-Y20). The funders were not involved in the study design, data collection and analysis, decision to be published, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
C.L. conceived the study. M.J. performed data analysis and NATs validation and drafted the manuscript. H.M.C. collected S. miltiorrhiza, extracted RNA for next-generation sequencing. J.T.L. performed qPCR validation experiments; S.L. reviewed the manuscript critically. Q.D. designed and revised the manuscript. All authors have read and agreed on the contents of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, M., Chen, H., Liu, J. et al. Genome-wide identification and functional characterization of natural antisense transcripts in Salvia miltiorrhiza. Sci Rep 11, 4769 (2021). https://doi.org/10.1038/s41598-021-83520-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83520-6
This article is cited by
-
Genome-wide view and characterization of natural antisense transcripts in Cannabis Sativa L
Plant Molecular Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.