Abstract
The increase of human population and associated increasing demand for agricultural products lead to soil over-exploitation. Biofertilizers based on lyophilized plant material containing living plant growth-promoting microorganisms (PGPM) could be an alternative to conventional fertilizers that fits into sustainable agricultural technologies ideas. We aimed to: (1) assess the diversity of endophytic bacteria in sugar and sea beet roots and (2) determine the influence of osmoprotectants (trehalose and ectoine) addition during lyophilization on bacterial density, viability and salt tolerance. Microbiome diversity was assessed based on 16S rRNA amplicons sequencing, bacterial density and salt tolerance was evaluated in cultures, while bacterial viability was calculated by using fluorescence microscopy and flow cytometry. Here we show that plant genotype shapes its endophytic microbiome diversity and determines rhizosphere soil properties. Sea beet endophytic microbiome, consisting of genera characteristic for extreme environments, is more diverse and salt resistant than its crop relative. Supplementing osmoprotectants during root tissue lyophilization exerts a positive effect on bacterial community salt stress tolerance, viability and density. Trehalose improves the above-mentioned parameters more effectively than ectoine, moreover its use is economically advantageous, thus it may be used to formulate improved biofertilizers.
Similar content being viewed by others
Introduction
Conventional agriculture practices negatively affect environment, e.g. by decreasing microbial diversity, soil quality, water supply and plant productivity1,2. Wide adoption of sustainable agricultural technologies, e.g. biofertilizers, may significantly decrease the use of chemical fertilizers, reducing negative consequences of agriculture on the environment2,3.
Biofertilizers are based on living plant growth-promoting microorganisms (arbuscular mycorrhizal fungi—AMF, plant growth-promoting rhizobacteria—PGPR, nitrogen fixing bacteria—NFB) and are key players in sustainable agriculture4. They can promote plant growth in several different ways (e.g. increasing availability of nutrients, synthesizing phytohormones or siderophores, fixing nitrogen), especially under unfavorable environmental conditions (e.g. drought or salinity)5,6,7. Most of commercially available biofertilizers are based on combination of two or more microbial beneficial strains, which is called ‘consortium’4. Compared to single strains, consortia display increased spectrum of beneficial effect of inoculum on plants. However, criteria of strain selection are the crucial factor influencing the inoculum success and should be considered not only based on plant genotype compatibility but also environmental factors.
The methods of biofertilizers production, storage and application are diverse. Inoculation techniques are based on microorganisms application in liquid (sprays and drenches) or solid form (lyophilizates delivered to soil/growth substrate). The most important problem in the preparation and storage technology of biofertilizers is maintaining high viability of microorganisms8. Lyophilization is well known and widely used technique extending microbial cell viability9. To alleviate negative effect of low temperature and desiccation on microorganisms in this technology, several different stabilizers can be used e.g. nonreducing disaccharides, glicerolglycerol or skim milk10.
Trehalose (α-d-glucopyranosyl-(1 → 1)-α-d-glucopyranoside) is a disaccharide present in almost all prokaryotic and eukaryotic organisms and exhibits high efficiency in protection of cells against low temperature, drying and osmotic stress10,11,12. Ectoine (1,4,5,6-tetrahydro-2-methyl-4-pyrimidinecarboxylic acid) is synthesized mostly by halotolerant and halophilic microorganisms and responsible for regulation of osmotic pressure in microbial cells, increasing their tolerance to osmotic stress (salinity)13,14,15,16. Application of trehalose and ectoine in the process of lyophilization of endophytic microbiomes was tested in our work for the first time.
Biofertilizer efficiency analyzed under laboratory conditions may not correspond to results obtained under field conditions2,17. This effect may be due to adverse effect of environmental conditions or autochthonic microorganisms on gene expression in microbial cells2 or low competitiveness of microorganisms used as biofertilizers, i.e. they may be outgrown by the autochthonic ones. This is why “plant microbiome” was proposed as the new generation of inoculants18. Inoculation of crops with microbiome and organic matter present in lyophilized plant roots seems to be a better solution to enrich microbial biodiversity of soil and crops with new endophytes.
Endophytes are bacteria and fungi that colonize the internal plants tissues without causing pathogenic symptoms19 and can directly (nitrogen fixation, phosphate solubilization, siderophore and phytohormone synthesis) and/or indirectly (biocontrol agents) promote plant growth and development, e.g. in crops20.
Recent data show that more than 7% of global land surface and 70% of all irrigated agricultural soils worldwide is affected by salinity21, and the problem is exacerbated by inorganic fertilization as well as by climate changes. Moreover, halophytes thriving in naturally saline environments are reservoirs of endophytes possessing high tolerance to salt stress22,23 that may be useful in alleviation of salt stress in crops. Application of halotolerant microbes in sustainable agriculture e.g. in the increasing salinity tolerance of non-halophytic crops, is well known and was extensively studied6,24,25,26,27.
Cultivated beets are one of the few crops whose direct ancestor (sea beet, Beta vulgaris ssp. maritima) still grows in the wild. This feature enables comparison of traits in plants that are very close genetically (ca. 0.5% difference28), but whose ecology differs considerably. Moreover, as sea beet is as a halophyte growing in nature29, it seems to be a good candidate for a source of microorganisms that could be useful for crop beets improvement. We analyzed both rhizosphere soil and plant roots to determine if the plant influence on the former is strong enough to make it the source of microbes for bioinoculant formulation.
The goal of our study was twofold: (i) to characterize rhizosphere and root microbiomes of cultivated and wild beet and choose the source of microorganisms for prospective bioinoculant formulation for growing beet in saline soils and (ii) to check if addition of osmoprotectants during lyophilization changes root bacterial community structure as well as microbiome salinity tolerance and viability. Specifically, we formulated two general hypotheses: (i) roots of different beet genotypes would filter rhizosphere microbes specifically and therefore they would be better source of microbes for bioinoculation than rhizosphere soils, (ii) addition of osmoprotectants would allow formulation of a better inoculant because of increased bacterial viability and microbiome salinity tolerance.
Results
Rhizosphere as a source of microbes in endosphere
Rhizosphere soil physicochemical parameters are different for sugar and sea beet
Majority of tested parameters was higher in sugar beet soil, but only in cases of CaCO3 and Na+ the difference was statistically significant. On the other hand, OC, P, Ca2+, Mg2+ and Nt were higher in sea beet soil and for the latter the difference was significant (Table 1). To check if these changed parameters influenced bacterial communities in rhizosphere soils, we sequenced 16S rRNA amplicon libraries.
Bacterial diversity in sugar beet roots is lower than in its wild ancestor
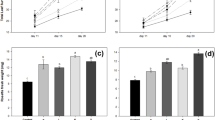
Bacterial diversity, evenness and species richness were the highest in 16S rRNA libraries coming from rhizosphere soil, regardless of plant genotype. Lyophilized sea beet roots harbored more diverse community than sugar beet (Fig. 1). The number of OTUs was ca. three times higher in the wild beet than in the crop (Fig. 1A,B), while the diversity was around 1.5 times higher (Fig. 1C), and evenness was ca. 1.3 times greater (Fig. 1D).
Species richness, evenness and diversity of bacterial communities in rhizosphere soils of sugar beet (Bh) and sea beet (Bm) and lyophilized roots of these plants untreated (C), and treated with ectoine (E) or trehalose (T). Means (n = 8–32) are presented, whiskers show standard error of the mean (SEM), and significant differences (ANOVA, p < 0.05) are denoted with different letters. Observed number of OTUs (A), estimated total number of OTUs (Chao1 index, B), Shannon's diversity index (H', C), Shannon's evenness (D).
Both endophytic and rhizosphere soil bacterial community is dominated by Proteobacteria
There were no significant differences in taxonomic composition of rhizosphere soil bacterial communities of sugar- and sea beet at the level of phylum (Fig. 2A). At the level of genus, three taxa were differentially represented, all of them belonging to Alphaproteobacteria: two Rhizobiales-belonging genera, Pedomicrobium and an unknown genus of JG34.KF.361 family as well as Woodsholea (Caulobacteraceae) were more abundant in the crop (Fig. 2C). Differences in lyophilized roots communities were more pronounced, although still there were no taxa significantly differentially represented between osmolyte treatments. At the level of phyla Proteobacteria-derived reads were more abundant in libraries from sugar beet lyophilized roots, while Actinobacteria, Bacteroidetes, Acidobacteria, Verrucomicrobia and rare phyla were more abundant in its wild ancestor (Fig. 2B). Among genera significant differences were observed for Stenotrophomonas and Bacillus that were more abundant in the crop and for proteobacterial genera Novosphingobium, Devosia (Alphaproteobacteria), Hydrogenophaga, Polaromonas (Betaproteobacteria), Rhizobacter and Tahibacter (Gammaproteobacteria) as well as for rare and unclassified genera being more abundant in sea beet (Fig. 2D).
Taxonomic composition of bacteria communities in rhizosphere soils of sugar beet (Bh) and sea beet (Bm) (A, C) and lyophilized roots of these plants (B, D) untreated (Bh_C and Bm_C) and treated with ectoine (Bh_E, Bm_E) or trehalose (Bh_T, Bm_T) at the phylum (A, B) and genus (C, D) levels. Means (n = 8–32) are presented, and significant differences between genotypes are marked either with m and h letters (panels A and C, significant differences between rhizosphere and endosphere in sea (m) or sugar (h) beet) or with asterisks (panels B and D, significant differences between genotypes, no differences due to osmolytes were found).
At the level of phyla , regardless of plant genotype, there were significant differences between soil and roots in all taxa but Firmicutes. Proteobacteria and Firmicutes were more abundant in roots than in soil, while abundance of the remaining phyla was lower in planta, and Gemmatimonadetes as well as Verrucomicrobia were absent from roots. At the level of genus, regardless of genotype, Pseudomonas and Rhizobium were significantly more abundant in roots than in soil, while Sphingomonas, Pedomicrobium, rare and unclassified bacteria were less frequent in roots than in rhizosphere. Novosphingobium, Pantoea, Hydrogenophaga, Polaromonas, Paenibacillus, Hyphomicrobium and Rhizobacter were significantly more abundant in wild beet roots than in soil, while Stenotrophomonas was the only genus that was more frequent in sugar beet roots than in soil. Variibacter, Chryseolinea, and Woodsholea were less abundant in wild beet roots than in soil, while Devosia and Hirschia were less frequent in sugar beet than in soil.
Effect of osmolytes on diversity, viability, and tolerance to salinity of bacterial communities in lyophilized beet roots
Bacterial cell density in lyophilized roots depends on host genotype but not on osmolyte
In total, 72 bacterial strains were isolated and identified, 35 coming from sugar beet and 37 from sea beet. Proteobacteria were the most frequent phylum in fresh roots of both sugar and sea beet, followed by Actinobacteria in the crop and Firmicutes in the wild plant. Pseudomonas and Sphingomonas were characteristic for fresh roots of sugar beet, while Bosea and Sphingopyxis were found exclusively in sea beet roots before lyophilization (Table 2). Density of culturable root endophytic bacteria was higher in sugar beet lyophilizates than in sea beet (ANOVA, p < 0.05, Fig. 3), regardless of the osmolytes addition. We observed no influence of osmolytes on sea beet endophytes density, while trehalose increased slightly, but significantly (ANOVA, p < 0.05) the density in sugar beet samples (Fig. 3).
Density of endophytic bacteria (expressed as log10 CFU per g of dry weight) isolated from lyophilized sugar- and sea beet roots. Means (n = 3) ± standard deviation are presented. Significant differences between variants (ANOVA, p < 0.05, with Tukey's HSD; C–control untreated with any osmolyte, ectoine (E) or trehalose (T) treated) were marked with different letters.
Sea beet endophytes are more salt tolerant than sugar beet ones
Increasing salinity negatively affected growth of culturable fraction of microbiome regardless of origin (sea- vs. sugar beet), however stronger effect was observed for sugar beet. In control treatment the growth was inhibited (final cell density below the critical level of 0.2 OD600) at 200 mM and 300 mM NaCl concentration for sugar and sea beet, respectively. Addition of osmolytes enhanced the growth in general and increased the inhibitory concentration to 400 and 700 mM, respectively (Supplementary Table 1). Influence of both osmolytes was similar, with trehalose performing slightly better at high NaCl concentrations., The effect was greater for sea beet, than for sugar beet (Fig. 4).
Osmolytes effect on growth of LB cultures inoculated with lyophilized sugar and sea beet roots untreated with any osmolyte (C), treated with ectoine (E) and trehalose (T). Means (n = 4–6) ± standard deviation are presented. Significant differences between treatments (ANOVA, p < 0.05, with Tukey's HSD) are marked with asterisks.
Bacterial viable cell density in lyophilized roots is associated with plant genotype and osmolyte
Cell viability in lyophilized beet roots was assessed by means of three, complementary methods: via plate counts, fluorescence microscopy and flow cytometry (Fig. 5). Bacterial viability in sugar beet was consistently higher than in roots of its wild relative, regardless of osmolyte treatment, storage time and measurement methodology. Both trehalose and ectoine increased the viability compared to control, regardless of genotype, but the effect of the former was more pronounced (Fig. 5).
Bacterial viability in lyophilized beet roots. Viability measured with BD Cell Viability kit under fluorescence microscope (AB) and using flow cytometer (C). Means are presented and statistically significant differences between treatments are marked with differing letters. Stars denote significant differences between genotypes.
Discussion
Bacterial diversity in beet rhizosphere
Differences in rhizosphere soil physicochemical properties observed in our study, may be due to greater nutritional demands of the two beet genotypes (TN, Na) or varying exudates composition (OC), as it was found that rhizodeposition is the primary organic carbon source in the rhizosphere30. Alternatively, they might be caused by changes in microbial activity resulting from microbial metabolic activity or interaction between microorganisms31,32.
Greater microbiome diversity in rhizosphere compared to endosphere was commonly observed, and resulted from natural plant selection mechanisms33,34,35. Accordingly, in our study, the higher bacterial diversity, evenness and species richness were noted in rhizosphere soil of both investigated genotypes, than in roots. At the same time, in spite of slightly different TN, OC and Na levels, microbiome composition and diversity were similar in rhizosphere soils of both studied plant genotypes. This observation could be explained by the use of the same starting substrate (garden soil) and short culture period (three months), not allowing the rhizosphere differences to fully manifest. Culture-independent analysis revealed that dominating bacterial phyla were the same as those observed in rhizosphere of many plant species e.g. barley, alfalfa or wheat36,37,38. Only a few differences between the genotypes were noted at the genus level, mainly concerning Alphaproteobacteria. Pedomicrobium as well as JG34.KF.361_ge, more frequent in sugar beet, represent Rhizobiales, an order known for organisms that establish beneficial interactions with plants and comprises numerous bacteria with nitrogen-fixing capability39. The observed lower TN level in the sugar beet rhizosphere may indicate higher demand for nitrogen. Tsurumaru and colleagues40 indicated that Mesorhizobium and Bradyrhizobium, also belonging to Rhizobiales, play an important ecological role in the taproot of sugar beet. Moreover, it was showed that higher levels of nitrogen (N) and potassium (K) significantly affect the growth parameters of sugar beet. Both elements were generally recognized as crucial for obtaining higher yields of this crop, favorably affecting organic metabolites biosynthesis and improving nutritional status41.
Bacterial diversity in beet roots
The higher diversity both in rhizo- and endosphere of the wild plant compared to its crop counterpart was observed42,43. It was hypothesized that beneficial endophytes associated with wild plants were absent or fewer in domesticated crops43. Sugar beet as a cultivated plant grows under more controlled conditions regulated by farmers, while sea beet grows mainly in highly saline and nutrients poor coastal soil28. Growth under adverse environmental conditions requires support of microorganisms with a wide range of beneficial metabolic properties tailored for specific plant needs23. The loss of high tolerance to salt stress during the process of sea beet domestication was demonstrated29 and might be associated with the loss of microbes that increased tolerance of this plant to salinity. Concordantly, despite the lack of differences in rhizosphere soil microbial composition, lower diversity of endophytes in sugar beet compared to its wild ancestor was noted in our study. This difference might be explained by varying root system architecture, with fibrous root system of sea beet providing more opportunities for bacteria to enter the endosphere33, which affects stochastic community assembly. On the other hand microbe selection can be driven by the genetic makeup of two studied subspecies. We observed that sea beet caused decrease in the soil Na level, suggesting accumulation of Na ions in wild plant tissues. Accordingly, there was an increase in community salinity resistance in this plant, which pointed at higher level of halotolerant and halophytic microorganisms.
In general, endophytic microbiome diversity and composition is related to soil properties as well as plant ecology and physiology44. Members of only three phyla (Proteobacteria, Actinobacteria and Firmicutes) were cultured in our experiment, this may be related to their high ability to grow on commercially available media5,6,44,45. It was emphasized that Proteobacteria distinctly predominate among culturable plant endophytes, then the presence of Firmicutes and Actinobacteria is common, and Bacteroidetes occur slightly less frequently44.
16S rRNA gene libraries generated in our study were dominated by the four phyla (Proteobacteria, Actinobacteria, Firmicutes, Bacteroidetes) commonly found in endosphere of glycophytes including maize (Zea mays L.46), Dactylis glomerata L., Festuca rubra L. and Lolium perenne L.47 as well as in halophytes such as Salicornia europaea23 or para grass (Urochloa mutica48). Sea beet was characterized by significantly higher frequency of Actinobacteria, Bacteroidetes, Acidobacteria, Verrucomicrobia and rare phyla compared to sugar beet, where Proteobacteria were observed more often. Zachow et al. observed greater frequency of Actinobacteria, Bacteroidetes and Verrucomicrobia in rhizosphere of wild beet cultivated in coastal soil than in sugar beet rhizosphere42. This fact, together with our results, may point at these bacterial taxa being preferred by sea beet regardless of soil.
Our 16S rRNA gene sequencing results also revealed significantly higher abundance of certain genera in sea beet endosphere, including: Novosphingobium, Devosia (Alphaproteobacteria), Hydrogenophaga, Polaromonas (Betaproteobacteria), Rhizobacter and Tahibacter (Gammaproteobacteria) as well as certain rare and unclassified bacteria. This set of microorganisms comprises extremophiles, e.g. Polaromonas49 or Hydrogenophaga50 and organisms modulating plant stress response, such as Novosphingobium25. In our study, only Stenotrophomonas and Bacillus genera were more frequent in roots of sugar beet than of sea beet. Stenotrophomonas and Pseudomonas sp. were identified in rhizospheric soil of sugar and sea beet, while the former together with Staphylococcus sp. were mainly observed in crop rhizosphere. Sea beet microbiome was found to be more diverse than that of sugar beet, which is explained by greater number of rare taxa. It was found that sugar beet rhizosphere was more frequently colonized by strains with antagonistic activity against plant pathogens and/or stress protection activity, while abiotic stress-releasing ones were more often found in sea beet’s rhizosphere42. These facts together with our results suggest that pre-adaptation to stress observed in sea beet transcriptome51 may also take place at the level of microbiome serving as a helper.
Osmoprotectants enhance bacterial viability and diversity in lyophilized beet roots
Significantly higher cell density of culturable bacteria observed in sugar beet lyophilized roots can be attributed to high content of sucrose. This sugar acts as a natural osmoprotectant, allowing better viability of microorganisms during lyophilization52. Another explanation of obtained results can be associated with higher ability of sugar beet endophytes to grow on solid medium.
Sea beet endophytic microbiome was found to be more resistant to salinity. Microorganisms present in a more saline sea beet tissue most likely developed mechanisms of adaptation to high salt level, which provided them ability to grow in higher NaCl concentrations compared to the sugar beet microbiome. This fact may be related to higher sodium accumulation in this plant tissues51, which caused decrease of soil sodium concentration observed in our study.
Salinity-induced changes in community structure and adverse effects on microbial density, activity, biomass were reported by many scientists53,54. The decrease in number of culturable microorganisms related to increasing NaCl concentration was noted even in the case of endophytes associated with halophytes (Aster tripolium, Salicornia europaea)5,6,55. Obtained results were in line with the above trend, but apart from negative effect of salinity on sugar and sea beet bacterial density, a beneficial impact of trehalose and ectoine on salt stress mitigation was demonstrated. Although ectoine is a major osmolyte in aerobic chemoheterotrophic bacteria and is considered as a marker for halophytic bacteria15, a slightly better effect of trehalose, was confirmed by the results of microscopic analyzes, flow cytometry and culture tests. Protective effect of trehalose is explained by “water replacement hypothesis” that states that the compound lowers the phase transition temperature of membrane phospholipids, by replacement of water molecules occurring around the lipid head groups56, thus protecting membrane structure57. This suggests that the use of trehalose is a better and more economic solution providing high viability of bacterial cells after lyophilization. In the case of sugar beet the above mentioned positive sucrose impact was enhanced by trehalose addition. Similar effect was observed for rhizobial strains, where trehalose worked better than sucrose/peptone mixture58. In general, 16S rRNA gene sequencing results considering diversity of endophytes associated with sea and sugar beet root did not show any effect of applied osmoprotectants neither on alpha nor beta diversity of bacteria. This observation can be explained by the presence of ‘relic DNA’, i.e. DNA coming from non-viable cells59 in lyophilized samples.
Bacillus sp. was the only species identified among the strains representing the Firmicutes phylum isolated from the lyophilized osmolytes-treated roots of both investigated genotypes. In the control variant the presence of Psychrobacillus sp. and Paenibacillus sp. inside sea and sugar beet root was additionally found, respectively. The viability of the above-mentioned bacteria after lyophilization was probably associated with their commonly known ability to form endospores and higher tolerance to environmental changes60,61,62. Actinobacteria proved to be sensitive to lyophilization, while Proteobacteria remarkably well tolerated it, and additional osmolytes promoted the incidence of culturable bacteria belonging to the latter phylum.
Conclusions
Our research revealed that plant genotype played a pivotal role in the shaping of its endophytic microbiome diversity and physicochemical rhizosphere soil properties, affecting soil sodium content, but not soil bacterial community structure. Bacterial diversity was lower in sugar beet roots than in its wild ancestor tissues. At the same time sea beet endophytic microbiome was more salt resistant and consisted of genera characteristic for extreme environments.
Supplementing osmoprotectants during root tissue lyophilization had a positive effect on bacterial salt stress tolerance, viability and density. Trehalose proved to improve these parameters more effectively than ectoine, moreover its use was economically advantageous.
Materials and methods
Experimental design
Sea beet (Beta vulgaris L. subsp. maritima L.) seeds were obtained from National Germplasm Resources Laboratory, Beltsville, MD, USA, while in the case of sugar beet (B. vulgaris subsp. vulgaris cv. 'Huzar') commercial seeds were bought from WHBC Poznań, Poland. Healthy and uniform-sized seeds were placed in 5 l pots filled with 2.5 kg of garden soil. From twenty plants, five representative ones (with two pairs of true leaves and similar in size) were chosen for analysis. Pot experiment was conducted from mid-March through mid-May 2017 in a greenhouse (Nicolaus Copernicus University in Toruń, Poland). Plants were grown under natural lighting conditions and temperature was maintained at 22–24 °C throughout the growth period. All plants were arranged randomly on the green house benches. The plants were watered with tap water every two days, amount depended on the plants demand. After three months plants and rhizosphere soil samples were collected and analyzed as shown in Fig. 6.
Soil analysis
Soil parameters (TOC, TN, CaCO3, Pcitr, pH, EC, Na, K, Ca, Mg, Cl, SO42−) were analyzed as described earlier in Furtado et al.27.
Plant and soil samples preparation
Plants were carefully uprooted, and 10 g of soil adhering to roots (rhizospheric soil) was collected, frozen at − 80 °C and lyophilized before DNA isolation for metagenomic analysis. Roots were washed with tap water to remove soil and were separated from shoots and leaves. Then, they were surface sterilized with 70% ethanol and 15% hydrogen peroxide mixture (1:1 v:v) for 5 min and subsequently rinsed six times with 0.9% NaCl. Efficiency of the sterilization process was evaluated by plating the last rinse on Luria–Bertani (Difco LB Agar, Miller) and potato dextrose extract (Lab A Neogen Company) media. Only properly sterilized plant material was used for subsequent analyzes. Approximately 100 g of fresh root material was homogenized in 100 ml of 0.9% NaCl by using surface sterilized (rinsed with 70% ethanol and UV-irradiated) blender. Homogenates were used to evaluate bacterial density and to prepare lyophilizates.
Roots lyophilization
Homogenized sugar and sea beet roots were used to prepare three variants of lyophilizates including (1) no osmolytes addition (control—C) (2) trehalose (T) and (3) ectoine (E) supplemented. Three biological replicates were prepared for each tested plant species (9 samples per plant species, in total 18 samples were used for downstream analyzes). Either 1 ml of 0.9% NaCl (control) or 1.0 mg of trehalose (Tre) or 1.0 mg of ectoine (Ect) were mixed with 50 g of homogenized roots. The mixtures were lyophilized in Telstar LyuQues (DanLab) until completely dry (approximately 24 h).
Estimation of bacterial density
Serial dilutions were prepared directly from the homogenized fresh roots and lyophilizates re-suspended in 0.9% NaCl (1:9 m:v). The dilutions (10−3 to 10−8) were plated in triplicates on LB plates supplemented with nystatin (Sigma, 100 µg/ml) to prevent fungal growth, and the plates were incubated for 5 days at 26 °C. Colony counts (expressed as CFU per 1 g of fresh or dry weight for homogenates and lyophilizates, respectively) were based on plates with 30–300 colonies. At least six bacterial isolates were purified per experimental variant.
Bacterial viability assessment: fluorescence microscopy and flow cytometry
Ten-miligram samples of ground lyophilized roots were mixed with 10 ml of PBS (pH = 7.4) and incubated for 2 days at 26 °C with mixing. The mixtures were filtered through a 40 µm cell strainer (Biologix) and 2 ml were centrifuged for 3 min at 1000 × g at RT to pellet the residual plant debris. Cells in the supernatant were stained with Cell Viability kit (BectonDickinson) as per the manufacturer's protocol, than bacterial viability was analyzed using fluorescence microscopy (after 6 and 12 months of storage) and flow cytometer (after 12 months storage). Preparations were photographed in red and green channel under 40 × magnification upon fluorescence excitation with 433 nm light on Axiostar plus fluorescence microscope (Zeiss) equipped with Delta Optical camera. Percentage of live cells was based on counts from at least 30 view fields per sample. Flow cytometric analysis was performed on samples stained as described above with FACS Aria III (BectonDickinson) using 488 nm laser for excitation. Fluorescence was collected at 530 ± 30 nm (for thiazole orange—TO) and 616 ± 26 nm (for propidium iodide—PI) bands and seventy-micrometer nozzle was used. Parameters were optimized basing on pure environmental strains and their mixtures analyses and autoclaved lyophilizate samples served as negative controls.
Salt tolerance assessment
Salt tolerance of root bacterial communities was measured as OD600 after 5 days incubation at 26 °C using 96-wells microtiter plate reader (Biolog Micro Station). 140 µl of LB medium supplemented NaCl to obtain final concentrations of 0, 50, 100, 150, 200, 300, 400, 500, 600, 700, 800, 900 mM were used per well. Inoculates were prepared by suspending 2 g of mortar-ground lyophilized roots in 18 ml of 0.9% NaCl and diluting the mixture ten times. The inoculates were filtered through 40 µm cell strainer (Biologix) to remove plant debris. Six test and two control wells were inoculated with 10 µl of filtered inoculate or 0.9% NaCl, respectively.
Isolates identification by 16S rRNA gene sequencing
Genomic DNA was isolated from purified strains using GeneMatrix Bacterial and Yeast Genomic DNA Purification Kit (EurX) according to the manufacturer's protocol with modified homogenization step (FastPrep-24 bead-beater, one cycle of 20 s at 4.0 m/s). The DNA was analyzed spectrophotometrically (NanoDrop 2000). 16S rRNA gene fragment was amplified using 27F and 1492R primers63, following the procedure described in Szymańska et al.6. The products were purified with GeneMatrix PCR/DNA Clean-Up DNA Purification Kit (EurX) according to the manufacturer’s protocol. Sanger sequencing was performed with BrightDye Cycle Sequencing kit (Nimagen), using 40 ng of template DNA, 1.5 pmol of primer and 1 µl of kit and 1.5 µl of BD buffer in 10 µl volume. The reactions were EtOH/NaAc precipitated and read out at IBB PAS, Warsaw, Poland.
16S rRNA gene fragment library construction and sequencing
Metagenomic DNA was isolated and V3-V4 16S rRNA gene fragment libraries for Illumina sequencing were prepared as described earlier64. They were sequenced on Illumina MiSeq using 600 cycles v.3 kit at CMIT NCU.
Statistical analysis and bioinformatics
Bioinformatics analyses of Illumina reads was performed as described earlier64. Briefly, the reads were denoised, merged and chimeras were removed with dada265, then amplicon variant sequences were exported together with abundance information and processed in Mothur v.1.3966: aligned against SILVA v.132 database, screened for those covering the 6428-22400 positions of the alignment, filtered to remove gap-only and terminal gap-containing positions, pre-clustered to remove residual noise and clustered into 0.03 dissimilarity OTUs. Representative OTU sequences were classified using naïve Bayesian classifier67 and SILVA database68. Sanger reads were manually inspected in Chromas to remove obvious errors, the corrected sequences were merged with CAP369, and classified using naïve Bayesian classifier with SILVA v.132 reference files.
Significance of differences between means was assessed with ANOVA test with Tukey's post-hoc analysis implemented in Statistica 10.0 (StatSoft). Normality of data was tested with Shapiro–Wilk's test and homogeneity of variance was assessed with Levene's test. When the assumptions were violated, non-parametric Kruskal–Wallis test with Dunn’s test as a post-hoc analysis was used. Significance level of 0.05 was assumed.
Data availability
Sequences generated during this study were deposited in the SRA repository and are accessible via BioProject no. PRJNA606174.
References
Gveroska, B., Miceska, G., Dimitrieski, M. & Korubin-Aleksoska, A. Use of biopreparates in Tobacco protection: contribution to sustainable agriculture. Türk Tarım ve Doğa Bilim. Derg. 1, 1509–1517 (2014).
Baez-Rogelio, A., Morales-García, Y. E., Quintero-Hernández, V. & Muñoz-Rojas, J. Next generation of microbial inoculants for agriculture and bioremediation. Microb. Biotechnol. 10, 19–21 (2017).
Pretty, J. Agricultural sustainability: concepts, principles and evidence. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 363, 447–465 (2008).
Malusá, E., Sas-Paszt, L. & Ciesielska, J. Technologies for Beneficial Microorganisms Inocula Used as Biofertilizers. Sci. World J. 2012, 1–12 (2012).
Szymańska, S. et al. Metabolic potential and community structure of endophytic and rhizosphere bacteria associated with the roots of the halophyte Aster tripolium L. Microbiol. Res. 182, 68–79 (2016).
Szymańska, S., Płociniczak, T., Piotrowska-Seget, Z. & Hrynkiewicz, K. Endophytic and rhizosphere bacteria associated with the roots of the halophyte Salicornia europaea L. – community structure and metabolic potential. Microbiol. Res. 192, 37–51 (2016).
Hrynkiewicz, K. & Patz, S. Salicornia europaea L. as an underutilized saline-tolerant plant inhabited by endophytic diazotrophs. J. Adv. Res. 19, 49–56 (2019).
Alori, E. T. & Babalola, O. O. Microbial inoculants for improving crop quality and human health in Africa. Front. Microbiol. 9, 2213 (2018).
Prakash, O., Nimonkar, Y. & Shouche, Y. S. Practice and prospects of microbial preservation. FEMS Microbiol. Lett. 339, 1–9 (2013).
Park, J. E., Lee, K. H. & Jahng, D. Effect of trehalose on bioluminescence and viability of freeze-dried bacterial cells. J. Microbiol. Biotechnol. 12, 349–353 (2002).
Reina-Bueno, M. et al. Role of trehalose in salinity and temperature tolerance in the model halophilic bacterium Chromohalobacter salexigens. PLoS ONE 7, e33587 (2012).
Lee, H.-J., Yoon, Y.-S. & Lee, S.-J. Mechanism of neuroprotection by trehalose: controversy surrounding autophagy induction. Cell Death Dis. 9, 712 (2018).
Oren, A. Microbial life at high salt concentrations: phylogenetic and metabolic diversity. Saline Systems 4, 2 (2008).
Han, J. et al. Transcriptomic and ectoine analysis of halotolerant Nocardiopsis gilva YIM 90087T under salt stress. Front. Microbiol. 9, 618 (2018).
Roberts, M. F. Organic compatible solutes of halotolerant and halophilic microorganisms. Saline Systems 1, 5 (2005).
Czech, L. et al. Role of the extremolytes ectoine and hydroxyectoine as stress protectants and nutrients: genetics, phylogenomics, biochemistry, and structural analysis. Genes (Basel). 9, 177 (2018).
Parnell, J. J. et al. From the lab to the farm: an industrial perspective of plant beneficial microorganisms. Front. Plant Sci. 7, 1110 (2016).
Compant, S., Samad, A., Faist, H. & Sessitsch, A. A review on the plant microbiome: ecology, functions, and emerging trends in microbial application. J. Adv. Res. 19, 29–37 (2019).
Hardoim, P. R. & van Elsas, J. D. Properties of Bacterial Endophytes Leading to Maximized Host Fitness. in Molecular Microbial Ecology of the Rhizosphere 405–411 (John Wiley & Sons, Inc., 2013). doi:https://doi.org/10.1002/9781118297674.ch37
Patle, P. et al. Endophytes in plant system: Roles in growth promotion, mechanism and their potentiality in achieving agriculture sustainability. Int. J. Chem. Stud. 6, 270–274 (2018).
Bencherif, K. et al. Impact of soil salinity on arbuscular mycorrhizal fungi biodiversity and microflora biomass associated with Tamarix articulata Vahll rhizosphere in arid and semi-arid Algerian areas. Sci. Total Environ. 533, 488–494 (2015).
Abbas, H., Patel, R. M. & Parekh, V. R. Culturable endophytic bacteria from halotolerant Salvadora persica L.: isolation and plant growth promoting traits. Indian J. Appl. Microbiol. 10, 1074 (2018).
Szymańska, S. et al. Bacterial microbiome of root-associated endophytes of Salicornia europaea in correspondence to different levels of salinity. Environ. Sci. Pollut. Res. 25, 25420–25431 (2018).
Yadav, A. N. & Saxena, A. K. Biodiversity and biotechnological applications of halophilic microbes for sustainable agriculture. J. Appl. Biol. Biotechnol. 6, 48–55 (2018).
Etesami, H. & Beattie, G. A. Mining halophytes for plant growth-promoting halotolerant bacteria to enhance the salinity tolerance of non-halophytic crops. Front. Microbiol. 9, 148 (2018).
Szymańska, S. et al. Boosting the Brassica napus L. tolerance to salinity by the halotolerant strain Pseudomonas stutzeri ISE12. Environ. Exp. Bot. 163, 55–68 (2019).
Furtado, B. U., Gołębiewski, M., Skorupa, M., Hulisz, P. & Hrynkiewicz, K. Bacterial and fungal endophytic microbiomes of Salicornia europaea. Appl. Environ. Microbiol. 85, (2019).
Dohm, J. C. et al. The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature 505, 546–549 (2014).
Rozema, J. et al. Comparing salt tolerance of beet cultivars and their halophytic ancestor: consequences of domestication and breeding programmes. AoB Plants 7, (2014).
Bashir, O. et al. Soil microbe diversity and root exudates as important aspects of rhizosphere ecosystem. in Plant, Soil and Microbes 337–357 (Springer International Publishing, 2016). https://doi.org/10.1007/978-3-319-29573-2_15
Nannipieri, P. et al. Effects of root exudates in microbial diversity and activity in rhizosphere soils. in 339–365 (Springer, Berlin, Heidelberg, 2008). https://doi.org/10.1007/978-3-540-75575-3_14
Shi, S. et al. Effects of selected root exudate components on soil bacterial communities. FEMS Microbiol. Ecol. 77, 600–610 (2011).
Kandel, S., Joubert, P. & Doty, S. Bacterial endophyte colonization and distribution within plants. Microorganisms 5, 77 (2017).
Liu, H. et al. Inner plant values: diversity, colonization and benefits from endophytic bacteria. Front. Microbiol. 8, 2552 (2017).
Cheng, D., Tian, Z., Feng, L., Xu, L. & Wang, H. Diversity analysis of the rhizospheric and endophytic bacterial communities of Senecio vulgaris L. (Asteraceae) in an invasive range. PeerJ 6, e6162 (2019).
Velázquez-Sepúlveda, I., Orozco-Mosqueda, M. C., Prieto-Barajas, C. M. & Santoyo, G. Bacterial diversity associated with the rhizosphere of wheat plants (Triticum aestivum): Toward a metagenomic analysis. Phyton (B. Aires). 81, 81–87 (2012).
Bulgarelli, D. et al. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17, 392–403 (2015).
Kumar, V. et al. Metagenomic analysis of rhizosphere microflora of oil-contaminated soil planted with barley and alfalfa. PLoS ONE 13, e0202127 (2018).
Erlacher, A. et al. Rhizobiales as functional and endosymbiontic members in the lichen symbiosis of Lobaria pulmonaria L. Front. Microbiol. 6, 53 (2015).
Tsurumaru, H. et al. Metagenomic analysis of the bacterial community associated with the taproot of sugar beet. Microbes Environ. 30, 63–69 (2015).
Abdel-Motagally, F. M. F. & Attia, K. K. Response of sugar beet plants to nitrogen and potassium fertilization in sandy calcareous soil. Int. J. Agric. Biol. 11, 695–700 (2009).
Zachow, C., Mueller, H., Tilcher, R. & Berg, G. Differences between the rhizosphere microbiome of Beta vulgaris ssp. maritima-ancestor of all beet crops-and modern sugar beets. Front. Microbiol. 5, 415 (2014).
Ofek-Lalzar, M. et al. Diversity of fungal endophytes in recent and ancient wheat ancestors Triticum dicoccoides and Aegilops sharonensis. FEMS Microbiol. Ecol. 92, fiw152 (2016).
Miliute, I., Buzaite, O., Baniulis, D. & Stanys, V. Bacterial endophytes in agricultural crops and their role in stress tolerance: a review. Zemdirbyste Agricu. 102, 465–478 (2015).
Brígido, C. et al. Diversity and Functionality of Culturable Endophytic Bacterial Communities in Chickpea Plants. Plants (Basel, Switzerland) 8, (2019).
Correa-Galeote, D., Bedmar, E. J. & Arone, G. J. Maize endophytic bacterial diversity as affected by soil cultivation history. Front. Microbiol. 9, 484 (2018).
Wemheuer, F. et al. Bacterial endophyte communities of three agricultural important grass species differ in their response towards management regimes. Sci. Rep. 7, 1–13 (2017).
Mukhtar, S. et al. Microbial diversity and metagenomic analysis of the rhizosphere of para grass (Urochloa mutica) growing under saline conditions. Pakistan J. Bot. 48, 779–791 (2016).
Gawor, J. et al. Evidence of adaptation, niche separation and microevolution within the genus Polaromonas on Arctic and Antarctic glacial surfaces. Extremophiles 20, 403–413 (2016).
Khan, M. & Goel, R. Principles, applications and future aspects of cold-adapted PGPR. in Plant-Bacteria Interactions 195–212 (Wiley-VCH Verlag GmbH & Co. KGaA, 2008). https://doi.org/10.1002/9783527621989.ch10
Skorupa, M. et al. Salt stress vs. salt shock-the case of sugar beet and its halophytic ancestor. BMC Plant Biol. 1–18 (2019).
Bircher, L., Geirnaert, A., Hammes, F., Lacroix, C. & Schwab, C. Effect of cryopreservation and lyophilization on viability and growth of strict anaerobic human gut microbes. Microb. Biotechnol. 11, 721–733 (2018).
Yan, N., Marschner, P., Cao, W., Zuo, C. & Qin, W. Influence of salinity and water content on soil microorganisms. Int. Soil Water Conserv. Res. 3, 316–323 (2015).
Zhang, K. et al. Salinity Is a key determinant for soil microbial communities in a desert ecosystem. mSystems 4, (2019).
Szymańska, S., Piernik, A. & Hrynkiewicz, K. Metabolic potential of microorganisms associated with the halophyte Aster tripolium L. in saline soils. Ecol. Quest. 18, 9–19 (2013).
Berninger, T., González López, Ó., Bejarano, A., Preininger, C. & Sessitsch, A. Maintenance and assessment of cell viability in formulation of non-sporulating bacterial inoculants. Microb. Biotechnol. 11, 277–301 (2018).
Nounjan, N. & Theerakulpisut, P. Effects of Exogenous proline and trehalose on physiological responses in rice seedlings during salt-stress and after recovery. Plant, Soil Environ. 58, 309–315 (2012).
Arraes Pereira, P. A., Oliver, A., Bliss, F. A., Crowe, L. & Crowe, J. Preservation of rhizobia by lyophilization with trehalose. Pesqui. Agropecu. Bras. 37, 831–839 (2002).
Carini, P. et al. Relic DNA is abundant in soil and obscures estimates of soil microbial diversity. Nat. Microbiol. 2, 16242 (2017).
Nicholson, W. L., Munakata, N., Horneck, G., Melosh, H. J. & Setlow, P. Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiol. Mol. Biol. Rev. 64, 548–572 (2000).
Pham, V. H. T., Kim, J. & Jeong, S.-W. Psychrobacillus soli sp. nov., capable of degrading oil, isolated from oil-contaminated soil. Int. J. Syst. Evol. Microbiol. 65, 3046–3052 (2015).
Sáez-Nieto, J. A. et al. Paenibacillus spp. isolated from human and environmental samples in Spain: detection of 11 new species. New Microbes New Infect. 19, 19–27 (2017).
Lane, D. J. 16S/23S rRNA sequencing. In Nucleic acid techniques in bacterial systematics (eds Stackebrandt, E. & Goodfellow, M.) 115–175 (Wiley, New York, 1991).
Thiem, D., Gołębiewski, M., Hulisz, P., Piernik, A. & Hrynkiewicz, K. How does salinity shape bacterial and fungal microbiomes of Alnus glutinosa roots?. Front. Microbiol. 9, 1–15 (2018).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 13, 581–583 (2016).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian Classifier for Rapid Assignment of rRNA Sequences into the New Bacterial Taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, 590–596 (2013).
Huang, X. & Madan, A. CAP3: A DNA Sequence Assembly Program. Genome Res. 9, 868–877 (1999).
Acknowledgements
This study was financed through grant from National Science Centre, Poland number 2016/21/B/NZ9/00840 to MG. The funder had no role in study design, analyzing data and writing the manuscript. We would like to thank Ada Błaszczyk and Anita Kowalczyk for their help in maintaining the plants for the experiments.
Author information
Authors and Affiliations
Contributions
S.S.: performed experiments, analyzed data, drafted the manuscript, M.S.: performed experiments, participated in writing the manuscript, K.H.: conceptualized the study, participated in writing the manuscript, J.T.: participated in writing the manuscript, A.T.: participated in writing the manuscript, M.G.: supervised the project, conceptualized the study, analyzed data, participated in writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szymańska, S., Sikora, M., Hrynkiewicz, K. et al. Choosing source of microorganisms and processing technology for next generation beet bioinoculant. Sci Rep 11, 2829 (2021). https://doi.org/10.1038/s41598-021-82436-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82436-5
This article is cited by
-
“Next-Generation Bioformulations” for Plant Growth Promotion and Stress Mitigation: A Promising Approach for Sustainable Agriculture
Journal of Plant Growth Regulation (2023)
-
Prokaryotic communities adapted to microhabitats on the Indian lotus (Nelumbo nucifera) growing in the high-altitude urban Dal Lake
International Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.