Abstract
Coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has spread worldwide as a pandemic throughout 2020. Since the virus uses angiotensin-converting enzyme 2 (ACE2) as a receptor for cellular entry, increment of ACE2 would lead to an increased risk of SARS-CoV-2 infection. At the same time, an association of the ABO blood group system with COVID-19 has also been highlighted: there is increasing evidence to suggest that non-O individuals are at higher risk of severe COVID-19 than O individuals. These findings imply that simultaneous suppression of ACE2 and ABO would be a promising approach for prevention or treatment of COVID-19. Notably, we have previously clarified that histone deacetylase inhibitors (HDACIs) are able to suppress ABO expression in vitro. Against this background, we further evaluated the effect of HDACIs on cultured epithelial cell lines, and found that HDACIs suppress both ACE2 and ABO expression simultaneously. Furthermore, the amount of ACE2 protein was shown to be decreased by one of the clinically-used HDACIs, panobinostat, which has been reported to reduce B-antigens on cell surfaces. On the basis of these findings, we conclude that panobinostat could have the potential to serve as a preventive drug against COVID-19.
Similar content being viewed by others
Introduction
Since the first patient was recognized in December 2019, coronavirus disease 2019 (COVID-19) has become a worldwide pandemic1, with more than 58.9 million cases and 1.3 million deaths as of late November 2020, according to the WHO online dashboard (https://covid19.who.int/). The disease is caused by novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which uses angiotensin-converting enzyme 2 (ACE2) as a receptor for cellular entry2,3. After binding of the SARS-CoV-2 spike (S) protein to ACE2, host proteases, principally transmembrane serine protease 2 (TMPRSS2), promote cellular entry of the virus3,4. These events are likely to occur in specific subsets of epithelial cells of the respiratory and gastrointestinal tracts, which express both ACE2 and TMPRSS24,5. Interestingly, ACE2 is suggested to be an interferon-stimulated gene and thus upregulated during inflammation, resulting in enhanced SARS-CoV-2 infection4. In addition, ACE2 is also considered to affect the pathophysiological process of multiple organ damage including acute lung injury6. These findings imply that increased expression of ACE2 would increase the risk of COVID-197,8, whereas reduction of ACE2 might be a promising therapeutic approach for COVID-199,10. However, no established method for reduction of ACE2 to prevent or relieve COVID-19 has been reported.
One of the other factors related to the risk of COVID-19 is the ABO blood group system11,12. The ABO system is composed of complex carbohydrate structures that are biosynthesized by A and B transferase encoded by the A and B alleles on the ABO gene, respectively13. While A- or B-antigens were originally identified on human red blood cells, they can also be expressed on epithelial cells of the respiratory and gastrointestinal tracts14. Although the precise mechanisms are still being investigated, accumulating reports suggest that individuals with the A blood group type are at increased risk for symptoms related to SARS-CoV-2 infection, such as acute respiratory syndrome and cardiovascular diseases, as well as severe outcomes including intubation and death15,16,17. Furthermore, a recent genome-wide association study has clarified that the 9q34.2 locus, which coincides with the ABO locus, is one of the two loci that are most significantly associated with severe COVID-19 with respiratory failure, representing a higher risk for blood group A individuals18. Studies so far suggest that O individuals are at lower risk from COVID-19 than non-O individuals. Taken together, these findings suggest that the ABO system could be another druggable target for alleviation of COVID-19 risk, i.e. reduction of A- and B-antigens might reduce the risks of COVID-19.
Previously, we had clarified that clinically used histone deacetylase inhibitors (HDACIs) suppress ABO expression in vitro, leading to a decrease of B-antigens on the surface of KATOIII cells, a gastric cancer cell line19. Therefore, we hypothesized that HDACIs could potentially serve as drugs to prevent severe outcomes of COVID-19. Against this background, in the present study, we further investigated the effect of HDACIs on the expression of ABO, as well as that of ACE2 and TMPRSS2, in cultured epithelial cell lines, to determine whether HDACIs could have a potential preventive effect against COVID-19. Given the fact that development of novel vaccines for SARS-CoV-2 would likely take considerable time20,21, our findings suggest that HDACIs would warrant clinical investigation to further evaluate their potential usage in this context.
Results
HDACIs such as sodium butyrate and panobinostat suppress ACE2 expression in KATOIII cells
We have previously reported that HDACIs such as sodium butyrate and panobinostat suppress ABO expression in the gastric cancer cell line KATOIII19. To examine whether the HDACIs also decrease ACE2 expression, we performed quantitative real-time PCR (qPCR) on KATOII cells treated with or without 1 mM sodium butyrate or 25 nM panobinostat for 6 or 24 h, targeting ABO, ACE2 and TMPRSS2 transcripts as well as β-actin (ACTB) as endogenous control. As we had shown before, these HDACIs suppressed ABO expression (Fig. 1A). In addition, it was also clarified that they suppressed the expression of ACE2 in a time-dependent manner (Fig. 1B). On the other hand, the HDACIs did not suppress TMPRSS2 (Fig. 1C), suggesting that the HDACI-related suppression was gene-specific.
ABO, ACE2 and TMPRSS2 expression in KATOIII cells cultured with or without sodium butyrate or panobinostat for 6 or 24 h. (A–C) Relative amounts of ABO (A), ACE2 (B) and TMPRSS2 (C) transcripts in KATOIII cells treated with or without HDACIs such as sodium butyrate and panobinostat. Clear bars indicate the basal expression levels in the absence of HDACIs, the gray bars represent the relative amounts of transcripts in the presence of 1 mM sodium butyrate, and the solid bars denote those in the presence of 25 nM panobinostat. The graphs express mean fold values relative to those without HDACI. In each panel, the left-hand bars show the results obtained 6 h after incubation with or without HDACIs, while the right-hand bars show data 24 h after treatment. The asterisks represent a significant reduction compared to the values without HDACI (p < 0.05).
Various HDACIs suppress ACE2 expression in KATOIII cells
To further evaluate the effect of HDACIs on ABO, ACE2 and TMPRSS2 expression, KATOIII cells were treated with several concentrations of various HDACIs including sodium valproate, vorinostat and trichostatin A for 24 h, and the relative amounts of these transcripts were evaluated by qPCR (Fig. 2A–O). In addition, the cell proliferations and viabilities were evaluated in each condition to reveal potential cytotoxicity of the HDACIs (Fig. 2P–T). As expected, sodium butyrate and panobinostat reduced the amount of both ABO and ACE2 transcripts simultaneously in a dose-dependent manner. In addition, sodium valproate and vorinostat caused similar suppression of both ABO and ACE2, except that a low concentration (0.5 mM) of sodium valproate increased the amount of ACE2 (Fig. 2H). On the other hand, trichostatin A rarely reduced the ABO and ACE2 transcripts (Fig. 2M,N), while moderately reducing the cell proliferations and viabilities (Fig. 2T). Those observations suggested the considerable cytotoxicity of trichostatin A. None of the HDACIs had a suppressive effect on TMPRSS2, except for a high concentration (50 nM) of panobinostat, which suppressed TMPRSS2 (Fig. 2F). On this basis, we concluded that the HDACIs used in the present study, except for trichostatin A, had the potential to suppress ABO and ACE2 concurrently on KATOIII cells, whereas such suppression was rarely observed for the TMPRSS2 transcript.
Alterations in the expression of ABO, ACE2 and TMPRSS2, cell proliferation, and viability by treatment of KATOIII cells with HDACIs. (A–O) Each panel represents the relative amount of ABO (A, D, G, J and M), ACE2 (B, E, H, K and N) or TMPRSS2 (C, F, I, L and O) transcripts in KATOIII cells incubated with or without various concentrations of sodium butyrate (A–C), panobinostat (D–F), sodium valproate (G–I), vorinostat (J–L) or trichostatin A (M–O) for 24 h. The amounts are expressed as mean fold values relative to those without HDACI. The asterisks represent a significant reduction compared to the values without HDACI (p < 0.05). (P–T) Each panel represents both the proliferation and viability of KATOIII cells incubated with or without various concentrations of HDACIs for 24 h. The bars indicate cell proliferation as the number of live cells relative to that without HDACI treatment on the left y-axes, while the dots and lines denote the absolute cell viability on the right y-axes.
Suppression of ABO and ACE2 by HDACIs on other cell lines
Next, we sought to evaluate the HDACI-related suppression of ABO and ACE2 on other cell lines. We first quantified the basal expression of ABO, ACE2 and TMPRSS2 on several cell lines that were available in our laboratory (Table 1). As a result, we found that, in addition to KATOIII cells, the gastric cancer cell line NUGC-4 also expressed considerable amounts of ABO, ACE2 and TMPRSS2 transcripts, while all the other cell lines including a lung derived cell line, HMVEC–L, expressed an insufficient amount of at least one of the three transcripts. Therefore, the NUGC-4 cell line was deemed relevant for further evaluation of the HDACI-related suppression of ABO and ACE2. Against this background, we performed similar qPCR experiments on NUGC-4 cells incubated with or without various concentrations of sodium butyrate or panobinostat (Fig. 3), resulting in similar suppression of ABO and ACE2 by the HDACIs, while TMPRSS2 was not suppressed. Thus, it was clarified that HDACIs such as sodium butyrate and panobinostat had the ability to suppress both ABO and ACE2 simultaneously in several epithelial cell lines.
Suppression of ABO and ACE2 in NUGC-4 cells treated with sodium butyrate or panobinostat for 24 h. (A–C) Relative amounts of ABO (A), ACE2 (B) and TMPRSS2 (C) transcripts in NUGC-4 cells treated with various concentrations of sodium butyrate for 24 h. (D–F) Relative amounts of ABO (D), ACE2 (E) and TMPRSS2 (F) transcripts in NUGC-4 cells treated with various concentrations of panobinostat for 24 h. The transcript amounts are expressed as mean fold values relative to those without HDACI. The asterisks represent a significant reduction compared to the untreated values (p < 0.05).
HDACIs decrease the ACE2 protein in cell lysates of KATOIII and NUGC-4
Whether or not the HDACI-related suppression of ACE2 would lead to a reduced amount of ACE2 protein in cultured cell lines was considered an intriguing issue. Notably, we had shown previously that panobinostat reduced the amount of B-antigens on KATOIII cells. Therefore, among the various HDACIs, we decided to focus on panobinostat, considering that it might serve as a preventive drug against COVID-19 by simultaneously diminishing A- or B-antigens and ACE2 proteins on the cell surface.
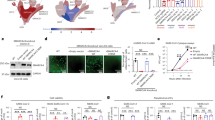
To this end, we performed enzyme-linked immunosorbent assays (ELISA) using cell lysates prepared from KATOIII and NUGC-4 cells incubated with 0, 25 or 50 nM panobinostat for 24 or 48 h (Fig. 4). For KATOIII cells, we confirmed that the amount of ACE2 was reduced at 48 h after incubation with 25 nM and 50 nM panobinostat (Fig. 4A). For NUGC-4 cells, ACE2 reduction was observed after both 24- and 48-h treatment with 25 nM and 50 nM panobinostat (Fig. 4B). There was no significant difference in the amount of ACE2 between the two concentrations of panobinostat. Furthermore, we carried out additional Western blot analysis of whole-cell lysates prepared from KATOIII and NUGC-4 cells cultured under the same conditions as those for the above ELISA. For KATOIII cells, however, we rarely detected ACE2 in the lysates even though no panobinostat was added (Fig. 4C,D), probably because of the relatively low level of ACE2 expression in KATOIII cells (Table 1). On the other hand, for NUGC-4 cells, it was clarified that panobinostat at both 25 nM and 50 nM decreased the amount of ACE2 after treatment for both 24 and 48 h (Fig. 4E–G). In conclusion, panobinostat was able to reduce the amount of ACE2 in cultured epithelial cells.
Reduction of the ACE2 amount in KATOIII and NUGC-4 cells treated with panobinostat. (A, B) Relative amounts of ACE2 in KATOIII (A) or NUGC-4 (B) cells incubated with 25 nM or 50 nM panobinostat for 24 or 48 h, analyzed by ELISA. The clear bars indicate the basal amount of ACE2 in the absence of panobinostat, the gray bars represent the relative amounts the ACE2 in the presence of 25 nM panobinostat, and the solid bars denote those in the presence of 50 nM panobinostat. The graphs express the mean fold values relative to those without HDACI. In each panel, the left-hand bars show the results obtained 24 h after incubation with or without panobinostat, while the right-hand bars show the data 48 h after treatment. The asterisks represent a significant reduction compared to the untreated values (p < 0.05). (C–F) Western blots for cell lysates derived from KATOIII (C, D) and NUGC-4 (E, F) cells treated with 0, 25 or 50 nM panobinostat for 24 or 48 h. Panels (C) and (E) represent ACE2 while panels (D) and (F) indicate actin as a loading control. The amount of protein applied to each lane was 100 or 40 μg for the KATOIII or NUGC-4 assays, respectively. In panel (C), no ACE2 is illustrated because of its low expression. The arrows in the panels (D) and (F) indicate actin with a band size of 42 kDa. The arrow in panel (E) represents ACE2 with a band size of 120 kDa. (G) Relative densitometric levels of ACE2 in NUGC-4 lysates divided by the amounts of actin. The clear bars indicate the basal amount of ACE2 in the absence of panobinostat, the gray bars represent the relative amounts of ACE2 in the presence of 25 nM panobinostat, and the solid bars denote those in the presence of 50 nM panobinostat. The left-hand bars show the results obtained 24 h after incubation with or without panobinostat, while the right-hand bars show the data 48 h after treatment.
Discussion
We revealed that several HDACIs suppressed ABO and ACE2 transcripts concurrently in KATOIII or NUGC-4 cells, while TMPRSS2 expression was rarely repressed. Notably, among the HDACIs used in the present study, panobinostat caused drastic suppression at the lowest concentrations, whereas trichostatin A barely reduced those transcripts probably because of its cytotoxicity. Finally, panobinostat reduced the amount of ACE2 proteins in both KATOIII and NUGC-4 cells. Considering together the findings that panobinostat decreases B-antigen on the KATOIII cells19, that non-O individuals have a higher risk of COVID-1918 and that higher expression of ACE2 is a risk factor for COVID-197,8,9,10, it seems plausible that panobinostat could have the potential to serve as a preventive drug against COVID-19.
Currently, the association of panobinostat with ACE2 expression is also being investigated by two other research groups22,23. Those two groups adopted very similar approaches; they re-analyzed the same publically available data, comprising gene expression profiles for thousands of perturbagens at a variety of time points, doses, and cell lines24 in order to identify drugs that could significantly modify ACE2 expression. Although their interpretations of the results differed, to our surprise, both of their analyses suggested that panobinostat might up-regulate ACE2 expression, contrary to our findings. Although the reason for this contradiction is unclear, we speculate that it might be attributable to differences in experimental conditions between our data and those publically available. For example, the former group, He and Garmire, analyzed expression profiles in the presence of 10 µM panobinostat, which was more than 200 times higher than the concentration we employed22. In addition, neither KATOIII nor NUGC-4 were featured in the publically available data24. As shown in Table 1, few cell lines seem to express a sufficient amount of ACE2, and thus most cultured cells are unsuitable for investigating regulation of the gene4,25. Therefore, the effects of HDACIs on gene expression in vitro need to be evaluated carefully. On the other hand, Xu et al. recently reported a novel approach whereby conventional molecular docking was computationally accelerated in combination with generative artificial intelligence, resulting in the identification of potential drug-repurposing candidates for COVID-1926. Surprisingly, though their approach was completely different from ours, panobinostat was identified as one of the six candidate drugs. Considering this accumulating evidence to suggest the preventive potential of panobinostat against COVID-19, further evaluations, including clinical trials, seem warranted. Notably, the in vitro potencies of valproic acid, panobinostat and vorinostat as therapeutics for various cancers were reported to be in the mM, µM and nM order, respectively27, whereas our data revealed that panobinostat might have preventive potential against COVID-19 at concentrations as low as the nM order. Although future in vivo or clinical studies will be needed to clarify the possible tolerable doses of HDACIs for potential prevention of COVID-19, our present data suggest that panobinostat in particular could be employed clinically at especially low doses, leading to less adverse effects.
In some experiments, we observed slight up-regulation of TMPRSS2 in KATOIII cells when the cells were treated with 1 mM sodium butyrate for 6 h (Fig. 1C) or 1 mM sodium valproate for 24 h (Fig. 2I). In addition, the NUGC-4 cells showed an apparent increase in TMPRSS2 expression in the presence of HDACIs such as sodium butyrate and panobinostat (Fig. 3C,F). While these results suggest potential activation of the host protease TMPRSS2 by HDACIs, this might be irrelevant to the risk of viral infection considering that TMPRSS2 is involved in the process of viral entry only after the virus has bound to ACE2 on epithelial cells3,4. Whether HDACIs actually inhibit the cellular entry of the virus will need to be clarified by future experiments using live virus.
An increasing number of reports have suggested an association between the ABO system and COVID-1911,12,14,15,16,17,18. This is not surprising, since involvement of the ABO system in viral infection, including SARS, has been documented28, although the precise mechanism remains unclear. It is controversial whether the association of the ABO system with COVID-19 could be attributable to the amount of A- or B-antigens on cells or anti-A or -B antibodies in serum11,12,14. Interestingly, Ladikou et al. have reported that patients with severe COVID-19 who developed venous thromboembolism had highly elevated levels of von Willebrand factor (vWF) and coagulation factor VIIIc29, whose serum levels are correlated with the ABO system and are higher in non-O individuals30,31. Considering that the presence of A- or B-antigens in vWF N-linked oligosaccharides plays a role in vWF levels32,33, reduction of the A- or B-antigen might be a reasonable approach for reducing plasma vWF levels as well as the risk of thrombopoietic symptoms of COVID-19. Notably, the ABO system is also associated with a number of other factors, including ACE plasma activity34 and interleukin levels35,36. Thus, the influence of the ABO system seems to be complex, highly diversified and much more significant than has been clarified33. Further investigations focusing on the ABO blood group system should help to reveal the hidden roles of this system that could significantly impact human health, disease and biology.
Materials and methods
Cell culture with or without HDACIs
The cell lines KATOIII (JCRB0611), K562 (JCRB0019), NUGC-4 (JCRB0834) and MKN1 (JCRB0252) were originally purchased from Japanese Collection of Research Bioresources Cell Bank (Osaka, Japan), SW480 (ATCC CCL-228), SV-HUC (ATCC CRL-9520), HMVEC-L (ATCC CC-2527) and SH-SY5Y (ATCC CRL-2266) were from American Type Culture Collection (Manassas, VA), 5637 (TKG 0605), KK47 (TKG 0663) and T24 (TKG 0443) were from Cell Resource Center for Biomedical Research (Miyagi, Japan) and LAN-5 (RCB0485) was from RIKEN BioResource Research Center (Ibaraki, Japan). The KATOIII and K562 cells were cultured as described previously19. The NUGC-4, MKN1, SW480, 5637, KK47 and LAN-5 cell lines were cultured in RPMI1640 medium containing 10% FCS, 100 U/ml penicillin and 100 µg/ml streptomycin. The culture medium for SV-HUC, T24, HMVEC-L or SH-SY5Y was Ham’s F12K, MEM, EGM-2MV Microvascular Endothelial Cell Growth Medium-2 or 1:1 mixture of MEM and F12, each with 10% FCS, 100 U/ml penicillin and 100 µg/ml streptomycin. For treatment with the HDACIs, the cells were seeded at a density of 2.5 × 105/ml one day before the experiment. On the following day, the cells were re-seeded at a density of 2.5 × 105/ml in new medium with or without HDACIs. The medium was not changed thereafter until harvest of the cells. The HDMCIs we used included sodium butyrate (#303410; Sigma-Akdrich), panobinostat (#13280; Cayman Chemical Company), sodium valproate (#13033; Cayman Chemical Company), vorinostat (#10009929; Cayman Chemical Company) and trichostatin A (#89730; Cayman Chemical Company). The solvent used for sodium butyrate and sodium valproate was deionized-distilled water, and that used for panobinostat, vorinostat and trichostatin A was dimethyl sulfoxide.
KATOIII cell proliferation and viability were evaluated by both manual and automatic counting of live or dead cells after trypan blue staining in each condition where the cells were treated with various concentrations of each HDACI for 24 h. The automatic counting was performed with a Countess II FL Automated Cell Counter (#AMQAF1000, Invitrogen). Every count was conducted at least twice.
Quantitative real-time PCR (qPCR)
RNA purification, cDNA preparation and quantification of ABO and ACTB transcripts were performed as described previously19. qPCR of the ACE2 and TMPRSS2 transcripts was performed with the specific primer sets “ACE2 Primer 2” and “TMPRSS2 Primer 2”, respectively, as described by Ma et al.37, under the following conditions: 95 °C for 3 min and 40 cycles at 95 °C for 3 s and at 60 °C for 30 s. Every assay was conducted at least twice, and the absolute amount of each transcript determined by qPCR was standardized by the amount of ACTB transcript.
Enzyme-linked immunosorbent assay (ELISA)
ELISA was performed using a Human ACE2 ELISA Kit (#ab235649, Abcam) following manufacturer’s instructions. Briefly, KAKTOIII or NUGC-4 cells were harvested 24 or 48 h after incubation with or without HDACIs, and solubilized in 1 × cell extraction buffer PTR. After centrifugation, the concentration of total protein in the supernatant was measured using a DC protein assay kit (#5000112JA, Bio-Rad), and 250 or 100 ng of total protein derived from KATOIII or NUGC-4 cell lysates, respectively, was applied to each well of a ready-to-use microplate provided in the kit. Then the ACE2 antibody cocktail was added to each well, followed by 1-h incubation at RT on a plate shaker set to 400 rpm. After the incubation, each well was rinsed three times, TMB Development Solution was added, and incubation was performed at RT on a plate shaker. Ten minutes later, the Stop Solution was added to each well, and the end point reading of OD at 450 nm was recorded using a iMark microplate absorbance reader (#168-1135, Bio-Rad). The concentration of the ACE2 protein in the sample was determined by interpolating the blank control subtracted absorbance values against the standard curve. Every assay was conducted in duplicate.
Western blotting
Whole-cell lysates were prepared from KATOIII and NUGC-4 cells incubated with 0, 25 or 50 nM panobinostat for 24 or 48 h. One hundred or 40 µg of total protein was applied to each lane for the KATOIII or NUGC-4 assays, respectively. The transferred membrane was reacted with a recombinant rabbit anti-ACE2 monoclonal antibody (#ab108252, Abcam), followed by treatment with anti-rabbit immunoglobulins/HRP (#P0448, DakoCytomation) and Amersham ECL Prime Western Blotting Detection Reagent (#RPM2232, GE Healthcare). Then, densitometry measurements were conducted with a LAS-3000 and MultiGauge v3.0 (FujiFilm, Tokyo, Japan). Subsequently, the same membrane was rinsed, processed with blocking buffer, reacted with mouse anti-β-actin monoclonal antibody (#017-24551, Wako) and anti-mouse IgG (H + L)-HRP conjugate (#172-1011, BioRad). Detection and densitometry measurements were performed again as described above.
References
Liu, S. L. New virus in China requires international control effort. Nature 577, 472 (2020).
Bao, L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature 583, 830–833 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271-280.e8 (2020).
Ziegler, C. G. K. et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell 181, 1016-1035.e19 (2020).
Xiao, F. et al. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 158, 1831-1833.e3 (2020).
Liu, M. et al. Potential role of ACE2 in coronavirus disease 2019 (COVID-19) prevention and management. J. Transl. Int. Med. 8, 9–19 (2020).
Fang, L., Karakiulakis, G. & Roth, M. Are patients with hypertension and diabetes mellitus at increased risk for COVID-19 infection?. Lancet Respir. Med. 8, e21 (2020).
Li, C. & Xu, B. H. The viral, epidemiologic, clinical characteristics and potential therapy options for COVID-19: a review. Eur. Rev. Med. Pharmacol. Sci. 24, 4576–4584 (2020).
Gue, Y. X. & Gorog, D. A. Reduction in ACE2 may mediate the prothrombotic phenotype in COVID-19. Eur. Heart J. 41, 3198–3199 (2020).
Mostafa-Hedeab, G. ACE2 as drug target of COVID-19 virus treatment, simplified updated review. Rep. Biochem. Mol. Biol. 9, 97–105 (2020).
Li, J. et al. Association between ABO blood groups and risk of SARS-CoV-2 pneumonia. Br. J. Haematol. 190, 24–27 (2020).
Gérard, C., Maggipinto, G. & Minon, J. M. COVID-19 and ABO blood group: another viewpoint. Br. J. Haematol. 190, e93–e94 (2020).
Schenkel-Brunnner, H. Syntax of referencing. in Human Blood Groups, 2nd edn. 47–97 (Springer-Verlag Wien, New York, 2000).
Yamamoto, F. ABO Blood Groups and SARS-CoV-2/COVID-19 Infection (version 3) by Fumiichiro Yamamoto. ResearchGate presentation. https://www.researchgate.net/publication/341576873_ABO_Blood_Groups_and_SARS-CoV-2COVID-19_Infection_version_2_by_Fumiichiro_Yamamoto. https://doi.org/10.13140/RG.2.2.19766.52802.
Wu, Y., Feng, Z., Li, P. & Yu, Q. Relationship between ABO blood group distribution and clinical characteristics in patients with COVID-19. Clin. Chim. Acta 509, 220–223 (2020).
Zietz, M. & Tatonetti, N. P. Testing the association between blood type and COVID-19 infection, intubation, and death. Preprint at https://www.medrxiv.org/content/https://doi.org/10.1101/2020.04.08.20058073v2 (2020).
Dai, X. ABO blood group predisposes to COVID-19 severity and cardiovascular diseases. Eur. J. Prev. Cardiol. 27, 1436–1437 (2020).
Severe Covid-19 GWAS Group; Ellinghaus, D. et al. Genomewide association study of severe covid-19 with respiratory failure. N. Engl. J. Med. 383, 1522–1534 (2020).
Takahashi, Y. et al. Histone deacetylase inhibitors suppress ABO transcription in vitro, leading to reduced expression of the antigens. Transfusion 57, 554–562 (2017).
Amanat, F. & Krammer, F. SARS-CoV-2 vaccines: status report. Immunity 52, 583–589 (2020).
Le Thanh, T. et al. The COVID-19 vaccine development landscape. Nat. Rev. Drug Discov. 19, 305–306 (2020).
He, B. & Garmire, L. Prediction of repurposed drugs for treating lung injury in COVID-19 [version 2; peer review: 2 approved]. F1000Research 9, 609 (2020).
Sinha, S., Cheng, K., Aldape, K., Schiff, E. & Ruppin, E. Systematic cell line-based identification of drugs modifying ACE2 expression. Preprint at https://www.preprints.org/manuscript/202003.0446/v1 (2020).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437-1452.e17 (2017).
Letko, M., Marzi, A. & Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 5, 562–569 (2020).
Xu, Z., Wauchope, O & Frank, A. T. Navigating chemical space by interfacing generative artificial intelligence and molecular docking. Preprint at https://www.biorxiv.org/content/https://doi.org/10.1101/2020.06.09.143289v1 (2020).
Mottamal, M., Zheng, S., Huang, T. L. & Wang, G. Histone deacetylase inhibitors in clinical studies as templates for new anticancer agents. Molecules 20, 3898–3941 (2015).
Cheng, Y. et al. ABO blood group and susceptibility to severe acute respiratory syndrome. JAMA 293, 1450–1451 (2005).
Ladikou, E.E. et al. Von Willebrand factor (vWF): marker of endothelial damage and thrombotic risk in COVID-19? Clin. Med. (Lond). clinmed.2020–0346 (2020).
Gill, J. C., Endres-Brooks, J., Bauer, P. J., Marks, W. J. Jr. & Montgomery, R. R. The effect of ABO blood group on the diagnosis of von Willebrand disease. Blood 69, 1691–1695 (1987).
Albánez, S. et al. Aging and ABO blood type influence von Willebrand factor and factor VIII levels through interrelated mechanisms. J. Thromb. Haemost. 14, 953–963 (2016).
Matsui, T., Titani, K. & Mizuochi, T. Structures of the asparagine-linked oligosaccharide chains of human von Willebrand factor. Occurrence of blood group A, B, and H(O) structures. J. Biol. Chem. 267, 8723–8731 (1992).
Liumbruno, G. M. & Franchini, M. Beyond immunohaematology: the role of the ABO blood group in human diseases. Blood Transfus. 11, 491–499 (2013).
Gassó, P., Ritter, M. A., Mas, S. & Lafuente, A. Influence of ABO genotype and phenotype on angiotensin-converting enzyme plasma activity. J. Renin Angiotensin Aldosterone Syst. 15, 580–584 (2014).
Naitza, S. et al. A genome-wide association scan on the levels of markers of inflammation in Sardinians reveals associations that underpin its complex regulation. PLoS Genet. 8, e1002480 (2012).
Johansson, Å., Alfredsson, J., Eriksson, N., Wallentin, L. & Siegbahn, A. Genome-wide association study identifies that the ABO Blood group system influences interleukin-10 levels and the risk of clinical events in patients with acute coronary syndrome. PLoS ONE 10, e0142518 (2015).
Ma, D. et al. Expression of SARS-CoV-2 receptor ACE2 and TMPRSS2 in human primary conjunctival and pterygium cell lines and in mouse cornea. Eye (Lond) 34, 1212–1219 (2020).
Acknowledgements
We thank Tomoko Miwa for preparation of this manuscript.
Funding
This work was supported in part by JSPS KAKENHI Grant Numbers 19K10678 to Y.T., 20K18977 to A.H., 20K10551 to R.S., 18K10118 to R.K. and 19H03916 to Y.K.
Author information
Authors and Affiliations
Contributions
Y.T. designed and performed the experiments, analyzed the data and wrote the manuscript. A.H., R.S., H.F., M.H., R.K. and T.O. performed the experiments. Y.K. designed the experiments and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takahashi, Y., Hayakawa, A., Sano, R. et al. Histone deacetylase inhibitors suppress ACE2 and ABO simultaneously, suggesting a preventive potential against COVID-19. Sci Rep 11, 3379 (2021). https://doi.org/10.1038/s41598-021-82970-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82970-2
This article is cited by
-
FragDPI: a novel drug-protein interaction prediction model based on fragment understanding and unified coding
Frontiers of Computer Science (2023)
-
Epigenetic perspectives associated with COVID-19 infection and related cytokine storm: an updated review
Infection (2023)
-
Epigenetic targeting of the ACE2 and NRP1 viral receptors limits SARS-CoV-2 infectivity
Clinical Epigenetics (2021)
-
Cellular host factors for SARS-CoV-2 infection
Nature Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.